Abstract
Alphaviruses cause severe human illnesses including persistent arthritis and fatal encephalitis. As alphavirus entry into target cells is the first step in infection, intensive research efforts have focused on elucidating aspects of this pathway, including attachment, internalization, and fusion. Herein, we review recent developments in the molecular understanding of alphavirus entry both in vitro and in vivo and how these advances might enable the design of therapeutics targeting this critical step in the alphavirus life cycle.
Introduction
Alphaviruses are enveloped, positive-sense, single-stranded RNA viruses in the Togaviridae family that are transmitted by arthropods and are responsible for emerging and reemerging diseases in humans. Some alphaviruses (e.g., Chikungunya (CHIKV), Ross River (RRV), Mayaro (MAYV), Semliki Forest (SFV), Sindbis (SINV), and O'nyong-nyong (ONNV)) cause acute inflammatory musculoskeletal and joint-associated syndromes, which can become chronic [1], whereas others (Eastern (EEEV), Western (WEEV), and Venezuelan (VEEV) equine encephalitis viruses) cause infection in the brain and neurological disease (Table 1). Although pathogenic alphaviruses are maintained in sylvatic transmission cycles in nature, their insect vectors and reservoir host species vary, which has implications for their geographic range and potential for causing outbreaks in humans.
Table 1. Diseases of pathogenic alphaviruses, mosquito vectors, and reservoir hosts.
| Medically relevant alphavirus | Antigenic clade | Confirmed or putative mosquito vector(s) species | Confirmed or putative reservoir host(s) | Disease manifestations |
|---|---|---|---|---|
| CHIKV | SFV | Aedes albopictus [128], Aedes aegypti, Aedes (subgenus Stegomyia) | Nonhuman primates [1] | Fever, polyarthralgia (frequently becomes recurrent), myalgia, rash, and headache |
| ONNV | SFV | Anopheles funestus, Anopheles gambiae [129] | Unknown | Similar to CHIKV with the addition of cervical lymphadenitis |
| MAYV | SFV | Haemagogus janthinomys [130], A. aegypti [131] | Nonhuman primates [132] | Identical to CHIKV |
| RRV | SFV | Culex annulirostris, Aedes vigilax [133] | Marsupials [134] | Identical to CHIKV |
| SFV | SFV | Aedes spp. [135] | Small mammals, birds, nonhuman primates [135] | Mild febrile illness in humans; infrequent myalgia and polyarthralgia; encephalitis can be induced in mice |
| EEEV | EEEV | Culiseta melanura [136], Culex erraticus [137] | Passeriformes birds [136] | Similar to CHIKV if there is no CNS involvement; encephalitic disease includes headache, vomiting, diarrhea, seizures, and coma |
| SINV | WEEV | Culex spp. [138] | Wild birds [139] | Arthralgia, rash, malaise |
| VEEV | VEEV | Culex (Melanoconion) spp. [140] | Small mammals [132] | Similar to EEEV; infection has lower mortality rate than EEEV |
| WEEV | WEEV | Culex tarsalis [141] | Wild birds [141] | Mainly subclinical or nonspecific febrile illness; can progress to encephalitis in rare cases |
Listed are the alphaviruses most frequently associated with disease outbreaks in humans. The reservoir hosts for these viruses include nonhuman primates, rodents, birds, and marsupials. Humans and equines represent either accidental hosts or are involved in epizootic transmission cycles. Mosquitoes from the Aedes and Culex genera are the major vectors of pathogenic alphaviruses, and at least 30 different species have been implicated.
CHIKV, Chikungunya; CNS, central nervous system; EEEV, Eastern equine encephalitis virus; MAYV, Mayaro; ONNV, O'nyong-nyong; RRV, Ross River; SFV, Semliki Forest; SINV, Sindbis; VEEV, Venezuelan equine encephalitis virus; WEEV, Western equine encephalitis virus.
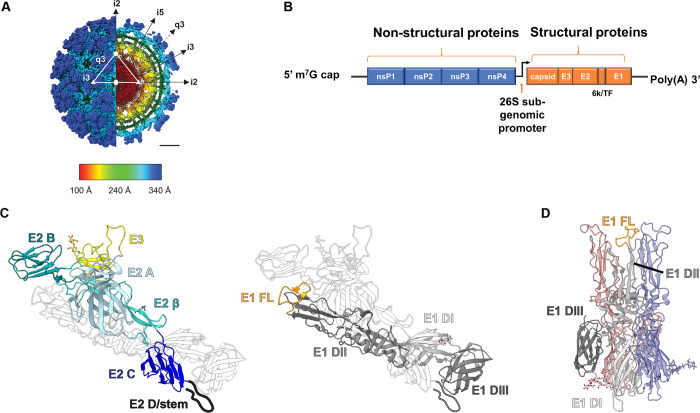
The alphavirus virion is approximately 70 nanometers in diameter and has T = 4 icosahedral symmetry (Fig 1A) [2,3]. The spherical virion is comprised of a single approximately 11.4 kb RNA genome encapsidated in a nucleocapsid core and surrounded by a host-derived lipid membrane. The genome encodes 4 nonstructural proteins, nsP1–4, which mediate viral translation, viral replication, and host subversion and evasion [4] and 6 structural proteins, capsid, E3, E2, 6K, transframe (TF), and E1 (Fig 1B). E1 and E2 are transmembrane proteins that interact to form a heterodimer (Fig 1C). Trimers of E1/E2 heterodimers assemble into higher order spikes (80 in total) on the virion surface. The alphavirus E2 protein facilitates receptor engagement [5], whereas E1 principally mediates membrane fusion after viral entry [5,6]. The carboxyl terminus of E2 also interacts with the capsid core, which stabilizes the virion [7,8]. The 6K protein is thought to promote glycoprotein maturation, spike assembly, and act as a viroporin [9]. The 6K gene produces 2 proteins, 6K and TF, the latter of which also contributes to virus particle assembly [9]. The TF product associates with E1/E2 and is detected on the virion surface, albeit at lower stoichiometric levels than other structural proteins [5]. TF also inhibits type I interferon (IFN) responses in cultured cells and in vivo through a mechanism dependent upon palmitoylation of the protein [10].
Fig 1. Alphavirus genome organization and molecular structure of the virion.
(A) Cryo-EM reconstruction of CHIKV VLP (EMDB: 9393) colored by radial distance and depicted from the surface (left half) and an equatorial cross section (right half). The white triangle indicates 1 icosahedral asymmetric unit, with the 5-fold (i5), 3-fold (i3), and 2-fold (i2) icosahedral axes of symmetry labeled with a pentagon, triangles, and an oval, respectively. Trimeric spikes are labeled “i3” if coincident with the i3 axes and “q3” if on a quasi-3-fold axis. The black arrows indicate the directions of the icosahedral symmetry axes. Radial distance color scheme: red, electron dense core and RNA; yellow, capsid; green, membrane lipid; cyan, E1; and dark blue, E2 spike. Scale bar: 100 A°. E1 and E2 are embedded in the viral membrane and assemble into a heterotrimer: E1 is responsible for membrane fusion, while E2 facilitates receptor engagement. E3 is also bound in some alphaviruses including CHIKV, SFV, and VEEV, but the significance of this is not fully understood. (B) The alphavirus genome is a single-stranded, plus-sense RNA molecule of approximately 11 kb and encodes 4 nonstructural proteins, nsP1–4 and 5 structural proteins, capsid, E3, E2, 6k/TF, and E1. NsP4 forms the primary RNA-dependent RNA polymerase, but the synthesis of the genome requires all 4 nonstructural proteins. The RNA is capped at the 5′ end and polyadenylated at the 3′ end. (C) The alphavirus structural proteins E2 and E3 are produced as a polyprotein termed p62 (left). P62 acts as a chaperone to ensure proper folding of E1 (right) in the ER and is proteolytically processed into the mature E2 and E3 proteins by host furin-like proteases. E3 binds E2 during transport of the protein complex to the cell surface and also remains bound in the mature virion for some alphaviruses. Structural image generated with Chimera software using structural data reported in [11] (PDB:3N40). Both glycoproteins are colored by domain. E3, yellow. E2: domain A, light blue; β-ribbon connector, light cyan; domain B, dark cyan; domain C, dark blue; cartoon of subdomain D/stem region, black. E1: DI, light gray; DII, medium gray; DIII, dark gray; FL, orange. (D) Upon exposure to low pH, E2 dissociates from E1, which drives E1 homotrimer formation (structural representation pictured). The FLs are exposed, which insert into the target membrane and facilitate membrane fusion in the early endosome. Image generated with Chimera software and structural data reported in [34] (PDB:1RER). One of the 3 E1 monomers is colored by domain. CHIKV, Chikungunya; cryo-EM, cryo-electron microscopy; ER, endoplasmic reticulum; FL, fusion loop; SFV, Semliki Forest; TF, transframe; VEEV, Venezuelan equine encephalitis virus; VLP, virus-like particle.
Over the past several decades, many groups have investigated the steps of alphavirus cellular entry given its implications for tropism. In this review, we summarize our understanding of alphavirus attachment and entry, as revealed by recent structural, biochemical, and genetic studies. We discuss how recent advances could be harnessed for possible therapeutic intervention.
Alphavirus structural proteins and their involvement in entry
E1/E2 structure and function
The E2 glycoprotein is translated in the infected cell in conjunction with E3 as a polyprotein termed p62, also called PE2. P62 co-translationally associates with E1 (Fig 1C) within the endoplasmic reticulum (ER), an interaction that is required for proper folding of E1 [11]. Subsequently, p62 is processed into the mature E2 and E3 proteins by furin-like proteases in the trans-Golgi network [12]. After furin cleavage, E3 remains associated with E2 at acidic pH to stabilize the heterodimer and prevent premature fusion within secretory vesicles [13–16]. For most alphaviruses, E3 dissociates from the virion in the neutral pH environment of the extracellular space. This coordinated binding and dissociation of E3 ensures the generation of a fusion-competent, infectious particle. Indeed, when the furin cleavage site of p62 is mutagenized, the resultant virion is less infectious and requires a lower pH to initiate fusion [17]. Moreover, structural analysis of immature CHIKV virus-like particles containing mutations in the furin cleavage site showed that E3 stabilizes domain B of E2 and prevents exposure of the fusion peptide on E1 [18]. However, for some alphaviruses (e.g., VEEV, SFV, and CHIKV), E3 may not fully dissociate from E2, which may depend in part on the pH of the medium in which the virus is produced [19–21]. Although the functional significance of retained E3 on the virion remains uncertain, it could impact receptor binding.
