Significance
Neurons of the cerebral cortex arise from a common progenitor pool that progressively generates different cell subtypes with distinct features, including deep layer (DL) and upper layer (UL) projection neurons. Several transcription factors (TFs) are known to specify whether a progenitor cell will become UL or DL; however, it is still unknown which upstream and downstream factors contribute to this fate decision. Here, we deeply characterize differential gene expression and chromatin accessibility of UL and DL neurons. We identify TFs that are highly differentially expressed and densely regulated and find that UL neurons retain the transcriptomic signature of their mother cells. We also discover a previously unknown role for the TF Myt1l in cell fate specification.
Keywords: transcription, gene regulation, cortical development, next-generation sequencing
Abstract
We are only just beginning to catalog the vast diversity of cell types in the cerebral cortex. Such categorization is a first step toward understanding how diversification relates to function. All cortical projection neurons arise from a uniform pool of progenitor cells that lines the ventricles of the forebrain. It is still unclear how these progenitor cells generate the more than 50 unique types of mature cortical projection neurons defined by their distinct gene-expression profiles. Moreover, exactly how and when neurons diversify their function during development is unknown. Here we relate gene expression and chromatin accessibility of two subclasses of projection neurons with divergent morphological and functional features as they develop in the mouse brain between embryonic day 13 and postnatal day 5 in order to identify transcriptional networks that diversify neuron cell fate. We compare these gene-expression profiles with published profiles of single cells isolated from similar populations and establish that layer-defined cell classes encompass cell subtypes and developmental trajectories identified using single-cell sequencing. Given the depth of our sequencing, we identify groups of transcription factors with particularly dense subclass-specific regulation and subclass-enriched transcription factor binding motifs. We also describe transcription factor-adjacent long noncoding RNAs that define each subclass and validate the function of Myt1l in balancing the ratio of the two subclasses in vitro. Our multidimensional approach supports an evolving model of progressive restriction of cell fate competence through inherited transcriptional identities.
The cerebral cortex is the region of the human brain responsible for perception, language, complex thinking, and motor control. Neurons in the cortex can be subdivided into two broad classes: Excitatory glutamatergic projection neurons and inhibitory GABAergic interneurons. Subclasses of cells exist within each of these broad classes. Recent efforts have sought to identify the complete catalog of cell types in the cortex using transcriptional profiling (1–4). At least 13 unique types of excitatory neurons have been identified in the developing mouse cortex (2), while the adult mouse cortex contains at least 52, a number that may rise with increased computational power to delineate cell types among single cells (1, 3).
While our ability to identify neuron subtypes contributes to our understanding of functional divisions within cortical circuits (5), how these circuits are specified during development remains unclear. Several recent studies have used single-cell RNA sequencing (scRNA-seq) to identify how differential gene expression over time establishes all of the identifiable cell subtypes in the developing brain (2, 6, 7). Delineating cell types using scRNA-seq profiles relies on dimensionality reduction with clustering and visualization tools, such as t-distributed stochastic neighbor embedding (t-SNE) (8) and uniform manifold approximation and projection (UMAP) (9). Default t-SNE, however, does not preserve global structure and therefore may miss biologically meaningful hierarchies in single-cell data (10). UMAP, on the other hand, was shown to preserve more local structure and continuity between cell subtypes than t-SNE, suggesting that UMAP improves on the utility of t-SNE for yielding insights into cell fate trajectories (9). A single cortical progenitor cell, for example, generates many daughter cells, each of which follows a specific trajectory to become a highly specialized cell type among a larger population with similar features. The cerebral cortex can be divided into two such large populations: Projection neurons of the deep layers (DL), layer 6 (L6) and L5, which are populated first, followed in order by projection neurons of the upper layers (UL), L4, then L2/3 (11). Although DL and UL subclasses are heterogeneous, cells within each subclass share common morphological and electrophysiological properties. The majority of DL neurons, for example, are corticofugal projection neurons (CFPNs), which send their axons to areas outside the cortex, including the thalamus and the spinal cord. The majority of UL neurons are cortico–cortical projection neurons (CPNs), many of which send their axons across the midline along the corpus callosum. While CPNs overwhelmingly occupy the ULs, they can also be found in the DLs.
A network of key transcription factors (TFs) expressed in early projection neurons determines whether a cell will acquire CFPN or CPN fate (12–14). Previous studies in the mouse revealed that TBR1 and FEZF2 regulate L6 and L5 CFPN development, respectively (15–22), while SATB2 is required for CPN fate (23–25). The upstream and downstream transcription cascade governing layer-specific maturation over time, however, is still unknown. No single study has directly assessed chromatin accessibility—an important indicator of gene regulation—and gene expression in CPNs or CFPNs during a specific stage of embryonic or early postnatal development. Moreover, it is still unknown how such transcriptional profiles differ between CPNs and CFPNs at comparable stages of development. One previous screen used bulk RNA-seq to identify genes that are differentially expressed between projection neuron subtypes on the same day of development in the mouse, when DL neurons are more developmentally mature than UL neurons (26).
Here, in order to identify transcriptional states that determine DL versus UL fate, we chose sequencing depth over sequencing breadth and analyzed cell type-specific gene expression plus differential chromatin accessibility over three defined stages of development in two sets of genetically defined neurons. We compared subclass- and stage-defining gene sets with subtype-defining gene sets identified using scRNA-seq and found broad agreement between our bulk-sequenced neuron populations and populations identified by scRNA-seq. Given the robustness of bulk sequencing, we also joined open chromatin with gene expression within these subclasses and identified densely regulated TFs with previously uncharacterized roles in specifying UL versus DL fate. In addition, we identified long-noncoding RNAs adjacent to important TF genes (TF-lncRNAs) with highly specific spatial and temporal expression patterns. Finally, we tested the ability of select candidate genes to modulate the ratio of CPN to CFPN neurons in vitro.
Results
Cell Subclass Is the Greatest Source of Variability in Gene Expression.
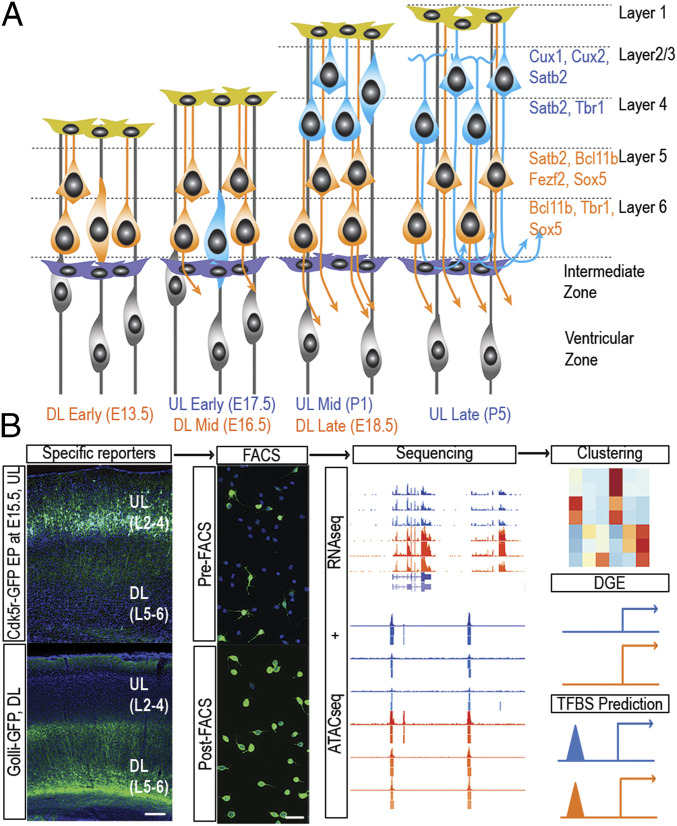
DL projection neurons (DL subclass, layers 5 and 6) are generated first and mature more quickly than UL projection neurons (UL subclass, layers 2 to 4) (Fig. 1A). Here, we focus on three stages of maturation common to DL and UL subclasses during the initial period after cell cycle exit: 1) “early,” when newly postmitotic cells are entering the cortical plate (CP); 2) “mid,” during primary axon extension; and 3) “late,” during axon collateral formation (27–29). We established a protocol for isolating DL and UL neurons at these three stages (Fig. 1B). For DL neurons, we used FACS of microdissected and dissociated mouse neocortex expressing the transgene golli-τ-EGFP (golli-EGFP), in which the 1.3-kb golli promoter of the myelin basic protein drives expression of EGFP in the subplate and in corticothalamic, corticospinal, corticocallosal, and corticocollicular neurons of layers 5 and 6 (30). FACS purification of GFP+ cells was performed on embryonic day (E) 13.5 (Theiler Stage [TS] 21; DL early), E16.5 (TS24; DL mid), and E18.5 (TS26; DL late). For UL neurons, we introduced a birth-dating plasmid containing the cyclin-dependent kinase 5 regulatory subunit p35 (Cdk5r) promoter driving GFP by in utero electroporation at E15.5, during the peak of UL neurogenesis. The Cdk5r promoter is expressed in neurons after cell cycle exit (31). We were therefore able to use GFP expression to FACS-purify UL (primarily L2/3) CPNs at E17.5 (TS25; UL early), postnatal day (P) 1 (UL mid), and P5 (UL late).
Fig. 1.
Excitatory projection neurons of the mouse neocortex can be subdivided into two broad classes, here called upper layer (UL) and deep layer (DL), each identified by a specific combination of TFs and labeled by an exclusive fluorescent reporter. (A) Schematic representation of neocortical development illustrating the stereotypical “inside-first, outside-last” sequence of cell fate specification and differentiation: DL neurons are generated first and reach milestones of development prior to UL neurons. (B) Experimental pipeline showing in vivo expression of subclass-specific reporters, FACS purification, and computational analysis. (Lower Left) Mouse cortex at P25 expressing Golli-GFP in DL neurons, L6 apical dendrites terminating in L4. (Upper Left) mouse cortex at P9 expressing Cdk5r-GFP in UL neurons occupying L2/3. (Scale bars, 100 μm for in vivo expression and 20 μm for dissociated cells.)
