Abstract
The persistence of cancer cells resistant to therapy remains a major clinical challenge. In triple-negative breast cancer, resistance to chemotherapy results in the highest recurrence risk among breast cancer subtypes. The drug-tolerant state seems largely defined by non-genetic features, but the underlying mechanisms are poorly understood. Here, by monitoring epigenomes, transcriptomes and lineages with single-cell resolution, we show that the repressive histone mark H3K27me3 regulates cell fate at the onset of chemotherapy. We report that a persister expression program is primed with both H3K4me3 and H3K27me3 in unchallenged cells, H3K27me3 being the lock to its transcriptional activation. We further demonstrate that depleting H3K27me3 enhances the potential of cancer cells to tolerate chemotherapy. Conversely, preventing H3K27me3 demethylation simultaneously to chemotherapy inhibits the transition to a drug-tolerant state, and delays tumor recurrence in vivo. Our results highlight how chromatin landscapes shape the potential of cancer cells to respond to initial therapy.
Introduction
Emergence of resistant phenotypes from initially responding or partially responding tumors has been modeled as a multi-step process1–4. Initially, after drug insult, only a pool of persister cells – also called drug-tolerant persister cells (DTPs) – manage to tolerate the cancer treatment and survive. These cells constitute a reservoir of cells from which drug-resistant cells, actively growing under cancer treatment, can ultimately emerge3,5,6. In triple-negative breast cancer (TNBC), both genetic and transcriptomic mechanisms have been proposed to drive cancer evolution towards chemoresistance, through combined selective and adaptive modes of evolution7. The recent identification of a multi-clonal reversible drug-tolerant state post neo-adjuvant chemotherapy in patient-derived xenografts (PDX)8 suggested that the earliest steps of chemoresistance in TNBC are not driven by genetic alterations, but rather by non-genetic plasticity in multiple cancer cells. Similarly, in other cancer types, drug-tolerant states have been identified solely characterized by changes in transcriptomic and epigenomic features in response to targeted therapies or chemotherapies9–11.
The genetic history of many cancer types has been extensively modeled thanks to both bulk and single-cell approaches7,12. In contrast, little is known about the epigenomic heterogeneity and dynamics of acquisition of epigenetic alterations. While recent studies have focused on the evolution of DNA methylation13,14 – among the most stable epigenetic locks to gene expression – contribution to tumor evolution of more versatile epigenetic modifications has remained poorly understood. Single-cell methods to map repressive and permissive histone modifications, key players of cellular plasticity, have emerged only recently15,16, enabling the study of epigenomic diversity within biological systems. Studying cases of acquired resistance in luminal and triple-negative cancer in PDX with single-cell chromatin profiling, we have identified examples of epigenomic clones within tumors, solely defined by H3K27me3 landscapes15. We detected resistant-like H3K27me3 landscapes in initial untreated tumors, proposing histone modifications as a molecular player of tumor evolution processes. Here, we focused on the earliest steps of cancer treatment prior to the emergence of resistant populations, to identify the initial epigenomic mechanisms driving drug tolerance. Thanks to a characterization of epigenomic and transcriptomic evolution under chemotherapy at single-cell resolution, we show that H3K27me3 landscapes, and not H3K4me3, are a proxy of cancer cell evolution upon initial chemotherapy insult. We discovered that the persister expression program is in a bivalent chromatin configuration in untreated cells, and locked by H3K27me3. Tracing cell lineages together with transcriptomics, we further reveal that H3K27me3 is a key determinant of cell fate upon initial chemotherapy exposure in TNBC: depriving cells of H3K27me3 enhances the potential of each tumor cell to tolerate chemotherapy. Finally, preventing cells under chemotherapy insult to demethylate H3K27me3 inhibits their potential to tolerate chemotherapy, and delays tumor recurrence in vivo.
