Abstract
Background
The XPEN60 CRP&SAA (hereafter XPEN60) is a new automated hematology analyzer that can rapidly detect C‐reactive protein (CRP), serum amyloid A (SAA), and blood cell counts (CBC), including the 5‐part differential of white blood cells (5‐DIFF). The aim of this study was to evaluate the analytical performance of XPEN60.
Methods
The analytical performance of XPEN60 was evaluated on the basis of several parameters, including the limit of blank (LoB), limit of detection (LoD), limit of quantitation (LoQ), precision, accuracy, carryover, linearity, clinical reportable range (CRR), and interference test. In addition, method comparisons between CBC and 5‐DIFF, CRP, and SAA were performed on several systems.
Results
Total imprecision and accuracy for all parameters fell within acceptable criteria, and excellent measurements were observed in the dilution linearity (coefficient of determination, R 2 > .99). LoBs and LoDs (0 and 0.21 mg/L for CRP, 1.1 and 2.27 mg/L for SAA) satisfy the manufacturer's statement. LoQs were 0.61 and 3.62 mg/L for CRP and SAA, respectively. No significant carryover or interference tests (<10%) were observed in this study. The comparison analysis demonstrated strong agreement between XPEN60 results and those of Sysmex–XN1000 (XN1000), except for basophils (Bas) and eosinophils (Eos). The data correlated well with E601 and Mindray CRP‐M100 for CRP.
Conclusion
XPEN60 was demonstrated satisfactory analytical performance, which made it well‐suited for use in clinical laboratories, emergency departments, and community hospitals.
Keywords: automated hematology analyzer, complete blood count, C‐reactive protein, performance evaluation, serum amyloid A
1. INTRODUCTION
A complete blood cell count (CBC) including a 5‐part differential of white blood cells (5‐DIFF), C‐reactive protein (CRP), and serum amyloid A (SAA) are routine indicators for patients with bacterial or viral infections. 1 , 2 In mild inflammation, SAA is commonly observed at medium concentrations (10‐100 mg/L). 3 A combination of CBC, CRP, and SAA was also a better indicator to determine disease severity. 4 , 5 Therefore, an automated hematology analyzer that can simultaneously measure CBC, CRP, and SAA is of significance in various clinical applications. However, most instruments can only test one parameter or CRP and CBC simultaneously at present. 6 , 7
XPEN60 was recently released by Shenzhen XPENARRAY Biotechnology Co., Ltd. (XPENARRAY). It is equipped with SAA mode, CRP mode, and CBC + DIFF mode and is currently the only automated hematology analyzer that can simultaneously test CRP, SAA, and CBC including a 5‐DIFF. Sample identification is achieved either by barcode or manual input. The instrument performs the sample from an open serum, plasma separation tube, or EDTA tube. It requires only 30 µL of whole blood or plasma for determination of CBC, CRP, or SAA, and its throughput is approximately 60 samples per hour. It takes about 1 minute from pressing the start button to complete the test, and results are displayed by figure and clear graphics. It should be noted that XPEN60 offers no reticulocyte or nucleated red blood cell (NRBC) count. It generates abnormal cell flags in the presence of abnormal cell lines, indicating that the results are unreliable. For the pathologic samples, re‐examination was tested with manual microscopic according to the recommendation 8 and good laboratory practice.
This study was designed to evaluate the performance of the new XPEN60 system in Department of Laboratory Medicine of Luohu District People's Hospital according to the Clinical and Laboratory Standard Institute's (CLSI) Guidelines and ICSH guidelines. 9 In particular, emphasis was placed on the assessment of CRP and SAA.
2. MATERIALS AND METHODS
2.1. Instruments
The CBC + DIFF mode of XPEN60 was calibrated using fresh whole blood against the site's Sysmex–XN1000 (Sysmex) instrument, according to the recommendations of the International Council for Standardization in Haematology (ICSH). 9 The CRP and SAA modes were calibrated to standard materials ERM®‐DA 474 IFCC and WHO‐standard materials (NIBSC code:92/680), respectively.
The instrument has the following dimensions: 700 mm width × 650 mm depth × 750 mm height. As it is a smaller size than the ADVIA analyzers, it is more feasible for the laboratory with limited space. XPEN60 uses various technologies, of which direct current impedance with hydrodynamic focusing is applied to count the red blood cells (RBC), platelets (PLT), and WBC. The 5‐DIFF is separated by cytochemical myeloperoxidase and laser light scattering principles. Hemoglobin (HGB) is determined through sodium lauryl sulfate (SLS) spectrophotometry. In addition, CRP and SAA are tested via latex‐enhanced scatter immunoturbidimetric technologies.
2.2. Reagents
As XPEN60 is a matching analytical system, the reagents and calibrators were obtained from the manufacturer. For quality control throughout the study and imprecision assessment, the compound serum control material (Lot 20190604), including CRP and SAA, was produced by the manufacturer also and stored at −20°C. The blood control material (Lot 20190612) was purchased from Mindray Medical International and stored at 2‐8°C.
