Significance
Phenoloxidase-catalyzed melanization and Spätzle-triggered Toll signaling are critical innate immune responses of insects that are triggered by specific proteolysis. It has been unclear whether separate protease cascades or an integrated network of serine proteases coordinates these responses. Here we present evidence that hemolymph protease HP5 acts in pathways eliciting activation of both phenoloxidase and Spätzle-1 in Manduca sexta. HP5 has a unique specificity by cleaving proHP6 at His112, resulting in HP6 activation. CLIP proteases related to HP6 from other insect species, including Drosophila Persephone, share this activation site feature with proHP6, and therefore, HP5 orthologs in other species are candidates for key roles in regulating and integrating protease cascade pathways in innate immune responses.
Keywords: insect immunity, serine protease cascade, clip domain, zymogen activation, hemolymph protein
Abstract
Proteolytic activation of phenoloxidase (PO) and the cytokine Spätzle during immune responses of insects is mediated by a network of hemolymph serine proteases (HPs) and noncatalytic serine protease homologs (SPHs) and inhibited by serpins. However, integration and conservation of the system and its control mechanisms are not fully understood. Here we present biochemical evidence that PO-catalyzed melanin formation, Spätzle-triggered Toll activation, and induced synthesis of antimicrobial peptides are stimulated via hemolymph (serine) protease 5 (HP5) in Manduca sexta. Previous studies have demonstrated a protease cascade pathway in which HP14 activates proHP21; HP21 activates proPAP2 and proPAP3, which then activate proPO in the presence of a complex of SPH1 and SPH2. We found that both HP21 and PAP3 activate proHP5 by cleavage at ESDR176*IIGG. HP5 then cleaves proHP6 at a unique site of LDLH112*ILGG. HP6, an ortholog of Drosophila Persephone, activates both proHP8 and proPAP1. HP8 activates proSpätzle-1, whereas PAP1 cleaves and activates proPO. HP5 is inhibited by Manduca sexta serpin-4, serpin-1A, and serpin-1J to regulate its activity. In summary, we have elucidated the physiological roles of HP5, a CLIPB with unique cleavage specificity (cutting after His) that coordinates immune responses in the caterpillar.
Serine protease cascades have evolved in vertebrates and invertebrates to mediate defense responses to pathogen attack and tissue damage (1, 2). Once microbial surfaces or aberrant tissues are sensed by host factors in the circulation, specific proteolysis occurs to activate sequentially the protease cascade members within minutes. In insects, the cascades include serine proteases with multiple modular regulatory domains (3) and proteases with amino-terminal clip domains (2). Terminal clip proteases in the pathways generate active phenoloxidases (POs) and the Toll ligand Spätzle by cleaving their inactive precursors. POs catalyze the formation of reactive compounds and melanin to kill and sequester pathogens (4). Spätzle binds the Toll receptor to induce synthesis of antimicrobial peptides (5). Noncatalytic serine protease homologs (SPHs) are also activated by limited proteolysis and become a cofactor for proPO activating proteases (PAPs) (6, 7). The proteases in these pathways are regulated by protease inhibitors from the serpin superfamily (8).
The immune serine protease-SPH systems and their regulation by serpins have been studied in Drosophila melanogaster, Tenebrio molitor, Manduca sexta, and a few other holometabolous insects (2, 8–10). Genetic investigations revealed a pathway for Toll activation in Drosophila that includes an initiating modular serine protease (ModSP) and terminal Spätzle processing enzyme (SPE), with the clip protease Grass between these two proteases, but the substrates of ModSP and Grass have not yet been determined experimentally (11). An alternative pathway that leads to Spätzle activation involves the clip protease Persephone but does not require ModSP (11). Persephone can be activated by a combination of microbial proteases and a hemolymph cathepsin (12), which cleaves after an unusual His residue at the activation site of Persephone. The melanization cascade in D. melanogaster includes the clip proteases MP1, MP2, and Hayan (13, 14). MP2 is a PAP for Drosophila proPO1 (15). There is not a well-characterized linkage between the protease cascades that activate proSpätzle and proPO in D. melanogaster immune responses. However, the serpins Necrotic and Spn28Da impede both Toll and proPO activation in D. melanogaster (16, 17). Biochemical studies in a beetle, T. molitor, revealed a serpin-regulated cascade of three proteases, modular serine protease (MSP), SPE activating enzyme, and SPE. SPE then cleaves and activates SPH1, PO, and Spätzle precursors (7). A transcriptome study (18) revealed the immune stimulated expression of additional clip proteases, suggesting that the protease cascade system in T. molitor has additional complexity.
In a lepidopteran insect, M. sexta, a ModSP ortholog, hemolymph protease 14 (HP14), activates hemolymph protease 21 (HP21), which in turn activates PAP2 and PAP3 (19–21). The PAPs cleave and activate SPH1, SPH2, and proPOs (Fig. 1). Hemolymph protease 6 (HP6), an ortholog of Drosophila Persephone, cleaves proPAP1 for proPO activation and also cleaves hemolymph protease 8 (HP8), which activates proSpätzle-1 (22, 23) (Fig. 1). Like Persephone, proHP6 has a His residue at its activation cleavage site. Some evidence indicates that hemolymph protease 1 (HP1) may activate proHP6 to form HP6 (24, 25), but it is still unclear whether proHP6 activation may be linked to HP14 or HP21. Here we report our results indicating that HP5 acts as a previously unknown link between HP21 and HP6, cleaving proHP6 at its activation site His residue. We also found that HP5 is inhibited by multiple serpins present in hemolymph.
Fig. 1.
A simplified model of the serine protease network in M. sexta. The part directly surrounding HP5 (in bold font) is elucidated in the current study. The system outputs (colored red) include PO and Spätzle-1. Black arrows denote proteolytic activation of substrate proteins. Serpin-1A, Serpin-1J, and Serpin-4 inhibit M. sexta HP5.
Results
Structural Features, Expression Pattern, and Production of Recombinant proHP5.
