Abstract
The coronavirus disease 2019 (COVID-19) pandemic is a global public health crisis. However, little is known about the pathogenesis and biomarkers of COVID-19. Here, we profiled host responses to COVID-19 by performing plasma proteomics of a cohort of COVID-19 patients, including non-survivors and survivors recovered from mild or severe symptoms, and uncovered numerous COVID-19-associated alterations of plasma proteins. We developed a machine-learning-based pipeline to identify 11 proteins as biomarkers and a set of biomarker combinations, which were validated by an independent cohort and accurately distinguished and predicted COVID-19 outcomes. Some of the biomarkers were further validated by enzyme-linked immunosorbent assay (ELISA) using a larger cohort. These markedly altered proteins, including the biomarkers, mediate pathophysiological pathways, such as immune or inflammatory responses, platelet degranulation and coagulation, and metabolism, that likely contribute to the pathogenesis. Our findings provide valuable knowledge about COVID-19 biomarkers and shed light on the pathogenesis and potential therapeutic targets of COVID-19.
Keywords: COVID-19, SARS-CoV-2, proteomics, biomarkers, plasma
Graphical Abstract

Highlights
-
•
We profile plasma proteomics of COVID-19 cases at distinct symptoms and time points
-
•
The alterations of host plasma proteins are linked with COVID-19 development
-
•
Machine-learning-based models distinguish patients with different severity
-
•
Biomarker combinations show the power to predict COVID-19 clinical outcomes
Proteomic quantifications and experimental validation of plasma samples from three cohorts of COVID-19 patients with distinct symptoms at different time points identify differentially expressed host proteins that correlate with disease severity and prioritize biomarker combinations for accurately predicting COVID-19 clinical outcomes.
Introduction
The pandemic of coronavirus disease 2019 (COVID-19) has caused over millions of confirmed cases and hundreds of thousands of deaths worldwide as reported by the World Health Organization (WHO) (WHO, 2020) and has shown tremendous impacts on globe health and economics. COVID-19 is caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), which is the third coronavirus to cause severe respiratory disease in humans in addition to SARS-CoV and Middle East respiratory syndrome coronavirus (MERS-CoV) (Lu et al., 2020; Wu et al., 2020b; Zhou et al., 2020b; Zhu et al., 2020). SARS-CoV-2 has been found to mainly infect human low respiratory tract and lung, although many other organs, including the liver, kidney, muscle, gastrointestinal tract, lymph node, central nervous system, and heart, have also been found or proposed to be attacked by this virus (De Felice et al., 2020; Varga et al., 2020; Zhang et al., 2020a). According to a large-scale cohort study conducted by the Chinese Center for Disease Control and Prevention, more than 19% of COVID-19 patients have been reported to develop severe or critical conditions. In addition, although most COVID-19 patients only show mild symptoms, the conditions can rapidly develop from mild to severe and even critical illness, particularly when adequate medical care is insufficient (Guan et al., 2020). Moreover, the mortality rate of critically ill COVID-19 cases can even reach more than 60% (Yang et al., 2020b). Among the broad symptoms of COVID-19, fever, pneumonia, sepsis, respiratory failure, acute respiratory distress syndrome (ARDS), and multiorgan injury are frequently observed complications and are usually associated with the pathophysiological changes, such as alveolar macrophage activation, lymphopenia, cytokine release syndrome, microthrombosis, and intravascular coagulation in severe COVID-19 patients (Chen et al., 2020; Guan et al., 2020; Huang et al., 2020; Jose and Manuel, 2020; Moore and June, 2020; Wang et al., 2020a; Yang et al., 2020a). However, despite rapid and extensive efforts made to study this emerging coronavirus disease, the molecular mechanisms underlying the pathogenesis of COVID-19, particularly under pathophysiological conditions, are still poorly understood.
Alterations of human plasma proteins have been well recognized as indicators of pathophysiological changes caused by various diseases, including viral infections. Here, we profiled the host responses to SARS-CoV-2 infection in humans by performing quantitative proteomics of the plasma samples from a cohort of COVID-19 patients, including the non-survivors (fatalities) as well as survivors recovered from mild or severe symptoms. Our study uncovered a number of COVID-19-associated alterations of host proteins, particularly ones involved in inflammation and coagulation. Moreover, to identify potential biomarkers for accurate classification of different samples, we developed a machine-learning-based pipeline, resulting in the identification of 11 biomarkers as well as a set of biomarker combinations that could accurately distinguish or predict different COVID-19 outcomes. These biomarkers and combinations were further validated by the proteomic data from an independent cohort. Some of the identified biomarkers were further examined for their plasma levels via enzyme-linked immunosorbent assay (ELISA) of a larger cohort of COVID-19 patients, which are in line with the proteomic results. These biomarkers include host proteins that play critical roles in major pathophysiological pathways, and the abnormal alterations of these proteins in patient plasma probably contribute to the pathogenesis of COVID-19. These striking findings provided valuable knowledge about plasma biomarkers associated with COVID-19, shed light on the pathogenesis of SARS-CoV-2 infection, and might reveal potential therapeutic targets.
Results
Study Design and Patients
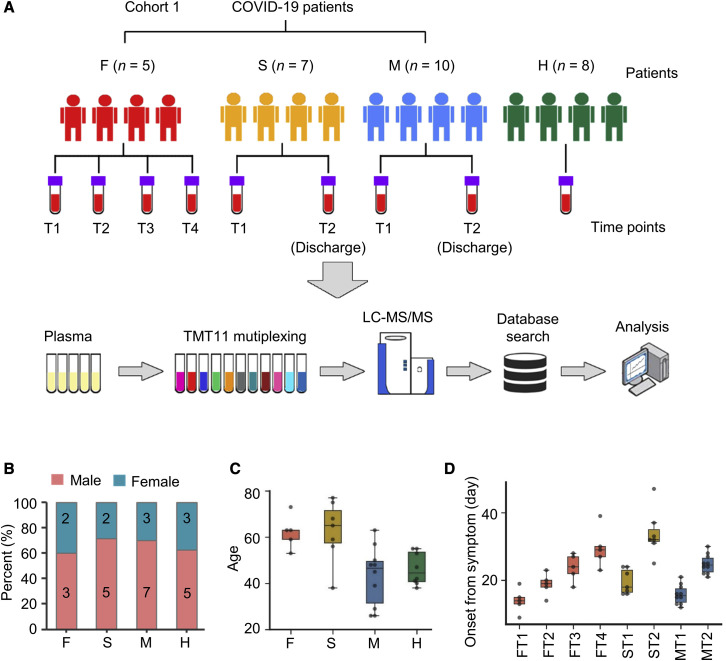
We collected the blood samples of a cohort of COVID-19 patients (cohort 1), including 5 patients with fatal (F) outcome, 7 patients diagnosed as severe (S) symptoms, and 10 patients diagnosed as mild (M) symptoms at Wuhan Jinyintan Hospital (Figures 1 A–1D; Table S1). The patients in the S and M groups had survived COVID-19 and been discharged from the hospital. Of note, blood samples were collected from the fatality patients along with the deterioration of this disease, as FT1 represents the first samples collected from this group of patients, and FT4 represents the last samples before any additional samples could be collected (Figure 1A). ST1 and MT1 represent the samples collected at the disease peak from the S and M groups, respectively, which were diagnosed based on National Health Commission (2020), although ST2 and MT2 represent the last samples collected from patients in each group shortly before they were discharged from the hospital (Figure 1A). Furthermore, blood samples from 8 healthy (H) subjects, whose throat swab and serological tests were negative for SARS-CoV-2, were collected for comparison.
Figure 1.
Study Design and Patients
(A) Overview of blood samples collection from COVID-19 patients (cohort 1), including F (n = 5), S (n = 7), M (n = 10) patients, and H volunteers (n = 8). T1–T4 mean different sample collection time points. The workflow for processing the proteomic data were shown, including the plasma separation, TMT 11-plex labeling, LC-MS/MS analysis, database search, and further computational analyses.
(B) The gender distribution of COVID-19 patients and H volunteers. The x axis represents different groups of cases, and the y axis represents the ratio of different genders (female or male).
(C) The age distribution of different groups.
(D) The number of days between symptom onset and the sample collection with different time points. Data points indicate the data of single patient at each time point and are presented as median with interquartile range (FT1–FT4, n = 5; ST1 and ST2, n = 7; MT1 and MT2, n = 10). The center line within each box shows the median, and the top and bottom of each box represent the 75th and 25th percentile values, respectively. The upper and lower whiskers extend from the hinge to the largest and smallest value no further than 1.5 times the distance between the first and third quartiles, respectively.
See also Table S1.
For each blood sample of cohort 1, plasma was separated and total proteins were extracted, denatured, and digested into peptides by trypsin (Figure 1A). Then, a total of 62 plasma samples were categorized into 7 batches and separately subjected to tandem mass tag (TMT) labeling (Table S2). For each batch, individual samples were labeled with TMT 11-plex reagents, and a pooling mixture of all the 62 samples was included and labeled as a standard control to eliminate the batch effect. After fractionation, each batch of peptide mixtures was analyzed by liquid chromatography with tandem mass spectrometry (LC-MS/MS). For database search, we constructed a human proteome database and also included a SARS-CoV-2 proteome database.