2.3. Specimen
In current study, all samples were remnants collected from the testing of clinical patients anticoagulated with EDTA in Luohu District People's Hospital Department of Laboratory Medicine. All samples were either used to test CBC counts or centrifuged (3200 × g for 10 minutes) to obtain plasma for SAA and CRP detection. Samples were stored at room temperature and tested within 4 hours after collection. According to CLSI EP9‐A3, 10 a total number of 120 samples were collected, from which every 40 samples were used for CBC, SAA, and CRP, respectively. Those samples were enrolled with the criteria of abnormally high value, low value as well as in the normal range. Besides, the samples contained hemato‐oncological cases and bacterial or viral infection cases. Over 50% of samples were collected from patients with infection that had concentrations >10 mg/L for CRP and SAA test.
2.4. Statistical analysis
Data were performed using GraphPad Prism (version 7.0), MedCalc (version 17.9.7), and SPSS (IBM SPSS, Version 24.0) for statistical analysis. Imprecision was calculated using one‐way analysis of variance and expressed as the standard deviation (SD) and coefficient of variation (%CV). Grubb's test was used to detect outliers. All data were processed with SPSS for normality test using the Shapiro‐Wilk. Pearson's correlation coefficient test and Passing‐Bablok regression were used to evaluate the correlation between methods, and a correlation coefficient (r) ≥ .90 was set as the threshold value for a very high correlation. A scatter plot was used to detect outliers, while the Bland‐Altman test was used to identify the mean difference and 95% limits of agreement.
2.5. Precision
Ten measurements were performed with a run on a single selected clinical sample (4.94 × 109/L for WBC, 6.12 × 1012/L for RBC, 151.3 g/L for HGB, 76.18 fL for MCV and 219.2 × 109/L for PLT, respectively) to evaluate repeatability (ie, within‐run precision) of CBC mode. The total imprecision (ie, within‐laboratory precision) was evaluated using whole‐blood control materials with low, medium, and high concentrations. The control materials had been analyzed five times per day for five consecutive days.
The repeatability and within‐laboratory precision of CRP and SAA were evaluated using plasma control materials with three identical levels (4.88, 38.45, and 106.37 mg/L for CRP and 10.2, 43.81, and 167.65 mg/L for SAA). In each day, duplicates for each control were tested per run for two runs (at least a 2‐hour separation between runs), which lasted for 20 days, according to EP5‐A3. 11 The control materials were separated into 40 aliquots and stored at − 20°C.
2.6. Carryover
Carryover was evaluated for WBC, RBC, HGB, PLT, CRP, and SAA. According to CLSI H26‐A2, 12 a clinical sample with high concentration was tested three times (H1, H2, and H3), followed by another low‐concentration sample tested with the same manner (L1, L2, and L3). The carryover was calculated with the following formula: carryover % = (L1‐L3)/(H3‐L3) × 100%. The samples for the carryover study were prepared with the following concentrations: WBC (high: 24.8 × 109/L, low: 1.7 × 109/L), HGB (high: 238.7 g/L, low: 44.3 g/L), RBC (high: 12.1 × 1012/L, low: 1.6 × 1012/L), PLT (high: 440 × 109/L, low: 21.7 × 109/L), CRP (high: 131.6 × 1012/L, low: 8.5 mg/L), and SAA (high: 204.8 × 1012/L, low: 4.3 mg/L).
2.7. Linearity
Linearity was assessed using several levels of pools that were diluted at fixed ratios from a high‐concentration sample. Each pool was measured in triplicates. Finally, the mean value of each pool was compared with the theoretical values using a regression equation.
2.8. Accuracy
The accuracy was evaluated using the WHO International standard (NIBSC code:92/680) for SAA, with a certified value of 156 mg/L, and the human serum (CRP)‐ERM®‐DA 474 IFCC, with a certified value of 41.1 mg/L. According to EP15‐A2, 13 the material was tested two times per day over five consecutive days. For the CRP assessment, accuracy was validated with the average value, standard deviation (SD), and 99th confidence interval. However, the accuracy assessment of SAA was calculated by percentage bias, as the uncertainty was not obtained.
2.9. Limits of blank, detection, and quantitation
The detection capabilities for CRP and SAA were estimated based on the CLSI document EP17‐A2. 14 To verify the limit of blank (LoB), blank samples were repeatedly tested 20 times per day for 3 days. The LoBs were calculated through nonparametric tests with following formula: LoB = PctB100‐α, (the value corresponding to the 95th percentile, α = 0.05).
The limit of detection (LoD) and limit of quantitation (LoQ) for CRP and SAA were determined using samples with five to seven different concentrations around the expected LoQ. These samples were diluted using the plasma with low concentrations and diluent solutions. Each pool was then measured four times per day for 5 days. The LoD value was calculated as follows: LoD = LoB + cβ * SDs, where cβ was the calibration factor for the value corresponding to the 95th percentile of the standard normal distribution, and SDs were defined as the estimates of the standard deviations of the low‐concentration samples.