M. sexta HP5 is a member of the CLIP family of proteases in subfamily B (2). The proHP5 cDNA sequence encodes a signal peptide and a group 2 clip domain, followed by a 96-residue linker region rich in Ser (11), Asp (10), Thr (9), Asn (9), Pro (9), and Gly (8) (SI Appendix, Fig. S1). The carboxyl-terminal protease domain contains a catalytic triad of His198-Asp260-Ser358, a primary specificity pocket of Asp352-Gly384-Gly395, which suggests a trypsin-like specificity, and a signature “75-loop” (5) of Cys225 through Cys234. The activation cleavage of proHP5 between R153 and I154 (see below) yields an amino-terminal light chain (16.7 kDa) and a carboxyl-terminal heavy chain (28.5 kDa) connected by an interchain disulfide bond.
HP5 is mainly expressed in larval fat body, epidermis, and trachea (SI Appendix, Fig. S2). The mRNA level in fat body increased dramatically after an immune challenge, suggesting a role in antimicrobial defense. Like some other immunity-related genes, HP5 mRNA levels increased in wandering and pupal stages in the absence of infection. We produced M. sexta proHP5 in a baculovirus–insect cell system. The purified proHP5 migrated as a 52-kDa band on the Coomassie-stained gel, which was recognized by HP5 antiserum produced using HP5 protein synthesized in Escherichia coli as antigen (Fig. 2A).
Fig. 2.
Involvement of M. sexta HP5 in melanization and antimicrobial peptide induction. (A) The 10% SDS/PAGE and immunoblot analysis of the purified proHP5 from Sf9 cells. Aliquots of the protein (1.0 µg for staining; 25 ng for immunodetection) were treated by SDS sample buffer, separated along with protein size markers, and detected by Coomassie brilliant blue (CBB; Left) or using 1:2,000 diluted HP5 antiserum (HP5 Ab; Right). Positions and sizes of the Mr makers are indicated. (B) Concentration-dependent proPO activation caused by proHP5 in the absence of a microbial elicitor. Aliquots of 1:10 diluted IP (5.0 µL) from day 2, fifth instar at 24 h after bacterial injection were incubated with various amounts of proHP5 for 45 min at 25 °C. PO activities were measured and plotted as mean ± SEM (n = 3). (C) Synergistic enhancement of M. luteus-elicited proPO activation by proHP5. Aliquots of 1:10 diluted IP (5.0 µL) were incubated with buffer A, M. luteus (0.1 µg), proHP5 (0.1 µg), or both for 45 min at 25 °C. PO activities were assayed and plotted as the bar graph (mean ± SEM, n = 3). Interactions of plasma with M. luteus and proHP5 both led to proPO activation. (D) Induction of antimicrobial peptide transcript levels. Fifty microliters of buffer A, proHP5 (4 μg), PAP3 (250 ng), or both in buffer A were incubated for 60 min at 25 °C and then injected into individual day 0, fifth instar larvae. At 24 h after injection, fat body RNA samples were prepared from three insects in each group for qRT-PCR analysis of the antimicrobial peptide mRNA levels in these samples (three biological samples, each pooled from three larvae, and three technical replicates per sample). After normalization against rpS3, relative antimicrobial peptide transcript levels after buffer A injection are taken as an internal control (i.e., adjusted to 1.00) for calculating antimicrobial peptide mRNA levels in other samples. Student’s t test is run to reveal statistical significance of the mRNA level changes. *P < 0.05; n.s. P > 0.05.
M. sexta HP5 Stimulates proPO Activation.
To test a possible role for HP5 in melanization, we incubated 100–500 ng proHP5 with diluted plasma from bacteria-induced larvae and detected increased PO activity that depended on the concentration of added proHP5 (Fig. 2B). Addition of 100 ng of recombinant proHP5 to plasma also significantly enhanced proPO activation triggered by M. luteus (Fig. 2C), to a degree more than twice the effect of the sum of the two individual treatments. Therefore, these results indicate that HP5 is a component of the proPO activation system in M. sexta that is triggered by recognition of bacteria.
ProHP5 Is Cleaved at Its Activation Site by M. sexta PAP3 and HP21.
ProHP5 must be cleaved by another protease at its zymogen activation site to elicit its activity during an immune response. We tested several hemolymph clip domain proteases (PAP1, PAP2, PAP3, HP2, and HP21) as proHP5 activators, based on their known specificity for cleavage after a basic residue, with the prediction that a proHP5 activator will cleave after Arg153 in proHP5. After incubation with active PAP3, the 52-kDa proHP5 was converted to a 30-kDa band visible with Coomassie staining and with detection by HP5 antibodies, consistent with the size of the C-terminal protease domain of HP5 (Fig. 3A). A 13-kDa band was visible after silver staining, corresponding to a light chain produced upon cleavage of proHP5. While HP5 and 6 × His antibodies recognized the C-terminal heavy chain, the HP5 antiserum did not detect the light chain including the clip domain, perhaps due to poor antigenicity. Automated Edman degradation showed that the amino-terminal sequence of the 30-kDa heavy chain was Ile-Ile-Gly-Gly-Asn-Arg, which verified cleavage of proHP5 between R153 and I154 and is consistent with PAP3’s known specificity for cleavage after Arg in its natural substrates (19–21, 26). In similar experiments we found HP21 can activate proHP5. We incubated the HP21 activated by HP14 with proHP5 and found that HP21 also produced a 30-kDa product of proHP5 (Fig. 3B). The modular protease HP14 itself did not cleave proHP5, which is consistent with its specificity for cutting after Leu387 in proHP14 (27) and Leu152 in proHP21 (19). In contrast, PAP1, PAP2, and HP2 did not activate proHP5 (SI Appendix, Fig. S3 A–C). Thus, HP21 and PAP3 are apparently specific activating enzymes of proHP5.
Fig. 3.
