SUMMARY
Natural killer (NK) cells play a critical role in controlling viral infections, coordinating the response of innate and adaptive immune systems. They also possess certain features of adaptive lymphocytes, such as undergoing clonal proliferation. However, it is not known whether this adaptive NK cell response can be modulated by other lymphocytes during viral exposure. Here, we show that the clonal expansion of NK cells during mouse cytomegalovirus infection is severely blunted in the absence of cytotoxic CD8+ T cells. This correlates with higher viral burden and an increased pro-inflammatory milieu, which maintains NK cells in a hyper-activated state. Antiviral therapy rescues NK cell expansion in the absence of CD8+ T cells, suggesting that high viral loads have detrimental effects on adaptive NK cell responses. Altogether, our data support a mechanism whereby cytotoxic innate and adaptive lymphocytes cooperate to ensure viral clearance and the establishment of robust clonal NK cell responses.
In Brief
NK cells undergo clonal proliferation during certain viral infections, similar to CD8+ T cells. However, the interdependence of NK and CD8+ T cell expansion remained unclear. Here, Diaz-Salazar and Sun show that CD8+ T cells promote NK cell expansion by modulating the degree and duration of viremia and host inflammation during MCMV infection.
Graphical Abstract

INTRODUCTION
Natural killer (NK) cells are lymphocytes that provide protection against intracellular pathogens and viruses, especially herpesviruses such as cytomegalovirus (CMV) (Biron et al., 1989; Bukowski et al., 1985). NK cells recognize and clear infected cells through a balance of germline-encoded inhibitory and activating receptors, which can bind both “self” and “non-self” ligands (Lanier, 2008; Yokoyama et al., 2004). Upon activation, NK cells release cytokines (including interferon γ [IFN-γ]) and cytotoxic molecules (such as perforin and granzymes) to lyse infected or stressed cells (Abel et al., 2018; Vivier et al., 2008). Given the tight balance of activating and inhibitory signals, NK cells play a pivotal role in coordinating the response of innate and adaptive immunity, often bridging these two host immune responses.
NK cells can communicate with and regulate a plethora of neighboring immune cells within their microenvironment. For example, NK cell production of IFN-γ is critical for the complete elimination of intracellular bacteria by myeloid cells (Dunn and North, 1991; Feng et al., 2006; Steinhoff et al., 1991). NK cells can also control macrophage polarization in adipose tissue (Lee et al., 2016; O’Sullivan et al., 2016), recruit dendritic cells (DCs) to tumors (Barry et al., 2018; Böttcher et al., 2018), and promote neutrophil infiltration into inflamed joints (Louis et al., 2020). On the other hand, NK cells have also been shown to modulate CD4+ T and CD8+ T cell responses in a variety of contexts, ranging from directly pruning activated CD4+ T cells to preventing exaggerated CD8+ T cell responses (Lang et al., 2012; Schuster et al., 2014; Waggoner et al., 2011). Whether adaptive lymphocytes can reciprocally regulate NK cell responses in inflammatory settings has been less studied.
Traditionally considered part of the innate immune system, NK cells are now appreciated to have features of adaptive lymphocytes, such as clonal expansion and maintenance of a pool of long-lived “memory” cells (Dokun et al., 2001; Geary and Sun, 2017; Gumá et al., 2004; Lopez-Vergès et al., 2011; Reeves et al., 2015; Sun et al., 2009; Vivier et al., 2011). In the case of mouse cytomegalovirus (MCMV), these adaptive responses are driven by a subset of NK cells expressing the activating receptor Ly49H, which recognizes and binds the viral glycoprotein m157 (Arase et al., 2002; Sun et al., 2009). For productive adaptive responses to occur, NK cells require receptor ligation and sensing of pro-inflammatory cytokines, mainly IL-12 and type I interferons (IFN) (Geary et al., 2018; Madera et al., 2016; Madera and Sun, 2015; Sun et al., 2012). However, although adaptive NK cell responses share similar clonal expansion kinetics with antigen-specific T cells, whether these adaptive lymphocytes directly impact the NK cell response to viral infections remains largely unexplored. In this study, we sought to investigate whether adaptive lymphocytes play a role in Ly49H+ NK cell expansion and determine the underlying mechanisms coordinating lymphocyte responses during MCMV infection.
RESULTS AND DISCUSSION
Optimal NK Cell Expansion Requires an Adaptive Immune System
To investigate the role of T and B cells in adaptive NK cell responses against MCMV infection, we compared Ly49H+ NK cell expansion in wild-type (WT) mice versus Rag2−/− mice, which lack adaptive lymphocytes. Whereas WT mice exhibited a robust NK cell response, peaking ~7 days post-infection (PI), Rag2−/− mice showed a diminished response, with significantly lower numbers of circulating Ly49H+ NK cells (Figure S1A). Since a history of Rag activity has been shown to modulate NK cell fitness (Karo et al., 2014), we assessed whether NK cells raised in a WT environment still had an expansion defect in the absence of adaptive lymphocytes. To do so, we transferred purified WT NK cells into Ly49h−/− × Rag2−/− mice or Ly49h−/− mice, both of which lack the activating receptor Ly49H, the main driver of adaptive NK cell response against MCMV. (However, whereas Ly49h−/− × Rag2−/− mice also lack adaptive lymphocytes, Ly49h−/− mice have a full lymphocyte compartment.) Compared to Ly49h−/− control mice, in which adoptively transferred Ly49H+ NK cells expanded robustly, the same cells transferred into Ly49h−/− × Rag2−/− mice failed to expand to the same degree, showing ~10-fold lower expansion (Figure 1A), suggesting that the adaptive immune system plays a critical role in promoting robust NK cell expansion.
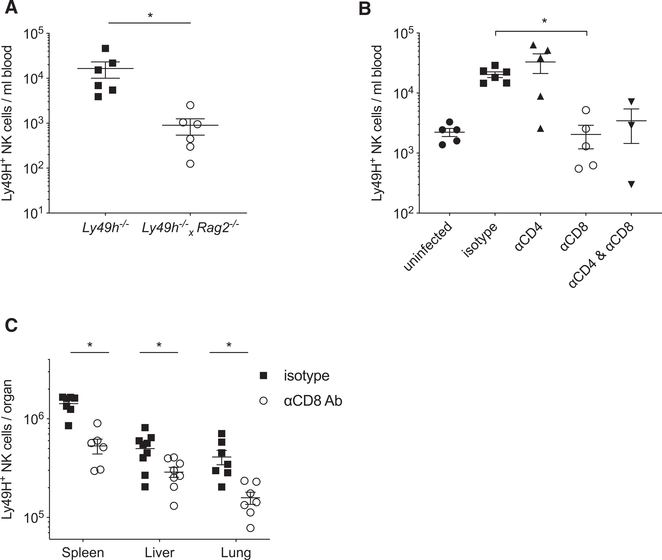
Figure 1. The Adaptive Immune System Is Required for Optimal NK Cell Expansion.
(A) 4 × 105 purified NK cells were transferred into congenically distinct Ly49H-deficient (Ly49h−/−) mice, or T-cell- and B-cell-deficient mice crossed with Ly49H-deficient mice (Ly49h−/− × Rag2−/−). One day after transfer, mice were challenged with MCMV, and the number of transferred Ly49H+ NK cells was quantified in peripheral blood 7 days post-infection (PI). Data are pooled from two independent experiments with three mice per group.