E2 is comprised of 3 principal ectodomains, A, B, and C [22,23]. A subdomain D within E2 also was identified in the VEEV crystal structure [19], seen in SINV [23], and contains key residues for SFV budding [24]. This subdomain has also been referred to as the E2 stem region [11]. Domain B is positioned furthest from the lipid bilayer, domain C is membrane proximal, and domain A is located between domains B and C [22]. E2 also contains a β-ribbon motif that connects domains A and B [11,22]. E1 is a class II fusion protein [25] that has 3 ectodomains, DI, DII, and DIII [26]. The hydrophobic fusion loop (FL) is located in DII [26]. E1 also contains a stem region that connects DIII to the transmembrane domain of the protein [11,26,27]. DIII adopts an immunoglobulin-like fold and is connected to DI through a linker region of approximately 28 amino acids [28].
Upon exposure to low pH in solution or in endosomes [6], E1 dissociates from E2 [29,30], which exposes the hydrophobic FL (Fig 1D). Subsequently, E1 forms a homotrimer, which triggers membrane fusion and enables nucleocapsid penetration into the cytosol. A computational study predicted that highly conserved histidine residues across 13 different alphavirus species located at the E1/E2 interface mediate the dissociation of E2 from E1 [31]. This model is consistent with structural [23] and biochemical analyses suggesting that conserved histidine residues stabilize E1/E2 interactions [32,33]. A recently reported 3.5 Å resolution cryo-electron microscopy (cryo-EM) structure of SINV corroborated observations from prior analyses [11,22] and provided new insights into the features governing the dissociation of E2 from E1. In addition to the role of conserved histidine residues, this study identified a novel hydrophobic pocket formed by E2 subdomain D and the E2 and E1 transmembrane helices [23]. Decreases in pH might disrupt this hydrophobic pocket, which, along with changes in hydrogen bonding between the conserved histidine residues, could destabilize E1/E2 interactions and promote E1 homotrimerization and membrane fusion [23].
The E1 DI and DII subdomains fold into a hairpin-like structure [11] following trimerization. Domain DIII packs against DI and DII [34] and participates in a “fold-back” mechanism that brings the viral envelope and target membranes in proximity [35]. E1-mediated membrane fusion is dynamic, with several intermediates described [22,36,37]. E1 initially engages the target membrane as 3 individual monomers, while E2 is still complexed as a trimer [36]. Using truncated forms of SFV E1 in vitro, a stable E1 trimer was shown to consist of only DI and DII at low pH (pH 5.7) in the absence of DIII or hairpin formation [36]. Thus, E1 trimerization likely occurs prior to the fold-back of DIII. Accordingly, exogenously expressed DIII can inhibit membrane fusion by acting as a dominant negative [38].
Alphavirus attachment
For alphaviruses to initiate infection, they must attach to target cells and engage an entry receptor. Over the years, alphavirus interactions with several ubiquitously expressed cell surface molecules, termed attachment factors, have been described [39]. We distinguish between an attachment factor, which allows the virus to make initial contact with the target cell and entry receptors, which facilitate internalization of the virus prior to endosomal fusion [40]. Although a consensus definition is not established, we suggest a protein is a bona fide virus receptor if the following features are confirmed: (1) direct and specific binding interaction between virus and receptor; (2) the receptor directly mediates and/or facilitates internalization of the virus; (3) virus infection is blocked by antibodies against the receptor, by soluble receptor decoy molecules, or through mutagenesis of the virus receptor binding domain; and (4) susceptibility of a permissive cell type correlates with receptor expression level [41]. While many virus receptors are expressed on the cell surface [41,42], some viruses such as those in the Filoviridae family require engagement of an endosomal receptor (e.g., Niemann–Pick C1 (NPC1)) to successfully complete the viral entry process [43].
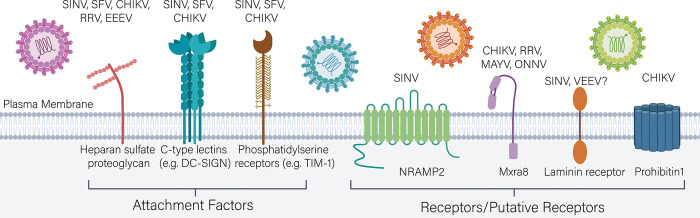
Attachment factors, in comparison, may display some but not all characteristics of receptors. However, virus binding and internalization generally are still observed in the absence of a given attachment factor, perhaps at a lower efficiency. Nevertheless, attachment factors can be important to viral pathogenesis, as they enhance target cell binding and decrease the amount of time the virus spends in the extracellular milieu, which, if prolonged, can lead to virus inactivation [44]. Below, we describe some of the best supported attachment factors and receptors for alphaviruses (Fig 2).
Fig 2. Alphavirus attachment factors entry receptors.
Alphaviruses utilize several cell surface molecules including HS, DC-SIGN/L-SIGN, and PS receptors to carry out an initial attachment to target cells. These molecules increase infectivity of multiple alphaviruses and may also enhance virus internalization. Alphavirus receptors that satisfy all criteria to describe a protein as a virus receptor include NRAMP2 and Mxra8. Other putative receptors requiring further corroboration include laminin receptor and PHB1. CHIKV, Chikungunya; DC-SIGN, dendritic cell-specific intercellular adhesion molecule-3-grabbing non-integrin; EEEV, Eastern equine encephalitis virus; HS, heparan sulfate; L-SIGN, liver-specific SIGN; MAYV, Mayaro; Mxra8, matrix remodeling associated protein 8; NRAMP2, natural resistance-associated macrophage protein 2; ONNV, O'nyong-nyong; PHB1, prohibitin1; PS, phosphatidylserine; RRV, Ross River; SFV, Semliki Forest; SINV, Sindbis; TIM-1, T cell immunoglobulin mucin domain 1.
Attachment factors
Heparan sulfate
Several alphaviruses use heparan sulfate (HS), a negatively charged glycosaminoglycan (GAG), as an attachment factor either naturally or as an adaptation after passage in culture. A curious feature of HS-mediated attachment is that it can increase alphavirus infectivity in cultured cells but either increase or decrease virulence in vivo depending on the virus and route of inoculation [45–48]. Initially, HS expression was shown to increase the infectivity of SINV [49,50] in a manner dependent on the extent of sulfation [48]. SFV also bound liposomes containing heparin after frequent passage and adaptation of the virus [51]. A more recent genome-wide, exon-trap screen in human HAP-1 cells identified the enzyme N-deacetylase and N-sulfotransferase 1 (NDST1), which facilitates N-sulfation of HS, as critical for infectivity of a pseudotyped virus bearing the E1 and E2 glycoproteins of a clinical CHIKV isolate [52]. The reduced infectivity of the pseudotyped virus was not rescued by the addition of chondroitin sulfate (CS), another cell surface GAG, which suggested that the phenotype was HS specific. A separate CRISPR/Cas9 screen in HAP-1 cells also identified GAG biosynthesis genes including B3GAT3, SLC35B2, and PAPSS1 as important factors for CHIKV infection [53].
The E2-82 residue in domain A of CHIKV E2 reportedly determines HS interactions. An arginine at this position (present in cell culture–adapted strains) results in enhanced HS interaction, and a glycine (present in clinical isolates) results in less binding to HS [54]. CHIKV strains containing the G82R mutation show reduced musculoskeletal disease and pathogenesis in mice [47]. The G82 residue also contributes to CHIKV persistence by allowing the virus to evade antibody neutralization and immune-mediated clearance [55].
The dependence of RRV on HS as an attachment factor was shown in the context of passage in avian cells [56] even though birds are not a reservoir species. Three mutations allowed RRV to replicate to higher titers [56]. Two of the variants had lysine substitutions at the surface-exposed residue E2-218 in domain B of E2 (N→K) [56], and a subsequent study showed that an arginine (N→R) at E2-218 also increased infectivity in an HS-dependent manner [57]. Cryo-EM experiments with the E2-N218R mutant of RRV revealed that HS binds the most distal portion of E2 [58,59]. In a separate study with an RRV E1 and E2 pseudotyped virus, a charge mutation to the adjacent residue E2-216 (T→R) also promoted the use of HS as an attachment factor [60]. As alluded to above, several HS-adapted arthritogenic alphaviruses show reduced virulence in vivo [48,54]. One explanation is that these virions become trapped by HS binding at the cell surface and bud less efficiently [60]. Alternatively, HS binding may target virus to cells that are inherently non- or less permissive and prevent productive spread [46].
Although the adaptation of HS binding for some alphaviruses leads to increased cell culture infectivity yet decreased pathogenicity in vivo, some natural alphavirus isolates use HS as a virulence factor in specific contexts. Non-passaged EEEV isolates use HS as an attachment factor [45]. Mutagenesis of 3 conserved lysine residues in domain A of E2 (E2-71, E2-74, and E2-77) abrogated EEEV binding to HS and decreased neurovirulence in mice [45]. Five naturally occurring EEEV strains with sequence variation at E2-71 and E2-72 showed differential dependence on HS as an attachment factor and distinct infectivity in mice [61]. EEEV strains that more readily bind HS are less pathogenic in mice after subcutaneous inoculation but display increased neurovirulence when delivered via intracranial injection [61]. Increased neurovirulence conferred by HS binding has also been reported with other neurotropic alphaviruses [62–64].
Initial cryo-EM analysis of HS analogs in complex with EEEV [61,65] suggested that the binding interface consists of the 3 key lysine residues at E2-71, E2-74, and E2-77, which are located within a surface-exposed β-strand and loop in E2 domain A [65]. However, a more recent, higher resolution 5.8 Å structure of EEEV in complex with HS showed that each spike contains 4 HS contact points, 3 along the quasi-3-fold axis of symmetry and 1 at the vertex of the spike [66]. While several basic amino acid residues were identified as part of the HS binding interface [66], the higher resolution structure did not identify the E2-71, E2-74, and E2-77 triad as directly interacting with HS. The basis for the disparity remains unclear, although the type of HS used might contribute to the differences [66].
