Abstract
Considering the issues of shortage of medical resources and the invasiveness and infection risk involved in the collection of nasopharyngeal swab specimens, there is a need for an effective alternative test specimen for SARS-CoV-2 RNA detection. Here, we investigated suitability of saliva as a non-invasively obtained specimen for molecular detection of SARS-CoV-2 RNA in Japanese patients with COVID-19. In total, 28 paired clinical specimens of saliva and nasopharyngeal swabs were collected from 12 patients at various time points after symptom onset. Each specimen was assayed using reverse transcription real-time polymerase chain reaction (rRT-PCR) on the BD MAX open system using primers and probes targeting the N-gene. The saliva and nasopharyngeal swab specimens showed 19 and 15 positive results, respectively. No invalid (PCR inhibition) result was observed for any specimen. The qualitative results of each specimen obtained in the period immediately after symptom onset were similar. Three convalescent patients presented saliva-positive results, whereas their nasopharyngeal swabs were negative at four different time points, suggesting that saliva may be superior to nasopharyngeal swabs in terms of obtaining stable assay result of SARS-CoV-2. In conclusion, our results suggest that saliva can potentially serve as an alternative to nasopharyngeal swabs as a specimen for SARS-CoV-2 rRT-PCR. As saliva can be collected by patients themselves, it may be an effective way to overcome the shortage of personal protective equipment and specimen sampling tools.
Keywords: SARS-CoV-2, Saliva, Nasopharyngeal swab, Reverse transcription real-time PCR
The recent rapid and wide spread of coronavirus disease (COVID-19) caused by SARS-CoV-2 has become a national concern in Japan [1]. Rapid and accurate diagnosis of COVID-19 is important to prevent spread of infections. For SARS-CoV-2 detection assays, the Japanese National Institute of Infectious Diseases recommends collecting sputum (first priority) [2]. However, as dry cough (reduced sputum) is common in COVID-19, nasopharyngeal specimens (secondary priority) are also collected widely. Unfortunately, nasopharyngeal sample collection has disadvantages such as high invasiveness, infection risk to health workers, and a need for technical skill (as an inappropriate procedure may lead to false-negative test results). Moreover, there is a shortage of medical resources such as personal protective equipment (PPE), sterile swabs, and virus transportation medium (VTM) in the country. Therefore, we considered it essential to find a new suitable test specimen as a substitute for nasopharyngeal swabs. Currently, saliva is attracting considerable interest as a specimen for SARS-CoV-2 detection [3]. Therefore, the present study aimed to compare nasopharyngeal swab and saliva specimens for the molecular detection of SARS-CoV-2 RNA in Japanese patients with COVID-19.
This study was approved by the Human and Animal Ethics Review Committee of Aichi Medical University Hospital, Nagakute, Japan (approval number 2020-027).
In total, 28 paired nasopharyngeal swab and saliva clinical specimens were collected from 12 patients at various time points after symptom onset, during the month of April 2020 at Aichi Medical University Hospital. Among them, five patients had been diagnosed with COVID-19 by reverse transcription real-time polymerase chain reaction (rRT-PCR) of nasopharyngeal swabs and were hospitalized before the first collection of paired specimens, whereas seven were outpatients suspected to have COVID-19 based on their clinical symptoms. All patients showed mild to moderate symptoms (i.e., did not require admission to ICU or artificial ventilation) at the time of the first rRT-PCR assay. After collection, the nasopharyngeal swabs were immediately placed in sterile tubes containing 3 mL of VTM. Saliva was collected in a sterile 50 mL tube using the passive drool method [4]. Then, 0.5 mL of saliva was re-suspended in a sterile 15 mL tube containing 3 mL of PBS. The re-suspended saliva was centrifuged at 500×g for 1 min, and the supernatant fluid was used for the assay.
