Abstract
Background
With cardiovascular disease increasing, substantial research has focused on the development of prediction tools. We compare deep learning and machine learning models to a baseline logistic regression using only ‘known’ risk factors in predicting incident myocardial infarction (MI) from harmonized EHR data.
Methods
Large-scale case-control study with outcome of 6-month incident MI, conducted using the top 800, from an initial 52 k procedures, diagnoses, and medications within the UCHealth system, harmonized to the Observational Medical Outcomes Partnership common data model, performed on 2.27 million patients. We compared several over- and under- sampling techniques to address the imbalance in the dataset. We compared regularized logistics regression, random forest, boosted gradient machines, and shallow and deep neural networks. A baseline model for comparison was a logistic regression using a limited set of ‘known’ risk factors for MI. Hyper-parameters were identified using 10-fold cross-validation.
Results
Twenty thousand Five hundred and ninety-one patients were diagnosed with MI compared with 2.25 million who did not. A deep neural network with random undersampling provided superior classification compared with other methods. However, the benefit of the deep neural network was only moderate, showing an F1 Score of 0.092 and AUC of 0.835, compared to a logistic regression model using only ‘known’ risk factors. Calibration for all models was poor despite adequate discrimination, due to overfitting from low frequency of the event of interest.
Conclusions
Our study suggests that DNN may not offer substantial benefit when trained on harmonized data, compared to traditional methods using established risk factors for MI.
Keywords: Myocardial infarction, Machine learning, Electronic health records
Introduction
Cardiovascular disease (CVD) has long been a leading cause of death in the United States, with more than 900,000 deaths in 2016, a substantial portion of which were attributable to myocardial infarction (MI) [1]. Although there have been dramatic improvements in public health that have spurred a decline in CVD related deaths over the past several decades, incidence of CVD mortality has remained steady in recent years. As a result, considerable effort has been placed in improving risk prediction of CVD-related events [2, 3]. Despite this, clinical practice has remained largely unchanged, relying on traditional scoring metrics such as Thrombolysis in Myocardial Infarction (TIMI) Risk Score [4] for risk stratification.
Machine learning (ML) has risen as a contemporary method of prediction. ML methods offer an approach that contrasts with typical statistical tools to extract relationships between variables in a training dataset to predict various outcomes, including mortality. Numerous studies have applied these techniques to predicting cardiac events primarily in patients presenting with acute coronary syndrome (ACS), in some cases showing outperformance of traditional risk methods and statistical techniques [4–10]. A specific class of these techniques, deep learning, has received significant interest in recent years. A form of representation learning, deep learning allows a machine to learn patterns based on raw input of data without any prior variable engineering [11]. Successful applications of deep learning have been diverse, from playing poker [12] to detecting mammographic lesions [13]. Studies have demonstrated deep learning can improve upon traditional ML methods in predicting mortality, by capturing non-linear relationships among predictor variables [14] and through the incorporation of ‘unstructured’ data such as word embeddings in discharge summaries [15].
Use of Electronic Health Record (EHR) data has increasingly been the focus of such prediction efforts. EHR data offers numerous advantages such as longitudinal follow up and large, more generalized cohorts for study [16]. Applications in cardiology have included prediction of lifetime costs associated with CVD, phenotyping patients most likely to benefit from lipid pharmacotherapy, and estimating 1 year mortality risk in the ICU setting. Perhaps most exciting is the use of EHR data to facilitate the development of learning healthcare systems, where data can be shared across institutions and used to enhance models for real-time clinical prediction. Prior work has demonstrated that clinical ‘phenotypes’ can be developed that describe institutional patient populations and are stable from institution to institution [17]. Harmonization is a method of sharing these representations across institutions in a way that is translatable and preserves patient privacy [18].
Here we explore the application of ML methods to a harmonized dataset to predict incident MI at 6 months. Prior work, particularly studies that have used deep learning, have largely limited the populations studied for MI prediction to those patients that present to the ED or cath lab for ACS [14, 15, 19, 20]. And while prior studies have used EHR data for MI prediction, none to our knowledge have used harmonized datasets [5, 21–23]. We aim to test the efficacy of ML methods in this context compared to more traditional statistical approaches. Our end goal is to develop a prediction system that not only meaningfully predicts MI incidence over a clinically-relevant time horizon in an undifferentiated patient but also can be integrated with EHR systems across institutions to facilitate the advent of learning healthcare systems. Among the characteristics we examined in this developmental process includes identification of the appropriate data resampling to manage dataset imbalance, and development of a classification algorithm based on training time and accuracy.
Methods
We conducted a systematic examination of EHR data sampled from over 2 million individuals, in whom we have harmonized 52,000 features, including diagnoses, medications, and procedures under the Observation Medical Outcomes Partnership common data model (OMOP-CDM). The code used for the analyses, as well as the model weights and mapping (OMOP-CDM input codes) for the final model, are available in the supplemental material. This investigation was approved by the University of Colorado Multiple Institutional Review Board (COMIRB), with permission for data access under the Health Data Compass honest-broker agreement. De-identified data was used for all analyses; no individual patient information was accessed in conduct of this investigation.