Although the duration of viremia and magnitude of virus dissemination is reduced for HS-binding alphaviruses, the potentially deleterious effect on virus spread is counterbalanced by the enhanced neurovirulence phenotype conferred by adaptation to HS. HS-binding neurotropic alphavirus isolates that can bypass the requirement for sustained viremia to access the central nervous system may have a greater potential for virulence. Further studies are warranted to better understand the pathogenic mechanisms mediated by HS binding during neurotropic alphavirus infections including interactions at the blood–brain barrier [62]. More studies are also needed to distinguish HS adaptations that occur in cell culture from those present in natural isolates and how these sequence changes differentially impact alphavirus infection, tropism, immunity, and pathogenesis in mammalian and mosquito vector hosts.
C-type lectins
C-type lectins, including dendritic cell-specific intercellular adhesion molecule-3-grabbing non-integrin (DC-SIGN) and liver-specific SIGN (L-SIGN), can act as an attachment factors for some alphaviruses [67,68]. In addition to their role in cell migration [69], these proteins function as pattern recognition receptors (PRRs) by binding high-mannose N-glycans on the surface of pathogens [70]. Alphaviruses, along with viruses from other families [71], can exploit this interaction to gain access to permissive cells. Cells transfected with either DC-SIGN or L-SIGN showed increases in SINV binding and infectivity [67]. Alphavirus binding to these lectins was observed only with virus generated in mosquito cells, which produce high- and pauci-mannose N-linked glycans compared to the complex N-linked glycans of vertebrates [72] and in mammalian cells lacking glycosyltransferases or treated with alpha-mannosidase inhibitors. These experiments demonstrated that differential mannose processing on N-linked glycans affects alphavirus interactions with DC-SIGN or L-SIGN. Consistent with these results, lentivirus pseudotyped with SFV E1 and E2 glycoproteins display enhanced transduction efficiencies in C-type lectin-expressing cells when the virus is generated under conditions that maintain high-mannose N-glycosylation [68]. Thus, high-mannose glycosylation of alphavirus glycoproteins can influence the tropism of target cells early during infection in mammalian hosts, which may impact the outcome of disease [73].
In addition to acting as alphavirus attachment factors, C-type lectins also function as PRRs. However, it is unclear whether their downstream signaling pathways are activated upon alphavirus engagement, which could impact post-entry steps in the alphavirus replication cycle [74]. C-type lectin signaling pathway activation could have functional implications for DC and/or macrophage maturation, including effects on major histocompatibility complex (MHC) class I presentation, adaptive immune responses, and pathogenesis [70]. Indeed, particle size and structure can dictate how antigens are internalized by DCs [75]. Upon binding to DC-SIGN, smaller, polymeric structures are routed to endosomes, whereas larger structures characteristic of viruses localize to non-endosomal compartments [75]. It remains to be determined how engagement of DC-SIGN and other C-type lectins affects the subcellular trafficking and infectivity of alphaviruses. Despite their effects on infectivity in cell culture, studies in mice deficient in DC-SIGN or L-SIGN expression did not show an impact on CHIKV pathogenesis [76]. In comparison, expression of the C-type lectin dendritic cell immunoreceptor (DCIR) limited CHIKV disease, an effect which may require direct binding of CHIKV [76]. The physiological significance of interactions with DC-SIGN/L-SIGN and other C-type lectin receptors on disease progression and immunity with other arthritogenic or encephalitic alphaviruses is undetermined.
Phosphatidylserine receptors
Phosphatidylserine (PS) is a component of the eukaryotic plasma membrane that is found in the host-derived lipid bilayer of many enveloped viruses [77]. PS receptors are increasingly recognized as attachment factors and/or receptors for viruses [77]. The most well-characterized function of PS receptors is the binding of surface-exposed PS during apoptosis and subsequent signaling of dying cells to be marked for phagocytosis [78]. Accordingly, this has led the viral hijacking of PS receptors to be termed “apoptotic mimicry” [79]. The T cell immunoglobulin mucin (TIM) domain family proteins were the first PS-binding receptors proposed as attachment factors for SINV, CHIKV, RRV, and EEEV [80,81]. Infection of pseudotyped viruses displaying the alphavirus E1/E2 glycoproteins was increased in cells expressing TIM-1 and decreased in the presence of PS-containing membranes, suggesting that infectivity depends on TIM-1 binding of PS as opposed to a direct interaction with the virus glycoproteins [80]. Ectopic expression of TIM-1 also increased RRV uptake and infection in cells, and this phenotype was blocked by incubation with an anti-TIM-1 antibody [80]. A separate study using pseudotyped viruses expressing SINV envelope proteins [82] expanded the list of PS-binding proteins used by alphaviruses as attachment factors to include milk fat globule-epidermal growth factor-factor 8 (MFG-E8) and growth arrest-specific gene 6 (Gas6), 2 soluble adaptor molecules that engage PS [82]. Another PS receptor, CD300a, also increased the binding of pseudotyped virus displaying SINV E1/E2 to cells but did not enhance infection rates [82]. The effect of PS receptor engagement by alphaviruses on apoptosis signaling pathways has not been extensively investigated, which independently may impact immunity and pathogenesis.
Many questions remain about the interactions between attachment factors and receptors in alphavirus entry. Some groups have speculated that an exceptionally high affinity virus receptor could preclude a requirement for PS binding or other attachment factors [81,83]. Thus, alphaviruses with lower affinities for their viral receptor might preferentially use other attachment factors. For most alphavirus attachment factors, the full picture of how they function to enhance viral infection in vivo is incomplete.
Receptors
The identification of bona fide receptors for alphaviruses historically has been elusive. One impediment has been the lack of a discernable interaction between putative receptors and purified E2 proteins [84]. Moreover, for many proposed receptor molecules, infection has still been shown to occur in cells lacking the protein, suggesting that either receptor usage is highly cell-type–specific and/or that the proposed molecule is a subordinate receptor on most cells or acts more as an attachment factor to enhance infectivity. In the following section, we describe recent progress on alphavirus receptors including different proteins with their varying degrees of supportive data.
NRAMP
Natural resistance-associated macrophage protein (NRAMP) proteins are divalent metal ion transporters that have been proposed as receptors for SINV in both insect and mammalian cells [85]. Gene silencing of dNRAMP (the Drosophila gene) resulted in decreased infection for both cell culture-adapted and wild-type SINV strains in fruit flies [86]. As transfection of SINV RNA directly into Drosophila cells bypassed a requirement for dNRAMP, this protein was hypothesized to function during alphavirus entry. Indeed, direct virus binding to and interaction with dNRAMP was demonstrated by co-immunoprecipitation and confocal microscopy assays. The mammalian NRAMP2 protein is ubiquitously expressed on the cell surface of neuronal cells and macrophages [85]. SINV infection also was reduced in NRAMP2-deleted mouse embryonic fibroblasts (MEFs) and in several mammalian (MEF and U2OS) and insect (Drosophila DL1 and Aedes aegypti Ag-2) cell lines treated with exogenous iron, which downregulates dNRAMP/NRAMP2 protein expression. In contrast, infection of a chimeric alphavirus displaying the RRV envelope proteins was insensitive both to iron treatment and to NRAMP2 deletion. It is unclear whether other alphaviruses use NRAMP2 or other conserved membrane transporter proteins as receptors for infection. It also will be important to determine the role of NRAMP2 in SINV pathogenesis in vivo, as this has not yet been tested. Finally, alphavirus infection studies with blocking antibodies against NRAMP2 or receptor decoy molecules could provide further evidence for this protein as a bona vide receptor.
Mxra8
In a genome-wide CRISPR/Cas9 screen, matrix remodeling associated protein 8 (Mxra8) was identified as a receptor for several arthritogenic alphaviruses, including CHIKV, RRV, MAYV, and ONNV [87]. Mxra8 is expressed on the surface of epithelial, mesenchymal, and myeloid cells [87], all of which are targets of infection by arthritogenic alphaviruses [88]. Several lines of evidence support Mxra8 as an alphavirus receptor: (1) ectopic expression of Mxra8 enhances alphavirus infection; (2) transfection of viral RNA into cells bypasses a requirement for Mxra8 expression; (3) CHIKV binding to and infection of cells is blocked with antibodies against Mxra8 or a soluble Mxra8-Fc decoy protein; and (4) Mxra8 binds directly to CHIKV viruses or virus-like particles by enzyme-linked immunosorbent assay (ELISA) and surface plasmon resonance. Mxra8 also is required for arthritogenic alphavirus pathogenesis, as infection in vivo was inhibited with blocking Mxra8-Fc treatment, anti-Mxra8 antibodies, or in Mxra8-deficient mice [87,89]. However, low levels of viral infection occurred in cell culture and in vivo in the absence of Mxra8 expression, suggesting the existence of an unidentified subordinate receptor for this group of viruses. Outstanding questions remain on how Mra8 facilitates alphavirus internalization. Given that the cytoplasmic tail of Mxra8 is not required for receptor function [87], it is possible that Mxra8 engagement triggers interaction with an unidentified co-receptor that activates the endocytic pathway.
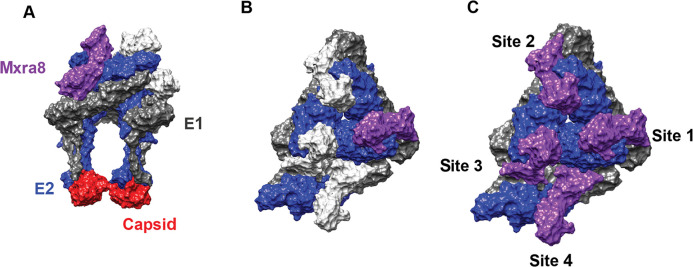
Structural analysis of Mxra8 in complex with either CHIKV virions [21] or E1/E2 glycoprotein complex [90] was recently described. Mxra8 ectodomain is comprised of 2 Ig-like domains arranged in an unusual head-to-head configuration [21,90]. A single monomer of Mxra8 can engage 3 different E1/E2 heterodimers on the virion surface [21,90]. Mxra8 wraps around the membrane-distal end of 1 E1/E2 dimer, makes intraspike contacts with a second heterodimer, and engages a third neighboring E1/E2 complex in an interspike interaction [21,90] (Fig 3A–3C). Mxra8 contacts residues in domains A and B of E2 and the FL and other sites in E1 DII. The binding occupancy of Mxra8 to infectious CHIKV particles is reduced by the presence of E3 on the virion [21]. Thus, retention of bound E3 even after cleavage and maturation could affect alphavirus binding and Mxra8 usage as a receptor. Future studies should determine whether the presence of E3 on some arthritogenic alphaviruses (e.g., SFV) explains their relatively weak dependence on Mxra8 for binding.
