Abstract
Chromosomal instability in cancer consists of dynamic changes to the number and structure of chromosomes1,2. The resulting diversity in somatic copy number alterations (SCNAs) may provide the variation necessary for tumour evolution1,3,4. Here we use multi-sample phasing and SCNA analysis of 1,421 samples from 394 tumours across 22 tumour types to show that continuous chromosomal instability results in pervasive SCNA heterogeneity. Parallel evolutionary events, which cause disruption in the same genes (such as BCL9, MCL1, ARNT (also known as HIF1B), TERT and MYC) within separate subclones, were present in 37% of tumours. Most recurrent losses probably occurred before whole-genome doubling, that was found as a clonal event in 49% of tumours. However, loss of heterozygosity at the human leukocyte antigen (HLA) locus and loss of chromosome 8p to a single haploid copy recurred at substantial subclonal frequencies, even in tumours with whole-genome doubling, indicating ongoing karyotype remodelling. Focal amplifications that affected chromosomes 1q21 (which encompasses BCL9, MCL1 and ARNT), 5p15.33 (TERT), 11q13.3 (CCND1), 19q12 (CCNE1) and 8q24.1 (MYC) were frequently subclonal yet appeared to be clonal within single samples. Analysis of an independent series of 1,024 metastatic samples revealed that 13 focal SCNAs were enriched in metastatic samples, including gains in chromosome 8q24.1 (encompassing MYC) in clear cell renal cell carcinoma and chromosome 11q13.3 (encompassing CCND1) in HER2+ breast cancer. Chromosomal instability may enable the continuous selection of SCNAs, which are established as ordered events that often occur in parallel, throughout tumour evolution.
Chromosomal instability (CIN) results from the occurrence and tolerance of chromosome segregation errors during cell division. CIN has been linked to poor prognosis5,6,7,8,9 and leads to SCNAs that may act as a substrate for selection1,3,4.
However, the prevalence of ongoing CIN later in tumour evolution2 and the temporal order of clonal and subclonal SCNAs in relation to whole-genome doubling (WGD) events and metastatic dissemination remain unclear.
Pan-cancer ongoing CIN and SCNA heterogeneity
We applied a multi-sample phasing SCNA analysis method (Extended Data Fig. 1a–c and Methods) to 1,421 cancer samples from 394 patients across 22 tumour subtypes (median 3 samples per tumour; range 2–16 samples per tumour) (Extended Data Fig. 1d, e and Supplementary Table 1), to obtain SCNA heterogeneity at haplotype resolution. We used MEDICC10 to estimate the copy number states of the most-recent common ancestor (MRCA) of each tumour, which reflects the SCNAs that were acquired before subclonal diversification. In our analysis, 1,019 out of 1,421 samples were from primary tumours, 32 were from post-treatment primary tumours, 7 samples were obtained after local relapse and 363 samples were of metastatic origin. In each case, there were at least two samples per tumour and 152 tumours had at least one primary and at least one metastatic sample.
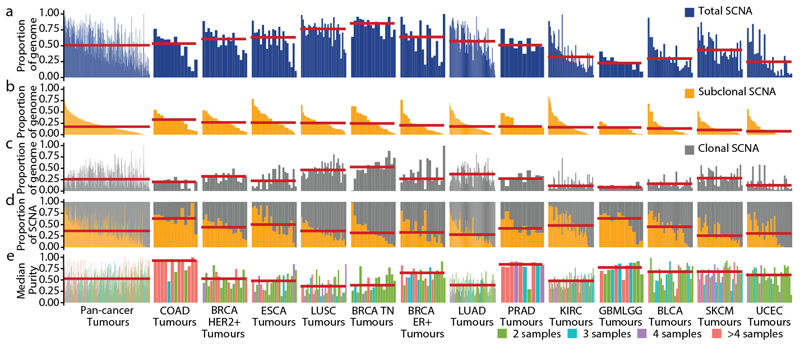
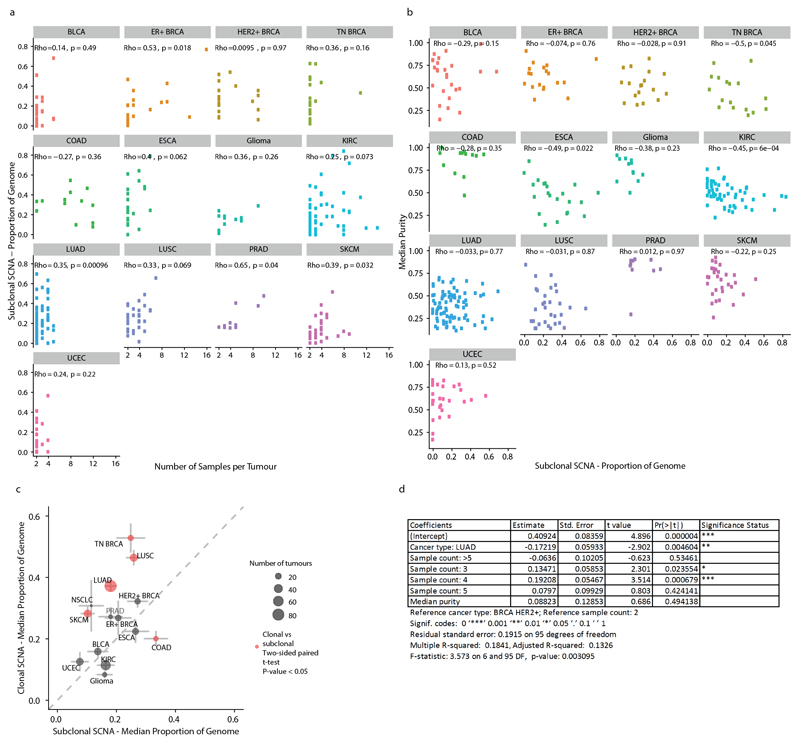
To explore CIN during cancer evolution, we quantified the total proportion of the genome affected by SCNAs and the proportion of clonal, early SCNAs, compared with subclonal, late SCNAs (Fig. 1a–d). We identified clonal SCNAs in every tumour (Fig. 1c) and found that 99% of tumours (390 out of 394) had at least one subclonal SCNA (Fig. 1b). A median of 26% of the genome was subject to clonal SCNAs and 18% to subclonal SCNAs. In 45% of tumours, more than 20% of the genome was subject to subclonal SCNAs, which highlights that ongoing CIN is pervasive. However, this is probably an underestimate of CIN as only a small proportion of each tumour is sequenced. Consistent with this, we observed a significant correlation between the number of samples per tumour and SCNA heterogeneity (Extended Data Fig. 2a). Analysis of triple-negative breast cancer, oesophageal adenocarcinoma and clear cell renal cell carcinoma showed a significant association between median purity (Fig. 1e) and the proportion of the genome that is affected by subclonal SCNAs in these tumour types (Extended Data Fig. 2b), indicating that tumour purity may interfere with the estimation of SCNA clonality.
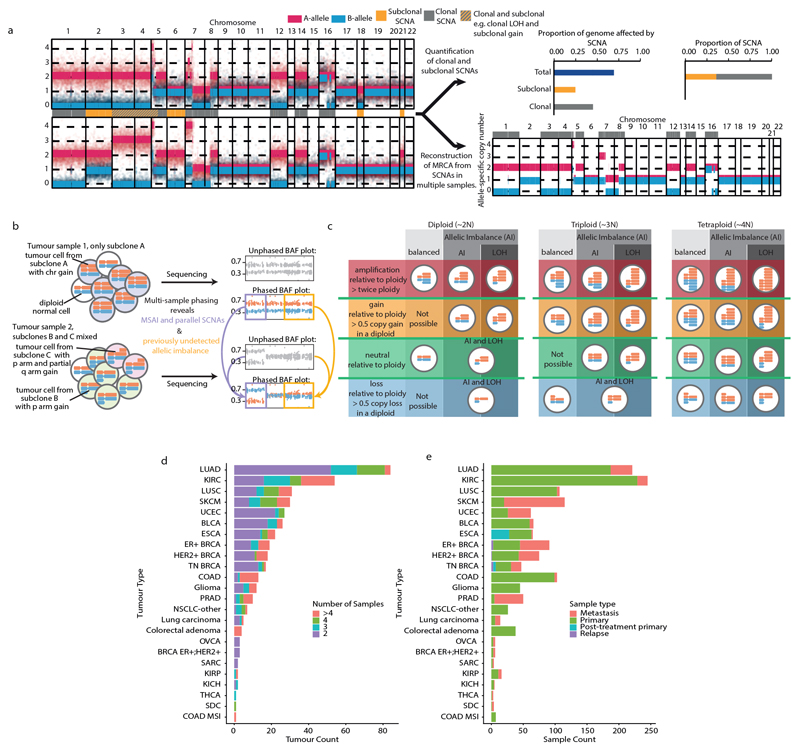
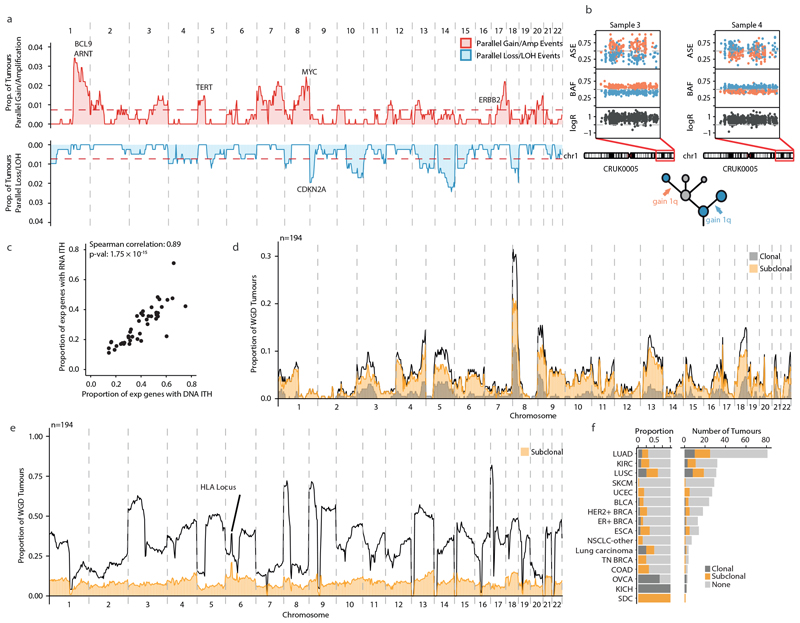
Fig. 1. Overview of somatic copy number heterogeneity across tumour types.
a, For each tumour, the proportion of the genome that is affected by SCNAs (both clonal and subclonal) is indicated. Tumour types with tumour samples from at least 10 patients were included: colorectal adenocarcinoma (COAD, n = 13), HER2+ breast cancer (HER2+ BRCA, n = 18), oesophageal adenocarcinoma (ESCA, n = 22), lung squamous cell carcinoma (LUSC,n = 31), triple-negative breast cancer (TN BRCA, n = 17), ER+ breast cancer (ER+ BRCA, n = 19), lung adenocarcinoma (LUAD, n = 84), prostate adenocarcinoma (PRAD, n = 10), clear cell renal cell carcinoma (KIRC, n = 54), glioma (n = 12), bladder urothelial carcinoma (BLCA, n = 26), melanoma (SKCM, n = 30) and endometrial carcinoma (UCEC, n = 27). Tumour types and tumours are ordered by the median proportion of the genome that is affected by subclonal SCNA—this order is maintained throughout the figure. Red lines indicate the median of the distribution. b, c, The proportion of the genome affected by subclonal (b) and clonal (c) SCNAs. d, The proportions of SCNAs that are subclonal and clonal are shown. The red line indicates the median proportion of SCNAs that are subclonal. e, The median purity and number of samples for each tumour.
The timing of SCNAs varied across tumour types (Fig. 1a–c and Extended Data Fig. 2c). Despite a comparable total proportion of the genome affected by SCNAs between lung adenocarcinoma (LUAD) and HER2+ breast cancer (57% compared with 58%, respectively; P = 0.81, effect size = 0.05), in LUADs a larger proportion of SCNAs were clonal, whereas HER2+ breast cancers showed a higher proportion of subclonal SCNAs (28% and 44% in LUAD and HER2+ breast cancer, respectively; P = 8.1 × 10−3, effect size = 0.59; the analysis was also controlled for sample number) (Extended Data Fig. 2d).
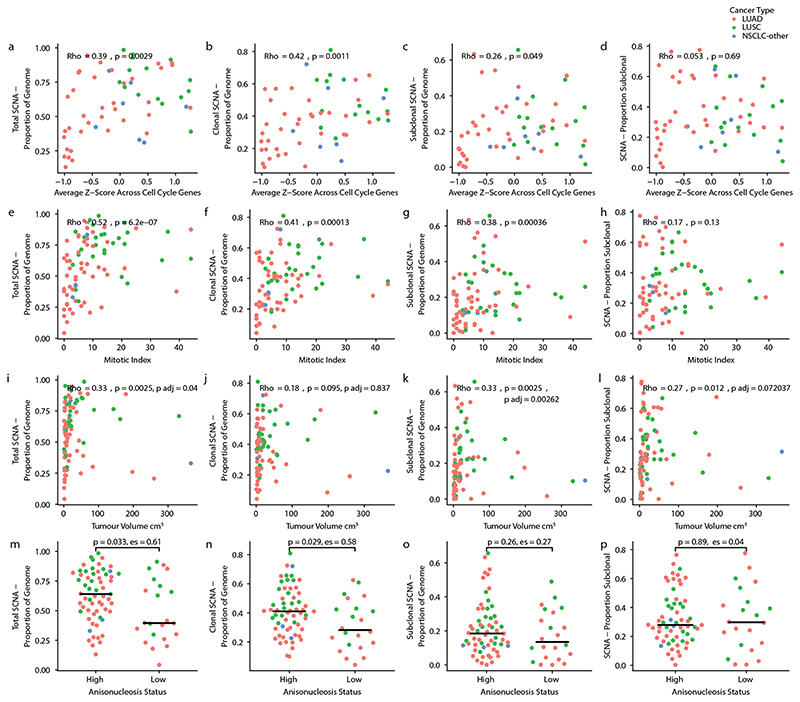
Consistent with increased proliferation in CIN tumours, the total, clonal and subclonal SCNA burden correlated with both increased cell cycle gene expression in 58 non-small cell lung cancers (NSCLCs) for which RNA sequencing data were available and with an increased mitotic index score in 83 NSCLCs for which digitized diagnostic slides were available (Methods, Extended Data Fig. 3a–h, Supplementary Table 2). Furthermore, in the 83 NSCLCs with mitotic index scores, the estimates of tumour volume, which were derived from preoperative computed tomography scans, were found to correlate with the total and subclonal proportion of the genome affected by SCNAs, and these associations remained significant when controlling for sample number (Extended Data Fig. 3i–l). Finally, anisonucleosis—a measure of variation in the size of the nucleus (Methods) that is prognostic in NSCLC11,12—was associated with increased total and clonal SCNA burden, but not with subclonal SCNA burden (Extended Data Fig. 3m–p and Supplementary Table 2).