Study population and case ascertainment
The UCHealth hospital system includes 3 large regional centers (North, Central, South) over the front range of Colorado that share a single Epic instance, which allows data from all centers to be pooled into a single data warehouse, a copy of which is located on the Google cloud platform. This warehouse of data was queried using Google BigQuery to create a dataset and conduct analyses directly on the Google cloud platform, where an array of machine-learning tools can be run on virtual machines. To create our study dataset, we applied a classification approach based on predicting risk of incident MI over a 6-month period. We performed a SQL query on the UCHealth EHR for subjects, first extracting patients that had MI, excluding patients who had a prior diagnosis of MI. For these patients with incident MI, the index date was 6 months prior to the MI event and all data prior to this date was used for medical history. For the cohort of control patients without MI, the index date used was 6 months prior to the last recorded encounter date in our dataset and all data prior to this date was again used for medical history. Data was gathered during the time period 2003 to 2018.
Common data model and data splitting
We used a common data model for EHR data, based on the Observational Health Data Sciences and Informatics (OHDSI) collaboration, which uses OMOP-CDM [24–26]. The OMOP CDM is a mapping of the raw EHR data to a harmonized dataset; we used this CDM with 52 k variables (i.e., features) from the EHR, including age, sex, diagnoses, procedures, and medications. The missing data assumption for this study was that if a given feature, other than age or sex, was absent then this feature was assumed to not be present for that person. Preliminary studies identified a substantial decrease in analytical time using the top 8500 most common (across the entire EHR) concepts, which were used as input into prediction models. The final dataset was composed of 2.27 million records, which was then split into training (80%), dev for hyperparameter tuning (0.3%), and testing (19.7%) sets to compare the models developed in this investigation.
Model development
For all models, hyperparameter tuning was performed using iterative random sampling of 10,000 records for manual grid search (neural networks), and 10-fold cross validation for automated grid search (for other machine learning approaches).
Due to the relative infrequency of the outcome (6-month incident MI) across the dataset (0.91%), there is substantial imbalance between the cases and controls in this investigation. The presence of such imbalance can produce classifiers that default to assigning new cases to the majority class, thus having very poor accuracy overall [27]. Re-sampling techniques have increasingly been studied as a potential solution to this, with both oversampling techniques - which increase frequency of rare cases relative to controls - and undersampling techniques - which reduce frequency of controls relative to cases - used. We examined several strategies for resampling, including random oversampling, synthetic minority oversampling technique (SMOTE) [27], random undersampling, and cluster centroid. To identify the best resampling approach, we used a deep neural network (DNN) (7 layers × 100 neurons/layer), as pilot analyses using a smaller dataset suggested this approach might be superior to other ML approaches. We also compared with a model using no resampling (imbalanced).
Once we identified an optimal resampling approach, we compared several classification algorithms, including naïve Bayesian classification, regularized logistic regression, random forest classification, boosted gradient classification, one-layer fully connected neural networks (shallow) and multiple layer fully connected neural networks (deep). Parameters for the DNN were obtained by hyperparameter tuning using manual grid search, with including the number of layers, number of neurons, activation function (tanh, relu, sigmoid and elu), learning rate, dropout and batch-size. Tuning yieleded a final DNN size of 7 layers and 100 neurons per layer. Model comparison was based on area-under-curve and F1 statistic. Loss function applied in this analysis was the cross-entropy loss. Computation time includes all prior data sampling and algorithm performance. Once an optimal model and resampling approach were identified, we conducted sensitivity analysis using several alternative resampling and modeling approaches in combination to ensure that the combination (dimensionality reduction, resampling, and classification algorithm) identified was indeed optimal. Precision-recall and receiver-operator characteristic curves, as well as feature importance plots, were created for the optimal model for manual inspection. Platt re-scaling and isotonic regression was employed to better calibrate predicted probabilities to expected distributions of the observed probabilities in the data.
Validation of developed model
The optimal model was then compared with a simple logistic regression model without regularization based on presence of known clinical predictors of MI, using diagnosis codes (ICD-10; ICD-9) to obtain inputs of diabetes (ICD 9: 250, ICD10 EO8 – E13), hypertension (I10x; 401.x), age, sex, smoking status (ICD 9305.1, ICD 10 F17.210), and fasting lipid level.
Computation and analysis
All analyses were run on Google Cloud Platform, using 96 CPUs and 620 GB of RAM. Scripts were composed in Python (version 3) and were run on Jupyter Notebook with Tensorflow platform on the Google Cloud Platform. Machine learning packages included scikit-learn and keras. Confidence intervals were calculated using Wald method [28, 29], although almost all were within the rounding error of the estimates due to the large testing sample size (N = ~ 2.27 M), and are not displayed.
Results
Across the entire dataset of ~ 2.27 million cases, approximately 21 thousand cases of MI (Table 1) were recorded. Among this group, the prevalence of cardiovascular risk factors was, as expected, significantly greater than controls. Patients with a first-MI within 6 months were older with higher rates of coronary artery disease, diabetes, hypertension, chronic kidney disease, obesity and heart failure.
Table 1.
