Abstract
Scalloped Hammerhead shark (Sphyrna lewini) is an endangered species which its populations have been declining globally including in Indonesia, the world’s top shark fishing country. However, there is a lack of information on the recent population structure of this species to promote proper management and its conservation status. This study aimed to investigate the genetic diversity, population structure, and connectivity of the S. lewini population, in three major shark landing sites: Aceh (n = 41), Balikpapan (n = 30), and Lombok (n = 29). Meanwhile, additional sequences were retrieved from West Papua (n = 14) and the Western Indian Ocean (n = 65) populations. From the analyses of the mitochondrial CO1 gene, a total of 179 sequences of S. lewini, with an average size of 594 bp, and 40 polymorphic loci in four and eight haplotypes for the Indonesian population and the Western Indian Ocean population were identified. The overall values of genetic diversity were high (h = 0.717; π = 0.013), with the highest values recorded in Aceh (h = 0.668; π = 0.002) and the lowest in Papua (h = 0.143; π = 0.000). On the contrary, the overall value was fairly low in the Western Indian Ocean (h = 0.232; π = 0.001). Furthermore, AMOVA and FST showed three significant subdivisions in Indonesia (FST = 0.442; P < 0.001), with separated populations for Aceh and West Papua, and mixed between Balikpapan and Lombok (FST = 0.044; P = 0.091). In contrast, genetic homogeneity was observed within the population of the Western Indian Ocean (FST = –0.013; P = 0.612). The establishment of a haplotype network provided evidence of a significantly different population and a limited genetic distribution between the Indonesian and the Western Indian Ocean populations (FST = 0.740; P < 0.001). This study showed the presence of a complex population of S. lewini with limited connectivity only in Indonesia separated from the Western Indian Ocean and requiring specific management measures based on the population structure at the regional level.
Introduction
The scalloped hammerhead shark, Sphyrna lewini is considered a coastal species because of its need for nursery areas. Globally, it is distributed in tropical waters as well as on the mainland, islands, and near the coastal region [1, 2]. This species has the unique modification of a lateral head which improves the ability to navigate and follow geomagnetic orientations across the ocean [3–5]. S. lewini can move in high rates of dispersal, and its female show allegiance to single nursery areas and exhibit no evidence of continued inter-oceanic migration. On the contrary, male is spread over a large area across ocean waters, with clear evidence of cross-reproduction and gamete transmission [6].
Scalloped hammerhead is one of the most exploited and threatened sharks. Around one to three million sharks are killed each year because of fishing and the shark fin trade around the world [7, 8]. This species was considered to be underexploited in 1999. However, in 2009 the International Union for Conservation of Nature (IUCN) listed the species on the Red List with Endangered (EN) status [9]. Five years later, the Convention on International Trade in Endangered Species (CITES) listed hammerhead sharks in Appendix II, and in 2019 the status was upgraded to Critically Endangered (CR) [10].
High exploitation of the S. lewini has an impact on its population structure, reducing the fecundity of the species and the genetic diversity [11]. Hammerhead sharks are viviparous with a yolk-sac placenta with an annual number of 12–30 young per litter. This species has a slow growth rate, late sexual maturity, long gestation period, and a long lifespan in nature [12–15]. The combination of high pressure and their biological properties makes this species vulnerable to overexploitation.
The study of population genetics has become an important tool for understanding population connectivity, supporting fisheries management, and improving conservation strategies. Furthermore, genetic information can be used to define the conservation effort and course of action by studying the structure of shark populations [16–19].
Shark fin product, including from scalloped hammerhead are very popular in Hong Kong [20], where trade regulations for endangered species and effective regulations are promoted [16, 20, 21]. The population structure of S. lewini, which is important for fisheries stock management, has been widely investigated in different coastal areas and ocean basins on a global and regional scale [17–19, 22, 23]. Duncan et al. [22] reported a global phylogeographic study of S. lewini that indicated that the Indo-West Pacific region is the center of diversity for tropical sharks, such as S. lewini, with a high and unique genetic diversity; however, no samples from Indonesia were included in that study, or in the study reported by Ovenden et al. [23], which included limited samples from Indonesia.
This study aimed to investigate the genetic diversity, population structure, and connectivity of S. lewini, where the populations of this species are affected by fishing activities at a regional scale in the Western Indian Ocean. Finally, the implications of these results for species management and conservation were examined.
Material and methods
Ethics statement
All samples were already dead when collected, and therefore, no approval from any institutional animal ethics committee was required. The sample collection and transportation followed the regulation of the Ministry of Marine Affairs and Fisheries of the Republic of Indonesia (Number 5/PERMEN-KP/2018) on the prohibition of cowboy and hammerhead shark export from Indonesia. Furthermore, this study was approved by the Ministry of Marine Affairs and Fisheries of the Republic of Indonesia, under permission numbers 276/BPSPL.03/PRL/X/2018 and 319/PNK/BPSPL.03/PK.230/REKOM/X/2018.
Tissue sample collection
From October 2017 to November 2018, a total of 100 tissue samples were obtained from S. lewini, including 41 from the fishing ports of Meulaboh and Aceh Jaya, 30 from a local shark landing in Manggar, and 29 from the fishing port of Tanjung Luar (Table 1). The samples (~0.5 cm3) were dissected and preserved in sample bottles containing 96% ethanol.