Results
Deriving persister cells from patient tumor models
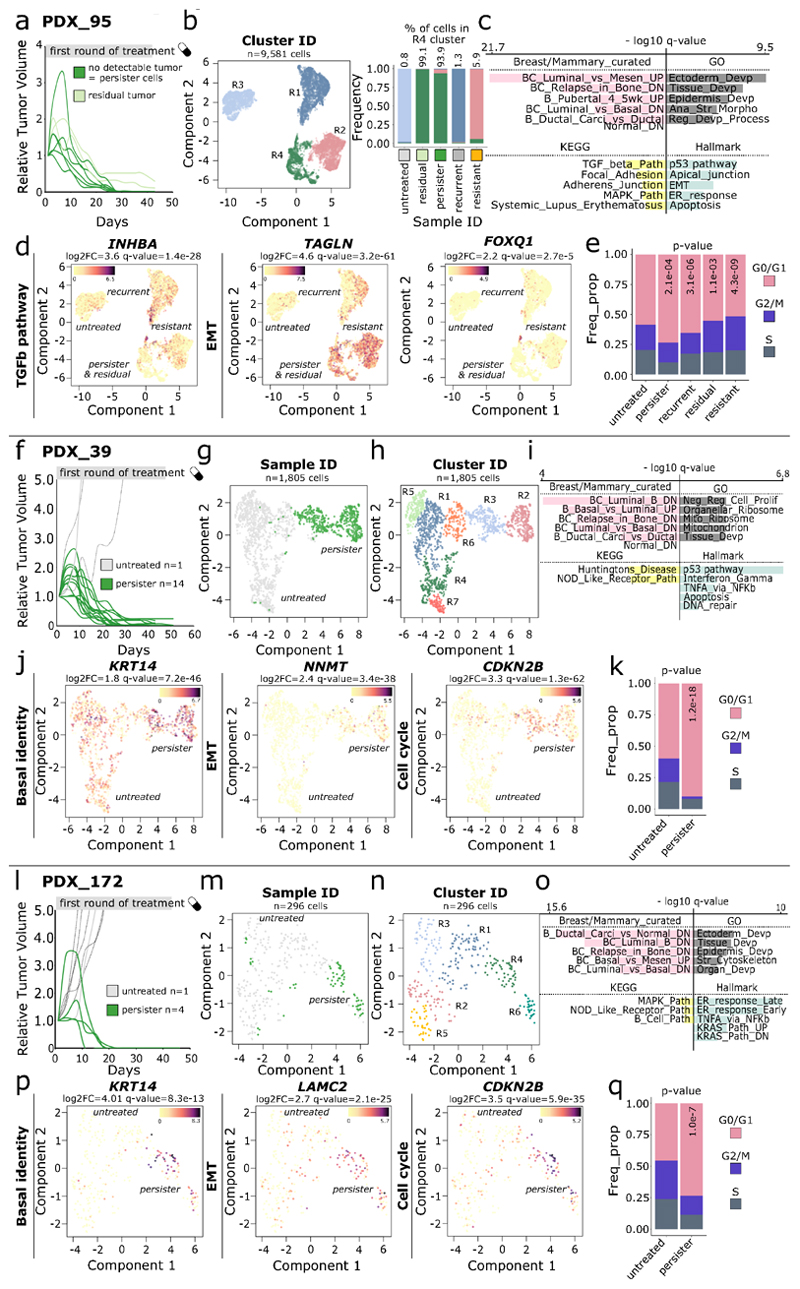
We focused on mechanisms of drug tolerance in residual TNBC, for which patients have the poorest outcome. Resistance to adjuvant chemotherapy cannot be easily studied as biopsies are not routinely performed when the disease progresses (Fig. 1a). To circumvent these limitations, we modeled, in vivo and in vitro, phenotypes of drug-response observed in patients. In vivo, we first treated three PDX models – PDX_95, PDX_39 & PDX_172 – established from patients with residual TNBC15,17, with Capecitabine, the standard of care for residual breast tumors (Fig. 1b, Extended Data Fig. 1). After the first round of chemotherapy treatment, mice displayed a pathological complete response (pCR), but tumors eventually recurred (‘recurrent’) and mice were treated again with chemotherapy, to which tumors responded to various extents, some maintaining constant tumor volume under treatment (‘resistant’) (Fig. 1b). These recurrent tumors potentially arose from persister cells, surviving initial chemotherapy treatment5. We isolated persister cells by pooling the fat pad from mice with pCR (from 4 to 14, Extended Data Fig. 1a, 1f & 1l). To phenocopy a clinical situation of partial response, we also generated ‘residual’ tumors for one model (PDX_95, n = 2) (Extended Data Fig. 1a) by administering half the dose of Capecitabine.