Proteolytic activation of M. sexta proHP5 by PAP3 and HP21. (A) PAP3 cleavage. Purified proHP5 (0.1 µg, 1.0 µL), PAP3 (20 ng, 1.0 µL), and buffer A (to 20 µL) were incubated at 25 °C for 1 h. The reaction mixture and controls lacking one of the proteins were separated by 10% SDS/PAGE under reducing condition and detected by immunoblotting using HP5 (Left) or 6×His (Right) antibodies or staining (Middle gel; lane 1, 0.8 µg proHP5; lane 2, 50 ng PAP3; and lane 3, both proHP5 and PAP3, CBB; lane 4, 800 ng proHP5 and 20 ng PAP3, silver stain, 15% gel, position of the 10 kDa marker in red). (B) HP21 cleavage. Purified proHP5 (25 ng), proHP21 (10 ng), HP14 (10 ng, from a mixture of preincubated 1.0 µg E. coli peptidoglycan with 0.3 µg PGRP1, 0.3 µg MBP, and 0.1 µg proHP14 in 24 µL buffer A at 37 °C for 1 h), and buffer A were incubated at 25 °C for 1 h. The reaction mixture and controls were subjected to 10% SDS/PAGE and immunoblot analysis using HP5 antibodies. Sizes and positions of the Mr markers are indicated on the left (except for the silver stained gel strip). The HP5 precursor, catalytic domain, and light chain are marked with circles, triangles, and asterisk, respectively.
ProHP6 and proSPH2 Are Substrates of Active HP5.
To investigate how HP5 stimulates proPO activation in hemolymph, we tested candidate substrate proteins known to act in this pathway, including proHP6, proSPHs, and proPO itself. HP6 activates proPAP1 (22); proteolytically activated SPH1 and SPH2 together act as a cofactor of the PAPs (28). We first tested whether HP5 can directly activate proHP6 (Fig. 4A). ProHP6 alone was detected as a 38-kDa band by immunoblot analysis using HP6 antiserum. After incubation with a mixture of PAP3 and proHP5, most of the proHP6 was converted to a 27-kDa band, consistent with the expected size of the carboxyl-terminal protease domain after activation cleavage. The amino-terminal sequence of this band was determined by automated Edman degradation to be Ile-Leu-Gly-Gly-Glu-Glu, indicating that HP5 cleaves proHP6 between His112 and Ile113, its predicted activation site. PAP3 in the absence of HP5 did not produce this cleavage of proHP6, indicating that HP5 activated by PAP3 was responsible for the observed cleavage of proHP6. In the control mixture of proHP5 and proHP6 without PAP3, a faint band at the same size as the HP6 protease domain was present, perhaps due to a small amount of active HP5 in the proHP5 preparation (Fig. 4A, lane 3). This is consistent with the PO activity increase after mixing proHP5 with induced plasma (Fig. 2 B and C).
Fig. 4.
Cleavage activation of proHP6 and proSPH2 by M. sexta HP5. (A) HP6. PAP3 (20 ng), proHP5 (25 ng), proHP6 (50 ng), and buffer A (to 12 µL) were incubated at 25 °C for 1 h. The reaction mixture and controls lacking one of the proteins were resolved by 10% SDS/PAGE under the reducing condition and detected by immunoblotting using HP6 antibodies. Sizes and positions of the Mr markers are indicated on the left. (B) SPH2. HP21 was produced in a mixture of 1.0 µg E. coli peptidoglycan with 0.3 µg PGRP1, 0.3 µg MBP, 0.1 µg proHP14, and 0.1 µg proHP21 in 30 µL buffer A at 37 °C for 1 h. HP21 (20 ng) was then incubated with proHP5 (25 µg), proSPH2 (0.1 µg), and buffer A (to 12 µL) at 25 °C for 1 h. This mixture and controls lacking one of the components were subjected to 10% SDS/PAGE and immunoblot analysis using SPH2 antibody. Arrowhead indicates HP6 catalytic domain and SPH2 heavy chain. (C) Function of SPH2 activated by HP5 along with proSPH1 as a PAP cofactor. A mixture of HP21 (20 ng), proHP5 (25 ng), PAP1 (20 ng), proSPH1 (0.1 µg), proSPH2 (0.1 µg), proPO (0.16 µg), and buffer A (to 20 µL) was incubated at 25 °C for 50 min. In addition to the negative controls lacking one of the components, a positive control included the natural complex of SPH1 and SPH2 (20 ng) isolated from hemolymph (30) reacted with PAP1 (20 ng) and proPO (0.16 µg) to generate active PO. PO activities were measured and plotted as mean ± SEM (n = 3). Student’s t test is run to reveal statistical significance of the PO activity differences between the indicated groups. *P < 0.05.
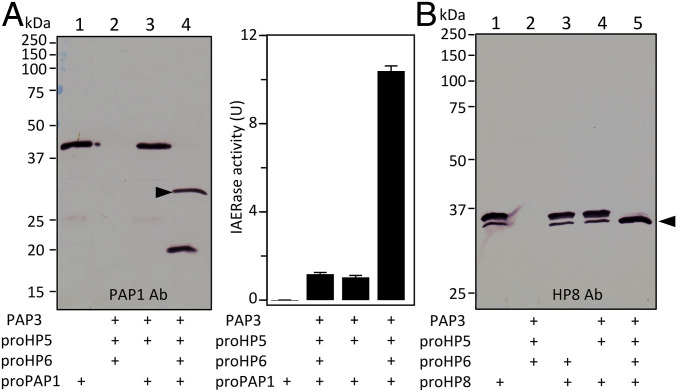
The finding that HP5 cleaves proHP6 after the His residue predicted to be its activation site led to experiments to test whether HP6 cleaved by HP5 is active against its natural substrates. We examined the proteolytic activation of proPAP1 and proHP8 by HP5-activated HP6 (Fig. 5). HP6 activated by HP5 was able to cleave and activate proPAP1, demonstrated by immunoblot results and PAP1 activity against a peptide substrate, IEAR-p-nitroanilide (Fig. 5A), and proHP8 (Fig. 5B). These results are very similar to those previously obtained using a proHP6 engineered to be activated Factor Xa (22). These data indicate that HP6 is likely activated by HP5 during immune responses, leading to stimulation by HP6 of melanization (by PAP1 activation) and the Toll pathway (by HP8 activation).