Table 1. Sampling collection sites at major shark landing sites in Indonesia.
| Site | Geographic coordinate | Number of samples |
|---|---|---|
| Aceh (ACH) | ||
| Meulaboh | N 04° 08’ 29” E 96° 07’ 55” | 33 |
| Aceh Jaya | N 04° 38’ 34” E 95° 34’ 58” | 8 |
| Balikpapan (BPN) | ||
| Manggar | S 01° 12’ 53” E 116° 58’ 24” | 30 |
| Lombok (LOM) | ||
| Tanjung Luar | S 08° 46’ 39” E 116° 31’ 01” | 29 |
| Total | 100 |
DNA extraction, amplification, and sequencing
DNA extraction was performed at the Biodiversity and Biosystematics Laboratory, IPB University, according to the protocol of the gSYNC DNA extraction kit product. A fragment of the mitochondrial cytochrome oxidase subunit 1 (CO1) gene was amplified using the forward primer fish-BCL (5'–TCA ACY AAT CAY AAA GAT ATY GGC AC–3′) and the reverse fish-BCH (5'–ACT TCY GGG TGR CCR AAR AAT CA–3′) [24, 25] in a 24 μL reaction mixture consisting of 3 μL of DNA template, 12.5 μL of MyTaq HS Red Mix, 9 μL of ddH2O, 1.25 μL each of forward and reverse primers. Meanwhile, the reaction mixture was processed in a polymerase chain reaction (PCR) on a thermocycler using modified cycling conditions [26, 27]: pre-denaturation at 94°C for 15 s. This process was followed by 38 cycles of denaturation at 94°C for 30 s, annealing at 50°C for 30 s, and extension at 72°C for 45 s; as well as a final extension at 72°C for 10 min. In addition, the amplicons were visualized by 1.5% agarose gel electrophoresis added with ethidium bromide at 100 V for 20 min. The gel was observed under UV light to identify bands showing the presence of DNA fragments. Sequencing was also performed using a machine with an optimized protocol of Sanger method [28].
All laboratory protocols on sampling and DNA identification methods were deposited in protocols.io platform with a digital object identifier (DOI) available at dx.doi.org/10.17504/protocols.io.bfwmjpc6.
Data analysis
Genetic diversity
Over 179 mitochondrial CO1 DNA sequences with an average length of 594 bp were edited and aligned using the ClustalW algorithm [29] implemented in MEGA 6.06 [30]. Genetic diversity parameters, such as the number of haplotypes and diversity (h) as well as nucleotide diversity (π), were calculated using the DNASp v6 [31] and Arlequin v.3.5 program [32]. Furthermore, additional CO1 sequencing data of S. lewini from West Papua retrieved from GenBank (Table 2) (n = 14) were reanalyzed. These days were obtained by Sembiring et al. [33], sequences from previous studies performed in India (n = 6) [34], the United Arab Emirates (n = 30) [35], and Madagascar (n = 29) [36] to assess the genetic diversity in Indonesian and Western Indian Ocean populations.
Table 2. Localities, the total number (n), and accession number of CO1 gene sequences of S. lewini from Aceh, Balikpapan, Lombok, and Western Papua (Indonesia), India, the United Arab Emirates, as well as Madagascar (Western Indian Ocean).
| Locality | n | Accession number | Author |
|---|---|---|---|
| Indonesia | |||
| Aceh | 41 | MT324149-156, MT324187-219 | This study |
| Balikpapan | 30 | MT324157-186 | This study |
| Lombok | 29 | MT324220-248 | This study |
| West Papua | 14 | KF590254-55, KF590271-76, KF793729, KF793738-42 | [33] |
| Western Indian Ocean | |||
| India | 6 | KF899746-51 | [34] |
| United Arab Emirates | 30 | KP177238-41, KP177241, KP177254, KP177262, KP177272, KP177285-99, KP177300-07 | [35] |
| Madagascar | 29 | HQ171735-47, HQ171761-76 | [36] |
Population structure
An analysis of molecular variance (AMOVA) and fixation index (FST) [37] was performed for three major groups: 1) within and among the four populations from Indonesia, 2) within and among the three populations from the Western Indian Ocean, and 3) comparison between populations from Indonesia and Western Indian Ocean using the Arlequin v.3.5 program (set up, 1000 permutations; significance level threshold, α = 0.05). These two analyses allowed the estimation of the overall extent of the genetic variation and differentiation level in Indonesia and the Western Indian Ocean. Furthermore, population differentiation and its significance between sampling sites were also calculated with pairwise estimates [38–40].
Genetic connectivity
A haplotype network was constructed with a median-joining method in the Network v5.1.1.0 program [41] for all haplotypes detected in Indonesia and the Western Indian Ocean. This network aimed to obtain haplotype connectivity following a broader spatial connection in the regional area of the Indian Ocean. The distribution of haplotypes for each location was also provided in a proper map to show the clear distributions and genetic connectivity among the populations.
Results
Genetic diversity
All sequences of S. lewini obtained were deposited in the BOLD System with the Barcode Index Number (BIN) registry of BOLD: AAA2403 and database of GenBank with accession numbers MT324149-248 (Table 2). A total of 179 sequences of 594 bp mitochondrial CO1 gene was obtained from three sampling sites (Aceh, Balikpapan, and Lombok). Meanwhile, additional sequences of samples from West Papua and the Western Indian Ocean region were used to generate a total of 11 haplotype variations with 40 polymorphic loci (Table 3).