The LoQ was defined as the lowest concentration that could be tested with a 20% imprecision 15 and was calculated by the imprecision profile curve. 16
2.10. Clinical reportable range
To verify the maximum dilution, two samples with high concentrations were diluted by two‐ to six fold within the linearity range. These serial dilutions were tested in triplicates, and the recovery rate was calculated with the measurements and the expected value. The recovery needed to meet an acceptable range (90%‐110%). Subsequently, the CRR was calculated by the maximum dilution, linearity (upper limit: 306 mg/L for CRP, and 316 mg/L for SAA), and LoQ.
2.11. Interference study
Common interfering substances, including conjugated bilirubin (CB, sigma), hemolysis (Hb, sigma), chyle (TG, sigma), and rheumatoid factor (RF, FRPON), were added into two plasma samples with different concentrations of CRP and SAA, respectively. Similarly, a diluent with the same volume as each interfering substance was added to the samples served as the control groups. All test and control groups were sufficiently mixed and measured in triplicate, and the percent biases relative to the control groups (paired‐difference testing) were calculated. A % bias ≤ 10% stated by the manufacturer was considered acceptable.
2.12. Method comparison
For CBC + DIFF comparison, XPEN60 was compared with the Sysmex–XN1000 (XN1000) automated hematology analyzer. For CRP comparison, the measurements obtained by XPEN60 were compared with those obtained by the Roche Cobas E601 (Roche, Basel, Switzerland, E601) and the Mindray CRP‐M100 (Mindray, Shenzhen, China, CRP‐M100) automated biochemical analyzers. For SAA comparison, the study was performed with a WeimiBio‐Tech AFS‐2000A (WeimiBio‐Tech, WBT).
3. RESULTS
3.1. Precision
In this study, no outliers were detected while the repeatability met the requirements for all evaluated analytical parameters (1.67% for WBC, 1.29% for RBC, 0.99% for HGB, 0.35% for MCV, and 3.35% for PLT). The total imprecision was within desirable specifications for the low‐, middle‐, and high‐level controls (see Table 1). The imprecision values of HGB counts with low and middle concentrations were equal to specifications, and the SD values were within the limit of validation (1.0 vs 3.0 g/L and 2.0 vs 3.0 g/L).
Table 1.
Total imprecision of CBC
| Total imprecision | Units | Low level | Medium level | High level | TEa% | |||
|---|---|---|---|---|---|---|---|---|
| Mean ± SD | CV% | Mean ± SD | CV% | Mean ± SD | CV% | |||
| WBC | ×109/L | 3.68 ± 0.18 | 4.89 | 8.31 ± 0.33 | 3.96 | 17.82 ± 0.41 | 2.30 | 6.00 |
| RBC | ×1012/L | 2.64 ± 0.06 | 2.27 | 4.78 ± 0.10 | 2.09 | 5.53 ± 0.09 | 1.63 | 2.50 |
| HGB | g/L | 57.8 ± 1.00 | 2.00 | 129.04 ± 2.00 | 2.00 | 164.68 ± 2.00 | 1.00 | 2.00 |
| MCV | fl | 88.06 ± 1.41 | 1.57 | 92.44 ± 1.45 | 1.60 | 92.21 ± 0.63 | 0.68 | 2.50 |
| PLT | ×109/L | 55.48 ± 4.00 | 7.00 | 269.16 ± 9.00 | 3.00 | 468.52 ± 13.00 | 3.00 | 8.00 |
Abbreviation: CV, coefficient of variation; HGB, hemoglobin; MCV, mean cellular volume; PLT, platelets; RBC, red blood cells; SD, standard deviation; WBC, white blood cells.
As shown in Table 2, the repeatability ranged from 4.63% to 5.37% for CRP and 5.37% to 7.14% for SAA, and the total imprecision varied from 5.39% to 6.36% for CRP and 5.54% to 10.87% for SAA.
Table 2.
Repeatability and total imprecision of CRP and SAA determinations on the XPEN60
| CRP | SAA | |||||
|---|---|---|---|---|---|---|
| Level1 | Level2 | Level3 | Level1 | Level2 | Level3 | |
| Repeatability | ||||||
| Mean value(mg/L) | 4.51 | 34.1 | 96.41 | 9.73 | 41.47 | 139.92 |
| SR(mg/L) | 0.22 | 1.83 | 4.46 | 0.68 | 2.23 | 9.99 |
| 95% lower limit (CV%) | 4 | 4.41 | 3.8 | 5.74 | 4.42 | 5.86 |
| 95% upper limit (CV%) | 6.24 | 6.87 | 5.92 | 8.94 | 6.89 | 9.14 |
| CV% | 4.84 | 5.37 | 4.63 | 7.03 | 5.37 | 7.14 |
| 1/4 TEa (CV%) | 14.15 | 9.28 | ||||
| Total imprecision | ||||||
| Mean value(mg/L) | 4.51 | 34.1 | 96.41 | 9.73 | 41.47 | 139.92 |
| SWL(mg/L) | 0.24 | 2.09 | 6.13 | 0.8 | 2.3 | 15.21 |
| 95% lower limit (CV%) | 4.94 | 5.54 | 5.43 | 7.33 | 5.12 | 8.54 |
| 95% upper limit (CV%) | 6.99 | 7.84 | 7.68 | 10.37 | 7.24 | 12.07 |
| CVWL% | 5.39 | 6.13 | 6.36 | 8.26 | 5.54 | 10.87 |
| 1/3 TEa (CV%) | 18.87 | 12.37 | ||||
95% verification limit determined as F×%CV (SR or SWL) according to the CLSI document EP5‐A3; SR, user estimate for repeatability; SWL, user estimate for total imprecision; TEa is from CLIA'88 deduced by Biological Variation.