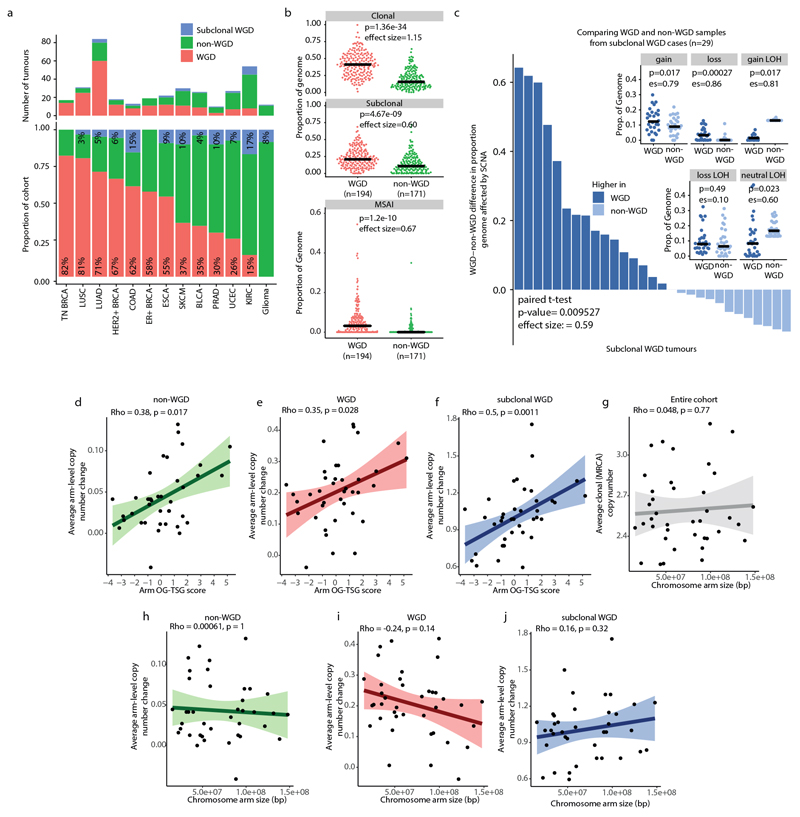
In total, 57% of tumours exhibited WGD (Methods), which occurred as a clonal event in 87% of these tumours (Extended Data Fig. 4a). WGD was associated with an increased burden of clonal and subclonal SCNAs compared with non-WGD tumours (clonal, P = 1.36 × 10−34, effect size = 1.15; subclonal, P = 4.67 × 10−9, effect size = 0.6) (Methods and Extended Data Fig. 4b). Using multi-sample phasing, we investigated the presence of mirrored subclonal allelic imbalance7, which results from SCNAs that disrupt the same genomic region but affect different parental alleles within separate tumour subclones (Methods). WGD tumours were enriched in mirrored subclonal allelic imbalance events compared with non-WGD tumours (P = 1.2 × 10−10, effect size = 0.67) (Methods and Extended Data Fig. 4b). In tumours with subclonal WGD, we observed a higher frequency of SCNAs in subclones that were affected by WGD compared with their non-WGD sister clones (P = 9.5 × 10−3, effect size = 0.59, paired Student’s t-test) (Extended Data Fig. 4c), thus accounting for germline and somatic alterations as confounding variables.
Evolution of the SCNA landscape
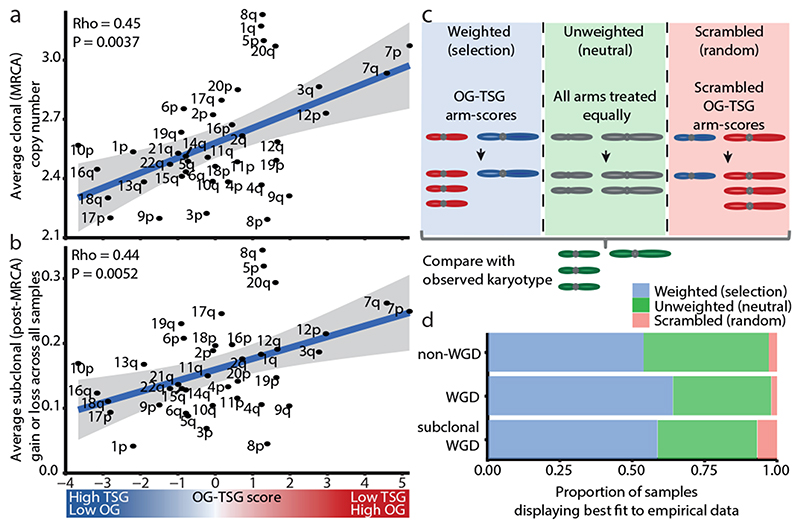
To investigate the degree to which the SCNA landscape is shaped by neutral evolution or selection, we analysed whether the propensity for each chromosome arm to be gained or lost during tumour evolution was related to the density of tumour-suppressor genes (TSGs) and oncogenes (OGs) that are encoded on each chromosome arm, as captured by the OG–TSG score3. Consistent with ongoing selection, the OG–TSG score significantly correlated with the burden of arm-level alterations in the MRCA (Fig. 2a) as well as with subclonal arm-level alterations (Fig. 2b and Extended Data Fig. 4d–f). No relationship between the average change in clonal or subclonal chromosome copy numbers and the size of the chromosome arm was observed (Extended Data Fig. 4g–j).
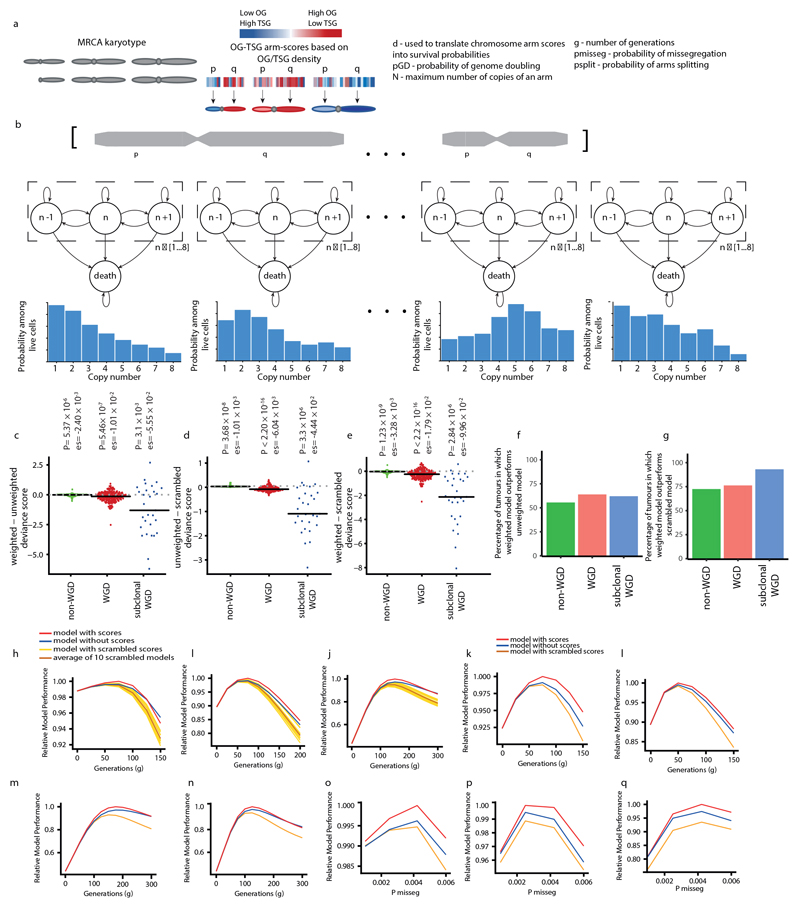
Fig. 2. Selection shapes the SCNA landscape.
a, There is a positive correlation between the average clonal copy number present in the MRCA and the OG–TSG score. n = 394 tumours. The grey shaded area represents the 95% confidence interval. ρ and P values are from a Spearman correlation test. b, There is a positive correlation between OG–TSG score and average change in SCNA (gain or loss) from the MRCA. n = 394 tumours. The grey shaded area represents the 95% confidence interval. ρ and P values are from a Spearman correlation test. c, The three conditions under which karyotype evolution was modelled: chromosome arms with OG–TSG scores included (weighted model); chromosome arms were treated equally (neutral model); OG–TSG scores were randomly permuted (scrambled model). d, For each context (WGD, n = 194 tumours; non-WGD, n = 171 tumours; and subclonal WGD, n = 29 tumours), the percentage of tumours for which each model condition best recapitulates the empirically observed data is shown.
To understand the subclonal SCNA dynamics within each tumour, we adapted our previous model that predicts population karyotypes over time13,14. We used arm-level copy number profiles from the MRCA of each tumour as the starting point and compared how different iterations of the model predicted the observed subclonal tumour karyotypes (Fig. 2c, Methods and Extended Data Fig. 5a, b). We compared three conditions; first, a condition in which karyotypes with a higher oncogenic or tumour-suppressive propensity were favoured or unfavoured, respectively, using the relative OG–TSG scores3 (weighted model); second, a model in which chromosome arms were treated equally (neutral model); and third, a condition in which the OG–TSG scores were randomly permuted (scrambled model). On average, the weighted model predicted the trajectory of subclonal SCNA more accurately, outperforming the two other models, as shown by significantly reduced deviance scores (Fig. 2c, d and Extended Data Fig. 5c–g) irrespective of the rate of chromosome missegregation or the number of cell divisions (Extended Data Fig. 5h–q).
Collectively, these data suggest that CIN enables continuous selection that is driven by the relative dosage imbalance of oncogenes and tumour-suppressor genes and that WGD may support further genome remodelling during later stages of tumour evolution. However, in 41% of our cohort the neutral or scrambled models outperformed the weighted model, which potentially reflects the evolution of a neutral karyotype or the need for tumour-type-specific chromosome arm weightings15,16. We found more evidence for subclonal selection in WGD tumours (the weighted model outperformed the neutral or scrambled models in 64% of WGD, 59% of subclonal WGD and 54% non-WGD tumours), which is consistent with WGD being a transformative event during tumour evolution13,14,17 (Fig. 2d and Extended Data Fig. 5f, g).
Evolution of clonal SCNAs
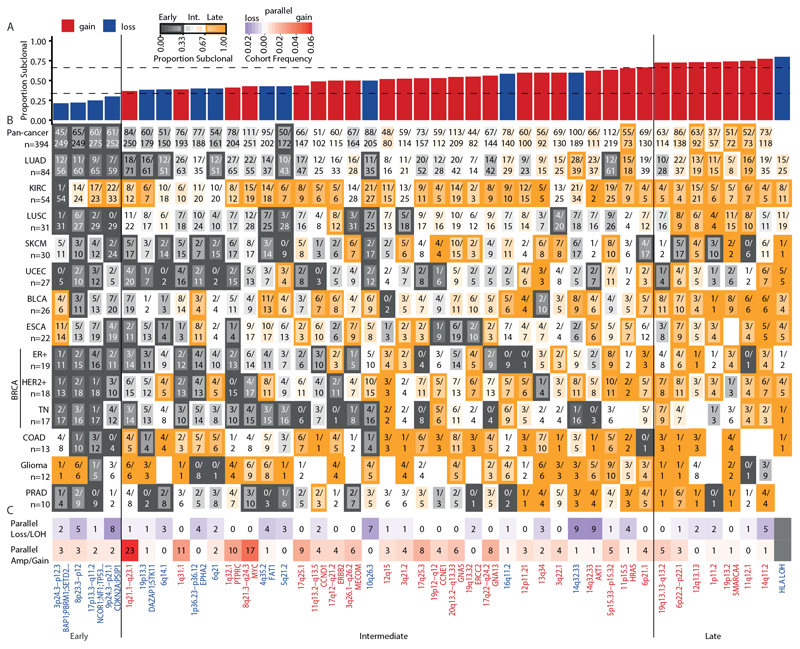
To decipher SCNA timing, we used GISTIC2.0 to identify recurrent SCNAs present in at least two tumour types (Methods, Extended Data Figs. 6a–h, 7a–e and Supplementary Table 3). We designated these as consensus peak regions and assigned each peak region to distinct evolutionary timing categories: early, intermediate or late (Fig. 3a, b and Methods). SCNAs that overlap with early peak regions may be implicated in tumorigenesis. SCNAs that overlap with intermediate or late peak regions may be involved in tumour maintenance and progression. Recurrent clonal and subclonal arm-level gain or loss SCNAs for each tumour type were identified using permutation testing (Methods and Supplementary Table 4).
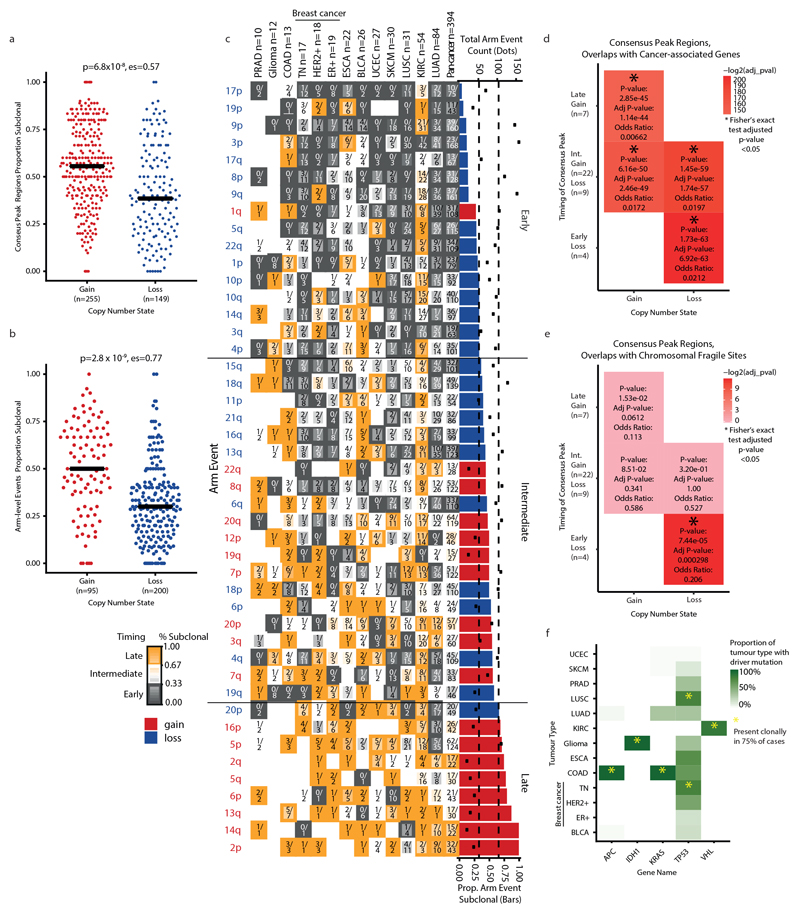
Fig. 3. Timing, recurrence and parallel evolution of subclonal SCNAs.
a, The consensus gain-peak (red) and loss-peak (blue) regions identified as subclonal across tumour types with ≥10 tumours. Data are sorted by the median proportion of SCNAs that is subclonal. Vertical lines indicate pan-cancer categories of early, intermediate and late events determined by median subclonal tumour type occurrence. b, For each consensus peak the proportion of SCNAs found to be subclonal within each tumour type. Orange, higher subclonality; grey, higher clonality. The border of each cell is classified according to early (grey border), intermediate (no border) or late (orange border) events. The numerator within each cell indicates the number of subclonal events; the denominator indicates the total number of clonal and subclonal events. Detection of HLA LOH was performed only in tumours with whole-exome sequencing. c, Consensus peak regions that show instances of parallel evolution of loss/LOH (purple) and gain/amplification (red). For full lists of cancer-associated genes within consensus peak regions see Supplementary Table 3.