UCHealth Population by MI diagnosis
| No MI | 6-month Incident MI | |
|---|---|---|
| Number (%) | 2.25 million(%) | 20,591 (%) |
| Age (Mean ± SD) | 43.32 ± 22.56 | 70.36 ± 14.02 |
| Female sex (%) | 1,228,689 (54.5%) | 7880 (37.7%) |
| Hypertension (%) | 376,371 (16.72%) | 12,314 (59.8%) |
| Coronary artery disease (%) | 59,199 (2.63%) | 6700 (32.53%) |
| Mitral valve disease (%) | 30,251 (1.34%) | 1668 (8.1%) |
| Heart failure (%) | 44,999 (2%) | 3038 (14.75) |
| Diabetes mellitus (%) | 131,059 (5.82%) | 5432 (26.38%) |
| Obesity (%) | 127,575 (5.67%) | 2343 (11.37%) |
| Chronic kidney disease (%) | 42,759 (1.9%) | 2128 (10.33%) |
Legend: Baseline demographics and relative frequency of typical MI risk factors. Diagnoses based on presence of diagnosis code (ICD-10; ICD-9) for each. Hypertension: I10x; 401.x, Coronary artery disease: I25.1; 414.01, Mitral Valve disease: I34.2, I34.0, 394.0, 424.0, Heart failure: I50.9, 428.0, Type II Diabetes Mellitus: E11.9, 250.00, Obesity: E66.9, 278.0, Chronic kidney disease: N18.9, 585.9
We next examined various re-sampling methods to address the substantial degree of imbalance in the dataset of cases compared to controls. Using a 7-layer DNN algorithm with hyperbolic tangent activation and 20% dropout, we found that random undersampling had the overall best performance by AUC and F1 (Table 2) and shortest time for algorithm time-to-completion. All re-sampling methods, including random oversampling, SMOTE, random undersampling and undersampling with cluster centroid, substantially outperformed the DNN model with no re-sampling methods used to address class imbalance (AUC 0.51, F1 0.01).
Table 2.
Comparison of Resampling Strategies
| F1 Score | AUC | Training time | |
|---|---|---|---|
| Oversampling | |||
| Random | 0.105 | 0.816 | 22 min |
| SMOTE | 0.109 | 0.786 | 34 min |
| Undersampling | |||
| Random | 0.091 | 0.839 | 2 min |
| Cluster centroid | 0.057 | 0.78 | 78 min |
| None | 0.01 | 0.51 | 3 min |
Sampling comparison from deep learning model