Table 3. Forty polymorphic loci of 11 haplotypes from 179 sequences of the mitochondrial CO1 gene of S. lewini samples from four localities in Indonesia and three localities in the Western Indian Ocean region.
| Haplotypes | Locus Position | |||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 15 | 24 | 27 | 33 | 45 | 48 | 63 | 69 | 105 | 120 | 138 | 141 | 156 | 165 | 174 | 186 | 198 | 229 | 279 | 289 | 303 | 306 | 325 | 339 | 342 | 346 | 366 | 396 | 420 | 453 | 456 | 459 | 483 | 495 | 498 | 522 | 531 | 534 | 543 | |
| H1* | A | T | A | C | T | T | G | T | T | A | T | C | G | T | T | C | C | T | T | C | G | T | T | C | C | A | C | T | T | C | C | T | T | C | T | C | C | T | C | T |
| H2* | . | . | . | A | . | . | . | . | . | G | . | A | T | . | C | T | T | C | C | T | . | C | C | T | T | C | . | C | C | T | . | . | . | A | . | T | . | C | . | C |
| H3 | . | . | . | A | . | . | . | . | C | G | . | A | T | . | C | T | T | C | C | T | . | C | . | T | T | C | . | C | C | T | . | . | . | A | . | T | . | C | . | . |
| H4* | . | . | . | A | . | . | . | . | . | G | . | A | T | . | C | T | T | C | C | T | . | C | C | T | T | C | . | C | C | T | . | . | . | A | . | T | . | C | . | . |
| H5 | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | G | . | G | . | . | . | . | . |
| H6 | . | . | . | . | . | C | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . |
| H7 | . | . | C | . | . | . | . | . | . | . | . | . | . | G | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . |
| H8 | . | C | . | . | C | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . |
| H9 | . | . | . | . | . | . | . | . | . | . | G | . | . | . | . | . | . | G | . | T | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | G | . | |
| H10 | G | . | . | . | . | . | . | G | . | . | . | . | . | G | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . |
| H11 | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | T | . | . | . | T | . | . | . | C | . | T | . | . | . |
Notes:
Nucleobase at each position is given for H1 while those different are written for all other haplotypes. Nucleobases identical to H1 are indicated with dots (.)
*Three original haplotypes of the S. lewini populations were obtained from the areas of study (Aceh, Balikpapan, and Lombok) in Indonesia. The remaining haplotypes were reanalyzed from previous studies.
The comparison of the genetic diversity of S. lewini following haplotype and nucleotide diversity showed the presence of variation (Table 4). The haplotype diversity (h) among the samples obtained from Indonesia ranged from 0.143 to 0.668, while the nucleotide diversity (π) ranged from 0.000 to 0.020. The highest genetic diversity was observed for the samples from Aceh (h = 0.668; π = 0.020), followed by the Balikpapan population, which exhibited a lower haplotype and nucleotide diversity (h = 0.646; π = 0.002). On the contrary, the lowest genetic diversity was detected in West Papua (h = 0.143, π = 0.000). Similarly, the S. lewini population from Lombok exhibited a fairly low genetic diversity (h = 0.362; π = 0.001). However, the overall diversity in Indonesia was relatively high (h = 0.717; π = 0.013) since the average in the Western Indian Ocean region was low and ranged from 0.000 to 0.467. Therefore, it is reasonable to conclude that the overall diversity in the Western Indian Ocean region was also low (h = 0.232; π = 0.001).
Table 4. Genetic diversity of S. lewini based on sample size (n), haplotype number (Hn), haplotype diversity (h), and nucleotide diversity (π) in samples from each site in Indonesia and the Western Indian Ocean region.
| Population | n | Genetic Diversity | ||
|---|---|---|---|---|
| Hn | h | Π | ||
| Indonesia | ||||
| Aceh (ACH) | 41 | 4 | 0.668 | 0.020 |
| Balikpapan (BPN) | 30 | 3 | 0.646 | 0.002 |
| Lombok (LOM) | 29 | 3 | 0.362 | 0.001 |
| West Papua (WEP) | 14 | 2 | 0.143 | 0.000 |
| Overall Indonesia | 114 | 4 | 0.717 | 0.013 |
| Western Indian Ocean | ||||
| India (IND) | 6 | 1 | 0.000 | 0.000 |
| United Arab Emirates (UAE) | 30 | 8 | 0.467 | 0.002 |
| Madagascar (MDG) | 29 | 1 | 0.000 | 0.000 |
| Overall Western Indian Ocean | 65 | 8 | 0.232 | 0.001 |
Population structure
The analysis of the fixation index (FST) and the corresponding P-values between and within the four S. lewini populations (ACH, BPN, LOM, and WEP) from Indonesia and three populations (IND, UAE, and MDG) from the Western Indian Ocean region are shown in Table 5. The overall FST value in Indonesia was significantly higher than that observed in other regions (FST = 0.442; P < 0.001) due to the presence of multiple subdivisions.