Fig. 1. Identification of a pool of basal persister cells in TNBC in vivo and in vitro.
a. Schematic representation of the standard of care for TNBC patients and the generation of patient-derived xenograft models. b. Graph of the relative tumor volumes (RTV) over time (days). Colored growth curves correspond to tumors that have been further studied by scRNA-seq. Black arrows indicate the start of the second round of Capecitabine treatment for the corresponding mice. c. (Up) Phenotypes and cell numbers are indicated, with the number of mice used to collect samples in brackets. (Down) UMAP representation of PDX scRNA-seq datasets, colored according to sample of origin (first panel - cluster ID are indicated) or log2 gene expression signal for differentially expressed genes between persister cells and untreated tumor cells (remaining panels), log2FC and adjusted P values are indicated above the graph. d. (Left) Venn diagram displaying the intersection of pathways activated in persister cells from the 3 PDX models, among MSigDB c2_curated Breast/Mammary and c7_Hallmark pathways. P value associated with the intersection is indicated below (exact test of multi-set intersections) (Right) Barplot displaying the top 5 pathways - for each category - activated in persister cells. x-axis corresponds to -log10 adjusted P values for the model PDX_95. e. Graph representation of the cell proliferation of triple negative breast cancer cell line MDA-MB-468 (MM468) treated with 5-FU (green for persister cells, and orange lines for resistant cells) or with DMSO (untreated - grey lines). f. (Up) Schematic view of the experimental design. Experiment number and corresponding passage of cells at D0 are indicated. (Down) UMAP representation of MDA-MB-468 cells scRNA-seq datasets, colored according to the sample of origin (first panel - cluster ID are indicated) or log2 gene expression signal for differentially expressed genes between persister cells (cluster R2) and untreated cells (cluster R10, KRT14 and TGFB1 panels) or for a differentially expressed gene between the two persisters clusters, i.e. clusters R4 vs R2 (CDH2 panel). Untreated population (in grey) corresponds to DMSO-D0-#1. g. (Left) UMAP representation of scRNA-seq as in 1f, restricted to cells with detected lineage barcode. Cells are colored according to lineage barcode and cluster membership is indicated (Extended Data Fig. 2c). R1, R2 correspond to RNA-inferred clusters. (Right) Scatter plot of the lineage barcode diversity detected in the scRNA-seq data across clusters and samples. Colors correspond to sample ID as in 1f. (Means are indicated for persister clusters R2 & R4, two-sided Wilcoxon rank test).
In vitro, we treated an initially chemosensitive TNBC cell line (MDA-MB-468), with Fluorouracil (5-FU)18, the active metabolite of Capecitabine, which is not converted in vitro. We drove independently three pools of cells to chemotolerance and subsequently to chemoresistance with prolonged 5-FU treatment (>15 weeks, Fig. 1e). After 3 weeks, only few cells survived drug insult (0.01% of the initial population, Extended Data Fig. 2a), out of which a minority (36%) divided after 10-15 additional days (Extended Data Fig. 2a); this group of cells, surviving initial chemotherapy, are referred to as ‘persister’ cells. Over 15 weeks, populations of resistant cells emerged, with doubling times comparable to untreated cells and an IC50 to 5-FU over 4-fold higher than untreated population (Extended Data Fig. 2b).
