Abstract

Infrared (IR) laser ablation-remote-electrospray ionization (LARESI) platform coupled to a tandem mass spectrometer (MS/MS) operated in selected reaction monitoring (SRM) or multiple reaction monitoring (MRM) modes was developed and employed for imaging of target metabolites in human kidney cancer tissue. SRM or MRM modes were employed to avoid artifacts that are present in full scan MS mode. Four tissue samples containing both cancerous and noncancerous regions, obtained from three patients with renal cell carcinoma (RCC), were imaged. Sixteen endogenous metabolites that were reported in the literature as varying in abundance between cancerous and noncancerous areas in various human tissues were selected for analysis. Target metabolites comprised ten amino acids, four nucleosides and nucleobases, lactate, and vitamin E. For comparison purposes, images of the same metabolites were obtained with ultraviolet (UV) desorption/ionization mass spectrometry imaging (UV-LDI-MSI) using monoisotopic silver-109 nanoparticle-enhanced target (109AgNPET) in full-scan MS mode. The acquired MS images revealed differences in abundances of selected metabolites between cancerous and noncancerous regions of the kidney tissue. Importantly, the two imaging methods offered similar results. This study demonstrates the applicability of the novel ambient LARESI SRM/MRM MSI method to both investigating and discovering cancer biomarkers in human tissue.
Introduction
Kidney cancer accounts for 2.2% of the total worldwide cancers and is the third most common cancer of the urinary tract after prostate and bladder cancer, whether measured by incidence or prevalence. In 2018, over 400 000 cases of kidney cancer were diagnosed and more than 170 000 deaths were due to this disease. Furthermore, its incidence is on the rise.1 There are three main histological subtypes of renal cancer, namely chromophobe RCC (cRCC), papillary RCC (pRCC), and clear cell RCC (ccRCC). Of these, ccRCC accounts for nearly 90% of all kidney tumors.2 Diagnosis of RCC is based on medical imaging (computed tomography, CT), magnetic resonance imaging (MRI), ultrasound). More than 50% of RCC’s are diagnosed incidentally, and approximately one-third of patients have metastatic tumors beyond the kidney at the time of diagnosis.3 Although the most effective treatment for localized RCC is radical nephrectomy with nephron-sparing surgery at an early stage, even after such optimal surgery, nearly one-third of patients experience disease recurrence after surgical resection.4 Various RCC biomarkers, most of which are proteins, (C-reactive protein (CRP), PTEN, carbonic anhydrase IX (CAIX), hypoxia-inducible factors (HIF-1α and HIF-1β), vascular endothelial growth factor (VEGF, CD44, E-cadherin, osteopontin, antigen Ki-67, and tumor protein p53) have been proposed, and their monitoring might promote timely prognosis of metastatic RCC. However, these biomarkers suffer from low sensitivity and specificity. Thus, further research to identify new RCC biomarkers is required for early detection, diagnosis, treatment guidance and assessment, monitoring of treatments, identifying relapses, as well as elucidation of molecular processes behind the disease states.5
The interest in mass spectrometry imaging (MSI) has grown steadily in recent years. MSI is used for visualization and analysis of spatial distributions of molecules, within a relatively wide molecular weight range, in complex biological systems with excellent molecular specificity.6 It is noteworthy that MSI has been employed almost exclusively in nontargeted mode, using full mass scan, MS1-spectrum mode.7 Matrix-assisted laser desorption/ionization (MALDI) is the chosen technique for molecular imaging of tissue samples. Indeed, MALDI MSI has been successfully employed to analyze the spatial distribution of proteins8 and lipids9 within kidney tissue of patients with RCC. However, in MALDI MSI, due to utilization of a near-UV laser for ablation, relatively large amounts of additional low-molecular-weight organic acids are added as an ablation matrix. This leads to the generation of numerous and abundant chemical background peaks in the low-mass region (under m/z 1000) and the ability to detect cellular metabolites is limited. Further, acidic matrices can cause acid-catalyzed hydrolysis of various biomolecules. MALDI also suffers from a highly variable sensitivity (the sweet spot effect) and low ionization efficiency for low-polarity compounds. To overcome the limitations of MALDI, a matrix-free, laser desorption–ionization method that uses a steel target covered with cationic silver or gold nanoparticles (AgNPs and AuNPs) has been developed.10,11 The latter represents a surface-assisted laser desorption ionization method (SALDI) that, unlike MALDI, is matrix-free. When Ag-109 nanoparticles are used, the method is referred to as 109AgNPET LDI.
MSI techniques that allow imaging of biological material under ambient ionization conditions have also been developed.12 To the best of our knowledge, desorption electrospray ionization (DESI) is the only ambient ionization technique that has been applied to the imaging of lipids and other low molecular compounds in kidney cancer tissue.13−15 With DESI, the main limitations include a relatively low spatial resolution, usually limited to 100 μm, a low ionization efficiency for some molecules, and a low desorption efficiency for molecules that are strongly bound to surfaces.16 Moreover, the DESI sampling depth is very shallow, which greatly reduces the amount of material that is available for analyses in the mass spectrometer. As in MALDI, this increases the risk that MS signals are obtained mainly from extracellular fluids.
Among ambient environment MS methods, the ones that use a mid-IR laser for sampling are the most suitable for the analysis of metabolites within biological tissues. A laser beam with a wavelength of about 2.94 μm effectively couple its energy into the O–H stretching mode in hydrogen-bonded water, present in any hydrated biological material.17 A main advantage of IR laser over UV laser, for ablation from tissue, is a greater depth of sampling, about 10 μm for a single laser pulse and arbitrarily deep for repeated pulses. Literature contains mentions regarding experimental setup examples that also use laser ablation and electrospray ionization, LAESI.18 This method was recently used to perform imaging of human hepatocarcinoma tissue samples.19 However, mentioned work differs considerably in technical design with the one presented herein and measurements were made in ToF-mode, without using ion fragmentation techniques.