Specimen volumes of 750 μL were assayed on the BD MAX open system (a fully-integrated, automated platform that performs nucleic acid extraction and real-time PCR) (Japan Becton Dickinson and Company, Japan), using the following reagents: BD MAX ExK TNA-3 (Swabs), BD MAX TNA MMK (SPC; containing Sample Processing Control to monitor PCR inhibition), and BD MAX PCR cartridges. The N and N2 primer-probe sets were used for rRT-PCR [5] and the cycling conditions were as follows: 95 °C for 5 min (activation), 60 °C for 30 min (reverse transcription), 95 °C for 1 min (denaturation), and 45 cycles of 95 °C for 5 s and 56 °C for 17.1 s (PCR). The fluorescence gains and thresholds were set at 50 and 50 for the targets (N and N2) and at 50 and 100 for SPC, respectively. The rRT-PCR results were interpreted according to the following criteria: when the fluorescence signal of SARS-CoV-2 was detected, the result was interpreted as “positive.” When the fluorescence signal of SARS-CoV-2 was not detected, the result was interpreted as “negative”; however, if SPC fluorescence signal was not detected, the result was interpreted as “invalid.” Qualitative results (number of positive, negative, or invalid) and threshold cycle (Ct) values on the days after symptom onset were compared between nasopharyngeal swab and saliva specimens.
The results obtained for the twelve test patients, including the seven patients with a confirmed diagnosis of COVID-19 (twenty-three paired specimens collected) and five non-COVID-19 patients (five paired specimens collected) are shown in Table 1 . No invalid result was obtained with any specimen. Fig. 1, Fig. 2 show the results of rRT-PCR and Ct values at each time point of specimen collection after symptom onset in the seven patients with COVID-19, respectively. The qualitative results of each specimen obtained in the period immediately after symptom onset were similar (refer to patient numbers 2–7 in the figures). As for the Ct values, the superior (lower Ct value) specimen differed based on the patient. Saliva presented lower Ct values in patients 3 and 4, whereas nasopharyngeal swabs presented lower Ct values in patients 5–7. Three convalescent patients (numbers 2–4) demonstrated positive results with the saliva at four different time points, whereas their nasopharyngeal swabs collected on the same day yielded a negative result.
Table 1.
Results of SARS-CoV-2 reverse transcription real-time polymerase chain reaction for each specimen.
| Specimens | Number of results |
||
|---|---|---|---|
| Positive | Negative | Invalid | |
| Nasopharyngeal swab | 15 | 13 | 0 |
| Saliva | 19 | 9 | 0 |
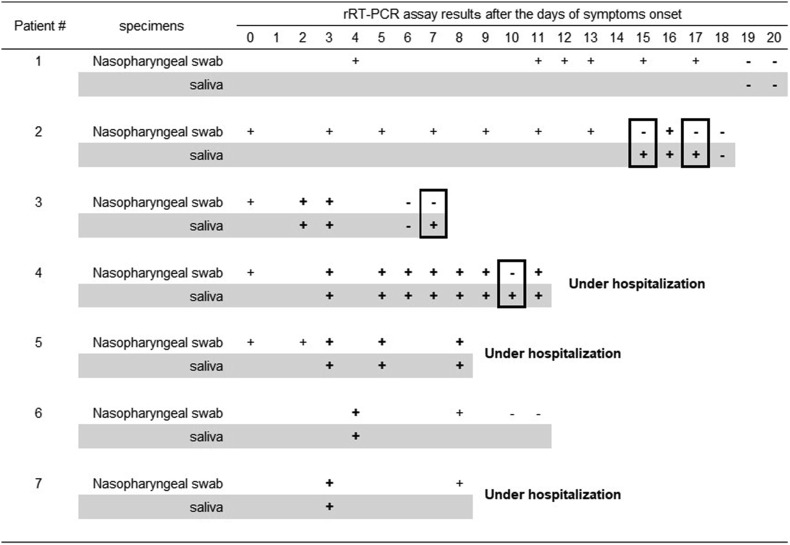
Fig. 1.
Comparison between SARS-CoV-2 detection from nasopharyngeal swab and saliva specimens using reverse transcription real-time polymerase chain reaction (rRT-PCR) at various time points after symptom onset in seven cases of COVID-19. “+” and “-” indicate “positive” and “negative”, respectively. Frames around indicates the difference in results between nasopharyngeal swab and saliva.