Identification of a pool of basal persister cells
To characterize transcriptomic evolution of untreated cells towards chemotolerance and subsequently chemoresistance, we performed single-cell RNA-seq (scRNA-seq) in both cell lines and PDX models (Fig. 1c, 1f, Extended Data Figs. 1 & 2). In vivo, scRNA-seq was key to identify the rare human persister cells among the vast majority of stromal mouse cells. Out of the fat pad, we typically isolated hundreds of persister cells per mouse. In vivo, persister cells from different mice grouped within one or two expression clusters (Extended Data Fig. 1b, 1g-h & 1m-n). In vitro, diverse cell populations (res #1, 2 and 3) with distinct expression programs, originated from the pool of persister cells, all grouped within common clusters across experiments (scRNA-based clusters R2/R4, Extended Data Fig. 2c). In vivo and in vitro, persister cells recurrently activated a set of common pathways compared to untreated cells (Fig. 1c-d-f, Extended Data Figs. 1c-d, 1i-j, 1o-p, 2d-e). Originating from KRT5-expressing cancer cells, persister cells recurrently activated sets of genes further establishing basal cell identity, such as KRT14 (Fig. 1c, 1f, Extended Data Fig. 1j & 1p). Compared to untreated cells, persister cells in vivo and in vitro also showed an activation of genes associated with the epithelial-to-mesenchymal transition (EMT) (Fig. 1c-f, Extended Data Figs. 1c-d, 1j & 1p, 2d & 2f) – such as TAGLN, which encodes an actin-binding protein, previously shown to promote metastasis through EMT19, and NNMT, characteristic of the metabolic changes that accompany EMT20–22. Persister cells also activated genes involved in the TNF-alpha/NF-κB pathway. Part of the persister expression program remained patient-specific: in PDX_95 and in vitro persister cells showed for example an activation of the TGF-β pathway with the expression of multiple players including ligands and receptors (Fig.1c & 1f, Supplementary Tables 1-4).
In vivo, we showed that persister and residual tumor cells actually clustered together (Fig. 1c, cluster R4), thereby sharing common expression features, suggesting similar mechanisms of chemotolerance independent of the residual burden. Yet we detected a higher number of cells in G0/G1 within persister populations than in residual or untreated tumors (Extended Data Fig. 1e-k-q), a characteristic recapitulated in vitro (Extended Data Fig. 2g) and in line with previous reports1,9,10,23. In vitro, we identified two clusters of persister cells (clusters R2 and R4), that differ by their expression of additional EMT markers such as CDH2 (Fig. 1f) and TWIST1. Early individual non-cycling persisters (day 33) solely belonged to cluster R2/CDH2- whereas dividing persisters could either belong to R2/CDH2- or R4/CDH2+ (Fig. 1f). Overall, we identified both in vivo and in vitro a reservoir of persister basal cells with EMT markers and activated NF-κB pathway. NF-κB signaling pathway and EMT-associated genes were proposed as potential drivers of chemoresistance in various tumor types, including lung24,25, pancreatic26, breast27,28. Here we pinpoint NF-κB and EMT pathway activation as the earliest common molecular events at the onset of chemotolerance in TNBC, defining a common Achilles’ heel to target chemotolerant cells before they phenotypically diversify.
Cell fate bias under chemotherapy
To follow clonal evolution under therapeutic stress, we had initially introduced unique genetic barcodes in untreated MDA-MB-468 cells prior to our experiments (Extended Data Fig. 3a). We leveraged our previous barcoding method29 to allow robust detection of barcodes within scRNA-seq data (Fig. 1g), as shown by the number of cells with a detected lineage barcode (Extended Data Fig. 3b). In addition, we verified that barcodes frequencies detected in scRNA-seq data recapitulated those detected in bulk, confirming the sensitivity of barcode detection in scRNA-seq data (Extended Data Fig. 3c). By combining detection of lineage barcode and expression programs at single-cell resolution, we were able to monitor clonal evolution over the course of the treatment and within each expression clusters (Fig. 1g). If non-cycling persisters (day 33) were multi-clonal (62% of unique barcodes), R6 and R8 chemoresistant clusters were constituted of few clones (3 and 4 respectively). According to bulk data (Extended Data Fig. 3d-e), barcode diversity in the cell population decreased with time both under 5-FU and DMSO exposure. Across experiments and time points, the fraction of unique barcodes within the CDH2-/R2 persister cluster was significantly higher than in CDH2+/R4 cluster (average of 41% versus 8% unique barcode, P = 1.6 × 10-2, Fig. 1g), demonstrating that if the drug-tolerant state is multi-clonal, only rare persister cells switch to the CDH2+ state.