(B) Approximately 1.5 × 105 splenic NK cells were transferred into congenically distinct Ly49h−/− mice, treated with αCD8-depleting antibody, αCD4-depleting antibody, both, or isotype control and challenged with MCMV 1 day after transfer. The number of transferred Ly49H+ NK cells was quantified in peripheral blood at day 7 PI. Data are pooled from two independent experiments with three mice per group.
(C) WT mice treated with αCD8-depleting antibody or isotype control antibody were challenged with MCMV, and the number of endogenous Ly49H+ NK cells was quantified in various peripheral organs at day 6 PI. Data are pooled from two independent experiments with three or four mice per group.
Graphs present individual data points as scatter dot plots, and error bars show mean ± SEM. Statistical differences were calculated with a two-way ANOVA (A and C) and a one-way ANOVA (B). A p value of <0.05 was used as the significance cut-off and is indicated with a single asterisk (*). See also Figure S1.
To further dissect the specific cellular component of the adaptive immune system that impacts NK cell expansion, we treated recipient Ly49h−/− mice with depleting antibodies against CD4+ T cells, CD8+ T cells, or both and assessed transferred Ly49H+ NK cell numbers in peripheral blood following MCMV infection. Whereas depletion of the CD4+ T cell compartment had no effect on circulating NK cell numbers, depleting CD8+ T cells had a profound effect on NK cell expansion (Figure 1B). Depletion of both CD4+ and CD8+ T cell compartments had the same effect as depletion of CD8+ T cells alone (Figure 1B). A similar result was observed in a direct infection setting, in which the numbers of endogenous Ly49H+ NK cells were severely reduced when WT mice were depleted of CD8+ T cells (Figure S1B). Similar to the peripheral blood, we observed consistently lower Ly49H+ NK cell numbers in both lymphoid and non-lymphoid tissues such as the spleen, liver, and lung (Figure 1C), suggesting that CD8+ T cells are required for general NK cell expansion in an organ-independent manner.
Cytotoxic CD8+ T Cells Are Required and Sufficient to Promote NK Cell Expansion
We next investigated the mechanisms by which CD8+ T cells promote NK cell expansion. Adoptive transfer of purified CD8+ T cells into Rag2−/− mice was sufficient to rescue the expansion defect observed in the endogenous Ly49H+ NK cell population (Figure 2A). Interestingly, we found a strong positive correlation between the amount of Ly49H+ NK cell expansion and the number of reconstituted CD8+ T cells in these experiments (Figure 2B), as there is some reconstitution variability in individual mice (by nature of the experiment), suggesting CD8+ T cells are sufficient and required to ensure strong adaptive NK cell responses during viral infection.
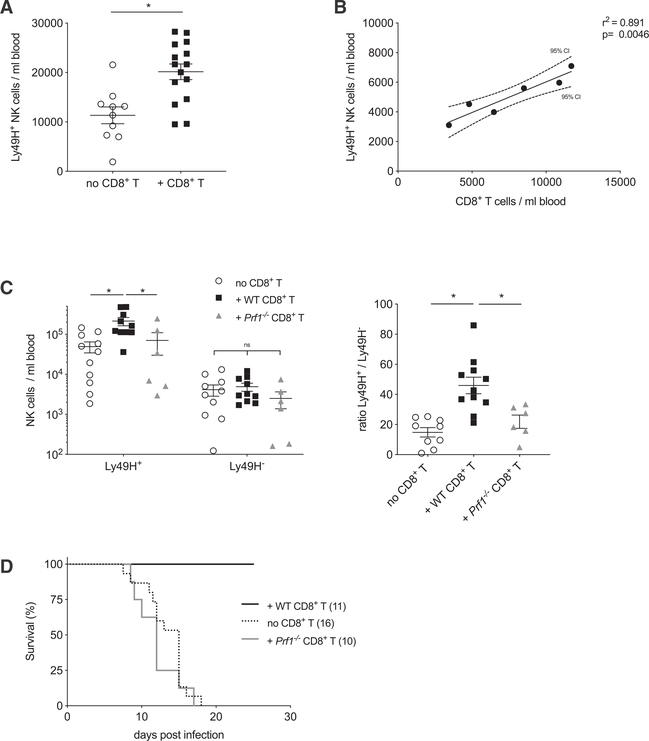
Figure 2. Cytotoxic CD8+ T Cells Are Sufficient and Required to Promote NK Cell Expansion.
(A) 3 × 106 purified CD8+ T cells were transferred into T-cell- and B-cell-deficient mice (Rag2−/−) and challenged with MCMV 10 days after transfer. The number of endogenous Ly49H+ NK cells was quantified in peripheral blood at day 14 PI. Data are pooled from three independent experiments with three to seven mice per group. Statistical differences were calculated with an unpaired t test.
(B) Linear correlation between the number of endogenous Ly49H+ NK cells and the number of CD8+ T cells transferred into Rag2−/− recipients. Data are representative of three independent experiments with six to eight mice per group. Linear correlation was determined using Pearson’s correlation coefficient.
(C) Rag2−/− × Il2rg−/− mice were reconstituted with 3 × 106 purified WT CD8+ T cells, perforin-deficient (Prf1−/−) CD8+ T cells, or no cells. Ten days after transfer, all recipients received 5 × 105 purified NK cells and were challenged with MCMV 1 day later. The number (left) and ratio (right) of transferred Ly49H+ and Ly49H− NK cells were quantified in peripheral blood at day 7 PI. Data are pooled from two independent experiments with three to five mice per group. Statistical differences were calculated with a two-way ANOVA.
(D) Same as in (C), but purified CD8+ T cells and NK cells were transferred into Rag2−/− × Il2rg−/− recipients and survival was calculated using the Kaplan- Meier estimator. Data are pooled from three independent experiments with three to five mice per group.
Graphs present individual data points as scatter dot plots, and error bars show mean ± SEM. A p value of <0.05 was used as the significance cut-off and is indicated with a single asterisk (*). See also Figure S1.
We tested whether CD8+ T cells drive optimal NK cell expansion through a direct mechanism (e.g., by secreting factors required for NK cell proliferation) or an indirect mechanism (e.g., via clearance of virally infected cells). We hypothesized that interleukin 2 (IL-2), a cytokine capable of driving NK cell expansion, and produced by a subset of effector CD8+ T cells in response to antigen stimulation (Feau et al., 2011), could contribute to NK cell expansion. However, Ly49h−/− × Rag2−/− mice reconstituted with IL-2-deficient CD8+ T cells drove similar numbers of co-transferred Ly49H+ NK cells during MCMV infection as mice reconstituted with WT CD8+ T cells (Figure S1C).
We next investigated whether joint control of MCMV by CD8+ T cells and NK cells would result in reduction of viral titers to levels that optimally drive antigen-specific (Ly49H+) NK cell expansion while leaving nonspecific (Ly49H−) NK cell responses unaffected. To this end, we reconstituted Rag2−/− × Il2rg−/− mice (which lack all lymphocytes) with perforin-deficient (Prf1−/−) CD8+ T cells, which cannot lyse virally infected target cells. Compared to WT CD8+ T cells, the Prf1−/− cells were unable to promote the expansion of co-transferred Ly49H+ NK cells (Figure 2C). Similarly, the ratio of Ly49H+ to Ly49H− NK cells was much higher in mice reconstituted with WT CD8+ T cells than in those receiving Prf1−/− CD8+ T cells or NK cells only (Figure 2C). Consistent with this finding, mice reconstituted with both NK cells and WT CD8+ T cells afforded the greatest survival benefit against MCMV challenge, whereas mice reconstituted with NK cells and Prf1−/− CD8+ T cells were as susceptible to virus as mice given NK cells alone (Figure 2D). Altogether, these findings suggest that the cooperative cytotoxic activity of virus-specific CD8+ T cells is required to ensure proper adaptive NK cell responses and protection against infection.
