Abstract
Melioidosis caused by the facultative intracellular pathogen Burkholderia pseudomallei is difficult to treat due to poor intracellular bioavailability of antibiotics and antibiotic resistance. In the absence of novel compounds, polymersome (PM)-encapsulation may increase the efficacy of existing antibiotics and reduce antibiotic resistance by promoting targeted, infection-specific intracellular uptake. In this study we developed PMs composed of widely available polyethylene oxide-polycaprolactone (PEO-PCL) block co-polymers and demonstrated their delivery to intracellular B. thailandensis infection using multispectral imaging flow cytometry (IFC) and coherent anti-Stokes Raman scattering (CARS) microscopy. Antibiotics were tightly sequestered in PMs and did not inhibit the growth of free-living B. thailandensis. However, on uptake of antibiotic-loaded PMs by infected macrophages, IFC demonstrated PM co-localisation with intracellular B. thailandensis and a significant inhibition of their growth. We conclude that PMs are a viable approach for the targeted antibiotic treatment of persistent intracellular Burkholderia infection.
Keywords: intracellular bacteria, antibiotics, nanoparticles, polymersomes, Raman spectroscopy, CARS imaging, imaging flow cytometry
Introduction
Melioidosis is a common but neglected tropical disease with a high disease burden caused by the Gram-negative bacterial pathogen Burkholderia pseudomallei. Melioidosis presents with non-specific signs and symptoms making diagnosis difficult, and may result in septicaemia, pneumonia and encephalitis, leading to severe illness and death.1 B. pseudomallei is endemic in northern Australia and south-east Asia with an estimated incidence of 165,000 cases/year and a mortality of 89,000 deaths/year.2 There is no vaccine for B. pseudomallei,3 and it is designated as a Centers for Disease Control and Prevention (CDC) ‘select agent’ - a potential target as a biowarfare agent.4 A feature of the pathogenicity of this species is its ability to survive intracellularly within non-phagocytic and phagocytic cells, especially macrophages.5 Following phagocytosis B. pseudomallei rapidly escapes from the phagolysosome and replicates in the host cell cytosol.6,7 The bacterium can move transcellularly in vitro actin-based motility and spreads between adjacent cells using a type VI secretory system, causing cell fusion and multi-nucleated giant cell formation, a hallmark of Burkholderia infection.8 This benefits the bacteria by avoidance of the innate immune system9 and protection from the adaptive immune system, particularly antibodies, which do not penetrate host cell membranes readily.10
B. pseudomallei infection is hard to treat due to the bacterium’s ability to survive intracellularly. Antibiotics, such as aminoglycosides and β-lactams, have poor intracellular bioavailability as they penetrate intact host cell membranes poorly.11 This is compounded by the intrinsic antibiotic resistance of B. pseudomallei, conferred by the expression of efflux pump proteins and β-lactamases.12 Treatment typically relies upon long-term antibiotic therapy with combinations of antibiotics, but even after prolonged antibiotic therapy there can be a re-emergence of infection, likely due to incomplete bacterial clearance.13 This creates its own challenges where access and adherence to prolonged antibiotic treatment regimens is difficult, leading to toxic side-effects, and the requirement for sustained antibiotic use, which in turn results in increased selection pressure for antibiotic resistance.14,15 Together, this highlights the need to develop new antimicrobial strategies for melioidosis, which improve the delivery, release and pharmacokinetics of already-available antibiotics. This strategy may also lead to the use of previously disregarded antibiotics in front-line melioidosis treatment due to their perceived poor pharmacokinetics.
We are investigating the utility of polymersome (PM) nanoparticles to address these challenges. PMs are composed of amphiphilic co-polymers, which self-assemble to form vesicles upon contact with aqueous solutions. These vesicles have a hydrophobic shell membrane, surrounding a hydrophilic aqueous core,16–18 enabling packaging of both hydrophobic and hydrophilic compounds. PMs have thick hydrophobic membranes, which reduces their permeability to encapsulated hydrophilic drug payloads and provides a large reservoir for otherwise insoluble hydrophobic payloads.16 PMs are more stable than other carriers, such as liposomes, which are rapidly cleared by the body.19 Furthermore, in contrast to other classes of polymeric nanoparticles20,21 their chemical properties can be controlled to enable responsiveness to stimuli such as temperature,22,23 pH,24,25 UV,26 and intracellular redox environments.27 These advantages have led to interest in PMs as drug-delivery vehicles for a range of applications, most frequently oncology.28–30 There are several polymeric micelle nanoparticle formulations in both Phase I and II clinical trials31 but, to the best of our knowledge, no PM preparations have yet been approved for use as a therapy.
PMs have only recently begun to be explored as a method of intracellular antibiotic delivery. PMs comprising pH-sensitive PMPC-PDPA di-block copolymers have been used to encapsulate metronidazole and doxycycline for delivery to Porphyromonas gingivalis in oral epithelial cells.32 More recently, rifampicin-loaded PMs composed of PEO-PCL copolymers were tested for delivery to Mycobacterium tuberculosis infected cells,33 work that has been extended using similar PM chemistries to deliver rifampicin for targeting M. tuberculosis infection in zebrafish34 and mice.35 Intracellular targeting and inhibition of Mycobacterium spp., Staphylococcus aureus and access to tuberculosis (TB) granulomas in zebrafish36 has been shown using antibiotics encapsulated in PMPC-PDPA PMs. The only published study to our knowledge to date that has focussed on developing PMs to target Burkholderia spp. used pH-dependent self-assembling polymer brushes for delivery of ceftazidine to infected macrophages.37 Such approaches, however, often rely on complex polymer chemistry to engineer functionality. It is notable that simpler and more translatable chemistries are yet to be exploited for use against Burkholderia spp.
Here, we hypothesised that PMs made from simple off-the-shelf, long circulating co-polymers could be used to stably sequester antibiotics and release them in direct proximity to intracellular Burkholderia thailandensis. We reasoned that the intracellular acidic and enzymatic conditions would be sufficient for drug release without complex chemistries for engineering co-polymers. To test this, we developed PEO-PCL PMs and used a suite of imaging techniques, including multispectral imaging flow cytometry (IFC) and coherent anti-Stokes Raman scattering (CARS) microscopy, to probe interactions between PMs and bacteria. IFC enables high-throughput, subcellular quantitative analysis of co-localisation between labelled molecules inside cells38 and is an attractive and unexplored technique to measure intracellular nanoparticle-bacteria interactions, while CARS is a label-free vibrational technique enabling direct and highly selective intracellular identification of endogenous and exogenous chemical species.39 Finally we test the efficacy of antibiotic-loaded PMs on subsequent inhibition of intracellular bacteria growth.
Results/Discussion
Doxycycline and rifampicin can be stably encapsulated in PEO-PCL polymersomes
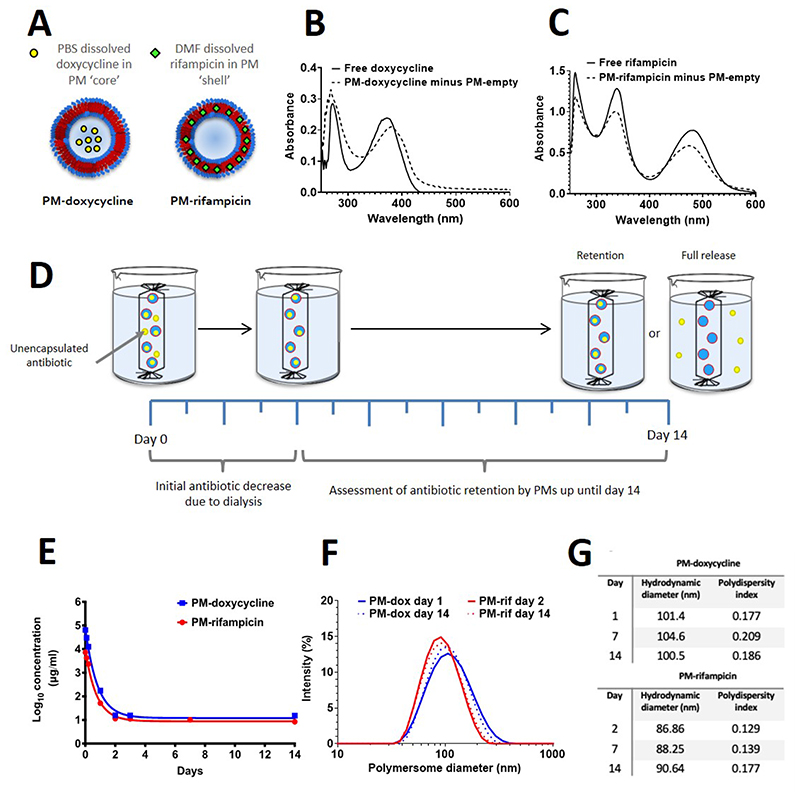
We first tested the hypothesis that PEO(5k)-PCL(18k) PMs can stably encapsulate antibiotic payloads. Stable, spherical PMs can be produced using this copolymer,18 and we have previously demonstrated incorporation of both hydrophilic or hydrophobic small molecules by solvent-switch nanoprecipitation.17,40 By similar means, doxycycline and rifampicin were loaded by solubilisation into either the aqueous phase (for doxycycline, PBS; ‘core’ loading), or into the organic solvent, (for rifampicin, DMF; ‘shell’ loading) prior to nanoprecipitation (Figure 1A). Doxycycline is used for the treatment of melioidosis,41 and rifampicin has been shown to have activity against B. pseudomallei 42 and is in use as an antibiotic for TB, which has similar latency to melioidosis. Encapsulation may provide benefits for tolerability and optimised drug regimens and so these antibiotics were chosen as examples of an antibiotic in use for melioidosis presently and one that may be re-purposed by encapsulation. A Rayleigh scattering peak was present in absorption spectra for all PM preparations at >300 nm, declining up to 500 nm (Supplementary Figure S1). Subtraction of spectra obtained from unloaded PM preparations (PM-empty) from either doxycycline loaded or rifampicin-loaded PM preparations (‘PM-doxycycline’ and ‘PM-rifampicin’) resolved spectra similar to free, solubilised doxycycline or rifampicin. This was reflected in characteristic peaks at 372 nm for doxycycline (Figure 1B) or 338 nm and 484 nm for rifampicin (Figure 1C), and indicates encapsulation of each antibiotic.
Figure 1. Encapsulation and retention of doxycycline and rifampicin by PMs.
