Abstract
Background
Anaplasma and Ehrlichia species are tick-borne pathogens of both veterinary and public health importance. The current status of these pathogens, including emerging species such as Ehrlichia minasensis and Anaplasma platys, infecting cattle in Kenya, remain unclear, mainly because of limitation in the diagnostic techniques. Therefore, we investigated the Anaplasma and Ehrlichia species infecting dairy cattle in Nairobi, Kenya using molecular methods.
Results
A total of 306 whole blood samples were collected from apparently healthy dairy cattle. Whole blood DNA was extracted and tested for presence of Anaplasma and Ehrlichia DNA through amplification and sequencing of the 16S rDNA gene. Sequence identity was confirmed using BLASTn analysis while phylogenetic reconstruction was performed to determine the genetic relationship between the Kenyan isolates and other annotated genotypes available in GenBank. Anaplasma and Ehrlichia species were detected in 19.9 and 3.3% of all the samples analyzed, respectively. BLASTn analysis of the sequences against non-redundant GenBank nucleotide database revealed infections with A. platys (44.8%), A. marginale (31%) and A. bovis (13.8%). All four sequenced Ehrlichia spp. were similar to Ehrlichia minasensis. Nucleotide polymorphism was observed for A. platys, A. bovis and E. minasensis. The Anaplasma species clustered in four distinct phylogenetic clades including A. marginale, A. platys, A. bovis and some unidentified Anaplasma spp. The Kenyan Ehrlichia minasensis clustered in the same clade with isolates from America and Australia but distant from E. ruminantium.
Conclusion
This study provides the first report of infection of dairy cattle in Kenya with A. platys and E. minasensis, which are emerging pathogens. We conclude that cattle in peri-urban Nairobi are infected with various species of Anaplasma and E. minasensis. To understand the extent of these infections in other parts of the country, large-scale screening studies as well as vector identification is necessary to inform strategic control.
Keywords: Tick-borne pathogens, Anaplasma platys, Ehrlichia minasensis, 16S rDNA, Phylogenetics, Molecular diagnostics, Ticks
Background
Anaplasma and Ehrlichia species of the family Anaplasmataceae are tick-borne pathogens of livestock with some species known to infect humans and therefore are of both veterinary and public health importance [1]. Anaplasma species documented to infect domestic ruminants including cattle are Anaplasma marginale (A. maginale), A. centrale, A. ovis, A. bovis, A. phagocytophilum and more recently A. platys [2, 3]. Anaplasma marginale transmitted by several tick vectors including some Rhipicephalus (boophilus) species is the most common in Kenya causing a severe hemolytic disease in the ruminants. Anaplasma centrale whose only known vector is Rhipicephalus simus [4] causes mild anaplasmosis in cattle [5] but has not been reported in Kenya possibly because the tick vector is absent. Anaplasma bovis transmitted by various species of Amblyomma and Rhipicephalus ticks [6] causes sub-clinical disease in cattle and has been recently been reported in indigenous calves in Kenya [7].
Anaplasma platys transmitted by Rhipicephalus sanguineus typically infects dogs resulting in canine infectious cyclic thrombocytopenia [8]. However, A. platys has also been isolated from cattle neutrophils [9]. To date, there is no information on cattle infection with this pathogen in Kenya. Anaplasma phagocytophilum is a zoonotic species that has been documented to infect cattle resulting in fever, respiratory signs, reduced milk production and infertility [10]. Anaplasma phagocytophilum was detected recently in questing ticks in a National park in Kenya [11]. For Ehrlichia species, E. ruminantium transmitted by ticks in the genus Amblyomma and the emerging Ehrlichia minasensis (E. minasensis) are the only species in that genera known to infect cattle [12, 13]. Infections of cattle with E. ruminantium and E. minasensis are mainly characterized by severe fever, anemia, thrombocytopenia and enlarged lymph nodes [14].
Infections of cattle with the pathogenic species of Anaplasma and Ehrlichia cause mortalities and morbidities with subsequent losses in production of dairy and beef products. The losses usually result in marked economic impact to dairy and beef farmers in the tropical and sub-tropical regions [15] contributing to poverty in the affected households.
In Kenya including the peri-urban Nairobi, anaplasmosis and theileriosis are some of the tick-borne diseases (TBDs) known to cause economic losses to dairy farmers [16, 17]. Although ehrlichiosis caused by Ehrlichia ruminantium has been reported in Kenya [7] and approximately 150 million animals are at risk of infection in Africa [18], quantification of its economic impact in Kenya has not been evaluated possibly because of the difficulty in confirming the diagnosis and the fact that this infection commonly co-infects with other TBDs such as East Coast Fever and anaplasmosis [7]. For a long time, the identification of cattle infected with pathogenic Anaplasma and Ehrlichia in Kenya has been based mainly on questionnaire data, clinical signs, microscopic examination [16, 19] and serological tests [20, 21]. Nevertheless, few studies in Kenya have used molecular techniques such as reverse line blot, polymerase chain reaction and sequencing for the confirmation and characterization of the tick-borne pathogens infecting cattle [7, 22]. Previously, there had been clinical cases reported to the University of Nairobi Veterinary Hospital, Kenya presenting with unspecific clinical signs such as unthriftness and loss of body conditions. On screening of cattle from the areas where the clinical cases had originated, E. ruminantium was identified by antigen detection using Enzyme-linked Immunosorbent Assay (ELISA) [23]. Additionally, microscopic examination of blood from these cattle revealed that some of them had Ehrlichia-like inclusion bodies in the white blood cells suggesting infections with other unknown potentially pathogenic haemoparasites [23]. Hence, there was need to identify and characterize those Ehrlichia-like inclusion bodies observed in these animals.
Therefore, to further characterize the Anaplasma and Ehrlichia species infecting dairy cattle in Kenya, this study used molecular tools to determine the prevalence and genetic profiles of various species circulating in apparently healthy dairy cattle in peri-urban Nairobi. Sequence identities and phylogenetic analysis were used to determine the presence of emerging pathogens such as A. platys and E. minasensis. Subsequently, we provide information that will aid in the understanding of molecular epidemiology of these emerging pathogens of livestock in Kenya and possible epidemiological factors contributing to their spread.
Results
Prevalence of Anaplasma and Ehrlichia species
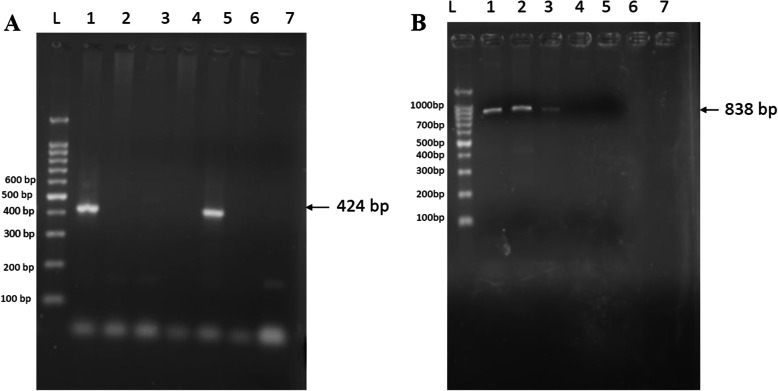
Of the 306 blood DNA samples analyzed, 61 (19.9%) [95% CI 15.6–24.9] were PCR positive for Anaplasma species while 10 (3.3%) [95% CI 1.6–5.9]) were positive for Ehrlichia. The Anaplasma species yielded a specific band corresponding to 424 bp (Fig. 1a) while primers targeting the 16S rDNA gene of Ehrlichia species produced a specific band corresponding to the expected size of 838 bp (Fig. 1b). The distribution of the positive samples in different sub-counties is shown in Table 1. The highest numbers of both Anaplasma 34 (55.7) and Ehrlichia 7(70) infections were found in Kasarani-Ruai Sub-County while Lang’ata had the least number of cattle positive for Anaplasma 6(9.8) infection. Ehrlichia infections were however not detected in cattle in Dagorreti Sub-County.
Fig. 1.