In tandem mass spectrometry (MS/MS), the selectivity for a targeted analyte is enhanced by monitoring a compound-unique ion fragmentation in selected reaction monitoring (SRM), or multiple reaction monitoring (MRM), mode. As the chemical noise is effectively removed, this usually enhances sensitivity despite the lower ion detection efficiency. For these reasons, MS/MS is a powerful tool for high-sensitivity detection and near-certain identification of targeted low-molecular-weight compounds. The method is also characterized by a wide dynamic range, extremely high speed, and suitability for quantitation.20
The aim of this study is to report the development of new experimental MSI setup referred to as “laser ablation-remote-electrospray ionization” (LARESI) and to present unique MS imaging results of human tissue, that is, to demonstrate LARESI SRM targeted MS/MS imaging of frozen kidney tissue containing cancerous and noncancerous regions. Images depict 16 target metabolites. Importantly, each metabolite was identified based on its MS/MS fragmentation using SRM measurement mode. No additional sample preparation other than rinsing was required.. The tissue was sampled employing laser ablation and all metabolites were detected from the same ablation event in any one pixel. The ion plume-gathering interface used LARESI platform. The ESI-produced ions were analyzed on a QTRAP mass spectrometer, and this is the first report of results from this MS imaging instrument.
Images of kidney tissue with RCC were also obtained with the 109AgNPET LDI” method.21 In the latter, ablation occurs in vacuum and ions are detected in full-scan mode on a TOF-mass spectrometer. Images of RCC kidney tissue obtained with these two markedly different MSI methods are compared, and it is found that the they are comparable.
Experimental Section
1. Participants
The study protocol was approved by the Bioethics Committee at the University of Rzeszow (Poland, permission no. 2018/04/10), and the research was performed in accordance with relevant guidelines and regulations. Specimens and clinical data from patients involved in the study were collected with informed consent. All laboratory test results for the cancer patients (complete blood count, kidney function tests, CRP, urine analysis, bleeding profile) were within normal limits. Whole tumors and small fragments of adjacent healthy tissue were resected (cancer and control tissue, respectively). Control tissue was recognized as normal based on pathological analysis. Patients characteristics are provided in Section S3 of the Supporting Information (SI).
2. Materials and Equipment
Silver-109 (min. 99.75% of 109Ag) isotope was purchased from BuyIsotope (Sweden) and transformed to trifluoroacetate salt using known methods (involving dissolution in HNO3; precipitation of 109AgOH; reaction with trifluoroacetic acid; and recrystallization from tetrahydrofuran/hexane mixed solvent system). 2,5-Dihydroxybenzoic acid (DHB) was purchased from Aldrich. Steel targets were machined from H17 stainless steel. All solvents were of HPLC quality, except for water (18 MΩ water produced locally) and methanol (LCMS grade, Fluka). The silver-109 nanoparticles were synthesized on the surface of steel targets11 Optical photographs of tissue samples were obtained on an Olympus SZ10 microscope, equipped with an 8 Megapixel Olympus camera.
3. Tissue Processing
Following surgery, tissue samples were immediately frozen in dry ice to minimize sample degradation and stored at −60 °C until analysis. For the LARESI SRM MSI imaging experiments, 100-μm-thick tissue sections were cut using a microtome, and the slices were mounted on the Peltier stage set to −18 °C to minimize lateral mixing of compounds in the sample surface.
Four different kidney tissue sections (Specimen 1–4 as described in SI Section S3) recovered from three different patients were examined. Specimens 2 and 3 were obtained from the same patient (SI Section S3). Each of the four tissue sections contained both a cancerous (RCC) and a noncancerous (control) region.
4. Laser Ablation, Remote-Electrospray Ionization, Selected Reaction Monitoring, Mass Spectrometry Imaging: LARESI SRM MSI of Kidney Cancer Tissue
4.1. Laser System
An Nd/YAG-pumped, tunable OPO laser (IR Opolette 2731-HE; Opotek, Carlsbad, CA) with 4 ns, mid-IR laser pulses with a maximum repetition frequency of 20 Hz, was tuned to 2.94 μm. The pulse energy was 3.5 mJ (measured using a pyroelectric energy meter PE25-SH-V2; Ophir-Spiricon, Logan, UT).
4.2. Imaging Sample Chamber and Imaging Procedure
The experiment was performed in an airtight chamber as depicted in Figure 1. The chamber is pressurized with nitrogen gas to produce nitrogen flow of 2 L/min. The specimen is placed on a 50 × 50 mm sample stage. A Peltier cooling plate (TE-127–1.4–1.5; TE Technology, Traverse City, M) maintains the sample at of −18 °C. Excess heat generated from the Peltier element is removed using circulating water and an external radiator. The temperature-controlled sample stage is mounted on a motorized XY-stage (MTS50-Z8; Thorlabs, Newton, NJ). The pulsed beam from the OPO laser enters the sample chamber through a 1” Infrasil window (Thorlabs) and is redirected toward the sample stage by a gold-plated mirror (PF10–03-M01; Thorlabs). The beam is focused onto the sample surface by a 40 mm focal length CaF2 spherical lens (Thorlabs), mounted on a Z-axis stage (Thorlabs). The incidence angle on the sample is 90°; the size of the laser focus is 60 ± 10 μm and the pulse energy measured at the sample surface is 2.5 mJ. During imaging, the laser focal point remains fixed in space, while the sample is moved by the computer-controlled XY-stage. A funnel, connected to a 4 mm I.D. PTFE tube, is positioned over the laser ablation site. The overpressure in the chamber drives a 2 L × atm/min nitrogen gas flow through the tube. The laser ablation plumes are entrained into the gas and transported to the electrospray ionization (ESI) source of the SCIEX QTRAP 5500 mass spectrometer. The outlet end of the Teflon tube is mounted axially with the MS sampling cone inside the ESI source, and the distance between the butt end of the tube and the tip of the cone is 20 mm. A binary HPLC pump (Agilent G1312A) provides a steady flow of a binary solvent mixture (2:1 IPA:water with 0.5% acetic acid; 20 μL/min) to the electrospray needle.
Figure 1.

Schematic representation of the LARESI SRM MSI experimental setup.
Samples were maintained at −18 °C during analysis by the Peltier module. Imaging of tissue sections was performed over a square or rectangular area of about 1 cm2. The spatial resolution was in a range from 175 to 300 μm. Each of 1.5 × 103 pixels was exposed to the laser for 2 s, at a laser pulse repetition rate of 15 Hz. Between pixels, the sample stage moved with a speed of 2 mm/s. The time delay between pixels, during which the stage was moving along a straight line (the horizontal direction in Figures 3–5), was 4 s, and the time delay between lines was 5 s. Control and analysis software was described recently.22
Figure 3.