We used RNA-seq to quantify gene expression at early, mid, and late stages of DL and UL development for two biological replicates per stage per subclass and generated an average of 2.35 × 107 reads per replicate (range, 1.65 × 107 to 2.86 × 107) (SI Appendix, Fig. S1A). We have reported previously that two biological replicates per condition is sufficient to identify statistically significant differences at the genome-wide level (32, 33). These data have been deposited in the Gene Expression Omnibus (GEO) and can be accessed under accession no. GSE116147. Previous reports have suggested that in the developing neocortex, cell subclass does not contribute as much variability to gene expression as does developmental time (26). Recent scRNA-seq data, however, have demonstrated that gene expression can distinguish both subclass and developmental stage (1–3, 6, 34). To test whether subclass is the greatest source of variability in gene expression, we used hierarchical clustering on principal components (HCPC) (35), t-SNE, UMAP, and unsupervised hierarchical clustering on highly variable transcripts. HCPC of all samples (n = 12), using Ensembl mm9 transcripts with average transcripts per million (TPM) > 10 (12,026 transcripts), showed that biological replicates clustered together, with the exception of DL late replicate 1, which formed its own cluster (Fig. 2 A and B). Moreover, all DL and UL samples segregated along principal component (PC) 1, while all stages (early, mid, and late), segregated along PC2 (Fig. 2B). As alternatives to PC analysis, t-SNE showed even clearer clustering of each subclass, while UMAP maintained some of the local structure visualized by PC analysis, placing one DL early replicate and one DL late replicate closer to the UL cluster (SI Appendix, Fig. S1B).
Fig. 2.
HCPC and naive hierarchical clustering show that gene expression is able to distinguish cell subclass. (A–D) HCPC of all transcripts (A and B) and of all TF transcripts (C and D) shows that variance between subclasses is captured in the first PC and variance between stage is captured in the second PC. (E) Unsupervised hierarchical clustering of the top 500 most variable Ensembl mm9 transcripts separates UL from DL neurons and identifies a cluster of UL-like or noisy transcripts in DL neurons. Clusters labeled to the right are listed in SI Appendix, Table S1. (F) Shared expression of subclass-defining genes identified by bulk RNA-seq or scRNA-seq. (G) Hypergeometric P values of the overlap of subclass-defining genes identified by bulk RNA-seq and stage-defining L4 genes identified by scRNA-seq. Color-filled cells indicate significance after correcting for multiple comparisons. Darker shades indicate more overlap. (H) Predicted cell types contained within each subclass.
Due to our interest in transcription cascades, we asked whether limiting the HCPC analysis to transcripts encoding TFs would result in different clustering. HCPC of transcripts encoding TFs with an average TPM > 10 (482 transcripts), improved clustering such that all replicates clustered together (Fig. 2C). Interestingly, HCPC identified four broad clusters—DL early, DL mid/late, UL early, and UL mid/late—suggesting that TF gene expression within a subclass is more similar between middle and late stages of differentiation than between early and mid/late stages. Subclass contributed the most variability: All samples segregated by subclass along PC1 and by stage along PC2 (Fig. 2D). Moreover, hierarchical clustering on all TF transcripts clearly segregated DL and UL subclasses (SI Appendix, Fig. S1C), further suggesting that TF gene expression is a strong predictor of cell identity.
To determine which transcripts contributed the most variability between subclasses, we performed unsupervised hierarchical clustering on the 500 most variable transcripts in the dataset. Hierarchical clustering revealed that two of the six DL replicates (one early and one late) clustered more closely with the UL replicates than with the other four DL replicates (Fig. 2E). Given that DL neurons consist of both CPNs and CFPNs, while UL neurons primarily consist of CPNs alone, one explanation for this uneven clustering is that a subcluster of highly variable transcripts defines a population of “UL-like” CPNs that populates the DLs. An alternative explanation is that these transcripts represent biological or technical noise. To distinguish between these two hypotheses, we identified clusters of highly variable transcripts that differentiated UL, DL, UL-like, and DL subset populations. Each cluster contained bona fide lamination markers (SI Appendix, Table S1), including the UL TFs Nfix and Cux1 in the UL cluster, the DL TFs Sox5 and Tle4 in the DL cluster, the DL CPN marker Ptn in the UL-like cluster (36), and the DL-enriched Lmo3 in the DL-subset cluster (37). The three genes that encode subunits of the platelet-activating factor acetylhydrolase 1B complex (PAFAH1B2), an enzyme involved in neuron migration and synaptic function, were also differentially expressed: Pafah1b1 (Lis1) and Pafah1b2 clustered with DL transcripts, and Pafah1b3 clustered with UL transcripts, consistent with a previous report (38).
We next compared these four clusters with clusters of transcripts found to define cortical cell types identified using scRNA-seq of brain cells from P2 and P11 mouse (SPLiT-seq cells) (2) (Fig. 2F). DL and UL clusters shared expression exclusively with SPLiT-seq DL and UL cell types, respectively. The UL-like cluster contained Mpped1, a gene expressed across many SPLiT-seq cell types, while the DL-subset cluster contained Lmo3 and Dlg2, similar to the SPLiT-seq L5/6 Npr3 cell type. Together, these data suggest that the UL subclass contained L2/3 and L4 cells, while the DL subclass contained L5 and L6 cells. UL-like transcripts may represent a common Ptn+ CPN population, given that Mpped1 is expressed in 6 of 10 SPLiT-seq types, while DL-subset transcripts may represent L5/6 Sox5−, Tle4−, Npr3+ cells (Fig. 2H). Moreover, when we compared these clusters with genes expressed during different stages, or “waves,” of UL L4 differentiation, as defined in Telley et al. (39), UL and UL-like transcripts significantly overlapped L4 genes (wave 2 and wave 5) (Fig. 2G). DL transcripts less significantly overlapped L4 genes, but only during wave 2. Together, these data demonstrate that bulk-sequenced UL and DL populations represent distinct projection neuron subclasses transcriptionally similar to scRNA-seq–identified subclasses.
DL and UL Neuron Subclasses Can Be Defined by Consistently Significant Differential Expression of TFs over Time.
We next asked which transcripts were significantly different between DL and UL neurons at all three stages and whether such transcripts were enriched for known biological functions. We first identified all transcripts that were significantly higher in DL or UL neurons at all three stages (Q value ≤ 0.05 for each pairwise comparison) (SI Appendix, Table S2). The 199 transcripts that were consistently higher in DL neurons were enriched for genes involved in neuron commitment in the forebrain (gene ontology GO:0021902) and mRNA splice site selection (GO:0006376), while the 235 transcripts that were consistently higher in UL neurons were enriched for genes involved in glucocorticoid receptor signaling (GO:2000324) and postsynaptic membrane organization (GO:1901628). Within these two groups of transcripts, we identified 26 TFs that were consistently higher in DL neurons and 21 TFs that were consistently higher in UL neurons, including known, recently identified (Rprm for DL and TNC for UL) (40), and novel (Zfp521 for DL and Bhlhe40 for UL) (41, 42) layer-specific markers (Fig. 3 A–C and SI Appendix, Fig. S1D and Table S2).
Fig. 3.
Specific expression of a set of TFs throughout development distinguishes UL from DL populations. (A and B) Heatmaps of relative expression of TFs that are significantly higher in DL (A) or UL (B) neurons at all three stages of development. FL, full length. Transcripts with a black dot are shown in C. Underlined gene names are referenced in the human dataset in F. (C) In situ hybridization of the DL TF Zfp521 and the UL TF Bhlhe40 on sections through the embryonic mouse brain. High-magnification images of the boxed regions show restricted expression in the deep (Zfp521) or superficial (Bhlhe40) layers of the CP. (Scale bar, 800 μm for low-magnification images and 100 μm for high-magnification images.) Image credit: © 2009 and 2012 Allen Institute for Brain Science. Mouse Brain. Available from: developingmouse.brain-map.org. (D) Heatmap of relative expression of DL- and UL-specific early TFs. For all three heatmaps, each column represents the average TPM of two replicates. (E) FISH of the DL early TF St18 on sections through the mouse cortex showing enriched expression in L6 at E13.5 and reduced expression at E16.5. (Scale bar, 50 μm.) (F) Unsupervised hierarchical clustering of the top 500 most variable transcripts between embryonic and adult human motor cortex reveals expression of DL and UL TFs in the embryonic cluster and two UL TFs in the adult cluster. PCW, postconception weeks.
Given our finding that TF gene expression within a subclass was most distinct at early stages, we asked which TFs were differentially expressed between early DL and UL populations. We strictly defined an “early” transcript as significantly higher in the early stage compared with both the middle and late stages. TFs that were early in only one subclass were highly restricted to the early stage in that subclass and thus highly class- and stage-specific (Fig. 3D). Of the 10 early DL TFs, St18, a member of the myelin transcription factor (MYT) family of TFs, was the most differentially expressed between early and middle stages and between DL and UL subclasses. We confirmed early DL expression of St18 using FISH and observed that St18 was highly expressed in L6 at E13.5 and reduced dramatically by E16.5 (Fig. 3E).
St18 was also enriched in embryonic (12 postconception weeks) human motor cortex compared with adult motor cortex (36 to 40 y) according to the BrainSpan atlas (43) (Fig. 3F). Unsupervised hierarchical clustering of the top 500 most variable BrainSpan transcripts between the embryonic and adult motor cortex showed clear segregation of embryonic and adult gene expression and embryonic enrichment of five additional mouse subclass-specific TFs (Sox5, Neurog2, Bcl11b, Nfib, and Fezf2). Surprisingly, two mouse UL TFs, Hopx and Bhlhe40, were enriched in the adult human motor cortex. Given its UL-specificity and enriched expression in the adult human motor cortex, we searched for potential protein binding partners of BHLHE40 using the STRING database (44). A regulator of circadian rhythm, BHLHE40 is known to interact with several clock proteins (SI Appendix, Fig. S2A), including the UL-enriched Arntl (SI Appendix, Fig. S2B and Table S8), and the UL-enriched TF Epas1 (SI Appendix, Table S2), both of which were found to be UL-expressed postnatally in the mouse, similar to the expression pattern of Bhlhe40 (SI Appendix, Fig. S2 C and D).