Next, using random undersampling on the training data, we tested the accuracy of various classification models on the held-out dataset and found a DNN outperformed Naïve Bayes, logistic regression with regularization, shallow NN, random forest, and boosted gradient descent models (Table 3). Model accuracy, measured by AUC, was quite similar between the DNN and a logistic regression with regularization (AUC 0.835 vs. 0.829). The final model had a sensitivity of 0.82, specificity of 0.733, precision of 0.05 and recall of 0.82 (Fig. 1).
Table 3.
Comparison of machine learning approaches
| F1 Score | AUC | Training time | |
|---|---|---|---|
| Naïve Bayes | 0.060 | 0.73 | 1 min |
| Logistic regression with L2 regularization | 0.084 | 0.829 | 1 min |
| Logstic regression with no regularization | 0.06 | 0.79 | 1 min |
| RF | 0.084 | 0.765 | 3 min |
| Shallow NN | 0.101 | 0.83 | 1 min |
| Deep NN | 0.092 | 0.835 | 2 min |
| GBM | 0.077 | 0.83 | 9 min |
Comparison of various models using Random Undersampling technique and all features. F1 and AUC calculated from model applied to held-out testing set (20%); training time is for training of training set (80%)
Fig. 1.
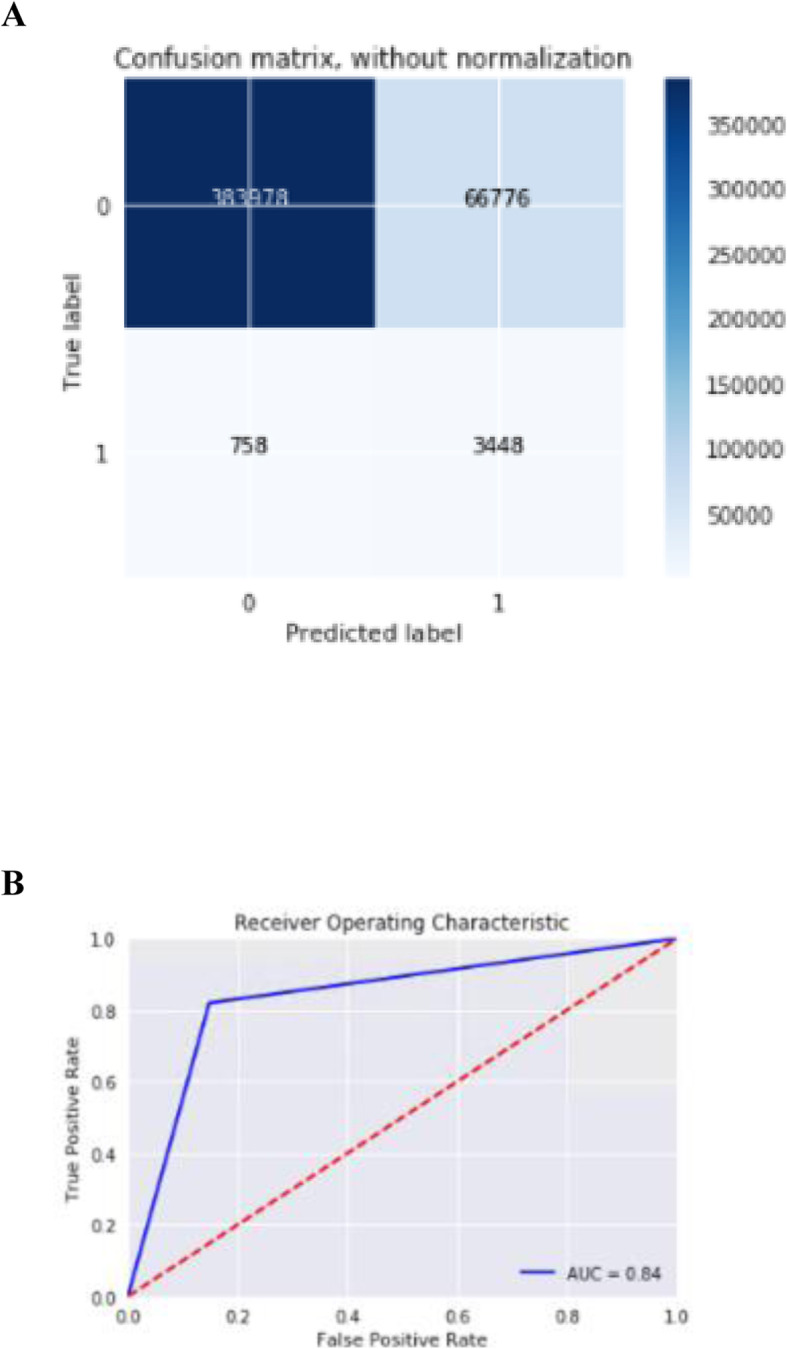
a. Precision-recall curve for optimal model. b. ROC curve for optimal model. A) Confusion matrix for the optimally performing DNN B) ROC curve for DNN model
Finally, the optimal model, a DNN, was compared to a logistic regression using only the known risk factors for MI listed above. The baseline model’s AUC, 0.79, and F score 0.06, both were near that of the optimal model.
Calibration plots for both the optimal DNN (Fig. 2) and simple logistic models (Fig. 3) were poor, with substantial discrepancy between observed distributions and predicted probabilities. Both models, however, appear to adequately discriminate positive cases of incident MI from controls. Both Platt’s rescaling and isotonic regression were employed with no improvement in model calibration.
Fig. 2.
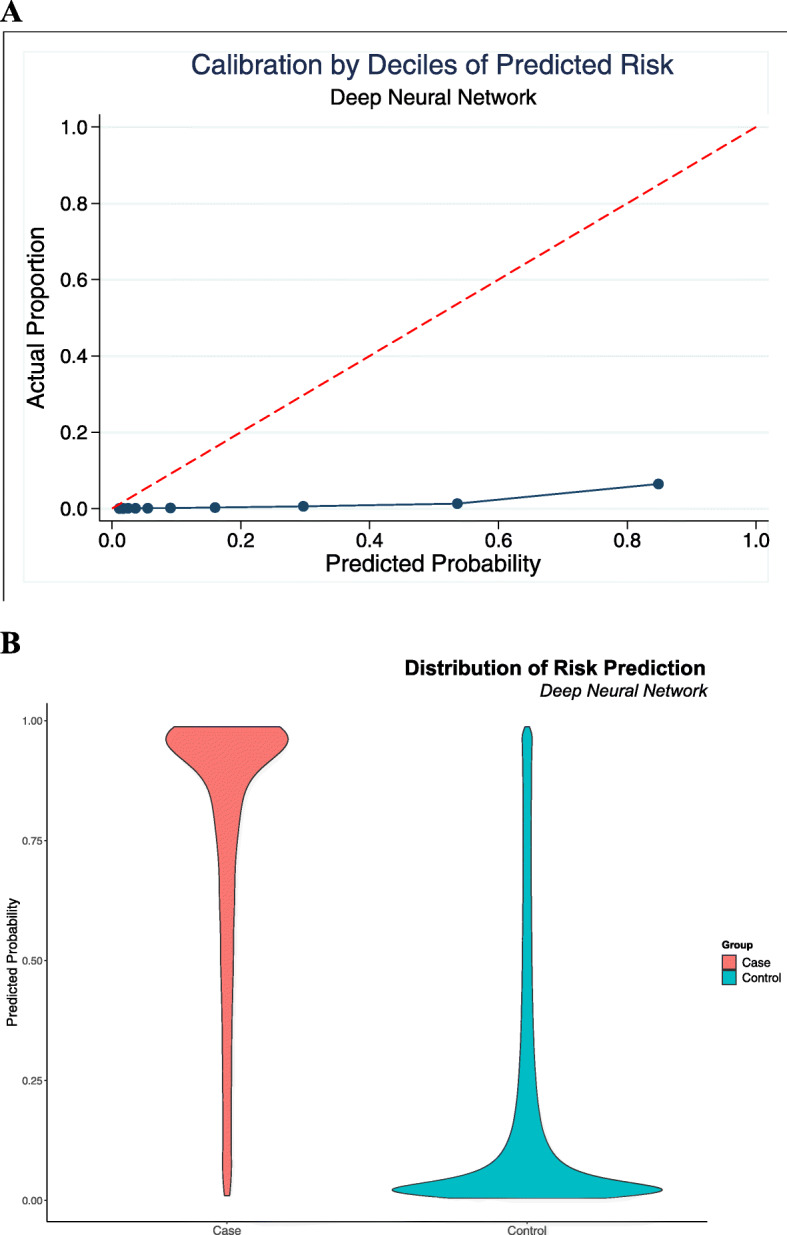
Calibration curve for optimal model. a. Calibration plot for the DNN, showing a wide discrepancy between actual observed distribution of outcomes vs. predicted probabilities from the model b. Distribution of distribution of predicted probabilities for cases vs. controls, showing good discrimination
Fig. 3.
