Summary
We meta-analyze amyotrophic lateral sclerosis (ALS) genome-wide association study (GWAS) data of European and Chinese populations (84,694 individuals). We find an additional significant association between rs58854276 spanning ACSL5-ZDHHC6 with ALS (p = 8.3 × 10−9), with replication in an independent Australian cohort (1,502 individuals; p = 0.037). Moreover, B4GALNT1, G2E3-SCFD1, and TRIP11-ATXN3 are identified using a gene-based analysis. ACSL5 has been associated with rapid weight loss, as has another ALS-associated gene, GPX3. Weight loss is frequent in ALS patients and is associated with shorter survival. We investigate the effect of the ACSL5 and GPX3 single-nucleotide polymorphisms (SNPs), using longitudinal body composition and weight data of 77 patients and 77 controls. In patients’ fat-free mass, although not significant, we observe an effect in the expected direction (rs58854276: −2.1 ± 1.3 kg/A allele, p = 0.053; rs3828599: −1.0 ± 1.3 kg/A allele, p = 0.22). No effect was observed in controls. Our findings support the increasing interest in lipid metabolism in ALS and link the disease genetics to weight loss in patients.
Keywords: amyotrophic lateral sclerosis, motor neuron disease, cross-ethnic meta-analysis, weight loss, genome-wide association study, genetics, genomics, eQTLs, fat-free mass, longitudinal study
Graphical Abstract

Highlights
-
•
Cross-ethnic meta-analysis finds an association between the ACSL5-ZDHHC6 locus and ALS
-
•
The ACSL5-ZDHHC6 association is replicated in an independent Australian cohort
-
•
ACSL5-ZDHHC6 lead SNP is in ACSL5 and is an eQTL of ZDHHC6 in brain tissues
-
•
ACSL5 SNPs might have an effect on fat-free mass in ALS patients
Using meta-analysis of European and Chinese ALS GWAS data, Iacoangeli et al. find an association between ACSL5-ZDHHC6 and ALS risk, with replication in an Australian cohort. They identify B4GALNT1, G2E3-SCFD1, and TRIP11-ATXN3 using a gene-based analysis. They also find a suggestive association between ACSL5 SNPs and lower fat-free mass in patients.
Introduction
Amyotrophic lateral sclerosis (ALS) is a neurodegenerative disease, primarily affecting upper and lower motor neurons, that results in progressive weakness and culminates in death from neuromuscular respiratory failure, typically 2–5 years after diagnosis (Brown and Al-Chalabi, 2017). Many genetic factors that drive and contribute to its development and progression have been identified (Iacoangeli et al., 2019b). Rare disruptive mutations have been shown to be responsible for approximately two thirds of the 5%–10% of patients with a family history of ALS (Renton et al., 2014) and for ~10%–15% of the remaining cases (Renton et al., 2014; Chia et al., 2018; Al-Chalabi, 2017) who do not report a family history of ALS. Genome-wide association studies (GWASs) have been particularly successful in the discovery of loci involved with the disease; for example, the initial identification of the C9orf72 locus (Shatunov et al., 2010) narrowed down a much larger locus identified through linkage (Vance et al., 2006; Morita et al., 2006) and led to the discovery of a pathogenic hexanucleotide-repeat expansion, the most common cause of ALS in the European population (Iacoangeli et al., 2019a). Moreover, GWASs have provided direct evidence of the major contribution of single-nucleotide polymorphisms (SNPs) to the heritability of ALS (Fogh et al., 2014; van Rheenen et al., 2016) and have identified a number of loci associated with the disease. To gain new insight into the genetics of ALS, we meta-analyzed the summary statistics from two ALS GWASs: the largest ALS study to date on >80,000 individuals of European ancestry (Nicolas et al., 2018) and a Chinese ALS study on >4,000 individuals (Benyamin et al., 2017). This cross-ethnic approach was previously used with an older European GWAS (van Rheenen et al., 2016) and led to the discovery of the association between the GPX3-TNIP1 locus and ALS (Benyamin et al., 2017).
Results
GWAS Meta-analysis
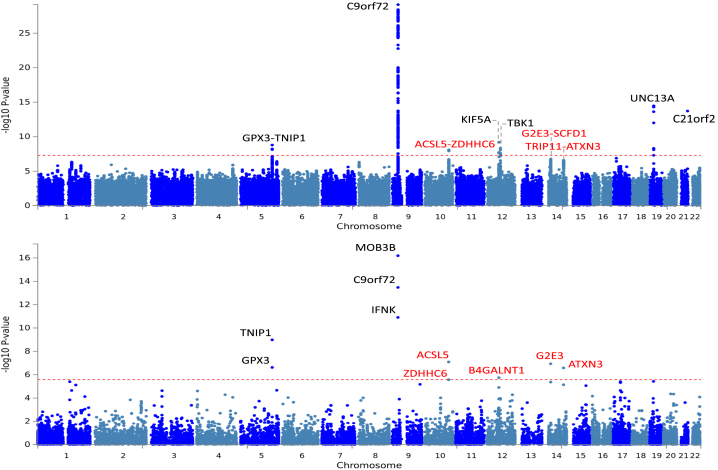
We meta-analyzed the summary statistics of the European and Chinese GWASs (STAR Methods). The total number of individuals was 84,694 (22,040 cases and 62,654 controls), and the total number of meta-analyzed SNPs was 5,356,204. No inflation of test statistics in the quantile-quantile plot (λgc = 1.042 and λ1,000 = 1.001) was observed (Figure S1). SNP-based meta-analysis (Figure 1A) replicated the previously identified loci C9orf72, UNC13A, and GPX3-TNIP1. C21orf2, TBK1, and KIF5A SNPs previously reported to be associated with ALS risk were present only in the European dataset. We identified an additional association in the ACSL5-ZDHHC6 locus (five significant SNPs in linkage disequilibrium [LD], lead SNP rs58854276 p = 8.3 × 10−9, A allele frequency 64.9%, odds ratio [OR] = 1.08, 95% confidence interval [CI] = 1.05–1.11; Table 1; Figure S2A) and two putative loci, G2E3-SCFD1 (lead SNP rs229247 p = 2.2 × 10−7, T allele frequency 47.9%, OR = 1.07, 95% CI = 1.04–1.10; Table 1; Figure S2B) and TRIP11-ATXN3 (lead SNP rs10143310 p = 2.6 × 10−7, C allele frequency 24.5%, OR = 1.08, 95% CI = 1.05–1.12; Table 1; Figure S2C). Using an independent Australian cohort (837 cases and 665 controls of European ancestry; STAR Methods; Table S1), we replicated the association between the ACSL5 lead SNP, rs58854276, and ALS (A allele frequency 66.1%, OR = 1.18, 95% CI = 1.01–1.38, p = 0.037). rs58854276 was not the lead ACSL5 SNP in either of the meta-analyzed studies. It was the second most significant ACSL5 SNP in the Chinese study and the fourth most significant ACSL5 SNP in the European study (p = 0.00015 and p = 1.1 × 10−6 respectively; Tables 1 and S2), highlighting the differences in terms of LD structure and frequencies in the two populations. ACSL5 and ZDHHC6 (gene-p = 8.1 × 10−8 and 2.7 × 10−6, respectively) and one gene in each of the two putative loci, ATXN3 (gene-p = 2.6 × 10−7) and G2E3 (gene-p = 1.2 × 10−7), were genome-wide significant in the gene-based analysis (Figure 1B; STAR Methods), consistent with the information contributed by the SNP associations, and recognizing that neighboring genes have overlapping boundaries in the gene-based analyses so that the same SNPs can contribute to more than one gene test. Another gene, B4GALNT1, was genome-wide significant in the gene-based analysis (gene-p = 1.8 × 10−6); however, no SNPs individually reached the putative threshold (lead SNP rs12320537, C allele frequency 20.5%, p = 6.2 × 10−6). Finally, using the summary statistics data of our previously published gene-burden analysis of disruptive, damaging, and missense variants on 4,389 ALS patients and 1,846 controls (van der Spek et al., 2019), for the seven genes in the four identified loci, i.e., ACSL5, ZDHHC6, B4GALNT1, SCFD1, G2E3, TRIP11, and ATXN3, we observed an association between rare variants in G2E3 and ALS (p = 0.0019, OR = 2.32, 95% CI = 1.27–4.23; Table 1).