Subclasses of Projection Neurons Have Common and Specific Expression Dynamics.
Common gene-expression dynamics may reveal shared biological function. To identify groups of genes that were common to both subclasses at a specific stage, we compared gene expression across time and space to arrive at sets of genes that defined early, mid, and late stages. We defined early and late as being significantly higher (Q value ≤ 0.05, log2 fold-change [log2FC] ≥ 1.5) in the early or late stage compared with the middle stage, regardless of expression at the other end. Similarly, we defined a middle transcript as being significantly higher in the middle stage compared with the early stage but not with the late stage. There were more common early TFs than common middle or late TFs, and many of these early TFs, such as Tcf12, were highly restricted to the early stage in both subclasses, although there was a noticeable difference between DL-enriched (i.e., Nhlh1) and UL-enriched (i.e., Zfp664) early genes (SI Appendix, Fig. S3A and Table S3). Moreover, several TFs with known roles in neural progenitor proliferation—such as Pax6, Eomes, Gli2, and Gli3—showed early-restricted expression in both DL and UL cells, suggesting that postmitotic neurons maintain residual transient expression of progenitor genes shortly after they exit the cell cycle. In contrast, only two middle TFs (Mef2c and Mkx) and one late TF (Thrb) were shared between subclasses. Similarly, DL and UL neurons shared early expression of eight axon guidance genes, but late expression of only one gene axon guidance gene (Cntn4) (SI Appendix, Fig. S3C). We also identified middle and late TFs specific to each subclass (SI Appendix, Fig. S3B and Table S4). Of the 16 late UL TFs, six (37.5%) have known roles in circadian control of gene expression (Per1, Npas2, Rora, Rorb, Nr1d2, and Arntl). Together with the above finding that Bhlhe40 is UL-enriched and interacts with other circadian clock proteins, these data suggest that a network of cortical circadian genes is expressed specifically in UL excitatory neurons from P5.
A recent study used scRNA-seq of embryonic mouse brain to identify cohorts of genes that define stages of neuron development and concluded that differentiation programs are conserved across cortical neuron cell types; early genes are enriched for nuclear-localized transcriptional and cell cycle regulators, while later-expressed genes are enriched for axon, dendrite, and synaptic functions (6). GO analysis (45) of our common stage-specific transcripts showed a similar shift from early transcriptional regulation by the Smoothened pathway to synaptic signaling and positive regulation of cell communication (interactions between a cell and its surroundings) over developmental time (SI Appendix, Fig. S3 C and D). We therefore asked if there was significant overlap of our stage-defining gene sets and those identified among single cells in the above scRNA-seq study (SI Appendix, Fig. S3E). We found that there was significant overlap, after correcting for multiple comparisons, of our early genes and genes enriched in late basal progenitors and 1-d-old neurons. Similarly, our middle genes most significantly overlapped genes enriched in 1-d-old neurons and 4-d-old neurons but not basal progenitors. Finally, our late genes overlapped two classes of 4-d-old neuron gene sets but not any of the younger gene sets. These data suggest that our early, middle, and late gene sets are consistent with early, middle, and late genes identified by scRNA-seq.
TF-lncRNAs Have Cell-Specific Expression Dynamics.
Highly specific expression of a transcript at a single stage of development may indicate a specific developmental function. In order to identify genes that were highly specific at one stage, we looked for transcripts that “spiked” at the middle stage of development, or showed high expression at the middle stage relative to early and late. We found only a few transcripts that spiked in either DL neurons or UL neurons using two different methods for differential expression analysis: 1) kallisto+Sleuth (Q value ≤ 0.05), consistent with the above analysis, and 2) STAR+Cuffdiff (Q value ≤ 0.05, log2FC ≥ 1.43), an approach that predated the development of kallisto+Sleuth. Both approaches yielded fewer spike transcripts for DL neurons compared with UL neurons, and while there was no overlap for DL neurons between Cuffdiff and Sleuth-identified spike transcripts, almost half of the Sleuth-identified transcripts for UL neurons were identified by the Cuffdiff (SI Appendix, Table S5).
We took a closer look at the 11 transcripts that spiked in DL neurons by either analysis and noticed a trend: The reads that tended to spike at E16 appeared enriched in noncoding regions (introns and 3′UTRs) or genes sharing an apparent bidirectional promoter with a TF gene. Three of the 11 transcripts encoded lncRNAs, each adjacent to a major cortical TF gene. Here, we name these transcripts after their respective adjacent TF genes: Satb2-lncRNA (9130024F11Rik), Pou3f3-lncRNA (2900092D14Rik), and Pou3f2-lncRNA (AK039117). In DL neurons, Satb2-lncRNA and Pou3f3-lncRNA expression paralleled the expression of their neighboring TF genes over time, supporting previous observations that lncRNA expression is often correlated with that of a proximal gene (46–49). Of these three transcripts, only Satb2-lncRNA spiked in both DL and UL neurons (Fig. 4A, blue dashed line).
Fig. 4.
Satb2-lncRNA is differentially expressed in cortical neurons over time and space. (A) TF-lncRNAs show increased expression (in TPM) during the middle stage in DL neurons, and Satb2-lncRNA shows a similar spike in UL neurons. (B–D′′) FISH showing parallel expression of Satb2-lncRNA (B–D) and Satb2 (B′–D′; merged channels in B′′–D′′) in the mouse neocortex at E13.5, E16.5, and P0. (Scale bar in D′′ is 200 μm for B–D′′) (E–H) FISH of Satb2-lncRNA colabeled with Satb2 RNA at P3 (E and F), SATB2 protein (G), or CTIP2 protein (H) showing expression in UL nuclei (arrow) and cell bodies (arrowhead) (E), DL nuclei (arrow) (F), and most SATB2+ (arrowhead in G), but few CTIP2+ (arrowhead showing colabeling versus arrow showing CTIP+/lncRNA− cell in H) in cultured mouse cortical cells. (Scale bar shown in H is 50 μm and applies to E–H.) (I and J) Fewer DL cells express Satb2-lncRNA compared with UL cells (I) (n = 3), and only about a quarter of CTIP2+ cells express Satb2-lncRNA, while almost all SATB2+ cells express the lncRNA (J) (n = 6). Error bars represent SD. *P < 0.05. ***P < 0.0001.
Two lncRNAs that flank Pou3f3 have been shown to regulate cortical progenitor proliferation and fate in mice (50, 51). We therefore hypothesized that Satb2-lncRNA may play a similar role in cell fate. We used two-color FISH to analyze the localization of Satb2-lncRNA relative to Satb2 (Fig. 4 B–D′′). Using a probe that targeted the first two exons of the lncRNA, and therefore both isoforms, we found that Satb2-lncRNA expression paralleled that of Satb2 in the neocortex. Both transcripts were expressed in the CP and ventricular zone at E13.5, becoming restricted to the CP and intermediate zone by E16.5 and enriched in UL cells by P0. Confocal imaging confirmed that individual cells coexpressed both Satb2 and Satb2-lncRNA; however, in DL neurons, their expression was restricted to the nucleus, but in UL neurons their expression was both nuclear and cytoplasmic (Fig. 4 E and F). While most DL cells expressed Satb2-lncRNA at P3, almost all UL cells expressed Satb2-lncRNA at P3, a difference that was statistically significant (Fig. 4I) (P = 0.04, n = 3). Moreover, in cultured mouse cortical neurons, Satb2-lncRNA was expressed in a significantly greater proportion of SATB2+ cells (∼95%) than CTIP2+ cells (∼24%) (Fig. 4 G, H, and J) (P = 3.2 × 10−7, n = 6), consistent with previous reports that ∼20% of CTIP2+ cells coexpress SATB2 between E16.5 and P4 (25).
These results suggest that Satb2-lncRNA may help regulate the ratio of SATB2+ CPNs to CTIP2+ CFPNs. To test this hypothesis, we designed two small-interfering RNA (siRNA) constructs targeting the 3′ end of the first exon of Satb2-lncRNA, which is shared by the short and long isoforms and does not overlap the first exon of Satb2 (SI Appendix, Fig. S4A). We confirmed diminished cytoplasmic localization and efficient knockdown (KD) of both isoforms in dissociated cortical neurons cultured for 72 h (SI Appendix, Fig. S4 B and C). Two-color FISH did not reveal changes to Satb2 transcript localization or relative levels after KD (SI Appendix, Fig. S4C). To evaluate the effect of KD on cell fate, we quantified the ratio of SATB2+ to CTIP2+ cells in control (scrambled siRNA) and KD samples (SI Appendix, Fig. S4 D and E). The number of cells expressing SATB2 relative to CTIP2 was reduced modestly but not significantly (P = 0.055) in KD cells compared with control cells. Together, these results suggest that Satb2-lncRNA may be involved in regulating the balance of CPNs to CFPNs; however, cytoplasmic-specific reduction of Satb2-lncRNA had a minor but statistically insignificant effect on cell fate.
ATAC-seq of Subclasses of Projection Neurons Reveals Few Persistent Subclass-Specific Regulatory Elements.
TFs can bind DNA and modulate gene expression in regions where chromatin is accessible to transposase insertion, at putative promoters, enhancers, and repressors. To identify additional pathways involved in regulating the balance of CPNs to CFPNs, we used an assay for transposase accessible chromatin (ATAC-seq) to compare chromatin accessibility between DL and UL neurons at early, middle, and late stages (Fig. 5) (52). To be consistent with the RNA-seq, we used two biological replicates per stage per subclass and generated an average of 7.11 × 107 reads after deduplication per replicate (range, 1.32 × 107 to 1.56 × 108) (SI Appendix, Fig. S5A). ATAC-seq datasets are available in the GEO (accession no. GSE116147).