To test if the lineages that persist were selected within the untreated population, we next compared barcode frequencies between the starting population and the 5-FU- or DMSO-treated cells using additional bulk experiments (Experiment #3, Extended Data Fig. 3d-g). If surviving cells had no particular predisposition then they should resemble a random draw of the initial untreated population (day 0, Extended Data Fig. 3f). In contrast to DMSO-treated cells, barcode frequencies of the 5-FU-treated cells deviated from this random scenario (Extended Data Fig. 3g, r = 0.68 and r = 0.29 respectively), indicating that a fraction of lineages present in the untreated population have a predisposition to tolerate chemotherapy. Using our single-cell barcoding dataset, we were able to identify within the untreated populations, future persisters (n = 143, for which the lineage barcode is found both in persisters and untreated cells) and compare them to ‘non-persisting’ cells (n = 201, Extended Data Fig. 3h-j). The only transcriptomic differences between those two cell populations were the overexpression of S100A2, which encodes a calcium binding protein, and LDHB, which encodes a lactate dehydrogenase, in future persister cells prior to treatment. These results suggest that metabolic differences could be an indicator of potential for persistence.
Dynamics of genomic and epigenomic features
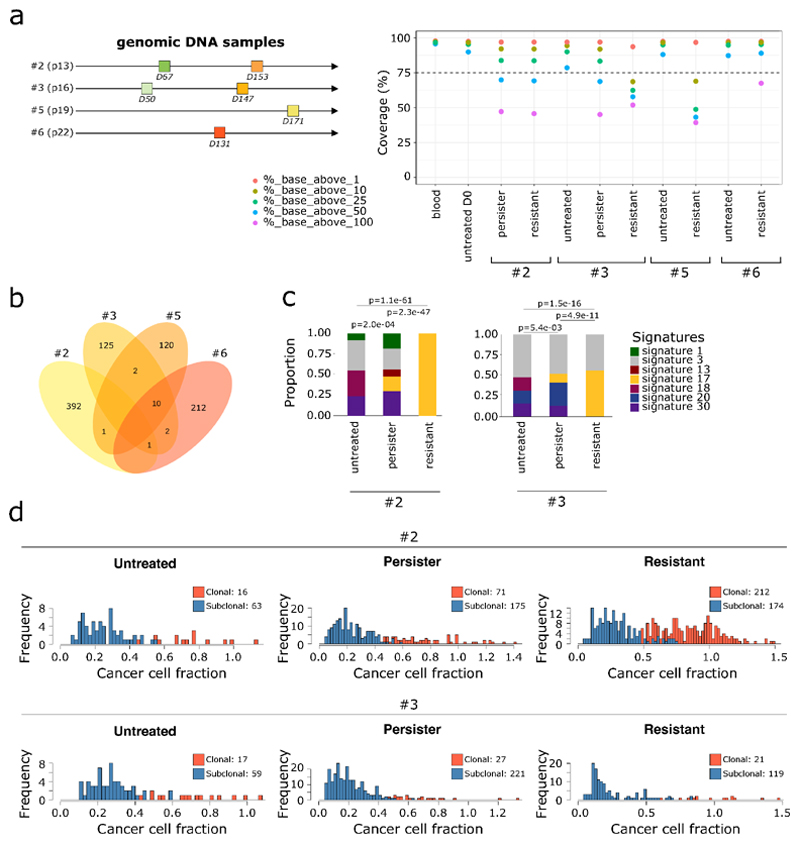
To hamper the chemotherapy-driven clonal evolution of cancer cells, we next investigated the molecular basis of such rapid phenotypic evolution. Using whole-exome sequencing (Extended Data Fig. 4a), we first analyzed mutations, copy-number alterations (CNA) and related mutational signatures acquired by persister and resistant cell populations since the onset of 5-FU treatment. We could not identify any recurrent mutations across experiments (Extended Data Fig. 4b), or any CNA (amplifications or homozygous deletions) or recurrent mutations affecting known driver genes of breast cancer in any population12. Only a minor fraction of mutations found in persister cells were attributed to 5-FU exposure (mutational signature 1730) in contrast to resistant cell populations where over 50% of acquired mutations are associated to 5-FU (P < 10-10, Extended Data Fig. 4c). These results indicated that chemotherapy-related mutations are acquired over a timeframe that is not compatible with the rapid phenotypic evolution seen in persister cells. Finally, computing cancer cell fractions for each mutation, we confirmed that persister populations are extensively multi-clonal (Extended Data Fig. 4d), in line with the lineage barcoding results.