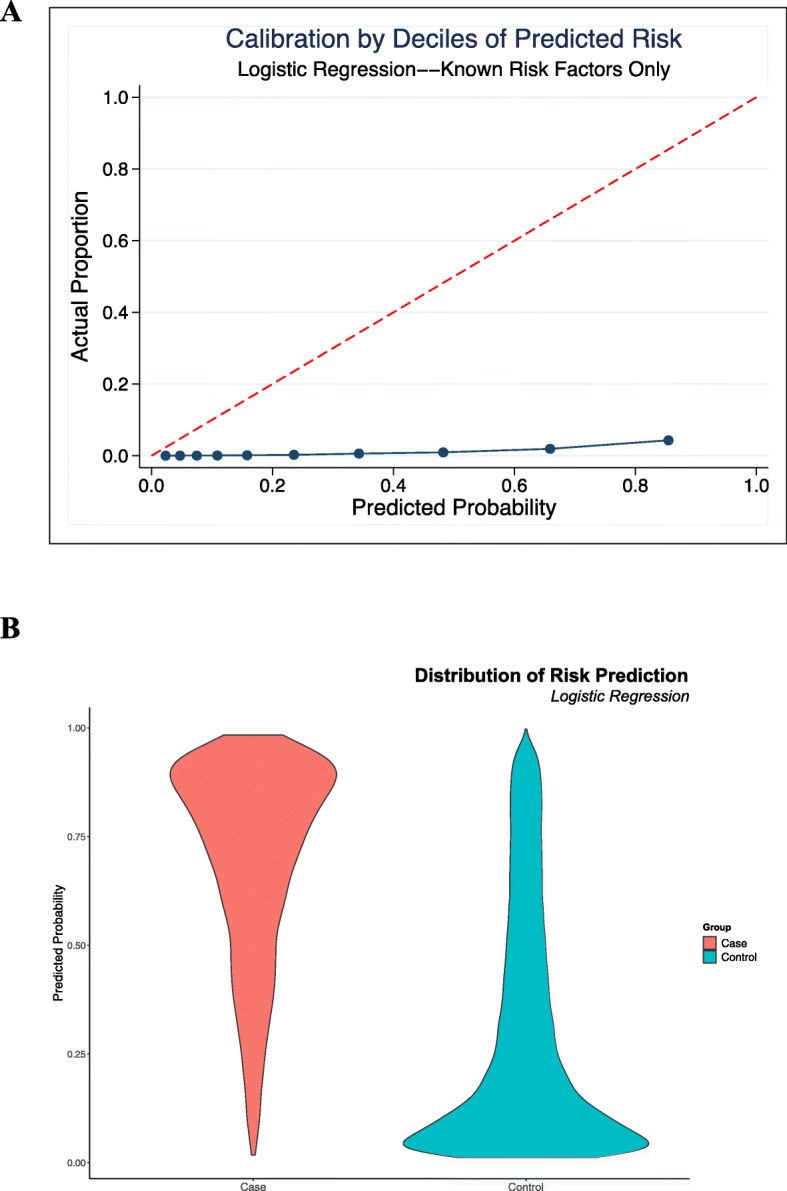
Calibration curve for comparison model. a. Calibration plot for the logistic model using only known risk factors, showing a similar discrepancy between actual observed distribution of outcomes vs. predicted probabilities b. Distribution of distribution of predicted probabilities for cases vs. controls, again with good discrimination
Discussion
In this investigation of using harmonized EHR data for prediction of incident MI at 6 months, we found a DNN with random undersampling had the most accurate classification, although a simpler logistic regression using a more limited set of ‘known risk factors’ performed nearly as well. All models were poorly calibrated. While our study was ultimately negative a variety of useful insights emerged that would help guide future research in integrating ML methods with real-world clinical decision-making. Our study with over 2 million subjects and 52,000 features has the largest sample size and feature space for MI prediction to date [4–6, 8–10, 14, 15, 19, 20, 23]. As prior studies that have applied deep learning to MI prediction have focused on predicting recurring events or mortality risk in patients that presented to the ED or cath lab with ACS [14, 15, 19, 20], to our knowledge, this is the first investigation to attempt to use deep learning to predict ‘first-MI’ events in an otherwise undifferentiated patient population. It is well established that after an initial MI, rates of subsequent MI are markedly higher, up to 30%, compared to the general population [30]. The population of those with ‘first-time MI’ represents a different phenotype than has been assessed in prior studies, with expected lower rates of MI and overall improved survival outcomes. Despite the DNN’s performance, with an AUC in range of prior studies, the degree of improvement in accuracy was not substantial compared to a logistic regression with a handful of ‘known’ predictors of MI. A key explanation for this is likely the data input itself, harmonized EHR data. Prior studies have shown advanced ML methods and even DNN improve in performance relative to controls when the numbers of features increase specifically when using EHR data [5, 15]. These studies, however, did not use harmonized data [31]. While harmonized data offers many benefits in real world applications, such as integration immediately with healthcare systems and allowing for direct application and validation to data mapped from a separate EHR, there are substantial challenges to successfully deploying complicated ML methods to learn from this data. Numerous variables have not been harmonized across datasets via the OMOP-CDM, many of which are more granular, ‘unstructured’ data that may specifically benefit deep learning methods. Payrovnaziri et all, in fact, noted that word embeddings derived from discharge summaries were critical training inputs to improve DNN’s predictions compared to controls when using EHR data [15]. Novel biomarkers [32, 33] and features of both echo [34] and cardiac MRI [35] imaging modalities are being developed as markers of acute MI and again, represent ‘unstructured data’ not currently included in OMOP-CDM. While it is not new that a logistic model can perform similarly to ML methods in the context of MI mortality prediction [8, 10], it is striking and distinct from prior studies that a logistic model with a feature space substantially reduced to only known risk factors performed similarly to the DNN trained on the full feature set. This reflects that most of the risk prediction ‘content’ of the harmonized dataset is already captured by known risk factors for MI. Deep learning, which thrives on uncovering complex, nonlinear relationships between variables, may ultimately show improved performance with more granular data, but such granular data would limit the key benefit of harmonization in that it allows models developed at individual institutions to speak to each other in the same global space [18]. This trade-off between use of data inputs that can be replicated across populations such as harmonized data with data types that are more granular and thus likely to be more predictive, is reflective of the bias-variance trade-off in ML. Moreover, as harmonization is a crucial step towards incorporating prediction models in real-world EHR systems at scale, these limitations of advanced ML methods, specifically DNN, are a very significant consideration for practical development of risk prediction tools deployed at scale across institutions.