We observed differences in evolutionary timing between peak regions that were associated with gains (gain peaks) and those with losses (loss peaks). Loss peaks were significantly more likely to be early compared with gain peaks (P = 6.8 × 10−8, effect size = 0.57; Extended Data Fig. 8a). Similarly, a higher proportion of recurrent arm-level losses were clonal compared with arm-level gains (P = 2.8 × 10−9, effect size = 0.77) (Extended Data Fig. 8b, c). Gain-peak regions were enriched in known oncogenes, whereas loss-peak regions were enriched in known tumour-suppressor genes (Extended Data Fig. 8d). Early loss-peak regions were also enriched in chromosomal fragile sites (Extended Data Fig. 8e), suggesting that some loss peaks may not be functionally important.
Frequencies of clonal SCNAs that affected early peak regions exceeded the frequency of clonal somatic driver point mutations and small insertions or deletions (indels) in cancer-associated genes (Fig. 3b and Extended Data Fig. 8f). The loss peak on chromosome 17p13.3−q11.2—which encompasses TP53—was classified as early in 9 out of 13 tumour types and classified as late only in KIRC (74% subclonal). In three tumour types (HER2+ breast cancer, lung squamous cell carcinoma (LUSC) and triple-negative breast cancer (TN BRCA)) more than 90% of tumours exhibited clonal loss of heterozygosity (LOH) at chromosome 17p13.1, which suggests that loss is required for tumorigenesis in these tumour types. Across tumour types, TP53 LOH was clonal rather than subclonal in 92% of WGD tumours when observed, indicating that TP53 LOH potentially enables tolerance for WGD18. In KIRC, loss or LOH of chromosome 3p26.3–p12.1, as well as LOH at the VHL locus, were early events (clonal LOH in 98% of KIRCs) (Extended Data Fig. 6h). Other high-frequency clonal peaks within individual tumour types included gains at chromosome 17q12−q21.2, which encompasses ERBB2, in HER2+ breast cancer (61% prevalence, 82% clonal), chromosome 3p LOH in LUSC (100% prevalence, 97% clonal) and gains in chromosome 7p11.2, which encompasses EGFR, in LUAD (63% prevalence, 72% clonal).
We reasoned that a genomic loss that occurred before WGD must lead to LOH with complete loss of the minor allele. Conversely, single losses that occurred after WGD will not lead to LOH. On average, across the cohort, 94% of clonal losses that overlapped early loss peaks involved LOH, which suggests that recurrent clonal loss events usually precede WGD.
The timing of other peak regions was variable between tumour types. For example, the loss peak at chromosome 4q35.2, which encompasses FAT1, was early in triple-negative breast cancer (88% prevalence, 80% clonal), intermediate in ER+ breast cancer (58% prevalence, 64% clonal) and late in HER2+ breast cancer (61% prevalence, 27% clonal) (Fig. 3b).
Evolution of subclonal SCNAs
We next analysed which specific subclonal SCNAs were recurrent during tumour evolution. The gain peaks with the highest frequencies, including chromosomes 1q21.1–q21.3 (which encompasses BCL9, MCL1, and ARNT) and 5p15.33–p15.32 (which includes TERT), varied in timing across tumour types. For example, in LUAD, 80% of gains in chromosome 5p15.33–p15.32 were clonal, whereas most gains in chromosome 5p15.33–p15.32 were subclonal in KIRC (76% subclonal), ER+ breast cancer (89% subclonal) and glioma (90% subclonal) (Fig. 3b). In LUSC, the timing of TERT gains was related to both its focality and amplitude; the majority of low-level gains were both clonal and arm-level (13 out of 21 tumours) whereas high-level TERT amplifications were often subclonal and focal (10 out of 11 tumours). This may reflect augmentation of gene dosage during evolution, with low-level TERT gain selected clonally, followed by a high-level amplification that is selected in a subset of cancer cells later in tumour evolution.
The gain peak in chromosome 19p12−q12 (which encompasses CCNE1) was late or intermediate in 10 out of 13 tumour types. High-level amplifications of CCNE1 (more than 2× ploidy), which was previously associated with WGD1,19, occurred exclusively in WGD tumours. CCNE1 amplification was subclonal in 9 out of 20 tumours with clonal WGD, which suggests that CCNE1 amplification may be selected for both before and after WGD.
Parallel evolution of SCNA events, which reflect events that occurred in distinct subclones within individual tumours and that converged on a similar evolutionary solution, was observed in 146 out of 394 (37%) tumours (Fig. 3c and Extended Data Fig. 9a). Allele-specific expression tracked parallel evolutionary events that originated from distinct haplotypes in samples with matched multi-sample RNA sequencing data (ρ = 0.89, P = 1.75 × 10−15, Spearman correlation) (Extended Data Fig. 9b, c).
Consistent with positive selection, parallel gains were significantly more focal than non-parallel subclonal gains (P = 7.1 × 10−3, effect size = 0.1). The most prominent parallel gains included those overlapping chromosomes 1q21.3–q44, which encompasses BCL9, MCL1 and ARNT, 5p15.33 which includes TERT, and 8q24.1, which encompasses MYC (Fig. 3c and Extended Data Fig. 9a). The most common parallel loss events included chromosomes 14q (14q32.33 (encompassing ASPP1) and 14q11.2 (encompassing NDRG2)), 10q and 9p (Extended Data Fig. 9a).
Subclonal LOH after a clonal WGD event occurs through more than one loss event of the same allele after the doubling event (Extended Data Fig. 9d). The HLA locus (chromosome 6p21.3) represented a clear peak of subclonal LOH in WGD samples, which affected 22% of the cohort, indicating that two loss events of the same alleles after WGD within the subclone occurred (Extended Data Fig. 9e). HLA LOH was prevalent as a subclonal event in KIRC, breast cancer, bladder urothelial carcinoma, endometrial carcinoma and oesophageal adenocarcinoma (Methods and Extended Data Fig. 9f) in addition to NSCLC as previously reported20. One exception was melanoma (SKCM), which is characterized by a high mutational burden and improved overall survival after checkpoint inhibitor blockade21. SKCM exhibited a low frequency of HLA LOH (0% clonal, 3% subclonal). The most prevalent recurrent clonal arm-level gain event in SKCM was 6p, which as well as encompassing the HLA locus, also contains the melanoma metastasis-associated gene NEDD9 22 at chromosome 6p24.2, which may constrain subsequent HLA loss (Extended Data Fig. 7d).
In a diploid cancer cell, any loss results in LOH. If this cell undergoes WGD, the LOH will be maintained and the remaining allele is duplicated, which leads to a total copy number of two. Notably, in the case of clonal chromosome 8p23.3–p12 loss, we observed a peak region of haploid LOH in WGD tumours, with only a single copy (Extended Data Fig. 9d). This haploid, single-copy LOH strongly suggests that a loss event of one of the two remaining copies occurred after WGD. Loss of chromosome 8p23.3–p12 was most prominent in breast cancer, in which this loss has been linked to a chromosome-dosage effect and has been shown to influence lipid metabolism and metastatic potential23.
Late-emerging subclones may seed metastases
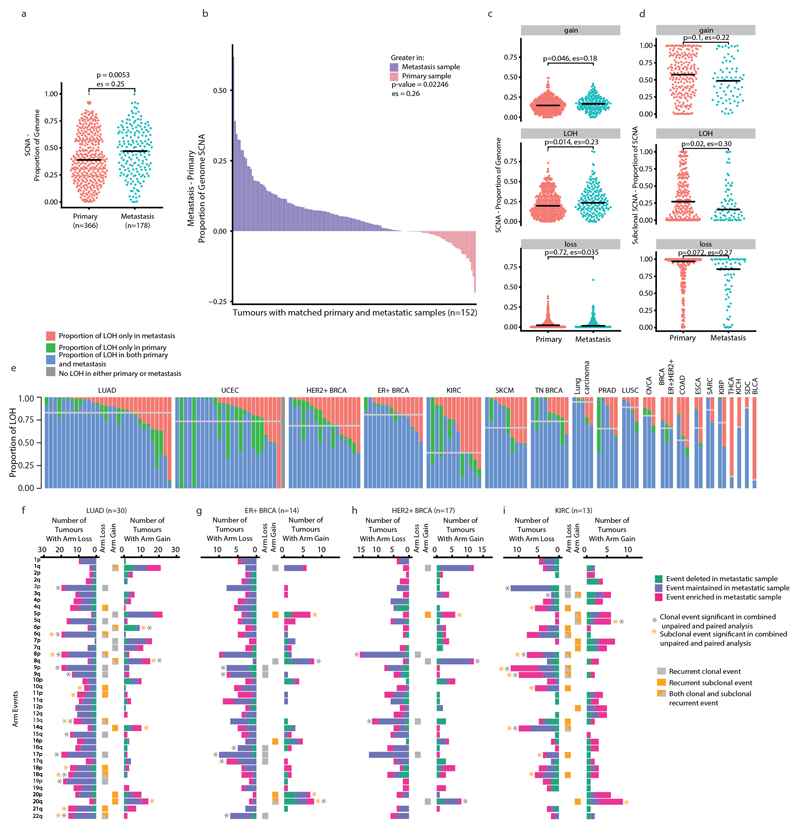
Finally, we explored associations between SCNAs and metastasis. Consistent with previous research24, a higher proportion of the genome was affected by SCNAs in metastatic samples (n = 178 patients) compared with primary tumour samples (n = 366 patients) (P = 5.3 × 10−3, effect size = 0.25) (Extended Data Fig. 10a). This remained significant after controlling for tumour type and when considering comparisons of both paired and unpaired primary tumours and metastases (Extended Data Fig. 10b) with LOH events showing the greatest increase from primary tumour to metastasis compared with gains or losses without LOH (Extended Data Fig. 10c). No significant increase in ploidy was observed between matched primary tumour and metastatic samples in the cohort as a whole, or in any individual tumour type.
Consistent with an evolutionary bottleneck, SCNAs were found to be more frequently clonal in metastases compared with primary tumours (Extended Data Fig. 10d). Indeed, in all 22 (5 ER+, 5 HER2+, and 2 TN BRCA as well as 5 KIRC, 2 LUAD, 1 SKCM, 1 papillary renal cell carcinoma and 1 lung carcinoma) tumours for which we had multiple primary tumour and matched metastatic samples, we identified SCNAs that were present as minor subclones within the primary tumour yet fully clonal in the metastasis. In 77% of tumours (116 out of 151) with at least one LOH event and paired primary tumour–metastasis samples, the majority of LOH was found to be shared between primary tumour and metastatic samples, with a median of 74% shared events. This suggests that there is a relatively late divergence of the metastatic clone relative to the MRCA in many tumours after WGD (Methods and Extended Data Fig. 10e).
To evaluate the relative importance of specific SCNAs in metastasis, we focused on recurrent SCNAs and performed a combined analysis using both paired analyses of 74 tumours with matched primary and metastatic samples, and unpaired analyses of 2,631 primary tumour samples from The Cancer Genome Atlas (TCGA) and 1,024 metastatic samples from the Hartwig Medical Foundation (HMF) for the four tumour types (HER2+ breast cancer, ER+ breast cancer, LUAD and KIRC) for which sufficient primary tumour–metastasis pairs were available. Distinct patterns of SCNA metastatic dissemination were observed in different tumour types. In ER+ breast cancer, HER2+ breast cancer and LUAD, the majority of the recurrent arm-level events that were enriched in metastasis relative to the primary tumours were clonal events (Extended Data Fig. 10f–h). Conversely, in KIRC, which also had the lowest proportion of shared LOH between primary tumour and metastatic samples, most recurrent arm-level events that were enriched in metastatic samples were subclonal events (Extended Data Fig. 10i), which suggests that these arm-level events are associated with metastatic potential in a limited number of cells within the primary tumour.
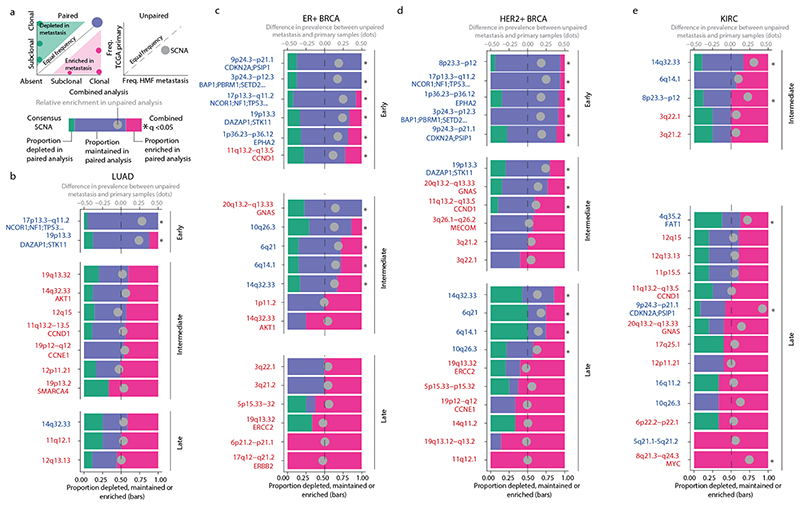
The early loss peak in chromosome 1p36.23–p36.12, which encompasses EPHA2, and the early loss peak in chromosome 17p13.3–q11.2, which encompasses TP53, were enriched in metastatic samples compared with primary tumour samples in ER+ breast cancer and HER2+ breast cancer (Fig. 4). In LUAD, two early loss consensus peak regions were significantly enriched in metastases (chromosomes 17p13.3–q11.2 (which encompasses TP53) and 19p13.3 (which encompasses STK11)), consistent with the idea that these early events in tumour evolution contribute to the metastatic potential of the tumour.