Fig. 5.
Densely regulated TFs suggest subclass-specific transcriptional programs. (A) Fragment length density for each ATAC-seq library; two biological replicates per stage represented by solid and dotted lines of the same color. (B) ATAC-seq peaks significantly overlap previously identified cortical enhancers and repressors. (C) Top conserved P300-bound enhancers and putative target genes in each subclass. (D) Binomial test showing that a subclass-specific ATAC-seq peak is more likely to occur in the regulatory domain of a gene more highly expressed in the same subclass. (E) Enriched motifs and TF consensus sites in ATAC-seq peaks of densely regulated DL and UL genes. (F) Differentially expressed TFs between DL and UL neurons plotted by log2(fold-change) in expression (x axis) and binomial P value (y axis). Each comparison is UL/DL. TFs with the greatest change in binomial P value or referenced in Figs. 5 and 6 are labeled. (G–J′) In situ hybridization of select densely regulated TFs. Image credit: (Right to Left) © 2008 and 2009 Allen Institute for Brain Science. Mouse Brain. Available from: developingmouse.brain-map.org. The arrowhead in G′ indicates residual Nhlh2 expression in the hindbrain. (Scale bar in J′ is 700 μm for G and H, 1 mm for G′ and H′, and 1.1 mm for I–J′.)
Our complete set of early, middle, and late DL and UL ATAC-seq peaks was significantly enriched for previously identified enhancers and repressors in the mouse brain: 93.2% of cortical enhancers bound by P300 (P < 1 × 10−4), 60.5% of cortical repressors marked by H3K27me3 (P < 1 × 10−4), and 6.1% of cortical repressors marked by H3K9me3 (P < 2 × 10−2) overlapped our set of ATAC-seq peaks, confirming that these peaks represented both activating and repressing chromatin states (Fig. 5B). After masking for blacklist regions, repeat regions, segmental duplications, and exons, we identified ATAC-seq peaks that were present in both replicates of each cell subclass at each stage (“replicate peaks”). The number of replicate peaks decreased between the early and middle/late stages for both DL and UL neurons (SI Appendix, Table S6). We identified 1,082 peaks that were present in DL neurons and absent in UL neurons at all three stages, and 185 peaks that were present in UL neurons and absent from DL neurons at all three stages (SI Appendix, Table S7). We used GREAT (genomic regions enrichment of annotations tool) regulatory domains to associate DL-specific regions with 1,472 nearby genes and UL-specific regions with 320 nearby genes and found that only 23 genes occurred in both lists (53) (SI Appendix, Table S7). We plotted by heatmap the mean TPM of all transcripts of genes with peaks that were specific to a subclass across all stages and found that genes with UL-specific peaks were consistently higher in UL cells, while genes with DL-specific peaks were consistently higher in DL cells at early and middle stages, but not late stages (SI Appendix, Fig. S5 B and C). Genes common to these two lists showed no consistently higher expression in one subclass (SI Appendix, Fig. S5D).
Within these subclass-specific sets, 31 DL peaks and 37 UL peaks overlapped repressors (H3K27me3 chromatin immunoprecipitation sequencing [ChIP-seq]), and 7 DL peaks and 9 UL peaks overlapped enhancers (P300 ChIP-seq) previously identified in mouse embryonic whole brain (32, 54). We hypothesized that subclass-specific enhancers may represent regions important for establishing DL versus UL fate. We therefore used the GREAT regulatory domains to associate these 13 subclass-specific enhancers with nearby genes (SI Appendix, Table S8) and identified a UL-specific enhancer just downstream of Khdrbs1/Sam68, which regulates alternative splicing of Nrxn1 in the mouse brain (55). These findings suggest that regulatory elements with a role in establishing cell fate may be more accessible in one subclass over another; however, such subclass-wide accessibility appears to be rare.
Despite the lack of sustained subclass-specific enhancer accessibility over developmental time, we hypothesized that subclass-specific ATAC-seq peaks may be enriched near subclass-specific genes at various stages throughout development. An analysis of fragment length density revealed similar size distributions between early DL and early UL ATAC-seq sets, both of which had the highest numbers of replicate peaks (Fig. 5A). TF gene expression was also the most distinct at early stages (Fig. 2 C and D). We therefore looked for subclass-specific regulatory elements in early DL and UL neurons by identifying early DL-specific and UL-specific ATAC-seq peaks overlapping cortical P300 ChIP-seq peaks and ranking them by conservation (SI Appendix, SI Materials and Methods). Of the top five most-conserved peaks for each class, two were in the regulatory domains of known fate-determining genes (Sox5 and Tbr1), while four were in the regulatory domains of genes with differential expression in the current study (Sox21, Zfhx4, St18, and Fbn1) (Fig. 5C). These findings suggest that subclass-specific gene expression may be densely regulated. To test this hypothesis, we identified genes that were significantly differentially expressed between early DL and early UL neurons (Q value < 0.05), establishing a set of 4,673 subclass-specific genes (1,967 DL and 2,706 UL). We next identified all subclass-specific ATAC-seq peaks in the regulatory domains of these genes, establishing a set of 1,869 DL-specific and 3,919 UL-specific peaks. Finally, we calculated the likelihood that a subclass-specific ATAC-seq peak would be found in the regulatory domain of a gene specific to its same subclass, using a binomial P value, which accounts for differences in regulatory domain sizes (Fig. 5D). DL-specific peaks were more likely to be found in the regulatory domain of a DL-specific gene (binomial P value: 0.11); likewise, UL-specific peaks were more likely to be found in the regulatory domain of a UL-specific gene (binomial P value: 5.06 × 10−16). Together, these data confirm that subclass-enriched genes are densely regulated in the cells in which they are more highly expressed.
Densely Regulated TFs Include Known and Novel Regulators of Cell Fate in the Cortex.
TFs that are highly differentially expressed and densely regulated in one subclass may play subclass-specific roles. We therefore searched for densely regulated TFs expressed in DL and UL cells. Using the above sets of subclass-specific genes and subclass-specific ATAC-seq peaks, we calculated two binomial P values for each gene—a binomial P value for the number of DL-specific ATAC-seq peaks in the gene’s regulatory domain, and a binomial P value for the number of UL-specific ATAC-seq peaks in the gene’s regulatory domain—and ranked genes by the log fold-change in these two binomial P values (Fig. 5F). When we limited the ranking to genes with a log fold-change in gene expression greater than 1 or less than −1, and looked for densely regulated DL and UL TFs (differentially expressed TFs with a corresponding difference in binomial P value), we found that densely regulated DL TFs included genes known to be involved in DL cell fate specification (Fezf2 and Nhlh2) (Fig. 5F and SI Appendix, Table S9). The DL TF with the greatest log fold-change in ATAC-seq P value was the imprinted gene Peg3 (56), although its expression in DL neurons was less than onefold higher than in UL neurons. The most densely regulated UL TFs included genes known to be involved in neural progenitor cell cycle regulation [Tcf3 (57), Tcf4, and Hes5 (58)], neuron differentiation (Nfix and Cux2), and positioning of UL neurons within the CP [Prdm16 (59)] (Fig. 5F).
We next searched for enriched TF binding motifs in ATAC-seq peaks in densely regulated DL- or UL-specific genes. We limited the analysis to genes with a log fold-change in binomial P value greater than 2 and ran seven different published motif discovery tools on each set of peaks (32). Near identical motif predictions from at least two different tools were combined, and Tomtom (60) was used to match the most frequently occurring motif with a set of known TF binding motifs. In each subclass, the top predicted motifs matched TFs found in our set of densely regulated genes (Fig. 5 E and F): For DL, the consensus motif for the bHLH TFs NEUROD and NHLH, both of which were more highly expressed and densely regulated in DL neurons, and the TGIF consensus motif were identified. Tgif2 is expressed in the ventricular zone of the neocortex (61), was a common early TF (Fig. 3E), and exhibited higher expression and chromatin accessibility in the UL subclass (Fig. 5E). The top predicted UL motif resembled the consensus motifs for the progenitor TF TEAD2 (61, 62), the intermediate progenitor retinoic acid cofactor ZFP423 (63), and the bHLH oligodendrocyte specification TF OLIG2 (64), all three of which were more highly expressed in the UL subclass (Fig. 5E).
The enrichment of progenitor genes, such as Tead2, Zhx2, and Olig2, in the emerging UL transcriptional network was particularly intriguing. Expression of bHLH TFs, such as Neurod1 and Nhlh2, typically peaks after cell cycle exit, with down-regulation of progenitor genes such as Tead2 in postmitotic neurons, suggesting that the UL population might be slightly behind the DL population in its developmental trajectory. On the other hand, a recent scRNA-seq study suggested that apical progenitor cells become more neuron-like over developmental time and placed the intermediate progenitor TF Eomes in the same developmental gene cluster as Neurod1, raising the intriguing possibility that neuron diversity is generated starting in apical progenitor mother cells, which transmit age-dependent transcriptional identities to their daughter cells (6). Daughter cells could therefore maintain a transient progenitor-like transcriptomic signature. To address the hypothesis that the enrichment of progenitor TFs in UL genes represents a UL-specific transcriptional program, we analyzed expression of select DL and UL genes in the publicly available Allen Developing Mouse Brain Atlas (https://developingmouse.brain-map.org/). We found that the DL gene Nhlh2 was more highly enriched in the CP at E13.5, during DL neurogenesis (Fig. 5G), than at E15.5, at the height of UL neurogenesis (Fig. 5G′). In contrast, the UL gene Zhx2, a known progenitor TF with similar log fold-change in expression to the UL TF Cux2 (65, 66), was more highly expressed in the CP at E15.5, at the height of UL neurogenesis, than at E13.5, near the end of DL neurogenesis (Fig. 5H vs. Fig. 5H′), suggesting a UL-specific role in cell fate. As a counter to the transient expression of Nhlh2 and Zhx2, the densely regulated DL gene Fezf2 and the densely regulated UL gene Nfix persisted in their respective layers from embryonic through postnatal stages (Fig. 5 I and J′). Together, these data are consistent with the evolving hypothesis that cell fate-specific developmental programs are inherited from specific progenitor TF networks in apical progenitor cells whose daughter neurons maintain progenitor-like and neuron-specific transcriptomic signatures.