Robust Adaptive NK Cell Responses Require Resolution of High Viral Load
Based on our findings above, we hypothesized that there may be an optimal viral load range that best induces Ly49H+ NK cell expansion. Thus, we investigated how viral titers would be impacted by the lack of CD8+ T cells early during MCMV infection, and whether this would subsequently perturb NK cell function. In WT mice depleted of CD8+ T cells, viral titers were not different at day 2 or 4 PI compared to control mice (Figure S2A), and NK cell number, activation, and effector functions (as measured by Ki67, granzyme B, and IFN-γ staining) were minimally affected (Figures S2B and S2C), consistent with previous reports showing that early NK cell activation following MCMV infection is independent of B and T cells (Welsh et al., 1991). However, CD8+ T cell depletion had a profound impact on the viral load at day 6 PI, where we measured ~10-fold higher amounts of virus in CD8+ T-cell-depleted mice compared to WT controls (Figure 3A).
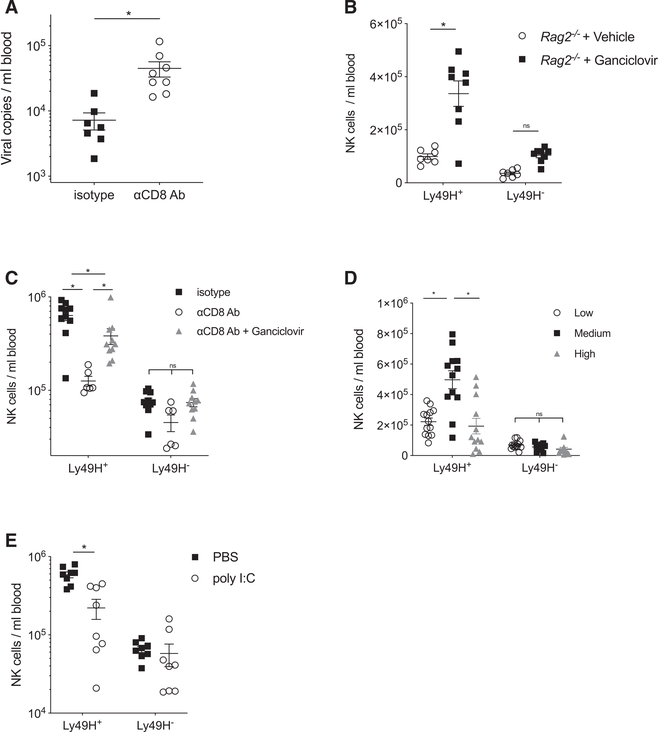
Figure 3. High Viral Load in CD8+ T-Cell-Depleted Mice Restricts NK Cell Expansion.
(A) WT mice treated with αCD8-depleting antibody or isotype control antibody were challenged with MCMV, and viral titers were quantified from peripheral blood at day 6 PI. Data are pooled from two independent experiments with four or five mice per group.
(B) Rag2−/− mice were challenged with MCMV and treated daily with the antiviral drug ganciclovir or vehicle control starting from day 3 PI. The number of endogenous Ly49H+ and Ly49H− NK cells was quantified in peripheral blood at day 7 PI. Data are representative of two independent experiments with seven or eight mice per group.
(C) WT mice treated with αCD8-depleting antibody or isotype control antibody were challenged with MCMV. αCD8-treated mice were treated daily with ganciclovir or vehicle control starting from day 3 PI. The number of endogenous Ly49H+ and Ly49H− NK cells was quantified in peripheral blood at day 7 PI. Data are pooled from three independent experiments with three to five mice per group.
(D) WT mice were treated with a regular viral dose (Medium) or a 5- to 10-fold higher (High) or lower (Low) dose, and the number of endogenous Ly49H+ and Ly49H− NK cells was quantified in peripheral blood at day 6 PI. Data are pooled from three independent experiments with four or five mice per group.
(E) WT mice were challenged with MCMV and treated daily with the TLR3 agonist poly I:C or vehicle control starting from day 4 PI. The number of endogenous Ly49H+ and Ly49H− NK cells was quantified in peripheral blood at day 7 PI. Data are representative of two independent experiments with three to five mice per group.
Graphs present individual data points as scatter dot plots, and error bars show mean ± SEM. Statistical differences were calculated with an unpaired t test (A) and with a two-way ANOVA (B–E). A p value of <0.05 was used as the significance cut-off and is indicated with a single asterisk (*). See also Figure S2.
To test whether higher viral titers were responsible for the impaired NK cell expansion observed in Rag2−/− mice, we infected these mice with MCMV and treated them daily with ganciclovir, a common antiviral treatment against cytomegalovirus. As expected, Rag2−/− mice treated with ganciclovir-controlled virus better than vehicle-treated controls (Figure S2D). Interestingly, Ly49H+ NK cell numbers in Rag2−/− mice treated with ganciclovir were greatly increased in comparison to controls, and a trend toward higher numbers was also observed in Ly49H− NK cells (Figure 3B). Similarly, ganciclovir treatment was also able to rescue adaptive NK cell expansion in WT mice depleted of CD8+ T cells (Figure 3C). Together, these data suggest that CD8+ T cells were not directly priming NK cells but rather indirectly promoting adaptive NK cell responses by targeting virally infected cells in the later stages of infection, where a high viral load has detrimental effects on NK cell expansion. To confirm this, we infected WT mice with a low, medium (regular), or high MCMV dose and measured viral titers at days 2, 4, and 6 PI and circulating Ly49H+ NK cell numbers at day 6 PI. Whereas both low and high viral doses resulted in suboptimal NK cell expansion, the medium dose drove the greatest adaptive NK cell response (Figures 3D and S2E), suggesting that robust NK cell responses require an optimal viral load following infection.
Sustained viral infections can lead to a state of heighten inflammation, which may negatively impact NK cell expansion. To test whether an increased pro-inflammatory milieu was able to dampen NK cell expansion irrespective of the viral load, we treated MCMV-infected WT mice with poly I:C, a strong inductor of pro-inflammatory cytokines, and quantified circulating endogenous NK cells at day 7 PI. Ly49H+ NK cell numbers were significantly lower in poly I:C-treated mice compared to vehicle-treated controls (Figure 3E). Next, we assessed whether CD8+ T-cell-depleted mice had increased levels of pro-inflammatory cytokines at day 6 PI, when viral titers are still high (Figure 3A). Compared to PBS-treated controls, the levels of IFN-α, IFN-γ, IL-18, and TNF-α were significantly increased in the serum of CD8+ T-cell-depleted mice (Figure S2F), suggesting that an increased pro-inflammatory milieu resulting from high-titer viral infection leads to dampened adaptive NK cell responses.