3.2. Carryover
The carryover was 0.04% for WBC, 0.09% for RBC, 0.93% for PLT, 0.26% for CRP, and 0.36% for SAA. No carryover for hemoglobin has been found. There was no clinical significance, and all values fell within the manufacturer's limits of 3.0%, 2.0%, 2.0%, 4.0%, and ≤1%, respectively.
3.3. Linearity
All results, including WBC (113.35 × 109/L), RBC (10.43 × 1012/L), HGB (226.67 g/L), PLT (1208.67 × 109/L), CRP (208.03 mg/L), and SAA (243.13 mg/L), showed excellent linearity. The slopes closely equaled to 1, and determination coefficients (R 2) were all >.99 (see Figure 1).
Figure 1.
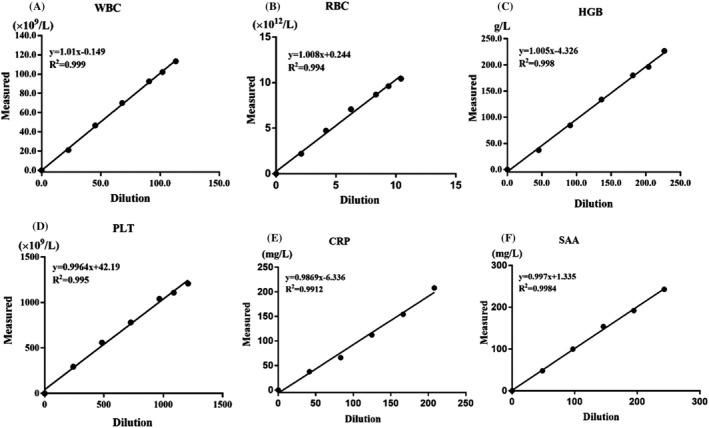
Linearity of complete blood cell count, CRP, and SAA observed with XPEN60. X‐axis represents the theoretical dilution concentration. Y‐axis represents the mean concentration of each sample
3.4. Accuracy
The certified value of 41.1 mg/L was within the 95% confidence interval (34.13‐44.45 mg/L). SAA was measured at 152.4 mg/L, and the calculated % bias was −2.31%, which is less than half of the permitted total error (8.24%) deduced from biological variation.
3.5. Detection capability
The observed LoBs were 0 and 1.1 mg/L for CRP and SAA. The LoD and LoQ values were 0.21 and 0.61 mg/L for CRP, and 2.27 and 3.62 mg/L for SAA, respectively (see Figure 2).
Figure 2.
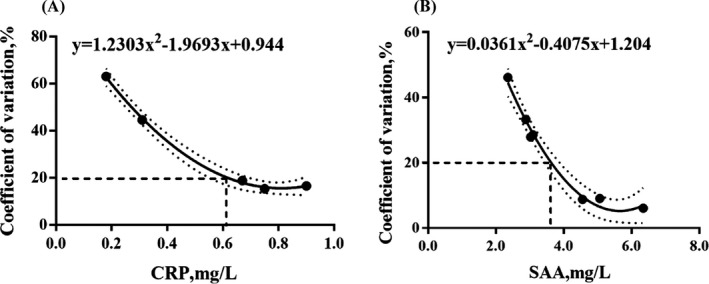
Assay imprecision profile, LoQ determination (with 20% imprecision); Each dot denotes the CV% of 20 measurements for a pool. The non‐linear relationship represents the imprecision (CV%, y‐axis) and concentration of the CRP and SAA, respectively (mg/L, x‐axis). Dashed lines around the curve showed predicted 95% confidence interval
3.6. Clinical reportable range
For both CRP and SAA, the maximum dilution ratio was within sixfold and the recovery rates ranged between 90% and 110%. The clinical reportable range of CRP and SAA was 0.61 ~ 1836 and 3.62 ~ 1896 mg/L, respectively (see Table S1).
3.7. Interference study
The % biases were within the expected bias range (±10%). Therefore, the interference substances did not significantly affect the measurements for CRP and SAA (see Table S2).
3.8. Method comparison
The agreement of XPEN60 with XN1000 was excellent for CBC and 5‐DIFF comparison, except for Eos and Bas. The observed correlation equations were y = 0.0834 + 0.848x and y = 0.08 + 0.800x (Figure 3 and Table 3). The Passing‐Bablok regression of correlation revealed very high consistency for CRP and SAA. The Pearson's correlation coefficient (r) ranged from .92 to 1.00 (P < .0001); however, a few outliers were observed for CRP and SAA. Bland‐Altman analysis showed negative mean % biases of −14.52% (−18.09% to −10.95%) and −8.86% (−2.11% to 19.84%) for CRP compared with E601 and CRP‐100, respectively, and a positive mean % bias of 19.33% (13.36% to 25.30%) for SAA compared with WBT (Figure 4 and Table 3).
