Abstract
We describe the first known relatives of rubella virus (Matonaviridae: Rubivirus)1 in Africa and Europe. Ruhugu virus, the closest relative of rubella virus, was found in apparently healthy cyclops leaf-nosed bats (Hipposideros cyclops) in Uganda. Rustrela virus, outgroup to the rubella/ruhugu clade of viruses, was found in acutely encephalitic placental and marsupial animals at a zoo in Germany and in wild yellow-necked field mice (Apodemus flavicollis) at and near the zoo. Ruhugu and rustrela viruses share an identical genomic architecture with rubella virus2,3. Amino acid sequences of rubella, ruhugu, and rustrela viruses are moderately to highly conserved within 4 putative B-cell epitopes in the fusion (EI) protein and, in the case of rubella and ruhugu viruses, within two putative T-cell epitopes in the capsid protein4–6. Modeling of E1 homotrimers in the post-fusion state predicts similar host-cell membrane fusion capacity for ruhugu and rubella viruses5. Together, these findings suggest show that some members of the Matonaviridae can cross wide host species barriers and that rubella virus likely had a zoonotic origin. Our findings raise concerns about future zoonotic transmission of rubella-like viruses but open doors for heretofore impossible comparative studies and novel animal models of rubella and congenital rubella syndrome.
Rubella, first described in 18147, is an acute, highly contagious human infectious disease typically characterized by rash, low-grade fever, adenopathy, and conjunctivitis1. Research from the 1940s to 1960s revealed that rubella (also called “German measles”) contracted during the first trimester of pregnancy was directly associated with severe congenital birth defects, miscarriage, and stillbirth8,9. Rubella virus (RuV), currently the sole recognized member of the riboviriad family Matonaviridae (genus Rubivirus), is the etiologic agent of rubella10,11 and causes fetal pathology after transplacental transmission12. Extensive rubella epidemics have occurred worldwide due to the high airborne transmissibility of RuV (R0 = 3.5–7.8)13. Safe, efficacious, live-attenuated RuV vaccines, including the measles/mumps/rubella (MMR) vaccine, are now deployed worldwide and have successfully decreased global rubella incidence. However, ≈100,000 cases of congenital rubella syndrome still occur annually1, and RuV can persist in immunologically privileged anatomic sites (e.g., the eye) for years14. Furthermore, RuV infection of adults is generally underreported, with 30–50% of adult cases being subclinical15. High-priority areas for rubella vaccination include the Western Pacific, Eastern Mediterranean, and African regions, where RuV circulates widely and primarily infects young children16. RuV elimination is considered rapidly achievable because of the effectiveness of available vaccines and the lack of known animal reservoirs17,18.
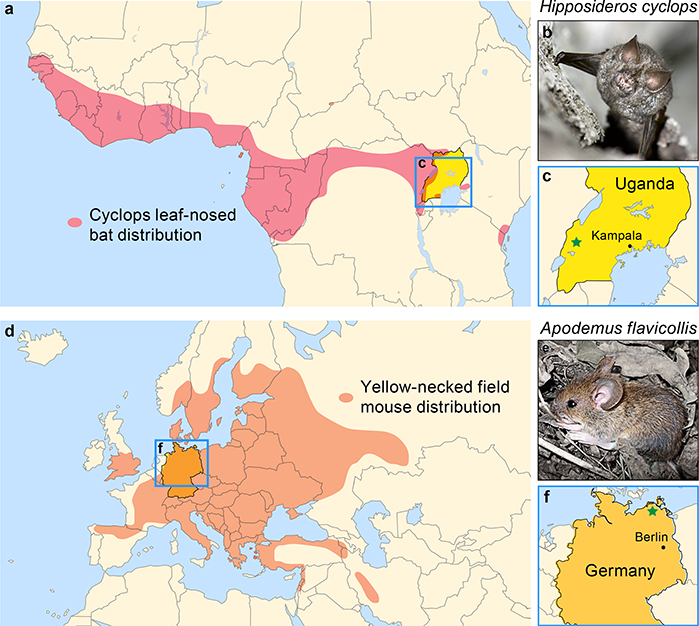
Here we report the discovery of ruhugu virus (RuhV) and rustrela virus (RusV), the first known relatives of RuV. RuhV was found in 10 of 20 oral swabs from apparently healthy cyclops leaf-nosed bats (Hipposideridae: Hipposideros cyclops Temminck, 1853) in Kibale National Park, Uganda (Fig. 1). RusV was found in brain tissues of three acutely ill animals at a zoo in Germany, all of which succumbed to severe, acute neurologic disease (Extended Data Table 2): a donkey (Equus asinus Linnaeus, 1758), a capybara (Hydrochoeris hydrochaeris [Linnaeus, 1766]), and a Bennett’s tree-kangaroo (Dendrolagus bennettianus [De Vis, 1886]). RusV was subsequently detected in brain tissues of 8 of 16 yellow-necked field mice (Muridae: Apodemus flavicollis [Melchior, 1834]) on the zoo grounds and within 10 km of the zoo (Fig. 1; Extended Data Table 1).
Fig. 1 |.
Geographic locations of viruses and their hosts. a) Summary map of estimated cyclops leaf-nosed bat distribution in Africa (red) and Uganda (blue box). b) Cyclops leaf-nosed bat in Kibale National Park, Uganda (photo credit: Caley Johnson). c) Location of bat sample collection and discovery of ruhugu virus (Kibale National Park, Uganda, green star). d) Summary map of estimated yellow-necked field mouse distribution in Eurasia (orange) and Germany (blue box). e) Yellow-necked field mouse in northeastern Germany (photo credit: Ulrike M. Rosenfeld). f) Location of zoo animals and discovery of rustrela virus in Germany (southern Baltic Sea region, green star).