Consistent with the hypothesis that late progenitor cells have distinct TF networks from early progenitor cells, we observed strong Olig1 and Olig2 signatures in the UL population, which are derived from the same progenitor population that gives rise to oligodendrocytes. RNA-seq read pile-ups that mapped to the Olig2 locus showed enriched expression in the 3′UTR in the middle and late stages, suggesting that cells fated to become UL neurons may inherit low expression of Olig2 mRNA after cell cycle exit and retain expression of the 3′UTR, as has been demonstrated for other transcripts in the nervous system (SI Appendix, Fig. S6A) (67). In order to determine whether a subset of Cdk5r-GFP+ cells were in fact oligodendrocyte progenitor cells (OPCs), we quantified the percentage of Olig2+ cells in the Cdk5r-GFP+ population and found that a small subset of Cdk5r-GFP+ cells (1.2 to 4.0% of the GFP+ population) (SI Appendix, Fig. S6B) maintained expression of Olig2 as late as P9 (SI Appendix, Fig. S6 C–F). A similarly small fraction expressed the progenitor markers Ki-67 and SOX2 (SI Appendix, Fig. S6 G and H), suggesting that the UL population inherits late progenitor TF signatures that confer OPC competence.
Myt1l Regulates the Ratio of UL to DL Neurons.
To identify previously untested TFs that contribute to transcription cascades that determine CFPN versus CPN fate, we searched for TFs with higher expression and enriched chromatin accessibility in a single-cell class. We found that members of the MYT family of TFs meet these criteria. MYT zinc-finger proteins include MYT1/NZF2, MYT1L/NZF1, and ST18/MYT3/NZF3. MYT1 and MYT1L play crucial roles in promoting neuronal differentiation in the telencephalon through repression of progenitor genes, thus antagonizing proliferation (68, 69). Above, we showed that St18 was expressed in early DL cells and had a highly conserved UL enhancer in its regulatory domain, Myt1 was a specific DL early gene (SI Appendix, Table S2), and Myt1l had an isoform that was exclusively expressed in UL cells at all three time points (SI Appendix, Table S4). Moreover, the Myt1l regulatory domain had a UL-specific ATAC-seq peak at all three stages (SI Appendix, Table S7). Layer-specific function of MYT family TFs in postmitotic neurons, however, has not been examined.
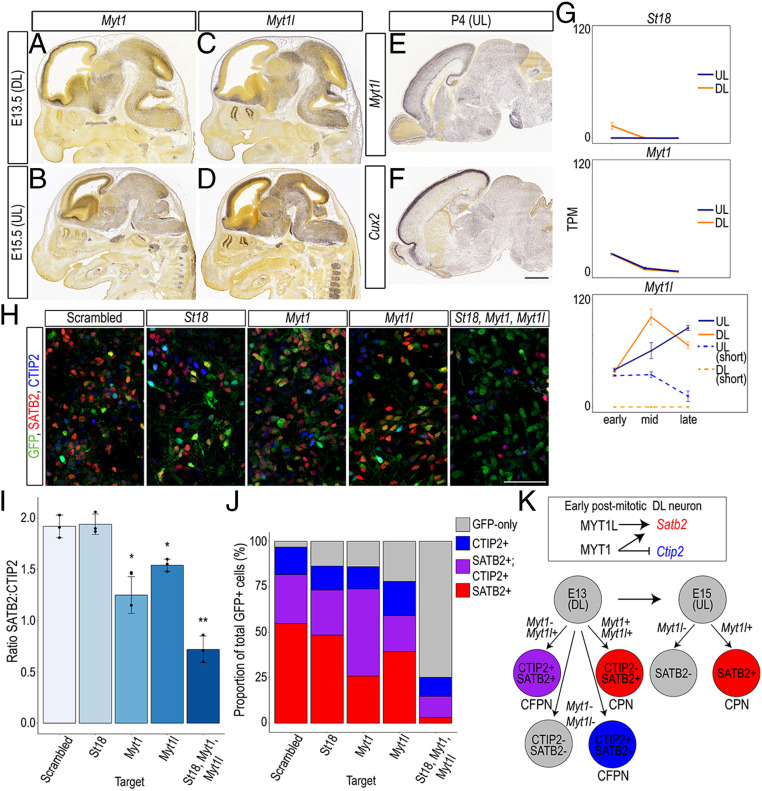
To determine whether MYT family TFs play a role in cortical cell fate specification, we first confirmed in vivo expression of Myt1 and Myt1l in the CP at E13.5 (Fig. 6 A and C) during DL neurogenesis. Myt1 expression was then excluded from the superficial CP at E15.5 (Fig. 6B), where Myt1l was enriched at E15.5 (Fig. 6D) and P4 (Fig. 6E), similar to Cux2 at P4 (Fig. 6F). These findings were consistent with our RNA-seq analysis showing decreasing Myt1 expression throughout DL and UL neuron differentiation, increasing Myt1l expression over DL and UL differentiation, consistently higher expression of a short Myt1l isoform (ENSMUST00000092649) in UL neurons (Fig. 6G), and a previous finding that Myt1l expression clusters with genes that are higher in UL neurons (cluster 15) (26).
Fig. 6.
MYT family TFs show stage- and subclass-specific expression in vivo and regulate subclass-specific genes in vitro. (A–F) The MYT family TFs Myt1 and Myt1l show early expression in the mouse CP (A and C) but diverge at E15.5 such that Myt1 expression is excluded from the superficial CP (B), while Myt1l is enriched in superficial layers (D). Myt1l remains enriched in UL cells through P4 (E) similar to that of the UL TF Cux2 (F). (Scale bar in F is 800 μm for A and C, and 1.2 mm for B and D–F.) Image credit: (Right to Left) © 2009 Allen Institute for Brain Science. Mouse Brain. Available from: developingmouse.brain-map.org. (G) St18 and Myt1 expression decline throughout neuron differentiation, while Myt1l expression increases throughout UL differentiation, and one isoform of Myt1l is consistently higher in UL cells. (H) pSico-GFP (green), SATB2 (red), and CTIP2 (blue) expression in cultured cortical neurons after lentiviral transfection of a shRNA targeting a scrambled control, St18, Myt1, Myt1l, or all three on E13.5 + 1 d in vitro. (Scale bar, 100 μm.) (I) shRNA KD of Myt1 or Myt1l but not of St18 significantly reduced the ratio of SATB2+ to CTIP2+ cortical neurons compared with KD of a scrambled control target. shRNA KD of all three Myt family TFs very significantly reduced the ratio of SATB2+ to CTIP+ neurons. *P < 0.02. **P < 0.001. n = 3 biological replicates. (J) The proportion of SATB2;CTIP2 double-positive cells was increased after Myt1 KD, causing a reduction in the proportion of SATB2-only cells (P = 0.02). The proportion of all SATB2+ cells was decreased after Myt1l KD (P = 0.05). KD of all three MYT family TFs resulted in a significant reduction of SATB2+ cells (P = 0.0005) and a very significant increase in GFP-only cells (P = 0.006). (K) Model of how MYT1 and MYT1L may coregulate Satb2 expression, and MYT1 may antagonize Ctip2 expression, in developing DL neurons, and how MYT1L may regulate Satb2 expression in developing UL neurons. Error bars represent SD.
We tested the function of all three MYT family TFs by short-hairpin RNA (shRNA) KD in cortical neurons cultured from E15.5 mice using previously validated shRNA lentiviral constructs (69). To evaluate a difference in the relative abundances of CPN and CFPNs, we counted the ratio of SATB2 to CTIP2 5 d after transfection (SI Appendix, Fig. S7 A and B). Compared with control cells transfected with scrambled shRNA, cells transfected with shRNA to St18 or Myt1 showed no significant difference in the ratio of SATB2 to CTIP2, after correcting for multiple comparisons. Cells transfected with shRNA to Myt1l, however, showed a significant decrease in the ratio of SATB2 to CTIP2 (P = 0.01), indicating a potential role in promoting Satb2 expression and CPN fate (SI Appendix, Fig. S7B). To further characterize the shift in the ratio of SATB2 to CTIP2, we quantified the proportion of transfected (GFP+) cells that was CTIP2+, SATB2+, or colabeled and found that there was a significant decrease in the proportion that was SATB2+ (P = 0.04) and a slight, although statistically insignificant, increase in the proportion that was CTIP2+ (P = 0.06), suggesting that MYT1L promotes Satb2 expression in early postmitotic UL neurons (SI Appendix, Fig. S7C).
Given that Myt1 and St18 were highest in early DL neurons (compared with middle and late), we asked whether MYT family TFs regulate cell fate at an earlier time point, during peak DL neurogenesis. We tested the function of all three MYT family TFs by shRNA KD in cortical neurons cultured from E13.5 mice (Fig. 6 H–J). KD of St18 again had no effect on the ratio of SATB2 to CTIP2; however, KD of Myt1 or Myt1l significantly reduced the ratio of SATB2 to CTIP2 (P = 0.009 and 0.016, respectively). Moreover, KD of all three MYT family TFs reduced the ratio even further (P = 0.0003). When we quantified the proportion of transfected cells that was SATB2+, CTIP2+, colabeled, or GFP-only, we found that Myt1 KD increased the proportion that was colabeled, which decreased the proportion that was SATB2+;CTIP2− (P = 0.02), while Myt1l KD decreased the total proportion that was SATB2+ (P = 0.05). KD of all three drastically reduced the total proportion that was SATB2+ (P = 0.0005) and increased the proportion that was GFP-only (P = 0.006). Collectively, these results suggest that MYT1L and MYT1 coregulate Satb2 and Ctip2 expression in early DL cells; while both promoted Satb2 expression, only Myt1 KD affected Ctip2 expression (Fig. 6K). We therefore propose a model of DL fate specification in which MYT1 and MYT1L promote Satb2+ and CPN (SATB2+;CTIP2−) fate, and MYT1 antagonizes CFPN (SATB2+;CTIP2+) fate. In UL cells, however, MYT1L was the primary MYT TF involved in promoting CPN fate (Fig. 6K).