An interesting observation from our analysis is that predicting an outcome over a short time frame appears to be poorly calibrated regardless of which predictive approach is used, even for models having good discrimination. Despite considerable efforts to improve calibration of the final model, including Platt rescaling and isontonic regression [36, 37], the final model’s calibration, or measure of how well predicted probabilities align with observed distributions of events, was extremely poor (Fig. 2). Calibration was similarly poor for the simpler logistic regression, despite its discrimination also appearing to be adequate (Fig. 3). The pattern of these calibration plots, while having good discrimination, suggests each of the models overpredicts events at all probabilities. This is consistent with the extremely high degree of imbalance in our dataset - as our approach aimed to be a screening mechanism integrated into EHR systems, the size of the dataset was substantially larger than any prior study, and as a result, the event of interest, i.e., a ‘first’ MI in 6 months, was very rare. Prior studies that have explicitly evaluated calibration when using ML methods for risk stratification [5, 38], for instance, had events of interest with 5–10 fold higher occurrence than in our study. Our prior study, another study with significant dataset imbalance in predicting a rare event, noted similar limits with accuracy and calibration of models in the context of prediction of first time occurrence of atrial fibrillation [39]. As all models in this investigation were poorly calibrated, regardless of complexity, it may be that the use of prediction methods has inherent limitations when applied as a screening tool for MI prediction in undifferentiated patients. Alternatively, it may simply be that the time horizon of 6 months, although a clinically relevant window, may be too short for such methods to show utility. In addition to highlighting these limitations, our study demonstrates the necessity of systematically evaluating model calibration. Few prior studies in MI prediction using ML have done so, despite calibrated models ultimately providing more interpretable probabilities that could be used as individual-level predictions and thus meaningfully impact clinical practice.
Limitations and future considerations
Our study has several limitations. For one, our study included a very simple method for the temporal relationships between features in our dataset, which did not account for time-varying effects or censoring. For instance, a diagnosis or medication that was given one month before a first MI event was weighted the same as one given 4 years prior. While we suggest a 6-month time frame is reasonable for short-term prediction, we did not rigorously test this assumption and additional information about temporal risk will be needed for more accurate prediction approaches. More sophisticated methods, such as recurrent neural networks or parametric survival functions could provide more accurate prediction in future investigations. Assessing the stability of model predictions across a range of time intervals would not only help with model selection but also provide clinically relevant stratification of patients that may not have immediate but do have heightened risks for MI.
Another weakness of our study, already noted above, is that we necessarily excluded some data elements, including lab values, diagnostic tests, imaging and even reports that may have proven useful for MI prediction. In part this is due to the lack of harmonization of some variables via OMOP-CDM.
A final limitation of our study is we necessarily limited the event of interest to first-MI. As MI events were obtained using ICD codes, it was impractical for us to obtain and model repeat MI events. Repeat MI’s, however, are of substantial clinical interest as risk and likelihood for MI heighten substantially after an initial event.
Conclusion
We studied the development of an ML model to predict 6-month occurrence of incident MI using harmonized EHR data and found that the combination of random undersampling and a DNN classification provided superior prediction than other models. However, a logistic model using a much smaller set of only ‘known’ risk factors for MI was nearly as accurate in prediction, while all final models remained poorly calibrated. ML methods, specifically DNN, may have limited benefit over more traditional MI risk prediction tools when using harmonized data until more granular data can readily be incorporated. Our methodology and use of harmonized data is an important step for developing prediction tools that can be scaled into real-world clinical practice. Future studies should also address prediction of varying time horizons, assessing for improvements in model calibration with more extended prediction intervals.
Supplementary information
Additional file 1 Python code used for analysis. Supplemental Figure 1. Calibration curve for optimal model with results from re-scaling methods
Acknowledgements
Data was provided by Health Data Compass University of Colorado Anschutz Medical Campus.