Fig 3. Mxra8 binding to CHIKV E1 and E2.
Side view (A) and top view (B and C) of Mxra8 bound to E1 and E2. Mxra8 (purple) wraps around 1 E1 (gray)-E2 (blue) heterodimer, contacts a second heterodimer within the same spike, and makes contacts with a third heterodimer on the adjacent spike. Capsid proteins are pictured in red. Mxra8 engages the CHIKV spike structure in a complex quaternary epitope. Image generated with Chimera software using [21] as a reference (PDB: 6NK6). CHIKV, Chikungunya virus; Mxra8, matrix remodeling associated protein 8.
Evolutionary and functional analyses established that most members of the Bovinae subfamily have a 15-amino-acid insertion in the Mxra8 ectodomain that blocks CHIKV binding. Introduction of this sequence into murine Mxra8 abolishes binding to virus particles and reduces CHIKV pathogenesis in vivo, whereas removal of the insertion in Bovinae Mxra8 enhances binding and infection [86]. This insertion likely evolved approximately 5 million years ago in the Miocene epoch [86], which could suggest that sequence acquisition was driven by positive selection against a primordial alphavirus. As alphaviruses are believed to have evolved much more recently (approximately 10,000 years ago) [91], this idea remains speculative.
Laminin receptor
Based on monoclonal antibody blocking studies, the laminin receptor was proposed as a possible receptor for SINV [92]. Laminin receptor is a cell surface-expressed protein that binds basement membrane laminin and has functions in both development and tumor metastasis [92]. SINV binds Chinese hamster ovary (CHO) cells overexpressing laminin receptor more readily than the parental control cell line, and the susceptibility of BHK cells increases with the amount of laminin receptor expressed on the cell surface [92]. Together with experiments showing inhibition of SINV infection by antibodies against laminin receptor, 3 of the criteria are met for establishing laminin receptor as a receptor. However, laminin receptor-mediated internalization of SINV (or any other alphavirus) or direct biochemical binding has yet to be documented. While a more recent study suggested that the carboxyl-terminal domain of laminin receptor interacts with VEEV E2 [93], an interaction with this encephalitic alphavirus has not been critically evaluated.
PHB1
Prohibitin1 (PHB1) is a protein that localizes to the plasma membrane and mitochondria and regulates cell proliferation and mitochondrial integrity [94]. One paper has described PHB1 as a receptor for CHIKV [95]. Using a proteomic approach, PHB1 was identified as a candidate binding partner for 2 CHIKV strains. Incubation of a microglia cell line with increasing concentrations of an anti-PHB1 antibody decreased the percentage of CHIKV-infected cells and viral yield. This observation was supported by gene silencing of PHB1, albeit this resulted in a relatively small reduction in CHIKV virus production. PHB1 also co-immunoprecipitated with CHIKV E2 and co-localized with CHIKV E2 at the plasma membrane by immunofluorescence microscopy. Although these data suggest that PHB1 might act as a receptor for CHIKV, no direct binding of CHIKV virions with soluble PHB1 has been demonstrated, and the effects of PHB1 on virus binding to and internalization in cells have not been reported. Of note, PHB1 reportedly also interacts with Dengue virus, a Flavivirus, to facilitate entry into insect cells [96].
Cellular uptake mechanisms of alphaviruses
Alphaviruses are internalized principally by clathrin-mediated endocytosis and delivered to the endosomal compartment where membrane fusion occurs, a process that has been reviewed extensively by others [39,97–99]. Live video tracking studies have shown that most CHIKV particles co-localize with clathrin prior to undergoing fusion [100]. A selective inhibitor of clathrin-mediated endocytosis, Pitstop, substantially reduced the number of CHIKV-infected cells, indicating a strong dependence on this entry pathway. CHIKV fusion primarily occurs in early endosomes as indicated by co-localization of virus particles with Rab5, a marker of this compartment [100]. Consistent with these data, a genome-wide RNA interference (RNAi) screen using SINV identified other host proteins important for clathrin-mediated endocytosis, including Fuzzy homologue (FUZ) and the tetraspanin membrane protein, TSPAN9 [101], which was specifically important for low-pH-triggered membrane fusion in the early endosome [101].
In some studies, alphaviruses are reported to internalize via alternate pathways including caveolae-dependent entry of MAYV into Vero cells [102], the direct delivery of SINV to target cells through a putative pore at the plasma membrane [103], and micropinocytosis-mediated uptake of CHIKV into muscle cells [104]. Considering the discovery of novel entry receptors for alphaviruses [85,87], and as yet unknown alternative receptor(s), it will be important to determine whether engagement by specific moieties on the surface facilitates distinct entry pathways of different alphaviruses in unique cell types.
Therapeutics targeting alphavirus cell entry
Currently, there are no Food and Drug Administration (FDA)-approved vaccines or antiviral drugs for pathogenic alphaviruses. Given that virus entry is the first step required to initiate a productive infection and can require highly specific interactions with receptors, it is an attractive target for the development of alphavirus antivirals.
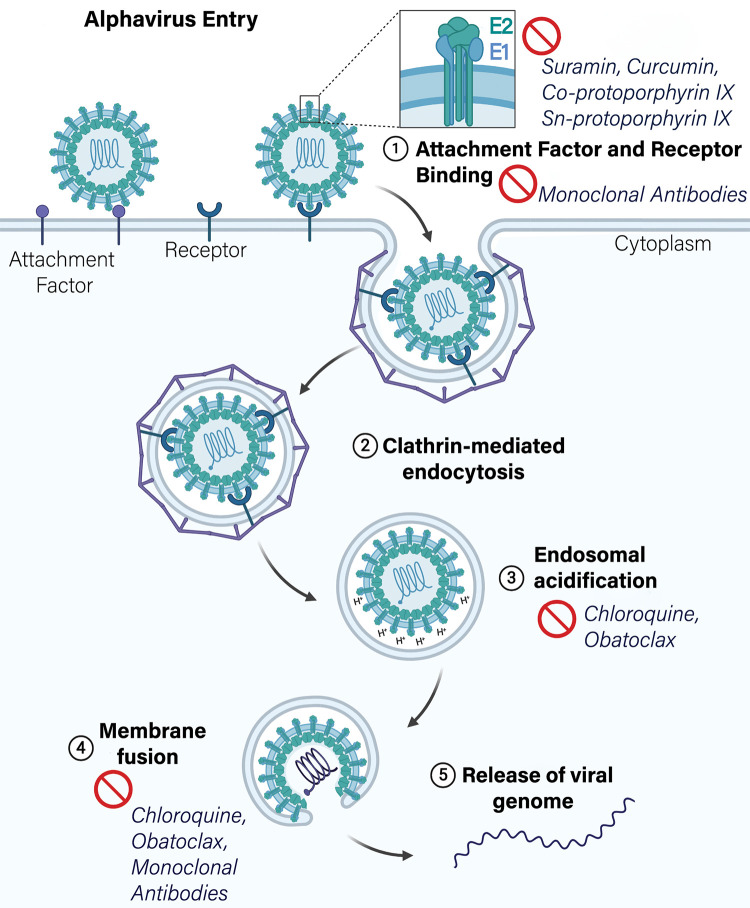
Candidate therapeutics that target alphavirus attachment and entry have recently been described (Fig 4). These include neutralizing monoclonal antibodies that block attachment, internalization, and pH-dependent fusion [105–107]. Monoclonal antibodies can be developed quickly during epidemics [107], have a rapid onset of protection [107], and facilitate viral clearance through multiple mechanisms including direct neutralization of virus [108] and indirect antibody-dependent effector functions including cell-mediated cytotoxicity [107,109], complement-dependent cytotoxicity, and phagocytosis [107]. A phase I/II clinical trial (NCT02230163) was initiated to determine the safety and efficacy of anti-CHIKV hyperimmune sera for treating neonatal infections resulting from vertical CHIKV transmission [110]. Another phase I trial was initiated to test the safety and tolerability of lipid-encapsidated mRNA encoding a neutralizing anti-CHIKV monoclonal antibody [111] (NCT03829384). Administration of the anti-inflammatory drug, CTLA4-Ig (also known as Abatacept), in conjunction with a neutralizing monoclonal antibody against CHIKV was highly protective against virus pathogenesis in mice [112]. The combination of antiviral and anti-inflammatory therapy may be a promising strategy for treating arthritogenic alphavirus infections and the ensuing immunopathology [113].
Fig 4. Schematic diagram of alphavirus entry pathway and inhibitors targeting each step.
Inhibitors that target alphavirus attachment and receptor binding, endosomal acidification, membrane fusion, and E1/E2 stability.
Other studies have identified small molecule inhibitors of alphavirus entry (Table 2) [114–116]. These include suramin, a known antiparasitic drug [117]. Molecular docking studies predict suramin intercalates between E1 DII and E2 domain C [114], potentially disrupting the stability of the E1/E2 heterotrimer and, in turn, its associated entry functions. Suramin also has activity against CHIKV in vivo and reduces virus burden and foot swelling in mice [118]. Other compounds that affect alphavirus entry include curcumin [116], a naturally occurring phenol, which reduces infectivity and blocks cell binding of CHIKV and Zika virus (ZIKV), an unrelated flavivirus [116]. Accordingly, transfection of viral RNA into cells in the presence of the drug bypasses the antiviral activity [116]. The flavonoid compound baicalin is also believed to affect CHIKV entry steps [119], although the exact mechanism of action is unknown. Co-protoporphyrin IX and Sn-protoporphyrin IX, 2 porphyrins that affect virus envelope integrity, impair adsorption of CHIKV, MAYV, SFV, and SINV to cells [120]. Obatoclax, another broad-spectrum antiviral against enveloped viruses, inhibits CHIKV and SFV infection by neutralizing endosomal pH and inhibiting fusion [121]. Chloroquine, a well-characterized antimalarial drug, has efficacy against CHIKV in cultured cells by neutralization of endosomal pH [122,123]. Although effective in cultured cells, studies in nonhuman primates showed that chloroquine treatment paradoxically resulted in higher viremia and delayed viral clearance [124], an effect that correlated with altered type I IFN responses. In human infections treated with chloroquine, viremia and persistent polyarthralgia were not improved [124]. Overall, these results with chloroquine highlight a need for caution when extrapolating from cell culture to in vivo systems.