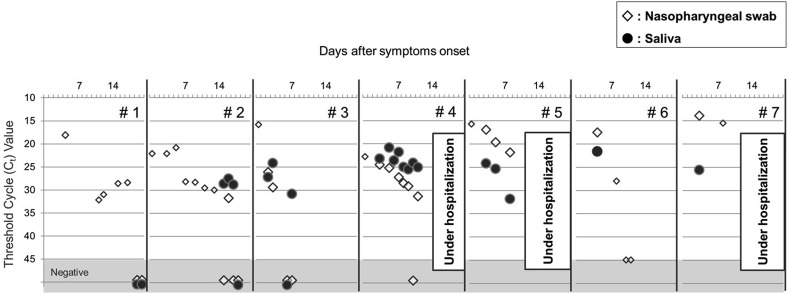
Fig. 2.
Comparison between threshold cycle values from nasopharyngeal swab and saliva specimens in SARS-CoV-2 reverse transcription real-time polymerase chain reaction (rRT-PCR) at various time points after symptom onset in seven cases of COVID-19. Small scale of diagram indicates that only nasopharyngeal swab was assayed.
The optimal specimen for SARS-CoV-2 detection by rRT-PCR thus requires further investigation. A previous study reported that nasopharyngeal swab specimens showed higher sensitivity than oropharyngeal swabs for SARS-CoV-2 rRT-PCR [6]. However, the infection risk at the time of specimen collection remains an issue. Saliva samples can be collected by patients themselves in a noninvasive manner. In the present study, we compared the potential for SARS-CoV-2 detection between paired nasopharyngeal swab and saliva specimens. Our results demonstrate that saliva specimens show higher sensitivity than nasopharyngeal swabs and support the findings of previous studies [3,7,8]. In COVID-19 cases, the assay results from both specimen types in the period immediately after symptom onset were in accordance with each other. Therefore, we propose that saliva may be used instead of nasopharyngeal swabs in the first diagnosis to identify a patient with COVID-19. Several previous studies reported that nasopharyngeal swab specimens showed higher sensitivity than saliva for SARS-CoV-2 rRT-PCR in the convalescence period [[9], [10], [11]]. However, in this study, saliva-positive and nasopharyngeal swab-negative cases were observed in three convalescent patients at four time points. Although several previous studies used the spitting method to collect saliva, we used the passive drool method to obtain a homogeneous specimen and to avoid the influence of inhibitory substances [9,11,12]. Our results indicate that saliva collected by the passive drool method may be superior to nasopharyngeal swabs to obtain stable assay results. Nonetheless, further evaluation is necessary to determine the most suitable saliva collection method.
In conclusion, our results suggest that saliva has potential as an alternative to nasopharyngeal swabs as a specimen for SARS-CoV-2 rRT-PCR and may present an effective approach to overcome the shortage of PPE and specimen sampling tools.
Author contribution
Contributor Mikamo H was responsible for the organization and coordination of the trial. Sakanashi D was the chief investigator and responsible for the data analysis. Hagihara M, Shiota A and Ohashi W developed the trial design. Mikamo H, Yamagishi Y, Asai N, Kurumiya A, Sakata M, Kato S, Muramatsu Y, Koizumi Y, and Kishino T contributed to the specimen collecting and transportation. Nakamura A, Miyazaki N, Kawamoto Y, Ohno T, Yamada A, Koita I, and Suematsu H contributed to perform rRT-PCR. Yamagishi Y contributed to obtain the ethical approval. All authors contributed to the writing of the final manuscript. All members of the Department of Infection Control and Prevention, and Department of Clinical Infectious Diseases contributed to the management or administration of the trial. All authors approved the manuscript to be published, and agree to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Funding
None.