Photographs and LARESI SRM MS ion images of nucleobases, nucleosides, and lactate in RCC Specimen 2 (SI Section S3). Optical photographs of the kidney cancer tissue prior to (A) and after imaging (B). Corresponding LARESI SRM images (C–G). The imaged area is 8 × 8 mm obtained with 40 × 40 pixels and at 200 × 200 μm resolution.
Figure 5.

Photographs and LDI-MSI analysis with 109AgNPET of the surface of the RCC Specimen 4 (SI Section S3). (A) An optical photograph of the imaged surface of the kidney cancer sample. (B) An optical image of the 109AgNPET surface with RCC tissue imprint. Dashed lines represent cancer-free and cancer regions. (C–R) Ion images (TIC normalization) for ions of m/z as specified below each image. All images are within ±0.05 m/z and spatial resolution of 250 × 250 μm.
4.3. Mass Spectrometer Parameters
A SCIEX QTRAP 5500 mass spectrometer was used in positive ion mode with selected reaction monitoring (SRM) measurement mode with Q1/Q3/DP/EP/CE and CXP settings as stated in Table 1. In MRM mode, each compound-specific fragmentation was monitored for specific time (Table 1) with a 5 ms delay to the next fragmentation. The settings of the ESI source were as follows: source temperature 500 °C, curtain gas 20 psi, ion source gas 1–30 psi, ion source gas 2–20 psi, ion-spray voltage −4500 V (for negative mode) and +5500 V (for positive mode), collision gas (nitrogen): medium. Images of extracted ion chromatograms for compounds studied with LARESI are provided in SI Section S4.
Table 1. Mass Spectrometry Parameters of LARESI SRM MSI Experiments (Q1, Q3, Scan Time, DP, EP, CE, CXP) and Observed Abundance Ratios of Selected Metabolites between Cancer and Cancer-Free Human Kidney Tissue Regions both for LARESI SRM MSI (“ESI”) and 109AgNPET LDI MSI (“LDI”).
| abundance
ratio |
|||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| compound name | polaritya | Q1 [m/z] | Q3 [m/z] | ESIb | LDIc | scantime [ms] | DP [V] | EP [V] | CE [V] | CXP [V] | image |
| alanine | + | 90.1 | 44.0 | 0.6 | 0.7 | 30 | 6 | 4 | 17 | 6 | Figure 2C |
| serine | + | 106.1 | 60.0 | 0.5 | 0.7 | 30 | 6 | 10 | 16 | 7 | Figure 2D |
| threonine | + | 120.1 | 103.2 | 0.8 | 0.8 | 30 | 20 | 14 | 27 | 10 | Figure 2E |
| lysine | + | 147.1 | 84.0 | 0.7 | 0.8 | 30 | 15 | 14 | 23 | 10 | Figure 2F |
| glutamic acid | + | 148.1 | 84.0 | 0.4 | 0.6 | 30 | 21 | 14 | 21 | 10 | Figure 2G |
| methionine | + | 150.2 | 104.0 | 0.7 | 0.6 | 30 | 6 | 12 | 15 | 12 | Figure 2H |
| histidine | + | 156.1 | 110.0 | 0.6 | 0.7 | 30 | 16 | 13 | 19 | 12 | Figure 2I |
| phenylalanine | + | 166.1 | 103.0 | 0.6 | 0.8 | 30 | 11 | 14 | 37 | 12 | Figure 2J |
| arginine | + | 175.2 | 70.0 | 1.2 | 1.1 | 30 | 40 | 11 | 27 | 8 | Figure 2K |
| tryptophan | + | 205.1 | 146.0 | 1.1 | 0.8 | 30 | 20 | 14 | 16 | 10 | Figure 2L |
| guanine | – | 150.0 | 108.0 | 1.2 | 0.7 | 50 | –80 | –10 | –18 | –13 | Figure 3C |
| uridine | – | 243.0 | 110.2 | 0.4 | 0.9 | 50 | –80 | –10 | –18 | –13 | Figure 3D |
| thymine | – | 125.0 | 42.0 | 1.8 | 1.3 | 50 | –80 | –10 | –14 | –13 | Figure 3E |
| inosine | – | 267.0 | 135.2 | 2.9 | 1.7 | 30 | –80 | –10 | –23 | –13 | Figure 3F |
| lactate | – | 89.0 | 43.1 | 2.3 | 1.0 | 30 | –80 | –10 | –15 | –13 | Figure 3G |
| vitamin E | + | 431.4 | 165.1 | 2.0 | 1.1 | 30 | 120 | 9 | 40 | 15 | Figure 4C |
| 137.1 | 30 | 120 | 9 | 68 | 19 | Figure 3D | |||||
“–” = negative ion mode; “+” = positive ion mode.
Averaged cancer-to-normal signal intensity ratio in LARESI SRM MSI (calculated by dividing averaged signal intensity from cancer region by value from normal region),
Averaged cancer-to-normal signal intensity ratio in 109AgNPET LDI MS imaging; DP = declustering potential; EP = entrance potential; CE = collision energy; CXP = cell exit potential.
4.4. LARESI Method Test in Enhanced Product Ion (EPI) MS/MS Mode
Volume of 10 μL of serine and aqueous lysine solution (0.1 mg/mL) was poured onto filter paper (2 × 2 cm) of 0.2 mm thickness. After drying, paper was attached to stainless steel plate (2 × 3 cm; 0.8 mm thickness) with sticky tape and inserted into LARESI chamber. Mass spectrometer was working in EPI mode set on protonated lysine (m/z 147) and deprotonated serine (m/z 104). Line resolution was 300 μm. Other parameters were as stated in Sections 4.2 and 4.3.
5. Monoisotopic Silver-109 Nanoparticles-Enhanced Target Laser Desorption/Ionization Mass Spectrometry Imaging: 109AgNPET LDI MSI of the Kidney Cancer Tissue
Silver-109 nanoparticle-enhanced target (109AgNPET) preparation and imaging of human kidney tissue with the use of nanoparticle-based methods were described in our recent works.10,11,23 For more details see SI Section S1. Studied specimen contained both normal and cancer regions and was 23 × 26 × 20 mm in size, and the MS imaging was made for imprint of tissue with 250 × 250 μm resolution.