Discussion
We have identified distinct transcriptional states of genetically defined DL and UL cortical projection neurons over time. The heterogeneity of each of these bulk populations allowed us to compare intragroup versus intergroup similarity over several stages of development, which revealed that gene expression alone can distinguish DL from UL identity, especially when limited to TF genes. Moreover, subclass, rather than developmental stage, contributed the highest source of variability in gene expression, and subclass and stage-specific gene expression significantly overlapped subclass- and stage-defining genes identified by scRNA-seq. Therefore, despite the intrinsic heterogeneity of bulk-sorted cells, neurons within each subclass retained significant intraclass similarity. We were therefore able to leverage sequencing depth over sequencing breadth to deeply compare DL and UL gene signatures.
All replicates within each subclass clustered together when limited to transcripts above a noise threshold. When limiting the analysis to the top 500 most variable transcripts, however, two samples of DL neurons clustered more closely with UL neurons. This result could suggest that DL neurons are more plastic early in development, consistent with the observation that progenitors fated to become DL neurons transplanted into an older host cortex have the capacity to become UL neurons, while UL progenitors transplanted into a younger host are fate-restricted (70–72). Alternatively, highly variable transcripts that cluster some DL neurons with UL neurons (UL-like transcripts) could represent technical noise or a population of CPNs that is similar across upper and deep layers. Indeed, the UL-like transcript Mpped1 was present in 60% of subtypes identified by scRNA-seq.
Very few regions of accessible chromatin were specific to one subclass at all three stages. This result is perhaps unsurprising given the heterogeneity of the two subclasses. In fact, that hundreds of regions were specific to a subclass throughout several developmental stages is unexpected when considering the vast diversity of cell types in the brain with distinct transcriptional programs (1, 73), and that the majority of open chromatin occupies promoter regions, which often remain open in cell types in which the downstream gene is not expressed. Our finding that later-born UL neurons retained a progenitor-like gene network signature enriched for genes involved in cell-extrinsic processes supports the recent finding that conserved postmitotic differentiation programs are applied to transcriptional “ground states” present in apical progenitor cells, and that these transcriptional ground states transition from cell-intrinsic to environment-sensing functions in both mother and daughter cells (6).
In contrast, from a cell-intrinsic view, DL and UL neurons expressed lncRNAs adjacent to major cortical TFs, suggesting a role for these lncRNAs in cell fate specification. Satb2-lncRNA expression spiked shortly after neurons reached the CP, and transcripts became differentially localized to the nucleus of SATB2+ DL neurons and the nucleus and cytoplasm of SATB2+ UL neurons. More evidence will be required to ascertain whether TF-lncRNAs play a cell type-specific role in cortical development, however. Numerous studies have suggested that lncRNAs can influence the expression of adjacent genes directly, through recruiting transcriptional or splicing machinery to the locus (74–76) or binding to chromatin to modulate enhancer–promoter contacts (77). Alternatively, lncRNA expression can regulate nearby genes indirectly, the process of transcription itself altering chromatin structure in a way that affects expression of the adjacent gene independently of the lncRNA transcript (49, 78–80). KD of the lncRNA at the mRNA level, therefore, would not affect expression of the nearby regulated gene, as we observed here. The statistically insignificant reduction in the number of SATB2+ cells relative to CTIP2+ cells after Satb2-lncRNA KD could indicate that Satb2-lncRNA is only a byproduct of transcription through the Satb2 locus, that other players in the transcriptional network compensate for the reduction in Satb2-lncRNA, that Satb2 splicing or transport is affected in a way that does not change total SATB2 levels, or that residual Satb2-lncRNA in the nucleus is sufficient.
Additional players in the transcriptional networks regulating cell fate could include TF families previously unexplored in this context. The enrichment of the NHLH binding motif in putative regulatory elements of DL genes suggests involvement of Nhlh1 and Nhlh2 in DL cell fate. A previous report, however, suggested that both Nhlh1 and Nhlh2 are dispensable for normal cortical development (81). Although this report did not examine expression of layer-specific markers in Nhlh1/Nhlh2 double-knockout mice, we decided to focus our validation experiments on a family of TFs with known roles in neural differentiation but previously unexplored in the context of cortical cell fate specification. Here, we show that MYT family TFs have dynamic expression patterns throughout cortical development: St18 expression in early DL mouse neurons reflected its expression in the developing human motor cortex; Myt1 and Myt1l were initially coexpressed in early DL neurons but, by E15.5, their expression appeared mutually exclusive, with Myt1 restricted to deep cortical layers and Myt1l enriched in superficial layers, suggesting opposing roles in cell fate. We found, however, that the function of MYT1, at least, may be stage-specific. The short isoform of Myt1l that was significantly higher in UL neurons at all three stages lacks several SIN3 interaction domains, which are important for repression, but retains most of its zinc finger domains, suggesting that its role in cortical neuron development may differ from its characterized role in repressing nonneural fates. Indeed, our data suggest that it may promote and maintain Satb2 expression in UL CPNs, where MYT1 function appeared dispensable for Satb2 expression.
Materials and Methods
FACS Purification of Cortical Neuron Subpopulations.
All animal work was carried out in compliance with Stanford University Institutional Animal Care and Use Committee under approved protocol #11499 and institutional and federal guidelines. The day of vaginal plug detection was designated as E0.5. The day of birth was designated as P0. For DL neurons, CD1 female mice were crossed with homozygous golli-τ-EGFP males, resulting in heterozygous offspring, and tissue was collected at E13.5, E16.5, and E18.5. For UL neurons, a plasmid encoding Cdk5r-promoter-GFP (a generous gift of Paola Arlotta, Harvard University, Cambridge, MA) was introduced into the developing neocortex of embryos of timed-pregnant CD1 dams on E15.5 (four to seven embryos per litter were injected) by in utero electroporation, as previously described (82), and tissue was collected at E17.5, P1, and P5.
For each biological replicate (two to five embryos from a single litter), the neocortex was microdissected in cold HBSS. Tissue was dissociated into a single-cell suspension using papain according to the manufacturer’s instructions (Worthington), and resuspended in cortico-spinal motor neuron medium (1 mM pyruvate, 2 mM l-glutamine, 5 μg/mL insulin, 100 U/100 μg/mL pen/strep, 1× Sato, 35 mM glucose, 0.34% BSA, and 800 μM kynurenic acid in 50% DMEM/50% Neurobasal) (83). FACS purifications were performed on a BD FACSAria II, BD FACSJazz, BD Influx, or BD FACS Aria Fusion in the Stanford Shared FACS Facility. Gates for FACS were set using non-GFP age-matched or littermate controls and confirmed using immunohistochemistry for GFP on pre- and postsorted cells. For RNA isolation, 75,000 to 500,000 cells were used per biological replicate. For accessible chromatin, 25,000 to 50,000 cells were used per biological replicate. Sorted cells were stored in cold cortico-spinal motor neuron medium and immediately processed for RNA or accessible chromatin isolation.
RNA-seq and ATAC-seq Library Preparation and Sequencing.
Bulk RNA was isolated from sorted cells using an RNAeasy Micro Kit (Qiagen) according to the manufacturer’s instructions. RNA quality was assessed using a Bioanalyzer 2100 (Agilent), and the RNA integrity number for each sample was above 9. At least 100 ng of total RNA input was used per library. Libraries for mRNA sequencing were prepared using a TruSeq Stranded mRNA Sample Prep Kit (Illumina), and libraries were sequenced on a HiSeq2500 (Illumina), generating an average of 2.35 × 107 100-bp paired-end reads per library (range, 1.65 × 107 to 2.86 × 107). Accessible chromatin was isolated as described in Buenrostro et al. (52), with the following modification: 12 rounds of PCR were used for all samples. Resulting ATAC-seq libraries were sequenced on a HiSeq2500 (Illumina), generating an average of 1.03 × 108 100-bp paired-end reads per library (range 2.51 × 107 to 1.91 × 108).
RNA-seq and ATAC-seq Analysis.
RNA-seq.
Paired-end reads were aligned to the mouse genome (mm9) using kallisto v0.43.1 (84) with default parameters and a transcriptome index built using Ensembl transcripts. Transcript abundance and differential expression were determined using Sleuth (85).
ATAC-seq.
Reads were trimmed using cutadapt v1.9 (86) and pairs were aligned to the mouse genome (mm9) using Bowtie2 v2.2.6 (87) for an average of 1.01 × 108 reads per biological replicate (range, 2.46 × 107 to 1.86 × 108). Duplicate reads were removed using the Picard Tools v1.140 MarkDuplicates function. Peaks were called using MACS v2.1.0 with a P value cutoff of 0.01. Replicated peaks were defined as overlapping regions within overlapping peaks and merged using bedtools. Peaks were masked for blacklist regions, segmental duplications, repeat regions, and exons. ATAC-seq peaks were associated with genes using basal regulatory domains defined by GREAT (v2.0.2) as 5-kb upstream and 1-kb downstream plus up to 1 Mb in both directions to the nearest gene’s basal regulatory domain (53).
Evolutionary Conservation and Motif Discovery.