In the case of RuhV in Uganda, all bats were captured and sampled from five tree roosts (hollow cavities in trees) each containing between 1 and 8 bats. Using molecular and metagenomic methods (Supplementary Methods), RuhV RNA was detected in 5/9 (55.6%) males and 5/11 (45.5%) females in 4 of 5 (80.0%) of roosts (50%; 95% confidence interval 29.9–70.1%). This high prevalence and frequency of positive roosts suggest that apparently healthy cyclops leaf-nosed bats are reservoir hosts, rather than incidental hosts, of RuhV. Cyclops leaf-nosed bats are insectivorous microbats primarily found in lowland rainforests from Senegal to Tanzania but also in coastal, montane, and swamp forests and disturbed and agricultural landscapes19–21 (Fig. 1a), and they host Plasmodium cyclopsi, an apicomplexan “bat malaria” parasite22,23. Whether RuhV can infect animals other than cyclops leaf-nosed bats remains unknown.
In the case of RusV in Germany, the donkey, capybara, and Bennett’s tree-kangaroo were submitted for post-mortem evaluation and testing (Supplementary Methods), which led to the identification of the virus (see below). Subsequent testing of rodents housed at the zoo and wild rodents on the zoo grounds and at two other locations within 10 km of the zoo revealed 8/16 (50%; 95% confidence interval 6.7–39.1%) yellow-necked field mice to be positive for RusV RNA in brain tissue. Surprisingly, the mice had no histologic evidence of encephalitis (7/8 mice investigated) and had only low concentrations of RusV RNA in peripheral organs (Extended Data Table 3). RusV RNA was not detected in any other small mammals collected simultaneously (n=38; Extended Data Table 1). Yellow-necked field mice are omnivorous rodents native to parts of Europe and Asia, occupying mature forests to agricultural and peridomestic habitats24 (Fig. 1d). They host tick-borne encephalitis virus (Flaviviridae: Flavivirus)25, Dobrava-Belgrade virus, genotype Dobrava (Hantaviridae: Orthohantavirus)26–28, Akhmeta virus (Poxviridae: Orthopoxvirus)29, and hepatitis E virus (Hepeviridae: Orthohepevirus)30. Routes of transmission of RuhV and RusV within reservoir hosts and to spillover hosts (in the case of RusV) remain unknown, but positive oral swabs and feces (Extended Data Table 3) suggest that contact with oral secretions and excreta could play a role.
Using molecular methods and in situ hybridization (Supplementary Methods), we confirmed the presence of RusV in brain tissues of all German zoo animals and in the liver of the donkey (Extended Data Table 2; Extended Data Figure 1). RusV RNA was detected within neuronal cell bodies and their processes by in situ RNA hybridization on brain tissue sections of the donkey (Extended Data Figure 1a), Bennett’s tree-kangaroo (Extended Data Figure 1b), and capybara (Extended Data Figure 1c). Histopathology revealed a non-suppurative meningoencephalitis in all 3 animals, characterized by perivascular cuffing (Fig. 2a–c), meningeal infiltrates (Fig. 2d), and glial nodules (Fig. 2e). Neuronal necrosis and degeneration with satellitosis were detected in the brain stem of the donkey (Fig. 2f). Immune cells in brain tissue consisted mainly of CD-3-positive T-lymphocytes, Iba-1-positive microglial cells and macrophages and CD79a-immunoreactive B-lymphocytes (Fig. 2g–l). In general, apoptosis was not a significant feature, with few active caspase-3-labelled cells distributed perivascularly and scattered within the gray and white matter (Fig. 2m–n). Multifocal perivascular red blood cells in brain samples from the donkey and Bennett’s tree-kangaroo were positive for iron in the Prussian Blue reaction, indicating intravital hemorrhages (Fig. 2o). In yellow-necked field mice, detection of viral RNA in samples collected between 2009 and 2020 and absence of inflammation (Extended Data Figure 1d–e) suggest this broadly distributed rodent to be the RusV reservoir.
Fig. 2 |.