Table 2. Inhibitors of alphavirus entry.
| Therapeutic target | Entry inhibitor [reference] | Mechanism of action |
|---|---|---|
| 1. Attachment factor and receptor binding | Monoclonal antibodies [107]; curcumin [116] Co-protoporphyrin IX and Sn-protoporphyrin IX [120] Suramin [114,115,118,142–144] |
Block receptor binding, ADCC; CDC Engage in hydrophobic interactions in the lipid bilayer to disrupt envelope integrity E1/E2 heterotrimer, may disrupt heterodimer assembly |
| 2. Clathrin-mediated endocytosis | Pitstop [145] | Clathrin inhibitor |
| 3. Endosomal acidification | Obatoclax [121], chloroquine [122–124] | Neutralizes endosomal pH |
| 4. Membrane fusion | Co-protoporphyrin IX and Sn-protoporphyrin IX [120] Monoclonal antibodies [107] |
Disrupt viral envelope integrity; block fusion |
Therapeutics that disrupt alphavirus entry include monoclonal antibodies and small molecule inhibitors. Steps in the entry pathway targeted (1) receptor binding, (2) clathrin-mediated endocytosis, (3) endosomal acidification, and (4) membrane fusion. ADCC, antibody-dependent cellular cytotoxicity; CDC, complement-dependent cytotoxicity.
Novel broad-spectrum antivirals that target fusion and internalization of several enveloped viruses including the alphavirus SFV were recently identified [125]. While these compounds reduced viral burden in cell culture and in vivo, they did not improve survival, at least in a mouse model of ZIKV infection. Moreover, inhibitors of entry that affect endosomal pH could have off-target effects given that this is a critical pathway in cellular homeostasis. The most effective entry inhibitors should block the attachment step, as this requires a highly virus-specific interaction between the receptor and viral attachment proteins. Indeed, most clinically approved entry inhibitors focus on this aspect of the life cycle [126] for viruses including human immunodeficiency virus (HIV), respiratory syncytial virus, varicella-zoster virus, herpes simplex virus, and hepatitis C virus. Other broad-spectrum antivirals that target the viral entry step include the envelope intercalating agent, LJ1001, which exploits differences in the biophysical properties of the viral and host membranes to prevent viral fusion while leaving host membranes unaffected [127].
Conclusions
Alphaviruses are rapidly emerging and reemerging human pathogens. Important structural, biochemical, and molecular insights into the entry step of the alphavirus life cycle have been made. These advances have enhanced our understanding of how alphaviruses attach to and invade target cells, how entry influences tissue tropism and virus pathogenesis, and importantly, highlight facets of the entry process for targeting with antiviral therapeutics. However, several key questions remain: What factors drive adaptation to HS attachment factor usage in cell culture versus natural adaptation? What are the alternative receptors for arthritogenic alphaviruses? What are the attachment and entry receptors for encephalitic alphaviruses? What are the entry pathways utilized by the currently described receptors for alphaviruses? How often and in what situations do non-clathrin-mediated entry pathways occur? How does the entry pathway affect tropism and immune evasion? How does receptor and/or attachment factor usage and their downstream signaling pathways contribute to alphavirus pathogenesis in vivo? Answering these fundamental questions will address gaps in our knowledge of the alphavirus entry pathway and may allow for the generation of countermeasures that more precisely target this critical first step in the alphavirus infection cycle.
Funding Statement
This work was supported by NIH grants R01AI114816, R01AI123348, R01AI143673, R01AI095436, and T32 AI007163. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Silva LA, Dermody TS. Chikungunya virus: epidemiology, replication, disease mechanisms, and prospective intervention strategies. J Clin Invest. 2017;127(3):737–749. 10.1172/JCI84417 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Fuller SD. The T = 4 envelope of Sindbis virus is organized by interactions with a complementary T = 3 capsid. Cell. 1987;48(6):923–934. 10.1016/0092-8674(87)90701-x [DOI] [PubMed] [Google Scholar]
- 3.Garoff H, Sjoberg M, Cheng RH. Budding of alphaviruses. Virus Res. 2004;106(2):103–116. 10.1016/j.virusres.2004.08.008 [DOI] [PubMed] [Google Scholar]
- 4.Rupp JC, Sokoloski KJ, Gebhart NN, Hardy RW. Alphavirus RNA synthesis and non-structural protein functions. J Gen Virol. 2015;96(9):2483–2500. 10.1099/jgv.0.000249 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Jose J, Snyder JE, Kuhn RJ. A structural and functional perspective of alphavirus replication and assembly. Future Microbiol. 2009;4(7):837–856. 10.2217/fmb.09.59 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Kielian M. Membrane fusion and the alphavirus life cycle. Adv Virus Res. 1995;45:113–151. 10.1016/s0065-3527(08)60059-7 [DOI] [PubMed] [Google Scholar]
- 7.Sjoberg M, Garoff H. Interactions between the transmembrane segments of the alphavirus E1 and E2 proteins play a role in virus budding and fusion. J Virol. 2003;77(6):3441–3450. 10.1128/jvi.77.6.3441-3450.2003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Byrd EA, Kielian M. The alphavirus E2 membrane-proximal domain impacts capsid interaction and glycoprotein lattice formation. J Virol. 2019;93(4). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Ramsey J, Mukhopadhyay S. Disentangling the frames, the state of research on the alphavirus 6K and TF proteins. Viruses. 2017;9(8). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Rogers KJ, Jones-Burrage S, Maury W, Mukhopadhyay S. TF protein of Sindbis virus antagonizes host type I interferon responses in a palmitoylation-dependent manner. Virology. 2020;542:63–70. 10.1016/j.virol.2020.01.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Voss JE, Vaney MC, Duquerroy S, Vonrhein C, Girard-Blanc C, Crublet E, et al. Glycoprotein organization of Chikungunya virus particles revealed by X-ray crystallography. Nature. 2010;468(7324):709–712. 10.1038/nature09555 [DOI] [PubMed] [Google Scholar]
- 12.de Curtis I, Simons K. Dissection of Semliki Forest virus glycoprotein delivery from the trans-Golgi network to the cell surface in permeabilized BHK cells. Proc Natl Acad Sci U S A. 1988;85(21):8052–8056. 10.1073/pnas.85.21.8052 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Sjoberg M, Lindqvist B, Garoff H. Activation of the alphavirus spike protein is suppressed by bound E3. J Virol. 2011;85(11):5644–5650. 10.1128/JVI.00130-11 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Fields W, Kielian M. Interactions involved in pH protection of the alphavirus fusion protein. Virology. 2015;486:173–179. 10.1016/j.virol.2015.08.028 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Uchime O, Fields W, Kielian M. The role of E3 in pH protection during alphavirus assembly and exit. J Virol. 2013;87(18):10255–10262. 10.1128/JVI.01507-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Snyder AJ, Mukhopadhyay S. The alphavirus E3 glycoprotein functions in a clade-specific manner. J Virol. 2012;86(24):13609–13620. 10.1128/JVI.01805-12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Zhang X, Fugere M, Day R, Kielian M. Furin processing and proteolytic activation of Semliki Forest virus. J Virol. 2003;77(5):2981–2989. 10.1128/jvi.77.5.2981-2989.2003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Yap ML, Klose T, Urakami A, Hasan SS, Akahata W, Rossmann MG. Structural studies of Chikungunya virus maturation. Proc Natl Acad Sci U S A. 2017;114(52):13703–13707. 10.1073/pnas.1713166114 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Zhang R, Hryc CF, Cong Y, Liu X, Jakana J, Gorchakov R, et al. 4.4 A cryo-EM structure of an enveloped alphavirus Venezuelan equine encephalitis virus. EMBO J. 2011;30(18):3854–3863. 10.1038/emboj.2011.261 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Garoff H, Simons K, Renkonen O. Isolation and characterization of the membrane proteins of Semliki Forest virus. Virology. 1974;61(2):493–504. 10.1016/0042-6822(74)90285-2 [DOI] [PubMed] [Google Scholar]
- 21.Basore K, Kim AS, Nelson CA, Zhang R, Smith BK, Uranga C, et al. Cryo-EM structure of Chikungunya virus in complex with the Mxra8 receptor. Cell. 2019;177(7):1725–1737.e16. 10.1016/j.cell.2019.04.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Li L, Jose J, Xiang Y, Kuhn RJ, Rossmann MG. Structural changes of envelope proteins during alphavirus fusion. Nature. 2010;468(7324):705–708. 10.1038/nature09546 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Chen L, Wang M, Zhu D, Sun Z, Ma J, Wang J, et al. Implication for alphavirus host-cell entry and assembly indicated by a 3.5A resolution cryo-EM structure. Nat Commun. 2018;9(1):5326 10.1038/s41467-018-07704-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Byrd EA, Kielian M. An alphavirus E2 membrane-proximal domain promotes envelope protein lateral interactions and virus budding. MBio. 2017;8(6). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Kielian M. Class II virus membrane fusion proteins. Virology. 2006;344(1):38–47. 10.1016/j.virol.2005.09.036 [DOI] [PubMed] [Google Scholar]
- 26.Lescar J, Roussel A, Wien MW, Navaza J, Fuller SD, Wengler G, et al. The Fusion glycoprotein shell of Semliki Forest virus: an icosahedral assembly primed for fusogenic activation at endosomal pH. Cell. 2001;105(1):137–148. 10.1016/s0092-8674(01)00303-8 [DOI] [PubMed] [Google Scholar]
- 27.Roussel A, Lescar J, Vaney MC, Wengler G, Wengler G, Rey FA. Structure and interactions at the viral surface of the envelope protein E1 of Semliki Forest virus. Structure. 2006;14(1):75–86. 10.1016/j.str.2005.09.014 [DOI] [PubMed] [Google Scholar]