Results and Discussion
LARESI Method Test in Enhanced Product Ion (EPI) MS/MS Mode
LARESI experimental setup was tested in order to provide data on variability of pixel-to-pixel MS/MS signal intensity. The ideal test object should have homogeneous distribution of a studied compound, but this alone does not provide perfect results as instrumental components of the setup, generate signal variability. As LARESI is capable of analysis of various biological and nonbiological objects due to application of relatively powerful pulsed laser, low-water content object—0.2 mm thick filter paper—was used as a matrix for test compounds. Data shown inSI Section S5 figure provide pixel intensity variation visualization for two test compounds,lysine and serine and their fragments in positive and negative MS/MS modes, respectively. Variabilities calculated as standard deviation of pixel maximum intensities were found to be 12% and 16% for protonated lysine and lysine fragment and 14% and 15% for deprotonated serine and serine fragment, respectively. The observed variabilities are very low when compared to the ones reported for MALDI MSI.24
LARESI SRM MS Imaging of Kidney Cancer Tissue
MS images for 16 selected metabolites in cancerous kidney tissue samples were obtained with two different MSI methods. IR-laser-ablation-based LARESI targeted MSI experiments utilizes tandem mass spectrometry (MS/MS) in fragmentation-based SRM or MRM modes, whereas 109AgNPET LDI11 employs a ToF/MS operating in the standard full scan MS1 mode. One of the study aims was to compare these two methods. It is recognized that both amino acid and nucleic acid metabolism are often upregulated in cancerous, relative to noncancerous tissue.25 Therefore, compounds representing the above pathways, namely 10 amino acids and 4 nucleosides or nucleobases were selected for imaging. Lactate, which is a key cancer metabolite,26 and vitamin E, were also included.
In this first reported application of the LARESI SRM MSI method, images for 16 selected metabolites (Table 1) in frozen human kidney tissue sections are presented. Three different sections were used, and they were obtained from Specimens 1, 2, and 3, respectively. Each section contains both cancerous and noncancerous regions. The images obtained for the Specimen 1, 2, and 3 sections are shown in Figures 2, 3, and 4, respectively. For each metabolite, the m/z values of the precursor/parent ion (Q1) and the product/daughter ion (Q3) are listed in Table 1, together with their optimized values for declustering potential (DP), entrance potential (EP), collision energy (CE), and cell exit potential (CXP).
Figure 2.
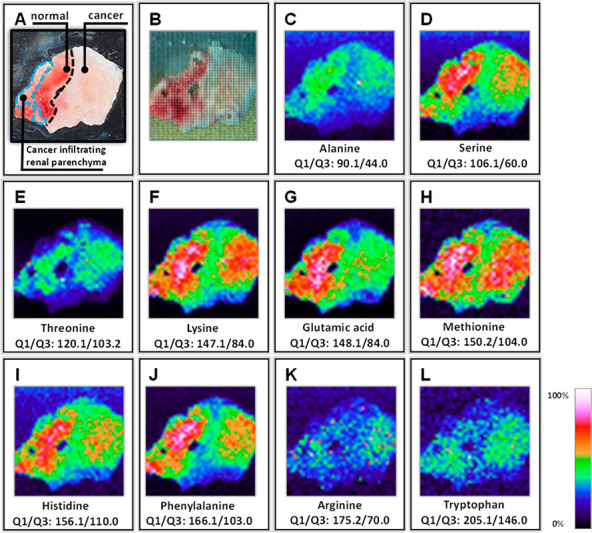
Photographs and LARESI SRM MSI ion images of selected amino acids in kidney tissue section from Specimen 1 (SI Section S3). Optical photographs of the imaged kidney tissue (A) prior to and (B) following imaging. LARESI SRM MSI ion images of the kidney tissue (C–L). The imaged area is 12 × 12 mm obtained with 40 × 40 pixels and at 300 × 300 μm resolution.
Figure 4.

Photographs and LARESI SRM ion images of kidney tissue representing specimen 3 (SI Section S3). Optical photographs of the sample (A) prior to and (B) after imaging; Images of vitamin E distributions (C, D) from two different SRM fragmentation reactions. The imaged area is 7 × 9 mm using 40 × 40 pixels with 175 × 225 μm resolution.
Images of the 10 selected amino acids were obtained from the Specimen 1 tissue section. A photograph of the tissue prior to imaging, with outlined cancerous and noncancerous regions is presented in Figure 2A. SRM ion images, (Figure 2C–L), reveal that several of the amino acids are detected with a lower abundance in the RCC than in the control. The cancer-to-control SRM signal abundances, averaged over the respective regions are listed in Table 1. In noncancerous tissue eight of the 10 amino acids (alanine, serine, threonine, lysine, glutamic acid, methionine, histidine, and phenylalanine) had a higher abundance than in cancerous tissue. Two of 10 amino acids (arginine, tryptophan) were detected with a slightly higher abundance in the RCC area. The largest differences between noncancerous and cancerous tissue was observed for serine, glutamic acid, histidine and phenylalanine.
The results obtained using LARESI imaging platform are consistent with the ones reported in several previous renal cancer studies. It has been demonstrated that serum levels of alanine, serine, lysine, glutamic acid, methionine, histidine, and phenylalanine were significantly decreased in patients with RCC while arginine levels were increased.27 A similar pattern of amino acid abundances within the kidney tissue is revealed with LARESI system (Figure 2). It should be noted that the phenomenon of amino acid up- and downregulation is not unique for renal cancers. Significant changes in the abundances of amino acids in plasma and tissue have indeed been observed for patients with many different types of cancers (lung, breast, gastric, colorectal, prostate, pancreatic, and colon).28−31 Owing its simplicity in sample preparation, LARESI imaging platform is, therefore, ideally suited for rapid investigation of other types of malignant tumors.
Nucleosides and their metabolites play an important role in the metabolism and growth of tumors, as rapidly proliferating tumor cells have a large need for nucleotides.32 Hsu et al. reported the increased levels of nucleosides, in particular of inosine, in urine from breast cancer patients compared to healthy persons,33 while the suppression of uridine in urine of patients with colorectal cancer has been observed.34 The authors proposed that nucleoside might serve as potential human tumor markers.
To demonstrate whether similar trends can be noted in tissues of kidney cancer patients, the spatial distributions of selected nucleosides (uridine, inosine), nucleobases (guanine, thymine), and organic metabolite, namely lactate, were studied in Specimen 2 (Figure 3). The SRM images for guanine, thymine, inosine, and lactate all showed higher abundances in the cancer region. In contrast, uridine abundances were lower in the cancer tissue. A decline of uridine concentrations in kidney cancer tissue is consistent with an earlier report.35 Apart from the differences between the cancer and cancer-free tissue regions, it is noteworthy that the images also demonstrate significant SRM abundance variations within each of the two regions. The MSI of guanine, uridine, thymine, and inosine with LARESI platform is the first report on the distribution of these metabolites within human kidney tissue.