Evolutionary conservation was determined using PhastCons (88) scores for each base pair within a 50-bp window (SI Appendix, SI Materials and Methods). TF binding site prediction was performed as described previously (32). Briefly, we ran seven different motif discovery tools on each set of correlated peaks, using the set of correlated peaks in the comparison cell subclass as the background set. Tools included AlignAce (89), CisFinder (90), MDscan (91), MEME (92), MoAn (93), MotifSampler (94), and Weeder (95). Near identical motif predictions from at least two different tools were combined and compared with a database of known TF binding motifs using TomTom v5.0.0 (60).
Primary Cell Culture, siRNA KD of lncRNA, and Transfection of Lentiviral Constructs.
For siRNA KD of Satb2-lncRNA, the neocortex was dissected on E15, cells were dissociated using trypsin, cultured in NeuroCult basal medium containing EGF, and proliferation supplement (Stemcell 05702) overnight, and switched to differentiation medium (Stemcell 05704) on the following day. Four hours after switching to differentiation medium, cells were transfected using Lipofectamine RNAiMax reagent (Invitrogen) according to the manufacturer’s instructions. After 4 d in culture, cells were fixed with cold 4% PFA for 5 min then processed for immunohistochemistry. The siRNA sequence for siRNA 1 was ACTCACTGACAAGCCGCAGAGAGAA, for siRNA 2 was GAGATGATTATTAGTTGCGTTGAGT, for scrambled 1 was ACTTCAGAC‐CCGGAGAAACGAA, and for scrambled 2 was GAGTTAGATTAGTTGTTGCGT‐AAGT (Stealth RNAi, ThermoFisher). KD was confirmed using qPCR and primers to Satb2-lncRNA (tgatcaAGACCGGTTCTGGAGAGAAAG) short isoform (ctcgaTGAATCATCATCAAATATATTTATTCACTG) or long isoform (ctcgagTATATATGTTTAATTACACAGTAGTAGAACAT).
For shRNA KD of Myt family TFs, dissections and infections were performed as described above with the following changes: shRNA directed to St18, Myt1, or Myt1l (SI Appendix, Table S10) cloned into a pSico-GFP lentiviral vector (generous gifts of Moritz Mall, Heidelberg University, Heidelberg, Germany) was introduced to cultured cells 4 h after switching to differentiation medium. Two different sequences were used for each target. After 4 to 6 d, cells were fixed with cold 4% PFA for 5 min and processed for immunohistochemistry.
Supplementary Material
Acknowledgments
We thank Dr. Chris Kaznowski and Melissa Lucero for technical help; Dr. Moritz Mall for generously sharing shRNA constructs; Dr. Paola Arlotta for the Cdk5r-GFP construct; Dr. Ben Barres for insights regarding oligodendrocyte progenitor cells; and members of the S.K.M. and G.B. laboratories for helpful comments, especially Dino Leone and Pushkar Joshi. This work was supported by a National Institute of Neurological Disorders and Stroke Epilepsy Training Grant (to W.E.H.), a Stanford Neurosciences Institute Interdisciplinary Scholar Postdoctoral Fellowship (to W.E.H.), National Institutes of Health Grant R01 MH51864 (to S.K.M.), and a Stanford Bio-X Interdisciplinary Seed grant (to G.B. and S.K.M.).
Footnotes
The authors declare no competing interest.
This article contains supporting information online at https://www.pnas.org/lookup/suppl/doi:10.1073/pnas.2008013117/-/DCSupplemental.
Data Availability.
Raw data have been deposited in the National Center for Biotechnology Information GEO (accession no. GSE116147).
References
- 1.Tasic B. et al., Shared and distinct transcriptomic cell types across neocortical areas. Nature 563, 72–78 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Rosenberg A. B. et al., Single-cell profiling of the developing mouse brain and spinal cord with split-pool barcoding. Science 360, 176–182 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Tasic B. et al., Adult mouse cortical cell taxonomy revealed by single cell transcriptomics. Nat. Neurosci. 19, 335–346 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bakken T. E. et al., A comprehensive transcriptional map of primate brain development. Nature 535, 367–375 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Zeng H., Sanes J. R., Neuronal cell-type classification: Challenges, opportunities and the path forward. Nat. Rev. Neurosci. 18, 530–546 (2017). [DOI] [PubMed] [Google Scholar]
- 6.Telley L. et al., Temporal patterning of apical progenitors and their daughter neurons in the developing neocortex. Science 364, eaav2522 (2019). [DOI] [PubMed] [Google Scholar]
- 7.Loo L. et al., Single-cell transcriptomic analysis of mouse neocortical development. Nat. Commun. 10, 134 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.van der Maaten L., Hinton G., Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008). [Google Scholar]
- 9.Becht E. et al., Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2018). [DOI] [PubMed] [Google Scholar]
- 10.Kobak D., Berens P., The art of using t-SNE for single-cell transcriptomics. Nat. Commun. 10, 5416 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Kwan K. Y., Sestan N., Anton E. S., Transcriptional co-regulation of neuronal migration and laminar identity in the neocortex. Development 139, 1535–1546 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Molyneaux B. J., Arlotta P., Menezes J. R. L., Macklis J. D., Neuronal subtype specification in the cerebral cortex. Nat. Rev. Neurosci. 8, 427–437 (2007). [DOI] [PubMed] [Google Scholar]
- 13.Leone D. P., Srinivasan K., Chen B., Alcamo E., McConnell S. K., The determination of projection neuron identity in the developing cerebral cortex. Curr. Opin. Neurobiol. 18, 28–35 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Greig L. C., Woodworth M. B., Galazo M. J., Padmanabhan H., Macklis J. D., Molecular logic of neocortical projection neuron specification, development and diversity. Nat. Rev. Neurosci. 14, 755–769 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Hevner R. F. et al., Tbr1 regulates differentiation of the preplate and layer 6. Neuron 29, 353–366 (2001). [DOI] [PubMed] [Google Scholar]
- 16.Chen J.-G., Rasin M.-R., Kwan K. Y., Sestan N., Zfp312 is required for subcortical axonal projections and dendritic morphology of deep-layer pyramidal neurons of the cerebral cortex. Proc. Natl. Acad. Sci. U.S.A. 102, 17792–17797 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Chen B., Schaevitz L. R., McConnell S. K., Fezl regulates the differentiation and axon targeting of layer 5 subcortical projection neurons in cerebral cortex. Proc. Natl. Acad. Sci. U.S.A. 102, 17184–17189 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Molyneaux B. J., Arlotta P., Hirata T., Hibi M., Macklis J. D., Fezl is required for the birth and specification of corticospinal motor neurons. Neuron 47, 817–831 (2005). [DOI] [PubMed] [Google Scholar]
- 19.Bedogni F. et al., Tbr1 regulates regional and laminar identity of postmitotic neurons in developing neocortex. Proc. Natl. Acad. Sci. U.S.A. 107, 13129–13134 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.McKenna W. L. et al., Tbr1 and Fezf2 regulate alternate corticofugal neuronal identities during neocortical development. J. Neurosci. 31, 549–564 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Han W. et al., TBR1 directly represses Fezf2 to control the laminar origin and development of the corticospinal tract. Proc. Natl. Acad. Sci. U.S.A. 108, 3041–3046 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Lodato S. et al., Gene co-regulation by Fezf2 selects neurotransmitter identity and connectivity of corticospinal neurons. Nat. Neurosci. 17, 1046–1054 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Alcamo E. A. et al., Satb2 regulates callosal projection neuron identity in the developing cerebral cortex. Neuron 57, 364–377 (2008). [DOI] [PubMed] [Google Scholar]
- 24.Britanova O. et al., Satb2 is a postmitotic determinant for upper-layer neuron specification in the neocortex. Neuron 57, 378–392 (2008). [DOI] [PubMed] [Google Scholar]
- 25.Leone D. P. et al., Satb2 regulates the differentiation of both callosal and subcerebral projection neurons in the developing cerebral cortex. Cereb. Cortex 25, 3406–3419 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Molyneaux B. J. et al., DeCoN: Genome-wide analysis of in vivo transcriptional dynamics during pyramidal neuron fate selection in neocortex. Neuron 85, 275–288 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.O’Leary D. D., Koester S. E., Development of projection neuron types, axon pathways, and patterned connections of the mammalian cortex. Neuron 10, 991–1006 (1993). [DOI] [PubMed] [Google Scholar]
- 28.Kadhim H. J., Bhide P. G., Frost D. O., Transient axonal branching in the developing corpus callosum. Cereb. Cortex 3, 551–566 (1993). [DOI] [PubMed] [Google Scholar]
- 29.Srivatsa S., Parthasarathy S., Molnár Z., Tarabykin V., Sip1 downstream Effector ninein controls neocortical axonal growth, ipsilateral branching, and microtubule growth and stability. Neuron 85, 998–1012 (2015). [DOI] [PubMed] [Google Scholar]
- 30.Jacobs E. C. et al., Visualization of corticofugal projections during early cortical development in a tau-GFP-transgenic mouse. Eur. J. Neurosci. 25, 17–30 (2007). [DOI] [PubMed] [Google Scholar]
- 31.Rouaux C., Arlotta P., Direct lineage reprogramming of post-mitotic callosal neurons into corticofugal neurons in vivo. Nat. Cell Biol. 15, 214–221 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Wenger A. M. et al., The enhancer landscape during early neocortical development reveals patterns of dense regulation and co-option. PLoS Genet. 9, e1003728 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Notwell J. H. et al., TBR1 regulates autism risk genes in the developing neocortex. Genome Res. 26, 1013–1022 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Bakken T. E. et al., Single-nucleus and single-cell transcriptomes compared in matched cortical cell types. PLoS One 13, e0209648 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Lê S., Josse J., Husson F., FactoMineR: An R package for multivariate analysis. J. Stat. Softw. 25, 10.18637/jss.v025.i01 (2008). [DOI] [Google Scholar]
- 36.Fame R. M., Dehay C., Kennedy H., Macklis J. D., Subtype-specific genes that characterize subpopulations of callosal projection neurons in mouse identify molecularly homologous populations in macaque cortex. Cereb. Cortex 27, 1817–1830 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Hinks G. L. et al., Expression of LIM protein genes Lmo1, Lmo2, and Lmo3 in adult mouse hippocampus and other forebrain regions: Differential regulation by seizure activity. J. Neurosci. 17, 5549–5559 (1997). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Escamez T. et al., Developmental dynamics of PAFAH1B subunits during mouse brain development. J. Comp. Neurol. 520, 3877–3894 (2012). [DOI] [PubMed] [Google Scholar]