Figure 3.
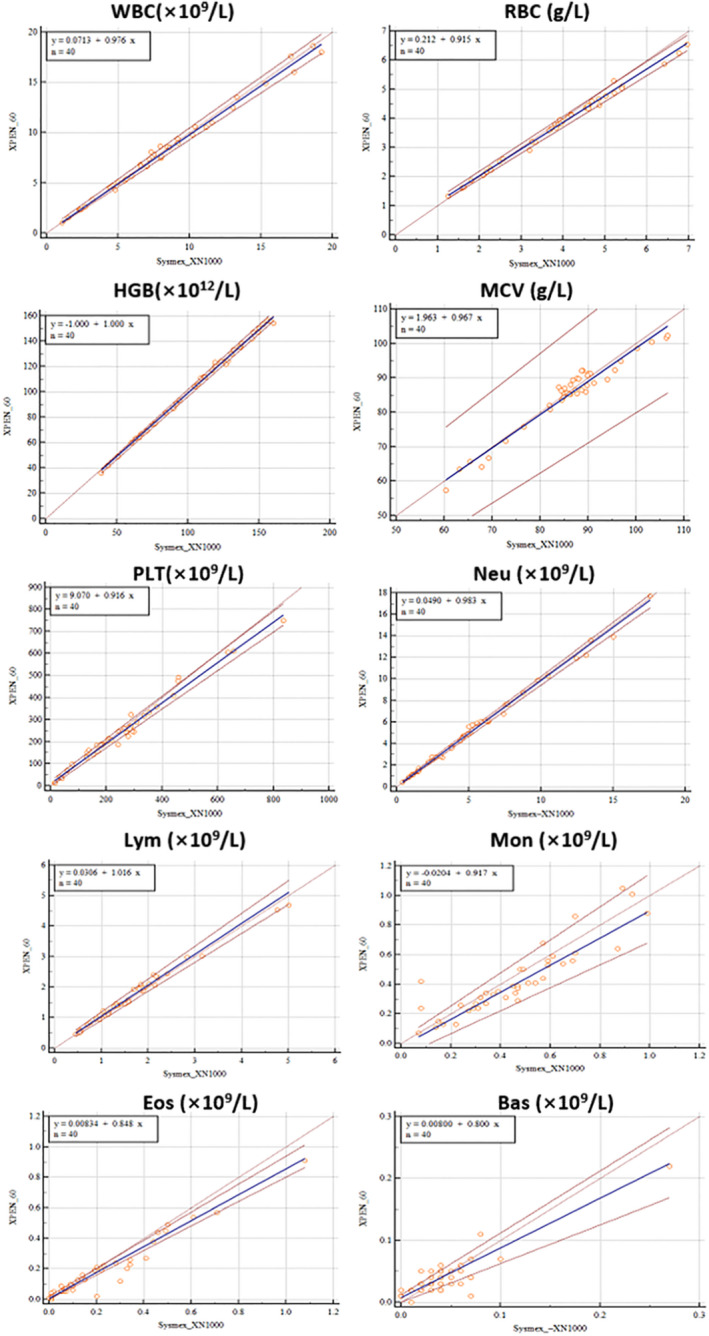
Comparison of the XPEN60 (ordinate) with the Symex XN‐1000 (abscissa) for 5‐Diff. The blue line shows the regression curve; the dashed brown lines indicate 95% confidence interval; the dashed red line shows the perfect correlation
Table 3.