Abbreviations
- CVD
Cardiovascular disease
- TIMI
Thrombolysis in Myocardial Infarction
- ML
Machine learning
- ACS
Acute coronary syndrome
- EHR
Electronic Health Record
- OMOP-CDM
Observation Medical Outcomes Partnership common data model
- SMOTE
Synthetic minority oversampling technique
- DNN
Deep neural network
- OHDSI
Observational Health Data Sciences and Informatics
Authors’ contributions
DM analyzed and interpreted the EHR data, model performance and calibration. PT performed analysis of the data, including implementation of in Python. MR guided project analysis, analyzed and interpreted model implementation and performance. SS and KC aided in result analysis and manuscript development. All authors read and approved the final manuscript.
Funding
This work was funded by grants from the National Institute of Health/NHLBI (MAR: 5 K23 HL127296).
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Ethics approval and consent to participate
Not applicable. Colorado’s IRB has declared this study not human subjects research (exempt) due to use of retrospective data and no human interaction with patients.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary information accompanies this paper at 10.1186/s12911-020-01268-x.
References
- 1.Global Burden of Cardiovascular Diseases Collaboration et al. The Burden of Cardiovascular Diseases Among US States, 1990–2016. JAMA Cardiol. 2018;3:375–389. doi: 10.1001/jamacardio.2018.0385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Wang Y, et al. Risk factors associated with major cardiovascular events 1 year after acute myocardial infarction. JAMA Netw Open. 2018;1:e181079. doi: 10.1001/jamanetworkopen.2018.1079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Yeh RW, Go AS. Rethinking the epidemiology of acute myocardial infarction: challenges and opportunities. Arch Intern Med. 2010;170:759–764. doi: 10.1001/archinternmed.2010.88. [DOI] [PubMed] [Google Scholar]
- 4.Liu N, et al. Prediction of adverse cardiac events in emergency department patients with chest pain using machine learning for variable selection. BMC Med Inform Decis Mak. 2014;14:75. doi: 10.1186/1472-6947-14-75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Steele AJ, Denaxas SC, Shah AD, Hemingway H, Luscombe NM. Machine learning models in electronic health records can outperform conventional survival models for predicting patient mortality in coronary artery disease. PLoS One. 2018;13:e0202344. doi: 10.1371/journal.pone.0202344. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Tay D, Poh CL, Van Reeth E, Kitney RI. The effect of sample age and prediction resolution on myocardial infarction risk prediction. IEEE J Biomed Health Inform. 2015;19:1178–1185. doi: 10.1109/JBHI.2014.2330898. [DOI] [PubMed] [Google Scholar]
- 7.Austin PC, Lee DS, Steyerberg EW, Tu JV. Regression trees for predicting mortality in patients with cardiovascular disease: what improvement is achieved by using ensemble-based methods? Biom J. 2012;54:657–673. doi: 10.1002/bimj.201100251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Mansoor H, Elgendy IY, Segal R, Bavry AA, Bian J. Risk prediction model for in-hospital mortality in women with ST-elevation myocardial infarction: a machine learning approach. Heart Lung. 2017;46:405–411. doi: 10.1016/j.hrtlng.2017.09.003. [DOI] [PubMed] [Google Scholar]
- 9.Shouval R, et al. Machine learning for prediction of 30-day mortality after ST elevation myocardial infraction: an acute coronary syndrome Israeli survey data mining study. Int J Cardiol. 2017;246:7–13. doi: 10.1016/j.ijcard.2017.05.067. [DOI] [PubMed] [Google Scholar]
- 10.Wallert J, Tomasoni M, Madison G, Held C. Predicting two-year survival versus non-survival after first myocardial infarction using machine learning and Swedish national register data. BMC Med Inform Decis Mak. 2017;17:99. doi: 10.1186/s12911-017-0500-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.LeCun Y, Bengio Y, Hinton G. Deep learning. Nature. 2015;521:436–444. doi: 10.1038/nature14539. [DOI] [PubMed] [Google Scholar]
- 12.Moravčík M, et al. DeepStack: expert-level artificial intelligence in heads-up no-limit poker. Science. 2017;356:508–513. doi: 10.1126/science.aam6960. [DOI] [PubMed] [Google Scholar]
- 13.Kooi T, et al. Large scale deep learning for computer aided detection of mammographic lesions. Med Image Anal. 2017;35:303–312. doi: 10.1016/j.media.2016.07.007. [DOI] [PubMed] [Google Scholar]
- 14.Song X, Mitnitski A, Cox J, Rockwood K. Comparison of machine learning techniques with classical statistical models in predicting health outcomes. Stud Health Technol Inform. 2004;107:736–740. [PubMed] [Google Scholar]
- 15.Payrovnaziri SN, Barrett LA, Bis D, Bian J, He Z. Enhancing prediction models for one-year mortality in patients with acute myocardial infarction and post myocardial infarction syndrome. Stud Health Technol Inform. 2019;264:273–277. doi: 10.3233/SHTI190226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Casey JA, Schwartz BS, Stewart WF, Adler NE. Using electronic health Records for Population Health Research: a review of methods and applications. Annu Rev Public Health. 2016;37:61–81. doi: 10.1146/annurev-publhealth-032315-021353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Chen Y, et al. Building bridges across electronic health record systems through inferred phenotypic topics. J Biomed Inform. 2015;55:82–93. doi: 10.1016/j.jbi.2015.03.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Huang Y, et al. Privacy-preserving predictive modeling: harmonization of contextual Embeddings from different sources. JMIR Med Inform. 2018;6:e33. doi: 10.2196/medinform.9455. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Barrett LA, Payrovnaziri SN, Bian J, He Z. Building computational models to predict one-year mortality in ICU patients with acute myocardial infarction and post myocardial infarction syndrome. AMIA Jt Summits Transl Sci Proc. 2019;2019:407–416. [PMC free article] [PubMed] [Google Scholar]