Table 5. Analysis of molecular variance (AMOVA) for the percentage of variation (%), FST value, and significance level (P-value) in S. lewini samples from Indonesian, the Western Indian Ocean, and between Indonesian and Western Indian Ocean populations.
| Source of variation | df | Percentage of variation (%) | FST value | P-value |
|---|---|---|---|---|
| Indonesia | ||||
| Among Populations | 3 | 44.15 | 0.442 | 0.000 |
| Within Populations | 110 | 55.85 | ||
| Total | 113 | |||
| Western Indian Ocean | ||||
| Among Populations | 2 | −1.31 | −0.013 | 0.612 |
| Within Populations | 62 | 101.31 | ||
| Total | 64 | |||
| Indonesia vs. Western Indian Ocean | ||||
| Among Populations | 1 | 74.04 | 0.740 | 0.000 |
| Within Populations | 177 | 25.96 | ||
| Total | 178 |
A genetic homogeneity was observed in the population from the Western Indian Ocean region (FST = –0.013; P = 0.612). However, a comparison of the population structure between Indonesia and the Western Indian Ocean region yielded significant differentiation (FST = 0.740; P < 0.001). The pairwise FST values between the populations from the four locations in Indonesia and the Western Indian Ocean population are shown in Table 6. Furthermore, the overall pairwise analysis through the distance method showed the presence of significant differentiation among the four populations. In contrast, BPN and LOM (FST = 0.044; P = 0.091) showed fairly low FST values and no significant P-values. Furthermore, among the populations from Indonesia, the ACH showed a trend of being closer to the Western Indian Ocean since it exhibited a lower FST and significant P-value (FST = 0.509; P < 0.001).
Table 6. Pairwise FST values (below the diagonal) and P-values (above the diagonal) between the S. lewini populations from Aceh (ACH), Balikpapan (BPN), Lombok (LOM), West Papua (WEP), and Western Indian Ocean (WIO).
| Sample Sites | ACH | BPN | LOM | WEP | WIO |
| ACH | - | 0.000 | 0.000 | 0.000 | 0.000 |
| BPN | 0.427 | - | 0.091 | 0.000 | 0.000 |
| LOM | 0.438 | 0.044 | - | 0.000 | 0.000 |
| WEP | 0.398 | 0.495 | 0.736 | - | 0.000 |
| WIO | 0.509 | 0.965 | 0.973 | 0.975 | - |
Genetic connectivity
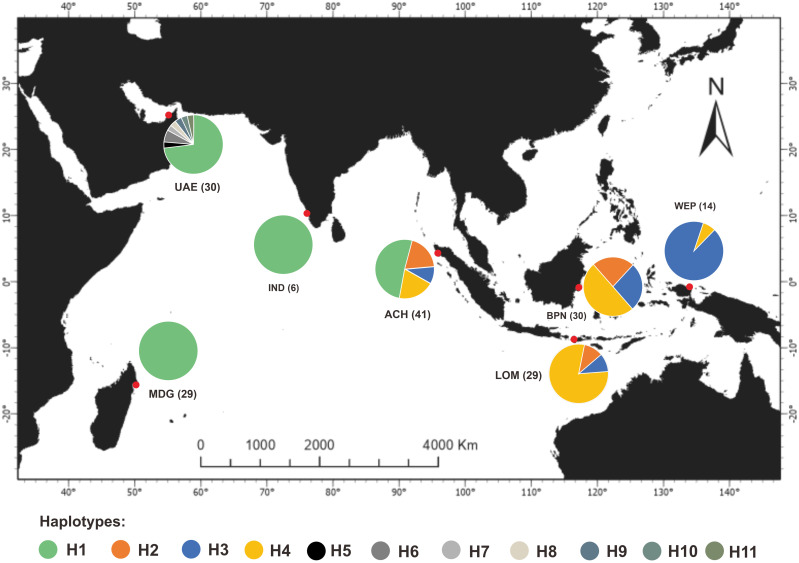
Network analysis of the haplotype identified two main groups of haplotypes (Fig 1) referred to as clade A and B. Clade A consisted of haplotype H1, which was observed in several regions, i.e., Aceh, India, United Arab Emirates, and Madagascar, while H5, H6, H7, H8, H9, H10, and H11 were present only in samples from the United Arab Emirates. Clade B consisted of three haplotypes (H2, H3, and H4), which were spread evenly in Indonesia. Haplotypes H1, H3, and H4 were predominant in Aceh, West Papua, and Balikpapan-Lombok, respectively (Fig 2).
Fig 1. Haplotype network of the S. lewini (n = 179) population from Indonesia and the Western Indian Ocean region, which was constructed using the median joining method.
Fig 2. Distribution of the 11 haplotypes of the S. lewini population from Indonesia and Western Indian Ocean at the regional scale.
Discussion
Genetic diversity
The overall genetic diversity of S. lewini at the haplotype and nucleotide levels was relatively high for populations in Indonesia. These findings are consistent with the results reported by Ovenden et al. [23] regarding the mitochondria control region from three localities in Indonesia [22]. The scalloped hammerhead sharks are a highly migratory species with a wide distribution in tropical and warm-temperate waters. This specie can move across oceanic waters to a distance of up to 1671 km [12]. Due to its migratory ability and broad ecological niches, this species tends to have higher genetic diversity than others [42]. Generally, high levels of genetic diversity are associated with large population size [43] and are promoted by several factors, such as local population sizes, fast generation times [44], high nucleotide substitution rates [45], and high gene flow between geographically distant populations.