Some of the PCR amplicons of Anaplasma and Ehrlichia 16S rDNA gene a) PCR product of Anaplasma species. Lane L: molecular ladder, lane lanes 1 and 5: positive samples showing amplicon at approximate 424 bp, lanes 2, 3, 4 and 6: no amplicons were observed, 7: negative control. b. PCR product of Ehrlichia species. Lane L: molecular ladder lane, lanes 1, 2 and 3: positive samples showing amplicon band at approximate 838 bp, lanes 4, 5 and 6: no amplicons were observed, 7: negative control
Table 1.
Distribution of Anaplasma and Ehrlichia spp. positive cattle in the four sub-counties of Nairobi County
| Sub-County | No. of Anaplasma spp. (%) [95% CI] | No. of Ehrlichia spp. (%) [95% CI] |
|---|---|---|
| Kasarani-Ruai | 34 (55.7) [42.4–68.5] | 7 (70.0) [34.8–83.3] |
| Westlands | 11 (18.0) [9.4–29.9] | 1 (10.0) [0.2–44.2] |
| Langata | 6 (9.8) [3.6–20.2] | 2 (20.0) [2.0–55.6] |
| Dagorreti | 10 (16.4) [8.2–28.0] | 0 (0.0) [0] |
| Total | 61 (100) | 10 (100) |
Sequence identities of the Anaplasma and Ehrlichia species detected
Twenty-nine PCR amplicons for Anaplasma and four for Ehrlichia were sequenced for confirmation of the identities of the detected pathogens. BLASTn analysis revealed that majority, 13(44.8%) of the Anaplasma 16S rDNA sequences were similar to A. platys with sequence identity of between 98.72 and 100% to annotated sequences in Genbank. Nine (31%) of the sequences were similar to A. marginale with a sequence identity of between 99.07 and 100%. Other sequences matched A. bovis 4(13.8%) with sequence identity of between 99.28 and 100% and unidentified Anaplasma species 3(10.3%) sequence identity of 97.85 to 100% (Table 2). All the four Ehrlichia sequences were similar to those of E. minasensis revealing a sequence identity of between 99.42 and 100% (Table 3).
Table 2.
Anaplasma species detected by BLASTn analysis of 16S rDNA gene sequences of the Kenyan isolates
| Isolate | Our accession number | Matching sequence | Accession no. of highest match | E-value | % Identity |
|---|---|---|---|---|---|
| 20 | MT163376 | A. platys | MN630836.1 | 0.0 | 100.00 |
| 46 | MT163377 | A. platys | MK408655.1 | 0.0 | 99.28 |
| 79 | MT163378 | A. platys | MN630836.1 | 0.0 | 100.00 |
| 85 | MT163379 | A. platys | MN630835.1 | 0.0 | 99.73 |
| 97 | MT163380 | A. platys | MN401150.1 | 0.0 | 99.76 |
| 100 | MT163381 | A. platys | MK408655.1 | 0.0 | 99.77 |
| 117 | MT163382 | A. platys | MN630836.1 | 0.0 | 100.00 |
| 173 | MT163387 | A. platys | MN630836.1 | 6e-154 | 98.72 |
| 175 | MT163388 | A. platys | MN401150.1 | 0.0 | 99.51 |
| 268 | MT163383 | A. platys | MN401150.1 | 0.0 | 100.00 |
| 318 | MT163384 | A. platys | MN159065.1 | 0.0 | 100.00 |
| 381 | MT163385 | A. platys | MN630836.1 | 0.0 | 100.00 |
| 425 | MT163386 | A. platys | MN861060.1 | 0.0 | 99.76 |
| 127 | MT163438 | A. marginale | MK310488.1 | 0.0 | 99.76 |
| 139 | MT163439 | A. marginale | MK310488.1 | 0.0 | 100.00 |
| 159 | MT163440 | A. marginale | MK016525.1 | 0.0 | 100.00 |
| 168 | MT163441 | A. marginale | MK310488.1 | 0.0 | 100.00 |
| 171 | MT163442 | A. marginale | MK310488.1 | 0.0 | 100.00 |
| 172 | MT163443 | A. marginale | MK016525.1 | 0.0 | 99.07 |
| 239 | MT163444 | A. marginale | MK310488.1 | 0.0 | 99.04 |
| 243 | MT163445 | A. marginale | MK016525.1 | 0.0 | 100.00 |
| 342 | MT163446 | A. marginale | MK310488.1 | 0.0 | 99.77 |
| 39 | MT160355 | A. bovis | MT036513.1 | 0.0 | 100.00 |
| 75 | MT160356 | A. bovis | MK028574.1 | 0.0 | 100.00 |
| 86 | MT160357 | A. bovis | MT036513.1 | 0.0 | 99.28 |
| 326 | MT160358 | A. bovis | MK028573.1 | 0.0 | 100.00 |
| 103 | MT163684 | Unidentified Anaplasma spp. | KY924885.1 | 0.0 | 100.00 |
| 112 | MT163683 | Unidentified Anaplasma spp. | KY924884.1 | 0.0 | 99.18 |
| 166 | MT163685 | Unidentified Anaplasma spp. | KY924884.1 | 0.0 | 97.85 |
Table 3.
Ehrlichia species detected by BLASTn analysis of 16S rDNA gene sequences
| Isolate | Our accession number | Matching sequence | Accession no. of highest match | E-value | % Identity |
|---|---|---|---|---|---|
| 32E | MT163429 | E. minasensis | MH500005.1 | 0.0 | 100.00 |
| 86E | MT163430 | E. minasensis | MH500005.1 | 0.0 | 99.42 |
| 175E | MT163431 | E. minasensis | MH500005.1 | 0.0 | 99.71 |
| 181E | MT163432 | E. minasensis | MH500005.1 | 0.0 | 100.00 |
Phylogenetic analysis
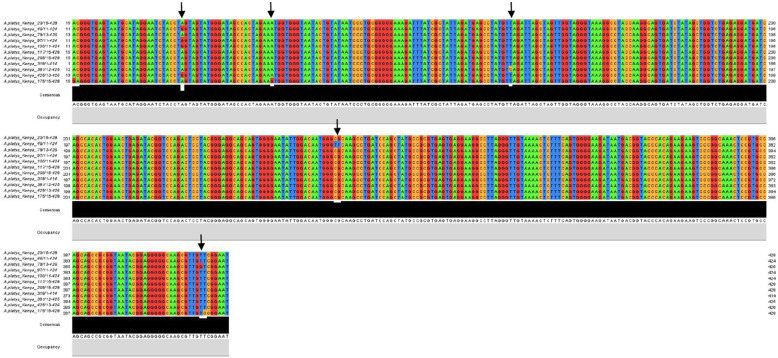
Multiple sequence alignment was done to assess the genetic similarity of the Kenyan isolates. The nucleotide sequences of three A.bovis isolates were conserved while one (MT160357) had three nucleotide polymorphisms at position 267, 268 and 332 (Table 4). Anaplasma platys sequences showed divergence of upto 4% (Table 5) with regions of nucleotide polmorphism (Fig. 2). The A. platys sequences MT163377 and MT163388 indicated multiple single nucleotide polymorphism while the other five isolates showed a single nucleotide polymorphism (SNP) (Table 4). All the Anaplasma marginale sequences from this study were however highly conserved sharing 97.6 to 100% nucleotide similarity (< 2.5% divergence) (Table 6). For E.minasensis, two isolates had conserved sequences while isolates MT163430 and MT163431 appeared to be genetically different showing multiple SNPs (Table 4). The multiple sequence nucleotide polymorphisms observed in the Kenyan isolates of A.bovis, A.platys and E.minasensis indicate that various strains of the pathogens may exist in the cattle in peri-urban Nairobi.
Table 4.