Figure 1.
Genome-wide Meta-analysis Results
Manhattan plots of the (A) SNP-based results and (B) gene-based results. Loci previously identified are in black. Additional loci identified by our meta-analysis are in red.
Table 1.
GWAS Results for the Four Identified Loci and Their Lead SNPs in Our SNP Meta-analysis, Gene-Based Analysis, and Project MinE Gene Burden Analysis of Rare Variants
| Lead SNP rs No. | Effect Allele | Effect Allele Frequency (%) | OR (95% CI) | Lead SNP p Value | Magma Gene p Value | Rare Variants Gene-Burden OR (CI) | Rare Variants Gene-Burden p Value | |
|---|---|---|---|---|---|---|---|---|
| ACSL5/ZDHHC6 (Benyamin et al. [2017] Chinese cohort) | rs58854276 | A | 51.1 | 1.20 (1.08–1.36) | 0.00015 | 3.1 × 10−5/6.5 × 10−5 | – | – |
| ACSL5/ZDHHC6 (Nicolas et al., [2018] European cohort) | rs58854276 | A | 65.6 | 1.07 (1.04–1.10) | 1.1 × 10−6 | 7.8 × 10−6/1.5 × 10−4 | – | – |
| ACSL5/ZDHHC6 (this study) | rs58854276 | A | 64.9 | 1.08 (1.05–1.11) | 8.3 × 10−9 | 8.1 × 10−8/2.7 × 10−6 | 1.78 (0.77–4.09)/0.84 (0.52–1.35) | 0.14/0.98 |
| ACSL5 (replication) | rs58854276 | A | 66.1 | 1.18 (1.01–1.38) | 0.037 | – | – | – |
| ACSL5 (joint) | rs58854276 | A | 64.9 | 1.09 (1.06–1.11) | 1.5 × 10−9 | – | – | – |
| B4GALNT1 (this study) | rs12320537 | C | 20.5 | 1.07 (1.04–1.11) | 6.2 × 10−6 | 1.8 × 10−6 | 0.65 (0.40–1.04) | 0.07 |
| G2E3/SCFD1 (this study) | rs229247 | T | 47.9 | 1.07 (1.04–1.10) | 2.2 × 10−7 | 1.2 × 10−7/4.2 × 10−6 | 2.32 (1.27–4.23)/0.84 (0.52–1.35) | 0.0019/0.46 |
| TRIP11/ATXN3 (this study) | rs10143310 | C | 24.5 | 1.08 (1.05–1.12) | 2.6 × 10−7 | 7.2 × 10−6/2.6 × 10−7 | 1.05 (0.86–1.28)/NA | 0.59/NA |
ACSL5 lead SNP results for the replication cohort and the two meta-analyzed GWAS are also reported. NA, not applicable.
Fine-Mapping of the Identified Loci
We attempted to fine-map the four identified loci by assessing whether their independent lead SNPs or LD proxies have cis expression quantitative trait locus (cis-eQTL) effects observed in brain and blood tissues in the GTEx data (eGTEx Project, 2017) (Table 2; STAR Methods). Only one lead SNP per locus was selected as no other significant SNP in the loci met our independence criterion (r2 < 0.80). All four lead SNPs were in LD with cis-eQTLs for several brain tissues (Table 2). rs12320537 (B4GALNT1 lead SNP) was in LD with rs2258877 (r2 = 0.88), a brain cerebellum and cerebellar hemisphere cis-eQTL of B4GALNT1 (p = 1.8 × 10−5 and 1.7 × 10−7, respectively). rs58854276 (ACSL5-ZDHHC6 lead SNP) was in LD with rs2419629 (r2 = 0.90), a brain cerebellum and brain nucleus accumbens basal ganglia cis-eQTL of ZDHHC6 (p = 2.1 × 10−7 and 1.9 × 10−8, respectively); with rs12414780 (r2 = 0.83), a brain frontal cortex cis-eQTL of ZDHHC6 (3.2 × 10−7); and with rs72821869 (r2 = 0.84), a brain cortex cis-eQTL of ZDHHC6 (7.6 × 10−6). rs229247 (SCFD1-G2E3 lead SNP) was in LD with cis-eQTLs of SCFD1 in brain cortex (rs229173, r2 = 0.95, p = 2.1 × 10−7), brain anterior cingulate cortex (rs7154847, r2 = 0.89, p = 3.3 × 10−7), brain cerebellar hemisphere (rs229231, r2 = 0.99, p = 3.1 × 10−16), brain cerebellum (rs229152, r2 = 0.94, p = 2.2 × 10−24), and brain frontal cortex (rs10130830, r2 = 0.91, p = 1.4 × 10−8). rs10143310 (TRIPP11-ATXN3 lead SNP) was in LD with rs2896190 (r2 = 1), a cis-eQTL of TRIP11 in brain cerebellum and cerebellar hemisphere (p = 2.5 × 10−8 and 6.8 × 10−7, respectively). The four lead SNPs were also in strong LD with blood cis-eQTLs. The complete results from this analysis are available in Table 2.
Table 2.