Proteomic Profiling of Plasma from COVID-19 Patients
From cohort 1, we obtained 8,472 peptides in total, with an average number range from 3,241.5 to 5,342.6 peptides in the 20 F, 14 S, 20 M, and 8 H samples (Figures 2 A, S1A, and S1B). We mapped these peptides to corresponding protein sequences, and the reporter ion MS2 module in the MaxQuant software package was used to quantify proteins (Tyanova et al., 2016). We found that 860 human proteins and 2 SARS-CoV-2 mature peptides or proteins were quantified in at least one sample (Table S3), with an average number of proteins ranging from 460.4 to 676.6 (Figures 2B and S1C). For cohort 1, lower numbers of peptides and proteins were identified in MT1, MT2, and H against other samples (Figures 2A and 2B), probably because of batch effect.
Figure 2.
Proteomic Profiling of Plasma from COVID-19 Patients and H Volunteers
(A and B) The distribution of numbers of quantified (A) peptides and (B) proteins in the 62 plasma samples. Error bars represent multiple independent samples, F (n = 5), S (n = 7), M (n = 10), H (n = 8).
(C) The distribution of MS/MS spectral counts of quantified peptides.
(D) The distribution of peptide numbers of quantified proteins.
(E) The distribution of protein numbers in plasma samples.
(F) The heatmap of the finally reserved proteins.
To evaluate the reliability of the proteomic data in cohort 1, we checked the raw MS/MS data and found that 6,705 peptides (79.1%) could be matched by ≥2 spectral counts (Figure 2C). The average spectral counts were calculated as 19.4 for all peptides, indicating the proteomic results are highly reliable at the peptide level. Also, we found that 472 proteins (54.8%) could be traced and supported by ≥2 peptides, with an average number of 10.0 peptides (Figure 2D). Thus, our quantification results are also highly reliable at the protein level. The distribution of proteins identified in different samples was analyzed, and we found that up to 348 (40.4%) proteins were simultaneously quantified in all the 62 samples (Figure 2E), indicating a high reproducibility of the proteomic profiling for cohort 1.
To ensure the data quality, only 530 proteins mutually quantified in >70% samples (≥44) were reserved for cohort 1. For each protein, multivariate normal imputation (MVNI) was applied to impute the missing values (Lee and Carlin, 2010). The principal-component analysis (PCA) of the 62 samples in the 7 batches was performed by using 351 proteins quantified in all samples with raw MS/MS values (Figure S1D), 348 proteins with normalized expression values (Figure S1E), or the 530 proteins after imputation (Figure S1F). Before normalization, it was found that the batches 5, 6, and 7 were very close and M and H samples were difficult to be separated (Figure S1D). After normalization, the M and H samples could be unambiguously distinguished, indicating the batch effect was greatly reduced (Figure S1E). The imputation did not influence the separation of different types of samples by normalization (Figure S1F). The F and S samples were not completely separated (Figures S1E and S1F), indicating the necessity of identifying potential biomarkers for classification of COVID-19 cases. Of note, this observation is consistent with the clinical observation that S and F COVID-19 patients presented certain overlapped manifestations (Chen et al., 2020; Guan et al., 2020; Huang et al., 2020; Wang et al., 2020a). The finally reserved proteins were analyzed by the hierarchical clustering method, and the results were visualized in a heatmap (Figure 2F). From the results, it could be found that a substantial number of proteins are differentially expressed in different types of plasma samples, indicating that single biomarkers or biomarker combinations might be found from the proteomic data.
Proteomic Alternations Associated with Clinical Symptoms of F and S COVID-19 Cases
We used our plasma proteomic data of cohort 1 to identify signatures of COVID-19 by analyzing plasma proteins that underwent significant fold changes (FCs) in F cases compared with those of H subjects (FT1–FT4 versus H, |log2(FC)| > 0.5; unpaired two-sided Welch’s t test; p < 0.05). A total of 195 differentially expressed proteins (DEPs) were identified under this condition, and the degree of differential expression of DEPs was obviously reduced in M group compared with S or F group (Figure S2A; Table S4), indicating that the alterations of plasma proteins became more extensive in more severe or deteriorated conditions. The DEPs were then subjected to Gene Ontology (GO) (The Gene Ontology Consortium, 2019) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway (Kanehisa et al., 2017) enrichment analyses. The GO terms and KEGG pathways of DEPs were highly enriched in processes involved in inflammation, immune cell migration and degranulation, complement system, coagulation cascades, and energy metabolism (Table S5). These results are consistent with the previous reports that acute inflammation and excessive immune cell infiltration are associated with the severity of COVID-19 patients (Chen et al., 2020; Guan et al., 2020; Huang et al., 2020; Wang et al., 2020a; Yang et al., 2020a).
We found that two processes, platelet degranulation and the complement and coagulation cascades, obtained the highest enrichment ratio (E ratio) scores in the GO and KEGG analyses, respectively (Figures 3A and 3B). Also, the proteins involved in these two processes were more dramatically altered in FT1–FT4 and ST1 compared to those in ST2, MT1, and MT2 (Figures 3C and 3D). These results were consistent with our clinical data that the coagulation tests, including D-dimer, the prothrombin time (PT), and the activated partial thromboplastin time (APTT), showed significant abnormality in FT1–FT4 and ST1 compared with those in ST2, MT1, MT2, and H (Figures 3E–3G; Table S6). These findings imply that the dysfunction of platelet degranulation and coagulation cascades are closely related with the severity of COVID-19.
Figure 3.
Proteomic Alternations Associated with Clinical Symptoms of F and S Cases
(A) GO-based enrichment analysis of DEPs shown in the term of biological processes (two-sided hypergeometric test; p < 0.001) and the number of counts (m > 10). GO terms were sorted by E-ratio.
(B) KEGG-based enrichment analysis of DEPs (two-sided hypergeometric test; p < 0.001) and the number of counts (m > 5). KEGG terms were sorted by E-ratio.
(C and D) Plasma levels of proteins in each group in relation to H group and the associated p values in the terms of platelet degranulation (C) and complement and coagulation cascades (D).
(E–G) Clinical data of D-dimer (E), prothrombin time (PT) (F), and activated partial thromboplastin time (APTT) (G) (y axis) in the indicated groups (x axis). Data points indicate the data of single patient at each time point and are presented as median with interquartile range (FT1–FT4, n = 5; ST1 and ST2, n = 7; MT1 and MT2, n = 10). The center line within each box shows the median, and the top and bottom of each box represent the 75th and 25th percentile values, respectively. The upper and lower whiskers extend from the hinge to the largest and smallest value no further than 1.5 times the distance between the first and third quartiles, respectively.
In addition to host proteins, we also identified two SARS-CoV-2-encoded proteins, nsP2 and nsP7, in the plasma samples of S (five out of seven S patients) and F groups (two out of five F patients), respectively, although neither of them could be found in the samples of M or H groups (Figure S2B), suggesting that the presence of these two viral proteins in patient plasma probably contributes to COVID-19 pathogenesis.
Machine-Learning-Based Selection of Biomarker Combinations for Classification of COVID-19 Cases
On the basis of the plasma proteomic data of cohort 1, we developed a new computational pipeline named Prioritization of Optimal biomarker Combinations for COVID-19 (POC-19) for identifying potential biomarker combinations to classify COVID-19 cases (Figure 4 A). POC-19 contains three steps, including differential protein reservation (DPR) to select 112 highly ranked DEPs, candidate biomarker selection (CBS) to generate 1,000 groups of initial biomarker combinations, and final biomarker determination (FBD) to get the protein combination with the highest area under the curve (AUC) value from the 5-fold cross-validation (Figure 4A). In the step of FBD, a widely used machine learning algorithm, penalized logistic regression (PLR) (Ning et al., 2020), was used for model training and parameter optimization.
Figure 4.
Identification of Potential Biomarker Combinations for the Classification of COVID-19 Patients and H Volunteers by Using a Machine-Learning Strategy
(A) The workflow of POC-19, including DPR, CBS, and FBD to prioritize highly potential biomarker combinations. In the step of FBD, LASSO and ridge regression penalties in PLR were adopted for model training and parameter optimization.
(B) From the 5-fold cross-validation, AUC values were calculated for the classification of F, S, and M COVID-19 patients and H volunteers, respectively.
(C) The confusion matrix of the 4-protein combination.
(D) The PCA analysis of the 4 proteins among different plasma samples.
(E) Overview of blood samples collection from cohort 2, including F (n = 9), S (n = 6), and M (n = 6) patients and H volunteers (n = 5).
(F) AUC values were calculated for the classification of F, S, and M COVID-19 patients and H volunteers based on cohort 2.
(G) The confusion matrix of the 4-protein combination in cohort 2.
(H) The PCA analysis of the 4 proteins among different plasma samples from cohort 2.