Fig. 4. Analysis of consensus peak regions in metastatic LUAD, ER+ and HER2+ breast cancers, and KIRC.
a, Schematics show the paired (left), unpaired (right) and combined (bottom) analyses of consensus peak regions. The schematic bar graph summarises the left graph for each event and indicates the proportion of paired primary tumour–metastasis samples in which a SCNA overlapping the event was enriched (pink), depleted (green) or maintained (purple) in metastatic samples. b–d, We restricted our analysis to tumour types with >10 paired primary tumour–metastatic samples: LUAD, paired n = 30, unpaired n = 844 TCGA and unpaired n = 315 HMF lung cancers (b); ER+ breast cancer, paired n = 14, unpaired n = 1,015 TCGA and unpaired n = 620 HMF breast cancers (c); HER2+ breast cancer, paired n = 17, unpaired n = 1,015 TCGA and unpaired n = 620 HMF breast cancers (d); and KIRC, paired n = 13, unpaired n = 772 TCGA and unpaired n = 89 HMF kidney cancers (e). These data were assessed using a two-sided binomial test. The grey circle in the schematic bar graph indicates the difference between the proportions of metastatic (HMF) and primary (TCGA) samples that contain the event in the unpaired primary tumour–metastasis analysis (two-sided test of equal or given proportions). A positive number indicates that the event was more prevalent in the metastatic (HMF) samples; a negative number indicates that the event was more prevalent in the primary tumour (TCGA) samples. The asterisks indicate whether an event was significantly enriched in metastatic samples as determined by a combined analysis of paired (multi-sample) and unpaired (HMF and TCGA) data using Fisher’s method after correction for multiple testing using the Benjamini–Hochberg method. The event timing classifications (early, intermediate or late) were determined based on the proportion of subclonal occurrence (Methods). Only losses (blue text) or gains (red text) that were either significant (q < 0.05) or exhibited ≥40% enrichment are shown. For full lists of cancer-associated genes within consensus peak regions, see Supplementary Table 3.
By contrast, other consensus peak regions that were enriched in metastases were classified as intermediate or late events (Fig. 4). Examples include the loss of chromosomes 14q32.33, 6q21 (which encompasses PRDM1), 6q14.1 and 10q26.3 (which encompasses MGMT) in HER2+ breast cancer, and loss of chromosomes 4q35.2 (which encompasses FAT1), 9p24.3–p21.1 and gain of chromosome 8q21.3–q24.3 in KIRC. In KIRC gain of chromosome 8q21.3–q24.3—which encompasses MYC—was highly enriched in our combined analysis as well as exclusively identified in the metastatic samples of our matched primary tumour–metastasis pairs. Notably, loss of chromosome 9p24.3–p21.1, which encompasses CDKN2A, was a late metastasis-associated event in KIRC, whereas in ER+ and HER2+ breast cancers, in which the loss of chromosome 9p24.3–p21.1 was also significantly associated with metastasis, this loss was predominantly early. Similarly, gain of chromosome 11q13.2–q13.5, which encompasses CCND1, was an early event in ER+ breast cancer, an intermediate event in HER2+ breast cancer and associated with metastasis in both tumour types.
Together, these results highlight the importance of early and continuous SCNA acquisition during tumour evolution and their potential importance during the transition to metastasis.
Discussion
Clonal and subclonal SCNAs are pervasive across tumour types and tend to occur as ordered events, which potentially reflects the continuous optimization of the fitness landscape throughout tumour evolution. WGD is a transformative event in tumour development, which is associated with the acquisition of clonal and subclonal SCNAs. LOH events that affected tumour-suppressor genes (including TP53) frequently preceded WGD, whereas recurrent gains (for example, in CCNE1) frequently followed WGD and were more likely to be subclonal.
The subclonal landscape of SCNAs is sculpted by both positive and negative selection, as well as neutral evolution. In a minority of tumours, our results are consistent with subclonal karyotypic evolution that reflected neutral growth15,16. However, particularly in tumours with WGD, SCNA evolution was better recapitulated using models that included positive and negative selection (Fig. 2d). Positive selection was further shown by recurrent peaks of subclonal amplifications, which were enriched in established oncogenes, subclonal losses that resulted in LOH, even after WGD, and parallel evolution of SCNAs. These data are consistent with documented parallel and convergent evolution of SCNAs7,25,26,27. Finally, recurrent focal subclonal SCNAs—including gains encompassing oncogenes such as CCND1 and MYC—were enriched at metastatic sites, suggesting that focal subclonal SCNAs have a potential role in metastasis. Consistent with this, MYC was recently described as a driver of brain metastasis in LUAD28. Certain early clonal SCNAs were enriched in metastases. These may be necessary but not sufficient for metastatic dissemination as most LOH events were shared between primary and metastatic samples, which suggests a late divergence of the metastatic clone, often after WGD.
Our work has limitations. Detection of recurrent SCNAs is not necessarily indicative of selection and may result from other processes that drive tumour progression, such as DNA repair dysfunction or the presence of adjacent fragile sites. Indeed, the higher frequency of recurrent SCNAs compared with driver point mutations may not reflect selection. However, we only found an association of fragile sites with early loss peak regions. Extrachromosomal DNA may also contribute to the subclonal SCNA amplification events that were observed29. The number of tumour samples, their sequencing depths and the lack of an extensive cohort of paired primary tumour and metastatic samples or single-cell sequencing data influence the degree to which subclonal heterogeneity can be deciphered, suggesting that the extent of diversity is underestimated. The lack of uniform clinical data collection and central pathology review prevented a detailed analysis of clinically relevant parameters. We are endeavouring to address these deficiencies within TRACERx7.
In conclusion, our work highlights the importance of ongoing CIN during tumour evolution and metastasis. As our functional understanding of the propensity for different chromosomes to missegregate30 and the extent to which chromosomal alterations may be deleterious or advantageous to the cancer cell improves17, it will be possible to refine the parameters of selection models and improve the ability to detect novel SCNA drivers, which may drive metastatic dissemination and death.
Methods
Statistical information
The experiments were not randomized and investigators were not blinded to allocation during experiments and outcome assessment. All statistical tests were performed in R version 3.6.1. No statistical methods were used to predetermine sample size. Tests involving correlations were done using the Spearman’s method. Tests involving comparisons of distributions were done using ‘wilcox.test’ or ‘t.test’ using the unpaired option, unless otherwise stated. For all statistical tests, the number of data points included are plotted or annotated in the corresponding figure legend. Effect sizes were calculated using the standardized means difference.
Whole-exome sequencing
All whole-exome sequencing (WES) data were processed from FASTQ, as previously described7. Copy number segmentation, tumour purity and ploidy for each sample were estimated using ASCAT31 version 2.3 and were used in our multi-sample SCNA clonality approach (see below). A subset of the WES cohort evaluated in this study comes from the first 100 patients prospectively analysed by the lung TRACERx study (https://clinicaltrials.gov/ct2/show/NCT01888601, approved by an independent research ethics committee, 13/LO/1546) and mirrors the previously described prospective 100 patient cohort7.
Whole-genome sequencing
Copy number segmentation, tumour purity and ploidy for each sample were estimated with Battenberg as previously described32,33,34,35,36,37 and used as input for downstream clonality analyses (see below).
Single-nucleotide polymorphism arrays
Copy number segmentation, tumour purity and ploidy for each sample assayed using single-nucleotide polymorphism (SNP) arrays15,38 were estimated using ASCAT31 version 2.3 and were then used for downstream clonality analyses (see below).
RNA sequencing
RNA sequencing data from 58 tumours from the TRACERx-100 cohort were used39. FASTQ data underwent quality control and were aligned to the hg19 genome using STAR40. Transcript quantification was performed using RSEM41 with default parameters.
Allele-specific expression
Allele-specific expression was obtained using phASER42. Allele-specific expression of heterozygous SNPs identified by Platypus43 version 0.8.1 analysis of WES data and with at least eight supporting RNA sequencing reads was used in allelic imbalance in expression analysis. Allelic imbalance in expression of each SNP was determined by a binomial test of allele-specific expression with a significance threshold of P < 0.05. Allelic imbalance in expression intratumour heterogeneity was calculated per gene, where allelic imbalance in expression intratumour heterogeneity is declared when some but not all samples of a tumour have allelic imbalance in expression. DNA allelic imbalance intratumour heterogeneity per gene was declared when some but not all samples of a tumour assayed with WES exhibited allelic imbalance.
Cancer-associated gene single-nucleotide variants and indel calls
Single-nucleotide variants (SNV) and indel calls and their clonality, classed as driver mutations in the respective publications that we have reanalysed as part of multi-sample cohort (Supplementary Table 1), were collated.
Definition of cancer-associated genes
Cancer-associated genes from a previous study44 (including oncogene and tumour suppressor classifications) that were defined on the basis of statistical analyses of only SNVs were used. Therefore, these can be considered orthogonal to cancer-associated genes identified through SCNA analysis.
Cancer-associated genes from COSMIC45 version 75 and genes from STOP and GO3 within consensus peaks of SCNA (see ‘GISTIC2.0 peak definition’, ‘GISTIC2.0 consensus peak definition’ and ‘Consensus peak timing’) were used for annotation but not enrichment analyses (see ‘Cancer-associated gene and fragile site enrichment’).
Definition of B-allele frequency
When analysing next-generation sequencing, the ‘B’ allele is the non-reference allele that is found at the position of a germline heterozygous SNP. The B-allele frequency (BAF) is defined as the proportion of the reads that carry the B allele (that is, the non-reference allele). In SNP arrays, BAF is defined as cases in which there are two probes (an A probe, which is generally the reference sequence, and a B probe) that cover a specific position and is a normalized measure of the allelic intensity ratio of the A and B probes.
SCNA estimation using multi-sample phasing
Multi-sample phasing uses the allelic imbalance that results from SCNAs causing an unequal copy number of homologous chromosomes at a genomic location to obtain a phasing of heterozygous SNPs. In regions of allelic imbalance, the heterozygous SNP BAF separates into two distributions. The identities of the heterozygous SNPs in each of these two distributions in the same genomic region will be consistent across samples from the same tumour as SCNAs will not alter the mapping of heterozygous SNPs to each original homologous chromosome. Our approach uses a phasing derived from an area of allelic imbalance in one sample and applies it to the same genomic region in another sample from the same tumour.
For all samples, manual verification of the automatically selected models for ploidy, purity and the resulting copy number segmentation that were produced by ASCAT31 or Battenberg33 was performed. Samples that had insufficient purity or unreliable copy number profiles were excluded. Only copy number segmentation from autosomes was included in the study. We then defined a tumour consensus segmentation profile, CS, by combining breakpoints from each SCNA segmentation profile of each individual tumour sample. For each segment csi of the CS from a tumour, we examined the allelic imbalance to determine whether multi-sample phasing could be applied if that genomic region was described to have allelic imbalance by ASCAT31 or Battenberg33 and it contained at least five heterozygous SNPs.
For each csi, the sample with the most bimodal distribution of BAF (ranked by the P value from Hartigans’ dip test statistic46 from the package ‘diptest’47 and then a measure of mean absolute deviation of the BAF in that segment from 0.5) is chosen as the reference sample that provides a phasing for all other samples.
We then estimate the phased A allele and the phased B allele copy number at each heterozygous SNP position, using the following equations, with the log2[R] value at the same position. Using these estimates, the phased allele specific copy number (cpn) is estimated for each csi of CS across all samples.
where ρ is tumour sample purity, ψ is tumour sample ploidy and γ accounts for. technological differences and refers to the compaction of log2[R] profiles.
SCNA classifications relative to sample ploidy
Three thresholds were used to identify four possible copy number states relative to ploidy: amplification, gain, neutral and loss. Each segment with ≥5 log2[R] values in all samples of a tumour was examined relative to log2[R] thresholds (termed log2[R]exp). These thresholds represent an expected ‘raw’ or continuous log2[R] estimate of total copy number adjusted to the values of purity and ploidy of that sample (see equations below).
Equations describe the ploidy- and purity-dependent copy number thresholds, where ρ is tumour sample purity and ψ is tumour sample ploidy.
The log2[R] values within a segment are then compared to each of these thresholds using a one-tailed Student’s t-test, ensuring that they are higher than the threshold when amplifications and gains are examined and lower when losses are examined with a P < 0.01 threshold. An amplification, gain or loss passing its respective threshold in a sample is considered to be clonal within that sample. The >2× ploidy threshold is the same threshold used for clinical decision making in HER2+ breast cancer using fluorescence in situ hybridization samples48.
To enable comparisons across tumours, segments were mapped to hg19 cytobands. If multiple segments mapped to a cytoband, the SCNA status of the segment with the largest overlap with the cytoband was chosen.
Detection of mirrored subclonal allelic imbalance
To detect subclonal allelic imbalance from independent SCNAs in distinct subclones, or mirrored subclonal allelic imbalance7, we used previously described methods 7. In brief, we used one tumour sample as a reference sample for multi-sample phasing, and explored whether multiple samples had the major allele—the haplotype with the higher frequency—which was derived from distinct haplotypes in two different samples from the tumour of a patient.
Detection of parallel SCNA evolution
We define parallel SCNA evolution as the same class of event (gain/amplification or loss/LOH) in multiple samples from an individual tumour but with major alleles from distinct haplotypes in the samples that had the event.
If SCNAs that affect the same genomic loci originate from different haplotypes within the tumour of the same patient, they are independent and therefore subclonal. A subset of these will also show parallel evolution when they result in the same class of copy number change relative to ploidy. We used SCNA classifications relative to sample ploidy (see ‘SCNA classifications relative to sample ploidy’) with our detection of mirrored subclonal allelic imbalance (see ‘Detection of mirrored subclonal allelic imbalance’) and identified tumours in which gains/amplifications from distinct haplotypes and loss/LOH events from distinct haplotypes in different samples were found. Manual review of events under one megabase in size was performed. The number of tumours with parallel events overlapping at least one cytoband within a consensus peak region was reported in Fig. 3c. Across-genome plots at the single cytoband level showing the proportion of the cohort affected by instances of parallel evolution overlapping each cytoband are shown in Extended Data Fig. 9a.
SCNA intratumour heterogeneity and clonality definitions
We quantified multi-sample phasing estimates of allele-specific SCNA clonality using our classifications relative to ploidy, mirrored subclonal allelic imbalance and LOH detection. The following definitions were used.
Clonal amplification, all tumour samples demonstrate amplification.
Subclonal amplification, at least one, but not all, samples of the tumour showed amplification.
Clonal gain, every sample analysed from the tumour showed gain or amplification.
Subclonal gain, one or more but not all samples analysed from the tumour had a relative to ploidy classification of gain or amplification.
Clonal loss, either all samples from the tumour had a loss relative to ploidy or all samples demonstrate LOH. A sample may have both LOH and a loss relative to ploidy and still count towards either of these definitions.
Subclonal loss, at least one or more but not all samples had a loss or at least one or more but not all regions had LOH.