(A) Model for the different PM loading strategies for the antibiotics rifampicin and doxycycline. (B) The absorption spectrum of free doxycycline is similar to that of its spectrum when loaded into PMs after subtraction of unloaded PMs (C) The same is true of rifampicin. (D) A schematic of the assay used to assess PM retention of the antibiotic after a 14-day period. Samples were left to dialyse for 14 days, with regular antibiotic concentration reading time points. (E) After initial burst release, PM-doxycycline maintained an encapsulated concentration of 15.2 μg/ml ± 2.2 μg/ml and PM-rifampicin 8.5 μg/ml ± 2.4 μg/ml. Data points plotted show the mean of three formulations, n = 3. Graph transformed using one phase decay analysis. (F) DLS data displaying the PM-antibiotic hydrodynamic radius distributions over 14 days. (G) The size and PdI values of PM-doxycycline and PM-rifampicin over a 14-day period. DLS data based on measurements from one batch of each type of PMs, performed in three technical repeats.
To determine whether these antibiotics could be stably retained by PMs, PMs were dialysed for 14 days in sink conditions and the amount of antibiotic retained in each preparation was measured by absorption spectroscopy at regular time-points (Figure 1D), with concentrations calculated from standard curves for free antibiotics. For both PM-rifampicin and PM-doxycycline, there was an initial sharp decrease in concentration between day 0 and day 3, which reflects removal of unincorporated antibiotic by dialysis (Figure 1E). After day 3, there was no further decrease in antibiotic concentration for up to 14 days, which indicates stable retention of antibiotic in each preparation. Confirming this, filtration of PMs by size-exclusion revealed negligible antibiotic in the filtrate, indicating all antibiotic detected by UV-vis spectroscopy was retained stably in PMs (Supplementary data, Figure S2). At 14 days of dialysis, the bulk concentration of doxycycline in PM-doxycycline preparations was 15.2 ± 2.2 μg/mL (34 μM), while the bulk concentration of rifampicin in PM-rifampicin preparations was 8.5 ± 2.4 μg /mL (10 μM) (n = 3). Assuming no polymer is lost in dialysis and using an estimated average formula mass of the polymer of 23 kDa, this results in a final molar ratio of doxycycline to di-block copolymer of 0.26 and for rifampicin of 0.07. Dynamic light scattering (DLS) analysis showed that both PM-doxycycline and PM-rifampicin nanoparticles were also stable in size over 14 days of dialysis, with no significant change in hydrodynamic radius for PM-doxycycline, but a small but significant increase in size for PM-rifampicin (n =3; p < 0.01) (Figure 1F and G). PM-doxycycline preparations were significantly larger in hydrodynamic radius than those of PM-rifampicin (n = 3; p < 0.001). Using multi-angle dynamic light scattering (MADLS), we found the concentrations of nanoparticles to be 9.26 × 1010/mL for PM-doxycycline preparations, and 3.98 × 1011/mL for PM-rifampicin, similar to that for PM-empty samples (4.19 × 1011/mL). In addition we tested the stability of PM-doxycycline after freezing at either – 20 °C or – 196 °C in the presence and absence of 30 % (w/v) sucrose, before thawing and exposure to 24 hours dialysis. Freeze-thawing had no significant effect on either PM diameter or on encapsulated antibiotic concentration when sucrose was present, whereas in the absence of sucrose there was visibly increase turbidity in samples frozen at 196 °C (Supplementary data, Figure 3). This indicates the potential for long term storage of this formulation.
It is important to note that bulk measurements of the concentration of antibiotics in PM preparations do not reflect the local concentration of antibiotic in suspended nanoparticles, as the distribution of the antibiotic in the bulk is partitioned in dispersed nanoparticles (Supplementary data, Figure S4A and B). To produce more concentrated preparations we used ultracentrifugation, a method to concentrate PMs that has been shown not to adversely affect the physical characteristics of PMs.43 By doing so, we were able to produce stable 1200-fold concentrated suspensions containing 38.1 and 29.1 mg/mL (85 mM and 35 mM) rifampicin and doxycycline, respectively (Supplementary data, Figure S4C). Size was not significantly affected by ultracentrifugation (Supplementary data, Figure S4D).
These data extend the findings of other groups who have attempted to encapsulate antibiotics in PMs. Moretton et al. demonstrated incorporation of rifampicin in PEO-PCL PMs with di-blocks ranging from 4k – 10k for PEO and 7.2k to 12.8k for PCL33 by solvent switching. These authors did not measure PM concentration, but were able to demonstrate final bulk concentrations of 1.9 mg/mL, a higher concentration than would be possible by solubilisation in aqueous buffer, but less than the concentration achieved in the current study. More recently, Trousil et al. formulated rifampicin in MPEO(5k)-PCL(4k) PMs and, although the concentration of rifampicin retained in PMs was not calculated, rifampicin retained activity in intracellular and in vivo infection assays.34,35 Antibiotic-containing PM preparations have been formulated using other di-block co-polymers including DMAEMA-HPMA37, PMPC-PDA32, or PMPC-PDPA44 which are pH-responsive, again with most extensive data available for rifampicin, likely due to PM incorporation improving its solubility in aqueous medium. Notably, in the latter study, antibiotic content per PM was calculated at 100-100,000 molecules per (PMPC-PDPA) PM. Our data are comparable with this study, and based on concentration measurements, we calculate 430 molecules of rifampicin per PM, which is broadly similar to these previous studies, and 6130 molecules of doxycycline per PM, which has not to our knowledge been successfully encapsulated in PMs previously.
PEO-PCL polymersomes are internalised by RAW 264.7 macrophages and partition to endosomes
Macrophages provide an intracellular niche for the proliferation, spread and survival of Burkholderia and therefore intracellular targeting of antibiotics to these cells is a means of treating human disease. There is extensive published evidence that PMs are taken up rapidly by macrophages.44,45 As expected, using confocal microscopy we found that fluorescent puncta became visible rapidly in RAW 264.7 cells incubated for 4 hours with DiI-labelled PMs (Figure 2A). There was negligible staining in the nucleus, indicating that PMs were primarily localised to the cytoplasm. PM-DiI nanoparticles were filtered to ensure there was no unencapsulated dye remaining within samples that might contribute to PM-independent membrane staining (Supplementary data, Figure S5).
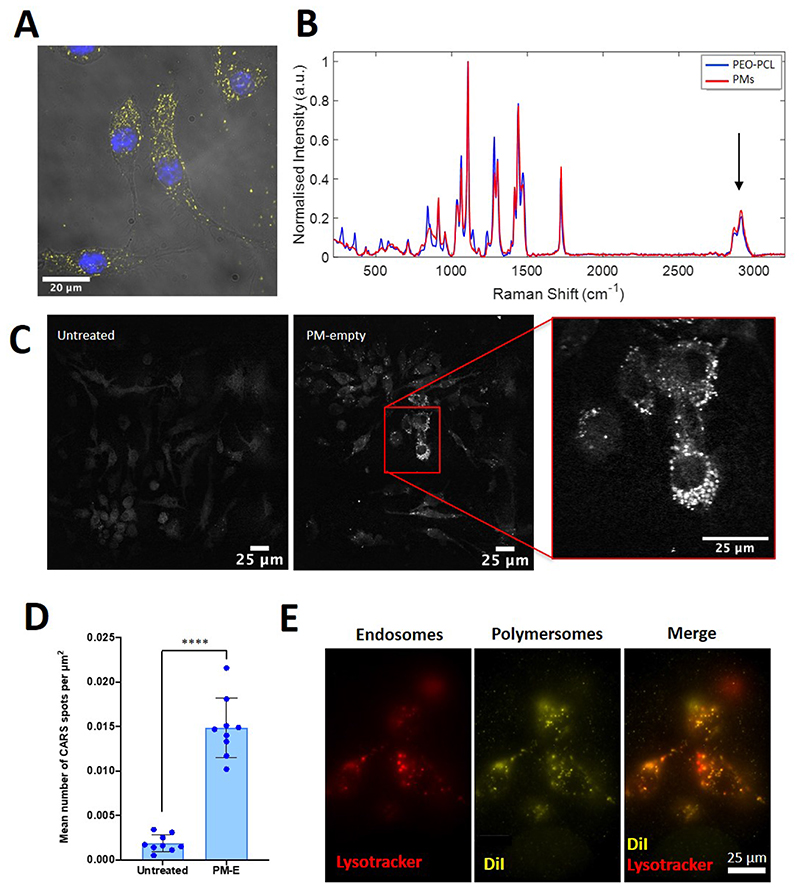
Figure 2. Uptake of PMs by RAW 264.7 macrophage cells using label-free imaging.
(A) After incubation with DiI-labelled PMs, RAW 264.7 cells show cytoplasmic fluorescent puncta. (B) The Raman spectra for both PMs and the PEO-PCL polymer. A characteristic peak is present at approximately 2800 cm-1 indicating C-H bonds (C) CARS label-free imaging showing a comparison of untreated RAW 264.7 cells versus RAW 264.7 cells exposed to PM-empty nanoparticles. (D) PM-empty treated RAW 264.7 cells showed a significantly higher mean number of CARS spots per μm2 compared to the untreated control, p < 0.0001. CARS quantitative analysis data shows the mean and SD of one biological repeat performed in triplicate. (E) DiI fluorescent puncta in RAW 264.7 cells co-localise with lysosomes labelled with Lysotracker.
One challenge of using fluorescent membrane stains to aid nanoparticle imaging is the possible dissociation of the stain from the nanoparticle, leading to non-specific uptake of dye. Coherent anti-Stokes Raman scattering (CARS) imaging is a vibrational technique that allows for label-free imaging of biological samples. CARS has previously been used to image nanomaterials in cells and tissues including ammonium palmitoyl glycol chitosan micelles,46 polystyrene nanoparticles,47,48 poly lactate-co-glycolide (PLGA)49 and methylmethacrylate nanoparticles,50 but to our knowledge there are no reports of the use of CARS for label free imaging of polymeric vesicles such as PMs. Typically, in order to visualise cellular details, a CARS frequency of 2845 cm-1 is used to excite C-H bonds in cellular lipids.51 We found that free PMs also give rise to a significant signal at this frequency due to the high number of C-H bonds present in the polymer (Figure 2B). Due to the polymer’s high density of C-H bonds, we hypothesised that label-free identification of intracellular PMs may be possible in cells, despite a contribution from biological molecules. After incubation of macrophages with PMs, we observed puncta (at the CARS frequency of 2845 cm-1 inside cells), which were not visible in control cells (Figure 2C). To highlight and visualise the puncta (bright spots) the intensity thresholds were defined for all images to remove background contributions. The pattern of staining is similar to that obtained by DiI-stained PMs, and likely reflects the PM’s C-H bonds in the PEO-PCL. Quantitative image analysis also confirmed that PM-empty treated cells had a significantly higher number of CARS spots than untreated cells, 0.015 spots/μm2 vs. 0.0019 spots/μm2, respectively (n = 9; p < 0.0001) (Figure 2D). Additionally, the mean area of CARS spots (μm2) per unit cell area (μm2) was also significantly higher in PM-empty treated cells compared to the untreated control at 3.6 × 10-3 compared to 3.6 × 10-4, respectively (n = 9; p < 0.0001).