High Viral Titers Maintain NK Cells in a Hyper-activated, Low-Proliferative State
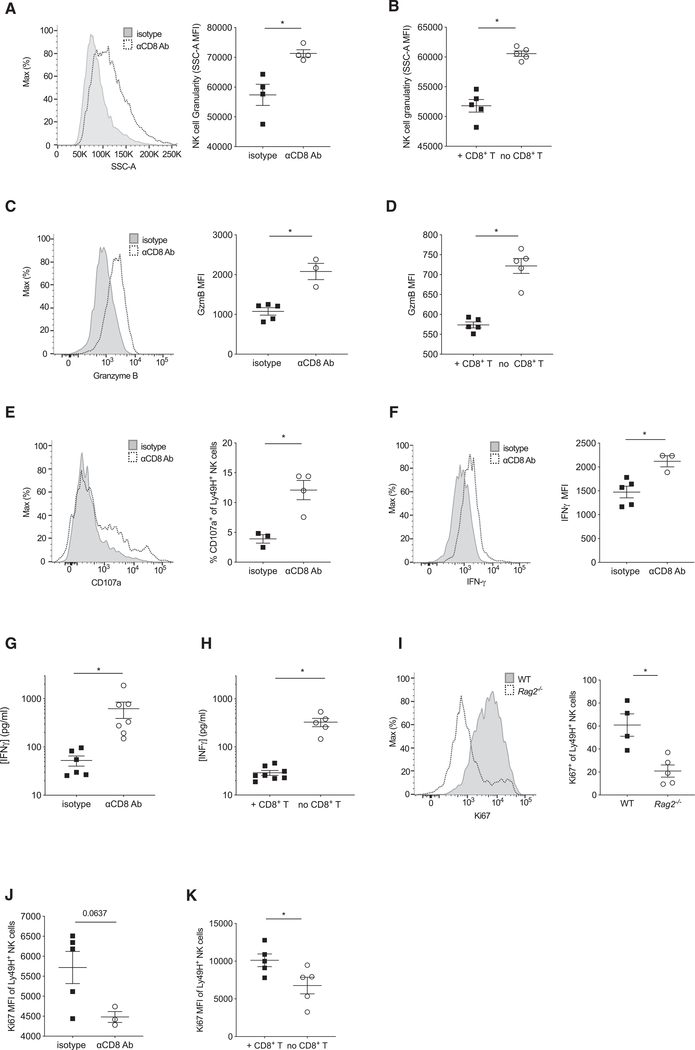
We next investigated how the high viral load limits NK cell expansion. Ly49H+ NK cells from CD8+ T-cell-depleted mice revealed a marked increase in granularity at day 7 PI compared to untreated controls (Figure 4A). Similarly, NK cells from Rag2−/− mice showed greater granularity compared to NK cells from Rag2−/− mice reconstituted with CD8+ T cells (Figure 4B). We thus hypothesized that the high viral load in CD8+ T-cell-depleted mice forces NK cells to stay in a prolonged effector function state, thus limiting other NK cell functions including their ability to undergo proliferation. Consistent with this hypothesis, NK cells from CD8+ T-cell-depleted mice exhibited higher levels of granzyme B compared to control mice at day 7 PI (Figures 4C and 4D), along with higher levels of degranulation and IFN-γ (Figures 4E and 4F). IFN-γ plasma levels were also ~10-fold elevated in CD8+ T-cell-depleted mice compared to control mice (Figures 4G and 4H).
Figure 4. Unchecked Viral Infection Locks NK Cells in an Overactivated State.
(A and C) WT mice treated with αCD8-depleting antibody or isotype control antibody were challenged with MCMV, and the granularity (A) and granzyme B content (C) of splenic NK cells was assessed at day 6 PI. Left: representative histogram. Right: quantification of these markers on Ly49H+ NK cells. Data are representative of three independent experiments with four or five mice per group.
(B and D) Lymphocyte-deficient mice reconstituted with NK cells, along with WT CD8+ T cells or no cells, were challenged with MCMV, and the granularity (B) and granzyme B content (D) of splenic Ly49H+ NK cells were assessed at day 10 PI. Data are representative of two independent experiments with four to six mice per group.
(E and F) Splenic NK cells from WT mice treated with αCD8-depleting antibody or isotype control antibody were assessed at day 6 PI for recent events of degranulation (as measured by CD107a expression) (E) and their ability to produce IFN-γ (F). Left: representative histogram. Right: quantification of these markers on Ly49H+ NK cells. Data are pooled from two independent experiments with three to five mice per group.
(G) WT mice treated with αCD8-depleting antibody or isotype control antibody were challenged with MCMV and IFN-γ quantified in serum at day 6 PI. Data are pooled from two independent experiments with four or five mice per group.
(H) Rag2−/− mice were reconstituted with 3 × 106 CD8+ T cells 10 days prior to MCMV infection, and IFN-γ was quantified in serum at day 7 PI. Data are representative of two independent experiments with five to eight mice per group.
(I–K) WT and Rag2−/− mice (I), WT mice treated with αCD8-depleting antibody or isotype control (J), and Rag2−/− mice reconstituted with CD8+ T cells (K) were challenged with MCMV, and the proliferative marker Ki67 was quantified in circulating Ly49H+ NK cells at day 7 PI. Data are representative of two independent experiments with three to five mice per group.
Graphs present individual data points as scatter dot plots, and error bars show mean ± SEM. Statistical differences were calculated with an unpaired t test. A p value of <0.05 was used as the significance cut-off and is indicated with a single asterisk (*). See also Figure S2.
Lastly, we investigated whether a state of hyper-activation and high viral titers restricted NK cell expansion by limiting their proliferation or exacerbating their activation-induced cell death. We did not observe any significant differences in NK cell apoptosis at later infection stages, as measured by caspase-3 staining (FLICA, Fluorescent Labeled Inhibitor of CAspases) and other measurements of cell death (Figures S2G and S2I; data not shown). However, NK cells from Rag2−/− mice showed a marked decrease in proliferation, as measured by Ki67 staining, compared to NK cells from WT mice (Figure 4I). A similar observation was made in CD8+ T-cell-depleted mice compared to isotype controls (Figure 4J) and in NK cells from Rag2−/− mice compared to NK cells from Rag2−/− mice reconstituted with CD8+ T cells (Figure 4K). Altogether, these data suggest that high viral loads restrict NK cell proliferation by locking them in a hyper-activated effector state for prolonged periods of time and that CD8+ T cells coordinate the clearance of infected cells to reduce the viral load and ensure optimal adaptive NK cell responses.
Numerous studies have provided evidence that adaptive NK cell responses can occur to a wide range of viral infections in different species, from mice to non-human primates to humans (Lopez-Vergès et al., 2011; Reeves et al., 2015; Sun et al., 2009). Although NK cell clonal expansion likely occurs concomitantly with T cell expansion, the interdependence between these adaptive responses has been largely unexplored. In this study, we show that virus-specific CD8+ T cells indirectly control the number and functionality of virus-specific NK cells by modulating the overall viral load.