WGD estimation
WGD estimation was performed as previously described7 (Supplementary Methods). All WGD estimates were manually reviewed.
Permutation test for recurrence of SCNA across tumours
A background rate was calculated and thresholds established for calling significance. Specifically, to determine significant clonal losses, for each tumour, the proportion of the genome subject to loss was determined. This value was taken as the probability of a loss event in each tumour. Based on this probability it was possible to separately generate an aberration state (loss or no loss) for each tumour and calculate the proportion of tumours that showed a loss. By repeating this process 1,000 times it was possible to obtain a background distribution that reflects the expected likelihood of loss events. Using this background distribution, a 0.05 significance loss threshold was established for which less than 5% of simulations exceeded that level of loss. The same procedure was used to establish thresholds for gains. Thresholds for each tumour type were established for (1) clonal SCNAs; (2) subclonal SCNAs; and (3) mirrored subclonal allelic imbalance.
Arm-level SCNA definition
Recurrent arm-level SCNAs were defined for four categories: clonal gain, subclonal gain, clonal loss/LOH and subclonal loss/LOH at P = 0.05 (see ‘Permutation test for recurrence of SCNA across tumours’). A significant arm-level event was defined as being present if at least 75% of the chromosome arm (defined at the cytoband level) was found to affect the cohort at a frequency above the significance threshold of 0.05 (see ‘Permutation test for recurrence of SCNA across tumours’).
For each tumour type, each arm-level SCNA was classified as one of three distinct evolutionary timing categories: early (clonal in more than two-thirds of tumours), late (subclonal in more than two-thirds of tumours) and intermediate timing (less than two-thirds clonal and less than two-thirds subclonal).
Post-WGD haploid LOH
Significantly recurrent areas of single-copy LOH were identified using the permutation test (see ‘Permutation test for recurrence of SCNA across tumours’) applied to copy number segments that showed single-copy LOH from WGD samples.
HLA LOH detection
The algorithm LOHHLA20 was used to identify LOH at the HLA locus. LOHHLA was applied to all WES data in the cohort, with default settings.
GISTIC2.0 peak definition
We generated summary SCNA profiles for each tumour that corresponded to either clonal SCNA or subclonal SCNA (Supplementary Methods).
The allele-specific copy number values present in the copy number segmentation for all samples were first transformed to match the non-allele-specific ‘seg_CN’ format expected by GISTIC2.0. Following the previously outlined procedure1, we normalized the total copy number by the ploidy of the corresponding sample.
This was performed with the following equation, where ψ represents tumour sample ploidy:
For details on the incorporation of LOH see Supplementary Methods.
GISTIC2.0 consensus peak definition
GISTIC2.049 was run on clonal and subclonal input from all tumour types with 10 or more tumours in our multi-sample cohort, with default settings (see ‘GISTIC2.0 peak definition’ and Supplementary Methods). The clonal and subclonal gain and loss peaks were mapped to the affected hg19 cytobands.
Cytobands that were identified as significant in both the clonal and subclonal GISTIC2.0 runs for the same tumour type were only included as significant subclonal events if they were also identified as a significant subclonal event in a separate permutation-based analysis to identify subclonal recurrence within that tumour type (see ‘Permutation test for recurrence of SCNA across tumours’).
Finally, a cytoband was identified as part of a consensus peak region of either gain or loss if it was present in at least four GISTIC2.0 clonal or subclonal peaks of the same type (gain/loss) as well as present in at least two tumour types.
Consensus peak timing
Consensus peak regions of SCNA were examined across all copy number data from all tumours in our cohort. For each tumour type, each consensus peak region was classified as one of three distinct evolutionary timing categories: early (SCNA overlapping a peak region that was clonal in more than two-thirds of tumours), late (SCNA overlapping a peak region that was subclonal in more than two-thirds of tumours) and intermediate timing (SCNA overlapping peak regions that were less than two-thirds clonal and less than two-thirds subclonal).
Ancestral reconstruction and phylogeny inference
We used MEDICC10 to reconstruct the phylogenetic trees of the tumour of each patient from allele-specific copy number profiles and to infer the allele-specific copy number profile of the MRCA.
Creation of arm-level input for Markov chain modelling
The allele-specific integer copy number profiles for all samples of a single tumour were used as input to MEDICC10 to create an integer copy number profile of the inferred MRCA. For each chromosome arm, the mean total copy number rounded to the nearest integer, weighted by segment size, was determined. This MRCA arm-level total copy number summary was used as the starting point for the Markov chain modelling for that tumour.
Description of the Markov chain model that incorporates arm-level events
We adapted a Markov chain model that we have described previously14 that keeps track of the distribution of the number of copies of a given chromosome arm (Supplementary Methods).
Markov chain model parameters
The values of the basic model parameters were based on the previously developed model14. Other parameter values, such as pGD (probability of WGD) and g (number of generations), were empirically derived to minimize the deviance between the predicted and actual copy numbers. Robustness analysis (Extended Data Fig. 5) indicated that our primary conclusion—that the model with scores outperforms the model without scores—is robust over a wide range of WGD rates pGD, number of generations g and chromosome missegregation rates pmisseg (Supplementary Methods).
Incorporation of OG–TSG scores in Markov chain modelling
At each generation, each cell in the colony dies spontaneously with certain probability 1 − Qsurv. To compute Qsurv, we use a formula similar to equation (1) of a previously published study14 (Supplementary Methods).
Investigation of Markov chain modelling results
For each sample, the model was run on the initial data of each tumour (the arm-level total copy number summaries for the MRCA), with parameter values for pmisseg (missegregation rate), pGD (probability of WGD), g (number of generations).
In order to assess the Markov chain modelling output, the weighted mean total copy number by segment size state of each chromosome arm was calculated for each sample of the tumour and the values rounded to the nearest integer. This produces arm-level total copy number summary profiles for each of the tumour samples from the observed SCNA data.
The output of the runs of the model that were weighted (incorporating arm OG–TSG scores)3 and unweighted were then scored versus the observed subclonal sample arm-level karyotype summaries. This was computed by looking at each sample as a separate event, with the error of the prediction (termed deviance score) measured as the sum over all samples of the squares of the differences between the final copy number in the sample and the average predicted copy number.
Differences in deviance score were compared across classifications of weighted, unweighted and scrambled runs of the model by subtracting the deviance score of tumours calculated using the results of one model run (for example, weighted) from a second model run (for example, unweighted). A negative deviance score difference in a comparison indicates the first model was closer to the observed subclonal SCNA data than the second model as it deviates less from the actual karyotype (Supplementary Methods).
Cancer-associated gene and fragile site enrichment
Enrichment for known cancer-associated genes (see ‘Definition of driver genes’) and fragile sites50 were assessed with Fisher’s exact tests. We examined the significance of the overlap at the level of cytobands of genes in gains and oncogenes, and the overlap of genes in losses and tumour-suppressor genes. Significant overlaps were those with P < 0.05.
TCGA data processing
Affymetrix SNP 6.0 profiles were obtained for paired tumour–normal samples from the TCGA (dataset ID, phs000178.v10.p8) and processed using PennCNV libraries51 to obtain BAFs and log2[R] values from each tumour–normal pair. log2[R] values and BAFs were processed with ASCAT31 version 2.4.2 using default parameters including correction for replication timing and GC-content biases52 to obtain copy number, purity and ploidy estimates.
Cell cycle gene expression signature
Transcripts per kilobase per million reads (TPM) expression values were obtained from our RNA sequencing data and 45 cell cycle genes53. A per-gene z-score was calculated to normalize comparisons across the gene set. For each sample, we calculated a mean z-score for all genes in the set and this score was compared with SCNA measures.
Mitotic index, anisonucleosis and tumour volume
The mitotic index and anisonucleosis (variation in nuclear size) were assessed from digitized diagnostic slides of the primary tumour (LUAD, n = 53 tumours; LUSC, n = 27 tumours; NSCLC-other, n = 3 tumours). The mitotic index was defined as the number of mitotic figures (the microscopic appearance of a cell undergoing mitosis) seen in 2.4 mm2 (equivalent to 10 high-power fields of an Olympus BX45 microscope) in the most mitotic region of the tumour. Anisonucleosis was scored from 1 to 3 and scores were assigned as follows: (1) tumours with minimal variation in nuclear size that could only be seen at high-power magnification; (2) moderate variation in nuclear size; (3) marked variation with numerous tumour nuclei that were more than double the diameter of other tumour nuclei. These categories were further grouped into ‘low’, which included those tumours with anisonucleosis scores of 1 and 2; and ‘high’, which included only those tumours with anisonucleosis scores of 3 (Supplementary Methods). Tumour volume estimates derived from diagnostic positron emission tomography–computed tomography scans for a subset of 83 tumours in our cohort were previously published54.
TCGA primary tumour and HMF metastatic data processing
Processed copy number segmentation, ploidy and purity information were downloaded from the HMF55. For processed copy number segmentation data from both the TCGA and HMF, for each segment in each sample, the total raw copy number (cpntotal) was determined as the sum of the major-allele copy number ‘cpnmajor’ and minor-allele copy number ‘cpnminor’ and processed as follows to assign a relative-to-ploidy copy number status using the following equations, in which ψ represents tumour ploidy.
Paired primary tumour–metastasis analysis
A paired analysis of matched primary and metastatic samples from 74 patients was performed. We designated lymph-node samples as metastases. For each case, for each consensus peak region and arm-level event for the corresponding tumour type we determined whether it was (1) maintained (that is, present in both primary tumour and metastatic samples); (2) enriched (that is, present only in metastatic sample(s)); (3) depleted (that is, present only in primary tumour samples(s)); or (4) absent (that is, not present in either primary tumour or metastatic samples).
Only samples that had primary tumour locations indicated as breast, lung or kidney were considered, as these were the tumour types for which we had sufficient (n > 10) paired primary tumour–metastatic samples and >50 unpaired metastatic samples.
For each tumour type, to determine whether an event was significantly enriched in metastatic samples, we performed a binomial test comparing the number of enriched versus depleted samples.
Unpaired primary tumour–metastasis analysis
We compared the frequency of each consensus peak region and arm-level event in primary tumour samples from the TCGA (n = 2,631; 1,015 breast cancer, 844 lung cancer and 772 kidney cancer samples) and metastatic samples from the HMF (n = 1,024: 620 breast cancer, 315 lung cancer and 89 kidney cancer samples). For each tumour type, to determine whether an event was significantly enriched in metastatic samples, we performed a test of equal or given proportions (prop.test in R) using the number of primary tumour samples that had the event, the number of metastatic samples that had the event, the total number of primary tumour samples and the total number of metastatic samples.
Paired and unpaired meta-analysis
To consider the results for each consensus peak region and arm-level event from the paired and unpaired analyses together, we performed a meta-analysis using the Fisher method (fisher.method from the metaseqR package56 version 1.26) with the P value generated from the binomial test on the paired data (see ‘Paired primary tumour–metastasis analysis’) and the P value generated from the prop.test on the unpaired data (see ‘Unpaired primary tumour–metastasis analysis). The resulting P value was then corrected for multiple testing using the Benjamini–Hochberg method to obtain q values. Events that were considered significantly enriched in this combined analysis were those with q < 0.05.
Primary tumour–metastasis shared and private LOH
All regions of LOH in each tumour with both primary tumour and metastatic samples were considered. Genomic regions that only demonstrated LOH in one or more primary tumour samples were classified as primary-tumour-only LOH, those that only demonstrated LOH in one or more metastatic samples were classified as metastasis-only LOH and those that demonstrated LOH in both at least one primary sample and at least one metastatic sample were classified as shared primary tumour–metastasis LOH. The total area of the genome subject to LOH was calculated by summing all three categories for each tumour and the relative proportion that each LOH category represented was calculated.
Extended Data
Extended Data Fig. 1. Measuring CIN across tumour types.
a, Schematic of the analyses of allele-specific copy number alterations. Left, the SCNA profiles across the genome for the two samples of a tumour (red, A allele; blue, B allele), with raw allele-specific copy number values for heterozygous SNPs shown as points and inferred allele-specific integer copy number states as lines. The clonality of the SCNAs across the two samples is indicated by a track between the two SCNA profiles, with clonal SCNAs indicated in grey, subclonal SCNAs in yellow and both clonal and subclonal SCNAs in dashed yellow and grey. All SCNA profile plots in the figure are scaled by the number of data points per chromosome. Top right, the approach to summarise SCNA timing (clonal versus subclonal) from the tumour. Bottom right, the integer SCNA profile across the genome of the inferred MRCA based on the integer SCNA profiles of the two samples of the tumour. b, c, Multi-sample phasing (b) and SCNA calling relative to ploidy (c). b, Multi-sample phasing is the method that we used to obtain allele-specific copy number profiles. This allowed us to identify previously undetected allelic imbalance (yellow boxes), and mirrored subclonal allelic imbalance and parallel SCNAs (purple boxes). c, Chromosomal illustrations and nomenclature of various SCNAs. As SCNAs are reported relative to ploidy, illustrations are provided for the diploid, triploid and tetraploid states. AI, allelic imbalance. d, e, Pan-cancer cohort characteristics. Our pan-cancer multi-sample cohort is summarised by tumour type in these bar plots, indicating the total number of patients (d) with the bar plot coloured according to the number of samples each tumour contributes, and tumour samples (e) with the bar plot coloured according to the type of sample.
Extended Data Fig. 2. SCNA correlates across tumour types.
a, Scatter plots indicating, for each tumour type, the association between the number of samples and the proportion of the genome affected by subclonal SCNAs. ρ and P values are from Spearman correlation tests. b, Scatter plots showing median purity per tumour versus the proportion of the genome affected by subclonal SCNA. ρ and P values are from Spearman correlation tests. c, Comparing the proportion of the genome affected by clonal and subclonal SCNAs. The median value for each tumour type is indicated. The size of the dots indicates the number of tumours in the corresponding tumour type. Red dots indicate tumour types with significant differences in the proportion of the genome affected by clonal versus subclonal SCNAs. A two-sided Student’s t-test was used to compare proportions of the genome affected by clonal and subclonal SCNAs. a–c, Tumour types with tumour samples from at least 10 patients were included: bladder urothelial carcinoma (BLCA, n = 26), ER+ breast cancer (ER+ BRCA, n = 19), HER2+ breast cancer (HER2+ BRCA, n = 18), triple-negative breast cancer (TN BRCA, n = 17), colorectal adenocarcinoma (COAD, n = 13), oesophageal adenocarcinoma (ESCA, n = 22), glioma (n = 12), clear cell renal cell carcinoma (KIRC, n = 54), lung adenocarcinoma (LUAD, n = 84), lung squamous cell carcinoma (LUSC, n = 31), prostate adenocarcinoma (PRAD, n = 10), melanoma (SKCM, n = 30) and endometrial carcinoma (UCEC, n = 27). d, The results of the linear regression analysis between LUAD and HER2+ breast cancer of the proportion of the genome subject to subclonal SCNAs along with the number of samples from each tumour and the median sample purity for each tumour.