Histopathology and immune reaction of rustrela virus in the brain of a capybara, Bennett’s tree-kangaroo and donkey. a–c) Non-suppurative meningoencephalitis with mononuclear, perivascular cuffing, brain, capybara (a), Bennett’s tree-kangaroo (b), and donkey (c). d) Mononuclear meningeal infiltrates, brain, donkey. e) Glial nodules, brain, donkey. f) Neuronal necrosis (arrow) and degeneration with satellitosis (arrow head), brain, donkey. HE stain; scale bar 20 μm (a–c, e–f), 50 μm (d). g–l) Immune reaction by immunohistochemistry, perivascular, brain, Bennett’s tree kangaroo; and in glial nodules, brain, donkey (j–l), numerous CD-3 labelled T- lymphocytes (g, j), Iba-1 positive microglial cells and macrophages (h, k), CD79a immunoreactive B- lymphocytes (i, l). Immunohistochemistry, AEC chromogen, Mayer’s haematoxylin counter stain, scale bar 20 μm. m–n) Apoptosis, few active Caspase-3 labelled cells (arrows), perivascular and scattered throughout the neuropil, brain, Bennett’s tree-kangaroo (m), brain, capybara (n). Immunohistochemistry, AEC chromogen, Mayer’s haematoxylin counter stain; scale bar 20 μm. o) Hemorrhage, Prussian Blue reaction demonstrates multiple iron deposits (arrows) within mononuclear cells found perivascularly, admixed with red blood cell accumulations, indicating intra-vital haemorrhage; scale bar 20μm. Immunohistochemistry was performed on at least 4 slides per animal, yielding comparable results in all cases. In each run, positive control slides and negative control primary antibodies were included. Evaluation and interpretation were performed by a board certified pathologist (DiplECVP) with more than 13 years experience
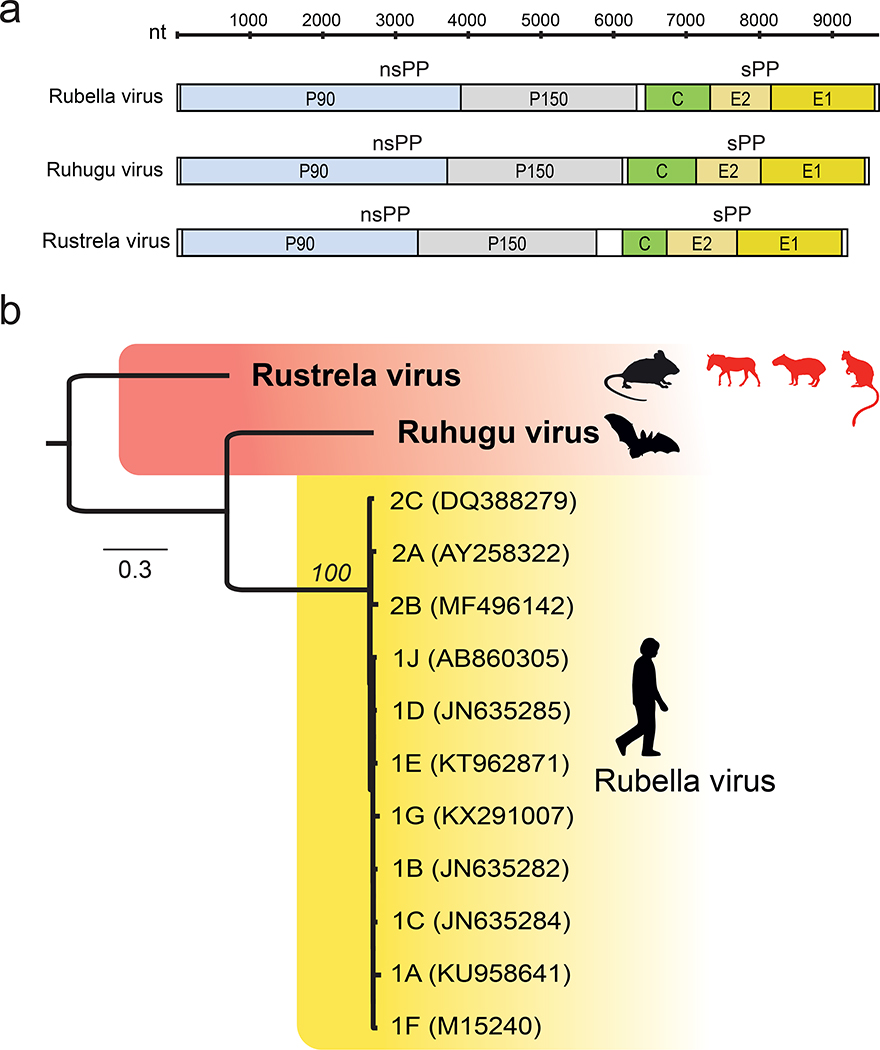
The genome organizations of RuV, RuhV, and RusV are identical, consisting of two large open reading frames (ORFs), two untranslated regions (UTRs) at 5′ and 3′ termini, and an intergenic region (IR) between the two ORFs (Fig. 3a). Across the nonstructural and structural coding regions, RuhV is more similar to RuV than is RusV (Extended Data Table 4). Genetic similarity varies within coding regions and is generally highest in a hyper-conserved region within the Y domain of P1502,31,32 (Extended Data Fig. 2). RusV contains a notably long IR (366 nt, versus 46 nt and 75 nt in RuV and RuhV, respectively) and a correspondingly short C protein (205 aa, versus 300 aa and 317 aa in RuV and RuhV, respectively; Extended Data Table 4). In addition, RuV and RuhV share a Gly-Gly-Gly amino acid sequence at the p150/p90 cleavage site, whereas RusV has a Gly-Gly-Ala amino acid sequence at this same site, which may impair cleavage in the case of RusV3.
Fig. 3 |.
Evolutionary relationships among viruses. a) Comparative genome architecture of RuV, RuhV, and RusV, showing five open reading frames (colored), two untranslated regions at the 5′ and 3′ termini (white), and an intergenic region (white) between the ORFs encoding the non-structural (nsPP) and structural (sPP) polyproteins. b) Maximum likelihood phylogenetic tree of rustrela virus, ruhugu virus, and rubella virus genotypes 1A–1J and 2A–2C. Black silhouettes represent natural hosts of each virus, and red silhouettes represent spillover hosts in the case of RusV. Numbers beside nodes indicate bootstrap values (%; only values for major branches are shown); the scale bar indicates amino acid substitutions per site.