Finally, to determine intracellular partitioning of labelled PMs we labelled PM-exposed cells with Lysotracker. It was evident at 30 minutes that some but not all DiI-labelled PMs localised to lysosomes (Figure 2E). These data are similar to that observed in other studies, for example, mesoporous silica52 or PLGA53 nanoparticles have been shown to partially partition to lysosomes within hours after uptake. Together these data confirm that PMs are taken up by RAW 264.7 macrophage cells and that they partition at least partially to intracellular endosomes.
B. thailandensis-infection of RAW 264.7 macrophages is heterogeneous and increases following exposure
Previous studies have shown that macrophages infected with M. bovis retain the ability to take up solid polymer nanoparticles53 or PMs,44 but there are no data available to our knowledge on Burkholderia-infected cells. In addition, there are conflicting data on whether intracellular bacteria and endocytosed nanoparticles occupy similar intracellular spaces. In some studies PLGA or poly(butyl cyanoacrylate) nanoparticles and M. bovis have been shown to remain in separate compartments53,54 whereas in other studies a significant degree of co-localisation has been shown to occur.34,55 These discrepancies may result from differences in the nanoparticle chemistries, physical properties or biological systems used, but may also result from subjective analysis using confocal microscopy and/or TEM. The fact that this latter, low throughput mode of analysis is most frequently used to determine PM uptake and subcellular distribution also means that it is challenging to quantify uptake and co-localisation.
To overcome this limitation, and to better characterise PM uptake and subcellular partitioning in B. thailandensis-infected macrophages, we employed ImageStream multispectral imaging flow cytometry (IFC), a high-throughput technique that allows rapid imaging of subcellular features of cells during flow cytometric analysis. We incubated RAW 264.7 cells with GFP-expressing B. thailandensis (a containment level 2 model for B. pseudomallei) to promote intracellular infection, before removing extracellular bacteria using kanamycin. Confocal microscopy revealed a heterogeneous pattern of cellular bacterial infection after exposure to bacteria (Figure 3A).
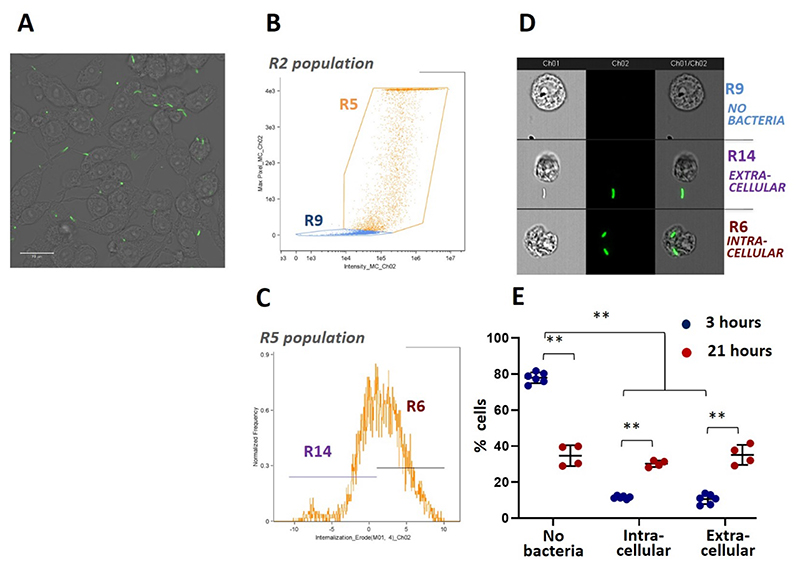
Figure 3. Intracellular infection of macrophages by B. thailandensis is heterogeneous and increased with time.
(A) Confocal imaging shows GFP-expressing B. thailandensis are present within RAW 264.7 cells after 3 hours incubation in the presence of kanamycin. Scale bar = 20 μm (B) Gating strategy for separating populations of B. thailandensis-positive (R5) and negative (R9) cells. (C) Gating strategy for separating RAW 264.7 cells with intracellular (R6) or extracellular (R14) B. thailandensis. (D) Representative ImageStream images of RAW264.7 cells with no bacteria present (R9), extracellular bacteria (R14) or intracellular bacteria (R6). Scale bar = 7 μM (E) There is a decrease in the percentage of cells with no bacteria present and an increase in those with either intracellular or extracellular bacteria between 3 and 21 hours.
To establish a strategy to quantify PM uptake based on intracellular localisation and burden of B. thailandensis we first used IFC to separate populations based on the following criteria: (1) cells in which bacteria were present intracellularly; (2) cells in which bacteria were present extracellularly; and (3) cells where no bacteria were present, either extracellularly or intracellularly. This stratification is not possible using conventional flow cytometry. Single, in-focus cells (defined as the R2 population) were first gated for positivity or negativity for bacteria within an ImageStream field of view (Figure 3B; R5 and R9, respectively). Cells positive for bacteria (R5) were subsequently defined as containing intracellular bacteria or extracellular bacteria (Figure 3C, R6 and R14) using the Erode function, as described in the methods. Examples of images obtained by this gating strategy are shown in Figure 3D. A detailed explanatory plan of the gating strategy used is shown in Supplementary Figure 6.
By quantifying events in each gated populations, we found that 3 hours following exposure to B. thailandensis, 77.9 ± 3.0 % of macrophages were not associated with bacteria, while intracellular bacteria were present in 11.5 ± 0.8 % of cells, and extracellular bacteria were present in 10.5 ± 2.8 % of cells (Figure 3E). At 21 hours, the proportion of macrophages with no bacteria associated had declined by a factor of ~ 0.4 to 34.7 ± 5.8 % of cells, whereas the proportion of cells with intra- or extracellular bacteria present had increased by a factor of ~3 to 30.2 ± 1.8 % and 35.1 ± 5.6 %, respectively. This indicates that during the course of culture there is an increase in the frequency of cells associated with bacteria, an unsurprising finding considering that B. thailandensis reproduce intracellularly and spread from cell-to-cell in macrophage culture.
B. thailandensis-infected RAW 264.7 macrophages take up PEO-PCL polymersomes
To quantify PM uptake in B. thailandensis-exposed macrophages, control and bacteria-exposed cells were incubated with DiD-labelled PMs. In B. thailandensis-exposed cells, DiD fluorescence was observed in regions separate from and directly adjacent or co-incident to areas containing B. thailandensis (green) (Figure 4A), indicating that antibiotic loaded PMs may be targeted to the same cellular compartments as the bacteria they are intended to inhibit, and that close interaction of PMs with bacteria during infection may occur. RAW 264.7 macrophages took up PMs rapidly, with almost all cells positive for DiD after 3 hours regardless of whether or not they had been exposed to B. thailandensis (Figure 4B). To determine the effect of intracellular bacterial burden on PM uptake, we measured PM uptake in each of the three populations defined above by measuring cellular DiD fluorescence intensity for each cell. There was a significant increase in cellular DiD fluorescence with respect to time in all populations, indicating increased PM uptake during the culture, regardless of bacterial infection (Figure 4C and D). In the bacteria-exposed population, the greatest increase in uptake was in cells with intracellular bacteria present, rather than in cells where there were no bacteria (by a factor of 2.7 ± 0.5 versus 2.0 ± 0.5, p =0.02, n = 6). In all cases there was significantly reduced uptake of PMs in bacteria-exposed macrophages compared to control, unexposed macrophages (by a factor of 0.44 ± 0.1, 0.78 ± 0.1, and 0.65 ± 0.1 for macrophages with no bacteria, intracellular bacteria or extracellular bacteria, respectively (p < 0.0001)).
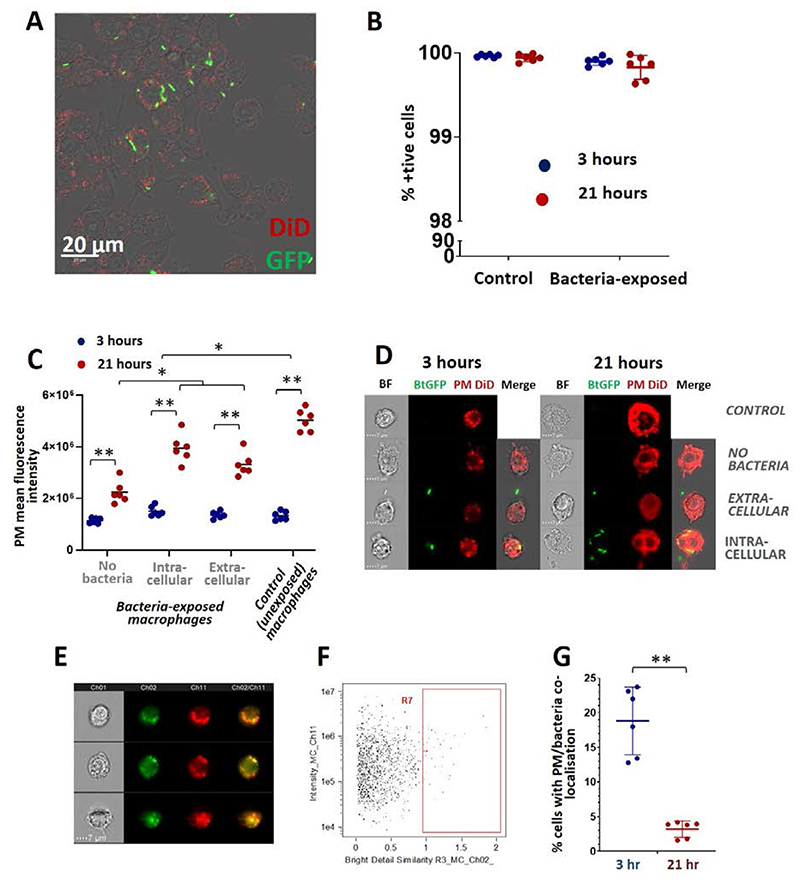
Figure 4. Imaging flow cytometry analysis of PM-DiD uptake by RAW 264.7 macrophage cells.