The response of NK and CD8+ T cells against viruses must be tightly regulated to ensure resolution of the infection while avoiding excessive cytotoxicity and inflammation, which may be detrimental to host survival. At high viral doses, NK cells have been described to provoke lung immunopathology in a model of influenza infection (Zhou et al., 2013). Do the same hyper-activation principles reported in the current study take place in this model, and as such, would adoptive transfer of influenza-specific CD8+ T cells prevent NK-cell-mediated lung pathology? In addition, given the recent finding of a tumor-specific axis involving NK cells, IL-18, and IFN-γ (Ni et al., 2020), it is tempting to hypothesize that the elevated IL-18 levels we observe in CD8+ T-cell-depleted mice may similarly be enhancing and/or prolonging IFN-γ secretion by the hyper-activated NK cells. On the other hand, NK cells have previously been shown to play a role as “rheostats” by altering the number and polyfunctionality of virus- specific CD8+ T cells and limiting CD8+ T-cell-mediated immunopathology (Waggoner et al., 2011). Such control of cytotoxic lymphocyte expansion would present a self-regulatory feedback loop in which NK cells limit the magnitude of CD8+ T cell expansion in a dose-dependent manner. One might hypothesize that such blunted CD8+ T cell responses would in turn tune down adaptive NK cell responses. Would such a mechanism present an efficient way to restrict aberrant cytotoxic lymphocyte responses and decrease immune-mediated immunopathology? Given the striking parallels between the cytotoxic CD8+ T cell and NK cell response to viral infection at a functional, transcriptional, and epigenetic level (Lau et al., 2018), it is compelling to speculate that such coordinated regulatory mechanisms are in place to ensure viral control while avoiding immunopathology.
The interactions of the cytotoxic lymphocytes with MCMV present a remarkable case of host-pathogen evolutionary adaptation. The viral protein m157, while recognized by the activating receptor Ly49H, likely evolved as a decoy major histocompatibility complex (MHC) class-I-like protein, since it is also recognized by the inhibitory receptor Ly49I. In further evidence of this evolutionary tug of war, MCMV strains isolated from wild mice exhibit high polymorphism in its m157 gene (Voigt et al., 2003), and in the absence of CD8+ T cells, MCMV eventually escapes NK cell control by mutating its m157 gene (French et al., 2005). This evolutionary perspective further adds another layer to the complex regulation of the response of NK and CD8+ T cells against MCMV.
Whether the mechanisms driving CD8+ T cell exhaustion are applicable to hyper-activated NK cells during viral infection remains to be determined. While we observed an increased pro-inflammatory milieu in CD8+ T-cell-depleted mice, whether chronic antigen exposure also contributes to the hyper-activated phenotype remains to be elucidated. Similarly, although we did not observe classic markers of CD8+ T cell exhaustion in this setting (data not shown), it will be intriguing to explore whether this hyper-activated NK cell phenotype results in irreversible epigenetic changes that then affect responses against subsequent viral challenge. In summary, a deeper understanding of the relationship and co-regulation of cytotoxic innate and adaptive lymphocytes during viral infection will contribute to the better design of antiviral therapies and vaccination strategies.
STAR★METHODS
RESOURCE AVAILABILITY
Lead Contact
Further information and requests for resources and reagents should be directed to and will be fulfilled by the Lead Contact, Joseph Sun (sunj@mskcc.org).
Materials Availability
This study did not generate new unique reagents.
Data and Code Availability
This study did not generate new datasets.
EXPERIMENTAL MODEL AND SUBJECT DETAILS
Mice
All the mouse strains used in this study were housed and bred at Memorial Sloan Kettering Cancer Center (MSKCC) under specific pathogen-free conditions. Mice used in this study (typically 7–9 weeks) were age- and gender-matched and randomized when possible, in accordance with approved institutional protocols from the Institutional Animal Care and Use Committee at MSKCC. Both genders were used indiscriminately throughout the study. The different strains on C57BL/6 background are listed below: C57BL/6 (CD45.2), B6.SJL (CD45.1), CD45.1 x CD45.2, Il-2fl/fl (Popmihajlov et al., 2012), Klra8−/− (ly49h−/−; (Fodil-Cornu et al., 2008)), Ly49h−/− x Rag2−/−, M38 TCR-transgenic (Torti et al., 2011), OT-I TCR transgenic (Hogquist et al., 1994), perforin−/− (Prf1−/−; (Walsh et al., 1994)), Rag2−/− (The Jackson Laboratory), Rag2−/− x Il2rg−/− (Taconic), and UbcreERT2 (Ruzankina et al., 2007). UbcreERT2 x Il-2fl/fl were generously provided by Dr. Alexander Rudensky (MSKCC).
METHOD DETAILS
Viral infections
Salivary gland stocks of MCMV (Smith strain) were prepared by passaging three times through BALB/c hosts. For direct infection studies, animals were infected intraperitoneally (i.p.) with ~104 PFU of MCMV. For adoptive transfer studies, animals were infected i.p. with ~2 × 103 PFU of MCMV.
Sample collection, organ processing, and flow cytometry
Peripheral blood was drawn retro-orbitally from isofluorane-anesthetised mice using pre-heparinized capillary tubes. Organs were harvested on ice in PBS plus 2% Fetal Bovine Serum, and dissociated to a single-cell suspension. Cell surface staining was performed using the indicated fluorophore-conjugated antibodies (BD Biosciences, BioLegend, eBioscience, R&D Systems, Tonbo). For staining of cytosolic proteins and cytokines, cells were fixed and permeabilized with the BD Cytofix/Cytoperm Fixation/Permeabilization Solution kit. For intranuclear staining, cells were fixed and permeabilized with the eBioscience Foxp3 / Transcription Factor Staining Buffer Set. The following fluorophore-conjugated antibodies were used: CD3ε (17A2), TCRβ (H57–597), CD4 (GK1.5), CD8α (53–6.7), CD8β (YTS156.7.7), CD19 (ID3), F4/80 (BM8.1), Ly6G (1A8), NK1.1 (PK136), NKp46 (29A1.4), CD49b (HMa2), Ly49H (3D10), CD45.1 (A20), CD45.2 (104), CD11b (M1/70), CD27 (LG.3A10), KLRG1 (2F1), CD25 (PC61), CD69 (H1.2F3), Granzyme B (GB11), IFN-γ (XMG1.2), CD107a (1D4B), and Ki67 (SOLA15). NK cell apoptosis was measured by staining cells with FAM-DEVD-FMK FLICA reagent (FLICA, Immunohistochemistry) to measure active Caspase-3/7 in live cells according to the manufacturer protocol, and by staining with Pacific Blue™ Annexin V (BioLegend) according to the manufacturer protocol. Flow cytometry was performed on an LSR II cytometer (BD Biosciences). Data was analyzed with FlowJo10 software (Tree Star).
Adoptive cell transfer
NK cell and CD8+ T cell adoptive transfer studies were performed as previously described (Beaulieu et al., 2014) with some modifications. Briefly, splenic NK cells and/or CD8+ T cells were enriched by negative selection using BioMag® Goat Anti-Rag IgG (QIAGEN), mixed with the following Antibody cocktail (αCD4 (GK1.5 clone, BioXcell), αCD19 (1D3 clone, BioXcell), αLy6G (1A8 clone, BioXcell)), and then sorted to > 95% purity on a BD FACSAria II cell sorter. Purified NK cells (1–4 × 105) or CD8+ T cells (1–3 × 106) were then injected into recipient mice retro-orbitally at least one day before treatment.
Antibody depletion, antiviral treatment, and poly I:C treatment
Depletion of CD8+ T cells and other lymphocyte populations was achieved by intraperitoneal injection of the following purified antibodies: αCD4 (GK1.5 clone, BioXcell, 200 μg/mouse), αCD8 (2.43 clone, BioXcell, 100 μg/mouse), or isotype control (LTF-2 clone, BioXcell, 100 μg/mouse) one day prior to infection. For experiments in which mice received antiviral treatment, mice were treated with daily doses of the nucleoside analog Ganciclovir (Sigma, catalog number 1288306) from day 3 PI to day 7 PI at 20 mg/kg per mouse, dissolved in warm in PBS. For experiments in which mice received polyinosinic:polycytidylic acid (poly I:C) treatment, mice were treated with daily doses of poly I:C (Invivogen) from day 4 PI to day 7 PI at 2.5 mg/kg per mouse.