eQTL Effect of the Lead SNPs in Brain and Whole-Blood Tissues
| Lead SNP Gene | Lead SNP | eQTL SNP | eQTL SNP Gene | r2 | Tissue | Ref | Alt | Ensebl Gene ID | Minor Allele Samples | Minor Allele Count | MAF | p Value | Slope | Slope SE |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ATXN3 | rs10143310 | rs2896190 | TRIP11 | 1.00 | brain cerebellar hemisphere | A | G | ENSG00000100815.12 | 78 | 90 | 0.26 | 6.8 × 10−7 | 0.24 | 0.05 |
| ATXN3 | rs10143310 | rs2896190 | TRIP11 | 1.00 | brain cerebellum | A | G | ENSG00000100815.12 | 89 | 104 | 0.25 | 2.5 × 10−8 | 0.29 | 0.05 |
| ATXN3 | rs10143310 | rs7142326 | ATNX3 | 0.56 | whole blood | T | C | ENSG00000066427.21 | 433 | 557 | 0.42 | 3.2 × 10−7 | 0.13 | 0.02 |
| ATXN3 | rs10143310 | rs76497846 | TRIP11 | 0.62 | whole blood | G | A | ENSG00000100815.12 | 407 | 516 | 0.39 | 3.0 × 10−5 | 0.09 | 0.02 |
| ATXN3 | rs10143310 | rs12587248 | NDUFB1 | 0.62 | whole blood | T | C | ENSG00000183648.9 | 356 | 423 | 0.32 | 2.5 × 10−8 | 0.10 | 0.02 |
| B4GALNT1 | rs12320537 | rs2258877 | B4GALNT1 | 0.88 | brain cerebellar hemisphere | A | G | ENSG00000135454.13 | 79 | 92 | 0.26 | 1.7 × 10−7 | 0.24 | 0.04 |
| B4GALNT1 | rs12320537 | rs2258877 | B4GALNT1 | 0.88 | brain cerebellum | A | G | ENSG00000135454.13 | 95 | 106 | 0.25 | 1.8 × 10−5 | 0.18 | 0.04 |
| B4GALNT1 | rs12320537 | rs12322482 | ATP23 | 0.99 | whole blood | G | A | ENSG00000166896.7 | 245 | 272 | 0.20 | 1.4 × 10−10 | −0.29 | 0.04 |
| SCFD1 | rs229247 | rs7154847 | SCFD1 | 0.89 | Brain anterior cingulate cortex BA24 | G | A | ENSG00000092108.20 | 82 | 101 | 0.34 | 3.3 × 10−7 | 0.33 | 0.06 |
| SCFD1 | rs229247 | rs229231 | SCFD1 | 0.99 | brain cerebellar hemisphere | G | A | ENSG00000092108.20 | 107 | 143 | 0.41 | 3.1 × 10−16 | 0.33 | 0.03 |
| SCFD1 | rs229247 | rs229152 | SCFD1 | 0.94 | brain cerebellum | T | C | ENSG00000092108.20 | 126 | 164 | 0.39 | 2.2 × 10−24 | 0.37 | 0.03 |
| SCFD1 | rs229247 | rs229173 | SCFD1 | 0.95 | brain cortex | T | C | ENSG00000092108.20 | 123 | 159 | 0.39 | 2.1 × 10−7 | 0.23 | 0.04 |
| SCFD1 | rs229247 | rs10130830 | SCFD1 | 0.91 | brain frontal cortex BA9 | A | G | ENSG00000092108.20 | 113 | 146 | 0.42 | 1.4 × 10−8 | 0.27 | 0.05 |
| SCFD1 | rs229247 | rs448175 | SCFD1 | 1.00 | whole blood | G | T | ENSG00000092108.20 | 415 | 536 | 0.40 | 1.9 × 10−56 | −0.28 | 0.02 |
| ACSL5 | rs58854276 | rs2419629 | ZDHHC6 | 0.90 | brain cerebellum | A | G | ENSG00000023041.11 | 113 | 144 | 0.34 | 2.1 × 10−7 | 0.28 | 0.05 |
| ACSL5 | rs58854276 | rs72821869 | ZDHHC6 | 0.84 | brain cortex | C | T | ENSG00000023041.11 | 121 | 152 | 0.37 | 7.6 × 10−6 | 0.29 | 0.06 |
| ACSL5 | rs58854276 | rs12414780 | ZDHHC6 | 0.83 | brain frontal cortex BA9 | C | G | ENSG00000023041.11 | 100 | 123 | 0.36 | 3.2 × 10−7 | 0.41 | 0.08 |
| ACSL5 | rs58854276 | rs2419629 | ZDHHC6 | 0.90 | brain nucleus accumbens basal ganglia | A | G | ENSG00000023041.11 | 120 | 151 | 0.37 | 1.9 × 10−8 | 0.40 | 0.07 |
| ACSL5 | rs58854276 | rs12414780 | ZDHHC6 | 0.83 | brain putamen basal ganglia | C | G | ENSG00000023041.11 | 101 | 129 | 0.38 | 1.3 × 10−5 | 0.34 | 0.08 |
| ACSL5 | rs58854276 | rs72821869 | ACSL5 | 0.84 | whole blood | C | T | ENSG00000197142.10 | 356 | 446 | 0.33 | 3.2 × 10−48 | −0.38 | 0.02 |
For each lead SNP, we reported the most significant eQTLs from GTEx in LD (r2 > 0.5) with the lead SNP. For each eQTL, we reported the the r2 with the corresponding lead SNP, the tissue in which the effect was observed, the corresponding regulated gene, the number of samples carrying the minor allele, the total number of minor alleles, the minor allele frequency, the p value, and regression slope and its standard error. The data are from GTEx version 8.
Investigation of the Effect of ACSL5 and GPX3 SNPs on Patients’ Body Weight and Composition
ACSL5 SNPs and their overexpression have been associated with rapid weight loss in humans (Adamo et al., 2007; Teng et al., 2009). Interestingly, another ALS gene, GPX3, was recently found to be associated with weight loss (Langhardt et al., 2018). To investigate the effect of the ALS-associated SNPs in ACSL5 and GPX3 on patients’ weight measures, we set out to test whether the SNPs (ACSL5 lead SNP rs58854276 and GPX3 lead SNP rs3828599) were associated with decline and difference in weight traits within the context of ALS (Table 3). We used a dataset from the metabolic exploration in neurodegenerative disease-motor neuron disease (MEND-MND) initiative that included 77 cases and 77 controls (STAR Methods; Table S4) for whom fat mass, fat-free mass, body weight, and body mass index (BMI) were available (complete results in Table S3). This dataset included longitudinal records for 67 of the 77 cases. In a linear regression analysis fitting sex as a covariate and using baseline measures, cases had significantly lower fat-free mass than controls (−4.2 ± 1.2 kg, p = 1.9 × 10−4); although cases had lower values for the other traits, they were not significantly different (pBMI = 0.58, pweight = 0.32, and pfat-mass = 0.22). For cases, using a repeated-measures linear mixed model, fat-free mass had the greatest decline over time (−2.2 kg/year, p = 2.6 × 10−18). Decline in weight (−2.2 kg/year, p = 1.2 × 10−9) and BMI (−0.73 year, p = 1.8 × 10−9) were also significant, but change in fat mass was not (0.091/year, p = 0.37). Therefore, we focused on fat-free mass for the genetic analyses. Using a linear regression model, for ACSL5 the A allele of rs58854276 was suggestively associated with lower fat-free mass at first visit in cases (−2.0 ± 1.3 kg/A allele, p = 0.14), but not in controls (−0.1 ± 1.0 kg/A allele, p = 0.89). For GPX3, the A allele of rs3828599 was not associated with lower fat-free mass at baseline visit in either cases (−1.0 ± 1.3 kg/A allele, p = 0.47) or controls (0.2 ± 1.2 kg/A allele, p = 0.89), although its effect was in the expected direction in cases. In the cases for whom longitudinal data were available, with a linear mixed model using the repeated-measures across individuals and fitting individuals as a random effect and time since first visit as a covariate, the association trend between the rs58854276 A allele and a lower fat-free mass showed a trend toward significance (−2.1 ± 1.3 kg/A allele, p = 0.053). A similar trend was observed for the rs3828599 A allele (−1.0 ± 1.3 kg/A allele, p = 0.22). Using a linear regression model, the mean change of fat-free mass between first and last visit was not associated with either rs58854276 or rs3828599 (p = 0.74 and 0.49, respectively). Recognizing that the MEND-MND sample lacks power (STAR Methods), we used the Sporadic ALS Australia (SALSA) cohort (217 cases; STAR Methods, related to the Experimental Model and Subject Details), which provided a larger sample size for BMI and weight at first visit, but not fat-free mass measurements. In this case cohort, we observed no association between the SNPs and weight (rs58854276 p = 0.97 and rs3828599 p = 0.50) or BMI (rs58854276 p = 0.47 and rs3828599 p = 0.33) at first visit. It is important to recognize that BMI and weight do not always accurately reflect changes in fat-free mass in ALS (Ioannides et al., 2017b; Kirk et al., 2019) and that in the case cohorts, individuals have weight measurements taken at cross-sectional times relative to their personal disease trajectory. Here, including time since diagnosis in the analysis did not offer further clarity.
Table 3.
Investigation of the Effect of the ACSL5 and GPX3 SNPs on Fat-Free Mass in the MEND-MND Cohorts
| Model | Sample Group | SNP/Allele | Effect (kg) | SE (kg) | p Value |
|---|---|---|---|---|---|
| Linear regression analysis at first visit | cases | rs58854276/A | −2.0 | 1.3 | 0.14 |
| Linear regression analysis at first visit | cases | rs3828599/A | −1.0 | 1.3 | 0.47 |
| Linear regression analysis at first visit | controls | rs58854276/A | −0.1 | 1.0 | 0.89 |
| Linear regression analysis at first visit | controls | rs3828599/A | 0.2 | 1.2 | 0.89 |
| Repeated-measures linear mixed model | cases | rs58854276/A | −2.1 | 1.3 | 0.053 |
| Repeated-measures linear mixed model | cases | rs3828599/A | −1.0 | 1.3 | 0.22 |
First visit refers to the time of blood sampling for controls. In all analyses, sex was used as a covariate.