The SRM image of lactate (Figure 3G) shows that the RCC cancer tissue contains approximately 2-fold higher level of lactate than does noncancerous tissue. Lactate is recognized as a key metabolite related to cancer progression and metastasis. It has long been known that cancer cells exhibit different metabolism than cancer-free cells. In 1920s Warburg found that cancer cells prefer aerobic glycolysis rather than oxidative respiration. In this process, tumor cells produce large amounts of lactate from glucose even in the presence of sufficient levels of cellular oxygen.36 In cervical cancer, accumulation of lactate within tumors was inversely correlated to patient survival.37 Similar correlations were observed in patients with neck and head squamous cell carcinoma, as the cancers in patients with short-term survivals had a significantly higher content of lactate than those in patients with long-term survival.38 Increased level of lactate were found also in RCC tissue,39,40 particularly for higher-grades of cancer.41 Hence, here is good evidence that tumor lactate metabolism and lactate levels are directly correlated with the aggressiveness of cancer. Conceivably, detection of high lactate levels would be a supportive tool for tumor diagnosis, prognosis, and for predicting probability of drug effectiveness.
Ion images in Figure 4 were obtained using LARESI SRM MSI of tissue from Specimen 3. Elevated levels of vitamin E (α-tocopherol) are observed in the cancer zone. The uptake of lipids and fatty acids is increased in cancer cells and the high concentration of vitamin E that is an antioxidant, may be explained by the need of cells to resist oxidative stress.42 Increased level of vitamin E in RCC tissue compared with a noncancer tissue has previously been reported.40,43
109AgNPET LDI MS Imaging of Kidney Cancer Tissue
All MSI methods are associated with matrix effects that influence the ratio of measured abundance to actual concentration of compounds. These effects are not trivial to quantify. The reliability of an abundance measurement in imaging as a predictor of relative concentrations will require substantial future work. In this study, a first step is taken by comparing MSI results from two very different MSI methods: LARESI SRM MSI, which uses an IR laser, and 109AgNPET LDI MSI, which employs a UV laser for ablation and/or desorption. The 109AgNPET LDI MSI experiment was performed by recording high-resolution TOF/MS spectra at 32 × 44 pixel-locations over an 8 × 11 mm size area, that is, with 250 × 250 μm resolution. Specimen 4 was used to make a tissue imprint on the 109Ag nanoparticle-covered target plate. The ion images are shown in Figure 5. The averaged ion intensities for both cancerous and noncancerous regions and normal regions presented in the sixth column of Table 1. The identity of some of compounds was confirmed with LIFT MS/MS experiments (SI Section S2).
Ion images of adduct-type ions of 10 amino acids presented in Figure 5 were obtained for the same selected metabolites as the ones investigated with LARESI SRM MSI system. The abundances of alanine, serine, threonine, lysine, methionine, histidine, phenylalanine, and glutamic acid ions are at slightly higher levels in the noncancer tissue region versus tumor tissue. Ions assigned to potassium adduct of arginine were present in higher abundance in cancer tissue compared to a noncancer tissue. A different result from that of LARESI SRM MS imaging, was obtained for sodiated tryptophan ion that was found to be in a higher abundance in noncancer tissue. The greatest difference in abundance within the examined tissue was observed for proton adduct of thymidine (Figure 5F) and silver-109 isotope adduct of inosine (Figure 5Q), which show higher intensity in the area of cancer tissue versus cancer-free region. Large variations can also be seen for ion putatively assigned to proton adduct of vitamin E. The latter were found dominating cancer-free tissue region compared to cancer one (Figure 5R). A number of recent scientific reports have suggested that potential biomarkers should have cancer to control fold change of at least 1.2 or under 0.8.44−46 Using this criteria for both tested MSI methods, the downregulated compounds, namely alanine, serine, glutamic acid, methionine, histidine, and upregulated ones such as thymine and inosine could be considered as kidney cancer biomarkers.
Methods Comparison
Imaging of tissue with the 109AgNPET LDI MSI method is an excellent alternative to the MALDI MSI method. The absence of an organic matrix and the option of performing internal mass calibration are two of the many advantages of this method. Important advantage of AgNPET-based family of methods is ease of use and also higher imaging speed (ca. 2–3-times), the latter providing images of much higher resolution in the same experiment time. Additional advantage is a smaller UV laser focal point diameter, compared to that of 2.9 μm IR laser. However, a drawback of the method is that intercellular fluid is preferentially transferred to the plate during imprinting, which may cause the loss of information from intracellular metabolism. In MALDI or similar LDI methods, for example, AgNPET, significant variations in sodium and potassium ion concentrations and in pH over the imaged tissue region may lead to quantitative problems. In this case, it is likely that analyte intensity may change more due to sodium/potassium local concentration than of metabolite itself. Moreover, in most MALDI instruments, the sample stage is at room temperature, which promotes tissue degradation and allows warping and cracking of high vacuum-dried tissue samples. These problems are virtually nonexistent when employing the LARESI SRM MSI method. It is an atmospheric pressure method that uses ablation of frozen sample material from a surface to the gas phase with a 2.9 μm IR laser, followed ionization by electrically charged solvent clusters generated through electrospray. In order to maximize the collection and ionization efficiencies in LARESI SRM MSI, a funnel-type device coupled with optics was positioned above the sample to confine the ablation plume and to guide the material through the PTFE tube and into the ESI source. The method has several advantages. LARESI SRM MSI enables direct analysis of samples of varying sizes, shape and physical form. There is no need for sample pretreatment steps such as application of a matrix or solvent, dehydration, or derivatization prior to analysis. Thus, it allows preserving the anatomical integrity of the sample, reducing the risk of delocalization of analytes and chemical contamination of the samples. The samples are not exposed to vacuum and this reduces the possibility of deformation. The latter would produce mass shifts of detected ions, a significant problem with vacuum, TOF-analyzer-based MS instruments. The cooling stage holds the specimen in a frozen state, which prevents water loss thus preserving sample integrity. In LARESI SRM MSI analytes are ionized by protonation, which greatly facilitates qualitative and quantitative analysis and makes the spectral results readily compatible with databases commonly used for metabolomics analyses. Finally, the ability of using SRM in MS/MS mode boosts both analytical sensitivity and selectivity. It is anticipated that here described LARESI setup will facilitate not only MS3 or higher order fragmentation MSI, but also direct ambient 3D imaging of tissues.