We next investigated changes in epigenomes during chemotherapy treatment. Using single-cell profiling (scChIP-seq), we observed that H3K27me3 dynamics captured the evolution of cell states with chemotherapy (Fig. 2a, Extended Data Fig. 5a and Supplementary Table 4). Persister cells shared a common H3K27me3 pattern (cluster E1, Fig. 2b, Extended Data Fig. 5b), in contrast to resistant cells split in clusters E1 and E3. In comparison to untreated cells, cells from cluster E1 showed recurrent redistribution of H3K27me3 levels, the highest changes (|log2FC| > 2 and adjusted P value < 10-1) occurring specifically at transcription start sites (TSS) and gene bodies (Fig. 2c) and corresponding to a loss of H3K27me3 enrichment (75 regions with log2FC < -2, and 2 regions with log2FC > 2). This depletion was associated with the highest changes in gene expression observed by scRNA-seq (Fig. 2d and Extended Data Fig. 5c) and to the transcriptional de-repression of 22% of persister genes – defined as genes overexpressed in persister versus untreated cells – such as TGFB1, FOXQ1, which encodes a transcription factor driving EMT (Fig. 2e, Supplementary Table 4). These epigenomic changes were not necessarily kept in resistant cell populations (Extended Data Fig. 5d), suggesting the existence of transient epigenomic features in persister cells.
Fig. 2. H3K27me3 represses the persister expression program prior to chemotherapy exposure.
All experiments were performed in MDA-MB-468 cells. a. UMAP representation of scChIP-seq H3K27me3 datasets, cells are colored according to the sample of origin. Persister and resistant samples correspond to 5-FU-treated cells, days of treatment are indicated. b. Same as in a. with cells colored according to cluster membership. E1, E2 correspond to epigenomic-based clusters. c. Enrichment of H3K27me3 significantly depleted peaks in persister cells compared to all peaks across various gene annotation categories with Fisher’s exact test (see Methods). Full bars indicate P value < 1.0 × 10-2. Empty bars indicate non-significant P values. “PC” indicates protein coding genes. d. Repartition of H3K27me3-depleted peaks within log2 expression fold-changes quantiles from scRNA-seq experiments. e. Cumulative H3K27me3 profiles over TGFB1 and FOXQ1 in untreated and persister cells (D33). Log2FC and adjusted P value correspond to differential analysis of cells from cluster E1 versus cells from clusters E2 + E4. f. Violin plot representation of the cell-to-cell inter-correlation scores between cells from clusters E1, E2 or E4 and cells from E1. Pearson’s correlation scores were compared using a two-sided Wilcoxon rank test, P value are indicated above plots. g. Dot plot representing log2 expression fold-change induced by 5-FU or EZH2i-1 at D33 versus D0. Pearson’s correlation scores and associated P value are indicated. h. Bulk H3K27me3 chromatin profiles for TGFB1 and FOXQ1 in cells treated with DMSO, 5-FU or EZH2i-1 at D33.