Discussion
We have identified one additional ALS locus, ACSL5-ZDHHC6, and three additional putative loci, B4GALNT1, G2E3-SCFD1, and TRIP11-ATXN3, with potential functional relevance for ALS. We achieved this by exploiting the GWAS summary statistics available from previously published studies. We meta-analyzed the largest ALS study to date on >80,000 individuals of European ancestry (Nicolas et al., 2018), and a Chinese ALS study on >4,000 individuals (Benyamin et al., 2017), under the hypothesis that common causal variants are ancient and will be shared across ethnicities. The signal in ACSL5-ZDHHC6 was genome-wide significant, and the association between the lead SNP and ALS was replicated in an independent Australian cohort. B4GALNT1, G2E3-SCFD1, and TRIP11-ATXN3 SNPs did not reach genome-wide significance, but the genes achieved significance in a gene-based analysis that combines all SNP association signals within a gene. Genes associated with disease may have different architectures with respect to the number and frequencies of causal variants and therefore gene-based tests can identify disease-associated genes that have multiple causal variants of small effect that individually are not genome-wide significant (Hägg et al., 2015). Such an approach was successfully used for other complex diseases including frontotemporal dementia (Mishra et al., 2017).
ACSL5 encodes an isozyme of the long-chain fatty-acid-coenzyme A (CoA) ligase family. All isozymes of this family convert free long-chain fatty acids into fatty acyl-CoA esters and thereby play a key role in lipid biosynthesis and fatty acid degradation. This gene functions in mediating fatty-acid-induced glioma cell growth. Weight loss is frequent in ALS patients and is a strong prognostic factor associated with shorter survival (Körner et al., 2013; Desport et al., 1999). Its causes cannot be entirely explained by the ALS phenotype (Körner et al., 2013); however, loss of appetite and hypermetabolism are thought to contribute (Ngo et al., 2019; Steyn et al., 2018). Interestingly, ACSL5 SNPs and its overexpression have been associated with rapid weight loss in humans, as has another ALS-associated gene, GPX3 (Adamo et al., 2007; Teng et al., 2009; Langhardt et al., 2018). Genetic factors, such as variants in the ACSL5 and GPX3 genes, could contribute to this phenomenon in ALS patients. Our investigation of the effect of ACSL5 and GPX3 SNPs on patients’ weight in the MEND-MND cohorts (77 cases and 77 controls) showed a potential association between the ACSL5 SNP rs58854276 and a lower fat-free mass in patients (−2.1 ± 1.3 kg/A allele, p = 0.053) and no effect in controls. Furthermore, no significant evidence for association in cases or controls for GPX3 SNP rs3828599 was shown, although for cases, the effect was in the expected direction (−1.0 ± 1.3 kg/A allele, p = 0.22). No association of ACSL5 and GPX3 SNPs with lower BMI or weight was observed in the SALSA dataset. In the interpretation of these results, it is important to recognize that given the size of the effect of our SNPs on fat-free mass in the MEND-MND cohorts, the dataset provided insufficient statistical power to reliably reject the null hypothesis. Furthermore, BMI and weight do not always accurately reflect changes in fat-free mass in ALS (Ioannides et al., 2017b; Kirk et al., 2019). Considering that other ACSL5 SNPs have been previously associated with weight loss outside the context of ALS, it may be that ACSL5 SNPs have a pleiotropic effect, influencing both ALS risk and weight independently. These previously reported SNPs are in weak LD with the SNPs identified in our study (r2 < 0.20), and the effect on fat-free mass we observed in cases was not observed in controls, suggesting that they might be on the same causal pathway. However, if weight loss and ALS risk were on the same causal pathway, its direction, i.e., whether weight loss is a consequence of ALS risk or ALS risk is a consequence of weight loss, cannot be clarified with the data currently available. Larger sample sizes with genotype data and multiple measurements of body weight and composition per individual are needed to draw more robust conclusions about the relationship between these genes, weight loss, and ALS. To maximize power for a given sample size, fat-free mass should be measured as this is most affected over the life course of ALS.
ZDHHC6 encodes for a DHHC enzyme (palmitoyltransferase), which localizes to the endoplasmic reticulum (ER) and controls stability, localization, trafficking, and function of a panel of key ER substrates (Abrami et al., 2017), a function common to other ALS genes (Johnson et al., 2010; Ferrara et al., 2018). Although all the genome-wide significant SNPs were in ACSL5, these were in strong LD (r2 > 0.80) with a number of ZDHHC6 SNPs, and ZDHHC6 was itself genome-wide significant in the gene-based analysis. Our attempt to fine-map the two genes using cis-eQTL data from GTEx showed that the lead SNP was in LD with cis-eQTLs of ZDHHC6 in brain tissues, suggesting that ZDHHC6 might be the relevant disease-associated gene. We considered testing the effect of ZDHHC6 SNPs on patients’ fat-free mass. However, given the strong LD between ACSL5 and ZDHHC6 SNPs, a large proportion of the observed effect of the ACSL5 SNPs is expected to be shared by the ZDHHC6 SNPs. Because of the very limited sample size of the MEND-MND dataset, we preferred to avoid increasing the number of tests and the consequential multiple testing burden.
The B4GALNT1 gene encodes an enzyme involved in the biosynthesis of complex gangliosides (beta-1,4-N-acetylgalactosaminyl transferase). Variants in this gene cause dramatic loss of series-a and series-b gangliosides in human brain and hereditary spastic paraplegias (Boukhris et al., 2013; Harlalka et al., 2013). G2E3 is a ubiquitin ligase (E3) that regulates the DNA damage response (DDR) (Brooks et al., 2007). Numerous diseases are associated with defects in the DDR, including neurodegenerative disorders, age-related diseases, and cancer (Jackson and Bartek, 2009). Interestingly, not only did common SNPs support its association with ALS but also we reported that rare variants in G2E3 might be risk factors for ALS (p = 0.0019) using the results of our previously published gene-burden analysis of rare variants in ALS (van der Spek et al., 2019). Such results, considering the role of G2E3 in the regulation of DDR, suggest that it could play an important role in the development of ALS. SCFD1 is also involved in vesicle transport (Hou et al., 2017); we previously reported the association between ALS and SCFD1 SNPs in a European GWAS using linear mixed model analysis (van Rheenen et al., 2016) and in our recently developed machine learning method for gene discovery in ALS (Bean et al., 2020). However, our attempts in both the same GWAS (van Rheenen et al., 2016) and successive GWASs (Nicolas et al., 2018; Benyamin et al., 2017) failed to replicate it. The ATXN3 gene provides instructions for making ataxin-3, an enzyme found in cells throughout the body. Ataxin-3 is involved in the ubiquitin-proteasome system that destroys and removes excess or damaged proteins. The protein encoded by the ATXN3 gene contains CAG repeats in the coding region, and the expansion of these repeats from the normal range of 13–36 to 68–79 is the cause of Machado-Joseph disease (Kawaguchi et al., 1994), also known as SCA type 3 (SCA3). Intermediate expansions of an identical repeat within ATXN1 or ATXN2 have been associated with an increased risk of ALS (Conforti et al., 2012; Tazelaar et al., 2020). However, ATXN3 repeat expansions were not shown to have the same effect (Gispert et al., 2012). ATXN3 expression controls and is essential for the recruitment of mutated SOD1 into toxic aggresomes (Wang et al., 2012), one of the most common causes of ALS. Also, ATXN3 was predicted by our machine learning method (Bean et al., 2020). TRIP11 encodes for a protein associated with the Golgi apparatus and is involved in vesicle transport (Follit et al., 2008). Mutations in this gene cause achondrogenesis type IA (Smits et al., 2010).