The finding of a relatively high genetic diversity for S. lewini appears to be inconsistent on the assumption that overexploitation of this species as both a target of fishing and bycatch led to the decline of its populations on a global scale [46]. However, the results obtained from Lombok may be relevant since the lower genetic diversity detected was probably driven by continuous fishing pressure. S. lewini species are the top three targeted sharks at the Tanjung Luar fishing port in Lombok and have faced high fishing pressure over more than 40 years with recent exploitation rates (E) reaching 0.59 [47].
Furthermore, according to the global fisheries information system on a global scale and Indonesia by FAO [48], the hammerhead sharks (Sphyrnidae), including S. lewini, are very important. These species were highly exploited in the last two decades, with an estimated rapid increase in global capture, from 220 tons per year in 1985 up to approximately 10,362 in 2016. During the same period, the capture level also increased significantly, reaching approximately 1,492 tons in 2016. Meanwhile, Indonesia recorded one of the highest numbers of sharks and rays caught on the global catches reported in 2000–2011 [49].
Clarke et al. [50] reported similar findings regarding the mitochondrial DNA of the silky shark, Carcharhinus falciformis, which exhibited a high genetic diversity under circumstances of overexploitation. Elasmobranchs exhibit adaptability to environmental and anthropogenic stresses, which causes genetic bottlenecks because of their particular life histories [8]. However, the population decline caused by recent fishery activities might be insufficient to reduce genetic diversity, particularly for species with a long life span (13–20 years), such as S. lewini [51, 52].
The persistent decline, as predicted for the Lombok population of S. lewini, correlated positively with the loss of genetic diversity and created a bottleneck [52, 53], as reported by Pinsky and Palumbi in a meta-analysis of several marine fish [54].
Population structure
The obtained results showed the presence of homogeneity among the S. lewini populations from Balikpapan and Lombok. The pairwise FST analysis detected no significant genetic differentiation in these two populations with the lowest value. These findings complement previous studies conducted in Indo-Australian waters. Similarly, Ovenden et al. [23] reported evidence for the mitochondrial control region regarding the structure, with no differentiation between two populations of S. lewini (Lombok and northern Australia). This pattern of single-stock population suggests that these localities are a migration zone of S. lewini and a reproductive movement may occur in their coastal areas. However, there are strong Indonesian through flow currents between Kalimantan and Sulawesi Island. Adult S. lewini specimens are highly migratory, with a large body supporting the high dispersal ability of this species. Consequently, they show the possibility of overcoming that geographical barrier. In contrast, different results from the comparison among these two populations from central Indonesia and one from Aceh (western Indonesia) as well as from West Papua (eastern Indonesia), which exhibited strong genetic differentiation were obtained. Furthermore, these regions were spatially separated by a long distance, with the possibility of the existence of more complex barriers. These barriers can be inter-island or anthropogenic factors on a high commercial and artisanal fishing pressure along the southern and northern coasts of Java.
Genetic connectivity
Regarding the significant value of FST among the population (FST = 0.740; P < 0.001) as well as haplotype network and distribution shown in Figs 1 and 2, two restricted haplogroups which separated the S. lewini population in Indonesian with the Western Indian Ocean population were observed. The two haplogroups were separated with 19 different nucleotide bases due to monomorphic and polymorphic mutations. Furthermore, limited gene flow that occurs among populations in Indonesia forms a different pool with the Western Indian Ocean population. However, this was expected because of the complex geographic barrier in Indonesia’s marine ecosystem, and the global distribution pattern of S. lewini, with significant separation population across ocean basin as well as discontinuous coastline habitat [6].
Generally, S. lewini populations from Balikpapan, Lombok, and West Papua appear to be isolated from the Western Indian Ocean and shared a haplotype network exclusively only in eastern Indonesian waters. However, an interesting result regarding genetic sharing between the populations from Aceh and the Indian Ocean population was obtained. H1 is a unique haplotype that was only obtained in Aceh. However, the FST value observed significant differences between the population in Aceh and the Western Indian Ocean, and there was an indication of genetic sharing between those localities (FST = 0.509; P < 0.001). The similarity between the predominant haplotype (H1) of S. lewini from Aceh and that of the populations from India, the United Arab Emirates as well as Madagascar reflected a genetic sharing process in the Indian Ocean region. This showed the presence of past historical gene flow between the populations in spatially separated regions driven by ancestral interaction [17]. However, recent studies reported that the scalloped hammerhead demonstrated a strong differentiation in population structure across ocean basins e.g. Indian Ocean and discontinuous continental coastlines, as shown by the separation between Aceh and Indian coastline [6, 22].
Conservation implications
The high diversity of the S. lewini populations in Indonesia shows that this species has not experienced a genetic loss because of exploitation pressure. However, the lower genetic diversity of S. lewini from Lombok and West Papua showed a higher risk of loss, which probably was the result of high fisheries pressure. Furthermore, the genetic assessment of S. lewini samples from four localities showed that a single stock exists between Lombok and Balikpapan. On the contrary, a separate stock was observed for Aceh and West Papua, showing that the management of this species should occur on a stock-based approach at least on three mitochondrial-stock conservation units. The complex population of S. lewini with limited connectivity observed in Indonesia and the Western Indian Ocean region demonstrated the importance of promoting specific collaborative management strategies among Indonesian, and in conjunction with Western Indian Ocean agencies at the regional scale.