Declaration of competing interest
H. Mikamo has received grant support from Asahi Kasei Pharma Corporation, Shionogi & Co. Ltd., Daiichi Sankyo Co., Ltd., Sumitomo Dainippon Pharma Co., Ltd., Pfizer Japan Inc. and FUJIFILM Toyama Chemical Co., Ltd., payment for lectures from Astellas Pharma Inc., MSD K.K., Daiichi Sankyo Co., Ltd., Sumitomo Dainippon Pharma Co., Ltd., MIYARISAN Pharmaceutical Co., Ltd. Becton, Dickinson and Company Japan, and FUJIFILM Toyama Chemical Co. Ltd.
The other authors declare that they have no conflicts of interest.
Acknowledgements
The authors would like to thank National Institute of Infectious Diseases Japan that assisted with provision of the primer-probe sets. We would like to thank Editage [http://www.editage.com] for editing and reviewing this manuscript for English language.
Footnotes
All authors meet the ICMJE authorship criteria.
References
- 1.Ministry of Health, Labour and Welfare Japan About coronavirus disease. 2019. https://www.mhlw.go.jp/stf/seisakunitsuite/bunya/newpage_00032.html
- 2.National Institute of Infectious Diseases Japan nCoV (new coronavirus) Sample collection and transportation manual. 2019. https://www.niid.go.jp/niid/images/pathol/pdf/2019-nCoV_200416.pdf.Japanese ver. 16 April 2020.
- 3.Khurshid Z., Asiri F.Y.I., Al Wadaani H. Human saliva: non-invasive fluid for detecting novel coronavirus (2019-nCoV) Int J Environ Res Publ Health. 2020;17 doi: 10.3390/ijerph17072225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Golatowski C., Salazar M.G., Dhople V.M., Hammer E., Kocher T., Jehmlich N., et al. Comparative evaluation of saliva collection methods for proteome analysis. Clin Chim Acta. 2013;419:42–46. doi: 10.1016/j.cca.2013.01.013. [DOI] [PubMed] [Google Scholar]
- 5.Shirato K., Nao N., Katano H., Takayama I., Saito S., Kato F., et al. Development of genetic diagnostic methods for novel coronavirus 2019 (nCoV-2019) in Japan. Jpn J Infect Dis. 2020 doi: 10.7883/yoken.JJID.2020.061. [DOI] [PubMed] [Google Scholar]
- 6.Zou L., Ruan F., Huang M., Liang L., Huang H., Hong Z., et al. SARS-CoV-2 viral load in upper respiratory specimens of infected patients. N Engl J Med. 2020;382:1177–1179. doi: 10.1056/NEJMc2001737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Azzi L., Carcano G., Gianfagna F., Grossi P., Gasperina D.D., Genoni A., et al. Saliva is a reliable tool to detect SARS-CoV-2. J Infect. 2020;S0163–4453(20):30213–30219. doi: 10.1016/j.jinf.2020.04.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.To K.K., Tsang O.T., Chik-Yan Yip C., Chan K.H., Wu T.C., Chan J.M.C., et al. Consistent detection of 2019 novel coronavirus in saliva. Clin Infect Dis. 2020;12 doi: 10.1093/cid/ciaa149. ciaa149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Iwasaki S., Fujisawa S., Nakakubo S., Kamada K., Yamashita Y., Fukumoto T., et al. Comparison of SARS-CoV-2 detection in nasopharyngeal swab and saliva. J Infect. 2020;81(2):e145–e147. doi: 10.1016/j.jinf.2020.05.071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Williams E., Bond K., Zhang B., Putland M., Williamson D.A. Saliva as a non-invasive specimen for detection of SARS-CoV-2. J Clin Microbiol. 2020 doi: 10.1128/JCM.00776-20. JCM.00776-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Jamal A.J., Mohammad M., Coomes E., Powis J., Li A., Paterson A., et al. Sensitivity of nasopharyngeal swabs and saliva for the detection of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) medRxiv. 2020 doi: 10.1101/2020.05.01.20081026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Bellagambi F.G., Lomonaco T., Salvo P., Vivaldi F., Hangouët M., Ghimenti S., et al. Saliva sampling: methods and devices. An overview. Trends Anal Chem. 2020;124:115781. doi: 10.1016/j.trac.2019.115781. [DOI] [Google Scholar]