Fig. 5.
Proteolytic processing of proHP8 and proPAP1 by HP5-generated HP6. (A) Activation of proPAP1. PAP3 (20 ng), proHP5 (25 ng), proHP6 (50 ng), proPAP1 (50 ng), and buffer A (to 20 µL) were incubated at 25 °C for 1 h. The reaction mixture and controls were subjected to 10% SDS/PAGE and immunoblot analysis (Left) using PAP1 antibodies. In a duplicated experiment, amidase activities in the reaction and control mixtures were determined using 150 µL of 25 µM of IEARpNA (46) and plotted as mean ± SD (n = 3). (B) Activation of proHP8. PAP3 (20 ng), proHP5 (25 ng), proHP6 (50 ng), proHP8 (60 ng), and buffer (to 20 µL) were incubated at 25 °C for 1 h. Half of the reaction mixture and controls were separated by 10% SDS/PAGE under reducing condition and detected by immunoblotting using HP8 antibodies. Sizes and positions of the Mr markers are indicated. Arrowhead indicates catalytic domain.
We then tested if HP5 can cleave and activate proSPHs, which act as a cofactor for PAPs in producing active PO (28). In this experiment, we used HP21 (activated by HP14) instead of PAP3 to activate proHP5 because PAP3 cleaves proSPH1 and proSPH2 (20). Recombinant proSPH2 was detected by immunoblot analysis as a 52-kDa band, using SPH2 antiserum (Fig. 4B). Its mobility was not affected by proHP5 or HP21 alone. After treatment with a mixture of HP21 and proHP5, the 52-kDa proSPH2 amount decreased, and a 36-kDa immunoreactive band appeared (Fig. 4B, lane 4). This result is consistent with cleavage of proSPH2 by HP5, producing a product similar to that previously observed after activation of proSPH2 by PAP3 (20). In contrast, proSPH1 did not appear to be a substrate for HP5 (SI Appendix, Fig. S3D).
We tested whether proSPH2 cleaved by HP5 can promote proPO activation by PAP1. PAP1 alone produced a low level of activation of proPO, and a mixture of proSPH1, proSPH2, HP21, and proHP5 (which includes proSPH2 cleaved by HP5) did not activate proPO (Fig. 4C). However, addition of the mixture including proSPH2 cleaved by HP5 resulted in 3.7-fold higher PO activity. This increase was not observed when either proSPH1 or proSPH2 was absent from the reaction. The results indicate that both SPH1 and SPH2 are required to form a functional cofactor for PAP1. The enhanced PAP1 activity was less than that observed when active complex of SPH1 and SPH2 purified from hemolymph was added to the reaction, perhaps because SPH1 was fully cleaved and active in this preparation (29).
Because the His residue at the activation cleavage site of proHP6 may not be charged at physiological pH, we investigated the effect of pH on the activity of HP5 in cleaving proHP6 (SI Appendix, Fig. S4). HP5 had little activity at pH 5, and its activity increased with increasing pH, with maximum activity at pH 7 to 8. This experiment is complicated by the titration of the His residue in the catalytic triad of HP5 but suggests that HP5 can efficiently cleave at an uncharged His112 in proHP6.
M. sexta HP5 Induces Antimicrobial Peptide Production.
The finding that HP6 activated by HP5 can cleave proHP8 led to testing if HP5 can trigger the Toll pathway via the pathway of HP6, HP8, and Spätzle-1 (Fig. 1). As shown in Fig. 2D, injection of buffer or PAP3 into larvae failed to induce antimicrobial peptide synthesis, whereas injection of proHP5 or PAP3-activated HP5 significantly increased mRNA levels of attacins, cecropin-6, moricin, gloverin, and lysozyme. These results are consistent with a hypothesis that HP5 cleaves and activates proHP6, leading to activation of HP8 and Spätzle-1, which can bind to the Toll receptor, stimulating downstream signal transduction leading to synthesis of antimicrobial peptides (22).
Inhibitory Regulation of M. sexta HP5 by Serpins.
To test whether HP5 is inhibited by a serpin, we incubated plasma for 30 min with M. luteus to activate immune protease cascades, in the presence of PO inhibitors to prevent quinone cross-linking of proteins. During this time, serine proteases are expected to be activated and then may also be inhibited by serpins in plasma. Then beads coupled with HP5 antibodies were added to the plasma. After washing the beads, the bound proteins were eluted at low pH, separated by SDS/PAGE, and subjected to immunoblot analysis. A 75-kDa band, consistent with the expected size of an HP5-serpin complex was recognized by antibodies to HP5, serpin-1, and serpin-4 (Fig. 6A) but not by antibodies to serpin-2, serpin-3, serpin-5, serpin-6, serpin-9, serpin-12, or serpin-13 (SI Appendix, Fig. S5A). We have shown that HP5 cleaves proHP6 after a His residue, a very unusual protease specificity. None of the M. sexta serpins has a His residue at the predicted P1 residue of the scissile bond of reactive center loop that determines primary selectivity of serpins in protease inhibition (30). Serpin-4 has an Arg residue at the P1 site (31). Alternate exon splicing in M. sexta serpin-1 can lead to 14 different serpin-1 isoforms with different reactive center loops that interact with target proteases (30). Serpin-1A, serpin-1J, and serpin-1Y have Arg residues at this position, like serpin-4. To confirm the results of immunopurification of HP5-serpin complexes from plasma, we tested whether purified serpin-1A, serpin-1J, and serpin-4 could form a complex with HP5, after its activation by PAP3. ProHP5 alone was detected as a 55-kDa band, and after treatment with PAP3, it was cleaved, releasing the band at 30 kDa containing the catalytic domain (Fig. 6B). When serpin-1J, serpin-1A, or serpin-4 were present in the reaction, the band representing the 30-kDa catalytic domain became much less intense, and a band at 75 kDa, consistent with the size of an SDS-stable HP5-serpin complex, appeared (Fig. 6C and SI Appendix, Fig. S6 A–C). In contrast, serpin-3 with K375 at the P1 position did not form a complex with HP5 (SI Appendix, Fig. S5B). Furthermore, in the presence of serpin-1J or serpin-4, HP5 did not cleave its substrate proHP6 (Fig. 6C and SI Appendix, Fig. S6D), indicating that these serpins inhibited the proteolytic activity of HP5. Therefore, it is likely that HP5 is specifically regulated by these serpin-1 variants and serpin-4 in hemolymph.