In the untreated cells, two epigenomic subclones (E2 & E4 clusters, Fig. 2b) were recurrently identified indicative of an epigenomic heterogeneity in this population (n = 3 experiments). In contrast to cells from cluster E4 (median correlation score r = -0.34), a fraction of cells within cluster E2 shared epigenomic similarities with cells from E1 (Fig. 2f, 49/381 cells over r = 0.20, P < 2.2 × 10-16). Yet these similarities did not affect persister genes (Extended Data Fig. 5e), and cells remained discernible from the pool of persister cells (no cells from E2 with r score over median r in E1, P < 2.2 × 10-16). This suggests that cells from E2 could fuel the persister population when exposed to chemotherapy, with the need of chemotherapy-induced chromatin changes to achieve tolerance. In addition, we also detected rare cells with a persister epigenomic signature, but in only one of our three experiments (60/976 cells – Extended Data Fig. 5b), suggesting that spontaneous transition to H3K27me3 drug-tolerant state rarely occurs in the absence of chemotherapy.
Persister expression program is locked by H3K27me3
To test whether H3K27me3 enrichment was the lock to the persister expression program in untreated cells, we treated cancer cells with the EZH2 inhibitor (EZH2i-1) UNC199931, to deplete H3K27me3 from cells, in the absence of chemotherapy. EZH2i-1 treatment phenocopied a drug-tolerant state as expression fold-changes induced by EZH2i-1 were specifically correlated to those induced by chemotherapy exposure at early time points in persister cells (Fig. 2g, Extended Data Fig. 5f, r = 0.71 versus r = 0.31 with changes in resistant cells). Furthermore, we observed that EZH2i-1 was sufficient to lead to the activation of 62% of persister genes with depletion of H3K27me3 upon 5-FU treatment (23/37 genes), suggesting that H3K27me3 was the main lock to their activation (Fig. 2h and Extended Data Fig. 5g). EZH2i-1 was also sufficient to lead to the overexpression of 60% of persister genes independently of any H3K27me3 enrichment in untreated cells (78/131 genes), such as KRT14, suggesting that these genes might be targets of H3K27me3-regulated persister genes. To further test this hypothesis, we explored the existence of potential ‘master’ transcription factors (TFs) within the persister genes. Using CheA332, we identified three transcription factors, FOXQ1, FOSL1 and N2RF2, that respectively target 34%, 42% and 29% of the persister genes (Extended Data Fig. 5h-i). All 3 are H3K27me3-regulated persister genes, with a loss of H3K27me3 upon 5-FU (Extended Data Fig. 5j) and re-expressed by EZH2i-1 treatment, nominating these H3K27me3-regulated TFs as potential drivers of the persister expression program.
Priming of the persister expression program
As we observed H3K27me3 changes upon 5-FU treatment precisely at TSSs, we further explored the evolution of chromatin modifications at TSS, focusing on H3K4me3, a permissive histone mark shown to accumulate over TSS with active transcription. In contrast to single-cell H3K27me3 landscapes, which were sufficient to separate cell states along treatment (Fig. 2a), individual H3K4me3 landscapes of untreated and persister cells were indiscernible (Fig. 3a-b). Sparse H3K4me3 enrichment was already observed in untreated cells at the TSS of persister genes (P < 10-15, compared to a set of non-expressed genes, Extended Data Fig. 6a-b). In individual persister cells, H3K4me3 enrichment was significantly more synchronous at the TSS of persister genes with more persister genes simultaneously marked by H3K4me3 in the same cell than in untreated cells (P = 4.0 × 10-2, Extended Data Fig. 6c-d). Based on these results, we reasoned that in untreated cells H3K4me3 and H3K27me3 could either accumulate on the same TSS but in different cells, or H3K4me3 could accumulate together with H3K27me3 in a subset of cells on the same TSS.