- 28.Zheng Y, Sanchez-San Martin C, Qin ZL, Kielian M. The domain I-domain III linker plays an important role in the fusogenic conformational change of the alphavirus membrane fusion protein. J Virol. 2011;85(13):6334–6342. 10.1128/JVI.00596-11 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Wahlberg JM, Bron R, Wilschut J, Garoff H. Membrane fusion of Semliki Forest virus involves homotrimers of the fusion protein. J Virol. 1992;66(12):7309–7318. 10.1128/JVI.66.12.7309-7318.1992 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Wahlberg JM, Garoff H. Membrane fusion process of Semliki Forest virus. I: low pH-induced rearrangement in spike protein quaternary structure precedes virus penetration into cells. J Cell Biol. 1992;116(2):339–348. 10.1083/jcb.116.2.339 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Zeng X, Mukhopadhyay S, Brooks CL III. Residue-level resolution of alphavirus envelope protein interactions in pH-dependent fusion. Proc Natl Acad Sci U S A. 2015;112(7):2034–2039. 10.1073/pnas.1414190112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Liao M, Kielian M. Functions of the stem region of the Semliki Forest virus fusion protein during virus fusion and assembly. J Virol. 2006;80(22):11362–11369. 10.1128/JVI.01679-06 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Qin ZL, Zheng Y, Kielian M. Role of conserved histidine residues in the low-pH dependence of the Semliki Forest virus fusion protein. J Virol. 2009;83(9):4670–4677. 10.1128/JVI.02646-08 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Gibbons DL, Vaney MC, Roussel A, Vigouroux A, Reilly B, Lepault J, et al. Conformational change and protein-protein interactions of the fusion protein of Semliki Forest virus. Nature. 2004;427(6972):320–325. 10.1038/nature02239 [DOI] [PubMed] [Google Scholar]
- 35.Roman-Sosa G, Kielian M. The interaction of alphavirus E1 protein with exogenous domain III defines stages in virus-membrane fusion. J Virol. 2011;85(23):12271–12279. 10.1128/JVI.05902-11 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Sanchez-San Martin C, Sosa H, Kielian M. A stable prefusion intermediate of the alphavirus fusion protein reveals critical features of class II membrane fusion. Cell Host Microbe. 2008;4(6):600–608. 10.1016/j.chom.2008.10.012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Cao S, Zhang W. Characterization of an early-stage fusion intermediate of Sindbis virus using cryoelectron microscopy. Proc Natl Acad Sci U S A. 2013;110(33):13362–13367. 10.1073/pnas.1301911110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Sanchez-San Martin C, Nanda S, Zheng Y, Fields W, Kielian M. Cross-inhibition of chikungunya virus fusion and infection by alphavirus E1 domain III proteins. J Virol. 2013;87(13):7680–7687. 10.1128/JVI.00814-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.van Duijl-Richter MK, Hoornweg TE, Rodenhuis-Zybert IA, Smit JM. Early events in Chikungunya virus infection—from virus cell binding to membrane fusion. Viruses. 2015;7(7):3647–3674. 10.3390/v7072792 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Jolly CL, Sattentau QJ. Attachment factors. Adv Exp Med Biol. 2013;790:1–23. 10.1007/978-1-4614-7651-1_1 [DOI] [PubMed] [Google Scholar]
- 41.Maginnis MS. Virus–receptor interactions: the key to cellular invasion. J Mol Biol. 2018;430(17):2590–2611. 10.1016/j.jmb.2018.06.024 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Dimitrov DS. Virus entry: molecular mechanisms and biomedical applications. Nat Rev Microbiol. 2004;2(2):109–122. 10.1038/nrmicro817 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Carette JE, Raaben M, Wong AC, Herbert AS, Obernosterer G, Mulherkar N, et al. Ebola virus entry requires the cholesterol transporter Niemann-Pick C1. Nature. 2011;477(7364):340–343. 10.1038/nature10348 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Pirtle EC, Beran GW. Virus survival in the environment. Rev Sci Tech. 1991;10(3):733–748. 10.20506/rst.10.3.570 [DOI] [PubMed] [Google Scholar]
- 45.Gardner CL, Ebel GD, Ryman KD, Klimstra WB. Heparan sulfate binding by natural eastern equine encephalitis viruses promotes neurovirulence. Proc Natl Acad Sci U S A. 2011;108(38):16026–16031. 10.1073/pnas.1110617108 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Bernard KA, Klimstra WB, Johnston RE. Mutations in the E2 glycoprotein of Venezuelan equine encephalitis virus confer heparan sulfate interaction, low morbidity, and rapid clearance from blood of mice. Virology. 2000;276(1):93–103. 10.1006/viro.2000.0546 [DOI] [PubMed] [Google Scholar]
- 47.Ashbrook AW, Burrack KS, Silva LA, Montgomery SA, Heise MT, Morrison TE, et al. Residue 82 of the Chikungunya virus E2 attachment protein modulates viral dissemination and arthritis in mice. J Virol. 2014;88(21):12180–12192. 10.1128/JVI.01672-14 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Byrnes AP, Griffin DE. Large-plaque mutants of Sindbis virus show reduced binding to heparan sulfate, heightened viremia, and slower clearance from the circulation. J Virol. 2000;74(2):644–651. 10.1128/jvi.74.2.644-651.2000 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Griffin P, Dimitry JM, Sheehan PW, Lananna BV, Guo C, Robinette ML, et al. Circadian clock protein Rev-erbalpha regulates neuroinflammation. Proc Natl Acad Sci U S A. 2019;116(11):5102–5107. 10.1073/pnas.1812405116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Klimstra WB, Ryman KD, Johnston RE. Adaptation of Sindbis virus to BHK cells selects for use of heparan sulfate as an attachment receptor. J Virol. 1998;72(9):7357–7366. 10.1128/JVI.72.9.7357-7366.1998 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Smit JM, Waarts BL, Kimata K, Klimstra WB, Bittman R, Wilschut J. Adaptation of alphaviruses to heparan sulfate: interaction of Sindbis and Semliki forest viruses with liposomes containing lipid-conjugated heparin. J Virol. 2002;76(20):10128–10137. 10.1128/jvi.76.20.10128-10137.2002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Tanaka A, Tumkosit U, Nakamura S, Motooka D, Kishishita N, Priengprom T, et al. Genome-wide screening uncovers the significance of N-sulfation of heparan sulfate as a host cell factor for Chikungunya virus infection. J Virol. 2017;91(13). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Meertens L, Hafirassou ML, Couderc T, Bonnet-Madin L, Kril V, Kummerer BM, et al. FHL1 is a major host factor for chikungunya virus infection. Nature. 2019;574(7777):259–263. 10.1038/s41586-019-1578-4 [DOI] [PubMed] [Google Scholar]
- 54.Silva LA, Khomandiak S, Ashbrook AW, Weller R, Heise MT, Morrison TE, et al. A single-amino-acid polymorphism in Chikungunya virus E2 glycoprotein influences glycosaminoglycan utilization. J Virol. 2014;88(5):2385–2397. 10.1128/JVI.03116-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Hawman DW, Fox JM, Ashbrook AW, May NA, Schroeder KMS, Torres RM, et al. Pathogenic Chikungunya virus evades B cell responses to establish persistence. Cell Rep. 2016;16(5):1326–1338. 10.1016/j.celrep.2016.06.076 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Kerr PJ, Weir RC, Dalgarno L. Ross River virus variants selected during passage in chick embryo fibroblasts: serological, genetic, and biological changes. Virology. 1993;193(1):446–449. 10.1006/viro.1993.1143 [DOI] [PubMed] [Google Scholar]
- 57.Heil ML, Albee A, Strauss JH, Kuhn RJ. An amino acid substitution in the coding region of the E2 glycoprotein adapts Ross River virus to utilize heparan sulfate as an attachment moiety. J Virol. 2001;75(14):6303–6309. 10.1128/JVI.75.14.6303-6309.2001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Zhang W, Heil M, Kuhn RJ, Baker TS. Heparin binding sites on Ross River virus revealed by electron cryo-microscopy. Virology. 2005;332(2):511–518. 10.1016/j.virol.2004.11.043 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Smith TJ, Cheng RH, Olson NH, Peterson P, Chase E, Kuhn RJ, et al. Putative receptor binding sites on alphaviruses as visualized by cryoelectron microscopy. Proc Natl Acad Sci U S A. 1995;92(23):10648–10652. 10.1073/pnas.92.23.10648 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Kesari AS, Sharkey CM, Sanders DA. Role of heparan sulfate in entry and exit of Ross River virus glycoprotein-pseudotyped retroviral vectors. Virology. 2019;529:177–185. 10.1016/j.virol.2019.01.022 [DOI] [PubMed] [Google Scholar]
- 61.Gardner CL, Choi-Nurvitadhi J, Sun C, Bayer A, Hritz J, Ryman KD, et al. Natural variation in the heparan sulfate binding domain of the eastern equine encephalitis virus E2 glycoprotein alters interactions with cell surfaces and virulence in mice. J Virol. 2013;87(15):8582–8590. 10.1128/JVI.00937-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Ferguson MC, Saul S, Fragkoudis R, Weisheit S, Cox J, Patabendige A, et al. Ability of the encephalitic arbovirus Semliki Forest virus to cross the blood-brain barrier is determined by the charge of the E2 glycoprotein. J Virol. 2015;89(15):7536–7549. 10.1128/JVI.03645-14 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Bear JS, Byrnes AP, Griffin DE. Heparin-binding and patterns of virulence for two recombinant strains of Sindbis virus. Virology. 2006;347(1):183–190. 10.1016/j.virol.2005.11.034 [DOI] [PubMed] [Google Scholar]