For the classification of COVID-19 patients and H volunteers, we identified a compact biomarker combination containing 4 proteins, including orosomucoid-1/alpha-1-acid glycoprotein-1 (ORM1/AGP1), ORM2, fetuin-B (FETUB), and cholesteryl ester transfer protein (CETP) (Figure 4B). Using cohort 1, the 5-fold cross-validation AUC values of this 4-protein combination to distinguish F, S, M, and H groups were calculated as 0.952 (95% confidence interval [CI] = 0.892–0.987), 0.917 (95% CI = 0.873–0.955), 0.974 (95% CI = 0.901–0.992), and 0.983 (95% CI = 0.916–1.000), respectively (Figure 4B). Moreover, the 5-fold cross-validation AUC values of each of the 4 proteins were determined, indicating that, even when being used alone, these proteins can still be informative to distinguish different groups in most cases (Figures S3A–S3D). To evaluate the reliability of the machine-learning strategy, confusion matrices were illustrated, and the results demonstrated that different samples could be correctly classified in a high accuracy (Figure 4C). Moreover, the PCA showed that the clustering of samples is clearly classified into different groups (Figure 4D), indicating the reliability of POC-19 for distinguishing different COVID-19 groups and H volunteers.
To validate the accuracy of the machine-learning-based classification of COVID-19 cases, we further collected 26 plasma samples of a new cohort (cohort 2), including 9 F, 6 S, and 6 M patients, together with 5 H volunteers (Figure 4E). The 26 plasma samples were categorized into 3 batches for LC-MS/MS analysis and database search, and the pooling mixture of the 62 samples of cohort 1 was used as the control for each batch (Table S2). The numbers of peptides and proteins were similar across different samples in cohort 2 (Figures S3E–S3G). From the proteomic data, we found that the 4 proteins, including ORM1, ORM2, FETUB, and CETP, were quantified in all samples (Table S3). Thus, we directly used the relative abundance values of the 4 proteins to evaluate the performance of POC-19. The AUC values were calculated as 0.941 (95% CI = 0.848–0.978), 0.825 (95% CI = 0.765–0.881), 0.842 (95% CI = 0.788–0.902), and 1.000 (95% CI = 1.000) for predicting F, S, M, and H cases, respectively (Figure 4F). The corresponding confusion matrix and PCA results also demonstrated that POC-19 exhibited a promising accuracy on the independent cohort (Figures 4G and 4H; Table S7).
The Biomarker Combinations for Prediction of Different Clinical Outcomes of COVID-19 Patients
We sought to utilize POC-19 to predict different clinical outcomes (e.g., S to F, M to S, and cases curable from the disease) on the basis of the DEPs from cohort 1. As a result, we selected a biomarker combination containing 3 proteins of CETP, S100A9, and C-reactive protein (CRP), which reached an AUC value of 0.929 (95% CI = 0.867–0.975) to predict severe COVID-19 patients with or without fatal outcome (Figure 5 A). The AUC values of each protein in this combination to discriminate F from S (Figures S4A–S4C) or each F time point (FT1–4) from ST1 (Figure S4D) were also determined, ranging from 0.757 to 0.914.
Figure 5.
Determination of Biomarker Combinations for Predicting Different COVID-19 Outcomes
(A–C) The receiver operating characteristic (ROC) curve (A), confusion matrix (B), and PCA plot (C) for the prediction of S to F outcome.
(D–F) The ROC curve (D), confusion matrix (E), and PCA plot (F) for the prediction of M to S outcome.
(G–I) The ROC curve (G), confusion matrix (H), and PCA plot (I) for the prediction of COVID-19 patients cured from the disease.
See also Figures S4 and S5 and Table S7.
To predict M to S outcome, POC-19 identified a 3-protein combination containing zinc-α2-glycoprotein 1 (AZGP1), ORM2, and complement factor I (CFI), either being used as a combination or separately, with AUC values of 1.000 (95% CI = 1.000) (Figures 5D and S4E–S4G). Of note, patients in M group were younger than those in S or F group (Figure 1C; Table S1), consistent with multiple clinical observations (Guan et al., 2020). If we consider age as a possible factor for discriminating S from M cases, its AUC value is 0.792 (Figure S4H), although the AUC value of the combination of the three proteins plus age is 1 (Figure S4I), same as that without considering age as a factor (Figure 5D). Thus, age is a factor, but not a decisive one, to interfere with the effectiveness and accuracy of this 3-protein combination in our model.
In addition, we further prioritized a biomarker combination containing serine proteinase inhibitor A3/α1-antichymotrypsin (SERPINA3/ACT), lymphocyte cytosolic protein 1/L-plastin (LCP1/LPL), and peptidase inhibitor 16 (PI16), with an AUC value 0.947 (95% CI = 0.887–0.985) for predicting convalescence (Figure 5G). The AUC values of individual proteins in this combination ranged from 0.832 to 0.941 (Figures S4J–S4L). Furthermore, the results of confusion matrices and PCA of these biomarker combinations showed high accuracy for classifying and clustering different groups (Figures 5B, 5C, 5E, 5F, 5H, and 5I). Normalized expression of each biomarker in different groups was shown in Figure S5.
The Alterations of Host Proteins in Plasma Are Linked with COVID-19 Development
In addition to identifying promising biomarkers for COVID-19, our work also revealed numerous alterations of host plasma proteins that might contribute to the pathogenesis of COVID-19. For instance, the plasma levels of ORM1, ORM2, S100A9, CRP, AZGP1, CFI, SERPINA3/ACT, and LCP1/LPL were significantly elevated in more severe COVID-19 conditions (Figures S5A–S5H). Among them, ORM1, ORM2, S100A9, CRP, and SERPINA3/ACT are acute-phase proteins (APPs) whose alterations in plasma are usually in response to inflammation, infection, or tissue injury (Gabay and Kushner, 1999), although LCP1/LPL and CFI are also involved in regulating immune responses. Besides, AZGP1 is an adipokine functionally implicated in lipid metabolism (Hassan et al., 2008; Liu et al., 2018), consistent with our previous observation that host metabolism was altered by COVID-19 (Wu et al., 2020a). Of note, the clinical data of CRP obtained from the medical record of this cohort were consistent with our proteomic results (Figure S5I; Table S6).
Besides these elevated plasma proteins, we also found the levels of FETUB, CETP, and PI16 were significantly reduced in more severe conditions (Figures S5J–S5L). FETUB is involved in fatty acid metabolism and can suppress inflammation via inhibiting merprins (Choi et al., 2012; Karmilin et al., 2019; Meex et al., 2015). CETP promotes lipid transfer between lipoproteins and can act as an inhibitor of prolonged inflammatory response, and the reduction of plasma CETP was associated with mortality in patients with severe sepsis (Martinelli et al., 2018; Venancio et al., 2016). PI6 has been found to suppress chemotaxis of some leukocytes, macrophages, and dendritic cells by inhibiting the chemokine chemerin (Regn et al., 2016). Therefore, the reduction of these proteins in patient plasma might also contribute to elevated inflammation and/or metabolic disorder.
Validation of the Biomarkers of Different COVID-19 Outcomes
To further verify the biomarkers obtained from the proteomic data, we expanded our findings in an additional cohort that includes 40 F, 40 S, and 40 M patients plus 40 H volunteers (cohort 3) (Figure 6 A). The number of days between symptom onset and sample collection of different groups was shown in Figure 6B and Table S8. Plasma samples from these patients were collected and then subjected to ELISA analyses for detecting the plasma levels of ORM1, AZGP1, CFI, FETUB, and S100A8/S100A9, respectively. Of note, S100A9 and its partner S100A8, which was also detected in our study, function as a heterodimer that can be detected together. The ELISA results showed that the plasma levels of ORM1, S100A8/S100A9, AZGP1, and CFI were significantly elevated in more severe COVID-19 conditions, although the level of FETUB was significantly reduced in more severe ones (Figures 6C–6G; Table S8), which are consistent with our proteomic findings.
Figure 6.
Serological Validation of COVID-19 Biomarkers
(A) Overview of blood samples collection from cohort 3, including F (n = 40), S (n = 40), and M (n = 40) patients and H volunteers (n = 40).
(B) The number of days between symptom onset and the sample collection with different time points.
(C–G) Plasma levels of the indicated proteins from the samples of cohort 3 were detected via ELISA. Data points indicate the data of single patient that are presented as median with interquartile range (F, n = 40; S, n = 40; M, n = 40; H, n = 40). The center line within each box shows the median, and the top and bottom of each box represent the 75th and 25th percentile values, respectively. The upper and lower whiskers extend from the hinge to the largest and smallest value no further than 1.5 times the distance between the first and third quartiles, respectively. Data were analyzed by unpaired two-sided Welch’s t test. ∗p < 0.05; ∗∗p < 0.01; ∗∗∗p < 0.001.
Furthermore, the statistical significance of each of the 11 proteins was further tested by using one-way analysis of variance (ANOVA) (p < 0.05). To control the familywise error rate (FWER), a permutation test (times = 100,000) was conducted, and the adjusted p values were calculated by Westfall and Young’s step-down min P approach to correct for multiple hypothesis testing (Oughtred et al., 2019). It could be found that all the proteins were significant to classify different types of samples (Table S9; p < 0.05; adjusted p < 0.05). One exception was S100A9 with a non-significant value (p = 0.108; adjusted p = 0.113) in distinguishing F and S samples, although the combination of CETP, S100A9, and CRP achieved a promising accuracy to predict severe COVID-19 patients with or without fatal outcome.
In summary, these results further confirmed the accuracy of our proteomic data and, more importantly, validated that the biomarkers identified in this study have promising potentials to clinically monitor and evaluate the progression of COVID-19.