(A) Fluorescent DiD puncta are visible in RAW 264.7 cells infected with GFP-labelled B. thailandensis. (B) Almost all RAW 264.7 cells are positive for DiD-labelled polymersomes after 3 hours (C) In bacteria-exposed populations, cells which contain intracellular bacteria take up the most PMs and those with no bacteria present take up the least, as measured by DiD fluorescence intensity. All exposed populations take up significant fewer PMs compared to unexposed macrophages. (* indicates p<0.01 and ** <0.001; data reflect means of 6 independent experiments). (D) Representative ImageStream images from the populations shown in (C). (E) Fluorescent puncta of DiD labelled PMs (Ch11) partially colocalise with GFP-positive bacteria, as shown by representative ImageStream images. (F) Gating strategy to separate RAW 264.7 cells with high co-localisation compared to low co-localisation. (G) The degree of PM/bacterial colocalisation declined from 3 hours to 24 hours. All ImageStream data is taken from three biological repeats, n=3, each performed in duplicate. ** = p < 0.0001.
These data suggest that exposure to B. thailandensis impairs the ability of RAW 264.7 cells to take up PMs. We were initially surprised by this observation as macrophages, activated by the presence of bacteria, become primed to engulf bacteria by phagocytosis. We hypothesised this activation would also promote nanoparticle uptake.56 However, there is evidence to suggest that nanoparticles smaller than 200 nm in diameter are preferentially taken up not by phagocytosis but by other endocytotic means, such as (macro)pinocytosis, in macrophages.57 As some data suggest that micropinocytosis is inhibited in activated macrophages,58 we suggest that this may be one explanation for the reduction in PM uptake in bacteria-exposed macrophages.
It is possible that the reason for the greater uptake in infected vs. uninfected cells (in cells that had been exposed to bacteria) may be related to a compromise in membrane integrity that occurs concurrent with B. thailandensis intracellular pathogenesis. Burkholderia spp. have diverse pathogenic effects on host cells, including co-option of actin polymerisation machinery, and induction of apoptotic genes or necrosis.59 Alternatively, penetration of the cell membrane by bacteria during cell-to-cell transmission may lead to membrane rupture, making cells more permeable in general to PMs. Supporting this, we found that there was a correlation between the intracellular bacteria burden and PM uptake in infected cells (r = 0.23; p< 0.0001; Supplementary Figure 7).
Together, these data indicate that exposure to bacteria causes a reduction in PM uptake regardless of whether intracellular infection has occurred, with the greatest reduction not in infected macrophages, but rather in macrophages which have no bacteria present. This supports the notion that bacteria-exposed cells are refractory to PM uptake.
PEO-PCL polymersome co-localise intracellularly with B. thailandensis
IFC allowed us to assess the intracellular association of PMs and B. thailandensis to determine if PMs and bacteria co-localise to the same intracellular compartment. Co-localisation was calculated by comparing fluorescence signal from both GFP and DiD channels from ImageStream images (Figure 4E).
Co-localisation in these images appears as orange puncta, highlighting an overlap of the green GFP fluorescence with the red DiD. Pearson’s correlation coefficient was used to generate a value between 0 and 1, representing no co-localisation or perfect co-localisation, respectively, and plotted as ‘Bright Detail Similarity’ and gated as the R7 populations (Figure 4F). Using this analysis we found there to be a significantly higher level of co-localisation at the early, 3-hour timepoint, with 18.8% of cells showing co-localisation, compared to the 21-hour timepoint, with only 3.2% of cells showing evidence of co-localisation (p < 0.0001; Figure 4G). This suggests that PMs occupy a similar intracellular niche to bacteria at 3 hours but thereafter this co-localisation declines.
This change in co-localisation may reflect early shuttling of PMs into lysosomes which later fuse with phagosomes containing bacteria. Supporting this, high-resolution electron microscopy analysis identified that solid nanoparticles localise on the surface of bacteria in phagosomes and phagolysosomes.60 Although it is known that PMs are trafficked to endosomes and lysosomes following cellular uptake,61 there is little data available on whether PMs may co-localise with bacteria in the same manner. Our data now provide clear evidence that PMs are trafficked to locations where bacteria are present in RAW264.7 cells, an advantageous feature for applications where delivery of a PM-containing antibiotic at high concentration is required. Furthermore, this determines a rationale for the use of IFC for determining co-localisation of different nanoparticles and bacteria in mammalian cells. IFC presents a number of advantages over conventional flow cytometry or automated fluorescent microscopy, including the ability to determine co-localisation of bacteria and nanoparticles in a quantitative, unbiased manner.
PM-doxycycline and PM-rifampicin preparations do not inhibit the growth of free living B. thailandensis
Our previous data indicated that doxycycline and rifampicin are tightly sequestered by PMs. To determine whether these antibiotics were bioavailable to free living B. thailandensis we tested growth inhibition of PM-antibiotic preparations compared to equivalent concentrations of antibiotic alone. While free antibiotics inhibited B. thailandensis growth, with rifampicin effective at concentrations above 12.5 μg/mL, and doxycycline above 1.6 μg/mL (Figure 5A), neither PM preparation had any effect on bacterial growth, even at the highest concentrations (Figures 5B). Since the bulk antibiotic concentration in PM-treated wells was above the MIC, we conclude that the antibiotics remain sequestered tightly and stably within PMs and are not bioavailable to bacteria. These findings are in contrast to data describing greater potency for PM-encapsulated rifampicin compared with free antibiotic.34 This was ascribed to an increase in bacterial permeability due to the surfactant nature of the co-polymer. It is possible that differences in the PM chemistries may explain this difference, as well as in the antibiotic used. But we also note that we performed stringent dialysis prior to use of PMs to ensure no free antibiotic remained, whereas previous reports34 separate the PMs using filtration, which may explain some of the observed differences.
Figure 5. PM-antibiotic preparations do not inhibit growth of free B. thailandensis.
(A) Both unloaded antibiotics were effective at inhibiting growth at the higher concentrations tested, at the 24-hour time point. Data is presented as the mean and SEM of one biological repeat tested in triplicate, n=1. (B) PM-rifampicin preparations and PM-doxycycline preparations do not inhibit the growth of B. thailandensis growing free in L-broth culture, at any concentration tested. Data is presented as the mean and SEM of one biological repeat tested in triplicate.
PM-doxycycline and PM-rifampicin preparations inhibit the growth of intracellular B. thailandensis
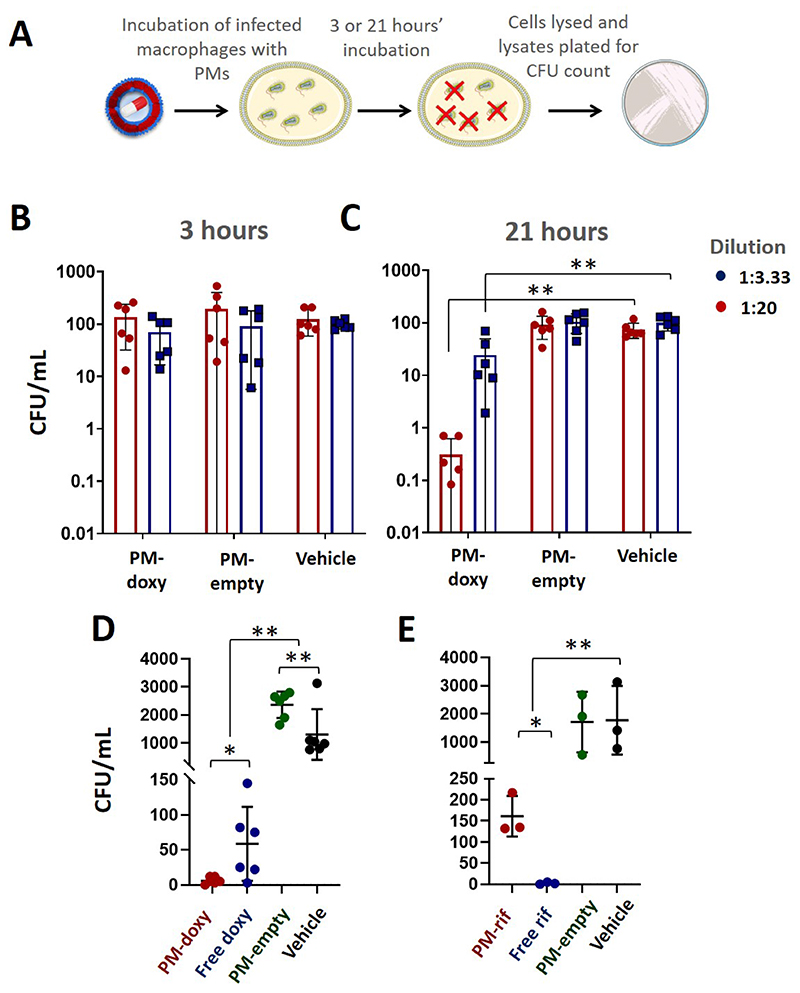
We next hypothesised that intracellular degradation of PMs may be required to release antibiotics and that macrophage uptake and intracellular lysosomal action may facilitate this release. PCL-PEG PMs contain a hydrolytically-sensitive PCL polymer block, and therefore are susceptible to degradation by lysozyme and other intracellular hydrolytic enzymes.62,63 We have previously demonstrated that an encapsulated hydrophilic fluorophore, fluorescein, is only released once PMs are taken up by mammalian cells, and that the PMs are otherwise stable in solution.64 We therefore tested whether PM-encapsulated doxycycline could inhibit the growth of intracellular bacteria by exposing B. thailandensis-infected macrophages to antibiotic PM preparations before lysing cells and measuring bacterial colony forming units (CFU; Figure 6A). PM-doxycycline was added at particle concentrations of 4.63 × 109/mL and 2.78 × 1010/mL (1:20 and 1:3.33 dilutions of neat, uncentrifuged suspensions), corresponding to final estimated antibiotic concentrations of 0.75 and 4.5 μg/mL, calculated from standard curves of absorbance. At 3 hours, there was no effect of test or control PMs in reducing recovered CFUs, which remained at ~100/mL of lysed cells, (Figure 6B). However, after 21 hours, there was a significant decrease in intracellular B. thailandensis burden compared to the bacteria only control group, where no PM preparation was added, with a 99.6% decrease at a 1:3.33 dilution, and a 75.5% decrease at a 1:20 dilution (p < 0.001, two-way ANOVA with Tukey’s test; Figure 6C). These data suggest that, similarly to fluorophores64 or kinase inhibitors,17 intracellular uptake of PMs leads to release of active compound which would otherwise be tightly sequestered. We consider it possible that the release of active doxycycline is facilitated by enzymatic degradation of the PMs in the cellular pathways of endocytosis. Most other studies on PM-based intracellular delivery have used pH-sensitive co-polymers, which undergo physical disassembly (rather than enzymatic degradation) upon exposure to the lower pH of lysosomes to trigger release.32,36 Our results indicate that pH sensitivity is certainly not a prerequisite for delivery, and are in agreement with those from other studies that demonstrate robust intracellular delivery33 and activity34 with simpler PEO-PEG chemistries. PCL is susceptible to the activity of esterases and/or lipases rather than hydrolysis, and therefore we consider it likely that lysosomal enzymes such as lysosomal acid lipase are responsible for this action65 although to our knowledge this is yet to be formally tested.