Nucleotide polymorphisms among 16SrDNA sequences of A. platys, A. bovis and E. minasensis Kenyan isolates
| aNucleotide position – Anaplasma platys | |||||||||
| Isolate | bAccession no. | 1 | 30 | 55 | 118 | 257 | 258 | 407 | 408 |
| 20 | MT163376 | A | A | A | T | C | G | T | T |
| 46 | MT163377 - MSNP | * | G | * | * | T | T | * | * |
| 79 | MT163379 -SNP | * | * | * | * | * | * | G | * |
| 97 | MT163380 -SNP | * | G | * | * | * | * | * | * |
| 100 | MT163381 -SNP | * | G | * | * | * | * | * | * |
| 175 | MT163388 - MSNP | G | G | G | * | * | * | * | C |
| 381 | MT163385 -SNP | * | * | * | C | * | * | * | * |
| aNucleotide position – Ehrlichia minasensis | |||||||||
| Isolate | bAccession no. | 1 | 130 | 257 | 652 | ||||
| 32E | MT163429 | G | C | A | G | ||||
| 181E | MT163432 | * | * | * | * | ||||
| 86E | MT163430 - MSNP | * | T | C | T | ||||
| 175E | MT163431-SNP | A | * | * | * | ||||
| aNucleotide position – Anaplasma bovis | |||||||||
| Isolate | bAccession no. | 267 | 268 | 332 | |||||
| 39 | MT160355 | C | G | G | |||||
| 75 | MT160356 | * | * | * | |||||
| 326 | MT160358 | * | * | * | |||||
| 86 | MT160357 - MSNP | T | T | A | |||||
Key: aNumbers denotes the nucleotide position on the sequence. Conserved nucleotide positions relative to the first sequence are indicated using asterisks while the specific nucleotide is indicated where a substitution occurred. MSNP- Multiple single sequence polymorphism, SNP-Single nucleotide polymorphism. Nucleotides: T-thymine, C-cytosine, G-guanine, A-adenine. bGenbank Accession numbers
Table 5.
Pairwise percent identity matches of 16SrDNA sequences of A. platys isolated from cattle in Kenya
| Isolates | ApN173 | ApN46 | ApN268 | ApN20 | ApN117 | ApN381 | ApN425 | ApN100 | ApN85 | ApN97 | ApN175 | ApN79 | ApN318 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ApN173 | 100.0 | 97.8 | 96.2 | 96.2 | 96.8 | 98.1 | 98.4 | 98.4 | 98.4 | 98.4 | 95.9 | 98.1 | 98.4 |
| ApN46 | 97.8 | 100.0 | 97.6 | 97.6 | 98.1 | 99.3 | 99.5 | 99.5 | 99.5 | 98.6 | 96.7 | 98.1 | 98.3 |
| ApN268 | 96.2 | 97.6 | 100.0 | 99.8 | 99.8 | 98.4 | 98.1 | 98.1 | 97.9 | 97.2 | 98.1 | 97.2 | 99.0 |
| ApN20 | 96.2 | 97.6 | 99.8 | 100.0 | 99.8 | 98.4 | 98.1 | 98.1 | 97.9 | 97.2 | 98.1 | 97.2 | 99.0 |
| ApN117 | 96.8 | 98.1 | 99.8 | 99.8 | 100.0 | 98.8 | 98.6 | 98.6 | 98.4 | 97.6 | 98.6 | 97.6 | 99.0 |
| ApN381 | 98.1 | 99.3 | 98.4 | 98.4 | 98.8 | 100.0 | 99.8 | 99.8 | 99.7 | 98.8 | 97.4 | 98.8 | 99.0 |
| ApN425 | 98.4 | 99.5 | 98.1 | 98.1 | 98.6 | 99.8 | 100.0 | 100.0 | 100.0 | 99.1 | 97.2 | 98.6 | 98.8 |
| ApN100 | 98.4 | 99.5 | 98.1 | 98.1 | 98.6 | 99.8 | 100.0 | 100.0 | 100.0 | 99.1 | 97.2 | 98.6 | 98.8 |
| ApN85 | 98.4 | 99.5 | 97.9 | 97.9 | 98.4 | 99.7 | 100.0 | 100.0 | 100.0 | 100.0 | 97.6 | 99.7 | 99.7 |
| ApN97 | 98.4 | 98.6 | 97.2 | 97.2 | 97.6 | 98.8 | 99.1 | 99.1 | 100.0 | 100.0 | 97.6 | 99.5 | 99.8 |
| ApN175 | 95.9 | 96.7 | 98.1 | 98.1 | 98.6 | 97.4 | 97.2 | 97.2 | 97.6 | 97.6 | 100.0 | 97.6 | 99.0 |
| ApN79 | 98.1 | 98.1 | 97.2 | 97.2 | 97.6 | 98.8 | 98.6 | 98.6 | 99.7 | 99.5 | 97.6 | 100.0 | 99.8 |
| ApN318 | 98.4 | 98.3 | 99.0 | 99.0 | 99.0 | 99.0 | 98.8 | 98.8 | 99.7 | 99.8 | 99.0 | 99.8 | 100.0 |
Key: Abbreviation: ApN Anaplasma platys Nairobi, followed by the isolate number
Fig. 2.
Multiple sequence alignment of A. platys 16S rDNA, indicating areas of sequence nucleotide polymorphism (black arrows). Numbers at the ends of each sequence indicate nucleotide lengths while the isolate names are indicated on the far left end of the nucleotide sequences
Table 6.
Pairwise percent identity matches of 16SrDNA sequences of A. marginale isolated from cattle in Kenya
| Isolates | AMN239 | AMN172 | AMN168 | AMN139 | AMN159 | AMN171 | AMN243 | AMN127 | AMN342 |
|---|---|---|---|---|---|---|---|---|---|
| AMN239 | 100.0 | 99.1 | 97.6 | 97.6 | 97.6 | 97.6 | 97.6 | 98.1 | 98.3 |
| AMN172 | 99.1 | 100.0 | 98.4 | 97.6 | 97.6 | 97.6 | 97.6 | 98.8 | 99.1 |
| AMN168 | 97.6 | 98.4 | 100.0 | 97.6 | 97.6 | 97.6 | 97.6 | 99.1 | 99.1 |
| AMN139 | 97.6 | 97.6 | 97.6 | 100.0 | 100.0 | 100.0 | 100.0 | 98.1 | 98.3 |
| AMN159 | 97.6 | 97.6 | 97.6 | 100.0 | 100.0 | 100.0 | 100.0 | 98.1 | 98.3 |
| AMN171 | 97.6 | 97.6 | 97.6 | 100.0 | 100.0 | 100.0 | 100.0 | 98.1 | 98.3 |
| AMN243 | 97.6 | 97.6 | 97.6 | 100.0 | 100.0 | 100.0 | 100.0 | 98.1 | 98.3 |
| AMN127 | 98.1 | 98.8 | 99.1 | 98.1 | 98.1 | 98.1 | 98.1 | 100.0 | 100.0 |
| AMN342 | 98.3 | 99.1 | 99.1 | 98.3 | 98.3 | 98.3 | 98.3 | 100.0 | 100.0 |
Key: The numbers denote the nucleotide identity rates found between the sequences. Abbreviation: AMN Anaplasma marginale Nairobi, followed by the isolate number
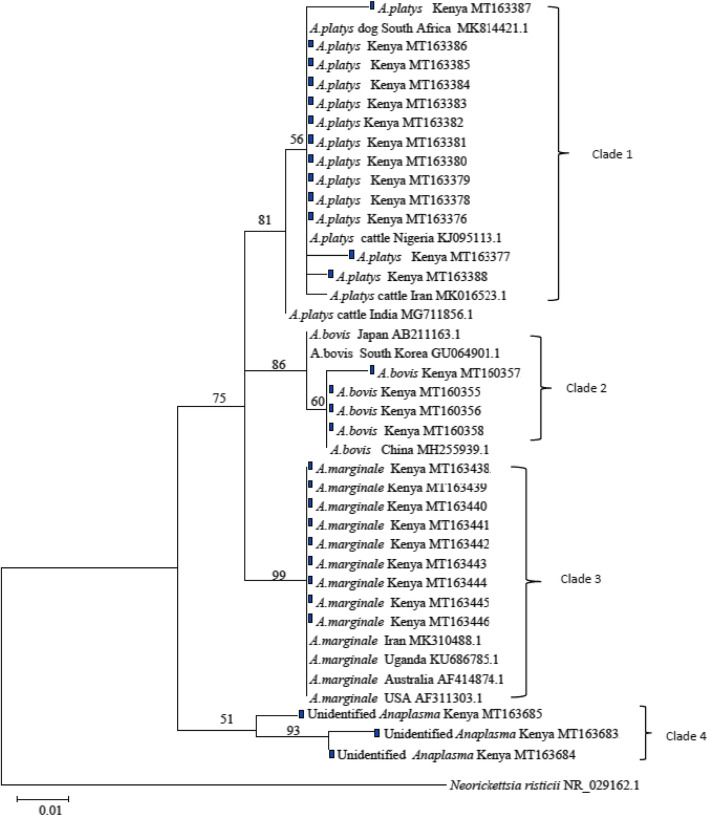
Phylogenetic analysis was done to understand genetic relatedness of the Kenyan isolates of the two genera with those of annotated sequences in GenBank (Figs. 3 and 4). The Kenyan isolates of A. platys clustered in the same clade as those of A. platys isolated from South Africa, Nigeria and Iran. They were however distinct from an isolate from India accession number MG711856.1 (Fig. 3-Clade 1). The Kenyan isolates of A. marginale were closely related to those from Uganda, USA, Australia and Iran (Clade 3). Anaplasma bovis isolates from Kenya were closely related to those from China but distantly related to those from South Korea and Japan (Clade 2). The unidentified Anaplasma species from this study clustered in their own clade separate from A. platys, A. marginale and A. bovis (Clade 4). For the Ehrlichia species, phylogeny was done to compare the detected E. minasensis genetic relatedness to other characterized species such as E. canis, the dog pathogen and the more common ruminant pathogen, E. ruminantium. The E. minasensis isolated in this study grouped in one clade with other isolates from USA, Australia and Brazil. These isolates were however closely related to E. canis than E. ruminantium (Fig. 4).