Quantification of IFN-γ and pro-inflammatory cytokines from plasma
Plasma was collected from peripheral blood, spun at 3000 g for 2 min and stored at −20°C until assayed. IFN-γ levels throughout infection were quantified using the IFN-γ ‘Femto-HS’ High Sensitivity Mouse Uncoated ELISA Kit (Invitrogen), according to manufacturer instructions. The concentration of pro-inflammatory cytokines at day 6 PI was assessed by cytometric bead array (Legendplex custom multi-analyte flow assay kit, Biolegend) according to the manufacturer’s protocol.
Quantification of viral titers from peripheral blood
Total DNA from peripheral blood was extracted using the QIAamp DNA Blood Mini Kit (QIAGEN). Viral copy number in eluted DNA was determined in triplicate by quantitative PCR using iQ SYBR mastermix (Bio-Rad) using the MCMV-specific primers IE-1 (forward: TCGCCCATCGTTTCGAGA, reverse: TCTCGTAGGTCCACTGACGGA). Viral copy number in DNA from samples was calculated by correlating the Cq values to a standard curve constructed with plasmid DNA containing the MCMV IE-1 target region, as previously described (Diaz-Salazar et al., 2020).
QUANTIFICATION AND STATISTICAL ANALYSIS
Statistical analysis
Unless otherwise indicated, all graphs present individual data points as scatter dot plots, and error bars show mean ± SEM. A P value of < 0.05 was used as the significance cut-off and is indicated with a single star (*). Statistical differences across two conditions were calculated using a two-tailed Student’s t test when comparing one variable, or a Sidak’s multiple comparisons test (two-way ANOVA) when comparing two variables. When comparing three or more conditions, a Dunnett’s multiple comparisons test (one-way ANOVA) was used when comparing one variable, and a Tukey’s multiple comparisons test (two-way ANOVA) was performed when comparing two variables. Linear correlation between two variables was determined using pearson’s correlation coefficient. Statistical differences in survival curves were determined by Mantel–Cox test analysis. Statistical analysis was performed using GraphPad Prism software.
Supplementary Material
KEY RESOURCES TABLE.
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| Anti-Mouse CD3ε (clone 17A2) | Tonbo Biosciences | Cat#25-0032; RRID:AB_2621619 |
| Anti-Mouse TCRβ (clone H57-597) | BioLegend | Cat#109220; RRID:AB_893624 |
| Anti-Mouse CD19 (clone 6D5) | BioLegend | Cat#115530; RRID:AB_830707 |
| Anti-Mouse F4/80 (clone BM8.1) | BioLegend | Cat#123117; RRID:AB_893489 |
| Anti-Mouse NK1.1 (clone PK136) | Tonbo Biosciences | Cat#65-5941; RRID:AB_2621910 |
| Anti-Mouse NKp46 (clone 29A1.4) | BioLegend | Cat#137604; RRID:AB_2235755 |
| Anti-Mouse CD49b (clone Hma2) | BioLegend | Cat# 103510; RRID:AB_492851 |
| Anti-Mouse Ly49H (clone 3D10) | eBioscience | Cat#11-5886-81; RRID:AB_1257160 |
| Anti-Mouse CD45.1 (clone A20) | BioLegend | Cat#110729; RRID:AB_1134170 |
| Anti-Mouse CD45.2 (clone 104) | BioLegend | Cat#109821; RRID:AB_493730 |
| Anti-Mouse/Human CD11b (clone M1/70) | BioLegend | Cat#101223; RRID:AB_755985 |
| Anti-CD27 (clone LG.7F9) | eBioscience | Cat#14-0271-81; RRID:AB_467182 |
| Anti-Mouse KLRG1 (clone 2F1) | Tonbo Biosciences | Cat#50-5893; RRID:AB_2621800 |
| Anti-Mouse CD69 (clone H1.2F3) | BioLegend | Cat#104524; RRID:AB_2074979 |
| Anti-Mouse CD25 (clone PC61) | BioLegend | Cat# 102007 RRID:AB_312856 |
| Anti-Human/Mouse Granzyme B (clone GB11) | BioLegend | Cat#515403; RRID:AB_2114575 |
| Anti-Mouse IFN gamma (clone XMG1.2) | Tonbo Biosciences | Cat#20-7311; RRID:AB_2621616 |
| Anti-Mouse CD107a (clone 1D4B) | BioLegend | Cat#121611; RRID:AB_1732051 |
| Anti-Mouse CD4 (clone GK1.5) | BioLegend | Cat# 100433; RRID: AB_893330 |
| Anti-Mouse CD8a (clone 53-6.7) | BioLegend | Cat#100730; RRID:AB_493703 |
| Anti-Mouse CD8b (clone YTS156.7.7) | BioLegend | Cat#126615; RRID: AB_2562776 |
| Anti-Mouse Ki67 (clone SOLA15) | eBioscience | Cat#48-5698-82; RRID: AB_11149124 |
| Anti-NK1.1 depletion antibody (clone PK136) | J. Sun (PI) | N/A |
| InVivoMAb rat IgG2b isotype control, anti-keyhole limpet hemocyanin (isotype control, clone LTF-2) | Bio X Cell | Cat# BE0090; RRID: AB_1107780 |
| InVivoMab Anti-Mouse CD8α (CD8α depletion antibody, NK cell enrichment, clone 2.43) | Bio X Cell | Cat#BE0061; RRID:AB_1125541 |
| InVivoMab Anti-Mouse CD4 (CD4 depletion antibody, NK cell enrichment, clone GK1.5) | Bio X Cell | Cat#BE0003-1; RRID:AB_1107636 |
| InVivoMab Anti-Mouse CD19 (NK cell enrichment, clone 1D3) | Bio X Cell | Cat#BE0150; RRID:AB_10949187 |
| InVivoMab Anti-Mouse Ter-119 (NK cell enrichment, clone TER-119) | Bio X Cell | Cat#BE0183; RRID:AB_10949625 |
| Bacterial and Virus Strains | ||
| Murine Cytomegalovirus (MCMV) | J.C. Sun (PI) | Smith Strain |
| Chemicals, Peptides, and Recombinant Proteins | ||
| Tamoxifen | Sigma-Aldrich | Cat#T5648 |
| Corn oil | Sigma-Aldrich | Cat#C8267 |
| Ganciclovir | Millipore-Sigma | Cat#1288306 |
| poly(I:C) (HMW) VacciGrade | InvivoGen | Cat#vac-pic |
| Critical Commercial Assays | ||
| QIAamp DNA Blood Mini Kit | QIAGEN | Cat#51106 |
| Foxp3 Transcription Factor Staining Buffer Set | Thermo Fisher Scientific | Cat#00-5523-00 |
| iQ SYBR Green Supermix | Bio-Rad | Cat#1708880 |
| BioMag Goat Anti-Rat IgG (NK cell enrichment) | QIAGEN | Cat#310107 |
| IFN gamma ‘Femto-HS’ Mouse Elisa Kit | Invitrogen | Cat#88-8314-22 |
| LEGENDplex Multi-Analyze Flow Assay Kit | BioLegend | (Custom Panel) |
| FAM FLICA Poly Caspase Kit | Bio-Rad | Cat#ICT092 |
| Pacific Blue Annexin V Apoptosis Detection Kit | BioLegend | Cat# 640926 |
| Experimental Models: Organisms/Strains | ||
| Mouse: WT or CD45.2: C57BL/6J | The Jackson Laboratory | Stock#000644; RRID:IMSR_JAX:000664 |
| Mouse: WT or CD45.1: B6.SJL-Ptprca Pepcb/BoyJ | The Jackson Laboratory | Stock#002014; RRID:IMSR_JAX:002014 |
| Mouse: CD45.1xCD45.2 | J. Sun (PI) | N/A |
| Mouse: UbcCre-ERT2: B6.Cg-Ndor1Tg(UBC-cre/ERT2)1Ejb/1J | The Jackson Laboratory | Stock#007001; RRID:IMSR_JAX:007001 |
| Mouse: IL-2fl/fl : IL2tm1.1Kasm | A. Rudensky (PI) (Popmihajlov et al., 2012) | N/A |
| Mouse: UbcCre-ERT2 IL-2fl/fl | A. Rudensky (PI) | N/A |
| Mouse: Rag2−/− IL2rg−/−: C;129S4-Rag2tm1.1Flv Il2rgtm1.1Flv/J | The Jackson Laboratory | Stock#014593; RRID:IMSR_JAX:014593 |
| Mouse: Klra8−/− or Ly49H-deficient | S. Vidal (PI) (Fodil-Cornu et al., 2008) | N/A |