Figure 6. PM-antibiotic preparations reduce intracellular B. thailandensis burden.
(A) A schematic showing the assay used to assess intracellular killing. (B) After a 3-hour incubation of infected cells with PM-doxycycline there was no inhibition of intracellular growth observed. (C) After a 21-hour incubation, there was a significant level of intracellular bacterial killing. Data is presented as the mean and SD of data from two independent experiments, n=2, performed in triplicate and normalised to the 1:20 control group. (D) PM-doxycycline (1:3.33 dilution) significantly reduced intracellular bacteria burden compared to free doxycycline (4.5 μg/mL; p <0.05), unloaded PMs or vehicle controls (p <0.01) n=6; two independent experiments, mean and SD shown. (E) PM-rifampicin significantly reduced intracellular bacteria burden compared to unloaded PMs or vehicle controls (p <0.01). Free rifampicin (2.6 μg/ml) reduced intracellular bacterial burden compared to PM-rifampicin, PM-empty or vehicle controls n=3, mean and SD shown.
Having established that PM-encapsulated doxycycline inhibits the intracellular growth of B. thailandensis, we compared its activity to PM-encapsulated rifampicin and free antibiotic controls. PM-rifampicin was used at a concentration of 1.20 × 1011 particles/mL, corresponding to a final bulk antibiotic concentration of 2.6 μg/mL. While PM-loaded doxycycline reduced intracellular bacteria burden more so than free antibiotic (Figure 6D), the opposite was true in the case of rifampicin (Figure 6E). There are two likely explanations for this. First, based on experiments on free living B. thailandensis, rifampicin has an order of magnitude lower minimum inhibitory concentration than doxycycline. Combined with the approximately 2-fold lower PM encapsulation of rifampicin compared to doxycycline, this has the corollary that less rifampicin may be available to bacteria following uptake and intracellular release. Second, rifampicin is much less soluble than doxycycline in aqueous solution and may as a result be more tightly associated with the PCL block of the co-polymer and therefore less efficiently release. Nevertheless, both preparations of PM-loaded antibiotics were effective in inhibiting the growth intracellular B. thailandensis providing compelling evidence for the development of these formulations for melioidosis treatment.
Conclusions
In summary, we have demonstrated that PEO-PCL PMs are a robust and stable method for the specific intracellular nanodelivery of antibiotics to B. thailandensis. PM preparations retain antibiotics in complex media but release active antibiotic upon uptake by macrophages, a process that is likely promoted by the enzymatic breakdown of PM components in endosomes and/or lysosomes. Imaging flow cytometry indicates that early and direct co-localisation between PMs and bacteria occurs intracellularly, with PM uptake confirmed by tracking intracellular distribution of polymer using the label-free method of coherent anti-Raman stokes scattering microscopy. As a result, both PM-doxycycline and PM-rifampicin preparations reduce the macrophage cell bacterial burden compared to control groups. Our data provide evidence that optimised, simple-to-produce PM formulations of antibiotics are a viable approach for development for clinical use.
Materials/Experimental
All reagents were purchased from Sigma Aldrich, UK, unless stated otherwise.
Polymersome formulation
PMs were made by first dissolving 6 mg of polyethylene oxide(5k)-b-polycaprolactone(18k) (PEO-PCL) (Polymer Source Inc., Canada) in 0.4 mL of dimethylformamide (DMF). The solution was then sonicated for ≈20-30 minutes to aid dissolution. The resulting solution was added dropwise using a syringe driver at 0.75 mL/min (≈ 1 drop every 8 seconds), under stirring, to a vial containing 1.6 mL of phosphate-buffered saline (PBS) (Fisher Scientific, UK). The final 2 mL solutions were then placed into sealed cellulose-membrane dialysis tubing prepared as per the manufacturer’s instructions (14,000 MWCO, Sigma Aldrich D9277-100FT), and dialysed in excess Dulbecco’s PBS (500 mL, pH 7.4, 0.1 mM, Sigma-Aldrich). Solutions were dialysed for a minimum of 48 hours, with at least 3 PBS changes. Dialysed samples were stored at 4 °C until ready to be used. For antibiotic-loaded PMs doxycycline or rifampicin were dissolved into the PBS fraction or DMF fractions to final preparation concentrations of 50 mg/mL or 10 mg/mL, respectively. Lipophilic membrane dyes DiI and DiD (Thermo Fisher Scientific, UK) were, as separate formulations, dissolved into the DMF fraction at a preparation concentration of 50 μM. PMs were concentrated by ultracentrifugation at 180,000 × g (55,000 RPM) for 2 hours (Optima MAX-XP, Beckman Coulter, USA). For studies addressing the effect of frozen storage, PM formulations were either frozen by placement in a -20 °C freezer or by flash-freezing in liquid nitrogen at -196 °C in the presence and absence of 30 % (w/v) sucrose. Frozen formulations were then stored for 24 hours (for -20 °C treatment) or within 2 hours (for flash-frozen samples) before defrosting at room temperature. For controls, PMs were stored at 4 °C for the same time period. Following defrosting, samples were dialysed overnight at 4 °C as above, before measurements using DLS or absorbance spectroscopy, as described below.
Antibiotic encapsulation and release
The concentration of antibiotic incorporated into the PMs was determined by measuring UV absorbance of samples in black-sided quartz cuvettes on a NanoDrop 2000c UV-vis spectrophotometer (Thermo Fisher Scientific, UK). Absorbance was measured at wavelengths between 190 nm and 840 nm to determine the absorbance spectrum of the free, unencapsulated antibiotic, and then characteristic peaks were used to construct standard curves. PM preparations were mixed with a 1:1 ratio of DMF to ensure disruption of PM structure in order to reduce interference from nanoparticle Rayleigh scattering. Encapsulated antibiotic concentrations were determined using the generated standard curves. Release of antibiotic from PMs was assessed under continuous dialysis. Samples were dialysed at room temperature over a 14-day period, with repeated buffer changes. Aliquots of PM samples were removed at the time-points indicated in the results, and residual antibiotic concentration was measured by UV-vis, as above. To measure undialysed and unencapsulated antibiotic in the preparations, PM samples were spun through a centrifugal spin filter with a molecular weight cut-off of 10 000 Da (Fisher Scientific, UK) to filter PMs from their surrounding buffer. The filtrate UV-vis spectra was then measured on the NanoDrop and assessed for residual antibiotic signals.
PM characterisation
PM size and polydispersity was measured using dynamic light scattering (DLS; Zetasizer Nano ZS ZEN3600, Malvern, UK). PM solutions were diluted 1:10 in PBS prior to measurement. Readings were taken at 20 °C, with light detected at a scattering angle of 173°. 10 data recordings were made per sample, with an acquisition of 10 seconds for each. Each sample measurement was repeated in triplicate. PM particle concentration was measured using a Zetasizer Ultra Multi-angle dynamic light scattering (MADLS), instrument (ZSU5700, Malvern, UK). Measurement angles were at 173°, 13° and 90°. Attenuator, cuvette position and sub runs determined automatically by machine.
Cell culture
RAW 264.7 murine macrophages (purchased from European Collection of Authenticated Cell Culture [ECACC], Porton Down, UK) were maintained in Dulbecco’s modified Eagle’s medium (DMEM) (Gibco, UK), supplemented with 10% (v/v) heat-inactivated fetal bovine serum (FBS) (Gibco, UK) and 2 mM L-glutamine (Gibco, UK). The cells were incubated at 37°C, 5% CO2, and approximately 95% humidity. RAW cells were subcultured by cell scraping at ≈ 80% confluence. During infection assays cells were grown in Leibovitz-L15 media (Gibco, UK) supplemented with 10% inactivated FBS and 2 mM glutamine and incubated at 37 °C without CO2.
Preparation of bacterial culture
B. thailandensis E555 pBHR4-groS-eGFP bacteria were routinely grown in 50 mL of Luria broth (L-broth) or on Luria agar (L-agar) plates both supplemented with 50 μg/mL chloramphenicol. Broth cultures were incubated for 16-18 hours at 37 °C with orbital shaking at 180 rpm. Plates were incubated at 37 °C for 48 hours to allow countable colonies.
Epifluorescence and confocal microscopy
For epifluorescence imaging, RAW 264.7 macrophage cells were seeded in glass-bottom culture dishes at 42,000 cells/cm2 and were then incubated overnight at 37 °C. DiI-labelled PMs were added to cultures at a dilution of 1:4 and incubated for 4 hours at 37 °C. Cells were then fixed in 4% (w/v) paraformaldehyde (PFA) before Hoechst stain was added for 10 minutes at 1 μg/mL. Cells were imaged on a Zeiss Axio Imager.M2m with a 63x oil immersion objective. The DAPI (Hoechst) excitation filter used was: FF01-357/44-25 with an emission filter of FF01-460/80-25; the rhodamine (DiI) excitation filter used was: FF01-536/40-25 with an emission filter of FF01-593/40-25.
For super resolution microscopy using Nanoimager S (Oxford Nanoimaging Ltd, Oxford, UK), cells were seeded as with the epifluorescence method, and PM-DiD were added to cells to a dilution of 1:16. Lysotracker dye (Thermo Fisher, UK) was added to a final concentration of 74 nM. Cells were imaged for a period of 25 minutes. The microscope was fitted with Olympus UPlanSApo 100x oil immersion objective (1.49 NA).