RuhV (named for Ruteete Subcounty, Uganda, and the Tooro word for insectivorous bat, obuhuguhugu) is an outgroup to all known RuV genotypes (Fig. 3b). RusV (named for its rubella virus-like genome and the Strela Sound of the Baltic Sea in Germany) is a close outgroup to the RuV/RuhV clade of viruses (Fig. 3b). This topology is consistent with the higher similarity of RuhV to RuV in each of the five mature polypeptides of the protein-coding viral genome (Extended Data Table 4, Extended Data Fig. 2). Nucleotide sequences of RusV were 97.4 to 100 % similar within p90 and E1 coding regions sequenced in the donkey, capybara, Bennett’s tree-kangaroo, and yellow-necked field mice in Germany (Extended Data Figure 3).
The RuV E1 protein, a receptor-binding, class II fusion protein5, contains an immune-reactive region (amino acid residue positions 202–283) with immunodominant T-cell epitopes6 and four linear, neutralizing B-cell epitopes (NT1–4)4 (Fig. 4a). The modeled tertiary and quaternary structures of trimeric RuhV and RusV E1 are homologous to RuV E133, and homology-based modeling of RuhV E1 quaternary structure predicts with high confidence that RuhV and RusV E1 proteins form homotrimers in the post-fusion state5 (Fig. 4b and c). One neutralizing epitope maps to amino acid positions 223–239 of the E1 protein at disulfide bond 8 (NT1)34. The mechanism of neutralization appears to involve blocking of E1 trimerization, which is necessary for virion fusion with the host-cell plasma membrane5. Surprisingly, only one amino acid residue (R237Q, near the C-terminus) differs between the RuV and RuhV NT1 epitope (Fig. 4a), despite higher amino acid-level divergence across E1 (Extended Data Figure 3). By contrast, RusV differs from RuV at 5 amino acid residues within the same region (Fig. 4a). T-cell epitopes are not well conserved in the capsid (Extended Data Table 5); however; the exposed RuhV and RusV NT3 and NT4 putative linear epitopes within E1 are moderately conserved in comparison to RuV (Fig. 4; Extended Data Table 5), suggesting that they also should be evaluated for cross-neutralization by anti-RuV antibodies.
Fig. 4 |.
Comparisons of the rubella (RuV), ruhugu (RuhV), and rustrela virus (RusV) E1 envelope glycoproteins. a) Amino acid alignment and sequence logo of an immunoreactive region of E1 for RuhV, RusV, and 13 RuV genotypes (GenBank accession numbers in parentheses). Lines indicate locatons of putative linear neutralizing B-cell epitopes NT1-NT4. b) Homology-based model of RuhV E1 homotrimer structure in the post-fusion state showing receptor-binding site view (left) and profile view (right). Global model quality estimates (QMEAN) indicate a good model fit relative to the crystal structure of the RuV E1 in the post-fusion form (Protein Data Bank biological assembly 4adg.1). c) Homology-based model of the RusV E1 homotrimer structure in the post-fusion state, as described above for RuhV. Key differences are seen in the modeled neutralizing epitopes NT3 and NT4 and in Fusion Loops 1 and 2 (FL1 and FL2). RuhV FL1 and FL2 residues are highly similar to those of RuV, whereas RusV FL2 residues differ from those of RuV FL2 to a greater extent. The color scale indicates normalized QMEAN local score.
The RuhV E1 fusion loops (FL1: residues 87–92; FL2: residues 130–136) are predicted to support the unusual metal ion complex necessary for E1-mediated RuV membrane fusion due to the presence of amino acids N87 and D135 (homologous to RuV N88 and N136, respectively; Fig. 4b)5. By contrast, FL2 of RusV is predicted to be less similar to RuV due to two amino acid residue replacements, P134A and T135A, the latter being a change from a polar to a non-polar residue (Fig. 4c). Across the RuV, RuhV, and RusV genomes, regions of marked conservation and stabilizing selection are evident immediately upstream of the putative methyltransferase domain of P150, in the RdRp domain of P90, and proximal to the aforementioned NT1 epitope of E1 (Extended Data Fig. 2).
The similarity or near identity of certain RuV, RuhV, and RusV B-cell epitopes (Extended Data Table 5) suggests that existing serologic assays for anti-rubella antibodies might detect RuhV, RusV, and other as-yet undiscovered rubella-like viruses. Future studies evaluating the performance of existing serologic tests for RuV infection in animals would be useful, as would the development of novel assays that can detect and differentiate among rubella-like viral infections in animals and humans. Implicating RuhV or RusV as zoonotic agents is currently speculative, but bats and rodents possess biological attributes that predispose them to hosting many zoonotic viruses35–37, so this scenario ought not to be dismissed. The ability of RusV to infect both placental and marsupial mammals and to cause clinical disease resembling severe encephalitic forms of rubella in humans38,39 reinforces such a precautionary stance.
The World Health Organization’s (WHO’s) Global Measles and Rubella Strategic Plan aims to control or eliminate rubella and congenital rubella syndrome in 5/6 WHO regions by the end of 202040. Our discoveries of relatives of RuV infecting asymptomatic bats and rodents suggests that rubella may have arisen as a zoonosis. Furthermore, the ability of RusV to infect mammals across wide taxonomic distances and to cause severe encephalitis in spillover hosts raises concern about the potential for zoonotic transmission of RuhV, RusV, or other as-yet undiscovered rubella-like viruses. Despite these concerns, our findings clearly augur well for comparative studies of RuV that were heretofore not possible, including the potential development of novel animal models for rubella and congenital rubella syndrome.
Extended Data
Extended Data Figure 1 |.