Discussion
The pandemic of COVID-19 has been one of the worst public health crises. It is of high priority to identify biomarkers that can monitor and predict the development of the disease and understand its pathogenesis. For this purpose, we conducted proteomics to profile plasma protein alterations in response to COVID-19 under different conditions and identified 11 host proteins and a set of biomarker combinations that can serve as biomarkers by using the machine-learning-based pipeline POC-19 developed by us. These biomarkers can classify and predict the outcomes of COVID-19. Moreover, the alternations of these host proteins provide very valuable insight for the pathogenesis of COVID-19. Strikingly, many biomarkers could be individually used to distinguish or predict COVID-19 outcomes, indicating that the alterations of these plasma proteins are closely linked with the disease. Moreover, the accuracy of these biomarkers to distinguish COVID-19 outcomes were further validated via proteomics and ELISA using the plasma samples from two additional cohorts of COVID-19 patients, respectively. These results confirmed that the altered plasma proteins identified in this study indeed reflect the authentic pathophysiological changes in response to COVID-19 and minimized the possibility that the host protein alternations were influenced by other factors. Therefore, these proteins show promising potentials to be further developed as clinical biomarkers, either individually or in combination, to closely monitor and evaluate the development of COVID-19, thereby providing timely advice for clinical treatment.
Intriguingly, the alterations of many plasma proteins uncovered here well correspond to the severity and pathophysiology of COVID-19. For instance, multiple APPs showed significant elevation in the plasma of patients with more severe outcomes. Among them, CRP, ORM1, ORM2, and SERPINA3 are inflammatory factors that are controlled by multiple cytokines, such as interleukin-1β (IL-1β), tumor necrosis factor-α (TNF-α), IL-6, and IL-6-related cytokines (Ceciliani and Lecchi, 2019; Cichy et al., 1995; Fournier et al., 2000; Luo et al., 2015; Tyagi et al., 2013; Yaprak et al., 2018), although S100A9 and its partner S100A8, also identified here, can induce inflammatory cytokines and immune cell migration (Wang et al., 2018). Moreover, CFI is a key component of complement system (Lachmann, 2019) and LCP1/LPL is a critical regulator of T cell and alveolar macrophage activation (Deady et al., 2014; Todd et al., 2016). On the other hand, some negative regulators of inflammation, such as FETUB, CETP, and PI16, showed a downward trend along with the deterioration of the disease in this study. Besides, many of these proteins, such as the APPs, CETP, and PI16, also participate in platelet dysfunction/aggregation and activation of coagulation cascades. Additionally, the significantly altered proteins identified in this study are generally involved in several major biological processes, of which the identified biomarkers are included in the processes of different immune responses, platelet degranulation and coagulation, and metabolism (Figure 7 ). These results are in accordance with previous clinical or autopsy observations that S COVID-19 cases are frequently associated with massive intravascular thrombus, hypoxemia, ARDS, sepsis, and multiorgan injury (Guan et al., 2020; Ranucci et al., 2020; The Novel Coronavirus Pneumonia Emergency Response Epidemiology Team, 2020; Xu et al., 2020b), which are pathophysiologically associated with cytokine release syndrome, alveolar macrophage activation, intravascular coagulation, and microthrombosis (Moore and June, 2020).
Figure 7.
A Plasma Protein Regulatory Network Associated with COVID-19
In the network, the 173 DEPs were classified into 9 groups on the basis of their major functions, including immune cell migration, complement activation, metabolic process, platelet/neutrophil degranulation, blood coagulation, cell cycle/proliferation, phagocytosis/endocytosis, transport/cell adhesion, and other immune response. The color-coded circular boxes represent the 11 proteins in the four biomarker combinations as indicated at the upper right corner.
Furthermore, based on our findings, it would be intriguing to speculate that some significantly altered proteins and related pathways could be promising therapeutic targets for COVID-19. For example, some clinically approved anticoagulants, such as proteinase-activated receptor-1 (PAR-1) antagonists, antithrombin, and antifactor Xa, might ameliorate COVID-19 severity associated with intravascular coagulation and inflammation. In addition, the S100A8/S100A9 complex is a danger-associated molecular pattern (DAMP) that promotes inflammation, and its extracellular functions include neutrophil and leukocyte recruitment, proinflammatory cytokine release, and apoptotic induction (Wang et al., 2018). Given that plasma S100A8 and S100A9 were significantly elevated in more severe COVID-19 and the S100A8/S100A9 inhibitors, quinoline-3-carboxamide compounds, have shown promising outcomes in treating inflammatory diseases with good safety records in clinical trials (Bengtsson et al., 2012; Björk et al., 2009), targeting S100A8/S100A9 might represent a promising strategy for treating severe or critically ill COVID-19 cases. Moreover, several clinically approved antibiotics, such as vancomycin, lincomycin, and erythromycin (Banères-Roquet et al., 2009), have been bound to inhibit the plasma levels of ORMs/AGPs, including ORM1/AGP1 and ORM2/AGP2, which might be helpful for alleviating symptoms. Future efforts should be made to test these possibilities.
Limitations of Study
There are some limitations in our study. First, most of the samples were collected from the patients in the early period of COVID-19 outbreak. Due to the priority to save patients’ lives at that time, the number of plasma samples for proteomic profiling was limited, and the samples of very early time points or early acute phase (within 1 week from onset) of this disease were omitted. Therefore, although we had used three different cohorts and two different approaches (i.e., mass spectrometry and ELISA) to generate consistent results, it would be ideal to involve more clinical samples, probably from earlier time points and multiple centers, to further validate the biomarkers we identified. Another possible drawback of this work is that different therapeutic strategies used during the treatment of different patients might affect the results, although the protein alterations uncovered here are quite consistent in different cohorts. Future studies should evaluate the impacts of different therapies on host responses. In addition, the data variance exists in all omics studies. Actually, the batch effect was not strong in our raw MS/MS data, because an identical internal reference was used for all the 10 batches of TMT labeling experiments. Besides statistical analyses, we also performed additional experiments for validation, which showed that the potential data variance did not affect our findings. Lastly, the detailed roles of the biomarker proteins in the pathogenesis of COVID-19 require further investigation or the potential therapeutic targets, such as S100A8/S100A9 and ORM1/ORM2, should be further elucidated or experimentally validated.
In summary, this work provides a highly valuable proteomics resource for the research community to better understand COVID-19-associated host responses, sheds light on the pathogenesis of SARS-CoV-2 infection, identifies a serial of valuable biomarker candidates, and provides hints of potential therapeutic strategies.
STAR★Methods
Key Resources Table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Biological Samples | ||
| Blood samples from human patients infected with SARS-CoV-2 | This paper | N/A |
| Blood samples from healthy donors | This paper | N/A |
| Deposited Data | ||
| Raw Proteomic data | This paper | iProX: PXD019106 |
| Critical Commercial Assays | ||
| Human Complement Factor I (CFI) ELISA Kit | CUSABIO | Cat#CSB-E13108h |
| Human Fetuin-B (FETUB) ELISA Kit | CUSABIO | Cat#CSB-EL008598HU |
| Human Zinc-Alpha-2-Glycoprotein (AZGP1) ELISA Kit | CUSABIO | Cat#CSB-EL002479HU |
| Human S100 Calcium-Binding Protein A8/A9 Complex (S100A8/A9) ELISA Kit | CUSABIO | Cat#CSB-E12149h |
| Human ORM1 ELISA Kit | ProteinTech | Cat#KE00137 |
| Software and Algorithms | ||
| MaxQuant V1.6.6 | https://www.maxquant.org/ | N/A |
| Python 3.7 | https://www.anaconda.com/ | N/A |
| Scikit-learn 0.22.1 | https://scikit-learn.org/stable/ | N/A |
| R package 4.0.2 | https://www.r-project.org/ | N/A |
| Multtest 2.44.0 | https://bioconductor.org/packages/release/bioc/html/multtest.html | N/A |
| Gene Ontology (GO) | https://www.geneontology.org/ | N/A |
| KEGG Database | https://www.genome.jp/kegg/ | N/A |
| HemI | https://hemi.biocuckoo.org/ | N/A |
| IBS | https://ibs.biocuckoo.org/ | N/A |
| UniProt | https://www.uniprot.org/ | N/A |
| NCBI | https://www.ncbi.nlm.nih.gov/ | N/A |
| BioGrid | https://thebiogrid.org/ | N/A |
| IID | https://iid.ophid.utoronto.ca/ | N/A |
| InBio Map™ | https://www.intomics.com/inbio/map | N/A |
| Mentha | https://mentha.uniroma2.it/ | N/A |
| HINT | https://hint.yulab.org/ | N/A |
| iRefIndex | https://irefindex.org/ | N/A |
| PINA | https://cbg.garvan.unsw.edu.au/pina | N/A |
| Cytoscape 3.6.1 software | https://cytoscape.org/ | N/A |
| GraphPad Prism v8.0 | https://www.graphpad.com | N/A |
Lead Contact and Materials Availability
Further information and request for resources and reagents should be directed to and will be fulfilled by the Lead Contact, Xi Zhou (zhouxi@wh.iov.cn).