For confocal imaging, RAW 264.7 cells were seeded at a density of 500,000 cells/mL (1.25 × 105/cm2), and incubated overnight at 37 °C. To infect cells with B. thailandensis bacteria were diluted in L-15 medium to a concentration of 5 × 107 bacteria/mL, measured using a Genesys 10 UV scanning spectrophotometer (Thermo Fisher, UK), and applied to cells for a period of 1 hour (multiplicity of infection (MOI) = 100). Following this, media was changed and kanamycin added at a concentration of 1 mg/mL to kill extracellular bacteria. DiD-loaded PMs were added added to the infected cell culture at a dilution of 1:10. Cells and PMs were incubated for 3 hours prior to imaging. Cells were imaged with a Zeiss LSM 710 confocal microscope with a x63 objective (Zeiss, Germany).
Coherent anti-Stokes Raman scattering (CARS) imaging
RAW 264.7 macrophage cells were seeded at 50,000 cm-2 on chambered coverslips and cultured overnight at 37 °C and 5 % CO2 in phenol-red-free DMEM. Medium was replaced with fresh phenol-red-free DMEM media in the presence or absence of 1.26 × 1011/mL PMs and incubated at 37 °C for 21 hours. A custom in-house built system CARS system was used for imaging as previously described in Moura et al..39 Briefly, a fundamental IR fibre laser (1031 nm, 2 picosecond, 80 MHz, Emerald Engine, APE) was coupled into a Nikon Ti eclipse inverted microscope with a 40× water immersion objection (1.15 NA). A portion of the 1031 nm IR laser was frequency doubled and used to synchronously pump an optical parametric oscillator (OPO) (APE, Levante Emerald, 650-950 nm). This created a tunable pump beam. Excitation wavelengths of 1031 nm (Stokes) and 797.2 nm (pump) were used. This made for a CARS imaging frequency of 2845 cm-1 (C-H stretch). The spectral resolution of the imaging system is approximately 10 cm-1. CARS signal was collected using a bandpass filter (643 ± 20 nm) and detected with a PMT (Hamamatsu H10722-20). Total power at the sample was 120 mW (80 mW pump, 40 mW Stokes). Cells were imaged live using a Chamlide TC live cell chamber (Quorum Technologies) with CU-109 incubation and FC-5 automatic CO2/air mixer at 37 °C and 5 % CO2. ScanImage (Vidrio Technologies) software was used for image acquisition. Images were acquired using a 3 by 3 tile scan, with 50 % offset.66 This covered an area of approximately 0.1 mm2. Three different fields of view were imaged for each culture. Experiments were performed in triplicate. Image tiles were stitched together in post-processing using the MIST plugin within ImageJ. Prior to quantitative analysis, stitched images were pre-processed so only the area within the cell boundaries was analysed. A binary image was created using pixel classification within Ilastik. This binary image was then applied as a mask to the stitched images to outline the cell boundary as well as measure total cell area. The spot detection plugin within ICY was then utilised for quantitative analysis of CARS puncta within processed images.67 CARS images were analysed to count the number and total area of white puncta present.
Raman spectroscopy
A home-built Raman microspectrometer was used to acquire Raman spectra of crystalline PEO-PCL and empty polymersome samples. The system was principally composed of an inverted microscope (Nikon Ti Eclipse), 785 nm CW laser (Laser 2000), 40x air objective (0.8 NA) and spectrometer with CCD (Andor SR303i). Each time prior to spectral acquisition the system was calibrated for spectral positions using a polystyrene standard. A total of 10 spectra were acquired and averaged using 10 s exposure, 10 μm slit width, 300 lines per mm grating and 40 mW of laser power at the sample. Raw spectra were baseline corrected using asymmetric least squares fitting.
Imaging flow cytometry (IFC)
Imaging flow cytometry assays were completed as previously described with minor modifications.38 Briefly, RAW 264.7 macrophages were seeded in 24-well plates at 2.5 × 105 cells/cm2 (5 × 105/mL) and incubated overnight to ensure a density ~ 1 × 106 after 24 hours. B. thailandensis bacteria were added to RAW cells for 1 hour at a concentration of 1×108 bacteria/mL ([MOI] = 100), as previously described, before replacement of DMEM with L-15 medium containing kanamycin at 1 mg/mL for 3 hours. DiD-labelled PMs were then added to the infected cell culture at a dilution of 1:10. Cells were incubated at 37 °C for a further 3 or 21 hours before fixation in 0.5% (w/v) PFA for imaging flow cytometry analysis. Control groups included infected cells in the absence of PMs, and cells containing PMs but with no bacterial infection. Data was acquired using an ImageStream X MkII (ISX, Amnis, Seattle, USA). Cells were imaged at a 60x magnification, with resolution of 0.3 μm/pixel, and a 2.5 μm optical slice image of cells. 10,000 cellular events were acquired for each sample set. Only data from the relevant channels was collected and these included Channel 01 (brightfield camera), Channel 02 (488 nm laser power: 100 mW, for B. thailandensis eGFP visualisation), and Channel 11 (642 nm laser power: 150 mW, to allow DiD visualisation). Single stain controls were used to calculate and apply a compensation matrix to all collected data presented, and all gates were kept consistant throughout all experiments and biological replicates. Imaging flow cytometry data was analysed using IDEAS® version 6.2 software (Luminex, USA). Only imaging events that were in focus (based on Gradient_M01_Ch01 > 50) and comprised a single cell (Gated using Area_M01 vs. Aspect Ratio_M01) were analysed. The functions Intensity_MC_Ch02 and Max Pixel_MC_Ch02 were used to gate cells positive for GFP-labelled bacteria, and the function Intensity_MC_Ch11 was used to gate cells positive for DiD (defined at 1.6 × 104), as indicated in the results. Erode (M01.4) masks of bright field images were used to define intracellular regions of captured cells. To distinguish extra- and intracellular GFP-labelled bacteria, an internalisation feature was used to calculate the ratio of GFP fluorescence intensity inside the cell (defined using the Erode (M01.4) mask) compared to outside the cell. Cells with an internalisation score of 1 or more were deemed to have intracellular B. thailandensis, whereas those with a score of less than 1 were deemed to have extracellular B. thailandensis. The Similarity Bright Detail Feature R3 feature was used to determine co-localisation of GFP-positive bacteria and DiD-labelled PMs, with a BDS value of 0.95 or greater indicating co-localisation. Full gating strategies are shown in the results.
Minimum inhibitory concentration (MIC) assays
B. thailandensis bacteria from overnight cultures were diluted in L-broth to 1 × 105 bacteria/mL (based on optical density (OD)), using a Genesys 10 UV scanning spectrophotometer (Thermo Fisher, UK)) and incubated in 96 well plates for 24 hours at 37 °C in the presence of doxycycline and rifampicin at concentrations of 0 – 100 μg/mL, or antibiotic-loaded PMs at seven 1:2 serial dilutions, as stated in the results. A Multiskan™ FC OD plate reader (Thermo Fisher Scientific, UK) was used to assess bacterial growth. For each treatment there were with three technical repeats.
Intracellular infection assays
RAW 264.7 macrophages were infected with B. thailandensis, as for imaging experiments. To test the efficacy of free and PM-encapsulated antibiotics, free doxycycline and rifampicin were included at concentrations in the range 0 – 4.5 μg/mL or 0 – 2.6 μL/mL respectively, as stated in the results. PM-doxycycline or PM-rifampicin preparations were added to infected RAW 264.7 cell cultures at dilutions of 1:3.33 or 1:20. PMs were incubated for 3 or 21 hours at 37 °C. Cells were then lysed using distilled water and the lysate plated onto agar plates for quantification of bacterial colony forming units. Negative controls included wells with PM-empty nanoparticles, and wells containing no antibiotics or PMs. Agar plates were incubated at 37 °C for 48 hours before visual inspection and manual colony counting.
Statistics
Statistical analysis was performed using GraphPad Prism 8 software (GraphPad, USA). Statistical tests performed were combinations of unpaired t-tests, and one- or two-way ANOVAs with Tukey’s post-hoc correction test, as stated in results in figure legends.
Supplementary Material
Acknowledgments
The authors acknowledge the kind financial support of the United Kingdom Ministry of Defence. NDE acknowledges studentship support from the Institute for Life Sciences University of Southampton, and the EPSRC (EP/K029150/1 and EP/M508147/1) for funding. SM acknowledges EP/N509747/1 and UKMoD for funding the studentship of JSPH. SM acknowledges funding from EP/T020997/1.
We acknowledge helpful advice from Prof Peter Roach, University of Southampton.
Footnotes
Associated Content
The authors confirm that they have no competing financial interests.