Conclusion
This study provided important findings on the population structure of S. lewini in Indonesia, with a high genetic diversity and three significant subdivisions. The results showed the capability of the population to adapt to rapid environmental changes and pressure, including fishing activities. In addition, the lower genetic diversity in Lombok and West Papua was also considered. The restricted genetic sharing detected among the species obtained from Indonesia showed unique features among these populations. Therefore, a specific collaborative action across regions is needed to promote sustainable management and conservation purposes, both in Indonesia and at the regional scale in the Western Indian Ocean area.
Acknowledgments
The authors are grateful to the institutions and individuals that have made the study possible: colleagues from Marine Biodiversity and Biosystematics Lab (BIODIVSI), Faculty of Marine Science and Technology, IPB University for their help during this study. The authors are further grateful to BPSPL Satker Balikpapan, Listian Nova, and Mr. Hery, Head of TPI Manggar, Balikpapan for their help during the sample’s collection, and to reviewers and proofreaders that have provided highly constructive suggestions for better writing of the study.
Data Availability
All relevant data are within the paper. All sequences of S. lewini obtained were deposited in the BOLD System with the Barcode Index Number (BIN) registry of BOLD: AAA2403 and database of GenBank with accession numbers MT324149-248.
Funding Statement
This research was supported by Wildlife Conservation Society (WCS)-Indonesia, in collaboration with Faculty of Fisheries and Marine Sciences IPB University. SH received a scholarship from Indonesia Endowment Fund for Education (LPDP). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Klimley AP, Butler SB. Immigration and emigration of a pelagic fish assemblage to seamounts in the Gulf of California related to water mass movements using satellite imagery. Mar Ecol Prog Ser. 1988;49: 11–20. 10.3354/meps049011 [DOI] [Google Scholar]
- 2.Compagno LJV. FAO Species Catalogue: Sharks of the world. Rome: Food and Agriculture Organization;1984. [Google Scholar]
- 3.Montgomery JC, Walker MM. Orientation and navigation in elasmobranchs: which way forward? Environ Biol Fishes. 2001;60: 109–116. [Google Scholar]
- 4.Kajiura SM, Holland KN. Electroreception in juvenile scalloped hammerhead and sandbar sharks. J Exp Biol. 2002;205: 3609–3621. [DOI] [PubMed] [Google Scholar]
- 5.Meyer CL, Holland KN, Papastamatiou YP. Sharks can detect changes in the geomagnetic field. J R Soc Interface. 2005;2: 129–130. 10.1098/rsif.2004.0021 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Daly-Engel TS, Seraphin KD, Holland KN, Coffey JP, Nance HA, Toonen RJ, et al. Global phylogeography with mixed-marker analysis reveals male-mediated dispersal in the endangered scalloped hammerhead shark (Sphyrna lewini). PLoS ONE. 2012;7(1): e29986 10.1371/journal.pone.0029986 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Clarke SC, McAllister MK, Milner-Gulland EJ, Kirkwoo GP, Michielsens CGJ, Agnew DJ, et al. Global estimates of shark catches using trade records from commercial markets. Ecol Lett. 2006;9(10): 1115–1126. 10.1111/j.1461-0248.2006.00968.x [DOI] [PubMed] [Google Scholar]
- 8.Chapman DD, Simpfendorfer CA, Wiley TR, Poulakis GR, Curtis C, Tringali M, et al. Genetic diversity despite population collapse in a critically endangered marine fish: the smalltooth sawfish (Pristis pectinata). J Hered. 2011;102: 643–652. 10.1093/jhered/esr098 [DOI] [PubMed] [Google Scholar]
- 9.Baum J, Clarke S, Domingo A, Ducrocq M, Lamónaca AF, Gaibor N, et al. Sphyrna lewini. The IUCN Red List of Threatened Species 2009. 2009; e.T39385A10190088.
- 10.Rigby CL, Dulvy NK, Barreto R, Carlson J, Fernando D, Fordham S, et al. Sphyrna lewini. The IUCN Red List of Threatened Species 2019. 2019; e.T39385A2918526.