Extended Data Fig. 3. NSCLC SCNAs correlate with cell cycle gene expression and tumour cell characteristics.
a, b, Scatter plots comparing the average cell cycle gene expression in LUAD tumours (n = 36), LUSC tumours (n = 15) and NSCLC-other tumours (n = 7) with the total proportion of the genome affected by SCNAs. Each dot is coloured according to tumour type. (a) and the proportion of the genome affected by clonal SCNAs (b). c, The proportion of the genome affected by subclonal SCNAs. d, The proportion of SCNAs that are subclonal. a–d, ρ and P values are from Spearman correlation tests. Associations between tumour cell characteristics and SCNA statistics for LUAD (n = 53), LUSC (n = 27) and NSCLC-other (n = 3). e–h, Mitotic index scores for each tumour are compared against total SCNAs (e), clonal SCNAs (f), subclonal SCNAs (g) and the proportion of SCNAs that are subclonal (h) in each tumour. Each dot is coloured according to tumour type. ρ and P values are from Spearman correlation tests. i–l, Association between tumour volume and SCNA metrics. For each tumour for which both digitized slides and tumour volume information were available (n = 83), we performed Spearman correlation tests comparing the tumour volume with the total proportion of the genome affected by SCNAs (i), the proportion of the genome affected by clonal SCNAs (j), the proportion of the genome affected by subclonal SCNAs (k) and the proportion of SCNAs that are subclonal (l). Padj values reflect P values from linear regression models incorporating the number of samples as well as estimated tumour volume and SCNA measure investigated. m–p, Associations between tumour cell characteristics and SCNA statistics for LUAD (n = 53), LUSC (n = 27) and NSCLC-other (n = 3). Anisonucleosis scores for each tumour are compared with the proportion of the genome affected by SCNAs (m), clonal SCNAs (n) or subclonal SCNAs (o) and the proportion of SCNAs that are subclonal (p) in each tumour. Each dot is coloured according to tumour type. The lines represent the median of each group. es, effect size.
Extended Data Fig. 4. WGD across tumour types.
a, Bar plots indicating the number and proportion of tumours of each tumour type that show WGD. Subclonal WGD tumours are indicated in blue. b, Beeswarm plots comparing the proportion of the genome affected by clonal or subclonal SCNAs and mirrored subclonal allelic imbalance (MSAI) in WGD and non-WGD tumours. Black bars indicate the median of each distribution. Two-sided Student’s t-tests were used for each comparison. c, Comparing the proportion of the genome affected by clonal or subclonal SCNAs in matched WGD and non-WGD samples from tumours with subclonal WGD. Bars indicate, for each patient with subclonal WGD, the difference between the median proportion of the genome affected by SCNAs in WGD and non-WGD samples. The inset beeswarm plots compare the proportion of the genome affected by different types of SCNAs in WGD and non-WGD samples. The black bars in the beeswarm plots represent the medians of each group. d–f, Impact of OG-TSG score on average arm-level copy number changes. Scatter plots showing the average subclonal arm-level change from MRCA in non-WGD (d; n = 171), WGD (e; n = 194) and subclonal WGD (f; n = 29) tumours versus arm OG–TSG score. Shaded areas indicate the 95% confidence interval. ρ and P values are from Spearman correlation tests. g, Scatter plot showing the average clonal (MRCA) copy number in the entire cohort (n = 394) versus chromosome arm size. h–j, Scatter plots showing the average subclonal arm-level change from MRCA in non-WGD (h; n = 171), WGD (i; n = 194) and subclonal WGD (j; n = 29) tumours versus chromosome size. Shaded areas indicate the 95% confidence interval. ρ and P values are from Spearman correlation tests.
Extended Data Fig. 5. Markov chain modelling of karyotype evolution.
a, List of parameters used for Markov chain modelling. b, Diagrams of simplified Markov chain for each chromosome arm and bar charts of the resulting probability distributions of arm-level copy number. c–e, Beeswarm plots showing the difference in deviance score on a per-tumour basis for non-WGD (n = 171), WGD (n = 194) and subclonal WGD (n = 29) tumours. Black horizontal bars indicate the median of the distribution. Paired two-tailed Student’s t-tests were performed between the deviance scores of the first and second model included in each comparison. es, effect size. c, Comparison between the unweighted (neutral) model and the weighted model that includes OG–TSG scores. d, Comparison between the unweighted model and the model with scrambled OG–TSG scores. e, Comparison between the weighted model that includes OG–TSG scores and the model with scrambled OG–TSG scores. f, g, For each context (non-WGD, WGD or subclonal WGD), the percentage of samples in which the OG–TSG-weighted model outperforms the unweighted model (f) or scrambled model (g) is shown. h–j, Robustness analysis of the Markov chain model of karyotype evolution. Graphs show the relative performance of the three iterations of the model with varying values of g with non-WGD (pGD = 0), WGD (pGD = 0.005) and subclonal WGD (pGD = 0.012) input. The model with scrambled scores has been run for 10 different random permutations of the chromosomes. k, l, Graphs show the performance of three iterations of the model with changing values of pGD (pGD = 0.003 in k and pGD = 0.007 in l) with WGD data. m, n, Graphs show the performance of three iterations of the model with changing values of pGD (pGD = 0.01 in m and pGD = 0.014 in n) with subclonal WGD data. o–q, Graphs show the performance of the three iterations of the model when varying pmisseg with non-WGD, WGD and subclonal WGD input data.
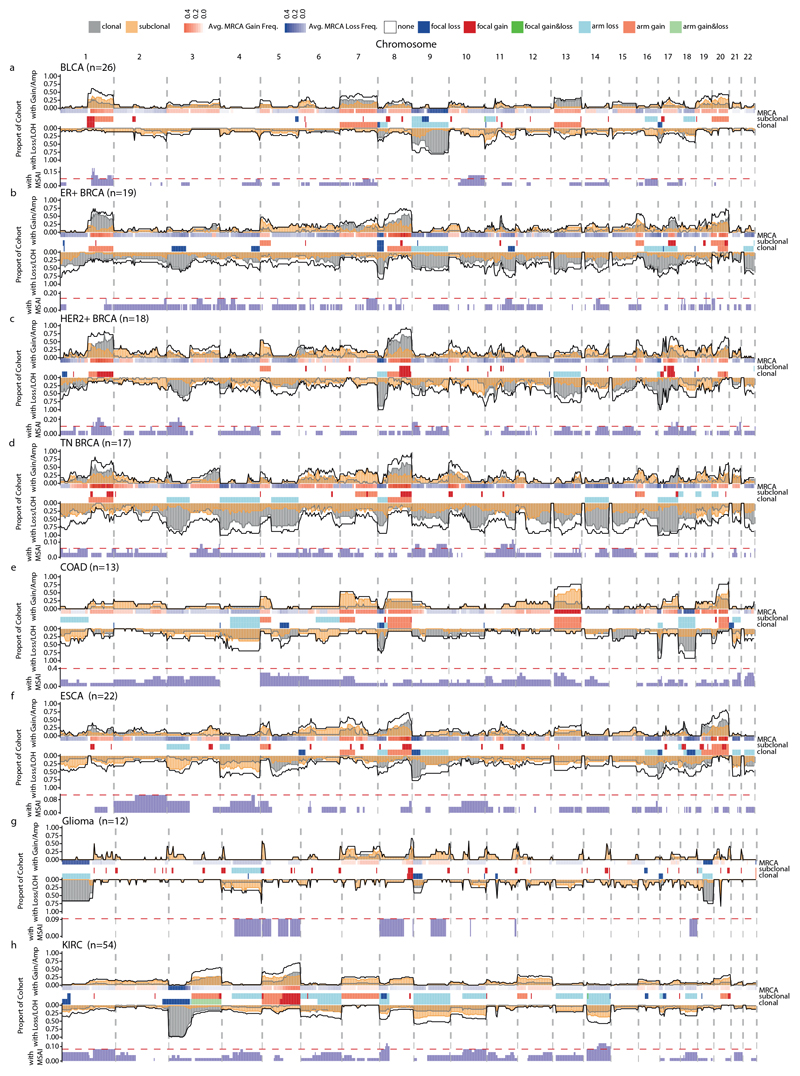
Extended Data Fig. 6. Subclonal SCNA landscape across tumour types.
a–h, The following tumour types were analysed: bladder urothelial carcinoma (a; n = 26), ER+ breast cancer (b; n = 19), HER2+ breast cancer (c; n = 18), triple-negative breast cancer (d; n = 17), colorectal adenocarcinoma (e; n = 13), oesophageal adenocarcinoma (f; n = 22), glioma (g; n = 12) and KIRC (h; n = 54). n numbers represent tumours. Across-genome plots show clonal and subclonal SCNAs. Within each tumour type for each chromosome, the following data are shown (top to bottom): the proportion of patients with gains or amplifications. The black line indicates the total proportion of patients with gains/amplifications; the yellow and grey lines or shades indicate the proportion of patients with subclonal and clonal gains, respectively. The MRCA was derived by phylogenetic analysis (see Methods, ‘Ancestral reconstruction and phylogeny inference’). For each locus, the frequency of gains (red) and losses (blue) found in the MRCAs of the tumours are indicated. The GISTIC2.0 events. These tracks indicate significant SCNA focal events that were identified by GISTIC2.0 (see Methods, ‘GISTIC2.0 peak definition’ and ‘GISTIC2.0 consensus peak definition’) and recurrent arm-level events (see Methods, ‘Arm-level SCNA definition’). The proportion of patients with loss/LOH events. The black line indicates the total proportion of patients with loss/LOH events; the yellow and grey lines or shades indicate the proportion of patients with subclonal and clonal losses, respectively. The black, yellow and grey lines indicate significance thresholds for total loss/LOH, subclonal loss/LOH and clonal loss/LOH, respectively. Proportion of patients with mirrored subclonal allelic imbalance (MSAI) originating from distinct haplotypes identified by multi-sample phasing. The red line indicates the significance threshold determined by a permutation test at the 0.05 level (see Methods, ‘Permutation test for recurrence of SCNA across tumours’).
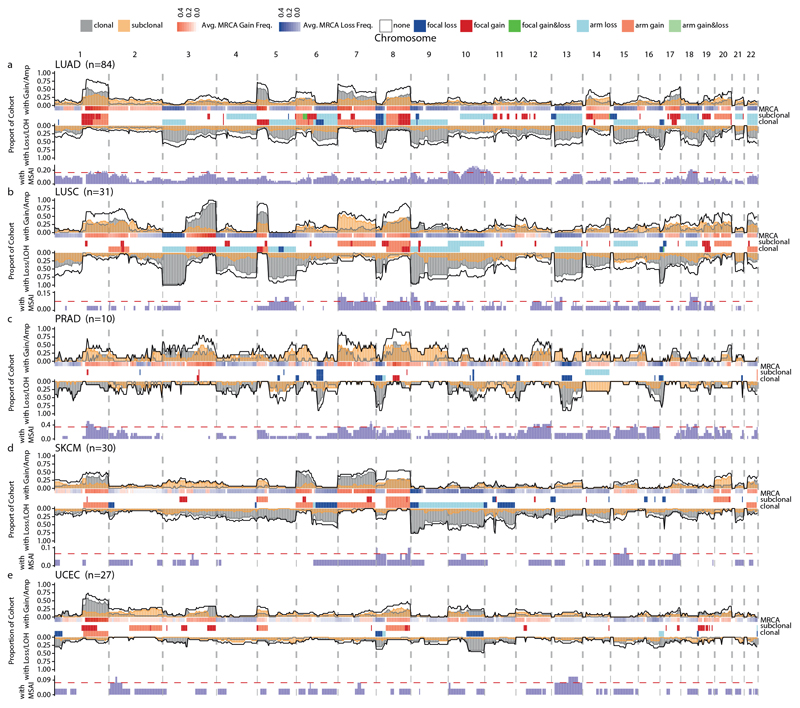
Extended Data Fig. 7. Subclonal SCNA landscape across tumour types.
a–e, The following tumour types were analysed: LUAD (a; n = 84), LUSC (b; n = 31), prostate adenocarcinoma (c; n = 10), SKCM (d; n = 30) and endometrial carcinoma (e; n = 27). Across-genome plots show clonal and subclonal SCNAs. Within each tumour type for each chromosome, the following data are shown (top to bottom): the proportion of patients with gains or amplifications. The black line indicates the total proportion of patients with gains/amplifications; the yellow and grey lines or shades indicate the proportion of patients with subclonal and clonal gains, respectively. The MRCA was derived by phylogenetic analysis (see Methods, ‘Ancestral reconstruction and phylogeny inference’). For each locus, the frequency of gains (red) and losses (blue) found in the MRCAs of the tumours are indicated. The GISTIC2.0 events. These tracks indicate significant SCNA focal events that were identified by GISTIC2.0 (see Methods, ‘GISTIC2.0 peak definition’ and ‘GISTIC2.0 consensus peak definition’) and recurrent arm-level events (see Methods, ‘Arm-level SCNA definition’). The proportion of patients with loss/LOH events. The black line indicates the total proportion of patients with loss/LOH events; the yellow and grey lines or shades indicate the proportion of patients with subclonal and clonal losses, respectively. The black, yellow and grey lines indicate significance thresholds for total loss/LOH, subclonal loss/LOH and clonal loss/LOH, respectively. Proportion of patients with mirrored subclonal allelic imbalance (MSAI) originating from distinct haplotypes identified by multi-sample phasing. The red line indicates the significance threshold determined by a permutation test at the 0.05 level (see Methods, ‘Permutation test for recurrence of SCNA across tumours’).
Extended Data Fig. 8. Recurrent SCNA across tumour types.
a, b, Difference in gains and losses in consensus-peak region gains (red, n = 255) and losses (blue, n = 149) (a) and chromosome arm gains (red, n = 95) and losses (blue, n = 200) across all tumour types (b). Black horizontal bars indicate the median of the distribution. Significance testing was performed using an unpaired Student’s t-test. c, Classification of chromosomal arm-level events according to timing. Left, heat map of the percentage of subclonal occurrence of all events in each tumour type. The numerator within each cell indicates, in that tumour type, the total number of subclonal occurrences of that event and the denominator indicates the total number of both clonal and subclonal occurrences of that event in that tumour type. Shading of each cell in the heat map indicates the percentage of subclonal occurrences of an event within a tumour type with orange indicating a higher subclonality and grey indicating a higher clonality. The border of each cell indicates the classification of that event in a tumour type as either early (grey border), intermediate (no border) or late (orange border). Right, bar plot of arm-level events ordered by median percentage of subclonal occurrences across tumour types (bottom axis). Bars representing gain events are coloured in red and loss events are coloured in blue. Horizontal black lines indicate separation of events into pan-cancer categories of early, intermediate and late, according to tertiles of the median proportion of SCNAs that is subclonal. Dots centred on the same axis positions indicate the total event count of each loss or gain event across tumour types (top axis). d, Enrichment of early, intermediate and late consensus peak events with known cancer-associated genes. Heat map indicating the resulting P values from two-sided Fisher’s exact tests comparing the overlap of genes in early, intermediate and late consensus peaks with previously reported oncogenes and tumour-suppressor genes. Gain peaks were investigated in relation to oncogenes, while loss peaks were investigated in relation to tumour-suppressor genes. Significant overlaps (Benjamini–Hochberg-adjusted P < 0.05) are indicated with an asterisk (see Methods, ‘Cancer-associated gene and fragile site enrichment’). e, Enrichment of early, intermediate and late consensus peak events with chromosome fragile sites. Heat map indicating the resulting P values from Fisher’s exact tests comparing the overlap of cytobands found in early, intermediate and late consensus peaks with cytobands from previously reported chromosome fragile sites. Significant overlaps (Benjamini–Hochberg-adjusted P < 0.05) are indicated with an asterisk (see Methods, ‘Cancer-associated gene and fragile site enrichment’). f, Prevalence of SNVs and indels in cancer-associated genes. Heat map displaying the proportion of samples from each tumour type with an SNV or indel in the corresponding cancer-associated gene. Yellow asterisks indicate where the SNVs and indels are present clonally in ≥75% of tumours in the corresponding tumour type.