STAR★Methods
Key Resources Table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Software and Algorithms | ||
| Magma toolkit | de Leeuw et al., 2015 | https://fuma.ctglab.nl, https://ctg.cncr.nl/software/magma |
| Fuma webserver | Watanabe et al., 2017 | https://fuma.ctglab.nl |
| METAL software | Willer et al., 2010 | https://genome.sph.umich.edu/wiki/METAL |
| R | The R Project for Statistical Computing | http://cran.r-project.org/mirrors.html |
| Plink 1.9 | Chang et al., 2015 | http://www.cog-genomics.org/plink/1.9 |
| LDlink | Machiela and Chanock, 2015 | https://ldlink.nci.nih.gov |
| Other | ||
| GTEx cis-eQTL data | eGTEx Project, 2017 | https://www.gtexportal.org/ |
| European ALS GWAS | Nicolas et al., 2018 | http://als.umassmed.edu |
| Chinese ALS GWAS | Benyamin et al., 2017 | https://cnsgenomics.com/ |
| Project MinE rare-variant GWAS | van der Spek et al., 2019 | http://databrowser.projectmine.com |
| 1000 Genomes phase 3 | Auton et al., 2015 | https://www.internationalgenome.org/ |
Resource Availability
Lead Contact
Further information and requests for data should be directed to and will be fulfilled by the Lead Contact, Dr. Alfredo Iacoangeli (alfredo.iacoangeli@kcl.ac.uk).
Materials Availability
This study did not generate new materials.
Data and Code Availability
The summary statistics of the final meta-analysis are available to download from the following link: https://github.com/KHP-Informatics/ALSMetaAnalysis2020.
Nicolas et al. GWAS (Nicolas et al., 2018) summary statistics can be downloaded from: http://als.umassmed.edu.
Benyamin et al. GWAS (Benyamin et al., 2017) summary statistics can be downloaded from: https://cnsgenomics.com/data/benyamin_et_al_2017_nc/BenyaminEtAl_NatComm_Data.zip.
Van Der Spek et al. GWAS (van der Spek et al., 2019) summary statistics can be downloaded from: http://databrowser.projectmine.com. Cis-eQTL data from GTEx version 8 is available on the consortium website: https://www.gtexportal.org/.
Experimental Model and Subject Details
Human subjects
Discovery datasets and cohorts
All samples and datasets involved in our discovery GWAS analyses were described previously (van Rheenen et al., 2016, Benyamin et al., 2017, Nicolas et al., 2018, van der Spek et al., 2019).
The Australian replication cohort
The Australian ALS cohort consisted of 837 cases and 665 controls of European ancestry (Table S1). Participants were genotyped on the Infinium CoreExome-24 v1.1 chip, and after standard QC steps, imputed to the Haplotype Reference Consortium (HRC) panel. The sample included patients and controls ascertained from the University of Sydney as part of the Australian MND DNA bank, which recruited participants from April 2000 to June 2011. Cases were white Australians older than 25 years recruited across Australia via state-based MND associations with diagnoses verified by neurologists. The study protocol was approved by the Sydney South West Area Health Service Human Research Ethics Committee (HREC). Other cases were recruited from clinics across Australia between 2015 and 2017. Control subjects were healthy individuals free of neuromuscular diseases, recruited as either partners or friends of patients with ALS or community volunteers. Written consent was obtained from all individuals enrolled in this study, and the study was approved by the corresponding HREC at the different sites: University of Sydney, Western Sydney Local Health District, The Royal Brisbane and Women Hospital (Metro North), South Metropolitan Health Service, and Macquarie University. All ALS cases were diagnosed with definite or probable ALS according to the revised El Escorial criteria. Those with a recorded family history of ALS were excluded. Additional controls were contributed from the Older Australian Twin Study(Sachdev et al., 2009) (OATS) comprising 90 monozygotic (MZ) twin pairs recruited at QIMR Berghofer Medical Research Institute, University of New South Wales and the University of Melbourne, with the studies being approved by their respective HRECs. The OATS study recruited MZ twins aged 65 years and over. Twin pair data helped in quality control checks but only one twin from each pair was used in our analyses.
The MEND-MND and the SALSA samples
Genotype data and weight-related measures were available in two additional Australian cohorts. These two datasets were used only to investigate the difference in body weight and composition between cases and controls, and the effect of the ACSL5 and GPX3 genotype on these measures. The MEND-MND cohort (Ioannides et al., 2017a) (Table S4) consisted of 77 ALS cases and 77 controls of European ancestry. For these samples, body weight, BMI, fat-free mass, and fat mass at first visit were available. Follow up data for the MEND-MND cohort was available for 67 cases. For the 77 cases, a total of 320 measurements per measure were obtained, with the mean time between follow up visits equal to 4.1 months (SD = 1.5 months), and the mean number of visits per participants equal to 4.2 (SD = 2.5). Fat-free mass and fat mass were determined by air displacement plethysmography using the BodPod system (Cosmed) (Dempster and Aitkens, 1995, Ioannides et al., 2017a). BMI was defined as body weight divided by the square of patient height (kg/m2). The SALSA cohort (Table S5) provided 217 European ancestry cases for whom genotypes, BMI and weight measures at first visit were available. The cases were independent of the MEND-MND cases and included 62 cases from the 837 cases used for the SNP association replication. SALSA cases were recruited from clinics across Australia, including those listed above. Additional sites included Calvary Health Care Bethlehem (Melbourne) and the Fiona Stanley Hospital (Perth). All SALSA participants were diagnosed with definite or probable ALS, according to the revised El Escorial criteria. In all cohorts, those with a family history of ALS, or those who had been tested positive for known SOD1 or C9orf72 mutations were excluded.
Method Details
Fine-mapping using GTEx eQTL data
To assess whether the SNPs that we identified to be associated with the risk of ALS modify gene expression, we used the cis-eQTL data from GTEx version 8 for brain and blood tissues (e, 2017). For the seven genes identified in our meta-analysis, ACSL5, ZDHHC6, SCFD1, G2E3, TRIP11, ATXN3 and B4GALNT1, we selected their independent lead SNPs (r2 < 0.80). Corresponding matching SNPs were extracted from the GTEx data for brain and blood tissues. If no matching SNP was available, we selected proxies in LD (r2 > 0.50).
Genome-wide meta-analysis
Inverse variance SNP-based meta-analysis was conducted using METAL (Willer et al., 2010). The SNP genome-wide p-value significance threshold was 5x10−8. Putative ALS genes were defined as those for which at least one SNP p-value was < 5x10−7 and genes that were genome-wide significant in the gene-based analysis. The gene-based association study was performed with the Magma ‘SNPtoGENE’ protocol (de Leeuw et al., 2015) on the FUMA webserver (Watanabe et al., 2017) to assess the overall association between all SNPs in a gene and a given phenotype. The SNPs were mapped to 18,067 protein coding genes. Therefore genome-wide significance was defined according to the conservative Bonferroni correction method at p-value = 0.05/18,067 = 2.8x10−6. We also investigated the association of rare variants (minor allele frequency < 0.01) in our candidate genes with ALS risk, using the summary statistics from a gene-burden analysis of disruptive, damaging and missense variants previously performed on 4,389 ALS patients and 1,846 controls (van der Spek et al., 2019). We used the Bonferroni correction based on the number of genes tested to assess significance (i.e., p = 0.05/7 = 0.0071). All annotation, genomic positions and variants refer to the reference human genome hg19/GRCh37. The 1000 genomes project Phase 3 Reference panel was used to compute r2 and minor allele frequency (Auton et al., 2015) with Plink 1.9 (Chang et al., 2015) and LDlink (Machiela and Chanock, 2015).