- 11.Ward RD. Genetics in fisheries management. Hidrobiologia. 2000;420: 191–201. [Google Scholar]
- 12.Kohler NE, Turner PA. Shark tagging: a review of conventional methods and studies. Environ Biol Fishes. 2001;60: 191–223. [Google Scholar]
- 13.White W, Barton C, Potter I. Catch composition and reproductive biology of Sphyrna lewini (Griffith & Smith) (Charcharhinifirnes, Sphyrnidae) in Indonesian waters. J Fish Biol. 2008;72: 1675–1689. [Google Scholar]
- 14.Ovenden J, Morgan JT, Street R, Tobin A, Simfendorfer CA, Macbeth W, et al. Negligible evidence for regional genetic population structure for two shark species Rhizoprionodon acutus (Rüppell, 1837) and Sphyrna lewini (Griffith & Smith, 1834) with contrasting biology. Mar Biol. 2011;158: 1497–1509. 10.1007/s00227-011-1666-y [DOI] [Google Scholar]
- 15.Bessudo S, Soler GA, Kimley PA, Ketchum J, Arauz R, Hearn A, et al. Vertical and horizontal movements of scalloped hamemerhead shark (Sphyrna lewini) around Malpelo and Cocos islands (Tropical Eastern Pacific) using satellite telemetry. Boletín de Investigaciones Marinas y Costeras. 2011;40: 91–106. [Google Scholar]
- 16.Cardeñosa D, Hyde J, Caballero S. Genetic diversity and population structure of the pelagic thresher shark (Alopias pelagicus) in the Pacific Ocean: evidence for two evolutionarily significant units. PLoS ONE. 2014;9(10): e110193 10.1371/journal.pone.0110193 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Castillo-olguin E, Aribe-Alcocer M, Diaz-Jaimes P. Assessment of the population genetic structure of Sphyrna lewini to identify conservation units in the Mexican Pacific. Ciencias Marinas. 2012;38(4): 635–652. 10.7773/cm.v38i4.2110 [DOI] [Google Scholar]
- 18.Quintanilla S, Gómez A, Mariño-Ramírez C, Sorzano C, Bessudo S, Soler G, et al. Conservation genetics of the scalloped hammerhead shark in the Pacific Coast of Colombia. J Hered. 2015;106: 448–458. 10.1093/jhered/esv050 [DOI] [PubMed] [Google Scholar]
- 19.Sukumaran S, Sebastian W, Mukundan LP, Menon M, Akhilesh K V., Zacharia PU, et al. Molecular analyses reveal a lack of genetic structuring in the scalloped hammerhead shark, Sphyrna lewini (Griffith & Smith, 1834) along the Indian coast. Mar Biodivers. 2020;50 10.1007/s12526-020-01040-4 [DOI] [Google Scholar]
- 20.Fields AT, Fischer GA, Shea SKH, Zhang H, Feldheim KA, Chapman DD. DNA Zip‐coding: identifying the source populations supplying the international trade of a critically endangered coastal shark. Animal Conservation. 2020; 10.1111/acv.12585 [DOI] [Google Scholar]
- 21.Jabado RW, Al Ghais SM, Hamza W, Henderson AC, Spaet JLY, Shivji MS, et al. The trade in sharks and their products in the United Arab Emirates. Biol Conserv. 2015;181: 190–198. 10.1016/j.biocon.2014.10.032 [DOI] [Google Scholar]
- 22.Duncan KM, Martin AP, Bowen BW, De Coute HG. Global phylogeography of the scalloped hammerhead shark (Sphyrna lewini). Mol Ecol. 2006;15:2239–51. 10.1111/j.1365-294X.2006.02933.x [DOI] [PubMed] [Google Scholar]
- 23.Ovenden JR, Kashiwagi T, Broderick D, Giles J, Salini J. The extent of population genetic subdivision differs among four co-distributed shark species in the Indo-Australian archipelago. BMC Evol Biol. 2009;9: 40 10.1186/1471-2148-9-40 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Baldwin CC, Mounts JH, Smith DG, Weigt LA. Genetic identification and color descriptions of early life-history stages of Belizean Phaeoptyx and Astrapogon (Teleostei: Apogonidae) with comments on identification of adult Phaeoptyx. Zootaxa. 2009;2008: 1–22. [Google Scholar]
- 25.Madduppa H, Ayuningtyas RU, Subhan B, Arafat D. Exploited but unevaluated: DNA barcoding reveals skates and stingrays (Chordata, Chondrichthyes) species landed in the Indonesian fish market. IJMS. 2016;21: 77–84. [Google Scholar]
- 26.Prehadi, Sembiring A, Kurniasih EM, Rahmad, Arafat D, Subhan B, et al. DNA barcoding and phylogenetic reconstruction of shark species landed in Muncar fisheries landing site in comparison with Southern Java fishing port. Biodiversitas. 2015;16: 55–61. [Google Scholar]
- 27.Hadi S, Anggraini NP, Muttaqin E, Simeon BM, Subhan B, Madduppa H. Genetic diversity of the endangered species Sphyrna lewini (Griffith and Smith 1834) in Lombok based on mitochondrial DNA. IOP Conf. Ser.: Earth Environ. Sci. 2019;236: 012024. [Google Scholar]
- 28.Sanger F, Nicklen S, Coulson AR. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci. 1977;74(12): 5463–5467. 10.1073/pnas.74.12.5463 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Thompson JD, Higgins DG, Gibson TJ. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994;22(22): 4673–4680. 10.1093/nar/22.22.4673 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Tamura K, Stecher G, Peterson D, Filipski A, Kumar S. Molecular Evolutionary Genetics Analysis Version 6.0. Mol Biol Evol. 2013;30(12): 2725–2729. 10.1093/molbev/mst197 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Rozas J, Ferrer-Mata A, Sánchez-DelBarriol JC, Guirao-Rico S, Librado P, Ramos-Onsins SE. DnaSP v6: DNA Sequence polymorphism analysis of large datasets. Mol Biol Evol. 2017;34: 3299–3302. 10.1093/molbev/msx248 [DOI] [PubMed] [Google Scholar]