RNA in situ hybridization of rustrela virus. a-e) Detection of rustrela virus RNA in brain tissues of a donkey (a), Bennett’s tree-kangaroo (b), capybara (c) and yellow-necked field mice (d, e). Chromogenic labelling (fast red) with probes to rustrela virus NSP-coding region are visible in neuronal cell bodies (arrow) but not in adjacent glial cells (arrow head). Mayer’s hematoxylin counter stain. Scale bar = 50 μm f). Negative control probe to bacterial gene dapB encoding dihydrodipicolinate reductase. Lack of chromogenic labelling (fast red). Mayer’s hematoxylin counter stain. Scale bar = 100 μm. RNAscope results were evaluated on at least 3 slides per animal, yielding comparable results in all cases. In situ hybridization was performed according to the manufacturer’s instructions, including a positive control probe, peptidylprolyl isomerase B (cyclophilin B, ppib), and a negative control probe, dihydrodipicolinate reductase (DapB). Evaluation and interpretation were performed by a board certified pathologist (DiplECVP) with more than 13 years experience.
Extended Data Figure 2 |. Average substitution rates at non-synonymous (dN; dashed lines) and synonymous (dS; grey lines) sites, and the ratio of dN/dS (solid lines), for aligned, concatenated amino acid sequences comparing RuV and RuhV (a), RuV and RusV (b), and RuhV and RusV (c) using sliding window analysis (100 residue window length, 10 residue steps).
Protein domains are labeled on the X axes: MT=methyltransferase; Y, Q, and X=domains of unknown function; Pro=protease; Hel=helicase; RdRp=RNA-directed RNA polymerase; NT1=neutralizing epitope 1.
Extended Data Figure 3 |. Phylogenetic analyses of the coding sequences of envelope glycoprotein E1 (a) and helicase and RNA-directed RNA polymerase p90 (b) of RusV and RuhV and RusV, including all sequences obtained in this study (GenBank accession numbers in parentheses).
Numbers above branches represent bootstrap values; scale bars indicate amino acid substitutions per site.
Extended Data Table 1 |.
Rustrela virus in small mammals from northeastern Germany assessed by RT-qPCR.
| Capture location |
||||
|---|---|---|---|---|
| Common name | Species | Zoo | Within 10 km of zoo | Total |
| Yellow-necked field mouse | Apodemus flavicollis [Melchior, 1834] | 6/11 (54.5 %) | 2/5 (40 %) | 8/16 (50 %) |
| Striped field mouse | Apodemus agrarius [Pallas, 1771] | 0/4 | 0/2 | 0/6 |
| Bank vole | Myodes glareolus [Schreber, 1780] | 0/3 | n/a | 0/3 |
| Brown rat | Rattus norvegicus [Berkenhout, 1769] | 0/13* | n/a | 0/13 |
| House mouse | Mus musculus Linnaeus, 1758 | 0/3* | 0/13 | 0/16 |
- = no material available.
Two brown rats and all three house mice were animals housed at the zoo.
Extended Data Table 2 |.
Rustrela virus distribution in zoo animal tissues assessed by RT-qPCR.
| Cq value | ||||
|---|---|---|---|---|
| Source | Donkey | Capybara | Bennett’s tree-kangaroo | |
| Central nervous system | Cerebrum (I)a | 22.9 | - | 30.2 |
| Cerebrum (II)b | 29.2 | 26.0 | - | |
| Cerebrum (III)b | 29.5 | 26.6 | - | |
| Cerebrum (IV)b | - | 30.9 | - | |
| Brain stemb | 30.5 | 29.1 | - | |
| Cerebellumb | 30.6 | - | - | |
| Medulla oblongatab | 33.9 | - | - | |
| Medullab | - | 34.6 | - | |
| Spinal cordb | - | 30.7 | - | |
| Peripheral organs | Liver (I)a | - | - | - |
| Liver (II)b | 35.9 | - | - | |
| Kidneyb | neg | neg | - | |
| Spleenb | neg | neg | - | |
| Small intestineb | - | neg | - | |
| Organ pool (I)a | neg | - | 35.5 | |
| Organ pool (II)a | - | - | - | |
fresh, unfixed tissues
formalin-fixed paraffin-embedded tissues
- = no material available; neg = negative. Cells are shaded in proportion to relative viral concentration (Cq value).
Extended Data Table 3 |.
Rustrela virus distribution in tissues of positive Apodemus flavicollis assessed by RT-qPCR.
| Cq value | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| KS19/923 | KS20/926 | KS19/928 | KS20/1296 | KS20/1340 | KS20/1341 | KS20/1342 | KS20/1343 | Mu09/1341 | |
| Cerebrum | 28.1 | neg | 22.9 | 24.1 | 26.3 | 21.1 | 20.8 | 20.4 | 25.9 |
| Heart | neg | neg | neg | neg | 31.9 | neg | neg | neg | - |
| Lungs | neg | neg | neg | neg | 36.7 | 35.0 | neg | neg | - |
| Liver | neg | neg | neg | neg | neg | neg | neg | neg | - |
| Kidneys | neg | neg | neg | neg | neg | neg | neg | neg | - |
| Spleen | neg | neg | neg | neg | neg | neg | neg | neg | - |
| Intestine/feces | neg | 36.7 | neg | neg | neg | neg | neg | neg | - |
| Thoracic lavage | neg | neg | neg | neg | 37.5 | neg | neg | neg | - |
| Oral swab | - | - | - | - | 36.2 | 37.5 | neg | neg | - |
- = no material available; neg = negative. Cells are shaded in proportion to relative viral concentration (Cq value).
Extended Data Table 4 |.