Fig. 3.
Maximum Likelihood tree of Anaplasma spp. constructed using partial sequences of 16S rDNA gene. The tree is drawn to scale with branch lengths measured in the number of substitutions per site. The analysis involved 29 nucleotide sequences from this study and 12 others obtained from Genbank. The tree shows the phylogenetic relatedness of Anaplasma isolates obtained from cattle blood in Kenya marked with dark box and sequences from other countries. Neorickettsia risticii was used as the outgroup. Sequence accession numbers are given at the end of each isolate
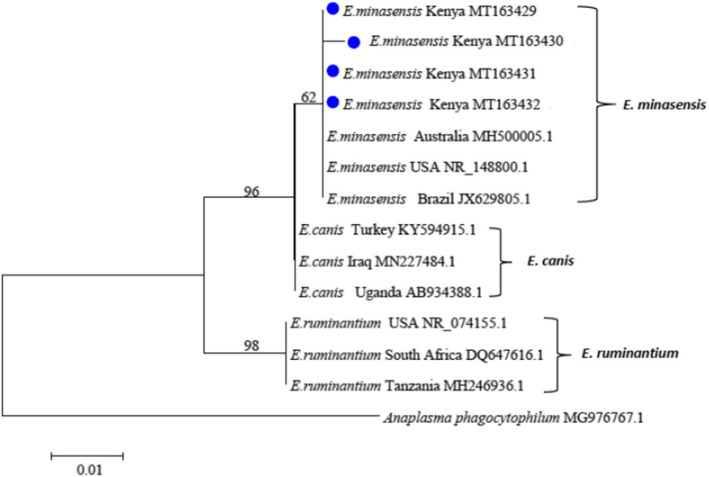
Fig. 4.
Maximum Likelihood tree of Ehrlichia spp. constructed using partial sequences of 16S rDNA gene. The tree is drawn to scale, with branch lengths measured in the number of substitutions per site. The analysis involved 4 nucleotide sequences from this study and 10 others obtained from Genbank. The phylogeny shows the relatedness of E. minasensis isolated from this study marked with blue dot with other isolates from USA, Brazil and Australia and its relation to E. canis and E. ruminantium. Anaplasma phagocytophilum was used as the outgroup
Discussion
This study aimed at detecting and characterizing Anaplasma and Ehrlichia species infecting dairy cattle in peri-urban Nairobi. This information is important in guiding development of control and preventive measures against the infections caused by these tick-borne pathogens.
Overall, we observed more Anaplasma infections than Ehrlichia in the study cattle population. Previous studies in Kenya [7], Sudan [24] and Ethiopia [25] also reported more Anaplasma pathogens than Ehlichia. This could be explained by the wide diversity of Anaplasma compared to Ehrlichia species that can potentially infect cattle [3]. Moreover, spatial occurrence of tick-borne pathogens has been associated with the presence of their tick vectors [26], so it could be that tick vectors which transmit Anaplasma species are more widespread in the study area than those of Ehrlichia pathogens.
The distribution of the infections varied across sample sub-counties. A higher proportion of cattle were infected with Anaplasma and Ehrlichia spp. in Kasarani-Ruai Sub-County compared to the other three study areas of Dagoretti, Westlands and Lang’ata. A possible explanation of this is that dairy farmers in Kasarani-Ruai practice mixed production system involving free and zero grazing unlike the other areas where farmers practiced exclusive zero-grazing. Various studies have shown that free grazing cattle have higher risk of tick-borne infections than zero grazed cattle because of high exposure to tick vectors [22].
Anaplasma platys pathogens were detected in the study cattle. Anaplasma platys has been considered an emerging Anaplasma species whose clinical disease is yet to be described [9, 27]. Previous studies in Algeria [2], Senegal [27] and Tunisia [28] similarly reported this pathogen in cattle. Yang et al. [29] suggested a possibility of domestic ruminants acting as alternative hosts or reservoirs for A. platys which is typically a canine pathogen [30]. Therefore, the detection of this pathogen in cattle raises questions of host specificity as earlier speculated [31].
Zobba et al. [9] noted that several domestic ruminants can harbor a number of strains of A. platys although these strains have different cell tropism compared to those infecting dogs. The ruminant strains infect neutrophils and are thought to be the ancestral pathogens that evolved to adopt to the canine platelets instead [9, 30]. We think that infection with A. platys may be associated with co-existence of dogs and cattle in the same households, a common practice observed in the dairy farms in peri-urban Nairobi. It is possible that tick-bites from Rhipicephalus sanguineus which are the main vectors of A. platys [32] may have played a role in the transmission. Screening of dogs for this pathogen can reveal if they are acting as maintenance hosts of the parasite.
Previous studies have documented the zoonotic potential of A. platys causing human disease characterized by headaches, intermittent edema and muscle pains [33]. In this regard, detection of A. platys pathogens in this study would indicate a possible zoonotic health risk to cattle owners who are in constant contact with their cattle.
In this study, cattle were also found to be infected with A. marginale, a common pathogen of cattle that has been reported in Eastern Africa [22], Southern Africa [34], North Africa [27] and West Africa [32]. It is not surprising to detect this pathogen in cattle in Kenya since Rhipicephalus (boophilus) tick species which are the documented vectors of A. marginale are widespread in Kenya. Anaplasma marginale causes a mild to severe anemia depending on the susceptibility of the cattle [35]. However, cattle in this study were apparently healthy suggesting a possible endemic situation or the animals had a persistent infection (PI) state which is known to occur in A. marginale infections [36]. Thus, infected animals can appear apparently healthy despite harboring the pathogen.
Anaplasma bovis which is a monocytic pathogen of ruminants was also detected in this study. Different tick species in the genera Amblyomma and Rhipicephalus have been documented to transmit this pathogen [6]. Similar studies have detected this pathogen in Kenya [7], China [37], South Korea [3] Tunisia [38] and Algeria [39]. Despite it causing a mild disease in cattle, some infected animals have been shown to manifest with decreased weight gain, fever and lymphadenopathy [40]. Other studies have documented sub-clinical infection with this parasite where animals don’t show clinical signs of the disease despite the infection [41] and this may have been the case in our study.