Correlation of XPEN60 and Sysmex XN1000 results for CBC and 5‐Diff, and correlation for CRP and SAA
| Measurand | Unit | Range(median) | Bland‐Altman statistics | Passing and Bablok regression | |||
|---|---|---|---|---|---|---|---|
| Mean Diff(95% CI) | 95%LoA | Slope(95% CI) | Intercept(95% CI) | Correlation coefficient( R) | |||
| WBC | 109/L | 1.03 ~ 18.61 (7.37) | −0.10 (−0.23 to 0.04) | −0.92 to 0.73 | 0.98 (0.94 to 1.01) | 0.07 (−0.04 to 0.33) | 0.99 (0.99 to 1.00) |
| RBC | 1012/L | 1.33 ~ 6.54 (3.93) | −0.13 (−0.19 to −0.08) | −0.45 to 0.19 | 0.92 (0.89 to 0.94) | 0.21 (0.14 to 0.30) | 1.00 (0.99 to 1.00) |
| HGB | g/L | 36 ~ 156 (114.5) | −0.97 (−1.58 to −0.37) | −4.67 to 2.72 | 1.00 (1.00 to 1.02) | −1.00 (−3.01 to −1.00) | 1.00 (0.99 to 1.00) |
| MCV | fL | 57.4 ~ 102.3 (86.7) | −0.67 (−1.39 to 0.05) | −5.11 to 3.77 | 0.97 (0.88 to 1.08) | 1.96 (−7.19 to 9.23) | 0.92 (0.85 to 0.96) |
| PLT | 109/L | 13 ~ 749 (232) | −11.72 (−20.14 to −3.31) | −63.31 to 39.86 | 0.92 (0.88 to 0.96) | 9.07 (−3.35 to 19.97) | 1.00 (0.99 to 1.00) |
| Neu | 109/L | 0.39 ~ 17.7 (4.84) | 0.05 (−0.05 to 0.16) | −0.59 to 0.69 | 0.98 (0.95 to 1.01) | 0.05 (−0.03 to 0.19) | 1.00 (0.99 to 1.00) |
| Lym | 109/L | 0.46 ~ 4.68 (1.49) | −0.05 (−0.08 to −0.01) | −0.28 to 0.19 | 1.02 (0.95 to 1.08) | 0.03 (−0.04 to 0.11) | 0.99 (0.97 to 0.99) |
| Mon | 109/L | 0.07 ~ 1.05 (0.38) | 0.03 (−0.00 to 0.06) | −0.17 to 0.24 | 0.92 (0.78 to 1.12) | −0.02 (−0.09 to 0.03) | 0.92 (0.85 to 0.96) |
| Eos | 109/L | 0.00 ~ 0.91 (0.13) | 0.03 (0.01 to 0.05) | −0.08 to 0.15 | 0.85 (0.80 to 0.92) | 0.01 (0.00 to 0.02) | 0.94 (0.89 to 0.97) |
| Bas | 109/L | 0.00 ~ 0.22 (0.03) | −0.00 (−0.01 to 0.00) | −0.04 to 0.04 | 0.80 (0.62 to 1.00) | 0.01 (0.00 to 0.01) | 0.69 (0.49 to 0.83) |
| CRP(E601) | mg/L | 0.58 ~ 320 (98.64) | −14.52 (−18.09 to −10.95) | −36.39 to 7.35 | 0.92 (0.88 to 0.96) | −4.77 (−7.94 to −0.92) | 0.99 (0.98 to 1.00) |
| CRP(CRP‐100) | mg/L | 0.58 ~ 320 (98.64) | −8.86 (−2.11 to 19.84) | −58.39 to 76.12 | 0.92 (0.88 to 0.96) | −15.97 (−31.33 to −6.94) | 0.97 (0.94 to 0.98) |
| SAA | mg/L | 8.9 ~ 127.2 (37.65) | 19.33 (13.36 to 25.30) | −17.23 to 55.89 | 0.69 (0.60 to 0.78) | 1.55 (−2.14 to 4.84) | 0.93 (0.88 to 0.96) |
Bland‐Altman (mean difference and limits of agreement) statistics and Passing and Bablok regression (slope, intercept, and correlation coefficient) are shown.
Abbreviations: 95% CI, 95% confidence interval; Bas, basophils; Eos, eosinophils; LoA, upper and lower limits of agreement; Lym, lymphocytes; Mean diff., mean difference; Mon, monocytes; Neu, neutrophils.
Figure 4.
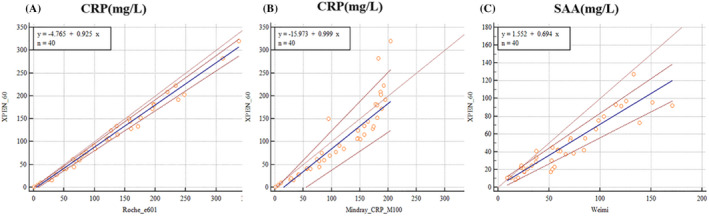
The comparison of XPEN60 with Roche E601 and Mindray CRP‐M100 in detecting for CRP, and with WeimiBio‐Tech AFS‐2000A for SAA. The blue line shows the regression curve; the dashed brown lines indicate 95% confidence interval; the dashed red line shows the perfect correlation
4. DISCUSSION
Currently, most instruments are capable of individually measuring CBC + 5‐DIFF, CRP, or SAA. XPEN60 introduces a medium‐sized automated hematology analyzer that could simultaneously measure the CBC with 5‐DIFF, CRP, and SAA. In this study, the performance characteristics of XPEN60 were evaluated according to CLSI protocols and ICSH guidelines.
XPEN60 showed acceptable imprecision and excellent linearity for CBC (Table 1 and Figure 1), which is comparable with other hematology analyzers. 17 , 18 The imprecision values for CRP and SAA were less than that of criteria derived from biological variation and the 95% verification limit determined (Table 2). No significant carryovers were observed for these parameters, and the carryovers of WBC, RBC, and HGB were even up to state‐of‐the‐art performance levels. 19 Although the carryover of PLT was slightly higher than the state‐of‐the‐art performance level (0.93% vs 0.5%), it was presumably limited by the concentrations of the samples (range: 21.7‐440 × 109/L). This range was dominated by the samples available, and clinical samples with both extremely low and high values were not able to obtain in this work. The observed LoBs and LoDs of CRP and SAA were less than those claimed by the manufacturers. In addition, the LoQs were established by ourselves in this study, which were not declared by the manufacturer. The LoQs were determined as 0.61 and 3.62 mg/L for CRP and SAA, which indicates high sensitivity for the detection of low‐level samples.