- 64.Ryman KD, Gardner CL, Burke CW, Meier KC, Thompson JM, Klimstra WB. Heparan sulfate binding can contribute to the neurovirulence of neuroadapted and nonneuroadapted Sindbis viruses. J Virol. 2007;81(7):3563–3573. 10.1128/JVI.02494-06 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Hasan SS, Sun C, Kim AS, Watanabe Y, Chen CL, Klose T, et al. Cryo-EM structures of eastern equine encephalitis virus reveal mechanisms of virus disassembly and antibody neutralization. Cell Rep. 2018;25(11):3136–3147.e5. 10.1016/j.celrep.2018.11.067 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Chen CL, Hasan SS, Klose T, Sun Y, Buda G, Sun C, et al. Cryo-EM structure of eastern equine encephalitis virus in complex with heparan sulfate analogues. Proc Natl Acad Sci U S A. 2020;117(16):8890–8899. 10.1073/pnas.1910670117 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Klimstra WB, Nangle EM, Smith MS, Yurochko AD, Ryman KD. DC-SIGN and L-SIGN can act as attachment receptors for alphaviruses and distinguish between mosquito cell- and mammalian cell-derived viruses. J Virol. 2003;77(22):12022–12032. 10.1128/jvi.77.22.12022-12032.2003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Froelich S, Tai A, Kennedy K, Zubair A, Wang P. Pseudotyping lentiviral vectors with aura virus envelope glycoproteins for DC-SIGN-mediated transduction of dendritic cells. Hum Gene Ther. 2011;22(10):1281–1291. 10.1089/hum.2010.196 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Erbacher A, Gieseke F, Handgretinger R, Muller I. Dendritic cells: functional aspects of glycosylation and lectins. Hum Immunol. 2009;70(5):308–312. 10.1016/j.humimm.2009.02.005 [DOI] [PubMed] [Google Scholar]
- 70.Zhou T, Chen Y, Hao L, Zhang Y. DC-SIGN and immunoregulation. Cell Mol Immunol. 2006;3(4):279–283. [PubMed] [Google Scholar]
- 71.Lozach PY, Burleigh L, Staropoli I, Amara A. The C type lectins DC-SIGN and L-SIGN: receptors for viral glycoproteins. Methods Mol Biol. 2007;379:51–68. 10.1007/978-1-59745-393-6_4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Rogers KM, Heise M. Modulation of cellular tropism and innate antiviral response by viral glycans. J Innate Immun. 2009;1(5):405–412. 10.1159/000226422 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Gardner CL, Burke CW, Tesfay MZ, Glass PJ, Klimstra WB, Ryman KD. Eastern and Venezuelan equine encephalitis viruses differ in their ability to infect dendritic cells and macrophages: impact of altered cell tropism on pathogenesis. J Virol. 2008;82(21):10634–10646. 10.1128/JVI.01323-08 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Kolokoltsova OA, Domina AM, Kolokoltsov AA, Davey RA, Weaver SC, Watowich SJ. Alphavirus production is inhibited in neurofibromin 1-deficient cells through activated RAS signalling. Virology. 2008;377(1):133–142. 10.1016/j.virol.2008.03.025 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Jarvis CM, Zwick DB, Grim JC, Alam MM, Prost LR, Gardiner JC, et al. Antigen structure affects cellular routing through DC-SIGN. Proc Natl Acad Sci U S A. 2019;116(30):14862–14867. 10.1073/pnas.1820165116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Long KM, Whitmore AC, Ferris MT, Sempowski GD, McGee C, Trollinger B, et al. Dendritic cell immunoreceptor regulates Chikungunya virus pathogenesis in mice. J Virol. 2013;87(10):5697–5706. 10.1128/JVI.01611-12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Moller-Tank S, Maury W. Phosphatidylserine receptors: enhancers of enveloped virus entry and infection. Virology. 2014;468–470:565–580. 10.1016/j.virol.2014.09.009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Lemke G. How macrophages deal with death. Nat Rev Immunol. 2019;19(9):539–549. 10.1038/s41577-019-0167-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Amara A, Mercer J. Viral apoptotic mimicry. Nat Rev Microbiol. 2015;13(8):461–469. 10.1038/nrmicro3469 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Moller-Tank S, Kondratowicz AS, Davey RA, Rennert PD, Maury W. Role of the phosphatidylserine receptor TIM-1 in enveloped-virus entry. J Virol. 2013;87(15):8327–8341. 10.1128/JVI.01025-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Jemielity S, Wang JJ, Chan YK, Ahmed AA, Li W, Monahan S, et al. TIM-family proteins promote infection of multiple enveloped viruses through virion-associated phosphatidylserine. PLoS Pathog. 2013;9(3):e1003232 10.1371/journal.ppat.1003232 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Morizono K, Chen IS. Role of phosphatidylserine receptors in enveloped virus infection. J Virol. 2014;88(8):4275–4290. 10.1128/JVI.03287-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Morizono K, Xie Y, Olafsen T, Lee B, Dasgupta A, Wu AM, et al. The soluble serum protein Gas6 bridges virion envelope phosphatidylserine to the TAM receptor tyrosine kinase Axl to mediate viral entry. Cell Host Microbe. 2011;9(4):286–298. 10.1016/j.chom.2011.03.012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Vancini R, Hernandez R, Brown D. Alphavirus entry into host cells. Prog Mol Biol Transl Sci. 2015;129:33–62. 10.1016/bs.pmbts.2014.10.002 [DOI] [PubMed] [Google Scholar]
- 85.Rose PP, Hanna SL, Spiridigliozzi A, Wannissorn N, Beiting DP, Ross SR, et al. Natural resistance-associated macrophage protein is a cellular receptor for sindbis virus in both insect and mammalian hosts. Cell Host Microbe. 2011;10(2):97–104. 10.1016/j.chom.2011.06.009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Kim AS, Zimmerman O, Fox JM, Nelson CA, Basore K, Zhang R, et al. An evolutionary insertion in the Mxra8 receptor-binding site confers resistance to alphavirus infection and pathogenesis. Cell Host Microbe. 2020;27(3):428–440.e9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Zhang R, Kim AS, Fox JM, Nair S, Basore K, Klimstra WB, et al. Mxra8 is a receptor for multiple arthritogenic alphaviruses. Nature. 2018;557(7706):570–574. 10.1038/s41586-018-0121-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Fox JM, Diamond MS. Immune-mediated protection and pathogenesis of Chikungunya virus. J Immunol. 2016;197(11):4210–4218. 10.4049/jimmunol.1601426 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Zhang R, Earnest JT, Kim AS, Winkler ES, Desai P, Adams LJ, et al. Expression of the Mxra8 receptor promotes alphavirus infection and pathogenesis in mice and Drosophila. Cell Rep. 2019;28(10):2647–2658.e5. 10.1016/j.celrep.2019.07.105 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Song H, Zhao Z, Chai Y, Jin X, Li C, Yuan F, et al. Molecular basis of arthritogenic alphavirus receptor MXRA8 binding to Chikungunya virus envelope protein. Cell. 2019;177(7):1714–1724.e12. 10.1016/j.cell.2019.04.008 [DOI] [PubMed] [Google Scholar]
- 91.Weaver SC, Hagenbaugh A, Bellew LA, Netesov SV, Volchkov VE, Chang GJ, et al. A comparison of the nucleotide sequences of eastern and western equine encephalomyelitis viruses with those of other alphaviruses and related RNA viruses. Virology. 1993;197(1):375–390. 10.1006/viro.1993.1599 [DOI] [PubMed] [Google Scholar]
- 92.Wang KS, Kuhn RJ, Strauss EG, Ou S, Strauss JH. High-affinity laminin receptor is a receptor for Sindbis virus in mammalian cells. J Virol. 1992;66(8):4992–5001. 10.1128/JVI.66.8.4992-5001.1992 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Malygin AA, Bondarenko EI, Ivanisenko VA, Protopopova EV, Karpova GG, Loktev VB. C-terminal fragment of human laminin-binding protein contains a receptor domain for venezuelan equine encephalitis and tick-borne encephalitis viruses. Biochemistry (Mosc). 2009;74(12):1328–1336. [DOI] [PubMed] [Google Scholar]
- 94.Merkwirth C, Langer T. Prohibitin function within mitochondria: essential roles for cell proliferation and cristae morphogenesis. Biochim Biophys Acta. 2009;1793(1):27–32. 10.1016/j.bbamcr.2008.05.013 [DOI] [PubMed] [Google Scholar]
- 95.Wintachai P, Wikan N, Kuadkitkan A, Jaimipuk T, Ubol S, Pulmanausahakul R, et al. Identification of prohibitin as a Chikungunya virus receptor protein. J Med Virol. 2012;84(11):1757–1770. 10.1002/jmv.23403 [DOI] [PubMed] [Google Scholar]
- 96.Kuadkitkan A, Wikan N, Fongsaran C, Smith DR. Identification and characterization of prohibitin as a receptor protein mediating DENV-2 entry into insect cells. Virology. 2010;406(1):149–161. 10.1016/j.virol.2010.07.015 [DOI] [PubMed] [Google Scholar]
- 97.Leung JY, Ng MM, Chu JJ. Replication of alphaviruses: a review on the entry process of alphaviruses into cells. Adv Virol. 2011;2011:249640 10.1155/2011/249640 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Kielian M, Jungerwirth S. Mechanisms of enveloped virus entry into cells. Mol Biol Med. 1990;7(1):17–31. [PubMed] [Google Scholar]
- 99.Sanchez-San Martin C, Liu CY, Kielian M. Dealing with low pH: entry and exit of alphaviruses and flaviviruses. Trends Microbiol. 2009;17(11):514–521. 10.1016/j.tim.2009.08.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Hoornweg TE, van Duijl-Richter MKS, Ayala Nunez NV, Albulescu IC, van Hemert MJ, Smit JM. Dynamics of Chikungunya virus cell entry unraveled by single-virus tracking in living cells. J Virol. 2016;90(9):4745–4756. 10.1128/JVI.03184-15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Ooi YS, Stiles KM, Liu CY, Taylor GM, Kielian M. Genome-wide RNAi screen identifies novel host proteins required for alphavirus entry. PLoS Pathog. 2013;9(12):e1003835 10.1371/journal.ppat.1003835 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Carvalho CAM, Silva JL, Oliveira AC, Gomes AMO. On the entry of an emerging arbovirus into host cells: Mayaro virus takes the highway to the cytoplasm through fusion with early endosomes and caveolae-derived vesicles. PeerJ. 2017;5:e3245 10.7717/peerj.3245 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Vancini R, Wang G, Ferreira D, Hernandez R, Brown DT. Alphavirus genome delivery occurs directly at the plasma membrane in a time- and temperature-dependent process. J Virol. 2013;87(8):4352–4359. 10.1128/JVI.03412-12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Lee CHR, Mohamed Hussain K, Chu JJH. Macropinocytosis dependent entry of Chikungunya virus into human muscle cells. PLoS Negl Trop Dis. 2019;13(8):e0007610 10.1371/journal.pntd.0007610 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Powell LA, Fox JM, Kose N, Kim AS, Majedi M, Bombardi R, et al. Human monoclonal antibodies against Ross River virus target epitopes within the E2 protein and protect against disease. PLoS Pathog. 2020;16(5):e1008517. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Pelfrene E, Mura M, Cavaleiro Sanches A, Cavaleri M. Monoclonal antibodies as anti-infective products: a promising future? Clin Microbiol Infect 2019;25(1):60–64. 