To date, bovine anaplasmosis in Kenya is mainly known to be caused by A. marginale and to some extent A. bovis [7, 16, 22]. However, we have detected A. platys pathogens for the first time in cattle in Kenya possibly contributing equally to the disease burden in dairy cattle. Further studies to investigate this pathogen using more specific genes such as membrane surface proteins (Msps) [42] are justified.
An emerging pathogen E. minasensis in the Ehrlichia genera was also detected in this study. This novel pathogen was initially reported in Canada [43] and Brazil [44] but has since been isolated in Ethiopia [45], South Africa [46], Pakistan [47] and China [48]. The clinical disease due to E. minasensis is variable with some reports of severe disease [14] and at times sub-clinical disease as observed in this study [45]. Screening of ticks collected from animals may confirm if the pathogen is common in cattle.
Although the specific ticks that transmit E. minasensis have not been well studied [49], its detection and transstadial transmission by Rhipicephalus microplus ticks has been documented [50]. Other tick species may still transmit the pathogen in areas where R. microplus is absent [45]. Indeed, Iweriebor et al. [46] detected this pathogen from R. appendiculatus, R. evertsi eversi, R. sanguineus and Ambylomma hebraeum ticks. Some of these tick species were observed to be infesting some animals in this study. It is possible that these ticks could be involved in transmission of this pathogen.
Similar to other studies [38, 51], phylogenetic analysis based on 16S rDNA was used to infer genetic diversity of Anaplasma and Ehrlichia species. However, other authors have used 16S rDNA gene in combination with other genes such as heat shock protein (groEL), citrate synthase (gltA), 23S rDNA and major surface protein 4 gene (msp4) [2, 52]. Combined gene assays have been used to enhance sensitivity since genes with multiple copies such as membrane surface proteins (MSP) are more sensitive for detection of Anaplasmataceae while more conserved genes (16S rDNA) are useful for database cross matching and sequence comparisons [53]. In this study, A. marginale isolates were found to be highly conserved indicating sequence divergence of less than 2.5% and clustered together with those from USA, Uganda, Iran and Australia similar to findings by Rjeibi et al. [39]. Anaplasma bovis, A. platys pathogens and E. minasensis strains detected in this study indicated certain levels of nucleotide polymorphism suggesting various strains of the pathogens may exist in the study cattle. This may be related to the increased cattle movement from other regions of the country for slaughter at the country’s major export abbatoirs located in Nairobi County. Extensive animal movement has been associated with development of new strains and introduction of the tick-borne pathogens to new geographic areas [38, 54].
In agreement with previous studies, phylogenetic analysis also indicated that E. minasensis is closely related to E. canis and distantly related to E. ruminantium despite infecting similar hosts [12, 55]. The clinical presentation of E. minasensis in cattle has been observed to be similar to the acute form of disease by E. canis in dogs [44, 56]. Cabezas-Cruz et al. [57] links the close relatedness of the two pathogens to possible evolution of E. minasensis from highly variable strains of E. canis. The detection of this novel Ehrlichia species suggests that it could be circulating in cattle in Kenya and its pathogenicity in the affected animals needs to be determined.
Despite the animals in this study not presenting with the clinical signs of the diseases caused by the pathogens they harbor, poor animal husbandry practices which are commonly practiced in smallholder dairy farms in peri-urban areas of Nairobi [58], causes stress to the animals consequently predisposing them to possible flaring up of clinical disease and mortalities [59]. The detection of these pathogen therefore highlights the importance of continued investigation into tick-borne diseases and the need for effective diagnosis and prevention.
Conclusion
The dairy cattle from peri-urban Nairobi, Kenya are infected with a range of Anaplasma species and Ehrlichia minasensis even though clinical disease was not evident. There is need for accurate diagnosis and effective tick control so as to reduce infection of cattle with these pathogens. To the best of our knowledge, this study provided first reports of cattle infected with A. platys and E. minasensis pathogens in Kenya. Extensive epidemiological studies would be necessary to determine the extent and pathogenicity of the newly detected A. platys and E. minasensis in cattle. Moreover, investigation into tick vectors involved in their transmission will be needed to inform strategic disease management and control. Moreover, there is need for further investigation of the unidentified Anaplasma species using more specific genes such as membrane surface proteins (Msps) [42].
Methods
Study area and design
The study area and design have been described in detail previously [23]. This was a cross-sectional study undertaken between January and May 2017. For purposes of data collection, Nairobi County was divided into four quadrants and for each quadrant purposive sampling was used to select the sub-county with high cattle population. Dairy farms in the four sub-counties in peri-urban Nairobi, namely; Dagoretti, Lang’ata, Kasarani-Ruai and Westlands were included to this study. These sub-counties are part of the peri-urban areas of the Nairobi County where dairy production has been established to meet the high milk demands of the urban population. Animals of different age-groups ranging from 3 months to 8 years were randomly selected and sampled for whole blood.
Collection of cattle blood samples
Three milliliter of whole blood were collected in ethylenediaminetetraacetic acid (EDTA)-coated vacutainers from the coccygeal vein of 306 apparently healthy dairy cattle. The samples were collected as follows: Dagoretti (n = 116), Lang’ata (n = 55), Kasarani-Ruai (n = 110) and Westlands (n = 25). The samples were then transported in an ice-box to the Molecular Laboratory at the Department of Public Health, Pharmacology and Toxicology of the University of Nairobi and stored at -20 °C pending subsequent analysis.
Extraction of Anaplasma and Ehrlichia DNAs
Whole blood genomic DNA (gDNA) was extracted from aliquots of 200 μl using QIAamp DNA Blood Mini Kit (Qiagen, Hilden, Germany) following manufacturer’s instructions. The DNA concentration and quality were assessed using QIAxpert (Qiagen, Hilden, Germany). DNA samples were then stored at − 200 C.
Primers design and PCR-amplification of Anaplasma and Ehrlichia DNA
A forward primer, ANAF 5′-TAGTGGCAGACGGGTGAGTA-3′ and a reverse ANAR 5′-AATTCCGAACAACGCTTGCC-3′ targeting an approximately 424 bp of Anaplasma 16S rDNA were designed using the Primer-BLAST tool of the National Center for Biotechnology Information [60] (NCBI) (www.ncbi.nlm.nih.gov/tools/primer-blast).
A forward primer EHRF 5′-AGCTGGTCTGAGAGGACGAT-3′ and a reverse primer EHRR 5′-GAGTGCCCAGCATTACCTGT-3′ targeting an approximately 838 bp of Ehrlichia 16S rDNA were also designed. PCR amplifications were performed using a thermal cycler (Applied biosystems Veriti 96 well, ThermoFisher). The Anaplasma and Ehrlichia 16S rDNA were amplified in a final volume of 20 μl reaction, each containing 3 μl of genomic DNA, 10 μl Master-mix (Taq PCR 2x mastermix, Qiagen, Germany) and 10 μM final concentration of each primer. The thermocycling conditions for Anaplasma involved a pre-denaturation at 95o C for 5 min followed by 40 cycles of denaturation at 95o C for 45 s, annealing at 57o C for 45 s and extension at 72o C for 45 s. A final cycle of extension at 72 °C for 7 min was performed. The amplification conditions for Ehrlichia 16S rDNA involved an initial denaturation cycle at 95o C for 5 min followed by 35 cycles of denaturation at 95 °C for 45 s, annealing at 62o C for 45 s and extension at 72o C for 45 s. The amplification cycles were followed by a final cycle of extension at 72o C for 7 min. Double distilled water was used as negative control for both assays. The amplified products were electrophoresed using 1.5% agarose gel in Tris-Borate-EDTA (TBE) buffer, pH 8, stained with Ethidium Bromide and visualized using UV-illuminator (GelMax® Imager, UK). The sizes of the amplicons were determined using molecular ladder (Gelpilot 1 kb plus ladder (100), Qiagen, Germany).