Genomic features of ruhugu virus (RuhV; GenBank MN547623) and rustrela virus (RusV; GenBank MN552442).
| Nucleotide position (5′→3′) |
Amino acid residues |
Amino acid sequence identity (%) |
GC content (%) |
|||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Genome featurec | RuhV | RusV | RuhV | RusV | RuhVa | RusVa | RuhV–RusV | RuhV | RusV | RuVb |
| Complete genome | 1–9621 | 1–9322 | 6296 | 5876 | 56.4 | 43.0 | 43.3 | 63.5 | 70.6 | 69.6 |
| Non-structural polyprotein | 44–6190 | 68–5833 | 2049 | 1921 | 59.0 | 45.9 | 47.5 | 62.2 | 70.2 | 70.0 |
| p150 protease | 44–3754 | 68–3391 | 1237 | 1108 | 48.6 | 34.5 | 35.7 | 63.1 | 72.0 | 71.4 |
| p90 replication complex | 3755–6190 | 3392–5830 | 812 | 813 | 75.7 | 65.5 | 66.6 | 60.9 | 67.7 | 67.8 |
| Structural polyprotein | 6266–9562 | 6193–9246 | 1099 | 1017 | 51.4 | 41.1 | 39.5 | 66.1 | 71.4 | 69.4 |
| Capsid protein | 6266–7216 | 6193–6807 | 317 | 205 | 51.7 | 46.6 | 43.0 | 66.6 | 74.5 | 73.1 |
| E2 envelope protein | 7217–8101 | 6808–7785 | 295 | 326 | 43.6 | 31.4 | 23.9 | 67.9 | 72.7 | 71.0 |
| E1 envelope protein | 8102–9562 | 7786–9243 | 487 | 486 | 56.3 | 51.0 | 50.6 | 64.8 | 69.3 | 66.3 |
Extended Data Table 5 |.
Conservation of B- and T-cell epitopes in the E1 fusion protein of the rubella wild-type virus 1B, ruhugu virus, and rustrela virus.
| Epitope | Rubella virus (JN635282) | Ruhugu virus (MN547623) | Rustrela virus (MN552442) | |
|---|---|---|---|---|
| Linear, neutralizing B-cell epitopes | NT1:E11221–239 | LGSPNCHGPDWASPVCQRHS | VGLPNCHGPDWASPVCQQHS | VPAPDCFGPAWASPVCARHM |
| NT2: E1245–251 | LVGATPE | LTGVPPE | LTGATPG | |
| NT3 :E1260–266 | ADDPLLR | ADDPRLT | ADDLGWH | |
| NT4 :E1274–285 | VWVTPVIGSQAR | VWAVAVKGTQPK | VWYQPVIGRQPR | |
| CD8+ T-cell epitopes | C9–22 | MEDLQKALEAQSRA | LADLQRLLEKQSAE | Deleted |
| C11–29 | DLQKALEAQSRALRAELAA | DLQRLLEKQSAELRAEMAR | Deleted | |
| C264–272 | RIETRSARH | KQDVKSDKV | RKEOLGATSGAA | |
Amino acid differences are in bold, insertions are underlined, and GenBank accession numbers are in parentheses.
Extended Data Table 6 |.
Immunohistochemical markers and applications.
| Marker | Antibody | Antigen Retrieval | Secondary reagents |
|---|---|---|---|
| Active caspase 3 | Anti-Active Caspase 3 (Promega, Walldorf, Germany), 1:200, overnight | n/a | ABC Kit Vectastain Elite PK 6100, 30 min (Dako) |
| CD79a | Mouse anti-CD79A (clone HM57) monoclonal, (LifeSpan BioSciences, Seattle, WA, USA), 1:50, overnight | HIER, 10 mM Tris/1mM EDTA buffer pH 9.0, 20 min | Dako EnVision+ System-HRP Labelled Polymer Anti-mouse, 30 min |
| CD3 | Rabbit anti-CD3 polyclonal (Dako), 1:100, overnight | HIER, 10 mM Tris/1mM EDTA buffer pH 9.0, 20 min | Dako EnVision+ System- HRP Labelled Polymer Anti-rabbit, 30 min |
| Iba-1 | Iba1 (Wako), 1:800, overnight | HIER, Citrate buffer pH 6.0, for 20 min | Dako EnVision+ System- HRP Labelled Polymer Anti-rabbit, 30 min |
HIER: Heat-induced epitope retrieval; HRP: horse-radish peroxidase. n/a = not applicable.
Supplementary Material
Acknowledgements
We thank David Hyeroba (Makerere University, Uganda, in memoriam), Katusabe Swaibu (Makerere University Biological Field Station, Uganda), and Jessica Carag (University of Wisconsin-Madison, Madison, WI, USA) for assistance in the field and Christoph Langner and the zoo in Germany for assistance with sampling and for implementing timely response strategies. We thank Laura Bollinger and Jiro Wada (NIH/NIAID Integrated Research Facility at Fort Detrick, Frederick, MD, USA) and Dennis Rubbenstroth (Friedrich-Loeffler-Institut, Greifswald-Insel Riems, Germany) for their help improving the manuscript and figures. We thank Gregory K. Rice (Leidos and Biological Defense Research Directorate, Naval Medical Research Center – Frederick, MD, USA) for advice and assistance with bioinformatics scripts, Patrick Zitzow and Jenny Lorke, Silvia Schuparis and Gabriele Czerwinski (Friedrich-Loeffler-Institut, Greifswald—Insel Riems, Germany) for invaluable technical assistance, and Christine Jelinek, Dörte Kaufmann, Julia Pöhlig and Christin Trapp for help with rodent trapping and dissection.