Extended Data Fig. 9. Recurrent parallel evolution and LOH across the genome.
a, Across-genome plot showing the frequency of parallel gain/amplification events in red and frequency of parallel LOH events in blue. The dashed red lines indicate the significance threshold determined by a permutation test. b, Example of parallel evolution on chromosome 1 in CRUK0005. log2[R], B-allele frequency (BAF) and allele-specific expression (ASE) plots are shown for chromosome 1 in samples 3 and 4. On the phylogenetic tree, we indicate the branches in which the parallel gains of chromosome 1 were identified. c, Correlating intra-tumour heterogeneity (ITH) for each gene at the DNA and RNA levels. The scatter plot shows that the percentage of expressed genes with allele-specific DNA intratumour heterogeneity correlates with the percentage of expressed genes with allele-specific RNA intratumour heterogeneity. Only the 43 tumours, for which we had paired multi-sample exome-sequencing and multi-sample RNA sequencing data, were included in this analysis. d, Prevalence of single haploid copies in WGD tumours. Across-genome plot showing the frequency of loss to a single haploid copy in WGD tumours at the cytoband level. Clonal loss to a single haploid copy is shown in grey. Subclonal loss to a single haploid copy is shown in orange. The solid black line indicates the total frequency, including both clonal and subclonal events, of loss to a single haploid copy. HLA LOH is not shown as only the whole-exome sequencing subset of our cohort could be analysed using the LOHHLA bioinformatics tool (see Methods, ‘HLA LOH detection’). e, Prevalence of LOH in WGD tumours. This across-genome plot at the cytoband level shows the proportion of tumours with LOH. The solid black line indicates the total proportion of tumours with either subclonal or clonal LOH; the yellow shading indicates the proportion of tumours with WGD in the cohort that had subclonal LOH at these cytobands. The dashed grey lines demarcate the borders between separate chromosomes. f, Prevalence of HLA LOH across tumour types. We indicate for each tumour type the count and proportion of tumours in which HLA LOH was observed. Dark grey and orange bars show tumours for which HLA LOH was observed clonally or subclonally, respectively; light grey bars show tumours for which no HLA LOH was observed.
Extended Data Fig. 10. SCNAs in metastatic samples.
a, Beeswarm plot indicating the total proportion of the genome affected by either clonal or subclonal SCNAs in primary tumour samples (red dots) or metastatic samples (blue dots). The black bars indicate the median of the distribution. A two-sided unpaired Student’s t-test was used in this comparison; the P value and effect size(es) are shown. b, Difference in the percentage of the genome affected by SCNAs between paired metastatic and primary tumour samples (n = 152). The waterfall plot shows whether a greater or lesser proportion of the genome was affected by total SCNAs in the primary or metastatic sample(s) of tumours with at least one primary tumour sample and at least one metastatic sample. Purple bars indicate that a greater proportion of the genome was affected by total SCNAs in the metastatic sample and pink bars indicate a greater proportion was affected in the primary tumour sample. A two-sided paired Student’s t-test was used for this comparison. c, Beeswarm plots indicating, for each primary tumour and metastatic sample, the proportion of the genome impacted by SCNAs. These are the same samples included in the analysis of a. The black bars indicate the median of the distribution. Two-sided unpaired Student’s t-tests were used for each comparison; P values are indicated at the top of each plot. d, Beeswarm plots indicating for each primary tumour and metastatic sample the proportion of SCNAs that is subclonal. These are the same samples included in the analysis of a. The black bars show the median of the distribution. Two-sided unpaired Student’s t-tests were used for each comparison; P values are indicated at the top of each plot. e, Shared and private primary tumour and metastatic LOH. Bar plots separated by tumour type with each stacked bar representing the LOH identified in a single tumour sample with both primary tumour and metastatic samples. Each bar is coloured according to the proportion of LOH identified in that tumour that is shared between the primary tumour and metastatic samples (blue), the proportion of LOH present only in primary tumour samples (green) or the proportion of LOH present only in metastatic samples (red). The grey horizontal lines show the median value of the proportion of LOH shared between primary tumour and metastatic samples for each tumour type. f–i, Chromosomal arm-level events enriched in metastatic samples. We included only the four tumour types with >10 tumours with paired primary tumour–metastatic samples: LUAD (f), ER+ breast cancer (g), HER2+ breast cancer (h) and KIRC (i). In each panel, all chromosome arms are featured. The bar plots show the number of tumours with arm-level SCNAs in each tumour type. The colour of the bars indicates whether that arm-level event was enriched, depleted or maintained in the metastatic sample when compared with the corresponding primary tumour sample from the disease of the same patient. Bars facing right represent gain SCNAs; bars facing left represent loss SCNAs. The rectangular blocks between the bar plots indicate whether the arm-level events were recurrent events. Orange blocks represent recurrent subclonal events; grey blocks represent recurrent clonal events; blocks that are partially grey and partially orange represent events that are clonally and subclonally recurrent. The asterisks indicate whether the arm-level event is significantly enriched in metastatic samples in the combined paired (two-sided binomial test) and unpaired (test of equal or given proportions) primary tumour–metastatic analysis.
Supplementary Material
Acknowledgements
T.B.K.W. was supported by the Francis Crick Institute, which receives its core funding from Cancer Research UK (FC001169), the UK Medical Research Council (FC001169) and the Wellcome Trust (FC001169) as well as the Marie Curie ITN Project PLOIDYNET (FP7-PEOPLE-2013, 607722), Breast Cancer Research Foundation (BCRF), Royal Society Research Professorships Enhancement Award (RP/EA/180007) and the Foulkes Foundation. E.L.L. receives funding from NovoNordisk Foundation (ID 16584). N.J.B. is a fellow of the Lundbeck Foundation and acknowledges funding from the Aarhus University Research Foundation. E.G. is funded by the European Research Council, FP7-THESEUS-617844 and PROTEUS-835297. J.D. is a postdoctoral fellow of the Research Foundation–Flanders (FWO) and the European Union’s Horizon 2020 research and innovation program (Marie Skłodowska-Curie grant agreement no. 703594-DECODE). R.R. is supported by Royal Society Research Professorships Enhancement Award (RP/EA/180007). K.L. is supported by a UK Medical Research Council Skills Development Fellowship Award (grant number MR/P014712/1). L.Y. was funded by a Wellcome Trust Clinical Career Development Fellowship 214584/Z/18/Z and CRUK Early Detection Pump Prime Award. B.C.B. is supported by an NCI Outstanding Investigatory Award (1R35CA220481). G.B.J. is supported by the Swedish Cancer Society, Swedish Research Council and the Berta Kamprad Foundation. S.L. is supported by the National Breast Cancer Foundation of Australia Endowed Chair and the Breast Cancer Research Foundation, New York. N.M.L. and G.D.C. were supported by the Francis Crick Institute, which receives its core funding from Cancer Research UK (FC010110), the UK Medical Research Council (FC010110) and the Wellcome Trust (FC010110). S.T. is funded by Cancer Research UK (grant number C50947/A18176), the National Institute for Health Research (NIHR) Biomedical Research Centre at The Royal Marsden Hospital and Institute of Cancer Research (grant number A109), the Kidney and Melanoma Cancer Fund of The Royal Marsden Cancer Charity, and The Rosetrees Trust (grant number A2204). M.J.-H. has received funding from Cancer Research UK, National Institute for Health Research, Rosetrees Trust, UKI NETs and NIHR University College London Hospitals Biomedical Research Centre. P.V.L. is supported by the Francis Crick Institute, which receives its core funding from Cancer Research UK (FC001202), the UK Medical Research Council (FC001202) and the Wellcome Trust (FC001202) and is a Winton Group Leader in recognition of the Winton Charitable Foundation’s support towards the establishment of The Francis Crick Institute. S.F.B. is supported by the Office of the Director, the National Institutes of Health under award number DP5OD026395 High-Risk High-Reward Program, the Department of Defense Breast Cancer Research Breakthrough Award W81XWH-16-1-0315 (project: BC151244), the Burroughs Wellcome Fund Career Award for Medical Scientists, the Parker Institute for Immunotherapy at MSKCC, the Josie Robertson Foundation and MSKCC core grant P30-CA008748. R.F.S. and M.P. thank the Helmholtz Association (Germany) for support. N.M. is a Sir Henry Dale Fellow, jointly funded by the Wellcome Trust and the Royal Society (Grant Number 211179/Z/18/Z) and also receives funding from Cancer Research UK, Rosetrees and the NIHR BRC at University College London Hospitals and the CRUK University College London Experimental Cancer Medicine Centre. C.S. is Royal Society Napier Research Professor. His work was supported by the Francis Crick Institute, which receives its core funding from Cancer Research UK (FC001169), the UK Medical Research Council (FC001169), and the Wellcome Trust (FC001169). C.S. is funded by Cancer Research UK (TRACERx, PEACE and CRUK Cancer Immunotherapy Catalyst Network), Cancer Research UK Lung Cancer Centre of Excellence, the Rosetrees Trust, Butterfield and Stoneygate Trusts, NovoNordisk Foundation (ID16584), Royal Society Research Professorships Enhancement Award (RP/EA/180007), the NIHR BRC at University College London Hospitals, the CRUK-UCL Centre, Experimental Cancer Medicine Centre and the Breast Cancer Research Foundation (BCRF). This research is supported by a Stand Up To Cancer-LUNGevity-American Lung Association Lung Cancer Interception Dream Team Translational Research Grant (SU2C-AACR-DT23-17). Stand Up To Cancer is a program of the Entertainment Industry Foundation. Research grants are administered by the American Association for Cancer Research, the Scientific Partner of SU2C. C.S. also receives funding from the European Research Council (ERC) under the European Union’s Seventh Framework Programme (FP7/2007-2013) Consolidator Grant (FP7-THESEUS-617844), European Commission ITN (FP7-PloidyNet 607722), an ERC Advanced Grant (PROTEUS) from the European Research Council under the European Union’s Horizon 2020 research and innovation programme (835297) and Chromavision from the European Union’s Horizon 2020 research and innovation programme (665233). The results published here are based in part on data generated by The Cancer Genome Atlas pilot project established by the NCI and the National Human Genome Research Institute. The data were retrieved through database of Genotypes and Phenotypes (dbGaP) authorization (accession number phs000178.v9.p8). Information about TCGA and the constituent investigators and institutions of the TCGA research network can be found at http://cancergenome.nih.gov/. This project was enabled through access to the MRC eMedLab Medical Bioinformatics infrastructure, supported by the Medical Research Council (MR/L016311/1). In particular, we acknowledge the support of the High-Performance Computing at the Francis Crick Institute as well as the UCL Department of Computer Science Cluster and the support team. This publication and the underlying study have been made possible partly on the basis of the data that the Hartwig Medical Foundation and the Center of Personalised Cancer Treatment (CPCT-02, NCT01855477) and DRUP clinical study (NCT02925234) have made available to the project.
Footnotes
Contributions
T.B.K.W. and E.L.L. created the genomics pipeline, designed and conducted bioinformatics analyses and wrote the manuscript. M.P. performed phylogenetic analyses and MRCA reconstructions. S.E. designed and performed the Markov-chain modelling and analysis with S.F.B. providing further analysis and comments. N.J.B., G.A.W., J.D., S.C.D, S.H., K. Haase, M.E., R.R., H.X., K.L., T.P.M. and M.D. provided considerable bioinformatics support. D.A.M. analysed pathology mitotic index and anisonucleosis measurements. E.G., A.R., D.B., S.M.D. and W.T.L. critically assessed the biological soundness of the methods and results. L.A., M.A.B. and L.S. helped to analyse patient clinical characteristics. G.D.C., P.L., I.N., K. Harbst, F.C.-G., L.R.Y., F.C., F.J., C.V., I.P.M.T., P.K.B., R.J.C., B.C.B., L.D., G.B.J., P.S., S.L. and F.A. helped with data access and avenues of enquiry related to individual tumour types. N.S. and V.C.G.T.-H. collated data for the Hartwig Medical Foundation. Z.S., N.M.L., P.J.C. and P.V.L. helped to direct the avenues of bioinformatics analysis and gave feedback on the manuscript. S.T. and M.J.-H. designed study protocols and helped to analyse patient clinical characteristics. R.F.S., N.M. and C.S. jointly designed and supervised the study and helped to write the manuscript.