Purification and sequencing of Anaplasma and Ehrlichia DNA
PCR amplicons in gels were excised and purified using QIAquick Gel Extraction Kit (Qiagen, GmbH, Germany) following the manufacturer’s protocol. The purified DNA were sequenced at Macrogen Europe Laboratories (Amsterdam, The Netherlands). Sequencing was done using the same forward and reverse primers as for the PCR reactions. The obtained sequences were viewed and manually verified using chromatogram peaks, edited and assembled using CLC Main Workbench 6.8.3 software (CLC bio, Qiagen GmbH, Germany).
Data analysis
Bioinformatics analysis of the parasite 16S rDNA sequences was done by using Basic Local Alignment Search Tooln (BLASTn), multiple sequence alignment and phylogenetic analyses. Sequence identities of the Anaplasma and Ehrlichia species were confirmed by BLASTn analysis [61] at https://blast.ncbi.nlm.nih.gov/Blast.cgi. Multiple sequence alignment was done using Log-Expectation (MUSCLE) v3.8.31 [62]. Sequence similarity was calculated using Clustal Omega to obtain identity matrixs [63]. A phylogenetic reconstruction was done using MEGA 6.0 [64]. The evolutionary history was inferred by using the Maximum Likelihood method based on the Tamura-Nei model [65]. Initial trees for the heuristic search were obtained automatically by applying Neighbor-Join and BioNJ algorithms to a matrix of pairwise distances estimated using the Maximum Composite Likelihood (MCL) approach and then selecting the topology with superior log likelihood value. All positions containing gaps and missing data were eliminated. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1000 replicates) was shown next to the branches.
Nucleotide sequence accession numbers
The partial 16S rDNA sequences obtained in this study were deposited in the GenBank under the following accession numbers; MT163376 to MT163388 for A. platys, MT160355 to MT160358 for A. bovis, MT163438 to MT163446 for A. marginale, MT163683 to MT163685 for unidentified Anaplasma species and MT163429 to MT163432 for E. minasensis.
Acknowledgements
We wish to acknowledge the support of Ann Wambui and Judy Barasa who were instrumental in the recruitment of the farms to the study. Our special gratitude goes to the farmers from Dagorreti, Lang’ata (Karen), Kasarani-Ruai and Westlands (Kabete and Roysambu) Sub-Counties who allowed their animals to be included into the study.
Abbreviations
- TBDs
Tick-borne diseases
- CI
Confidence Interval
- ELISA
Enzyme-linked immunosorbent Assay
- BLAST
Basic Local Alignment Search Tool
- MEGA
Molecular Evolutionary Genetics Analysis
- PI
Persistent Infection
Authors’ contributions
SGP, GOA, DWG, NM and CMM designed the study, SGP and DWG collected data, SGP, GOA and AMO designed, optimized the primers and ran PCR assays, SGP, HWK and EGK analyzed the sequence data, all the authors were involved in the drafting of the manuscript and approved the final submitted manuscript.
Funding
“This research was supported by the Consortium for Advanced Research Training in Africa (CARTA). CARTA is jointly led by the African Population and Health Research Center and the University of the Witwatersrand and funded by the Carnegie Corporation of New York (Grant No--B 8606.R02), Sida (Grant No: 54100113), the DELTAS Africa Initiative (Grant No: 107768/Z/15/Z) and Deutscher Akademischer Austauschdienst (DAAD). The DELTAS Africa Initiative is an independent funding scheme of the African Academy of Sciences (AAS)‘s Alliance for Accelerating Excellence in Science in Africa (AESA) and supported by the New Partnership for Africa’s Development Planning and Coordinating Agency (NEPAD Agency) with funding from the Wellcome Trust (UK) and the UK government. The statements made and views expressed in this research work are solely the responsibility of the fellow”. The role of the funding body was to finance the consumables used in this study. They had no role in the design of the study, collection, analysis and interpretation of the data and in the writing of the manuscript.
Availability of data and materials
Dataset used and analyzed during the current study are available from the corresponding author on reasonable request. Sequences obtained and analyzed from this study are available in GenBank database on the accession numbers indicated.
Ethics approval and consent to participate
The study was approved by the Biosecurity, Animal Use and Ethics Committee (BAUEC) at the Faculty of Veterinary Medicine, University of Nairobi, Kenya (FVM BAUEC/2016/122). Blood collection in cattle was carried out in accordance with the guidelines laid down by the BAUEC requiring keen adherence to animal welfare and strict observance of biosecurity measures during handling of infectious materials. The study was described in full details to the cattle owners after which they gave informed consent verbally for them to participate and for their animals to be included in the study.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no conflict of interest.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Ismail N, McBride JW. Tick-borne emerging infections: Ehrlichiosis and Anaplasmosis. Clin Lab Med. 2017;37:317–340. doi: 10.1016/j.cll.2017.01.006. [DOI] [PubMed] [Google Scholar]
- 2.Dahmani M, Davoust B, Benterki MS, Fenollar F, Raoult D, Mediannikov O. Development of a new PCR-based assay to detect Anaplasmataceae and the first report of Anaplasma phagocytophilum and Anaplasma platys in cattle from Algeria. Comp Immunol Microbiol Infect Dis. 2015;39:39–45. doi: 10.1016/j.cimid.2015.02.002. [DOI] [PubMed] [Google Scholar]
- 3.Park J, Han DG, Ryu JH, Chae JB, Chae JS, Yu DH, et al. Molecular detection of Anaplasma bovis in Holstein cattle in the Republic of Korea. Acta Vet Scand. 2018;60:1–5. doi: 10.1186/s13028-018-0370-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Potgieter FT, van Rensburg L. Tick transmission of Anaplasma Centrale. Onderstepoort J Vet Res. 1987;54:5–7. [PubMed] [Google Scholar]
- 5.Kocan KM, de la Fuente J, Cabezas-Cruz A. The genus Anaplasma: new challenges after reclassification Introduction and current classification. Rev Sci Tech Off Int Epiz. 2015;34:577–586. doi: 10.20506/rst.34.2.2381. [DOI] [PubMed] [Google Scholar]
- 6.Omondi D, Masiga DK, Fielding BC, Kariuki E, Ajamma YU, Mwamuye MM, et al. Molecular Detection of Tick-Borne Pathogen Diversities in Ticks from Livestock and Reptiles along the Shores and Adjacent Islands of Lake Victoria and Lake Baringo, Kenya. Front Vet Sci. 2017;4:1–15. doi: 10.3389/fvets.2017.00073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Njiiri NE, Bronsvoort BM d C, Collins NE, Steyn HC, Troskie M, Vorster I, et al. The epidemiology of tick-borne haemoparasites as determined by the reverse line blot hybridization assay in an intensively studied cohort of calves in western Kenya. Vet Parasitol. 2015;210:69–76. doi: 10.1016/j.vetpar.2015.02.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Ybañez AP, Inokuma H. Anaplasma species of veterinary importance in Japan. Vet World. 2016;9:1190–1196. doi: 10.14202/vetworld.2016.1190-1196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Zobba R, Anfossi AG, Parpaglia MLP, Dore GM, Chessa B, Spezzigu A, et al. Molecular investigation and phylogeny of anaplasma spp. in mediterranean ruminants reveal the presence of neutrophil-tropic strains closely related to a. platys. Appl Environ Microbiol. 2014;80:271–280. doi: 10.1128/AEM.03129-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Noaman V, Shayan P. Molecular detection of Anaplasma phagocytophilum in carrier cattle of Iran - first documented report. Iranian J Microbiol. 2009;1:37–42. [Google Scholar]
- 11.Mwamuye MM, Kariuki E, Omondi D, Kabii J, Odongo D, Masiga D, et al. Novel rickettsia and emergent tick-borne pathogens: a molecular survey of ticks and tick-borne pathogens in Shimba Hills National Reserve, Kenya. Ticks Tick-borne Dis. 2017;8:208–218. doi: 10.1016/j.ttbdis.2016.09.002. [DOI] [PubMed] [Google Scholar]
- 12.Cabezas-Cruz A, Zweygarth E, Aguiar DM. Ehrlichia minasensis, an old demon with a new name. Ticks Tick-borne Dis. 2019;10:828–829. doi: 10.1016/j.ttbdis.2019.03.018. [DOI] [PubMed] [Google Scholar]
- 13.Allsopp BA. Heartwater--Ehrlichia ruminantium infection. Revue Sci Tech (International Office of Epizootics) 2015;34:557–568. doi: 10.20506/rst.34.2.2379. [DOI] [PubMed] [Google Scholar]
- 14.de Aguiar DM, Junior JPA, Nakazato L, Bard E, Aguilar-Bultet L, Vorimore F, et al. Isolation and characterization of a novel pathogenic strain of Ehrlichia minasensis. Microorganisms. 2019;7:528. doi: 10.3390/microorganisms7110528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kivaria FM. Estimated direct economic costs associated with tick-borne diseases on cattle in Tanzania. Trop Anim Health Prod. 2006;38:291–299. doi: 10.1007/s11250-006-4181-2. [DOI] [PubMed] [Google Scholar]
- 16.Gakuya DW, Mulei CM. An investigation of the incidences of East Coast fever (ECF), Anaplasmosis and Babesiosis in the bovine cases managed by ambulatory services of the faculty of veterinary medicine, Kabete. Kenya Veterinarian. 2005;29:21–24. [Google Scholar]
- 17.Wesonga FD, Kitala PM, Gathuma JM, Njenga MJ, Ngumi PN. An assessment of tick-borne diseases constraints to livestock production in a smallholder livestock production system in Machakos District, Kenya. Livestock Res Rural Dev. 2010;22 https://www.lrrd.cipav.org.co/lrrd22/6/weso22111.htm. Accessed 13 Mar 2020.