- 20.Sherazi SWA, Jeong YJ, Jae MH, Bae J-W, Lee JY. A machine learning-based 1-year mortality prediction model after hospital discharge for clinical patients with acute coronary syndrome. Health Informatics J. 2019:1460458219871780. 10.1177/1460458219871780. [DOI] [PubMed]
- 21.Hu D, et al. Evidential MACE prediction of acute coronary syndrome using electronic health records. BMC Med Inform Decis Mak. 2019;19:61. doi: 10.1186/s12911-019-0754-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Asaria M, et al. Using electronic health records to predict costs and outcomes in stable coronary artery disease. Heart. 2016;102:755–762. doi: 10.1136/heartjnl-2015-308850. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Weiss JC, Natarajan S, Peissig PL, McCarty CA, Page D. Machine learning for personalized medicine: predicting primary myocardial infarction from electronic health records. AI Mag. 2012;33:33. doi: 10.1609/aimag.v33i4.2438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Jiang G, Kiefer RC, Sharma DK, Prud’hommeaux E, Solbrig HR. A consensus-based approach for harmonizing the OHDSI common data model with HL7 FHIR. Stud Health Technol Inform. 2017;245:887–891. [PMC free article] [PubMed] [Google Scholar]
- 25.Jiang G, et al. Harmonization of detailed clinical models with clinical study data standards. Methods Inf Med. 2015;54:65–74. doi: 10.3414/ME13-02-0019. [DOI] [PubMed] [Google Scholar]
- 26.Makadia R, Ryan PB. Transforming the Premier Perspective Hospital Database into the Observational Medical Outcomes Partnership (OMOP) Common Data Model. EGEMS (Washington, DC) 2014;2:1110. doi: 10.13063/2327-9214.1110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Blagus R, Lusa L. SMOTE for high-dimensional class-imbalanced data. BMC Bioinform. 2013;14:106. doi: 10.1186/1471-2105-14-106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Agresti A, Coull BA. Approximate is better than ‘exact’ for interval estimation of binomial proportions. Am Stat. 1998;52:119–126. [Google Scholar]
- 29.Lakhani P, Langlotz CP. Automated detection of radiology reports that document non-routine communication of critical or significant results. J Digit Imaging. 2010;23:647–657. doi: 10.1007/s10278-009-9237-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Johansson S, Rosengren A, Young K, et al. Mortality and morbidity trends after the first year in survivors of acute myocardial infarction: a systematic review. BMC Cardiovasc Disord. 2017;17:53. doi: 10.1186/s12872-017-0482-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Weiss JC, Page D, Peissig PL, Natarajan S, McCarty C. Statistical relational learning to predict primary myocardial infarction from electronic health records. Proc Innov Appl Artif Intell Conf. 2012;2012:2341–2347. [PMC free article] [PubMed] [Google Scholar]
- 32.Xue S, et al. Circulating MiR-17-5p, MiR-126-5p and MiR-145-3p are novel biomarkers for diagnosis of acute myocardial infarction. Front Physiol. 2019;10:123. doi: 10.3389/fphys.2019.00123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Cross DS, McCarty CA, Steinhubl SR, Carey DJ, Erlich PM. Development of a multi-institutional cohort to facilitate cardiovascular disease biomarker validation using existing biorepository samples linked to electronic health records. Clin Cardiol. 2013;36:486–491. doi: 10.1002/clc.22146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Kusunose K, et al. A deep learning approach for assessment of Regional Wall motion abnormality from echocardiographic images. JACC Cardiovasc Imaging. 2019. 10.1016/j.jcmg.2019.02.024. [DOI] [PubMed]
- 35.Androulakis AFA, et al. Entropy as a novel measure of myocardial tissue heterogeneity for prediction of ventricular arrhythmias and mortality in post-infarct patients. JACC Clin Electrophysiol. 2019;5:480–489. doi: 10.1016/j.jacep.2018.12.005. [DOI] [PubMed] [Google Scholar]
- 36.Zadrozny, B. & Elkan, C. Transforming Classifier Scores into Accurate Multiclass Probability Estimates. Proceed Eighth ACM SIGKDD Int Conf Knowl Discov Data Mining 694–699 (ACM, 2002). 10.1145/775047.775151.
- 37.Niculescu-Mizil, A. & Caruana, R. Obtaining Calibrated Probabilities from Boosting.
- 38.Gibson WJ, et al. Machine learning versus traditional risk stratification methods in acute coronary syndrome: a pooled randomized clinical trial analysis. J Thromb Thrombolysis. 2019. 10.1007/s11239-019-01940-8. [DOI] [PubMed]
- 39.Tiwari, Premanand, Colborn, Katie, Smith, Derek, Xing, Fuyong, Gosh, Debashis, Rosenberg Michael. Development of a Prediction Model for Incident Atrial Fibrillation using Machine Learning Applied to Harmonized Electronic Health Record Data. BioRxiv [Preprint]. January 18, 2019. Available from: 10.1101/520866.
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Additional file 1 Python code used for analysis. Supplemental Figure 1. Calibration curve for optimal model with results from re-scaling methods
Data Availability Statement
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.