Experimental Model and Subject Details
Ethics and Human Subjects
All work performed in this study was approved by the Wuhan Jinyintan Hospital Ethics Committee and written informed consents were obtained from patients. Diagnosis of SARS-CoV-2 infection was based on the New Coronavirus Pneumonia Prevention and Control Program (6th edition) published by the National Health Commission of China (National Health Commission, 2020). H subjects were recruited from healthcare workers and laboratory workers at Wuhan Jinyintan Hospital and Wuhan Institute of Virology, CAS, none of whom had previously experienced SARS-CoV-2 infection.
Patient Samples
SARS-CoV-2-positive patients were enrolled in the study after diagnosis. The severity of COVID-19 was determined by the attending doctors based on the clinical diagnostic guideline of Chinese Health Commission (6th edition) and previous studies (Xu et al., 2020a; Zhou et al., 2020a), which reveal the clinical courses of COVID-19 patients.
Blood sample (≤3 mL) from F patients were collected over the course of their disease at intervals of 3-5 days. Blood sample (≤3 mL) from the patients with S and M symptoms were collected at the time when the disease was most serious (3-7 days after hospitalization) and the time before discharged. Single samples were collected from the patients in Cohorts 2 and 3 as well as H volunteers. All the patients in Cohorts 1 and 2 were also included in Cohort 3 for ELISA validation. The throat swabs and serological testing of H volunteers were negative for SARS-CoV-2. All blood samples were collected after fasting overnight and by added with ethylene diamine tetraacetic acid (EDTA) plus potassium (K+). All the blood samples were treated according to the biocontainment procedures of the processing of SARS-CoV-2-positive samples.
Method Details
Biosafety
All the blood samples were treated according to the biocontainment procedures of the processing of SARS-CoV-2-positive sample.
Sample preparation
Ten microliters of plasma were mixed with 190 μL reaction solution (1% SDC, 10 mM TCEP, and 40 mM CAA). The reaction was performed at 60°C for 30 min for protein denaturation, disulfide bond reduction, and cysteine -SH alkylation. Protein concentration was measured by Bradford method. The samples were diluted with equal volume of H2O. Trypsin was added at a ratio of 1:50 (enzyme: protein, w/w) for overnight digestion at 37°C. After centrifugation (12,000 g, 15 min), the supernatant was subjected to peptide purification using self-made desalting columns filled with Poly(styrene-divinylbenzene) copolymer (SDB) materials as described (Rappsilber et al., 2007). The peptide eluate was vacuum dried and stored at −20°C for later use.
TMT labeling was performed according to manufacturer’s instructions. Briefly, peptides were reconstituted in TMT reagent buffer, and the samples were separately labeled with different TMT labeling reagents. The internal reference sample pooled from all the 62 samples of Cohort 1 (equal contribution) was labeled using channel 126 for each batch of TMT labeling experiment (Both Cohort 1 and 2), allowing comparison of relative protein abundance across different TMT experiments. The labeled samples were then mixed and subjected to Sep-Pak C18 desalting. The labeling efficiency of each labeled mixture was examined by mass spec identification of ∼2 μg of the mixture with TMT (N-terminal/K) as variable modifications. The labeling efficiency (calculated from the ratio of number of TMT labeled sites divided by number of all the potential labeling sites) had to pass the threshold of 95% before proceeding to the fractionation step. The remaining mixture for each group of TMT experiment was fractionated using high pH reverse phase chromatography into 60 fractions and further concatenated into 20 fractions (by combining of fractions 1, 21 and 41; fractions 2, 22 and 42; and so on) (Batth et al., 2014). Each fraction was vacuum-dried and stored at −80°C until MS analysis.
Before the protein denaturation, 20 μg of total plasma proteins were mixed with 5 × SDS-PAGE loading dye (10% SDS, 500 mM DTT, 50% Glycerol, 500 mM Tris-HCL and 0.05% bromophenol blue, pH 6.8), boiled for 10 min and then analyzed by 12% SDS-PAGE. According to the SDS-PAGE results (Figures S1A and S3E), all bands were clear and homogeneous without protein degradation. All the protein extractions of the 62 plasma samples were classified as Class A which represents the highest quality and the proteomic profiling could be performed at least twice for each sample (Wang et al., 2020b).
LC-MS/MS analysis
LC-MS/MS data acquisition was carried out on a Q Exactive HF-X mass spectrometer coupled with an Easy-nLC 1200 system (both Thermo Scientific) (Miao et al., 2019; Zhang et al., 2020b). Peptides were first loaded onto a C18 trap column (75 μm × 2 cm, 3 μm particle size, 100 Å pore size, Thermo) and then separated in a C18 analytical column (75 μm × 250 mm, 3 μm particle size, 100 Å pore size, Thermo). Mobile phase A (0.1% formic acid) and mobile phase B (80% acetonitrile, 0.1% formic acid) were used to establish a 90 min separation gradient (0 min – 8% B; 67 min – 30% B; 82 min – 45% B; 83 min – 90% B; 90 min – 90% B). A constant flow rate was set at 300 nL/min. For the analysis in data-dependent acquisition (DDA) mode, each scan cycle consisted of one full-scan mass spectrum (R = 120 K, AGC = 3e6, max IT = 50 ms, scan range = 350–1800 m/z) followed by 20 MS/MS events (R = 45 K, AGC = 1e5, max IT = 86 ms). High energy collision dissociation (HCD) collision energy was set to 32. Isolation window for precursor selection was set to 1.2 Da. Former target ion exclusion was set for 45 s.
Database search
MS raw data were analyzed with MaxQuant (V1.6.6) using the Andromeda database search algorithm (Miao et al., 2019; Tyanova et al., 2016). The human proteome database contained 20,366 Swiss-Prot/reviewed human protein sequences downloaded from the UniProt database (https://www.uniprot.org/proteomes/UP000005640, on March 17, 2020) (UniProt Consortium, 2019), whereas the SARS-CoV-2 proteome database contained 12 protein sequences including ORF1ab (YP_009724389.1), ORF1a (YP_009725295.1), S (YP_009724390.1), ORF3a (YP_009724391.1), E (YP_009724392.1), M (YP_009724393.1), ORF6 (YP_009724394.1), ORF7a (YP_009724395.1), ORF7b (YP_009725318.1), ORF8 (YP_009724396.1), ORF9 (YP_009724397.2), and ORF10 (YP_009725255.1) derived from its CDS regions, including (https://www.ncbi.nlm.nih.gov/nuccore/NC_045512.2, on March 17, 2020) (Wu et al., 2020b). The two databases were concatenated and reverse decoy sequences were generated. Then, spectra files were searched against the merged database using the following parameters: Type, TMT; Variable modifications, Oxidation (M), Deamidation (NQ), Acetyl (Protein N-term); Fixed modifications, Carbamidomethyl (C); Digestion, Trypsin/P. The MS1 match tolerance was set as 20 parts per million (ppm) for the first search and 4.5 ppm for the main search; the MS2 tolerance was set as 20 ppm. Search results were filtered with 1% false discovery rate (FDR) at both protein and peptide levels. Proteins denoted as decoy hits, contaminants, or only identified by sites were removed, and the remaining proteins were used for further analysis.
Enzyme-linked immunosorbent assay (ELISA)
Human protein ELISA kits were used to quantify plasma levels of endogenous protein according to manufacturers’ instructions. Briefly, plasma samples were diluted according to the manufacturers’ dilution guideline. Plasma were diluted at 1: 400 for detection of AZGP1 and CF1, 1:10 or 1:40 for FETUB, 1:10 for S100A8/S100A9 and 1: 100000 for ORM1. Then, a total of 100 μL of fixed dilution plasma sample were added to the precoated plates, and the plates were incubated at 37°C for 2 hr. After washing, 100 μL biotinylated-specific antibody was added to each well, and the plates were incubated at 37°C for 1 hr. Followed by washing, 100 μL sterptravidin-HRP was added and incubated at 37°C for 1 hr. Finally, the OD value at 450 nm were determined after addition of 100 μL Tetramethyl-benzidine (TMB) reagent and stop solution. The standard curve of each protein was generated by determination of OD values from serially dilutions of the standard samples with known protein concentrations provided by the manufacturers.
Quantification and Statistical Analysis
Proteomic data normalization and imputation
For each batch of the plasma proteomic data, the abundance of a protein in one patient sample was normalized against its corresponding abundance in the control sample to get the relative protein abundance, which was used for further analyses across different batches. For each batch, proteins not detected in control were discarded. The normalized expression values of proteins in Cohorts 1 and 2 were present in Tables S3.
To ensure the data quality and maximally use the proteomic data, proteins quantified in < 70% samples of Cohort 1 were discarded. To impute missing values of remaining proteins, we used a model-based method named MVNI, which assumes all data points jointly following a multivariate normal distribution (Lee and Carlin, 2010). For each protein, the multivariate normal distribution was modeled, and the missing values were imputed with the maximum likelihood estimation. The proteomic data imputation was performed by the multivariate_normal function in scipy.stats, a powerful Python module for data statistics.