- 18.Allsopp BA. Natural history of Ehrlichia ruminantium. Vet Parasitol. 2010;167:123–135. doi: 10.1016/j.vetpar.2009.09.014. [DOI] [PubMed] [Google Scholar]
- 19.Muraguri GR, McLeod A, McDermott JJ, Taylor N. The incidence of calf morbidity and mortality due to vector-borne infections in smallholder dairy farms in Kwale District, Kenya. Vet Parasitol. 2005;130:305–315. doi: 10.1016/j.vetpar.2004.11.026. [DOI] [PubMed] [Google Scholar]
- 20.Kiara H, Jennings A, Bronsvoort BMDC, Handel IG, Mwangi ST, Mbole-Kariuki M, et al. A longitudinal assessment of the serological response to Theileria parva and other tick-borne parasites from birth to one year in a cohort of indigenous calves in western Kenya. Parasitology. 2014;141:1289–1298. doi: 10.1017/S003118201400050X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Wesonga FD, Gachohi JM, Kitala PM, Gathuma JM, Njenga MJ. Seroprevalence of Anaplasma marginale and Babesia bigemina infections and associated risk factors in Machakos County, Kenya. Trop Anim Health Prod. 2017;49:265–272. doi: 10.1007/s11250-016-1187-2. [DOI] [PubMed] [Google Scholar]
- 22.Adjou Moumouni PF, Aboge GO, Terkawi MA, Masatani T, Cao S, Kamyingkird K, et al. Molecular detection and characterization of Babesia bovis, Babesia bigemina, Theileria species and Anaplasma marginale isolated from cattle in Kenya. Parasites Vectors. 2015;8:1–14. doi: 10.1186/s13071-015-1106-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Peter SG, Gakuya DW, Maingi N, Mulei CM. Prevalence and risk factors associated with Ehrlichia infections in smallholder dairy cattle in Nairobi City County, Kenya. Vet World. 2019;12:1599–1607. doi: 10.14202/vetworld.2019.1599-1607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Eisawi NM, El Hussein ARM, Hassan DA, Musa AB, Hussien MO, Enan KA, et al. A molecular prevalence survey on Anaplasma infection among domestic ruminants in Khartoum State, Sudan. Trop Anim Health Prod. 2020;52:1845–1852. doi: 10.1007/s11250-019-02176-7. [DOI] [PubMed] [Google Scholar]
- 25.Teshale S, Geysen D, Ameni G, Dorny P, Berkvens D. Survey of Anaplasma phagocytophilum and Anaplasma sp. “Omatjenne” infection in cattle in Africa with special reference to Ethiopia. Parasites Vectors. 2018;11:162. doi: 10.1186/s13071-018-2633-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Wikel SK. Ticks and tick-borne infections: complex ecology, agents, and host interactions. Vet Sci. 2018;5:60. doi: 10.3390/vetsci5020060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Dahmani M, Davoust B, Sambou M, Bassene H, Scandola P, Ameur T, et al. Molecular investigation and phylogeny of species of the Anaplasmataceae infecting animals and ticks in Senegal. Parasites Vectors. 2019;12:1–15. doi: 10.1186/s13071-019-3742-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Ben Said M, Belkahia H, El Mabrouk N, Saidani M, Alberti A, Zobba R, et al. Anaplasma platys-like strains in ruminants from Tunisia. Infect Genet Evol. 2017;49:226–233. doi: 10.1016/j.meegid.2017.01.023. [DOI] [PubMed] [Google Scholar]
- 29.Yang J, Han R, Niu Q, Liu Z, Guan G, Liu G, et al. Occurrence of four Anaplasma species with veterinary and public health significance in sheep, northwestern China. Ticks Tick Borne Dis. 2018;9:82–85. doi: 10.1016/j.ttbdis.2017.10.005. [DOI] [PubMed] [Google Scholar]
- 30.Ybañez AP, Ybañez RHD, Yokoyama N, Inokuma H. Multiple infections of Anaplasma platys variants in Philippine dogs. Veterinary World. 2016;9:1456–1460. doi: 10.14202/vetworld.2016.1456-1460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Bastos ADS, Mohammed OB, Bennett NC, Petevinos C, Alagaili AN. Molecular detection of novel Anaplasmataceae closely related to Anaplasma platys and Ehrlichia canis in the dromedary camel (Camelus dromedarius) Vet Microbiol. 2015;179:310–314. doi: 10.1016/j.vetmic.2015.06.001. [DOI] [PubMed] [Google Scholar]
- 32.Lorusso V, Wijnveld M, Majekodunmi AO, Dongkum C, Fajinmi A, Dogo AG, et al. Tick-borne pathogens of zoonotic and veterinary importance in Nigerian cattle. Parasites Vectors. 2016;9:1–13. doi: 10.1186/s13071-016-1504-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Breitschwerdt EB, Hegarty BC, Qurollo BA, Saito TB, Maggi RG, Blanton LS, et al. Intravascular persistence of Anaplasma platys, Ehrlichia chaffeensis, and Ehrlichia ewingii DNA in the blood of a dog and two family members. Parasites Vectors. 2014;7:1–7. doi: 10.1186/1756-3305-7-298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Fernandes S d J, Matos CA, Freschi CR, de Souza Ramos IA, Machado RZ, André MR. Diversity of Anaplasma species in cattle in Mozambique. Ticks Tick Borne Dis. 2019;10:651–664. doi: 10.1016/j.ttbdis.2019.02.012. [DOI] [PubMed] [Google Scholar]
- 35.Aktas M, Özübek S. Outbreak of anaplasmosis associated with novel genetic variants of Anaplasma marginale in a dairy cattle. Comp Immunol Microbiol Infect Dis. 2017;54:20–26. doi: 10.1016/j.cimid.2017.07.008. [DOI] [PubMed] [Google Scholar]
- 36.Kocan KM, de la Fuente J, Blouin EF, Coetzee JF, Ewing SA. The natural history of Anaplasma marginale. Vet Parasitol. 2010;167:95–107. doi: 10.1016/j.vetpar.2009.09.012. [DOI] [PubMed] [Google Scholar]
- 37.Yang J, Li Y, Liu Z, Liu J, Niu Q, Ren Q, et al. Molecular detection and characterization of Anaplasma spp. in sheep and cattle from Xinjiang, Northwest China. Parasites Vectors. 2015;8:1–7. doi: 10.1186/s13071-014-0608-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Belkahia H, Ben Said M, Alberti A, Abdi K, Issaoui Z, Hattab D, et al. First molecular survey and novel genetic variants’ identification of Anaplasma marginale, A. centrale and A. bovis in cattle from Tunisia. Infect Genet Evol. 2015;34:361–371. doi: 10.1016/j.meegid.2015.06.017. [DOI] [PubMed] [Google Scholar]
- 39.Rjeibi MR, Ayadi O, Rekik M, Gharbi M. Molecular survey and genetic characterization of Anaplasma Centrale, A. marginale and A. bovis in cattle from Algeria. Transbound Emerg Dis. 2018;65:456–464. doi: 10.1111/tbed.12725. [DOI] [PubMed] [Google Scholar]
- 40.Noaman V, Shayan P. Molecular detection of Anaplasma bovis in cattle from central part of Iran. Vet Res Forum. 2010;1:117–122. [Google Scholar]
- 41.Aktas M, Altay K, Dumanli N. Molecular detection and identification of Anaplasma and Ehrlichia species in cattle from Turkey. Ticks Tick Borne Dis. 2011;2:62–65. doi: 10.1016/j.ttbdis.2010.11.002. [DOI] [PubMed] [Google Scholar]