References
- (1).Limmathurotsakul D, Peacock SJ. Melioidosis: A Clinical Overview. British Medical Bulletin. 2011;99(1):125–139. doi: 10.1093/bmb/ldr007. [DOI] [PubMed] [Google Scholar]
- (2).Limmathurotsakul D, Golding N, Dance DA, Messina JP, Pigott DM, Moyes CL, Rolim DB, Bertherat E, Day NP, Peacock SJ, Hay SI. Predicted Global Distribution of Burkholderia pseudomallei and Burden of Melioidosis. Nat Microbiol. 2016;1(1) doi: 10.1038/nmicrobiol.2015.8. [DOI] [PubMed] [Google Scholar]
- (3).Titball RW, Burtnick MN, Bancroft GJ, Brett P. Burkholderia pseudomallei and Burkholderia mallei Vaccines: Are We Close to Clinical Trials? Vaccine. 2017;35(44):5981–5989. doi: 10.1016/j.vaccine.2017.03.022. [DOI] [PubMed] [Google Scholar]
- (4).Gilad J, Harary I, Dushnitsky T, Schwartz D, Amsalem Y. Burkholderia mallei and Burkholderia pseudomallei as Bioterrorism Agents: National Aspects of Emergency Preparedness. Israel Medical Association Journal. 2007;9(7):499–503. [PubMed] [Google Scholar]
- (5).Willcocks SJ, Denman CC, Atkins HS, Wren BW. Intracellular Replication of the Well-Armed Pathogen Burkholderia pseudomallei . Current Opinion in Microbiology. 2016;29:94–103. doi: 10.1016/i.mib.2015.11.007. [DOI] [PubMed] [Google Scholar]
- (6).Ozanic M, Marecic V, Abu Kwaik Y, Santic M. The Divergent Intracellular Lifestyle of Francisella tularensis in Evolutionarily Distinct Host Cells. PLoS Pathogens. 2015;11(12):1–8. doi: 10.1371/journal.ppat.1005208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (7).Cheng A, Currie B. Melioidosis: Epidemiology, Pathophysiology, and Management. Clinical microbiology reviews. 2005;18(2):383–416. doi: 10.1128/CMR.18.2.383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (8).Kespichayawattana W, Rattanachetkul S, Wanun T, Utaisincharoen P, Sirisinha S. Burkholderia pseudomallei Induces Cell Fusion and Actin-Associated Membrane Protrusion: A Possible Mechanism for Cell-to-Cell Spreading. Infect Immun. 2000;68(9):5377–5384. doi: 10.1128/iai.68.9.5377-5384.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (9).Ernst RK, Guina T, Miller SI. How Intracellular Bacteria Survive: Surface Modifications That Promote Resistance to Host Innate Immune Responses. The Journal of infectious diseases. 1999;179(Suppl):S326–S330. doi: 10.1086/513850. [DOI] [PubMed] [Google Scholar]
- (10).Leon-Sicairos N, Reyes-Cortes R, Guadrón-Llanos AM, Madueña-Molina J, Leon-Sicairos C, Canizalez-Román A. Strategies of Intracellular Pathogens for Obtaining Iron from the Environment. BioMed Research International. 2015;2015 doi: 10.1155/2015/476534. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (11).McOrist S. Obligate Intracellular Bacteria and Antibiotic Resistance. Trends in Microbiology. 2000;8(11):483–486. doi: 10.1016/S0966-842X(00)01854-0. [DOI] [PubMed] [Google Scholar]
- (12).Larsen JC, Johnson NH. Pathogenesis of Burkholderia pseudomallei and Burkholderia mallei . Military medicine. 2009;174(6):647–651. [PubMed] [Google Scholar]
- (13).Currie BJ, Fisher DA, Anstey NM, Jacups SP. Melioidosis: Acute and Chronic Disease, Relapse and Re-Activation. Trans R Soc Trop Med Hyg. 2000;94(3):301–304. doi: 10.1016/s0035-9203(00)90333-x. [DOI] [PubMed] [Google Scholar]
- (14).Dethlefsen L, Relman DA. Incomplete Recovery and Individualized Responses of the Human Distal Gut Microbiota to Repeated Antibiotic Perturbation. Proceedings of the National Academy of Sciences of the United States of America. 2011;(Suppl 1):4554–4561. doi: 10.1073/pnas.1000087107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (15).Zaman SBin, Hussain MA, Nye R, Mehta V, Mamun KT, Hossain N. A Review on Antibiotic Resistance: Alarm Bells Are Ringing. Cureus. 2017;9(6) doi: 10.7759/cureus.1403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (16).Discher BM, Won Y-Y, Ege DS, Lee JC-M, Bates FS, Discher DE, Hammer DA. Polymersomes: Tough Vesicles Made from Diblock Copolymers. Science. 1999;284(5417):1143–1146. doi: 10.1126/science.284.5417.1143. [DOI] [PubMed] [Google Scholar]
- (17).Scarpa E, Janeczek AA, Hailes A, de Andrés MC, De Grazia A, Oreffo RO, Newman TA, Evans ND. Polymersome Nanoparticles for Delivery of Wnt-Activating Small Molecules. Nanomedicine: Nanotechnology, Biology, and Medicine. 2018;14(4):1267–1277. doi: 10.1016/j.nano.2018.02.014. [DOI] [PubMed] [Google Scholar]
- (18).Qi W, Ghoroghchian PP, Li G, Hammer DA, Therien MJ. Aqueous Self-Assembly of Poly(ethylene Oxide)-Block-Poly(ε-Caprolactone) (PEO-b-PCL) Copolymers: Disparate Diblock Copolymer Compositions Give Rise to Nano- and Meso-Scale Bilayered Vesicles. Nanoscale. 2013;5(22):10908–10915. doi: 10.1039/c3nr03250g. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (19).Allen TM, Cullis PR. Liposomal Drug Delivery Systems : From Concept to Clinical Applications ☆. Advanced Drug Delivery Reviews. 2013;65(1):36–48. doi: 10.1016/j.addr.2012.09.037. [DOI] [PubMed] [Google Scholar]
- (20).Youssef M, Fattal E, Alonso MJ, Roblot-Treupel L, Sauzières J, Tancrède C, Omnès A, Couvreur P, Andremont A. Effectiveness of Nanoparticle-Bound Ampicillin in the Treatment of Listeria monocytogenes Infection in Athymic Nude Mice. Antimicrob Agents Chemother. 1988;32(8):1204–1207. doi: 10.1128/aac.32.8.1204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (21).Balland O, Pinto-Alphandary H, Pecquet S, Andremont A, Couvreur P. The Uptake of Ampicillin-Loaded Nanoparticles by Murine Macrophages Infected with Salmonella typhimurium . Journal of Antimicrobial Chemotherapy. 1994;33(3):509–522. doi: 10.1093/jac/33.3.509. [DOI] [PubMed] [Google Scholar]
- (22).Gandhi A, Paul A, Sen SO, Sen KK. Studies on Thermoresponsive Polymers : Phase Behaviour, Drug Delivery and Biomedical Applications. Asian Journal of Pharmaceutical Sciences. 2015;10(2):99–107. doi: 10.1016/j.ajps.2014.08.010. [DOI] [Google Scholar]
- (23).Lee JS, Zhou W, Meng F, Zhang D, Otto C, Feijen J. Thermosensitive Hydrogel-Containing Polymersomes for Controlled Drug Delivery. Journal of Controlled Release. 2010;146(3):400–408. doi: 10.1016/j.jconrel.2010.06.002. [DOI] [PubMed] [Google Scholar]
- (24).Du J, Tang Y, Lewis AL, Armes SP. PH-Sensitive Vesicles Based on a Biocompatible Zwitterionic Diblock Copolymer. 2005:17982–17983. doi: 10.1021/ja056514l. [DOI] [PubMed] [Google Scholar]
- (25).Lomas H, Canton I, MacNeil S, Du J, Armes SP, Ryan AJ, Lewis AL, Battaglia G. Biomimetic PH Sensitive Polymersomes for Efficient DNA Encapsulation and Delivery. Advanced Materials. 2007;19(23):4238–4243. doi: 10.1002/adma.200700941. [DOI] [Google Scholar]
- (26).Cabane E, Malinova V, Menon S, Palivan CG, Meier W. Photoresponsive Polymersomes as Smart, Triggerable Nanocarriers. Soft Matter. 2011;7(19):9167–9176. doi: 10.1039/c1sm05880k. [DOI] [Google Scholar]
- (27).Cerritelli S, Velluto D, Hubbell JA. PEG-SS-PPS: Reduction-Sensitive Disulfide Block Copolymer Vesicles for Intracellular Drug Delivery. Biomacromolecules. 2007;8(6):1966–1972. doi: 10.1021/bm070085x. [DOI] [PubMed] [Google Scholar]
- (28).Egli S, Nussbaumer MG, Balasubramanian V, Chami M, Bruns N, Palivan C, Meier W. Biocompatible Functionalization of Polymersome Surfaces : A New Approach to Surface Immobilization and Cell Targeting Using Polymersomes. 2011:4476–4483. doi: 10.1021/ja110275f. [DOI] [PubMed] [Google Scholar]
- (29).Meng F, Cheng R, Deng C, Zhong Z. Intracellular Drug Release Nanosystems in Order to Elicit Therapeutic Effects, Many Drugs Including Small Molecule. Materials Today. 2012;15(10):436–442. doi: 10.1016/S1369-7021(12)70195-5. [DOI] [Google Scholar]
- (30).Sanson C, Schatz C, Meins JLe, Soum A, Thévenot J, Garanger E, Lecommandoux S. A Simple Method to Achieve High Doxorubicin Loading in Biodegradable Polymersomes. Journal of Controlled Release. 2010;147(3):428–435. doi: 10.1016/j.jconrel.2010.07.123. [DOI] [PubMed] [Google Scholar]
- (31).Anselmo AC, Mitragotri S. Nanoparticles in the Clinic: An Update. Bioengineering & Translational Medicine. 2019;4(3):1–16. doi: 10.1002/btm2.10143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (32).Wayakanon K, Thornhill MH, Douglas CWI, Lewis AL, Warren NJ, Pinnock A, Armes SP, Battaglia G, Murdoch C, Porphyromo T. Polymersome-Mediated Intracellular Delivery of Antibiotics to Treat Porphyromonas gingivalis-Infected Oral Epithelial Cells. 2013;27(11):4455–4465. doi: 10.1096/fj.12-225219. [DOI] [PubMed] [Google Scholar]
- (33).Moretton MA, Cagel M, Bernabeu E, Gonzalez L, Chiappetta DA. Nanopolymersomes as Potential Carriers for Rifampicin Pulmonary Delivery. Colloids and Surfaces B: Biointerfaces. 2015;136:1017–1025. doi: 10.1016/j.colsurfb.2015.10.049. [DOI] [PubMed] [Google Scholar]
- (34).Trousil J, Syrová Z, Dal N-JK, Rak D, Konefał R, Pavlova E, Matĕjková J, Cmarko D, Kubíčková P, Pavliš O, Urbánek T, et al. Rifampicin Nanoformulation Enhances Treatment of Tuberculosis in Zebrafish. Biomacromolecules. 2019;20(4):1798–1815. doi: 10.1021/acs.biomac.9b00214. [DOI] [PubMed] [Google Scholar]
- (35).Trousil J, Pavliš O, Kubíčková P, Škorič M, Marešová V, Pavlova E, Knudsen KD, Dai Y-S, Zimmerman M, Dartois V, Fang J-Y, et al. Antitubercular Nanocarrier Monotherapy: Study of in Vivo Efficacy and Pharmacokinetics for Rifampicin. J Control Release. 2020;321:312–323. doi: 10.1016/j.jconrel.2020.02.026. [DOI] [PubMed] [Google Scholar]