- 39.Telley L. et al., Sequential transcriptional waves direct the differentiation of newborn neurons in the mouse neocortex. Science 351, 1443–1446 (2016). [DOI] [PubMed] [Google Scholar]
- 40.Klingler E. et al., A translaminar genetic logic for the circuit identity of intracortically projecting neurons. Curr. Biol. 29, 332–339.e5 (2019). [DOI] [PubMed] [Google Scholar]
- 41.Jiang X. et al., BHLHB2 controls Bdnf promoter 4 activity and neuronal excitability. J. Neurosci. 28, 1118–1130 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Kamiya D. et al., Intrinsic transition of embryonic stem-cell differentiation into neural progenitors. Nature 470, 503–509 (2011). [DOI] [PubMed] [Google Scholar]
- 43.Miller J. A. et al., Transcriptional landscape of the prenatal human brain. Nature 508, 199–206 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Szklarczyk D. et al., STRING v10: Protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43, D447–D452 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Ashburner M. et al.; The Gene Ontology Consortium , Gene ontology: Tool for the unification of biology. Nat. Genet. 25, 25–29 (2000). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Kapranov P. et al., RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science 316, 1484–1488 (2007). [DOI] [PubMed] [Google Scholar]
- 47.Guil S., Esteller M., Cis-acting noncoding RNAs: Friends and foes. Nat. Struct. Mol. Biol. 19, 1068–1075 (2012). [DOI] [PubMed] [Google Scholar]
- 48.Uesaka M. et al., Bidirectional promoters are the major source of gene activation-associated non-coding RNAs in mammals. BMC Genomics 15, 35 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Engreitz J. M. et al., Local regulation of gene expression by lncRNA promoters, transcription and splicing. Nature 539, 452–455 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Sauvageau M. et al., Multiple knockout mouse models reveal lincRNAs are required for life and brain development. eLife 2, e01749 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Goff L. A. et al., Spatiotemporal expression and transcriptional perturbations by long noncoding RNAs in the mouse brain. Proc. Natl. Acad. Sci. U.S.A. 112, 6855–6862 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Buenrostro J. D., Giresi P. G., Zaba L. C., Chang H. Y., Greenleaf W. J., Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.McLean C. Y. et al., GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.ENCODE Project Consortium , An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Iijima T. et al., SAM68 regulates neuronal activity-dependent alternative splicing of neurexin-1. Cell 147, 1601–1614 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Perera B. P. U., Kim J., Alternative promoters of Peg3 with maternal specificity. Sci. Rep. 6, 24438 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Pfurr S. et al., The E2A splice variant E47 regulates the differentiation of projection neurons via p57(KIP2) during cortical development. Development 144, 3917–3931 (2017). [DOI] [PubMed] [Google Scholar]
- 58.Bansod S., Kageyama R., Ohtsuka T., Hes5 regulates the transition timing of neurogenesis and gliogenesis in mammalian neocortical development. Development 144, 3156–3167 (2017). [DOI] [PubMed] [Google Scholar]
- 59.Baizabal J.-M. et al., The epigenetic state of PRDM16-regulated enhancers in radial glia controls cortical neuron position. Neuron 98, 945–962.e8 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Gupta S., Stamatoyannopoulos J. A., Bailey T. L., Noble W. S., Quantifying similarity between motifs. Genome Biol. 8, R24 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Vied C. M. et al., A multi-resource data integration approach: Identification of candidate genes regulating cell proliferation during neocortical development. Front. Neurosci. 8, 257 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Zhao B., Lei Q.-Y., Guan K.-L., The Hippo-YAP pathway: New connections between regulation of organ size and cancer. Curr. Opin. Cell Biol. 20, 638–646 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Massimino L. et al., TBR2 antagonizes retinoic acid dependent neuronal differentiation by repressing Zfp423 during corticogenesis. Dev. Biol. 434, 231–248 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Yue T. et al., A critical role for dorsal progenitors in cortical myelination. J. Neurosci. 26, 1275–1280 (2006). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Eckler M. J. et al., Cux2-positive radial glial cells generate diverse subtypes of neocortical projection neurons and macroglia. Neuron 86, 1100–1108 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Franco S. J. et al., Fate-restricted neural progenitors in the mammalian cerebral cortex. Science 337, 746–749 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Kocabas A., Duarte T., Kumar S., Hynes M. A., Widespread differential expression of coding region and 3′ UTR sequences in neurons and other tissues. Neuron 88, 1149–1156 (2015). [DOI] [PubMed] [Google Scholar]
- 68.Vasconcelos F. F. et al., MyT1 counteracts the neural progenitor program to promote vertebrate neurogenesis. Cell Rep. 17, 469–483 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Mall M. et al., Myt1l safeguards neuronal identity by actively repressing many non-neuronal fates. Nature 544, 245–249 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.McConnell S. K., Kaznowski C. E., Cell cycle dependence of laminar determination in developing neocortex. Science 254, 282–285 (1991). [DOI] [PubMed] [Google Scholar]
- 71.Frantz G. D., McConnell S. K., Restriction of late cerebral cortical progenitors to an upper-layer fate. Neuron 17, 55–61 (1996). [DOI] [PubMed] [Google Scholar]
- 72.Desai A. R., McConnell S. K., Progressive restriction in fate potential by neural progenitors during cerebral cortical development. Development 127, 2863–2872 (2000). [DOI] [PubMed] [Google Scholar]
- 73.Gray L. T. et al., Layer-specific chromatin accessibility landscapes reveal regulatory networks in adult mouse visual cortex. eLife 6, e21883 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Aprea J. et al., Transcriptome sequencing during mouse brain development identifies long non-coding RNAs functionally involved in neurogenic commitment. EMBO J. 32, 3145–3160 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Tripathi V. et al., Long noncoding RNA MALAT1 controls cell cycle progression by regulating the expression of oncogenic transcription factor B-MYB. PLoS Genet. 9, e1003368 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Nagano T. et al., The Air noncoding RNA epigenetically silences transcription by targeting G9a to chromatin. Science 322, 1717–1720 (2008). [DOI] [PubMed] [Google Scholar]
- 77.Yin Y. et al., Opposing roles for the lncRNA haunt and its genomic locus in regulating HOXA gene activation during embryonic stem cell differentiation. Cell Stem Cell 16, 504–516 (2015). [DOI] [PubMed] [Google Scholar]
- 78.Bassett A. R. et al., Considerations when investigating lncRNA function in vivo. eLife 3, e03058 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Paralkar V. R. et al., Unlinking an lncRNA from its associated cis element. Mol. Cell 62, 104–110 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Wei W., Pelechano V., Järvelin A. I., Steinmetz L. M., Functional consequences of bidirectional promoters. Trends Genet. 27, 267–276 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Krüger M., Braun T., The neuronal basic helix-loop-helix transcription factor NSCL-1 is dispensable for normal neuronal development. Mol. Cell. Biol. 22, 792–800 (2002). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Ohtsuka T. et al., Visualization of embryonic neural stem cells using Hes promoters in transgenic mice. Mol. Cell. Neurosci. 31, 109–122 (2006). [DOI] [PubMed] [Google Scholar]
- 83.Dugas J. C. et al., A novel purification method for CNS projection neurons leads to the identification of brain vascular cells as a source of trophic support for corticospinal motor neurons. J. Neurosci. 28, 8294–8305 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Bray N. L., Pimentel H., Melsted P., Pachter L., Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016). [DOI] [PubMed] [Google Scholar]
- 85.Pimentel H., Bray N. L., Puente S., Melsted P., Pachter L., Differential analysis of RNA-seq incorporating quantification uncertainty. Nat. Methods 14, 687–690 (2017). [DOI] [PubMed] [Google Scholar]
- 86.Martin M., Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 17, 10–12 (2011). [Google Scholar]
- 87.Langmead B., Salzberg S. L., Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Siepel A. et al., Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res. 15, 1034–1050 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Roth F. P., Hughes J. D., Estep P. W., Church G. M., Finding DNA regulatory motifs within unaligned noncoding sequences clustered by whole-genome mRNA quantitation. Nat. Biotechnol. 16, 939–945 (1998). [DOI] [PubMed] [Google Scholar]
- 90.Sharov A. A., Ko M. S. H., Exhaustive search for over-represented DNA sequence motifs with CisFinder. DNA Res. 16, 261–273 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Liu X. S., Brutlag D. L., Liu J. S., An algorithm for finding protein-DNA binding sites with applications to chromatin-immunoprecipitation microarray experiments. Nat. Biotechnol. 20, 835–839 (2002). [DOI] [PubMed] [Google Scholar]
- 92.Bailey T. L., Elkan C., Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 2, 28–36 (1994). [PubMed] [Google Scholar]
- 93.Valen E., Sandelin A., Winther O., Krogh A., Discovery of regulatory elements is improved by a discriminatory approach. PLOS Comput. Biol. 5, e1000562 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Thijs G. et al., A Gibbs sampling method to detect overrepresented motifs in the upstream regions of coexpressed genes. J. Comput. Biol. 9, 447–464 (2002). [DOI] [PubMed] [Google Scholar]
- 95.Pavesi G., Mauri G., Pesole G., An algorithm for finding signals of unknown length in DNA sequences. Bioinformatics 17, S207–S214 (2001). [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Raw data have been deposited in the National Center for Biotechnology Information GEO (accession no. GSE116147).