Quantification and Statistical Analysis
Body weight and composition analyses in the MEND-MND and SALSA datasets
We investigated body weight and composition differences between cases and controls and the effect of the ACSL5 and GPX3 SNPs on the individuals’ body weight and composition in an additive model. For the MEND-MND cohort, using a linear regression model (Chambers, 1992), we first tested the differences at baseline (first visit) in body weight and composition measures between cases and controls and within cases and controls with different ACSL5 and GPX3 SNP alleles. Second, we tested the difference in terms of the mean decline over time (change between first visit and last visit divided by time between first and last visit) in cases with different ACSL5 and GPX3 SNP alleles. Using a repeated-measures linear mixed model (Bates et al., 2014) for cases, we assessed the effect of time on body weight and composition, and the effect of our selected SNPs on patients’ fat-free mass. This uses the repeated-measures nature of the data to improve precision of the association of SNP with our measures. Taking fat-free mass as an example, we regressed fat-free mass on days since first visit, fitting individuals as a random effect. The 217 samples of the SALSA cohort had weight and BMI at first visit available. We used a linear regression model to test the difference of BMI and weight in patients with different ACSL5 and GPX3 SNP alleles. Sex was fitted as a covariate in all analyses. We used the lm R function for the linear regression model and the lmer function from the lme4 R package (Bates et al., 2014) for the repeater-measures linear mixed model. Power calculation was performed using the Genetic Power Calculator (Purcell et al., 2003) with type I error rate equal to 0.05 and the allele frequencies of rs58854276 and rs3828599 A alleles from the 1000 genomes project phase 3. Using these parameters, a dataset of 77 individuals provides 80% power to detect an effect > = 5.9 kgs of fat-free mass per allele in a linear additive model.
Acknowledgments
This is an EU Joint Programme-Neurodegenerative Disease Research (JPND) project. The project is supported through the following funding organizations under the aegis of JPND-http://www.neurodegenerationresearch.eu/ (United Kingdom, Medical Research Council MR/L501529/1 to A.A.-C., principal investigator [PI] and MR/R024804/1 to A.A.-C., PI]; Economic and Social Research Council ES/L008238/1 to A.A.-C. [co-PI]) and through the Motor Neurone Disease Association. This study represents independent research partly funded by the National Institute for Health Research (NIHR) Biomedical Research Centre at South London and Maudsley NHS Foundation Trust and King’s College London. The work leading up to this publication was funded by the European Community’s Horizon 2020 Programme (H2020-PHC-2014-two-stage; grant 633413). Sequence data used in this research were, in part, obtained from the UK National DNA Bank for MND Research, funded by the MND Association and the Wellcome Trust. N.R.W. acknowledges funding from the National Health and Medical Research Council (NHMRC) (1078901, 1113400, and 1151854) and the Motor Neurone Disease Research Institute of Australia Ice Bucket Challenge Grant. We acknowledge the Motor Neurone Disease Research Institute of Australia Cunningham Collaboration MND Research Grant (to P.A.M., R.H., and S.T.N.) and the Cunningham Family MND Research Grant (to F.J.S., P.A.M., R.H., and S.T.N.). The Older Australian Twin Study was facilitated through Twins Research Australia, a national resource supported in part by a Centre for Research Excellence from the Australian NHMRC. Funding for this study was awarded by the NHMRC/Australian Research Council Strategic Award (grant 401162) and the NHMRC project grant 1405325. S.T.N. is supported by the Scott Sullivan Fellowship (MND and Me Foundation, Royal Brisbane and Women's Hospital Foundation, and the Queensland Brain Institute) and the Australian Institute for Bioengineering and Nanotechnology at the University of Queensland. We acknowledge use of the research computing facility at King’s College London, Rosalind (https://rosalind.kcl.ac.uk), which is delivered in partnership with the National Institute for Health Research (NIHR) Biomedical Research Centres at South London & Maudsley and Guy’s & St. Thomas’ NHS Foundation Trusts and part-funded by capital equipment grants from the Maudsley Charity (award 980) and Guy's and St Thomas' Charity (TR130505). The views expressed are those of the author(s) and not necessarily those of the NHS, the NIHR, King’s College London, or the Department of Health and Social Care. We thank Professor Nigel Laing for his contribution in providing DNA samples from the Western Australia cohort. We thank all of the people who participated in this project, including those with MND, their families, and those who served as controls.
Author Contributions
Analysis of the data, A.I. and T.L.; Conception of the study and writing the manuscript, A.I.; Study design and interpretation of data, A.I., T.L., S.T.N., F.J.S., N.R.W., and A.A.-C.; Acquisition of data, A.I., A.A.K., T.L., S.T.N., and F.J.S.; Revising the article, all authors; Reading and approving the final manuscript, all authors.
Declaration of Interests
The authors declare no competing interests.
Published: October 27, 2020
Footnotes
Supplemental Information can be found online at https://doi.org/10.1016/j.celrep.2020.108323.
Supplemental Information
References
- Abrami L., Dallavilla T., Sandoz P.A., Demir M., Kunz B., Savoglidis G., Hatzimanikatis V., van der Goot F.G. Identification and dynamics of the human ZDHHC16-ZDHHC6 palmitoylation cascade. eLife. 2017;6:e27826. doi: 10.7554/eLife.27826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Adamo K.B., Dent R., Langefeld C.D., Cox M., Williams K., Carrick K.M., Stuart J.S., Sundseth S.S., Harper M.E., McPherson R., Tesson F. Peroxisome proliferator-activated receptor γ 2 and acyl-CoA synthetase 5 polymorphisms influence diet response. Obesity (Silver Spring) 2007;15:1068–1075. doi: 10.1038/oby.2007.630. [DOI] [PubMed] [Google Scholar]
- Al-Chalabi A. Perspective: Don’t keep it in the family. Nature. 2017;550:S112. doi: 10.1038/550S112a. [DOI] [PubMed] [Google Scholar]
- Auton A., Brooks L.D., Durbin R.M., Garrison E.P., Kang H.M., Korbel J.O., Marchini J.L., McCarthy S., McVean G.A., Abecasis G.R., 1000 Genomes Project Consortium A global reference for human genetic variation. Nature. 2015;526:68–74. doi: 10.1038/nature15393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bates D., Mächler M., Bolker B., Walker S. Fitting linear mixed-effects models using lme4. arXiv. 2014;1406:5823. https://arxiv.org/abs/1406.5823 [Google Scholar]