Statistical analysis of the quantitative proteomic data
For Cohort 1, we identified potential DEPs that were significantly altered in F cases against H volunteers. The mean values of the relative abundances of each protein were calculated for FT1-FT4 and H samples, respectively. The FC value was calculated based on the ratio of FT1-FT4/H, and proteins with |log2(FC)| > 0.5 were reserved. Because the variances might not be equal in FT1-FT4 and H samples, the statistical significance was calculated for reserved proteins, using the unpaired two-sided Welch’s t test (p < 0.05) (Table S4). For these proteins, the FC values of FT1/H, FT2/H, FT3/H, FT4/H, ST1/H, ST2/H, MT1/H and MT2/H were also calculated, and corresponding p values were computed by the unpaired two-sided Welch’s t test (Table S4). The statistical analyses were conducted by the ttest_ind function in scipy.stats. The multiple testing correction was not performed.
The enrichment analyses
The two-sided hypergeometric test was adopted for the enrichment analysis of the 195 DEPs. Here, we defined:
N = number of human proteins annotated by at least one term
n = number of human proteins annotated by term t
M = number of the 195 proteins annotated by at least one term
m = number of the 195 proteins annotated by term t
Then, the E-ratio was calculated, and the p value was computed with the hypergeometric distribution as below:
In this study, only statistically enriched GO terms (p < 0.001, m > 10) and KEGG pathways (p < 0.001, m > 5) were considered. GO annotation files (on 03 January 2020) were downloaded from the Gene Ontology Consortium Web site (http://www.geneontology.org/), and we obtained 19,714 human proteins annotated with at least one GO biological process term. KEGG annotation files (released on 4 December 2017) were downloaded from the ftp server of KEGG (ftp://ftp.bioinformatics.jp/), which contained 6,956 human annotated genes.
Performance evaluation
To evaluate the accuracy of POC-19, true positive (TP), true negative (TN), false positive (FP) and false negative (FN) numbers were counted. Then, we calculated six measurements, including sensitivity (Sn), specificity (Sp), accuracy (Ac), positive predictive value (PPV), negative predictive value (NPV), Mathew correlation coefficient (MCC) as below:
The 5-fold cross-validation was performed on Cohort 1, and Sn, Sp, Ac, PPV, NPV and MCC values were calculated (Table S7). The ROC curve was illustrated based on Sn and 1-Sp scores. For each AUC value, the 95% CI was computed with 1000 stratified bootstrap replicates. The PCA analysis was implemented in Scikit-learn 0.22.1 (https://scikit-learn.org/stable/), a powerful package for data mining and analysis.
The prioritization of biomarker combinations by POC-19
For the identification of different types of biomarker combinations, we first classified the proteomic datasets of FT1-FT4, ST1, ST2, MT1, MT2 and H into different groups. (i) For the classification of COVID-19 patients, FT1-FT4 of F patients, ST1 of S patients, MT1 of M patients and H were included. ST2 and MT2 were not included because these blood samples were collected from the patients shortly before discharged from the hospital and could not faithfully reflect the disease peak. For each group, its corresponding proteomic data were taken as positive data, where the remaining data were regarded as negative data. (ii) For the identification of the biomarker combination to predict S to F outcome, we took FT1-FT4 that reflect the clinical deterioration of F patients as positive data, while ST1 that reflect the disease peak of S patients was taken as negative data. (iii) To predict M to S outcome, we took ST1 and FT1 as positive data, and MT1 as negative data, respectively. The clinical characteristics of S patients at ST1 and F cases at FT1 were similar, which reflect the disease peak of S and/or more S patients. MT1 reflect the disease peak of M patients. (iv) To predict COVID-19 patients curable from the disease, we took MT2 as positive data, and MT1, ST1 and FT1 as negative data, respectively.
POC-19 is a three-step pipeline, including DPR, CBS, and FBD. In the step of DPR, we compared the proteomic data of FT1-FT4 to H, and reserved highly potential DEPs as a candidate reservoir (|log2(FC)| > 0.8, unpaired two-sided Welch’s t test, p < 0.01). To avoid over-fitting, the number of proteins in a combination should be much smaller than the sample size. Thus, CBS was implemented to select and optimize different sets of biomarker combinations with ≤ 5 proteins. From the candidate reservoir, we randomly select 5 proteins to form a potential combination, and the initial weight value of each protein was set to 1. For each type of biomarker combination identifications, 1000 candidate combinations were prepared, respectively.
In the last step, the 5-fold cross-validation was conducted for model training, parameter optimization, and performance evaluation. For each candidate combination, we randomly generated a training dataset and a testing dataset with a ratio of approximately 4:1. The testing dataset was only used to evaluate the performance but not for training. The least absolute shrinkage and selection operator (LASSO, L1 regularization) penalty and the ridge regression (L2 regularization) penalty in PLR (Ning et al., 2020), were iteratively used to optimize the weight values of the 5 proteins. To simplify the composition of a combination, one or multiple protein was randomly dropped if the 5-fold cross-validation AUC value was increased. Such a procedure was repeatedly performed until the AUC value was not increased any longer. Then, the AUC values of all the 1000 candidate combinations were determined, and the final combination was determined based on the highest AUC value. The PLR algorithm was implemented in Python 3.7 with Scikit-learn 0.22.1. The source code of POC-19 is available at: https://github.com/Ning-310/POC-19http://ictcf.biocuckoo.cn/HUST-19.php. To test the statistical significance of each of proteins prioritized by POC-19, one-way ANOVA was conducted using the f_oneway function in scipy.stats (p < 0.05). Then, a permutation-based approach, Westfall and Young’s step-down min P correction for multiple hypothesis testing (Oughtred et al., 2019), was adopted to calculate permutation adjusted p values (times = 100,000), using the minP function in R package multtest (Oughtred et al., 2019).
Re-construction of a COVID-19-associated plasma protein network
Based on the functional annotations in UniProt, we classified the 195 DEPs into 9 classes, including platelet/neutrophil degranulation, complement activation, immune cell migration, metabolic process, blood coagulation, other immune response, phagocytosis/endocytosis, transport/cell adhesion, and cell cycle/proliferation. Human known protein-protein interactions (PPIs) were integrated from 7 public databases, including BioGrid (Oughtred et al., 2019), IID (Kotlyar et al., 2019), InBio Map™ (Li et al., 2017), Mentha (Calderone et al., 2013), HINT (Das and Yu, 2012), iRefIndex (Razick et al., 2008) and PINA (Cowley et al., 2012). In total, we collected 1,771,193 PPIs of 18,839 human proteins from these databases. For the 195 DEPs, we extracted 1113 PPIs for 173 unique proteins, and the plasma protein network modulated by COVID-19 was Constructed and visualized with Cytoscape 3.6.1 software package (Shannon et al., 2003).
Data and Code Availability
The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org) via the iProX partner repository (Ma et al., 2019) with the dataset identifier PXD019106 (https://www.iprox.org//page/project.html?id=IPX0002173000).
Acknowledgments
We thank the patients and the nurses and clinical staff who are providing care for these patients. We also thank many staff members at Wuhan Jinyintan Hospital and SpecAlly Life Technology Co., Ltd. for their contributions and assistance in this study and Dr. Zhimei Ma for technical support. We thank Mr. An Wang and other members of Zhou group for their assistance. We sincerely pay tribute to our colleagues who have strived in the forefront of treating COVID-19 patients and are studying this coronavirus in Wuhan and around the world. This work was supported by the Strategic Priority Research Program of CAS (XDB29010300 to X. Zhou.), the National Science and Technology Major Project (2020ZX09201-001 to D.-Y.Z. and 2018ZX10101004 to X. Zhou.), National Natural Science Foundation of China (81873964 to Y.Q.; 81971818 and 81772047 to Y.S.; 31930021, 31970633, and 34671360 to Y.X.; 82002155 to T.S.; 32000131 to D.W.; and 31670161 to X. Zhou.), grant from the CAS Youth Innovation Promotion Association (2020332 to Y.Q.), the program for HUST Academic Frontier Youth Team (Y.X.), and grant from Clinical Research Center for Anesthesiology of Hubei Province (2019ACA167).
Author Contributions
T.S., D.W., J.X., and Q.H. performed experiments with the help of M.H., X. Zou., Q.Y., Y.Y., Y.B., S.P., J.M., Y.H., X.Y., H.Z., R.L., Y.R., X.C., and S.Y.; W.N., Y.X., and Y.Q. analyzed the proteomics data with the help of T.S. and D.W.; T.S., W.N., D.W., Q.H., Y.Q., D.-Y.Z., Y.X., and X. Zhou. performed the experimental design and data interpretation; X. Zhou., Y.Q., Y.X., D.-Y.Z., and Y.S. analyzed the data and wrote the paper; and X. Zhou., Y.Q., Y.S., and D.-Y.Z. designed and supervised the overall study.
Declaration of Interests
Wuhan Institute of Virology and Wuhan Jinyintan Hospital on behalf of the authors X. Zhou., Y.S., D.-Y.Z., Y.X., Y.Q., T.S., D.W., and M.H. have filed three Chinese patent applications (202010478392.8, 202010476095.X, and 202010476805.9) related to the biomarkers for predicting the different outcomes of COVID-19 patients.
Published: October 20, 2020
Footnotes
Supplemental Information can be found online at https://doi.org/10.1016/j.immuni.2020.10.008.