Fig. 6.
Formation of an SDS-stable complex of HP5 complex with M. sexta serpins. (A) Composition of the serpin-protease complex (SPC) isolated from induced plasma (IP). Protease cascades in IP were stimulated by addition of M. luteus in the presence of 1-phenyl-2-thiourea to prevent melanization at 25 °C for 30 min. Following phenylmethylsulfonyl fluoride treatment to inhibit remaining protease activity, the HP5-serpin complexes (along with proHP5 and its cleavage products) were isolated by affinity chromatography on an HP5 antibody column. Elution fraction 4 was subjected to 10% SDS/PAGE followed by immunoblot analysis (Left) or CBB staining (Right). Arrowhead indicates SPC. (B) proHP6 activation blocked by inhibition of HP5 by serpin-1J. PAP3 (20 ng), proHP5 (25 ng), proHP6 (50 ng), and serpin-1J (1 µg) were incubated at 25 °C for 10 min. The samples were separated by 10% SDS/PAGE and detected by immunoblot analysis using HP6 antibodies. (C) SPC formation using purified proteins. PAP3-treated proHP5 was reacted with serpin-1J at 25 °C for 10 min. The mixture and controls were subjected to 10% SDS/PAGE and immunoblotting. Sizes of the molecular mass markers are indicated.
Discussion
Integration of the Serine Protease-SPH Network in M. sexta Larval Hemolymph.
Although initial data suggested distinct protease cascade pathways for activation of prophenoloxidase and the Toll pathway in insect hemolymph, it is becoming apparent that these immunity-activating pathways occur as interacting networks or webs. M. sexta HP6 induces antimicrobial peptide synthesis via a pathway which includes HP8 and Spätzle-1, and it stimulates melanization via PAP1, SPH2, and POs (Fig. 1) (22, 23). In this study, we demonstrated that these pathways are linked at other levels: proHP5 is activated by HP21 or PAP3. HP5 then cleaves proHP6 to generate active HP6, which can activate the PO pathway (by cleaving PAP1) or the Toll pathway (by cleaving HP8). The discovery of the HP14–HP21–HP5–HP6–(HP8, PAP1) pathway links activation of HP6 to microbe recognition through HP14 and HP21. HP14 is an initiation protease that self-activates after proHP14 associates with bacterial peptidoglycans or fungal β-1,3-glucan via binding to specific recognition proteins (28, 32, 33).
M. sexta HP5, a CLIPB Protease That Cleaves after His.
We have not previously known what protease can activate proHP6 by cleaving at its unique histidine activation site residue (2). Likewise, the endogenous proteases that can activate Drosophila Persephone and Hayan by cleavage after histidine are unknown (although some microbial proteases can activate Persephone). The branch-point enzyme proHP6 has a predicted activation site of LDLH*ILGG, which was precisely cleaved by HP5 at His112 to generate active HP6 (Fig. 4A). Since there is currently no protease in the MEROPS protease database known to preferentially cleave after a histidine residue (34), it is a unique finding that an endogenous insect clip-domain serine protease can cleave after His112 to activate proHP6, leading to PO and Spätzle-1 production. It may be surprising that the neutral charge status of His112 at physiological pH does not affect proHP6 cleavage (SI Appendix, Fig. S4), as the substrate binding pocket residues of HP5 suggest a trypsin-like specificity.
Another CLIP protease, HP1, can hydrolyze LDLH-p-nitroanilide, which corresponds to the LDLH P4−P1 region before the proteolytic activation site of proHP6, and addition of the catalytic domain of HP1 generated active HP6 (24). ProHP1 can be activated by incubation with a cationic detergent, cetylpyridinium chloride (CPC), without proteolytic cleavage, suggesting that a conformational change in proHP1, not induced by proteolysis, may allow proHP1* (asterisk indicates active conformation) to cleave proHP6 between H112 and I113 (25). This indicates an alternate pathway for activating proHP6 during immune responses, perhaps through interaction of proHP1 with hydrophobic surfaces.
Identification of HP5 as an activator of HP6 by cleavage at a His residue at its activation site may help to make predictions regarding activation of similar proteases in other insects. D. melanogaster Persephone and Hayan, Anopheles gambiae CLIPC4−6 and CLIPC10, Apis mellifera cSP14, and Tribolium castaneum cSP66 are all CLIP proteases that have a putative activation site of H*I (9). These apparent HP6 orthologs (9) may be activated by the HP5 orthologs present in these species. Grass/cSP1, one of the two Drosophila HP5 orthologs, may be involved in the activation of Persephone (cSP28), Hayan (cSP31), and Ser7 (cSP3) to induce the Toll pathway and melanization (35). While genetic evidence is lacking, our data suggest a possible mechanism of direct activation of cSP28 and cSP31 by cSP1. The unusual specificity of HP5 for cleavage at His residues may be conserved in orthologs in holometabolous insects, and this may provide a unique selectivity for control at a key position of the protease cascades that stimulate immune responses.
Materials and Methods
Detailed materials and methods are provided in SI Appendix.
Immune Challenge, Plasma and Tissue Collection, cDNA Synthesis, and qRT-PCR Analysis.