This work was supported in part through the US National Institute of Allergy and Infectious Diseases (NIAID) Virology Training Grant No. T32 AI078985 (to University of Wisconsin-Madison) and GEIS P0062_20_NM_06 (to K.A.B-L.), and by the Federal Ministry of Education and Research within the research consortium “ZooBoCo” (Grant No. 01KI1722A). This work was also partially supported through the prime contract of Laulima Government Solutions, LLC, with NIAID under contract no. HHSN272201800013C and Battelle Memorial Institute’s former prime contract with NIAID under contract no. HHSN272200700016I. J.H.K. performed this work as a former employee of Battelle Memorial Institute and a current employee of Tunnell Government Services (TGS), a subcontractor of Laulima Government Solutions, LLC, under contract no. HHSN272201800013C. Additional support was provided through the German Center for Infection Research (DZIF) TTU “Emerging Infections,” and by the University of Wisconsin-Madison Global Health Institute, Institute for Regional and International Studies, and John D. MacArthur Professorship Chair (to T.L.G).
The views and conclusions contained in this document are those of the authors and should not be interpreted as necessarily representing the official policies or positions, either expressed or implied, of the US Department of Health and Human Services, Department of the Navy, Department of Defense, US government, or any of the institutions and companies affiliated with the authors. In no event shall any of these entities have any responsibility or liability for any use, misuse, inability to use, or reliance upon the information contained herein. The US departments do not endorse any products or commercial services mentioned in this publication. J.H.K and K.A.B-L. are employees of the United States government. This work was prepared as part of their official duties. Title 17 U.S.C. 105 provides that ‘copyright protection under this title is not available for any work of the United States Government.’ Title 17 U.S.C. 101 defines a US Government work as work prepared by a military service member or employee of the US Government as part of that person’s official duties. The study protocol was reviewed and approved by the University of Wisconsin-Madison Institutional Animal Care and Use Committee in compliance with all applicable federal regulations governing the protection of animals and research.
Footnotes
Competing interest declaration. The authors declare no competing interests.
Reprints and permissions information is available at www.nature.com/reprints.
References
- 1.Lambert N, Strebel P, Orenstein W, Icenogle J & Poland GA Rubella. Lancet 385, 2297–2307, doi: 10.1016/S0140-6736(14)60539-0 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Zhou Y, Ushijima H & Frey TK Genomic analysis of diverse rubella virus genotypes. J Gen Virol 88, 932–941, doi: 10.1099/vir.0.82495-0 (2007). [DOI] [PubMed] [Google Scholar]
- 3.Chen J-P, Strauss JH, Strauss EG & Frey TK Characterization of the rubella virus nonstructural protease domain and its cleavage site. J Virol 70, 4707–4713 (1996). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Perelygina L et al. Infectious vaccine-derived rubella viruses emerge, persist, and evolve in cutaneous granulomas of children with primary immunodeficiencies. PLoS Pathog 15, e1008080, doi: 10.1371/journal.ppat.1008080 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.DuBois RM et al. Functional and evolutionary insight from the crystal structure of rubella virus protein E1. Nature 493, 552–556, doi: 10.1038/nature11741 (2013). [DOI] [PubMed] [Google Scholar]
- 6.McCarthy M, Lovett A, Kerman RH, Overstreet A & Wolinsky JS Immunodominant T-cell epitopes of rubella virus structural proteins defined by synthetic peptides. J Virol 67, 673–681 (1993). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Maton WG Some account of a rash liable to be mistaken for scarlatina. Med Trans R Coll Physicians 5, 149–165 (1815). [Google Scholar]
- 8.Cooper LZ The history and medical consequences of rubella. Rev Infect Dis 7 Suppl 1, S2–10 (1985). [DOI] [PubMed] [Google Scholar]
- 9.Gregg NM Congenital cataract following German measles in the mother. Aust N Z J Ophthalmol 3, 35–46 (1941). [PubMed] [Google Scholar]
- 10.Parkman PD, Buescher EL & Artenstein MS Recovery of rubella virus from army recruits. Proc Soc Exp Biol Med 111, 225–230, doi: 10.3181/00379727-111-27750 (1962). [DOI] [PubMed] [Google Scholar]
- 11.Weller TH & Neva FA Propagation in tissue culture of cytopathic agents from patients with rubella-like illness. Proc Soc Exp Biol Med 111, 215–225 (1962). [Google Scholar]
- 12.Swan C, Tostevin AL & Black GH Final observations on congenital defects in infants following infectious diseases during pregnancy, with special reference to rubella. Med J Aust 2, 889–908 (1946). [PubMed] [Google Scholar]
- 13.Edmunds WJ, Gay NJ, Kretzschmar M, Pebody RG & Wachmann H The pre-vaccination epidemiology of measles, mumps and rubella in Europe: implications for modelling studies. Epidemiol Infect 125, 635–650 (2000). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Gonzales JA et al. Association of ocular inflammation and rubella virus persistence. JAMA Ophthalmol 137, 435–438, doi: 10.1001/jamaophthalmol.2018.6185 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Grant GB, Reef SE, Patel M, Knapp JK & Dabbagh A Progress in rubella and congenital rubella syndrome control and elimination - worldwide, 2000–2016. MMWR Morb Mortal Wkly Rep 66, 1256–1260, doi: 10.15585/mmwr.mm6645a4 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Namuwulya P et al. Phylogenetic analysis of rubella viruses identified in Uganda, 2003–2012. J Med Virol 86, 2107–2113, doi: 10.1002/jmv.23935 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Kretsinger K, Strebel P, Kezaala R & Goodson JL Transitioning lessons learned and assets of the global polio eradication initiative to global and regional measles and rubella elimination. J Infect Dis 216, S308–S315, doi: 10.1093/infdis/jix112 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Wolfe ND, Dunavan CP & Diamond J Origins of major human infectious diseases. Nature 447, 279–283 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Fahr J in Mammals of Africa, vol. IV: hedgehogs, shrews and bats (eds Happold M & Happold DCD) 380–383 (Bloomsbury, 2013). [Google Scholar]