- 32.Excoffier L, Lischer H.E. L. Arlequin suite ver 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour. 2010;10: 564–567. 10.1111/j.1755-0998.2010.02847.x [DOI] [PubMed] [Google Scholar]
- 33.Sembiring A, Pertiwi NPD, Mahardini A, Wulandari R, Kurniasih EM, Kuncoro AW, et al. DNA barcoding reveals targeted fisheries for endangered sharks in Indonesia. Fisheries Research. 2015;164: 130–134. 10.1016/j.fishres.2014.11.003 [DOI] [Google Scholar]
- 34.Bineesh KK, Gopalakrishnan A, Akhilesh KV, Sajeela KA, Abdussamad EM, Pillai NGK, et al. DNA barcoding reveals species composition of sharks and rays in the Indian commercial fishery. Mitochondrial DNA. 2016. 10.3109/19401736.2015.1137900 [DOI] [PubMed] [Google Scholar]
- 35.Jabado RW, Ghais SMA, Hamza W, Henderson AC, Spaet JLY, Shivji MS, et al. The trade in sharks and their products in the United Arab Emirates. Biol Conserv. 2015;181: 190–198. 10.1016/j.biocon.2014.10.032 [DOI] [Google Scholar]
- 36.Doukakis P, Hanner R, Shivji M, Bartholomew C, Chapman D, Wong E, et al. Applying Genetic Techniques to Study Remote Shark Fisheries in Northeastern Madagascar. Mitochondrial DNA. 2011;22(S1): 15–20. [DOI] [PubMed] [Google Scholar]
- 37.Wright S. The genetical structure of populations. Annals of Eugenics. 1951;15: 323–354. 10.1111/j.1469-1809.1949.tb02451.x [DOI] [PubMed] [Google Scholar]
- 38.Weir BS, Cockerham CC. Estimating F-Statistics for the Analysis of Population Structure. Evol. 1984;38(6): 1358 10.2307/2408641 [DOI] [PubMed] [Google Scholar]
- 39.Excoffier L, Smouse PE, Quattro JM. Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics. 1992;131(2): 479‐491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Weir BS. Genetic Data Analysis II: Methods for Discrete Population Genetic Data. Sunderland, Mass: Sinauer Associates; 1996. [Google Scholar]
- 41.Bandelt H-J, Forster P, Röhl A. Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol. 1999;16: 37–48. 10.1093/oxfordjournals.molbev.a026036 [DOI] [PubMed] [Google Scholar]
- 42.Habel JC, Schmitt T. Vanishing of the common species: Empty habitats and the role of genetic diversity. Biol. Conserv. 1996;218: 211–216. 10.1016/j.biocon.2017.12.018 [DOI] [Google Scholar]
- 43.Frankham R. Relationship of Genetic Variation to Population Size in Wildlife. Conserv. Biol. 1996;10(6): 1500–1508. 10.1046/j.1523-1739.1996.10061500.x [DOI] [Google Scholar]
- 44.Hague MTJ, Routman EJ. Does population size affect genetic diversity? A test with sympatric lizard species. Heredity. 2016;116: 92–98. 10.1038/hdy.2015.76 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Martin AP, Naylor GJP, Palumbi SR. Rates of mitochondrial DNA evolution in sharks are slow compared with mammals. Nature. 1992;357: 153–157. 10.1038/357153a0 [DOI] [PubMed] [Google Scholar]
- 46.Compagno LJV, Dando M, Fowler SL. Sharks of the World. Princeton: Princeton University Press; 2005. [Google Scholar]
- 47.Simeon BM, Agustina S, Muttaqin E, Yulianto I, Ichsan M, Muhsin M. Laporan teknis: Pemanfaatan Hasil Tangkap Hiu dan Pari di Tanjung Luar, Provinsi Nusa Tenggara Barat. Bogor: Wildlife Conservation Society-Indonesia Program; 2017.
- 48.Food and Agriculture Organization (FAO). Fisheries and Aquaculture Information and Statistics Service: Global Capture Production 1950–2017; Database: Indonesian hammerhead shark capture 2017 [cited 2020 Mar 5]. http://www.fao.org/fishery/statistics/global-capture-production/query/es.
- 49.Clarke Dent F. State of the Global Market for Shark Products. Rome: Food and Agriculture Organization; 2015. [Google Scholar]
- 50.Clarke CR, Karl SA, Horn RL, Bernard AM, Lea JS, Hazin FH, et al. Global mitochondrial DNA phylogeography and population structure of the silky shark, Carcharhinus falciformis. Mar Biol. 2015;162(5): 945–955. 10.1007/s00227-015-2636-6 [DOI] [Google Scholar]
- 51.Sánchez-de Ita JA, Quiñónez-Velázquez C, Galván-Magaña F, Bocanegra-Castillo N, Félix-Uraga R. Age and growth of the silky shark Carcharhinus falciformis from the west coast of Baja California Sur, Mexico. J App Ichthyol. 2011;27: 20–24. [Google Scholar]
- 52.Nei M, Maruyama T, Chakraborty R. The bottleneck effect and genetic variability in populations. Evolution. 1975;29(1): 1 10.1111/j.1558-5646.1975.tb00807.x [DOI] [PubMed] [Google Scholar]
- 53.Beebee T, Rowe G. An Introduction to Molecular Ecology. 2nd ed New York: Oxford University Press Inc; 2008. [Google Scholar]
- 54.Pinsky ML, Palumbi SR. Meta-analysis reveals lower genetic diversity in overfished populations. Mol Ecol. 2013;23(1): 29–39. 10.1111/mec.12509 [DOI] [PubMed] [Google Scholar]