In this study, the accuracy of CBC was not evaluated; instead, comparisons were carried out with a routine hematology analyzer (reference method) that had previously been demonstrated to exhibit good performance. 20 , 21 The comparison with XN1000 revealed high consistency and accuracy of the new instrument (Table 3 and Figure 3); however, a poor slope and a few outliers in Eos and Bas comparison were observed (Figure 3). This phenomenon may be related to the different type of analyzers equipped (XPEN60: cytochemical staining analyzer, XN1000: fluorescence staining and flow cytometry analyzer 20 , 22 ), or, the extremely low values of Eos and Bas tested. The similar results were also found in other previous reports. 7 , 18 , 23
For CRP and SAA, the accuracy was evaluated using standard materials and adequately met the requirement of practical work. The results from XPEN60 showed a strong correlation with those from E601 and Mindray. It is worth noting that there was a systematic deviation. The results measured by XPEN60 were all lower in comparison with E601, with a % bias of −14.5%. This was probably referred to the different reference material used for calibration in each system. We found that ERM‐DA474 was used for XPEN60, while ERM‐DA470 was applied to E601. Therefore, it is easy to make a conclusion that the importance of traceability should be taken seriously to ensure interchangeability. In this work, a wide intercept of −15.97 and % bias of −8.86% were also observed on the comparison of CRP‐M100 and XPEN60. This may refer to the upper detection limit of the CRP‐M100 for CRP declared by the manufacturer (160 mg/L), which is far less than that of the XPEN60 (306 mg/L). Given this, it is suggested that laboratories should establish a reference interval for their individual instruments. For SAA comparison, there was a % bias of 19.33% compared with the WBT. This was originated to the different detection principle between two platforms (immunofluorescence principle vs latex‐enhanced scatter immunoturbidimetric technologies). Therefore, the sample from one patient should avoid to be tested separately on two different systems in the same laboratory.
There are several limitations in current study: A strong agreement in a broad concentration range was shown, but the sample size of the comparison study was insufficient. In addition, although the accuracy of the SAA analysis was verified using the standard material, further evaluation of the comparability of SAA with a reference method would be useful. Another limitation of the study is that some of the items, including morphology flags and manual differential results, were not performed, so these parameters could be the subjects of future research.
In conclusion, this study demonstrated that XPEN60 delivers satisfactory analytical performance for CBC, CRP, and SAA determination. XPEN60 is a convenient laboratory analyzer that can simultaneously measure CRP, SAA, and CBC, including 5‐DIFF. Especially, it is suitable for patients with inflammatory diseases at emergency departments and community hospitals.
CONFLICT OF INTEREST
The authors state that there are no financial, personal, or professional conflicts of interests that may hinder this work.
Supporting information
Table S1‐S2
Zong Z, Kan L, Luo Y, et al. Performance evaluation of the XPEN60 CRP&SAA, a novel automated hematology analyzer, in detecting complete blood count, C‐reactive protein, and serum amyloid A. J Clin Lab Anal. 2020;34:e23395 10.1002/jcla.23395
Zengyan Zong and Lijuan Kan contributed equally to this paper.
Funding information
This study was supported by The National Natural Science Foundation of China (Grant NO. 81772921, 81502344). A grant from The Science and Technology Planning Project of Shenzhen City of China (NO. JCYJ20180306172209668). The discipline construction ability promotion project of Shenzhen health and population family planning commission (NO. SZXJ2017018, SZXJ2018031). Sanming Project of Medicine in Shenzhen (NO. SZSM201601062).
Contributor Information
Dan Xiong, Email: sunny543@126.com.
Xiuming Zhang, Email: zhangxiuming0760@163.com.