Day 2, fifth instar larvae of M. sexta were injected with a mixture of E. coli, M. luteus, and curdlan (32). Hemolymph was collected 24 h later and centrifuged to remove hemocytes and obtain induced plasma (IP). Hemocytes, fat body, and other tissues were dissected from day 3, naïve larvae for RNA isolation. Similarly, fat body tissues were collected at various life stages to prepare total RNA. cDNA synthesis and qRT-PCR analysis of HP5 and antimicrobial peptide mRNA levels were performed using specific primers for them and rpS3. Amplification efficiencies of the cDNA fragments were determined as described before (36). Relative HP5 and antimicrobial peptide mRNA levels were calculated as (1 + ErpS3)Ct, rpS3/(1 + Ex)Ct, x.
Protein Preparation.
M. sexta PGRP1, microbe binding protein (MBP), proPAP1, proSPH1, proSPH2, proHP21, proHP6, and proHP8 were expressed in baculovirus-infected Sf9 cells (19, 37–41). Serpin-1A, Serpin-1J, Serpin-1J′, and Serpin-4 were produced in E. coli as soluble proteins and purified on Ni-NTA agarose columns (31, 42). M. sexta proHP14, PAP1, PAP3, and proPO were isolated from hemolymph and stored at −80 °C (26, 27, 43, 44).
M. sexta HP5 cDNA Cloning, E. coli Expression, and Antiserum.
A fragment of the HP5 cDNA was amplified from clone C2 (45) by PCR and then labeled with α-32P-dCTP to screen a cDNA library for full-length clones. The longest HP5 cDNA was isolated, sequenced, and amplified using primers j550 and j015. The DNA was subcloned into H6pQE60 for expression in E. coli. The recombinant protein was solubilized in 8 M urea and purified by nickel affinity chromatography, followed by preparative SDS/PAGE. The resulting protein was used as antigen to generate a rabbit antiserum (45).
Production of M. sexta proHP5 in the Baculovirus Expression System.
Primers j1353 and j591 were used to amplify a cDNA fragment encoding the mature proHP5 with an amino-terminal extension of GIR, followed by proHP5 residues 2 to 412 and then a carboxyl-terminal tag with the sequence LEHHHHHH. The PCR product was cloned into pMFH6 (40), in frame with an amino-terminal secretion peptide sequence encoded in this vector. The plasmid was used to produce a bacmid and a high-titer viral stock for infecting Sf9 cells (37). According to the same paper, proHP5 was purified from the conditioned media, concentrated, and stored at −80 °C.
Roles of HP5 in proPO Activation and Induced Antimicrobial Peptide Synthesis.
Aliquots of 1:5 diluted IP (5 μL) were incubated with various amounts of proHP5 in 20 μL buffer A for 45 min prior to PO activity assay using dopamine as substrate (46). In another experiment, 1:10 diluted IP was incubated with buffer A and then mixed with M. luteus, proHP5, or both for 45 min before PO activity assay. To test a possible role of HP5 in antimicrobial peptide gene expression, buffer A, proHP5, PAP3, and both were incubated for 60 min, and these samples were injected into each of day 0, fifth instar larvae. Total RNA samples were prepared from fat body tissues 24 h later for cDNA synthesis and qRT-PCR analysis.
Proteolytic Activation of proHP5 by PAP3 and HP21.
To test if PAP3 can activate proHP5, proHP5 was incubated with PAP3 and buffer A for 1 h at 25 °C. In the controls, proHP5, PAP3, or both was replaced by equal volume of buffer A. After separation by SDS/PAGE, the gels were visualized by Coomassie blue or silver staining and immunoblot analysis using HP5 or 6 × His antibody. Specific proteolysis was revealed by changes in gel mobility. Similarly, to test if HP21 can activate proHP5, proHP5 was incubated with proHP21, HP14, or both for 1 h (19). HP14 was generated in a mixture of E. coli peptidoglycan, MBP, PGRP1, and proHP14 (32, 33).
Cleavage Activation of proHP6 and proSPH2 by HP5.
To test proHP6 activation by HP5, a mixture of PAP3, proHP5, proHP6, and buffer A were incubated for 1 h, along with controls that lack one or two of the proteins. Since PAP3 directly activates proSPH1 and proSPH2 (20), HP5-generated by HP21 instead of PAP3 was used to test the activation of proSPH2. After the cleavage of proSPH2 was confirmed, the cofactor activity of SPH2 and proSPH1 was examined using PAP1 and proPO (40). The reactions and controls were analyzed by SDS/PAGE and immunoblotting using antibodies against HP6 or SPH2.
Determination of the Newly Exposed N-Terminal Sequences of HP5 and HP6.
HP5 (1.0 µg) after cleavage by PAP3 as described above was resolved by SDS/PAGE and transferred to a PVDF membrane. After staining, the band suspected to be the catalytic domain of HP5 was subjected to six cycles of automated Edman degradation. The cleavage site in proHP5 was deduced by comparing the experimental data with the HP5 sequence. To locate the position in proHP6 cleaved by HP5, PAP3 was used to activate HP5, which was then incubated with proHP6 in the same way, and the resulting 30-kDa band was analyzed by Edman degradation.
Cleavage Activation of proPAP1 and proHP8 by HP5-Activated HP6.
To confirm if HP6 cleaved by HP5 was active, PAP3, proHP5, proHP6, and buffer A were incubated with proPAP1 or proHP8 for 1 h at 25 °C. The reaction mixtures and controls that lacked one or two of the proteins were analyzed by SDS/PAGE and immunoblotting using PAP1 or HP8 antibody. To detect PAP1’s amidase activity, the duplicated reaction mixture and controls were incubated with 25 µM acetyl-Ile-Glu-Ala-Arg-p-nitroanilide in a microplate assay (46).
Identification of the Components of HP5-Serpin Complexes.