Conclusions
A novel MSI platform termed LARESI SRM has been developed that allows direct, pretreatment-free, ambient chemical imaging of complex biological samples, such as heterogeneous human tissues. The potential of the LARESI SRM MSI method for rapid and accurate detection of metabolites that could serve as tumor markers in human tissues was demonstrated in kidney samples from cancer patients. For comparison purposes, a markedly different MSI method, namely 109AgNPET LDI in full scan mode, was employed for untargeted analysis of the same tissue. The study is the first one to offer the comparison of molecular images from two different methods that depict spatial distribution of metabolites that included 10 amino acids, two nucleosides, and four nucleobases, as well as lactate and vitamin E, relevant to cancer diagnostics within human tissue from patients with RCC. Ion images of cancerous kidney tissue obtained for 16 selected metabolites revealed striking abundance differences and patterns versus cancer-free tissue regions. In almost every case, the abundance patterns observed with the two MSI methods were similar, thus, mutually supportive and clearly differed between cancerous and noncancerous tissues.
Employing LARESI SRM/MRM MSI platform, biomarkers discussed in this work, together with other low molecular weight compounds described in literature can aid identification of cancerous tissues. Conceivably, LARESI method can become an integral part of cancer diagnostic methods that are based on analyses of biopsy samples.
Acknowledgments
The study was supported by National Science Centre (Poland) research project OPUS Number 2016/23/B/ST4/00062.
Supporting Information Available
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.analchem.9b04580.
S1 Monoisotopic silver-109 nanoparticles-enhanced target laser desorption/ionization mass spectrometry imaging: 109AgNPET LDI MSI of the kidney cancer tissue. S2 LIFT MSMS analysis results. S3 Clinical characteristics of study group, and controls. S4 Extracted chromatograms (EICs) for LARESI experiments. S5 Test data of LARESI MS/MS pixel-to-pixel intensity variability (PDF)
The authors declare no competing financial interest.
Supplementary Material
References
- Bray F.; Ferlay J.; Soerjomataram I.; Siegel R. L.; Torre L. A.; Jemal A. Global Cancer Statistics 2018: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. Ca-Cancer J. Clin. 2018, 68 (6), 394–424. 10.3322/caac.21492. [DOI] [PubMed] [Google Scholar]
- Linehan W. M.; Walther M. M.; Zbar B. The Genetic Basis of Cancer of the Kidney. J. Urol. 2003, 170 (6), 2163–2172. 10.1097/01.ju.0000096060.92397.ed. [DOI] [PubMed] [Google Scholar]
- Janzen N.; Kim H.; Figlin R.; Belldegrun A. Surveillance after Radical or Partial Nephrectomy for Localized Renal Cell Carcinoma and Management of Recurrent Disease. Urol. Clin. North Am. 2003, 30 (4), 843–852. 10.1016/S0094-0143(03)00056-9. [DOI] [PubMed] [Google Scholar]
- Kuijpers Y. A. M.; Meijer R. P.; Jonges G. N.; de Jong J.; Bosch J. L. H. R.; Horenblas S.; Bex A. Potentially Curable Recurrent Disease after Surgically Managed Non-Metastatic Renal Cell Carcinoma in Low-, Intermediate- and High-Risk Patients. World J. Urol. 2016, 34 (8), 1073–1079. 10.1007/s00345-016-1822-6. [DOI] [PubMed] [Google Scholar]
- Sanganeria B. S.; Misra R.; Shukla K. K.. Molecular Diagnostics in Renal Cancer. In Molecular Diagnostics in Cancer Patients; Springer Singapore: Singapore, 2019; pp 199–218. 10.1007/978-981-13-5877-7_13. [DOI] [Google Scholar]
- Richard M.; Caprioli; Farmer T. B.; Gile J.. Molecular Imaging of Biological Samples: Localization of Peptides and Proteins Using MALDI-TOF MS. Anal. Chem. 1997. 694751. 10.1021/ac970888i. [DOI] [PubMed] [Google Scholar]
- Lippincott-Schwartz J.; Snapp E.; Kenworthy A. Studying Protein Dynamics in Living Cells. Nat. Rev. Mol. Cell Biol. 2001, 2 (6), 444–456. 10.1038/35073068. [DOI] [PubMed] [Google Scholar]
- Seeley E. H.; Oppenheimer S. R.; Mi D.; Chaurand P.; Caprioli R. M. Enhancement of Protein Sensitivity for MALDI Imaging Mass Spectrometry After Chemical Treatment of Tissue Sections. J. Am. Soc. Mass Spectrom. 2008, 19 (8), 1069–1077. 10.1016/j.jasms.2008.03.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hájek R.; Lísa M.; Khalikova M.; Jirásko R.; Cífková E.; Študent V.; Vrána D.; Opálka L.; Vávrová K.; Matzenauer M.; et al. HILIC/ESI-MS Determination of Gangliosides and Other Polar Lipid Classes in Renal Cell Carcinoma and Surrounding Normal Tissues. Anal. Bioanal. Chem. 2018, 410 (25), 6585–6594. 10.1007/s00216-018-1263-8. [DOI] [PubMed] [Google Scholar]
- Nizioł J.; Ossoliński K.; Ossoliński T.; Ossolińska A.; Bonifay V.; Sekuła J.; Dobrowolski Z.; Sunner J.; Beech I.; Ruman T.. Surface-Transfer Mass Spectrometry Imaging of Renal Tissue on Gold Nanoparticle Enhanced Target. Anal. Chem. 2016, 88 ( (14), ). 7365. 10.1021/acs.analchem.6b01859. [DOI] [PubMed] [Google Scholar]
- Nizioł J.; Rode W.; Laskowska B.; Ruman T.. Novel Monoisotopic 109AgNPET for Laser Desorption/Ionization Mass Spectrometry. Anal. Chem. 2013, 85 ( (3), ). 1926. 10.1021/ac303770y. [DOI] [PubMed] [Google Scholar]