| Mouse: Rag2−/−: B6(Cg)-Rag2tm1.1Cgn/J | The Jackson Laboratory | Stock# 008449; RRID:IMSR_JAX:008449 |
| Mouse: Ly49h−/−x Rag2−/− | J. Sun (PI) | N/A |
| Mouse: Prf1−/−: C57BL/6-Prf1tm1Sdz/J | The Jackson Laboratory | Stock# 002407; RRID: IMSR_JAX:002407 |
| Oligonucleotides | ||
| Primers against MCMV IE-1, F: TCGCCCATC GTTTCGAGA | Diaz-Salazar et al., 2020 | N/A |
| Primers against MCMV IE-1, R: TCTCGTAGGTCC ACTGACCGA | Diaz-Salazar et al., 2020 | N/A |
Highlights.
CD8+ T cells promote the clonal expansion of NK cells by controlling viremia
Antiviral therapy rescues NK cell expansion in the absence of CD8+ T cells
Extended inflammation maintains antiviral NK cells in a hyper-activated state
ACKNOWLEDGMENTS
We thank members of the Sun lab for technical support, experimental assistance, insightful comments, and helpful discussions. Tiger Zhang provided critical experimental assistance. Saskia Hemmers provided valuable feedback and critical mice for these studies. C.D.-S. was supported by a Fulbright fellowship by the Commission for Cultural, Educational and Scientific Exchange between the United States of America and Spain. J.C.S. was supported by the Ludwig Center for Cancer Immunotherapy, the American Cancer Society, the Burroughs Wellcome Fund, and the NIH (grants AI100874, AI130043, and P30CA008748).
Footnotes
DECLARATION OF INTERESTS
The authors declare no competing interests.
SUPPLEMENTAL INFORMATION
Supplemental Information can be found online at https://doi.org/10.1016/j.celrep.2020.108186.
REFERENCES
- Abel AM, Yang C, Thakar MS, and Malarkannan S (2018). Natural killer cells: development, maturation, and clinical utilization. Front. Immunol 9, 1869. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arase H, Mocarski ES, Campbell AE, Hill AB, and Lanier LL (2002). Direct recognition of cytomegalovirus by activating and inhibitory NK cell receptors. Science 296, 1323–1326. [DOI] [PubMed] [Google Scholar]
- Barry KC, Hsu J, Broz ML, Cueto FJ, Binnewies M, Combes AJ, Nelson AE, Loo K, Kumar R, Rosenblum MD, et al. (2018). A natural killer-dendritic cell axis defines checkpoint therapy-responsive tumor microenvironments. Nat. Med. 24, 1178–1191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beaulieu AM, Zawislak CL, Nakayama T, and Sun JC (2014). The transcription factor Zbtb32 controls the proliferative burst of virus-specific natural killer cells responding to infection. Nat. Immunol. 15, 546–553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Biron CA, Byron KS, and Sullivan JL (1989). Severe herpesvirus infections in an adolescent without natural killer cells. N. Engl. J. Med. 320, 1731–1735. [DOI] [PubMed] [Google Scholar]
- Böttcher JP, Bonavita E, Chakravarty P, Blees H, Cabeza-Cabrerizo M, Sammicheli S, Rogers NC, Sahai E, Zelenay S, and Reis E Sousa C (2018). NK cells stimulate recruitment of cDC1 into the tumor microenvironment promoting cancer immune control. Cell 172, 1022–1037.e14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bukowski JF, Warner JF, Dennert G, and Welsh RM (1985). Adoptive transfer studies demonstrating the antiviral effect of natural killer cells in vivo. J. Exp. Med. 161, 40–52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Diaz-Salazar C, Bou-Puerto R, Mujal AM, Lau CM, von Hoesslin M, Zehn D, and Sun JC (2020). Cell-intrinsic adrenergic signaling controls the adaptive NK cell response to viral infection. J. Exp. Med. 217, e20190549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dokun AO, Kim S, Smith HR, Kang HS, Chu DT, and Yokoyama WM (2001). Specific and nonspecific NK cell activation during virus infection. Nat. Immunol. 2, 951–956. [DOI] [PubMed] [Google Scholar]
- Dunn PL, and North RJ (1991). Early gamma interferon production by natural killer cells is important in defense against murine listeriosis. Infect. Immun. 59, 2892–2900. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feau S, Arens R, Togher S, and Schoenberger SP (2011). Autocrine IL-2 is required for secondary population expansion of CD8(+) memory T cells. Nat. Immunol. 12, 908–913. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feng CG, Kaviratne M, Rothfuchs AG, Cheever A, Hieny S, Young HA, Wynn TA, and Sher A (2006). NK cell-derived IFN-γ differentially regulates innate resistance and neutrophil response in T cell-deficient hosts infected with Mycobacterium tuberculosis. J. Immunol. 177, 7086–7093. [DOI] [PubMed] [Google Scholar]
- Fodil-Cornu N, Lee SH, Belanger S, Makrigiannis AP, Biron CA, Buller RM, and Vidal SM (2008). Ly49h-deficient C57BL/6 mice: a new mouse cytomegalovirus-susceptible model remains resistant to unrelated pathogens controlled by the NK gene complex. J. Immunol. 181, 6394–6405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- French AR, Pingel JT, Kim S, Yang L, and Yokoyama WM (2005). Rapid emergence of escape mutants following infection with murine cytomegalovirus in immunodeficient mice. Clin. Immunol. 115, 61–69. [DOI] [PubMed] [Google Scholar]
- Geary CD, and Sun JC (2017). Memory responses of natural killer cells. Semin. Immunol. 31, 11–19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Geary CD, Krishna C, Lau CM, Adams NM, Gearty SV, Pritykin Y, Thomsen AR, Leslie CS, and Sun JC (2018). Non-redundant ISGF3 components promote NK cell survival in an auto-regulatory manner during viral infection. Cell Rep. 24, 1949–1957.e1946. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gumá M, Angulo A, Vilches C, Gómez-Lozano N, Malats N, and López-Botet M (2004). Imprint of human cytomegalovirus infection on the NK cell receptor repertoire. Blood 104, 3664–3671. [DOI] [PubMed] [Google Scholar]
- Hogquist KA, Jameson SC, Heath WR, Howard JL, Bevan MJ, and Carbone FR (1994). T cell receptor antagonist peptides induce positive selection. Cell 76, 17–27. [DOI] [PubMed] [Google Scholar]