- 42.de la Fuente J, Lew A, Lutz H, Meli ML, Hofmann-Lehmann R, Shkap V, et al. Genetic diversity of Anaplasma species major surface proteins and implications for anaplasmosis serodiagnosis and vaccine development. Anim Health Res Rev. 2005;6:75–89. doi: 10.1079/ahr2005104. [DOI] [PubMed] [Google Scholar]
- 43.Gajadhar AA, Lobanov V, Scandrett WB, Campbell J, Al-Adhami B. A novel Ehrlichia genotype detected in naturally infected cattle in North America. Vet Parasitol. 2010;173:324–329. doi: 10.1016/j.vetpar.2010.06.034. [DOI] [PubMed] [Google Scholar]
- 44.Aguiar DM, Ziliani TF, Zhang X, Melo ALT, Braga ÍA, Witter R, et al. A novel Ehrlichia genotype strain distinguished by the TRP36 gene naturally infects cattle in Brazil and causes clinical manifestations associated with ehrlichiosis. Ticks Tick Borne Dis. 2014;5:537–544. doi: 10.1016/j.ttbdis.2014.03.010. [DOI] [PubMed] [Google Scholar]
- 45.Hailemariam Z, Krücken J, Baumann M, Ahmed JS, Clausen PH, Nijhof AM. Molecular detection of tick-borne pathogens in cattle from southwestern Ethiopia. PLoS One. 2017;12:1–16. doi: 10.1371/journal.pone.0188248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Iweriebor BC, Mmbaga EJ, Adegborioye A, Igwaran A, Obi LC, Okoh AI. Genetic profiling for Anaplasma and Ehrlichia species in ticks collected in the eastern Cape Province of South Africa. BMC Microbiol. 2017;17:1–8. doi: 10.1186/s12866-017-0955-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Rehman A, Conraths FJ, Sauter-Louis C, Krücken J, Nijhof AM. Epidemiology of tick-borne pathogens in the semi-arid and the arid agro-ecological zones of Punjab province, Pakistan. 2019. [DOI] [PubMed] [Google Scholar]
- 48.Li J, Liu X, Mu J, Yu X, Fei Y, Chang J, et al. Emergence of a novel ehrlichia minasensis strain, harboring the major immunogenic glycoprotein trp36 with unique tandem repeat and c-terminal region sequences, in haemaphysalis hystricis ticks removed from free-ranging sheep in Hainan Province, China. Microorganisms. 2019;7:396. doi: 10.3390/microorganisms7090369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Cabezas-Cruz A, Pollet T, Estrada-Peña A, Allain E, Bonnet SI, Moutailler S. Ticks and Tick-Borne Pathogens. 2019. Handling the Microbial Complexity Associated to Ticks; pp. 5–10. [Google Scholar]
- 50.Cabezas-Cruz A, Zweygarth E, Vancová M, Broniszewska M, Grubhoffer L, Passos LMF, et al. Ehrlichia minasensis sp. nov., isolated from the tick Rhipicephalus microplus. Int J Syst Evol Microbiol. 2016;66:1426–1430. doi: 10.1099/ijsem.0.000895. [DOI] [PubMed] [Google Scholar]
- 51.Iqbal N, Mukhtar MU, Yang J, Sajid MS, Niu Q, Guan G, et al. First molecular evidence of Anaplasma bovis and Anaplasma phagocytophilum in bovine from Central Punjab, Pakistan. Pathogens. 2019;8:155. doi: 10.3390/pathogens8030155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Ybañez AP, Sashika M, Inokuma H. The phylogenetic position of Anaplasma bovis and inferences on the phylogeny of the genus Anaplasma. J Vet Med Sci. 2014;76:307–312. doi: 10.1292/jvms.13-0411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Silaghi C, Santos AS, Gomes J, Christova I, Matei IA, Walder G, et al. Guidelines for the direct detection of Anaplasma spp. in diagnosis and epidemiological studies. Vector Borne Zoonotic Dis. 2017;17:12–22. doi: 10.1089/vbz.2016.1960. [DOI] [PubMed] [Google Scholar]
- 54.Ogden NH, Mechai S, Margos G. Changing geographic ranges of ticks and tick-borne pathogens: Drivers, mechanisms and consequences for pathogen diversity. Front Cell Infect Microbiol. 2013;4:1–11. doi: 10.3389/fcimb.2013.00046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Aguiar DM, Araujo JP, Nakazato L, Bard E, Cabezas-Cruz A. Complete genome sequence of an Ehrlichia minasensis strain isolated from cattle. Microbiol Resource Announc. 2019;8:18–20. doi: 10.1128/MRA.00161-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Kitaa JM. Clinical, haematological, treatment and molecular evaluation of Ehrlichial infections in dogs in Nairobi and its environs. Kenya: University of Nairobi; 2014. http://erepository.uonbi.ac.ke/handle/11295/76466. Accessed 4 Apr 2019.
- 57.Cabezas-Cruz A, Valdés JJ, De La Fuente J. The glycoprotein TRP36 of Ehrlichia sp. UFMG-EV and related cattle pathogen Ehrlichia sp. UFMT-BV evolved from a highly variable clade of E. canis under adaptive diversifying selection. Parasites and Vectors. 2014;7:1–7. doi: 10.1186/s13071-014-0584-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Nguhiu-Mwangi J, Aleri JW, Mogoa EG, Mbithi PM. Indicators of Poor Welfare in Dairy Cows Within Smallholder Zero-Grazing Units in the Peri-Urban Areas of Nairobi, Kenya. In: Insights from Veterinary Medicine: InTech; 2013. p. 50–87. 10.5772/53678.
- 59.Garry F. Rebhun’s Diseases of Dairy Cattle. W.B. Saunders. 2008. Miscellaneous infectious diseases; pp. 606–639. [Google Scholar]
- 60.Ye J, Coulouris G, Zaretskaya I, Cutcutache I, Rozen S, Madden TL. Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinform. 2012;13:134. doi: 10.1186/1471-2105-13-134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Schäffer AA, Aravind L, Madden TL, Shavirin S, Spouge JL, Wolf YI, et al. Improving the accuracy of PSI-BLAST protein database searches with composition-based statistics and other refinements. Nucleic Acids Res. 2001;29:2994–3005. doi: 10.1093/nar/29.14.2994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32:1792–1797. doi: 10.1093/nar/gkh340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Madeira F, Park YM, Lee J, Buso N, Gur T, Madhusoodanan N, et al. The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res. 2019;47:W636–W641. doi: 10.1093/nar/gkz268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Tamura K, Stecher G, Peterson D, Filipski A, Kumar S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol. 2013;30:2725–2729. doi: 10.1093/molbev/mst197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Tamura K, Nei M. Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol. 1993;10:512–526. doi: 10.1093/oxfordjournals.molbev.a040023. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Dataset used and analyzed during the current study are available from the corresponding author on reasonable request. Sequences obtained and analyzed from this study are available in GenBank database on the accession numbers indicated.