- Perez C. J.; Bagga A. K.; Prova S. S.; Yousefi Taemeh M.; Ifa D. R. Review and Perspectives on the Applications of Mass Spectrometry Imaging under Ambient Conditions. Rapid Commun. Mass Spectrom. 2019, 33 (S3), 27–53. 10.1002/rcm.8145. [DOI] [PubMed] [Google Scholar]
- Alfaro C. M.; Jarmusch A. K.; Pirro V.; Kerian K. S.; Masterson T. A.; Cheng L.; Cooks R. G. Ambient Ionization Mass Spectrometric Analysis of Human Surgical Specimens to Distinguish Renal Cell Carcinoma from Healthy Renal Tissue. Anal. Bioanal. Chem. 2016, 408 (20), 5407–5414. 10.1007/s00216-016-9627-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dill A. L.; Eberlin L. S.; Zheng C.; Costa A. B.; Ifa D. R.; Cheng L.; Masterson T. A.; Koch M. O.; Vitek O.; Cooks R. G. Multivariate Statistical Differentiation of Renal Cell Carcinomas Based on Lipidomic Analysis by Ambient Ionization Imaging Mass Spectrometry. Anal. Bioanal. Chem. 2010, 398 (7–8), 2969–2978. 10.1007/s00216-010-4259-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tamura K.; Horikawa M.; Sato S.; Miyake H.; Setou M.. Discovery of Lipid Biomarkers Correlated with Disease Progression in Clear Cell Renal Cell Carcinoma Using Desorption Electrospray Ionization Imaging Mass Spectrometry. Oncotarget 2019; Vol. 10. 10.18632/oncotarget.26706 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takáts Z.; Wiseman J. M.; Gologan B.; Cooks R. G. Mass Spectrometry Sampling under Ambient Conditions with Desorption Electrospray Ionization. Science 2004, 306 (5695), 471–473. 10.1126/science.1104404. [DOI] [PubMed] [Google Scholar]
- Dreisewerd K.; Draude F.; Kruppe S.; Rohlfing A.; Berkenkamp S.; Pohlentz G.; Molecular Analysis of Native Tissue and Whole Oils by Infrared Laser Mass Spectrometry. Anal. Chem. 2007. 10.1021/AC070191P. [DOI] [PubMed] [Google Scholar]
- Nemes P. N.; Vertes A.. Laser ablation electrospray ionization for atmospheric pressure, in vivo, and imaging mass spectrometry. Anal. Chem. 2007. 798098. 10.1021/ac071181r. [DOI] [PubMed] [Google Scholar]
- Zhou W.; Hong Y.; Huang C.; Shen C.; Chu Y.. Laser ablation electrospray ionization time-of-flight mass spectrometry for direct analysis of biological tissue. J. Anal. Methods Chem. 2019, 2019. 1. 10.1155/2019/1417035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kovarik P.; Grivet C.; Bourgogne E.; Hopfgartner G. Method Development Aspects for the Quantitation of Pharmaceutical Compounds in Human Plasma with a Matrix-Assisted Laser Desorption/Ionization Source in the Multiple Reaction Monitoring Mode. Rapid Commun. Mass Spectrom. 2007, 21 (6), 911–919. 10.1002/rcm.2912. [DOI] [PubMed] [Google Scholar]
- Nizioł J.; Misiorek M.; Ruman T. Mass Spectrometry Imaging of Low Molecular Weight Metabolites in Strawberry Fruit (Fragaria x Ananassa Duch.) Cv. Primoris with 109Ag Nanoparticle Enhanced Target. Phytochemistry 2019, 159, 11–19. 10.1016/j.phytochem.2018.11.014. [DOI] [PubMed] [Google Scholar]
- Brauer J. I.; Beech I. B.; Sunner J. Mass Spectrometric Imaging Using Laser Ablation and Solvent Capture by Aspiration (LASCA). J. Am. Soc. Mass Spectrom. 2015, 26 (9), 1538–1547. 10.1007/s13361-015-1176-0. [DOI] [PubMed] [Google Scholar]
- Nizioł J.; Misiorek M.; Ruman T. Mass Spectrometry Imaging of Low Molecular Weight Metabolites in Strawberry Fruit (Fragaria x Ananassa Duch.) Cv. Primoris with 109Ag Nanoparticle Enhanced Target. Phytochemistry 2019, 159, 11–19. 10.1016/j.phytochem.2018.11.014. [DOI] [PubMed] [Google Scholar]
- Porta T.; Lesur A.; Varesio E.; Hopfgartner G. Quantification in MALDI-MS Imaging: What Can We Learn from MALDI-Selected Reaction Monitoring and What Can We Expect for Imaging?. Anal. Bioanal. Chem. 2015, 407 (8), 2177–2187. 10.1007/s00216-014-8315-5. [DOI] [PubMed] [Google Scholar]
- Phan L. M.; Yeung S. C. J.; Lee M. H.. Cancer metabolic reprogramming: Importance, main features, and potentials for precise targeted anti-cancer therapies. Cancer Biol. Med. 2014, 1–19. 10.7497/j.issn.2095-3941.2014.01.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hirschhaeuser F.; Sattler U. G. A.; Mueller-Klieser W. Lactate: A Metabolic Key Player in Cancer. Cancer Res. 2011, 71 (22), 6921–6925. 10.1158/0008-5472.CAN-11-1457. [DOI] [PubMed] [Google Scholar]
- Mustafa A.; Gupta S.; Hudes G. R.; Egleston B. L.; Uzzo R. G.; Kruger W. D. Serum Amino Acid Levels as a Biomarker for Renal Cell Carcinoma. J. Urol. 2011, 186 (4), 1206–1212. 10.1016/j.juro.2011.05.085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leichtle A. B.; Nuoffer J. M.; Ceglarek U.; Kase J.; Conrad T.; Witzigmann H.; Thiery J.; Fiedler G. M. Serum Amino Acid Profiles and Their Alterations in Colorectal Cancer. Metabolomics 2012, 8 (4), 643–653. 10.1007/s11306-011-0357-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miyagi Y.; Higashiyama M.; Gochi A.; Akaike M.; Ishikawa T.; Miura T.; Saruki N.; Bando E.; Kimura H.; Imamura F.; et al. Plasma free amino acid profiling of five types of cancer patients and its application for early detection. PLoS One 2011, 6 ( (9), ). e24143. 10.1371/journal.pone.0024143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dereziński P.; Klupczynska A.; Sawicki W.; Pałka J. A.; Kokot Z. J. Amino Acid Profiles of Serum and Urine in Search for Prostate Cancer Biomarkers: A Pilot Study. Int. J. Med. Sci. 2017, 14 (1), 1–12. 10.7150/ijms.15783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simińska E.; Koba M. Amino Acid Profiling as a Method of Discovering Biomarkers for Early Diagnosis of Cancer. Amino Acids 2016, 48 (6), 1339–1345. 10.1007/s00726-016-2215-2. [DOI] [PubMed] [Google Scholar]