REFERENCES
- 1. Berger C, Uehlinger J, Ghelfi D, Blau N, Fanconi S. Comparison of C‐reactive protein and white blood cell count with differential in neonates at risk for septicaemia. Eur J Pediatr. 1995;154(2):138‐144. [DOI] [PubMed] [Google Scholar]
- 2. Zhang Y, Zhang J, Sheng H, Li H, Wang R. Acute phase reactant serum amyloid A in inflammation and other diseases. Adv Clin Chem. 2019;90:25‐80. [DOI] [PubMed] [Google Scholar]
- 3. Jovanovic DB. Clinical importance of determination of serum amyloid A. Srp Arh Celok Lek. 2004;132(7–8):267‐271. [DOI] [PubMed] [Google Scholar]
- 4. Ahout IML, Brand KH, Zomer A, et al. Prospective observational study in two Dutch hospitals to assess the performance of inflammatory plasma markers to determine disease severity of viral respiratory tract infections in children. BMJ Open. 2017;7(6):e014596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Liu J, Li S, Liu J, et al.Longitudinal characteristics of lymphocyte responses and cytokine profiles in the peripheral blood of SARS‐CoV‐2 infected patients. medRxiv. 2020:2020.2002.2016.20023671. [DOI] [PMC free article] [PubMed]
- 6. Inaba T, Nomura N, Ishizuka K, et al. Basic evaluation of Pentra MS CRP, a new automated hematology analyzer for rapid 5‐part WBC differential and CRP using a small volume of whole blood. Int J Lab Hematol. 2015;37(2):208‐216. [DOI] [PubMed] [Google Scholar]
- 7. Nomura N, Saito K, Ikeda M, et al. Evaluation of the Microsemi CRP, an automated hematology analyzer for rapid 3‐part WBC differential and CRP using whole blood. Int J Lab Hematol. 2015;37(4):466‐473. [DOI] [PubMed] [Google Scholar]
- 8. Barnes PW, McFadden SL, Machin SJ, Simson E, International Consensus Group for h . The international consensus group for hematology review: suggested criteria for action following automated CBC and WBC differential analysis. Lab Hematol. 2005;11(2):83‐90. [DOI] [PubMed] [Google Scholar]
- 9. International Council for Standardization in Haematology WG , Briggs C, Culp N, et al. ICSH guidelines for the evaluation of blood cell analysers including those used for differential leucocyte and reticulocyte counting. Int J Lab Hematol. 2014;36(6):613‐627. [DOI] [PubMed] [Google Scholar]
- 10. CLSI . Method Comparison and Bias Estim Ation Usingpatient Samples: Approved Guideline Seeond Edition. In. CLSI Document EP9‐A3. Wayne, PA: CLSI; 2002. [Google Scholar]
- 11. CLSI . Evaluation of Precision of Quantitative Measurement Procedures; Approved Guideline—Third Edition. In. CLSI document EP5‐A3. Wayne, PA: Clinical and Laboratory Standards Institute; 2014. [Google Scholar]
- 12. CLSI . Validation, Verification, and Quality Assurance of Automated Hematology Analyzers. In. CLSI document H26‐A2. A. Rabinovitch. Wayne, PA: Clinical and Laboratory Standards Institute; 2010. [Google Scholar]
- 13. CLSI . User verification of precision and estimation of bias; Appruved guideline‐Third Edition. In. CLSI document EP15‐A2. Wayne. PA: Clinical and Laboratory Standards Institute; 2014. [Google Scholar]
- 14. CLSI . Protocols for Determination of Limits of Detection and Limits of Quantitation. Approved Guideline‐Second Edition. In. CLSI document EP17‐A2. Wayne PA: Clinical and Laboratory Standards Institute; 2014. [Google Scholar]
- 15. Armbruster DA, Pry T. Limit of blank, limit of detection and limit of quantitation. Clin Biochem Rev. 2008;29(Suppl 1):S49‐52. [PMC free article] [PubMed] [Google Scholar]
- 16. Ceriotti F, Marino I, Motta A, Carobene A. Analytical evaluation of the performances of Diazyme and BRAHMS procalcitonin applied to Roche Cobas in comparison with BRAHMS PCT‐sensitive Kryptor. Clin Chem Lab Med. 2017;56(1):162‐169. [DOI] [PubMed] [Google Scholar]
- 17. Schapkaitz E, Raburabu S. Performance evaluation of the new measurement channels on the automated Sysmex XN‐9000 hematology analyzer. Clin Biochem. 2018;53:132‐138. [DOI] [PubMed] [Google Scholar]
- 18. Slim CL, Wevers BA, Demmers M, et al. Multicenter performance evaluation of the Abbott Alinity hq hematology analyzer. Clin Chem Lab Med. 2019;57(12):1988‐1998. [DOI] [PubMed] [Google Scholar]
- 19. Vis JY, Huisman A. Verification and quality control of routine hematology analyzers. Int J Lab Hematol. 2016;38(Suppl 1):100‐109. [DOI] [PubMed] [Google Scholar]
- 20. Aguadero V, Cano‐Corres R, Berlanga E, Torra M. Evaluation of biological fluid analysis using the sysmex XN automatic hematology analyzer. Cytometry B Clin Cytom. 2018;94(5):680‐688. [DOI] [PubMed] [Google Scholar]
- 21. Becker PH, Fenneteau O, Da Costa L. Performance evaluation of the Sysmex XN‐1000 hematology analyzer in assessment of the white blood cell count differential in pediatric specimens. Int J Lab Hematol. 2016;38(1):54‐63. [DOI] [PubMed] [Google Scholar]
- 22. Amundsen EK, Henriksson CE, Holthe MR, Urdal P. Is the blood basophil count sufficiently precise, accurate, and specific?: three automated hematology instruments and flow cytometry compared. Am J Clin Pathol. 2012;137(1):86‐92. [DOI] [PubMed] [Google Scholar]
- 23. Siekmeier R, Bierlich A, Jaross W. The white blood cell differential: three methods compared. Clin Chem Lab Med. 2001;39(5):432‐445. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Table S1‐S2