- Bean D.M., Al-Chalabi A., Dobson R.J.B., Iacoangeli A. A Knowledge-Based Machine Learning Approach to Gene Prioritisation in Amyotrophic Lateral Sclerosis. Genes (Basel) 2020;11:668. doi: 10.3390/genes11060668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benyamin B., He J., Zhao Q., Gratten J., Garton F., Leo P.J., Liu Z., Mangelsdorf M., Al-Chalabi A., Anderson L. Cross-ethnic meta-analysis identifies association of the GPX3-TNIP1 locus with amyotrophic lateral sclerosis. Nat. Commun. 2017;8:611. doi: 10.1038/s41467-017-00471-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boukhris A., Schule R., Loureiro J.L., Lourenço C.M., Mundwiller E., Gonzalez M.A., Charles P., Gauthier J., Rekik I., Acosta Lebrigio R.F. Alteration of ganglioside biosynthesis responsible for complex hereditary spastic paraplegia. Am. J. Hum. Genet. 2013;93:118–123. doi: 10.1016/j.ajhg.2013.05.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brooks W.S., Banerjee S., Crawford D.F. G2E3 is a nucleo-cytoplasmic shuttling protein with DNA damage responsive localization. Exp. Cell Res. 2007;313:665–676. doi: 10.1016/j.yexcr.2006.11.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown R.H., Al-Chalabi A. Amyotrophic lateral sclerosis. N. Engl. J. Med. 2017;377:162–172. doi: 10.1056/NEJMra1603471. [DOI] [PubMed] [Google Scholar]
- Chambers J. Statistical Models in S, J. Chambers and T.J. Hastie. Wadsworth & Brooks/Cole Advanced Books & Software; 1992. Linear models; pp. 95–144. [Google Scholar]
- Chang C.C., Chow C.C., Tellier L.C., Vattikuti S., Purcell S.M., Lee J.J. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience. 2015;4 doi: 10.1186/s13742-015-0047-8. s13742-015-0047-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chia R., Chiò A., Traynor B.J. Novel genes associated with amyotrophic lateral sclerosis: diagnostic and clinical implications. Lancet Neurol. 2018;17:94–102. doi: 10.1016/S1474-4422(17)30401-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Conforti F.L., Spataro R., Sproviero W., Mazzei R., Cavalcanti F., Condino F., Simone I.L., Logroscino G., Patitucci A., Magariello A. Ataxin-1 and ataxin-2 intermediate-length PolyQ expansions in amyotrophic lateral sclerosis. Neurology. 2012;79:2315–2320. doi: 10.1212/WNL.0b013e318278b618. [DOI] [PubMed] [Google Scholar]
- de Leeuw C.A., Mooij J.M., Heskes T., Posthuma D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput. Biol. 2015;11:e1004219. doi: 10.1371/journal.pcbi.1004219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dempster P., Aitkens S. A new air displacement method for the determination of human body composition. Med. Sci. Sports Exerc. 1995;27:1692–1697. [PubMed] [Google Scholar]
- Desport J.C., Preux P.M., Truong T.C., Vallat J.M., Sautereau D., Couratier P. Nutritional status is a prognostic factor for survival in ALS patients. Neurology. 1999;53:1059–1063. doi: 10.1212/wnl.53.5.1059. [DOI] [PubMed] [Google Scholar]
- eGTEx Project Enhancing GTEx by bridging the gaps between genotype, gene expression, and disease. Nat. Genet. 2017;49:1664–1670. doi: 10.1038/ng.3969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ferrara D., Pasetto L., Bonetto V., Basso M. Role of Extracellular Vesicles in Amyotrophic Lateral Sclerosis. Front. Neurosci. 2018;12:574. doi: 10.3389/fnins.2018.00574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fogh I., Ratti A., Gellera C., Lin K., Tiloca C., Moskvina V., Corrado L., Sorarù G., Cereda C., Corti S., SLAGEN Consortium and Collaborators A genome-wide association meta-analysis identifies a novel locus at 17q11.2 associated with sporadic amyotrophic lateral sclerosis. Hum. Mol. Genet. 2014;23:2220–2231. doi: 10.1093/hmg/ddt587. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Follit J.A., San Agustin J.T., Xu F., Jonassen J.A., Samtani R., Lo C.W., Pazour G.J. The Golgin GMAP210/TRIP11 anchors IFT20 to the Golgi complex. PLoS Genet. 2008;4:e1000315. doi: 10.1371/journal.pgen.1000315. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gispert S., Kurz A., Waibel S., Bauer P., Liepelt I., Geisen C., Gitler A.D., Becker T., Weber M., Berg D. The modulation of Amyotrophic Lateral Sclerosis risk by ataxin-2 intermediate polyglutamine expansions is a specific effect. Neurobiol. Dis. 2012;45:356–361. doi: 10.1016/j.nbd.2011.08.021. [DOI] [PubMed] [Google Scholar]
- Hägg S., Ganna A., Van Der Laan S.W., Esko T., Pers T.H., Locke A.E., Berndt S.I., Justice A.E., Kahali B., Siemelink M.A., GIANT Consortium Gene-based meta-analysis of genome-wide association studies implicates new loci involved in obesity. Hum. Mol. Genet. 2015;24:6849–6860. doi: 10.1093/hmg/ddv379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harlalka G.V., Lehman A., Chioza B., Baple E.L., Maroofian R., Cross H., Sreekantan-Nair A., Priestman D.A., Al-Turki S., McEntagart M.E. Mutations in B4GALNT1 (GM2 synthase) underlie a new disorder of ganglioside biosynthesis. Brain. 2013;136:3618–3624. doi: 10.1093/brain/awt270. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hou N., Yang Y., Scott I.C., Lou X. The Sec domain protein Scfd1 facilitates trafficking of ECM components during chondrogenesis. Dev. Biol. 2017;421:8–15. doi: 10.1016/j.ydbio.2016.11.010. [DOI] [PubMed] [Google Scholar]
- Iacoangeli A., Al Khleifat A., Jones A.R., Sproviero W., Shatunov A., Opie-Martin S., Morrison K.E., Shaw P.J., Shaw C.E., Fogh I., Alzheimer’s Disease Neuroimaging Initiative C9orf72 intermediate expansions of 24-30 repeats are associated with ALS. Acta Neuropathol. Commun. 2019;7:115. doi: 10.1186/s40478-019-0724-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iacoangeli A., Al Khleifat A., Sproviero W., Shatunov A., Jones A.R., Opie-Martin S., Naselli E., Topp S.D., Fogh I., Hodges A. ALSgeneScanner: a pipeline for the analysis and interpretation of DNA sequencing data of ALS patients. Amyotroph. Lateral Scler. Frontotemporal Degener. 2019;20:207–215. doi: 10.1080/21678421.2018.1562553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ioannides Z., Steyn F., Mi J., Henderson R., Mccombe P., Ngo S. Predictions of resting energy expenditure in amyotrophic lateral sclerosis are greatly impacted by reductions in fat free mass. Cogent. Med. 2017;4:1343000. [Google Scholar]
- Ioannides Z.A., Steyn F.J., Henderson R.D., Mccombe P.A., Ngo S.T. Anthropometric measures are not accurate predictors of fat mass in ALS. Amyotroph. Lateral Scler. Frontotemporal Degener. 2017;18:486–491. doi: 10.1080/21678421.2017.1317811. [DOI] [PubMed] [Google Scholar]
- Jackson S.P., Bartek J. The DNA-damage response in human biology and disease. Nature. 2009;461:1071–1078. doi: 10.1038/nature08467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson J.O., Mandrioli J., Benatar M., Abramzon Y., Van Deerlin V.M., Trojanowski J.Q., Gibbs J.R., Brunetti M., Gronka S., Wuu J., ITALSGEN Consortium Exome sequencing reveals VCP mutations as a cause of familial ALS. Neuron. 2010;68:857–864. doi: 10.1016/j.neuron.2010.11.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kawaguchi Y., Okamoto T., Taniwaki M., Aizawa M., Inoue M., Katayama S., Kawakami H., Nakamura S., Nishimura M., Akiguchi I. CAG expansions in a novel gene for Machado-Joseph disease at chromosome 14q32.1. Nat. Genet. 1994;8:221–228. doi: 10.1038/ng1194-221. [DOI] [PubMed] [Google Scholar]