- Romero-Garcia S.; Lopez-Gonzalez J. S.; Báez-Viveros J. L.; Aguilar-Cazares D.; Prado-Garcia H. Tumor Cell Metabolism: An Integral View. Cancer Biol. Ther. 2011, 12 (11), 939–948. 10.4161/cbt.12.11.18140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsu W. Y.; Lin W. De; Tsai Y.; Lin C. T.; Wang H. C.; Jeng L. Bin; Lee C. C.; Lin Y. C.; Lai C. C.; Tsai F. J. Analysis of Urinary Nucleosides as Potential Tumor Markers in Human Breast Cancer by High Performance Liquid Chromatography/Electrospray Ionization Tandem Mass Spectrometry. Clin. Chim. Acta 2011, 412 (19–20), 1861–1866. 10.1016/j.cca.2011.06.027. [DOI] [PubMed] [Google Scholar]
- Zheng Y. F.; Yang J.; Zhao X. J.; Feng B.; Kong H. W.; Chen Y. J.; Lv S.; Zheng M. H.; Xu G. W. Urinary Nucleosides as Biological Markers for Patients with Colorectal Cancer. World J. Gastroenterol. 2005, 11 (25), 3871–3876. 10.3748/wjg.v11.i25.3871. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koshida K.; Harmenberg J.; Borgström E.; Wahren B.; Andersson L. Pseudouridine and Uridine in Normal Kidney and Kidney Cancer Tissues. Urol. Res. 1985, 13 (5), 219–221. 10.1007/BF00261579. [DOI] [PubMed] [Google Scholar]
- Warburg O.; Wind F.; Negelein E. The Metabolism of Tumors in the Body. English Reprint. J. Gen. Physiol. 1927, 8 (6), 519–530. 10.1085/jgp.8.6.519. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walenta S.; Wetterling M.; Lehrke M.; Schwickert G.; Sundfør K.; Rofstad E. K.; Mueller-Klieser W.. High Lactate Levels Predict Likelihood of Metastases, Tumor Recurrence, and Restricted Patient Survival in Human Cervical Cancers. Cancer Res. 2000, 60 ( (4), ). [PubMed] [Google Scholar]
- Ziebart T.; Walenta S.; Kunkel M.; Reichert T. E.; Wagner W.; Mueller-Klieser W. Metabolic and Proteomic Differentials in Head and Neck Squamous Cell Carcinomas and Normal Gingival Tissue. J. Cancer Res. Clin. Oncol. 2011, 137 (2), 193–199. 10.1007/s00432-010-0875-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao H.; Dong B.; Jia J.; Zhu H.; Diao C.; Yan Z.; Huang Y.; Li X. Application of Ex Vivo 1H NMR Metabonomics to the Characterization and Possible Detection of Renal Cell Carcinoma Metastases. J. Cancer Res. Clin. Oncol. 2012, 138 (5), 753–761. 10.1007/s00432-011-1134-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hakimi A. A.; Reznik E.; Lee C.-H.; Creighton C. J.; Brannon A. R.; Luna A.; Aksoy B. A.; Liu E. M.; Shen R.; Lee W.; et al. An Integrated Metabolic Atlas of Clear Cell Renal Cell Carcinoma. Cancer Cell 2016, 29 (1), 104–116. 10.1016/j.ccell.2015.12.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wettersten H. I.; Hakimi A. A.; Morin D.; Bianchi C.; Johnstone M. E.; Donohoe D. R.; Trott J. F.; Aboud O. A.; Stirdivant S.; Neri B.; et al. Grade-Dependent Metabolic Reprogramming in Kidney Cancer Revealed by Combined Proteomics and Metabolomics Analysis. Cancer Res. 2015, 75 (12), 2541–2552. 10.1158/0008-5472.CAN-14-1703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rodrigues D.; Monteiro M.; Jerónimo C.; Henrique R.; Belo L.; Bastos M. de L.; Guedes de Pinho P.; Carvalho M.. Renal Cell Carcinoma: A Critical Analysis of Metabolomic Biomarkers Emerging from Current Model Systems. Translational Research; ,Mosby Inc., February 1, 2017; pp 1–11. 10.1016/j.trsl.2016.07.018. [DOI] [PubMed] [Google Scholar]
- Catchpole G.; Platzer A.; Weikert C.; Kempkensteffen C.; Johannsen M.; Krause H.; Jung K.; Miller K.; Willmitzer L.; Selbig J.; et al. Metabolic Profiling Reveals Key Metabolic Features of Renal Cell Carcinoma. J. Cell. Mol. Med. 2011, 15 (1), 109–118. 10.1111/j.1582-4934.2009.00939.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsai C. K.; Yeh T. Sen; Wu R. C.; Lai Y. C.; Chiang M. H.; Lu K. Y.; Hung C. Y.; Ho H. Y.; Cheng M. L.; Lin G. Metabolomic Alterations and Chromosomal Instability Status in Gastric Cancer. World J. Gastroenterol. 2018, 24 (33), 3760–3769. 10.3748/wjg.v24.i33.3760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kalantari S.; Nafar M.; Samavat S.; Parvin M. 1 H NMR-Based Metabolomics Study for Identifying Urinary Biomarkers and Perturbed Metabolic Pathways Associated with Severity of IgA Nephropathy: A Pilot Study. Magn. Reson. Chem. 2017, 55 (8), 693–699. 10.1002/mrc.4573. [DOI] [PubMed] [Google Scholar]
- Xiao H.; Langerman A.; Zhang Y.; Khalid O.; Hu S.; Cao C. X.; Lingen M. W.; Wong D. T. W. Quantitative Proteomic Analysis of Microdissected Oral Epithelium for Cancer Biomarker Discovery. Oral Oncol. 2015, 51 (11), 1011–1019. 10.1016/j.oraloncology.2015.08.008. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