- (36).Fenaroli F, Robertson JD, Scarpa E, Gouveia VM, Di Guglielmo C, De Pace C, Elks PM, Poma A, Evangelopoulos D, Canseco JO, Prajsnar TK, et al. Polymersomes Eradicating Intracellular Bacteria. ACS Nano. 2020;14(7):8287–8298. doi: 10.1021/acsnano.0c01870. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (37).Lane DD, Su FY, Chiu DY, Srinivasan S, Wilson JT, Ratner DM, Stayton PS, Convertine AJ. Dynamic Intracellular Delivery of Antibiotics in Vitro PH-Responsive Polymersomes. Polym Chem. 2015;6(8):1255–1266. doi: 10.1039/C4PY01249F. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (38).Jenner D, Ducker C, Clark G, Prior J, Rowland CA. Using Multispectral Imaging Flow Cytometry to Assess an in Vitro Intracellular Burkholderia thailandensis Infection Model. Cytometry A. 2016;89(4):328–337. doi: 10.1002/cyto.a.22809. [DOI] [PubMed] [Google Scholar]
- (39).Moura CC, Lanham SA, Monfort T, Bourdakos KN, Tare RS, Oreffo ROC, Mahajan S. Quantitative Temporal Interrogation in 3D of Bioengineered Human Cartilage Using Multimodal Label-Free Imaging. Integr Biol. 2018;10(10):635–645. doi: 10.1039/C8IB00050F. [DOI] [PubMed] [Google Scholar]
- (40).Scarpa E, Bailey JL, Janeczek AA, Stumpf PS, Johnston AH, Oreffo ROC, Woo YL, Cheong YC, Evans ND, Newman TA. Quantification of Intracellular Payload Release from Polymersome Nanoparticles. Sci Rep. 2016;6:29460. doi: 10.1038/srep29460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (41).White NJ. Melioidosis. Lancet. 2003;361(9370):1715–1722. doi: 10.1016/s0140-6736(03)13374-0. [DOI] [PubMed] [Google Scholar]
- (42).Ross BN, Myers JN, Muruato LA, Tapia D, Torres AG. Evaluating New Compounds to Treat Burkholderia pseudomallei Infections. Front Cell Infect Microbiol. 2018;8:210. doi: 10.3389/fcimb.2018.00210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (43).Vauthier C, Cabane B, Labarre D. How to Concentrate Nanoparticles and Avoid Aggregation? European Journal of Pharmaceutics and Biopharmaceutics. 2008;69(2):466–475. doi: 10.1016/j.ejpb.2008.01.025. [DOI] [PubMed] [Google Scholar]
- (44).Fenaroli F, Robertson JD, Scarpa E, Gouveia VM, Di Guglielmo C, De Pace C, Elks PM, Poma A, Evangelopoulos D, Canseco JO, Prajsnar TK, et al. Polymersomes Eradicating Intracellular Bacteria. ACS Nano. 2020 doi: 10.1021/acsnano.0c01870. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (45).Brož P, Benito SM, Saw C, Burger P, Heider H, Pfisterer M, Marsch S, Meier W, Hunziker P. Cell Targeting by a Generic Receptor-Targeted Polymer Nanocontainer Platform. Journal of Controlled Release. 2005;102(2):475–488. doi: 10.1016/j.jconrel.2004.10.014. [DOI] [PubMed] [Google Scholar]
- (46).Garrett NL, Lalatsa A, Begley D, Mihoreanu L, Uchegbu IF, Schätzlein AG, Moger J. Label-Free Imaging of Polymeric Nanomedicines Using Coherent Anti-Stokes Raman Scattering Microscopy. Journal of Raman Spectroscopy. 2012;43(5):681–688. doi: 10.1002/jrs.3170. [DOI] [Google Scholar]
- (47).Tong L, Lu Y, Lee RJ, Cheng J-X. Imaging Receptor-Mediated Endocytosis with a Polymeric Nanoparticle-Based Coherent Anti-Stokes Raman Scattering Probe. J Phys Chem B. 2007;111(33):9980–9985. doi: 10.1021/jp073478z. [DOI] [PubMed] [Google Scholar]
- (48).McKenzie Z, Kendall M, Mackay R-M, Whitwell H, Elgy C, Ding P, Mahajan S, Morgan C, Griffiths M, Clark H, Madsen J. Surfactant Protein A (SP-A) Inhibits Agglomeration and Macrophage Uptake of Toxic Amine Modified Nanoparticles. Nanotoxicology. 2015;9(8):952–962. doi: 10.3109/17435390.2014.992487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (49).Xu P, Gullotti E, Tong L, Highley CB, Errabelli DR, Hasan T, Cheng J-X, Kohane DS, Yeo Y. Intracellular Drug Delivery by Poly(Lactic-Co-Glycolic Acid) Nanoparticles, Revisited. Mol Pharm. 2009;6(1):190–201. doi: 10.1021/mp800137z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (50).Belsey NA, Garrett NL, Contreras-Rojas LR, Pickup-Gerlaugh AJ, Price GJ, Moger J, Guy RH. Evaluation of Drug Delivery to Intact and Porated Skin by Coherent Raman Scattering and Fluorescence Microscopies. J Control Release. 2014;174:37–42. doi: 10.1016/j.jconrel.2013.11.002. [DOI] [PubMed] [Google Scholar]
- (51).Steuwe C, Patel II, Ul-Hasan M, Schreiner A, Boren J, Brindle KM, Reichelt S, Mahajan S. CARS Based Label-Free Assay for Assessment of Drugs by Monitoring Lipid Droplets in Tumour Cells. J Biophotonics. 2014;7(11-12):906–913. doi: 10.1002/jbio.201300110. [DOI] [PubMed] [Google Scholar]
- (52).Clemens DL, Lee B-Y, Xue M, Thomas CR, Meng H, Ferris D, Nel AE, Zink JI, Horwitz MA. Targeted Intracellular Delivery of Antituberculosis Drugs to Mycobacterium tuberculosis-Infected Macrophages in Vitro Functionalized Mesoporous Silica Nanoparticles. Antimicrob Agents Chemother. 2012;56(5):2535–2545. doi: 10.1128/AAC.06049-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (53).Kalluru R, Fenaroli F, Westmoreland D, Ulanova L, Maleki A, Roos N, Madsen MP, Koster G, Egge-Jacobsen W, Wilson S, Roberg-Larsen H, et al. Poly(Lactide-Co-Glycolide)-Rifampicin Nanoparticles Efficiently Clear Mycobacterium bovis BCG Infection in Macrophages and Remain Membrane-Bound in Phago-Lysosomes. Journal of Cell Science. 2013;126(14):3043–3054. doi: 10.1242/jcs.121814. [DOI] [PubMed] [Google Scholar]
- (54).Kisich KO, Gelperina S, Higgins MP, Wilson S, Shipulo E, Oganesyan E, Heifets L. Encapsulation of Moxifloxacin within Poly(butyl Cyanoacrylate) Nanoparticles Enhances Efficacy against Intracellular Mycobacterium tuberculosis . Int J Pharm. 2007;345(1-2):154–162. doi: 10.1016/j.ijpharm.2007.05.062. [DOI] [PubMed] [Google Scholar]
- (55).de Faria TJ, Roman M, de Souza NM, De Vecchi R, de Assis JV, dos Santos ALG, Bechtold IH, Winter N, Soares MJ, Silva LP, De Almeida MV, et al. An Isoniazid Analogue Promotes Mycobacterium tuberculosis-Nanoparticle Interactions and Enhances Bacterial Killing by Macrophages. Antimicrob Agents Chemother. 2012;56(5):2259–2267. doi: 10.1128/AAC.05993-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (56).Qie Y, Yuan H, Von Roemeling CA, Chen Y, Liu X, Shih KD, Knight JA, Tun HW, Wharen RE, Jiang W, Kim BYS. Surface Modification of Nanoparticles Enables Selective Evasion of Phagocytic Clearance by Distinct Macrophage Phenotypes. Scientific Reports. 2016 Jan;6:1–11. doi: 10.1038/srep26269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (57).Foroozandeh P, Aziz AA. Insight into Cellular Uptake and Intracellular Trafficking of Nanoparticles. Nanoscale Res Lett. 2018;13 doi: 10.1186/s11671-018-2728-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (58).Redka DS, Gütschow M, Grinstein S, Canton J. Differential Ability of Proinflammatory and Anti-Inflammatory Macrophages to Perform Macropinocytosis. Mol Biol Cell. 2018;29(1):53–65. doi: 10.1091/mbc.E17-06-0419. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (59).Allwood EM, Devenish RJ, Prescott M, Adler B, Boyce JD. Strategies for Intracellular Survival of Burkholderia pseudomallei . Frontiers in Microbiology. 2011 Aug;2:1–19. doi: 10.3389/fmicb.2011.00170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (60).Pinto-Alphandary H, Balland O, Laurent M, Andremont A, Puisieux F, Couvreur P. Intracellular Visualization of Ampicillin-Loaded Nanoparticles in Peritoneal Macrophages Infected in vitro with Salmonella typhimurium . Pharm Res. 1994;11(1):38–46. doi: 10.1023/a:1018985308984. [DOI] [PubMed] [Google Scholar]
- (61).Roobol SJ, Hartjes TA, Slotman JA, de Kruijff RM, Torrelo G, Abraham TE, Bruchertseifer F, Morgenstern A, Kanaar R, van Gent DC, Houtsmuller AB, et al. Uptake and Subcellular Distribution of Radiolabeled Polymersomes for Radiotherapy. Nanotheranostics. 2020;4(1):14–25. doi: 10.7150/ntno.37080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (62).Woodward SC, Brewer PS, Moatamed F, Schindler A, Pitt CG. The Intracellular Degradation of Poly(E-Caprolactone) Journal of Biomedical Materials Research. 1985;19(4):437–444. doi: 10.1002/jbm.820190408. [DOI] [PubMed] [Google Scholar]
- (63).Arakawa CK, DeForest CA. Biology and Engineering of Stem Cell Niches. Elsevier Inc; London: 2017. Polymer Design and Development; pp. 295–314. [DOI] [Google Scholar]
- (64).Scarpa E, Bailey JL, Janeczek AA, Stumpf PS, Johnston AH, Oreffo ROC, Woo YL, Cheong YC, Evans ND, Newman TA. Quantification of Intracellular Payload Release from Polymersome Nanoparticles. Scientific Reports. 2016 Jul;6:1–13. doi: 10.1038/srep29460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (65).Peng H, Ling J, Liu J, Zhu N, Ni X, Shen Z. Controlled Enzymatic Degradation of Poly(ε-Caprolactone)-Based Copolymers in the Presence of Porcine Pancreatic Lipase. Polymer Degradation and Stability. 2010;95(4):643–650. doi: 10.1016/j.polymdegradstab.2009.12.005. [DOI] [Google Scholar]
- (66).Pologruto TA, Sabatini BL, Svoboda K. ScanImage: Flexible Software for Operating Laser Scanning Microscopes. BioMedical Engineering OnLine. 2003;2(1):13. doi: 10.1186/1475-925X-2-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (67).de Chaumont F, Dallongeville S, Chenouard N, Hervé N, Pop S, Provoost T, Meas-Yedid V, Pankajakshan P, Lecomte T, Le Montagner Y, Lagache T, et al. Icy: An Open Bioimage Informatics Platform for Extended Reproducible Research. Nature Methods. 2012;9(7):690–696. doi: 10.1038/nmeth.2075. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.