Competing interests
G.A.W. has consulted for and has stock options in Achilles Therapeutics. D.A.M. reports speaker fees from AstraZeneca. M.A.B. has consulted for Achilles Therapeutics. C.V. has received travel expenses from Astellas, Roche and Pfizer, and grant support from Bristol Myers Squibb. R.R. has consulted for and has stock options in Achilles Therapeutics. K.L. reports speaker fees from Roche Tissue Diagnostics. P.K.B. has consulted for Angiochem, Roche-Genentech, Eli Lilly, Tesaro, ElevateBio, Pfizer (Array), and received grant or research support from Merck, Bristol Myers Squibb and Eli Lilly and honoraria from Merck, Roche-Genentech and Eli Lilly. L.D. has sponsored research agreements with C2i-genomics, Natera, AstraZeneca and Ferring, and has an advisory/consulting role at Ferring. P.S. serves an uncompensated consultant for Roche-Genentech. S.L. receives research funding to her institution from Novartis, Bristol Myers Squibb, Merck, Roche-Genentech, Puma Biotechnology, Pfizer, Eli Lilly and Seattle Genetics, has acted as consultant (not compensated) to Seattle Genetics, Pfizer, Novartis, Bristol Myers Squibb, Merck, AstraZeneca and Roche-Genentech and has acted as consultant (paid to her institution) to Aduro Biotech, Novartis, GlaxoSmithKline and G1 Therapeutics. F.A. is a member of the Advisory Boards for Pfizer, AstraZeneca, Eli Lilly, Roche-Genentech, Novartis and Daiichi Sankyo, acknowledges grant support from Pfizer, AstraZeneca, Eli Lilly, Novartis and Daiichi Sankyo and is a co-founder of Pegacsy. V.C.G.T.-H. reports grants and personal fees from Pfizer, Roche, Novartis and Eli Lilly, grants from Eisai and personal fees from Accord. S.T. has received funding from Ventana Medical Systems Inc (grant numbers 10467 and 10530), has received speaking fees from Roche, AstraZeneca, Novartis and Ipsen and has the following European and US patent filed: Indel mutations as a therapeutic target and predictive biomarker (PCTGB2018/051892) and European patent: Clear Cell Renal Cell Carcinoma Biomarkers (P113326GB). M.J.-H. is a member of the Advisory Board for Achilles Therapeutics. S.F.B. holds a patent related to some of the work described targeting CIN and the cGAS-STING pathway in advanced cancer, owns equity in, receives compensation from and serves as a consultant and on the Scientific Advisory Board and Board of Directors of Volastra Therapeutics, and has also consulted for Sanofi, received sponsored travel from the Prostate Cancer Foundation, and both travel and compensation from Cancer Research UK. N.M. has stock options in and has consulted for Achilles Therapeutics and holds a European patent in determining HLA LOH (PCT/GB2018/052004). C.S. acknowledges grant support from Pfizer, AstraZeneca, Bristol Myers Squibb, Roche-Ventana, Boehringer-Ingelheim, Archer Dx Inc (collaboration in minimal residual disease sequencing technologies) and Ono Pharmaceutical, is an AstraZeneca Advisory Board Member and Chief Investigator for the MeRmaiD1 clinical trial, has consulted for Pfizer, Novartis, GlaxoSmithKline, MSD, Bristol Myers Squibb, Celgene, AstraZeneca, Illumina, Genentech, Roche-Ventana, GRAIL, Medicxi and the Sarah Cannon Research Institute, has stock options in Apogen Biotechnologies, Epic Bioscience, GRAIL, and has stock options and is co-founder of Achilles Therapeutics. C.S. holds European patents relating to assay technology to detect tumour recurrence (PCT/GB2017/053289); to targeting neoantigens (PCT/EP2016/059401), identifying patent response to immune checkpoint blockade (PCT/EP2016/071471), determining HLA LOH (PCT/GB2018/052004), predicting survival rates of patients with cancer (PCT/GB2020/050221), identifying patients who respond to cancer treatment (PCT/GB2018/051912), a US patent relating to detecting tumour mutations (PCT/US2017/28013) and both a European and US patent related to identifying insertion/deletion mutation targets (PCT/GB2018/051892).
Contributor Information
Roland F. Schwarz, Email: roland.schwarz@mdc-berlin.de.
Nicholas McGranahan, Email: nicholas.mcgranahan.10@ucl.ac.uk.
Charles Swanton, Email: charles.swanton@crick.ac.uk.
Data availability
TRACERx sequencing datasets used in this paper are described in previous studies7,39. Details of all other datasets obtained from third parties used in this study can be found in Supplementary Table 1. Clinical trial information (if applicable) is also available within the associated publications described in Supplementary Table 1.
Code availability
All code used for analyses was written in R version 3.6.1 and is available at: https://bitbucket.org/schwarzlab/refphase/. The Markov-chain modelling code and associated data can be found here: https://math.dartmouth.edu/~sergi/mathbio.php.
References
- 1.Zack TI, et al. Pan-cancer patterns of somatic copy number alteration. Nat Genet. 2013;45:1134–1140. doi: 10.1038/ng.2760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Bolhaqueiro ACF, et al. Ongoing chromosomal instability and karyotype evolution in human colorectal cancer organoids. Nat Genet. 2019;51:824–834. doi: 10.1038/s41588-019-0399-6. [DOI] [PubMed] [Google Scholar]
- 3.Davoli T, et al. Cumulative haploinsufficiency and triplosensitivity drive aneuploidy patterns and shape the cancer genome. Cell. 2013;155:948–962. doi: 10.1016/j.cell.2013.10.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Turajlic S, et al. Deterministic evolutionary trajectories influence primary tumor growth: TRACERx Renal. Cell. 2018;173:595–610. doi: 10.1016/j.cell.2018.03.043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.McGranahan N, et al. Cancer chromosomal instability: therapeutic and diagnostic challenges. ‘Exploring aneuploidy: the significance of chromosomal imbalance’ review series. EMBO Rep. 2012;13:528–538. doi: 10.1038/embor.2012.61. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Schwarz RF, et al. Spatial and temporal heterogeneity in high-grade serous ovarian cancer: a phylogenetic analysis. PLoS Med. 2015;12:e1001789. doi: 10.1371/journal.pmed.1001789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Jamal-Hanjani M, et al. Tracking the evolution of non-small-cell lung cancer. N Engl J Med. 2017;376:2109–2121. doi: 10.1056/NEJMoa1616288. [DOI] [PubMed] [Google Scholar]
- 8.Hieronymus H, et al. Tumor copy number alteration burden is a pan-cancer prognostic factor associated with recurrence and death. eLife. 2018;7:e37294. doi: 10.7554/eLife.37294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Carter S, et al. A signature of chromosomal instability inferred from gene expression profiles predicts clinical outcome in multiple human cancers. Nat Genet. 2006;38:1043–1048. doi: 10.1038/ng1861. [DOI] [PubMed] [Google Scholar]
- 10.Schwarz RF, et al. Phylogenetic quantification of intra-tumour heterogeneity. PLOS Comput Biol. 2014;10:e1003535. doi: 10.1371/journal.pcbi.1003535. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.von der Thüsen JH, et al. Prognostic significance of predominant histologic pattern and nuclear grade in resected adenocarcinoma of the lung: potential parameters for a grading system. J Thorac Oncol. 2013;8:37–44. doi: 10.1097/JTO.0b013e318276274e. [DOI] [PubMed] [Google Scholar]
- 12.Kadota K, et al. Comprehensive pathological analyses in lung squamous cell carcinoma: single cell invasion, nuclear diameter, and tumor budding are independent prognostic factors for worse outcomes. J Thorac Oncol. 2014;9:1126–1139. doi: 10.1097/JTO.0000000000000253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Laughney AM, Elizalde S, Genovese G, Bakhoum SF. Dynamics of tumor heterogeneity derived from clonal karyotypic evolution. Cell Rep. 2015;12:809–820. doi: 10.1016/j.celrep.2015.06.065. [DOI] [PubMed] [Google Scholar]
- 14.Elizalde S, Laughney AM, Bakhoum SF. A Markov chain for numerical chromosomal instability in clonally expanding populations. PLOS Comput Biol. 2018;14:e1006447. doi: 10.1371/journal.pcbi.1006447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Sottoriva A, et al. A Big Bang model of human colorectal tumor growth. Nat Genet. 2015;47:209–216. doi: 10.1038/ng.3214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Williams MJ, Werner B, Barnes CP, Graham TA, Sottoriva A. Identification of neutral tumor evolution across cancer types. Nat Genet. 2016;48:238–244. doi: 10.1038/ng.3489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.López S, et al. Interplay between whole-genome doubling and the accumulation of deleterious alterations in cancer evolution. Nat Genet. 2020;52:283–293. doi: 10.1038/s41588-020-0584-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Fujiwara T, et al. Cytokinesis failure generating tetraploids promotes tumorigenesis in p53-null cells. Nature. 2005;437:1043–1047. doi: 10.1038/nature04217. [DOI] [PubMed] [Google Scholar]
- 19.Bielski CM, et al. Genome doubling shapes the evolution and prognosis of advanced cancers. Nat Genet. 2018;50:1189–1195. doi: 10.1038/s41588-018-0165-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.McGranahan N, et al. Allele-specific HLA loss and immune escape in lung cancer evolution. Cell. 2017;171:1259–1271. doi: 10.1016/j.cell.2017.10.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Snyder A, et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N Engl J Med. 2014;371:2189–2199. doi: 10.1056/NEJMoa1406498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kim M, et al. Comparative oncogenomics identifies NEDD9 as a melanoma metastasis gene. Cell. 2006;125:1269–1281. doi: 10.1016/j.cell.2006.06.008. [DOI] [PubMed] [Google Scholar]
- 23.Cai Y, et al. Loss of chromosome 8p governs tumor progression and drug response by altering lipid metabolism. Cancer Cell. 2016;29:751–766. doi: 10.1016/j.ccell.2016.04.003. [DOI] [PubMed] [Google Scholar]
- 24.Bakhoum SF, et al. Chromosomal instability drives metastasis through a cytosolic DNA response. Nature. 2018;553:467–472. doi: 10.1038/nature25432. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Lackner C, et al. Convergent evolution of copy number alterations in multi-centric hepatocellular carcinoma. Sci Rep. 2019;9:4611. doi: 10.1038/s41598-019-40843-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Jakubek YA, et al. Large-scale analysis of acquired chromosomal alterations in non-tumor samples from patients with cancer. Nat Biotechnol. 2020;38:90–96. doi: 10.1038/s41587-019-0297-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Zaccaria S, Raphael BJ. Characterizing the allele- and haplotype-specific copy number landscape of cancer genomes at single-cell resolution with CHISEL. Nat Biotechnol. 2020 doi: 10.1038/s41587-020-0661-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Shih DJH, et al. Genomic characterization of human brain metastases identifies drivers of metastatic lung adenocarcinoma. Nat Genet. 2020;52:371–377. doi: 10.1038/s41588-020-0592-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Turner KM, et al. Extrachromosomal oncogene amplification drives tumour evolution and genetic heterogeneity. Nature. 2017;543:122–125. doi: 10.1038/nature21356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Worrall JT, et al. Non-random mis-segregation of human chromosomes. Cell Rep. 2018;23:3366–3380. doi: 10.1016/j.celrep.2018.05.047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Van Loo P, et al. Allele-specific copy number analysis of tumors. Proc Natl Acad Sci USA. 2010;107:16910–16915. doi: 10.1073/pnas.1009843107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Howie BN, Donnelly P, Marchini J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 2009;5:e1000529. doi: 10.1371/journal.pgen.1000529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Nik-Zainal S, et al. The life history of 21 breast cancers. Cell. 2012;149:994–1007. doi: 10.1016/j.cell.2012.04.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Gundem G, et al. The evolutionary history of lethal metastatic prostate cancer. Nature. 2015;520:353–357. doi: 10.1038/nature14347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Yates LR, et al. Subclonal diversification of primary breast cancer revealed by multiregion sequencing. Nat Med. 2015;21:751–759. doi: 10.1038/nm.3886. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Yates LR, et al. Genomic evolution of breast cancer metastasis and relapse. Cancer Cell. 2017;32:169–184. doi: 10.1016/j.ccell.2017.07.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Mitchell TJ, et al. Timing the landmark events in the evolution of clear cell renal cell cancer: TRACERx Renal. Cell. 2018;173:611–623. doi: 10.1016/j.cell.2018.02.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Martinez P, et al. Parallel evolution of tumour subclones mimics diversity between tumours. J Pathol. 2013;230:356–364. doi: 10.1002/path.4214. [DOI] [PubMed] [Google Scholar]
- 39.Rosenthal R, et al. Neoantigen-directed immune escape in lung cancer evolution. Nature. 2019;567:479–485. doi: 10.1038/s41586-019-1032-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Dobin A, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21. doi: 10.1093/bioinformatics/bts635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Li B, Dewey CN. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinformatics. 2011;12:323. doi: 10.1186/1471-2105-12-323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Castel SE, Mohammadi P, Chung WK, Shen Y, Lappalainen T. Rare variant phasing and haplotypic expression from RNA sequencing with phASER. Nat Commun. 2016;7:12817. doi: 10.1038/ncomms12817. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Rimmer A, et al. Integrating mapping-, assembly- and haplotype-based approaches for calling variants in clinical sequencing applications. Nat Genet. 2014;46:912–918. doi: 10.1038/ng.3036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Bailey MH, et al. Comprehensive characterization of cancer driver genes and mutations. Cell. 2018;173:371–385. doi: 10.1016/j.cell.2018.02.060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Forbes SA, et al. COSMIC: exploring the world’s knowledge of somatic mutations in human cancer. Nucleic Acids Res. 2015;43:D805–D811. doi: 10.1093/nar/gku1075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Hartigan JA, Hartigan PM. The dip test of unimodality. Ann Stat. 1985;13:70–84. [Google Scholar]
- 47.Maechler M. diptest: Hartigan’s dip test statistic for unimodality—corrected. R package version 0.75-7. 2015 https://cran.r-project.org/package=diptest.
- 48.Wolff AC, et al. Recommendations for human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists clinical practice guideline update. J Clin Oncol. 2013;31:3997–4013. doi: 10.1200/JCO.2013.50.9984. [DOI] [PubMed] [Google Scholar]
- 49.Mermel CH, et al. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 2011;12:R41. doi: 10.1186/gb-2011-12-4-r41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Fungtammasan A, Walsh E, Chiaromonte F, Eckert KA, Makova KD. A genome-wide analysis of common fragile sites: what features determine chromosomal instability in the human genome? Genome Res. 2012;22:993–1005. doi: 10.1101/gr.134395.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Wang K, et al. PennCNV: an integrated hidden Markov model designed for high-resolution copy number variation detection in whole-genome SNP genotyping data. Genome Res. 2007;17:1665–1674. doi: 10.1101/gr.6861907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Cheng J, et al. Single-cell copy number variation detection. Genome Biol. 2011;12:R80. doi: 10.1186/gb-2011-12-8-r80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Whitfield ML, et al. Identification of genes periodically expressed in the human cell cycle and their expression in tumors. Mol Biol Cell. 2002;13:1977–2000. doi: 10.1091/mbc.02-02-0030.. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Abbosh C, et al. Phylogenetic ctDNA analysis depicts early-stage lung cancer evolution. Nature. 2017;545:446–451. doi: 10.1038/nature22364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Priestley P, et al. Pan-cancer whole-genome analyses of metastatic solid tumours. Nature. 2019;575:210–216. doi: 10.1038/s41586-019-1689-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Moulos P, Hatzis P. Systematic integration of RNA-seq statistical algorithms for accurate detection of differential gene expression patterns. Nucleic Acids Res. 2015;43:e25. doi: 10.1093/nar/gku1273. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
TRACERx sequencing datasets used in this paper are described in previous studies7,39. Details of all other datasets obtained from third parties used in this study can be found in Supplementary Table 1. Clinical trial information (if applicable) is also available within the associated publications described in Supplementary Table 1.
All code used for analyses was written in R version 3.6.1 and is available at: https://bitbucket.org/schwarzlab/refphase/. The Markov-chain modelling code and associated data can be found here: https://math.dartmouth.edu/~sergi/mathbio.php.