IgG-coupled protein A-Sepharose beads were prepared using HP5 antiserum (10). Induced plasma was mixed with M. luteus to activate immune protease cascades. HP5 and associated proteins were isolated using the antibody beads and subjected to SDS/PAGE, staining, and immunoblotting. Formation of serpin–protease complexes (SPCs) was verified in vitro by incubating PAP3-activated HP5 with the serpins. Immunoblotting was performed to detect SPCs using antibodies to serpin-1 or HP5. To test the effect of serpin-1J inhibition on activation of proHP6 by HP5, PAP3 and proHP5 were preincubated for 1 h to activate HP5. After HP5 had been treated with serpin-1J for 15 min, proHP6 was added to the reaction for 1 h incubation. The reaction mixture and controls were analyzed by immunoblotting using HP6 antibody.
Supplementary Material
Acknowledgments
This study was supported by NIH grants GM58634 and AI139998. The paper was approved for publication by the Director of Oklahoma Agricultural Experimental Station and supported in part under project OKL03054. We thank Dr. Steve Hartson for his assistance in the mass spectrometric analysis.
Footnotes
The authors declare no competing interest.
This article is a PNAS Direct Submission. D.S.S. is a guest editor invited by the Editorial Board.
This article contains supporting information online at https://www.pnas.org/lookup/suppl/doi:10.1073/pnas.2004761117/-/DCSupplemental.
Data Availability.
The nucleotide sequence reported in this paper had been deposited in the GenBank/EBI Data bank under accession no. MN364869. All study data are included in the article and SI Appendix.
References
- 1.Krem M. M., Di Cera E., Evolution of enzyme cascades from embryonic development to blood coagulation. Trends Biochem. Sci. 27, 67–74 (2002). [DOI] [PubMed] [Google Scholar]
- 2.Kanost M. R., Jiang H., Clip-domain serine proteases as immune factors in insect hemolymph. Curr. Opin. Insect Sci. 11, 47–55 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Takahashi D., Garcia B. L., Kanost M. R., Initiating protease with modular domains interacts with β-glucan recognition protein to trigger innate immune response in insects. Proc. Natl. Acad. Sci. U.S.A. 112, 13856–13861 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Kanost M. R., Gorman M. J., “Phenoloxidases in insect immunity” in Insect Immunology, Beckage N., Ed. (Academic Press, San Diego, CA, 2008), pp. 69–96. [Google Scholar]
- 5.Veillard F., Troxler L., Reichhart J. M., Drosophila melanogaster clip-domain serine proteases: Structure, function and regulation. Biochimie 122, 255–269 (2016). [DOI] [PubMed] [Google Scholar]
- 6.Jiang H., Vilcinskas A., Kanost M. R., Immunity in lepidopteran insects. Adv. Exp. Med. Biol. 708, 181–204 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Park J. W. et al., Beetle immunity. Adv. Exp. Med. Biol. 708, 163–180 (2010). [DOI] [PubMed] [Google Scholar]
- 8.Meekins D. A., Kanost M. R., Michel K., Serpins in arthropod biology. Semin. Cell Dev. Biol. 62, 105–119 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Cao X., Jiang H., Building a platform for predicting functions of serine protease-related proteins in Drosophila melanogaster and other insects. Insect Biochem. Mol. Biol. 103, 53–69 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Wang Y. et al., Inhibition of immune pathway-initiating hemolymph protease-14 by Manduca sexta serpin-12, a conserved mechanism for the regulation of melanization and Toll activation in insects. Insect Biochem. Mol. Biol. 116, 103261 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Buchon N. et al., A single modular serine protease integrates signals from pattern-recognition receptors upstream of the Drosophila Toll pathway. Proc. Natl. Acad. Sci. U.S.A. 106, 12442–12447 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Issa N. et al., The circulating protease Persephone is an immune sensor for microbial proteolytic activities upstream of the Drosophila Toll pathway. Mol. Cell 69, 539–550.e6 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Tang H., Kambris Z., Lemaitre B., Hashimoto C., Two proteases defining a melanization cascade in the immune system of Drosophila. J. Biol. Chem. 281, 28097–28104 (2006). [DOI] [PubMed] [Google Scholar]
- 14.Nam H. J., Jang I. H., You H., Lee K. A., Lee W. J., Genetic evidence of a redox-dependent systemic wound response via Hayan protease-phenoloxidase system in Drosophila. EMBO J. 31, 1253–1265 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.An C., Zhang M., Chu Y., Zhao Z., Serine protease MP2 activates prophenoloxidase in the melanization immune response of Drosophila melanogaster. PLoS One 8, e79533 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Levashina E. A. et al., Constitutive activation of toll-mediated antifungal defense in serpin-deficient Drosophila. Science 285, 1917–1919 (1999). [DOI] [PubMed] [Google Scholar]
- 17.Scherfer C. et al., Drosophila Serpin-28D regulates hemolymph phenoloxidase activity and adult pigmentation. Dev. Biol. 323, 189–196 (2008). [DOI] [PubMed] [Google Scholar]
- 18.Johnston P. R., Makarova O., Rolff J., Inducible defenses stay up late: Temporal patterns of immune gene expression in Tenebrio molitor. G3 4, 947–955 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Wang Y., Jiang H., Reconstitution of a branch of the Manduca sexta prophenoloxidase activation cascade in vitro: Snake-like hemolymph proteinase 21 (HP21) cleaved by HP14 activates prophenoloxidase-activating proteinase-2 precursor. Insect Biochem. Mol. Biol. 37, 1015–1025 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Wang Y., Lu Z., Jiang H., Manduca sexta proprophenoloxidase activating proteinase-3 (PAP3) stimulates melanization by activating proPAP3, proSPHs, and proPOs. Insect Biochem. Mol. Biol. 50, 82–91 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Gorman M. J., Wang Y., Jiang H., Kanost M. R., Manduca sexta hemolymph proteinase 21 activates prophenoloxidase-activating proteinase 3 in an insect innate immune response proteinase cascade. J. Biol. Chem. 282, 11742–11749 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.An C., Ishibashi J., Ragan E. J., Jiang H., Kanost M. R., Functions of Manduca sexta hemolymph proteinases HP6 and HP8 in two innate immune pathways. J. Biol. Chem. 284, 19716–19726 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.An C., Jiang H., Kanost M. R., Proteolytic activation and function of the cytokine Spätzle in the innate immune response of a lepidopteran insect, Manduca sexta. FEBS J. 277, 148–162 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Yang F., Wang Y., He Y., Jiang H., In search of a function of Manduca sexta hemolymph protease-1 in the innate immune system. Insect Biochem. Mol. Biol. 76, 1–10 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.He Y., Wang Y., Yang F., Jiang H., Manduca sexta hemolymph protease-1, activated by an unconventional non-proteolytic mechanism, mediates immune responses. Insect Biochem. Mol. Biol. 84, 23–31 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Jiang H., Wang Y., Yu X.