- Kirk S.E., Tracey T.J., Steyn F.J., Ngo S.T. Biomarkers of metabolism in amyotrophic lateral sclerosis. Front. Neurol. 2019;10:191. doi: 10.3389/fneur.2019.00191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Körner S., Hendricks M., Kollewe K., Zapf A., Dengler R., Silani V., Petri S. Weight loss, dysphagia and supplement intake in patients with amyotrophic lateral sclerosis (ALS): impact on quality of life and therapeutic options. BMC Neurol. 2013;13:84. doi: 10.1186/1471-2377-13-84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Langhardt J., Flehmig G., Klöting N., Lehmann S., Ebert T., Kern M., Schön M.R., Gärtner D., Lohmann T., Dressler M. Effects of Weight Loss on Glutathione Peroxidase 3 Serum Concentrations and Adipose Tissue Expression in Human Obesity. Obes. Facts. 2018;11:475–490. doi: 10.1159/000494295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Machiela M.J., Chanock S.J. LDlink: a web-based application for exploring population-specific haplotype structure and linking correlated alleles of possible functional variants. Bioinformatics. 2015;31:3555–3557. doi: 10.1093/bioinformatics/btv402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mishra A., Ferrari R., Heutink P., Hardy J., Pijnenburg Y., Posthuma D., International FTD-Genomics Consortium Gene-based association studies report genetic links for clinical subtypes of frontotemporal dementia. Brain. 2017;140:1437–1446. doi: 10.1093/brain/awx066. [DOI] [PubMed] [Google Scholar]
- Morita M., Al-Chalabi A., Andersen P.M., Hosler B., Sapp P., Englund E., Mitchell J.E., Habgood J.J., de Belleroche J., Xi J. A locus on chromosome 9p confers susceptibility to ALS and frontotemporal dementia. Neurology. 2006;66:839–844. doi: 10.1212/01.wnl.0000200048.53766.b4. [DOI] [PubMed] [Google Scholar]
- Ngo S.T., van Eijk R.P.A., Chachay V., van den Berg L.H., McCombe P.A., Henderson R.D., Steyn F.J. Loss of appetite is associated with a loss of weight and fat mass in patients with amyotrophic lateral sclerosis. Amyotroph. Lateral Scler. Frontotemporal Degener. 2019;20:497–505. doi: 10.1080/21678421.2019.1621346. [DOI] [PubMed] [Google Scholar]
- Nicolas A., Kenna K.P., Renton A.E., Ticozzi N., Faghri F., Chia R., Dominov J.A., Kenna B.J., Nalls M.A., Keagle P. Genome-wide analyses identify KIF5A as a novel ALS gene. Neuron. 2018;97:1268–1283. doi: 10.1016/j.neuron.2018.02.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Purcell S., Cherny S.S., Sham P.C. Genetic Power Calculator: design of linkage and association genetic mapping studies of complex traits. Bioinformatics. 2003;19:149–150. doi: 10.1093/bioinformatics/19.1.149. [DOI] [PubMed] [Google Scholar]
- Renton A.E., Chiò A., Traynor B.J. State of play in amyotrophic lateral sclerosis genetics. Nat. Neurosci. 2014;17:17–23. doi: 10.1038/nn.3584. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sachdev P.S., Lammel A., Trollor J.N., Lee T., Wright M.J., Ames D., Wen W., Martin N.G., Brodaty H., Schofield P.R., OATS research team A comprehensive neuropsychiatric study of elderly twins: the Older Australian Twins Study. Twin Res. Hum. Genet. 2009;12:573–582. doi: 10.1375/twin.12.6.573. [DOI] [PubMed] [Google Scholar]
- Shatunov A., Mok K., Newhouse S., Weale M.E., Smith B., Vance C., Johnson L., Veldink J.H., van Es M.A., van den Berg L.H. Chromosome 9p21 in sporadic amyotrophic lateral sclerosis in the UK and seven other countries: a genome-wide association study. Lancet Neurol. 2010;9:986–994. doi: 10.1016/S1474-4422(10)70197-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smits P., Bolton A.D., Funari V., Hong M., Boyden E.D., Lu L., Manning D.K., Dwyer N.D., Moran J.L., Prysak M. Lethal skeletal dysplasia in mice and humans lacking the golgin GMAP-210. N. Engl. J. Med. 2010;362:206–216. doi: 10.1056/NEJMoa0900158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steyn F.J., Ioannides Z.A., van Eijk R.P.A., Heggie S., Thorpe K.A., Ceslis A., Heshmat S., Henders A.K., Wray N.R., van den Berg L.H. Hypermetabolism in ALS is associated with greater functional decline and shorter survival. J. Neurol. Neurosurg. Psychiatry. 2018;89:1016–1023. doi: 10.1136/jnnp-2017-317887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tazelaar G.H.P., Boeynaems S., De Decker M., van Vugt J.J.F.A., Kool L., Goedee H.S., McLaughlin R.L., Sproviero W., Iacoangeli A., Moisse M., Project MinE ALS Sequencing Consortium ATXN1 repeat expansions confer risk for amyotrophic lateral sclerosis and contribute to TDP-43 mislocalization. Brain Commun. 2020;2:a064. doi: 10.1093/braincomms/fcaa064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Teng A.C., Adamo K., Tesson F., Stewart A.F. Functional characterization of a promoter polymorphism that drives ACSL5 gene expression in skeletal muscle and associates with diet-induced weight loss. FASEB J. 2009;23:1705–1709. doi: 10.1096/fj.08-120998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van der Spek R.A.A., van Rheenen W., Pulit S.L., Kenna K.P., van den Berg L.H., Veldink J.H., Project MinE ALS Sequencing Consortium The project MinE databrowser: bringing large-scale whole-genome sequencing in ALS to researchers and the public. Amyotroph. Lateral Scler. Frontotemporal Degener. 2019;20:432–440. doi: 10.1080/21678421.2019.1606244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Rheenen W., Shatunov A., Dekker A.M., McLaughlin R.L., Diekstra F.P., Pulit S.L., van der Spek R.A., Võsa U., de Jong S., Robinson M.R., PARALS Registry; SLALOM Group; SLAP Registry; FALS Sequencing Consortium; SLAGEN Consortium; NNIPPS Study Group Genome-wide association analyses identify new risk variants and the genetic architecture of amyotrophic lateral sclerosis. Nat. Genet. 2016;48:1043–1048. doi: 10.1038/ng.3622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vance C., Al-Chalabi A., Ruddy D., Smith B.N., Hu X., Sreedharan J., Siddique T., Schelhaas H.J., Kusters B., Troost D. Familial amyotrophic lateral sclerosis with frontotemporal dementia is linked to a locus on chromosome 9p13.2-21.3. Brain. 2006;129:868–876. doi: 10.1093/brain/awl030. [DOI] [PubMed] [Google Scholar]
- Wang H., Ying Z., Wang G. Ataxin-3 regulates aggresome formation of copper-zinc superoxide dismutase (SOD1) by editing K63-linked polyubiquitin chains. J. Biol. Chem. 2012;287:28576–28585. doi: 10.1074/jbc.M111.299990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Watanabe K., Taskesen E., van Bochoven A., Posthuma D. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun. 2017;8:1826. doi: 10.1038/s41467-017-01261-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Willer C.J., Li Y., Abecasis G.R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics. 2010;26:2190–2191. doi: 10.1093/bioinformatics/btq340. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The summary statistics of the final meta-analysis are available to download from the following link: https://github.com/KHP-Informatics/ALSMetaAnalysis2020.
Nicolas et al. GWAS (Nicolas et al., 2018) summary statistics can be downloaded from: http://als.umassmed.edu.
Benyamin et al. GWAS (Benyamin et al., 2017) summary statistics can be downloaded from: https://cnsgenomics.com/data/benyamin_et_al_2017_nc/BenyaminEtAl_NatComm_Data.zip.
Van Der Spek et al. GWAS (van der Spek et al., 2019) summary statistics can be downloaded from: http://databrowser.projectmine.com. Cis-eQTL data from GTEx version 8 is available on the consortium website: https://www.gtexportal.org/.