- Karo JM, Schatz DG, and Sun JC (2014). The RAG recombinase dictates functional heterogeneity and cellular fitness in natural killer cells. Cell 159, 94–107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lang PA, Lang KS, Xu HC, Grusdat M, Parish IA, Recher M, Elford AR, Dhanji S, Shaabani N, Tran CW, et al. (2012). Natural killer cell activation enhances immune pathology and promotes chronic infection by limiting CD8+ T-cell immunity. Proc. Natl. Acad. Sci. USA 109, 1210–1215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lanier LL (2008). Up on the tightrope: natural killer cell activation and inhibition. Nat. Immunol. 9, 495–502. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lau CM, Adams NM, Geary CD, Weizman OE, Rapp M, Pritykin Y, Leslie CS, and Sun JC (2018). Epigenetic control of innate and adaptive immune memory. Nat. Immunol. 19, 963–972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee BC, Kim MS, Pae M, Yamamoto Y, Eberlé D, Shimada T, Kamei N, Park HS, Sasorith S, Woo JR, et al. (2016). Adipose natural killer cells regulate adipose tissue macrophages to promote insulin resistance in obesity. Cell Metab. 23, 685–698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lopez-Vergès S, Milush JM, Schwartz BS, Pando MJ, Jarjoura J, York VA, Houchins JP, Miller S, Kang SM, Norris PJ, et al. (2011). Expansion of a unique CD57+NKG2Chi natural killer cell subset during acute human cytomegalovirus infection. Proc. Natl. Acad. Sci. USA 108, 14725–14732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Louis C, Guimaraes F, Yang Y, D’Silva D, Kratina T, Dagley L, Hediyeh-Zadeh S, Rautela J, Masters SL, Davis MJ, et al. (2020). NK cell–derived GM-CSF potentiates inflammatory arthritis and is negatively regulated by CIS. J. Exp. Med. 217, e20191421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Madera S, and Sun JC (2015). Cutting edge: stage-specific requirement of IL-18 for antiviral NK cell expansion. J. Immunol. 194, 1408–1412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Madera S, Rapp M, Firth MA, Beilke JN, Lanier LL, and Sun JC (2016). Type I IFN promotes NK cell expansion during viral infection by protecting NK cells against fratricide. J. Exp. Med. 213, 225–233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ni J, Wang X, Stojanovic A, Zhang Q, Wincher M, Bühler L, Arnold A, Correia MP, Winkler M, Koch PS, et al. (2020). Single-cell RNA sequencing of tumor-infiltrating NK cells reveals that inhibition of transcription factor HIF-1α unleashes NK cell activity. Immunity 52, 1075–1087.e8. [DOI] [PubMed] [Google Scholar]
- O’Sullivan TE, Rapp M, Fan X, Weizman OE, Bhardwaj P, Adams NM, Walzer T, Dannenberg AJ, and Sun JC (2016). Adipose-resident group 1 innate lymphoid cells promote obesity-associated insulin resistance. Immunity 45, 428–441. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Popmihajlov Z, Xu D, Morgan H, Milligan Z, and Smith KA (2012). Conditional IL-2 gene deletion: consequences for T cell proliferation. Front. Immunol. 3, 102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reeves RK, Li H, Jost S, Blass E, Li H, Schafer JL, Varner V, Manickam C, Eslamizar L, Altfeld M, et al. (2015). Antigen-specific NK cell memory in rhesus macaques. Nat. Immunol. 16, 927–932. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruzankina Y, Pinzon-Guzman C, Asare A, Ong T, Pontano L, Cotsarelis G, Zediak VP, Velez M, Bhandoola A, and Brown EJ (2007). Deletion of the developmentally essential gene ATR in adult mice leads to age-related phenotypes and stem cell loss. Cell Stem Cell 1, 113–126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schuster IS, Wikstrom ME, Brizard G, Coudert JD, Estcourt MJ, Manzur M, O’Reilly LA, Smyth MJ, Trapani JA, Hill GR, et al. (2014). TRAIL+ NK cells control CD4+ T cell responses during chronic viral infection to limit autoimmunity. Immunity 41, 646–656. [DOI] [PubMed] [Google Scholar]
- Steinhoff U, Wand-Württenberger A, Bremerich A, and Kaufmann SH (1991). Mycobacterium leprae renders Schwann cells and mononuclear phagocytes susceptible or resistant to killer cells. Infect. Immun. 59, 684–688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun JC, Beilke JN, and Lanier LL (2009). Adaptive immune features of natural killer cells. Nature 457, 557–561. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun JC, Madera S, Bezman NA, Beilke JN, Kaplan MH, and Lanier LL (2012). Proinflammatory cytokine signaling required for the generation of natural killer cell memory. J. Exp. Med. 209, 947–954. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Torti N, Walton SM, Brocker T, Rülicke T, and Oxenius A (2011). Nonhematopoietic cells in lymph nodes drive memory CD8 T cell inflation during murine cytomegalovirus infection. PLoS Pathog. 7, e1002313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vivier E, Tomasello E, Baratin M, Walzer T, and Ugolini S (2008). Functions of natural killer cells. Nat. Immunol. 9, 503–510. [DOI] [PubMed] [Google Scholar]
- Vivier E, Raulet DH, Moretta A, Caligiuri MA, Zitvogel L, Lanier LL, Yokoyama WM, and Ugolini S (2011). Innate or adaptive immunity? The example of natural killer cells. Science 331, 44–49. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Voigt V, Forbes CA, Tonkin JN, Degli-Esposti MA, Smith HR, Yokoyama WM, and Scalzo AA (2003). Murine cytomegalovirus m157 mutation and variation leads to immune evasion of natural killer cells. Proc. Natl. Acad. Sci. USA 100, 13483–13488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waggoner SN, Cornberg M, Selin LK, and Welsh RM (2011). Natural killer cells act as rheostats modulating antiviral T cells. Nature 481, 394–398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walsh CM, Matloubian M, Liu CC, Ueda R, Kurahara CG, Christensen JL, Huang MT, Young JD, Ahmed R, and Clark WR (1994). Immune function in mice lacking the perforin gene. Proc. Natl. Acad. Sci. USA 91, 10854–10858. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Welsh RM, Brubaker JO, Vargas-Cortes M, and O’Donnell CL (1991). Natural killer (NK) cell response to virus infections in mice with severe combined immunodeficiency. The stimulation of NK cells and the NK cell-dependent control of virus infections occur independently of T and B cell function. J. Exp. Med. 173, 1053–1063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yokoyama WM, Kim S, and French AR (2004). The dynamic life of natural killer cells. Annu. Rev. Immunol. 22, 405–429. [DOI] [PubMed] [Google Scholar]
- Zhou G, Juang SW, and Kane KP (2013). NK cells exacerbate the pathology of influenza virus infection in mice. Eur. J. Immunol. 43, 929–938. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
This study did not generate new datasets.