-Q., Zhu Y., Kanost M., Prophenoloxidase-activating proteinase-3 (PAP-3) from Manduca sexta hemolymph: A clip-domain serine proteinase regulated by serpin-1J and serine proteinase homologs. Insect Biochem. Mol. Biol. 33, 1049–1060 (2003). [DOI] [PubMed] [Google Scholar]
- 27.Wang Y., Jiang H., Interaction of β-1,3-glucan with its recognition protein activates hemolymph proteinase 14, an initiation enzyme of the prophenoloxidase activation system in Manduca sexta. J. Biol. Chem. 281, 9271–9278 (2006). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Yu X. Q., Jiang H., Wang Y., Kanost M. R., Nonproteolytic serine proteinase homologs are involved in prophenoloxidase activation in the tobacco hornworm, Manduca sexta. Insect Biochem. Mol. Biol. 33, 197–208 (2003). [DOI] [PubMed] [Google Scholar]
- 29.Wang Y., Jiang H., Prophenoloxidase (proPO) activation in Manduca sexta: An analysis of molecular interactions among proPO, proPO-activating proteinase-3, and a cofactor. Insect Biochem. Mol. Biol. 34, 731–742 (2004). [DOI] [PubMed] [Google Scholar]
- 30.Li M. et al., The Manduca sexta serpinome: Analysis of serpin genes and proteins in the tobacco hornworm. Insect Biochem. Mol. Biol. 102, 21–30 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Tong Y., Jiang H., Kanost M. R., Identification of plasma proteases inhibited by Manduca sexta serpin-4 and serpin-5 and their association with components of the prophenol oxidase activation pathway. J. Biol. Chem. 280, 14932–14942 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Wang Y., Jiang H., Prophenoloxidase activation and antimicrobial peptide expression induced by the recombinant microbe binding protein of Manduca sexta. Insect Biochem. Mol. Biol. 83, 35–43 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Wang Y., Jiang H., Binding properties of the regulatory domains in Manduca sexta hemolymph proteinase-14, an initiation enzyme of the prophenoloxidase activation system. Dev. Comp. Immunol. 34, 316–322 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Rawlings N. D., et al. , The MEROPS database of proteolytic enzymes, their substrates and inhibitors in 2017 and a comparison with peptidases in the PANTHER database. Nucleic Acids Res. 46, D624−D632 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Dudzic J. P., Hanson M. A., Iatsenko I., Kondo S., Lemaitre B., More than black or white: Melanization and Toll share regulatory serine proteases in Drosophila. Cell Rep. 27, 1050–1061.e3 (2019). [DOI] [PubMed] [Google Scholar]
- 36.Schrag L. G. et al., Expression and characterization of Manduca sexta stress responsive peptide-1, an inducer of antimicrobial peptide synthesis. Biochem. Mol. Biol. 4, 42–52 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Sumathipala N., Jiang H., Involvement of Manduca sexta peptidoglycan recognition protein-1 in the recognition of bacteria and activation of prophenoloxidase system. Insect Biochem. Mol. Biol. 40, 487–495 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Wang Y., Sumathipala N., Rayaprolu S., Jiang H., Recognition of microbial molecular patterns and stimulation of prophenoloxidase activation by a β-1,3-glucanase-related protein in Manduca sexta larval plasma. Insect Biochem. Mol. Biol. 41, 322–331 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Wang Y., Jiang H., Kanost M. R., Expression and purification of Manduca sexta prophenoloxidase-activating proteinase precursor (proPAP) from baculovirus-infected insect cells. Protein Expr. Purif. 23, 328–337 (2001). [DOI] [PubMed] [Google Scholar]
- 40.Lu Z., Jiang H., Expression of Manduca sexta serine proteinase homolog precursors in insect cells and their proteolytic activation. Insect Biochem. Mol. Biol. 38, 89–98 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Wang Y., Jiang H., A positive feedback mechanism in the Manduca sexta prophenoloxidase activation system. Insect Biochem. Mol. Biol. 38, 763–769 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Jiang H., Kanost M. R., Characterization and functional analysis of 12 naturally occurring reactive site variants of serpin-1 from Manduca sexta. J. Biol. Chem. 272, 1082–1087 (1997). [DOI] [PubMed] [Google Scholar]
- 43.Gupta S., Wang Y., Jiang H., Purification and characterization of Manduca sexta prophenoloxidase-activating proteinase-1, an enzyme involved in insect immune responses. Protein Expr. Purif. 39, 261–268 (2005). [DOI] [PubMed] [Google Scholar]
- 44.Jiang H., Wang Y., Ma C., Kanost M. R., Subunit composition of pro-phenol oxidase from Manduca sexta: Molecular cloning of subunit ProPO-P1. Insect Biochem. Mol. Biol. 27, 835–850 (1997). [DOI] [PubMed] [Google Scholar]
- 45.Jiang H. et al., Molecular identification of a bevy of serine proteinases in Manduca sexta hemolymph. Insect Biochem. Mol. Biol. 35, 931–943 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Jiang H., Wang Y., Yu X. Q., Kanost M. R., Prophenoloxidase-activating proteinase-2 from hemolymph of Manduca sexta. A bacteria-inducible serine proteinase containing two clip domains. J. Biol. Chem. 278, 3552–3561 (2003). [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The nucleotide sequence reported in this paper had been deposited in the GenBank/EBI Data bank under accession no. MN364869. All study data are included in the article and SI Appendix.