- 20.Jetz W, McPherson JM & Guralnick RP Integrating biodiversity distribution knowledge: toward a global map of life. Trends Ecol Evol 27, 151–159, doi: 10.1016/j.tree.2011.09.007 (2012). [DOI] [PubMed] [Google Scholar]
- 21.O’Shea TJ, Bogan MA & Ellison LE Monitoring trends in bat populations of the United States and territories: status of the science and recommendations for the future. Information and Technology Report USGS/BRD/ITR–2003–0003. (U.S. Department of the Interior, U.S. Geological Survey; Washington, DC, USA, 2003). [Google Scholar]
- 22.Landau I & Chabaud A-G Description de Plasmodium cyclopsi n. sp. parasite du microchiroptère Hipposideros cyclops a Makokou (Gabon). Ann Parasitol Hum Comp 53, 247–253, doi: 10.1051/parasite/1978533247 (1978). [DOI] [PubMed] [Google Scholar]
- 23.Schaer J et al. High diversity of West African bat malaria parasites and a tight link with rodent Plasmodium taxa. Proc Natl Acad Sci U S A 110, 17415–17419, doi: 10.1073/pnas.1311016110 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Michaux JR, Libois R & Filippucci M-G So close and so different: comparative phylogeography of two small mammal species, the yellow-necked fieldmouse (Apodemus flavicollis) and the woodmouse (Apodemus sylvaticus) in the Western Palearctic region. Heredity (Edinb) 94, 52–63, doi: 10.1038/sj.hdy.6800561 (2005). [DOI] [PubMed] [Google Scholar]
- 25.Labuda M et al. Tick-borne encephalitis virus transmission between ticks cofeeding on specific immune natural rodent hosts. Virology 235, 138–143, doi: 10.1006/viro.1997.8622 (1997). [DOI] [PubMed] [Google Scholar]
- 26.Klempa B et al. Complex evolution and epidemiology of Dobrava-Belgrade hantavirus: definition of genotypes and their characteristics. Arch Virol 158, 521–529, doi: 10.1007/s00705-012-1514-5 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Sibold C et al. Dobrava hantavirus causes hemorrhagic fever with renal syndrome in central Europe and is carried by two different Apodemus mice species. J Med Virol 63, 158–167 (2001). [PubMed] [Google Scholar]
- 28.Oktem IM et al. Dobrava-Belgrade virus in Apodemus flavicollis and A. uralensis mice, Turkey. Emerg Infect Dis 20, 121–125, doi: 10.3201/eid2001.121024 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Doty JB et al. Isolation and characterization of Akhmeta virus from wild-caught rodents (Apodemus spp.) in Georgia. J Virol 93, e00966–00919, doi: 10.1128/JVI.00966-19 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Prpić J et al. First evidence of hepatitis E virus infection in a small mammal (yellow-necked mouse) from Croatia. PLoS One 14, e0225583, doi: 10.1371/journal.pone.0225583 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Hofmann J, Renz M, Meyer S, von Haeseler A & Liebert UG Phylogenetic analysis of rubella virus including new genotype I isolates. Virus Res 96, 123–128, doi: 10.1016/s0168-1702(03)00180-1 (2003). [DOI] [PubMed] [Google Scholar]
- 32.Abernathy E et al. Analysis of whole genome sequences of 16 strains of rubella virus from the United States, 1961–2009. Virol J 10, 32, doi: 10.1186/1743-422X-10-32 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Kelley LA, Mezulis S, Yates CM, Wass MN & Sternberg MJE The Phyre2 web portal for protein modeling, prediction and analysis. Nat Protoc 10, 845–858, doi: 10.1038/nprot.2015.053 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Wolinsky JS et al. An antibody- and synthetic peptide-defined rubella virus E1 glycoprotein neutralization domain. J Virol 67, 961–968 (1993). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Guy C, Thiagavel J, Mideo N & Ratcliffe JM Phylogeny matters: revisiting ‘a comparison of bats and rodents as reservoirs of zoonotic viruses’. R Soc Open Sci 6, 181182, doi: 10.1098/rsos.181182 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Luis AD et al. A comparison of bats and rodents as reservoirs of zoonotic viruses: are bats special? Proceedings. Biological sciences 280, 20122753, doi: 10.1098/rspb.2012.2753 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Olival KJ et al. Host and viral traits predict zoonotic spillover from mammals. Nature 546, 646–650, doi: 10.1038/nature22975 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Frey TK Neurological aspects of rubella virus infection. Intervirology 40, 167–175, doi: 10.1159/000150543 (1997). [DOI] [PubMed] [Google Scholar]
- 39.Bharadwaj SD et al. Acute encephalitis with atypical presentation of rubella in family cluster, India. Emerg Infect Dis 24, 1923–1925, doi: 10.3201/eid2410.180053 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Grant GB et al. Accelerating measles and rubella elimination through research and innovation - findings from the Measles & Rubella Initiative research prioritization process, 2016. Vaccine 37, 5754–5761, doi: 10.1016/j.vaccine.2019.01.081 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.