10.1016/j.cmi.2018.04.024 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Kumar R, Shrivastava T, Samal S, Ahmed S, Parray HA. Antibody-based therapeutic interventions: possible strategy to counter chikungunya viral infection. Appl Microbiol Biotechnol. 2020;104(8):3209–3228. 10.1007/s00253-020-10437-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Fox JM, Long F, Edeling MA, Lin H, van Duijl-Richter MKS, Fong RH, et al. Broadly neutralizing alphavirus antibodies bind an epitope on E2 and inhibit entry and egress. Cell. 2015;163(5):1095–1107. 10.1016/j.cell.2015.10.050 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Fox JM, Roy V, Gunn BM, Huang L, Edeling MA, Mack M, et al. Optimal therapeutic activity of monoclonal antibodies against chikungunya virus requires Fc-FcgammaR interaction on monocytes. Sci Immunol. 2019;4(32). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Contopoulos-Ioannidis D, Newman-Lindsay S, Chow C, LaBeaud AD. Mother-to-child transmission of Chikungunya virus: a systematic review and meta-analysis. PLoS Negl Trop Dis. 2018;12(6):e0006510 10.1371/journal.pntd.0006510 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Kose N, Fox JM, Sapparapu G, Bombardi R, Tennekoon RN, de Silva AD, et al. A lipid-encapsulated mRNA encoding a potently neutralizing human monoclonal antibody protects against chikungunya infection. Sci Immunol. 2019;4(35). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Miner JJ, Cook LE, Hong JP, Smith AM, Richner JM, Shimak RM, et al. Therapy with CTLA4-Ig and an antiviral monoclonal antibody controls chikungunya virus arthritis. Sci Transl Med. 2017;9(375). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Amaral JK, Bilsborrow JB, Schoen RT. Chronic Chikungunya arthritis and rheumatoid arthritis: what they have in common. Am J Med. 2020;133(3):e91–e97. 10.1016/j.amjmed.2019.10.005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Ho YJ, Wang YM, Lu JW, Wu TY, Lin LI, Kuo SC, et al. Suramin inhibits Chikungunya virus entry and transmission. PLoS ONE. 2015;10(7):e0133511 10.1371/journal.pone.0133511 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Albulescu IC, White-Scholten L, Tas A, Hoornweg TE, Ferla S, Kovacikova K, et al. Suramin inhibits Chikungunya virus replication by interacting with virions and blocking the early steps of infection. Viruses. 2020;12(3). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Mounce BC, Cesaro T, Carrau L, Vallet T, Vignuzzi M. Curcumin inhibits Zika and chikungunya virus infection by inhibiting cell binding. Antivir Res. 2017;142:148–157. 10.1016/j.antiviral.2017.03.014 [DOI] [PubMed] [Google Scholar]
- 117.McGeary RP, Bennett AJ, Tran QB, Cosgrove KL, Ross BP. Suramin: clinical uses and structure-activity relationships. Mini Rev Med Chem. 2008;8(13):1384–1394. 10.2174/138955708786369573 [DOI] [PubMed] [Google Scholar]
- 118.Kuo SC, Wang YM, Ho YJ, Chang TY, Lai ZZ, Tsui PY, et al. Suramin treatment reduces chikungunya pathogenesis in mice. Antivir Res. 2016;134:89–96. 10.1016/j.antiviral.2016.07.025 [DOI] [PubMed] [Google Scholar]
- 119.Oo A, Rausalu K, Merits A, Higgs S, Vanlandingham D, Bakar SA, et al. Deciphering the potential of baicalin as an antiviral agent for Chikungunya virus infection. Antivir Res. 2018;150:101–111. 10.1016/j.antiviral.2017.12.012 [DOI] [PubMed] [Google Scholar]
- 120.Neris RLS, Figueiredo CM, Higa LM, Araujo DF, Carvalho CAM, Vercoza BRF, et al. Co-protoporphyrin IX and Sn-protoporphyrin IX inactivate Zika, Chikungunya and other arboviruses by targeting the viral envelope. Sci Rep. 2018;8(1):9805 10.1038/s41598-018-27855-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Varghese FS, Rausalu K, Hakanen M, Saul S, Kummerer BM, Susi P, et al. Obatoclax inhibits alphavirus membrane fusion by neutralizing the acidic environment of endocytic compartments. Antimicrob Agents Chemother. 2017;61(3). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Khan M, Santhosh SR, Tiwari M, Lakshmana Rao PV, Parida M. Assessment of in vitro prophylactic and therapeutic efficacy of chloroquine against Chikungunya virus in vero cells. J Med Virol. 2010;82(5):817–824. 10.1002/jmv.21663 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Marti-Carvajal A, Ramon-Pardo P, Javelle E, Simon F, Aldighieri S, Horvath H, et al. Interventions for treating patients with chikungunya virus infection-related rheumatic and musculoskeletal disorders: a systematic review. PLoS ONE. 2017;12(6):e0179028 10.1371/journal.pone.0179028 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Roques P, Thiberville SD, Dupuis-Maguiraga L, Lum FM, Labadie K, Martinon F, et al. Paradoxical effect of chloroquine treatment in enhancing Chikungunya virus infection. Viruses. 2018;10(5). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Mazzon M, Ortega-Prieto AM, Imrie D, Luft C, Hess L, Czieso S, et al. Identification of broad-spectrum antiviral compounds by targeting viral entry. Viruses. 2019;11(2). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.De Clercq E, Li G. Approved antiviral drugs over the past 50 years. Clin Microbiol Rev. 2016;29(3):695–747. 10.1128/CMR.00102-15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Vigant F, Lee J, Hollmann A, Tanner LB, Akyol Ataman Z, Yun T, et al. A mechanistic paradigm for broad-spectrum antivirals that target virus-cell fusion. PLoS Pathog. 2013;9(4):e1003297 10.1371/journal.ppat.1003297 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Tsetsarkin KA, Vanlandingham DL, McGee CE, Higgs S. A single mutation in chikungunya virus affects vector specificity and epidemic potential. PLoS Pathog. 2007;3(12):e201 10.1371/journal.ppat.0030201 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Rezza G, Chen R, Weaver SC. O'nyong-nyong fever: a neglected mosquito-borne viral disease. Pathog Glob Health. 2017;111(6):271–275. 10.1080/20477724.2017.1355431 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Mackay IM, Arden KE. Mayaro virus: a forest virus primed for a trip to the city? Microbes Infect. 2016;18(12):724–734. 10.1016/j.micinf.2016.10.007 [DOI] [PubMed] [Google Scholar]
- 131.Long KC, Ziegler SA, Thangamani S, Hausser NL, Kochel TJ, Higgs S, et al. Experimental transmission of Mayaro virus by Aedes aegypti. Am J Trop Med Hyg. 2011;85(4):750–757. 10.4269/ajtmh.2011.11-0359 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Auguste AJ, Liria J, Forrester NL, Giambalvo D, Moncada M, Long KC, et al. Evolutionary and ecological characterization of Mayaro virus strains isolated during an outbreak, Venezuela, 2010. Emerg Infect Dis. 2015;21(10):1742–1750. 10.3201/eid2110.141660 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Shocket MS, Ryan SJ, Mordecai EA. Temperature explains broad patterns of Ross River virus transmission. Elife. 2018;7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Stephenson EB, Peel AJ, Reid SA, Jansen CC, McCallum H. The non-human reservoirs of Ross River virus: a systematic review of the evidence. Parasit Vectors. 2018;11(1):188 10.1186/s13071-018-2733-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Ylosmaki E, Martikainen M, Hinkkanen A, Saksela K. Attenuation of Semliki Forest virus neurovirulence by microRNA-mediated detargeting. J Virol. 2013;87(1):335–344. 10.1128/JVI.01940-12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Soghigian J, Andreadis TG, Molaei G. Population genomics of Culiseta melanura, the principal vector of Eastern equine encephalitis virus in the United States. PLoS Negl Trop Dis. 2018;12(8):e0006698 10.1371/journal.pntd.0006698 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137.Mukherjee S, Moody EE, Lewokzco K, Huddleston DB, Huang J, Rowland ME, et al. Eastern equine encephalitis in Tennessee: 2002–2008. J Med Entomol. 2012;49(3):731–738. 10.1603/me11151 [DOI] [PubMed] [Google Scholar]
- 138.Hesson JC, Lundin E, Lundkvist A, Lundstrom JO. Surveillance of mosquito vectors in Southern Sweden for Flaviviruses and Sindbis virus. Infect Ecol Epidemiol. 2019;9(1):1698903 10.1080/20008686.2019.1698903 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Ziegler U, Fischer D, Eiden M, Reuschel M, Rinder M, Muller K, et al. Sindbis virus—a wild bird associated zoonotic arbovirus circulates in Germany. Vet Microbiol. 2019;239:108453 10.1016/j.vetmic.2019.108453 [DOI] [PubMed] [Google Scholar]
- 140.Navarro JC, Medina G, Vasquez C, Coffey LL, Wang E, Suarez A, et al. Postepizootic persistence of Venezuelan equine encephalitis virus. Emerg Infect Dis. 2005;11(12):1907–1915. 10.3201/eid1112.050533 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Kenney JL, Adams AP, Weaver SC. Transmission potential of two chimeric western equine encephalitis vaccine candidates in Culex tarsalis. Am J Trop Med Hyg. 2010;82(2):354–359. 10.4269/ajtmh.2010.09-0092 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Lu JW, Hsieh PS, Lin CC, Hu MK, Huang SM, Wang YM, et al. Synergistic effects of combination treatment using EGCG and suramin against the chikungunya virus. Biochem Biophys Res Commun. 2017;491(3):595–602. 10.1016/j.bbrc.2017.07.157 [DOI] [PubMed] [Google Scholar]
- 143.Henss L, Beck S, Weidner T, Biedenkopf N, Sliva K, Weber C, et al. Suramin is a potent inhibitor of Chikungunya and Ebola virus cell entry. Virol J. 2016;13:149 10.1186/s12985-016-0607-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144.Albulescu IC, van Hoolwerff M, Wolters LA, Bottaro E, Nastruzzi C, Yang SC, et al. Suramin inhibits chikungunya virus replication through multiple mechanisms. Antivir Res. 2015;121:39–46. 10.1016/j.antiviral.2015.06.013 [DOI] [PubMed] [Google Scholar]
- 145.Lee RC, Hapuarachchi HC, Chen KC, Hussain KM, Chen H, Low SL, et al. Mosquito cellular factors and functions in mediating the infectious entry of chikungunya virus. PLoS Negl Trop Dis. 2013;7(2):e2050 10.1371/journal.pntd.0002050 [DOI] [PMC free article] [PubMed] [Google Scholar]