Supplemental Information
References
- Banères-Roquet F., Gualtieri M., Villain-Guillot P., Pugnière M., Leonetti J.P. Use of a surface plasmon resonance method to investigate antibiotic and plasma protein interactions. Antimicrob. Agents Chemother. 2009;53:1528–1531. doi: 10.1128/AAC.00971-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Batth T.S., Francavilla C., Olsen J.V. Off-line high-pH reversed-phase fractionation for in-depth phosphoproteomics. J. Proteome Res. 2014;13:6176–6186. doi: 10.1021/pr500893m. [DOI] [PubMed] [Google Scholar]
- Bengtsson A.A., Sturfelt G., Lood C., Rönnblom L., van Vollenhoven R.F., Axelsson B., Sparre B., Tuvesson H., Ohman M.W., Leanderson T. Pharmacokinetics, tolerability, and preliminary efficacy of paquinimod (ABR-215757), a new quinoline-3-carboxamide derivative: studies in lupus-prone mice and a multicenter, randomized, double-blind, placebo-controlled, repeat-dose, dose-ranging study in patients with systemic lupus erythematosus. Arthritis Rheum. 2012;64:1579–1588. doi: 10.1002/art.33493. [DOI] [PubMed] [Google Scholar]
- Björk P., Björk A., Vogl T., Stenström M., Liberg D., Olsson A., Roth J., Ivars F., Leanderson T. Identification of human S100A9 as a novel target for treatment of autoimmune disease via binding to quinoline-3-carboxamides. PLoS Biol. 2009;7:e97. doi: 10.1371/journal.pbio.1000097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Calderone A., Castagnoli L., Cesareni G. mentha: a resource for browsing integrated protein-interaction networks. Nat. Methods. 2013;10:690–691. doi: 10.1038/nmeth.2561. [DOI] [PubMed] [Google Scholar]
- Ceciliani F., Lecchi C. The immune functions of α1 acid glycoprotein. Curr. Protein Pept. Sci. 2019;20:505–524. doi: 10.2174/1389203720666190405101138. [DOI] [PubMed] [Google Scholar]
- Chen N., Zhou M., Dong X., Qu J., Gong F., Han Y., Qiu Y., Wang J., Liu Y., Wei Y. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet. 2020;395:507–513. doi: 10.1016/S0140-6736(20)30211-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choi J.W., Liu H., Mukherjee R., Yun J.W. Downregulation of fetuin-B and zinc-α2-glycoprotein is linked to impaired fatty acid metabolism in liver cells. Cell. Physiol. Biochem. 2012;30:295–306. doi: 10.1159/000339065. [DOI] [PubMed] [Google Scholar]
- Cichy J., Potempa J., Chawla R.K., Travis J. Stimulatory effect of inflammatory cytokines on alpha 1-antichymotrypsin expression in human lung-derived epithelial cells. J. Clin. Invest. 1995;95:2729–2733. doi: 10.1172/JCI117975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- The Gene Ontology Consortium The Gene Ontology resource: 20 years and still GOing strong. Nucleic Acids Res. 2019;47(D1):D330–D338. doi: 10.1093/nar/gky1055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- UniProt Consortium UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res. 2019;47(D1):D506–D515. doi: 10.1093/nar/gky1049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cowley M.J., Pinese M., Kassahn K.S., Waddell N., Pearson J.V., Grimmond S.M., Biankin A.V., Hautaniemi S., Wu J. PINA v2.0: mining interactome modules. Nucleic Acids Res. 2012;40:D862–D865. doi: 10.1093/nar/gkr967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Das J., Yu H. HINT: high-quality protein interactomes and their applications in understanding human disease. BMC Syst. Biol. 2012;6:92. doi: 10.1186/1752-0509-6-92. [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Felice F.G., Tovar-Moll F., Moll J., Munoz D.P., Ferreira S.T. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and the central nervous system. Trends Neurosci. 2020;43:355–357. doi: 10.1016/j.tins.2020.04.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deady L.E., Todd E.M., Davis C.G., Zhou J.Y., Topcagic N., Edelson B.T., Ferkol T.W., Cooper M.A., Muenzer J.T., Morley S.C. L-plastin is essential for alveolar macrophage production and control of pulmonary pneumococcal infection. Infect. Immun. 2014;82:1982–1993. doi: 10.1128/IAI.01199-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fournier T., Medjoubi-N N., Porquet D. Alpha-1-acid glycoprotein. Biochim. Biophys. Acta. 2000;1482:157–171. doi: 10.1016/s0167-4838(00)00153-9. [DOI] [PubMed] [Google Scholar]
- Gabay C., Kushner I. Acute-phase proteins and other systemic responses to inflammation. N. Engl. J. Med. 1999;340:448–454. doi: 10.1056/NEJM199902113400607. [DOI] [PubMed] [Google Scholar]
- Guan W.J., Ni Z.Y., Hu Y., Liang W.H., Ou C.Q., He J.X., Liu L., Shan H., Lei C.L., Hui D.S.C., China Medical Treatment Expert Group for Covid-19 Clinical characteristics of coronavirus disease 2019 in China. N. Engl. J. Med. 2020;382:1708–1720. doi: 10.1056/NEJMoa2002032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hassan M.I., Waheed A., Yadav S., Singh T.P., Ahmad F. Zinc α 2-glycoprotein: a multidisciplinary protein. Mol. Cancer Res. 2008;6:892–906. doi: 10.1158/1541-7786.MCR-07-2195. [DOI] [PubMed] [Google Scholar]
- Huang C., Wang Y., Li X., Ren L., Zhao J., Hu Y., Zhang L., Fan G., Xu J., Gu X. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet. 2020;395:497–506. doi: 10.1016/S0140-6736(20)30183-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jose R.J., Manuel A. COVID-19 cytokine storm: the interplay between inflammation and coagulation. Lancet Respir. Med. 2020;8:e46–e47. doi: 10.1016/S2213-2600(20)30216-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kanehisa M., Furumichi M., Tanabe M., Sato Y., Morishima K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 2017;45(D1):D353–D361. doi: 10.1093/nar/gkw1092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karmilin K., Schmitz C., Kuske M., Körschgen H., Olf M., Meyer K., Hildebrand A., Felten M., Fridrich S., Yiallouros I. Mammalian plasma fetuin-B is a selective inhibitor of ovastacin and meprin metalloproteinases. Sci. Rep. 2019;9:546. doi: 10.1038/s41598-018-37024-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kotlyar M., Pastrello C., Malik Z., Jurisica I. IID 2018 update: context-specific physical protein-protein interactions in human, model organisms and domesticated species. Nucleic Acids Res. 2019;47(D1):D581–D589. doi: 10.1093/nar/gky1037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lachmann P.J. The story of complement factor I. Immunobiology. 2019;224:511–517. doi: 10.1016/j.imbio.2019.05.003. [DOI] [PubMed] [Google Scholar]
- Lee K.J., Carlin J.B. Multiple imputation for missing data: fully conditional specification versus multivariate normal imputation. Am. J. Epidemiol. 2010;171:624–632. doi: 10.1093/aje/kwp425. [DOI] [PubMed] [Google Scholar]
- Li T., Wernersson R., Hansen R.B., Horn H., Mercer J., Slodkowicz G., Workman C.T., Rigina O., Rapacki K., Stærfeldt H.H. A scored human protein-protein interaction network to catalyze genomic interpretation. Nat. Methods. 2017;14:61–64. doi: 10.1038/nmeth.4083. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu M., Zhu H., Dai Y., Pan H., Li N., Wang L., Yang H., Yan K., Gong F. Zinc-α2-glycoprotein is associated with obesity in Chinese people and HFD-induced obese mice. Front. Physiol. 2018;9:62. doi: 10.3389/fphys.2018.00062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu R., Zhao X., Li J., Niu P., Yang B., Wu H., Wang W., Song H., Huang B., Zhu N. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet. 2020;395:565–574. doi: 10.1016/S0140-6736(20)30251-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luo Z., Lei H., Sun Y., Liu X., Su D.F. Orosomucoid, an acute response protein with multiple modulating activities. J. Physiol. Biochem. 2015;71:329–340. doi: 10.1007/s13105-015-0389-9. [DOI] [PubMed] [Google Scholar]
- Ma J., Chen T., Wu S., Yang C., Bai M., Shu K., Li K., Zhang G., Jin Z., He F. iProX: an integrated proteome resource. Nucleic Acids Res. 2019;47(D1):D1211–D1217. doi: 10.1093/nar/gky869. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martinelli A.E.M., Maranhão R.C., Carvalho P.O., Freitas F.R., Silva B.M.O., Curiati M.N.C., Kalil Filho R., Pereira-Barretto A.C. Cholesteryl ester transfer protein (CETP), HDL capacity of receiving cholesterol and status of inflammatory cytokines in patients with severe heart failure. Lipids Health Dis. 2018;17:242. doi: 10.1186/s12944-018-0888-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meex R.C., Hoy A.J., Morris A., Brown R.D., Lo J.C., Burke M., Goode R.J., Kingwell B.A., Kraakman M.J., Febbraio M.A. Fetuin B is a secreted hepatocyte factor linking steatosis to impaired glucose metabolism. Cell Metab. 2015;22:1078–1089. doi: 10.1016/j.cmet.2015.09.023. [DOI] [PubMed] [Google Scholar]
- Miao M., Yu F., Wang D., Tong Y., Yang L., Xu J., Qiu Y., Zhou X., Zhao X. Proteomics profiling of host cell response via protein expression and phosphorylation upon dengue virus infection. Virol. Sin. 2019;34:549–562. doi: 10.1007/s12250-019-00131-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moore J.B., June C.H. Cytokine release syndrome in severe COVID-19. Science. 2020;368:473–474. doi: 10.1126/science.abb8925. [DOI] [PubMed] [Google Scholar]
- National Health Commission . 2020. Protocol on prevention and control of COVID-19 (edition 6)https://www.chinadaily.com.cn/pdf/2020/2.COVID-19.Prevention.and.Control.Protocol V6.pdf. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ning W., Jiang P., Guo Y., Wang C., Tan X., Zhang W., Peng D., Xue Y. GPS-Palm: a deep learning-based graphic presentation system for the prediction of S-palmitoylation sites in proteins. Brief. Bioinform. 2020 doi: 10.1093/bib/bbaa038. Published online April 3, 2020. [DOI] [PubMed] [Google Scholar]
- Oughtred R., Stark C., Breitkreutz B.J., Rust J., Boucher L., Chang C., Kolas N., O’Donnell L., Leung G., McAdam R. The BioGRID interaction database: 2019 update. Nucleic Acids Res. 2019;47(D1):D529–D541. doi: 10.1093/nar/gky1079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ranucci M., Ballotta A., Di Dedda U., Bayshnikova E., Dei Poli M., Resta M., Falco M., Albano G., Menicanti L. The procoagulant pattern of patients with COVID-19 acute respiratory distress syndrome. J. Thromb. Haemost. 2020;18:1747–1751. doi: 10.1111/jth.14854. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rappsilber J., Mann M., Ishihama Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2007;2:1896–1906. doi: 10.1038/nprot.2007.261. [DOI] [PubMed] [Google Scholar]
- Razick S., Magklaras G., Donaldson I.M. iRefIndex: a consolidated protein interaction database with provenance. BMC Bioinformatics. 2008;9:405. doi: 10.1186/1471-2105-9-405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Regn M., Laggerbauer B., Jentzsch C., Ramanujam D., Ahles A., Sichler S., Calzada-Wack J., Koenen R.R., Braun A., Nieswandt B., Engelhardt S. Peptidase inhibitor 16 is a membrane-tethered regulator of chemerin processing in the myocardium. J. Mol. Cell. Cardiol. 2016;99:57–64. doi: 10.1016/j.yjmcc.2016.08.010. [DOI] [PubMed] [Google Scholar]
- Shannon P., Markiel A., Ozier O., Baliga N.S., Wang J.T., Ramage D., Amin N., Schwikowski B., Ideker T. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13:2498–2504. doi: 10.1101/gr.1239303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- The Novel Coronavirus Pneumonia Emergency Response Epidemiology Team Vital surveillances: the epidemiological characteristics of an outbreak of 2019 novel coronavirus diseases (COVID-19) — China, 2020. China CDC Weekly. 2020;2:113–122. [PMC free article] [PubMed] [Google Scholar]
- Todd E.M., Zhou J.Y., Szasz T.P., Deady L.E., D’Angelo J.A., Cheung M.D., Kim A.H., Morley S.C. Alveolar macrophage development in mice requires L-plastin for cellular localization in alveoli. Blood. 2016;128:2785–2796. doi: 10.1182/blood-2016-03-705962. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tyagi E., Fiorelli T., Norden M., Padmanabhan J. Alpha 1-antichymotrypsin, an inflammatory protein overexpressed in the brains of patients with Alzheimer’s disease, induces Tau hyperphosphorylation through c-Jun N-terminal kinase activation. Int. J. Alzheimers Dis. 2013;2013:606083. doi: 10.1155/2013/606083. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tyanova S., Temu T., Cox J. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat. Protoc. 2016;11:2301–2319. doi: 10.1038/nprot.2016.136. [DOI] [PubMed] [Google Scholar]
- Varga Z., Flammer A.J., Steiger P., Haberecker M., Andermatt R., Zinkernagel A.S., Mehra M.R., Schuepbach R.A., Ruschitzka F., Moch H. Endothelial cell infection and endotheliitis in COVID-19. Lancet. 2020;395:1417–1418. doi: 10.1016/S0140-6736(20)30937-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Venancio T.M., Machado R.M., Castoldi A., Amano M.T., Nunes V.S., Quintao E.C., Camara N.O., Soriano F.G., Cazita P.M. CETP lowers TLR4 expression which attenuates the inflammatory response induced by LPS and polymicrobial sepsis. Mediators Inflamm. 2016;2016:1784014. doi: 10.1155/2016/1784014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang D., Hu B., Hu C., Zhu F., Liu X., Zhang J., Wang B., Xiang H., Cheng Z., Xiong Y. Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus-infected pneumonia in Wuhan, China. JAMA. 2020;323:1061–1069. doi: 10.1001/jama.2020.1585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang S., Song R., Wang Z., Jing Z., Wang S., Ma J. S100A8/A9 in inflammation. Front. Immunol. 2018;9:1298. doi: 10.3389/fimmu.2018.01298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Y., Zhang D., Du G., Du R., Zhao J., Jin Y., Fu S., Gao L., Cheng Z., Lu Q. Remdesivir in adults with severe COVID-19: a randomised, double-blind, placebo-controlled, multicentre trial. Lancet. 2020;395:1569–1578. doi: 10.1016/S0140-6736(20)31022-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- WHO . 2020. Coronavirus disease 2019 (COVID-19): situation report - 128.https://www.who.int/docs/default-source/coronaviruse/situation-reports/20200527-covid-19-sitrep-128.pdf?sfvrsn=11720c0a_2 [Google Scholar]
- Wu D., Shu T., Yang X., Song J.-X., Zhang M., Yao C., Liu W., Huang M., Yu Y., Yang Q. Plasma metabolomic and lipidomic alterations associated with COVID-19. Natl. Sci. Rev. 2020;7:1157–1168. doi: 10.1093/nsr/nwaa086. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu F., Zhao S., Yu B., Chen Y.M., Wang W., Song Z.G., Hu Y., Tao Z.W., Tian J.H., Pei Y.Y. A new coronavirus associated with human respiratory disease in China. Nature. 2020;579:265–269. doi: 10.1038/s41586-020-2008-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu J., Yang X., Yang L., Zou X., Wang Y., Wu Y., Zhou T., Yuan Y., Qi H., Fu S. Clinical course and predictors of 60-day mortality in 239 critically ill patients with COVID-19: a multicenter retrospective study from Wuhan, China. Crit. Care. 2020;24:394. doi: 10.1186/s13054-020-03098-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu Z., Shi L., Wang Y., Zhang J., Huang L., Zhang C., Liu S., Zhao P., Liu H., Zhu L. Pathological findings of COVID-19 associated with acute respiratory distress syndrome. Lancet Respir. Med. 2020;8:420–422. doi: 10.1016/S2213-2600(20)30076-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang X., Yang Q., Wang Y., Wu Y., Xu J., Yu Y., Shang Y. Thrombocytopenia and its association with mortality in patients with COVID-19. J. Thromb. Haemost. 2020;18:1469–1472. doi: 10.1111/jth.14848. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang X., Yu Y., Xu J., Shu H., Xia J., Liu H., Wu Y., Zhang L., Yu Z., Fang M. Clinical course and outcomes of critically ill patients with SARS-CoV-2 pneumonia in Wuhan, China: a single-centered, retrospective, observational study. Lancet Respir. Med. 2020;8:475–481. doi: 10.1016/S2213-2600(20)30079-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yaprak E., Kasap M., Akpinar G., Islek E.E., Sinanoglu A. Abundant proteins in platelet-rich fibrin and their potential contribution to wound healing: an explorative proteomics study and review of the literature. J. Dent. Sci. 2018;13:386–395. doi: 10.1016/j.jds.2018.08.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang C., Shi L., Wang F.S. Liver injury in COVID-19: management and challenges. Lancet Gastroenterol. Hepatol. 2020;5:428–430. doi: 10.1016/S2468-1253(20)30057-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Y., Wang Y., Feng Y., Tu Z., Lou Z., Tu C. Proteomic profiling of purified rabies virus particles. Virol. Sin. 2020;35:143–155. doi: 10.1007/s12250-019-00157-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou F., Yu T., Du R., Fan G., Liu Y., Liu Z., Xiang J., Wang Y., Song B., Gu X. Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study. Lancet. 2020;395:1054–1062. doi: 10.1016/S0140-6736(20)30566-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou P., Yang X.L., Wang X.G., Hu B., Zhang L., Zhang W., Si H.R., Zhu Y., Li B., Huang C.L. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020;579:270–273. doi: 10.1038/s41586-020-2012-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu N., Zhang D., Wang W., Li X., Yang B., Song J., Zhao X., Huang B., Shi W., Lu R., China Novel Coronavirus Investigating and Research Team A novel coronavirus from patients with pneumonia in China, 2019. N. Engl. J. Med. 2020;382:727–733. doi: 10.1056/NEJMoa2001017. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org) via the iProX partner repository (Ma et al., 2019) with the dataset identifier PXD019106 (https://www.iprox.org//page/project.html?id=IPX0002173000).