Fig. 3. Epigenomes of untreated cells are primed with co-accumulation of H3K27me3 and H3K4me3.
a. UMAP representation of scChIP-seq H3K4me3 datasets, cells are colored according to the sample of origin - untreated cells (D0) and persister cells (D60). b. (Up) Cumulative H3K4me3 enrichment profiles over FOXQ1 in untreated cells (D0) and persister cells (D60). ‘ns’ stands for not significant after differential testing comparing untreated and persister cells. (Down) H3K27me3→H3K4me3 and H3K27me3→IgG sequential ChIP-seq profiles of FOXQ1 in the untreated population. Comparative tracks show enrichment over IgG control with associated odd ratio and adjusted P value from Fisher’s exact test adjusted for multiple testing. c. Doughnut chart displaying the fraction of persister genes (n = 168) with H3K27me3 loss upon 5-FU treatment or with bivalent chromatin at TSS in untreated cells. Candidate master TFs - among persister genes - are indicated with the number of persister genes potentially regulated by the corresponding TF in parentheses. d. (Left) H3K27me3 and H3K4me3 chromatin profiles of human tumor samples for candidate master TFs (Patient_39, Patient_95 and Patient_172). The percentage of tumoral cells are indicated for each sample. (Right) H3K4me3→H3K27me3 and H3K4me3→IgG sequential ChIP-seq profiles of untreated population of the three derived PDX_models (PDX_39, PDX_95 and PDX_172). Comparative tracks show enrichment over IgG control with associated odds ratio and adjusted P value from Fisher’s exact test adjusted for multiple testing. e. H3K27me3 and H3K4me3 chromatin profiles for FOXQ1 of 6 additional human tumor samples. The percentage of tumoral cells are indicated for each sample. f. Dotplot showing the top pathways enriched in genes displaying a dual H3K27me3 and H3K4me3 enrichment in human tumor samples. Color of the dot corresponds to adjusted P values - calculated with hypergeometric test adjusted for multiple testing - and the size of the dot corresponds to the gene ratio, i.e. the fraction of bivalent genes belonging to this pathway. Stars indicate human tumor samples used to establish our PDX models.
To test whether H3K4me3 could co-exist with H3K27me3 in the same individual cells prior to chemotherapy exposure, we performed successive immunoprecipitation of H3K27me3 with H3K4me3 (or vice-versa) or H3K27me3 (or H3K4me3) with isotype control (IgG) on mono-nucleosome chromatin. In MDA-MB-468 cells, we detected 1,547 TSSs significantly enriched in DNA immunoprecipitated with both H3K27me3 and H3K4me3, compared to the control (H3K27me3/IgG) precipitated fraction (peak-ratio > 0.15, q-value < 10-3, Extended Data Fig. 6e-h). We found that bivalent chromatin in untreated cells was detected at TSSs of genes associated to mammary stem cell and EMT pathways, as well as various developmental pathways (e.g. Hedgehog pathway, Extended Data Fig. 6h-i). The majority of H3K27me3-regulated persister genes (26 out of 37), including the 3 candidate master TFs, were found in a bivalent chromatin configuration in the untreated cell population (e.g. TGF-β1, FOXQ1, FOSL1, Fig. 3b-c, Extended Data Fig. 6d and Supplementary Table 4). We next studied bivalent chromatin landscapes in additional TNBC cell lines (HCC38 & BT20, Extended Data Fig. 6i), in our 3 PDX models and n = 9 patient samples – 3 of which correspond to the tumors used for PDX derivation (Patient_95, Patient_39 & Patient_172) (Fig. 3d-f, Extended Data Fig. 7). We confirm the existence of bivalent programs in all patient tumors, and show that paired PDX models recapitulate bivalency of patient samples (Extended Data Fig. 7a-f). Bivalent chromatin is also found at genes implicated in EMT and mammary stem cell identity, and developmental pathways (Fig. 3f, Extended Data Fig. 7h) and at candidate master TFs of the persister programs – FOXQ1, KLF4 or TFCP2L1 – (Fig. 3d-e, Extended Data Fig. 7g). With a continuum of epigenomic datasets, from patients to PDX, we demonstrate that cancer cells display a set of bivalent TSSs, which could lead to the rapid activation of an entire persister expression program upon therapeutic stress.
Depriving cells of H3K27me3 enhances chemotolerance
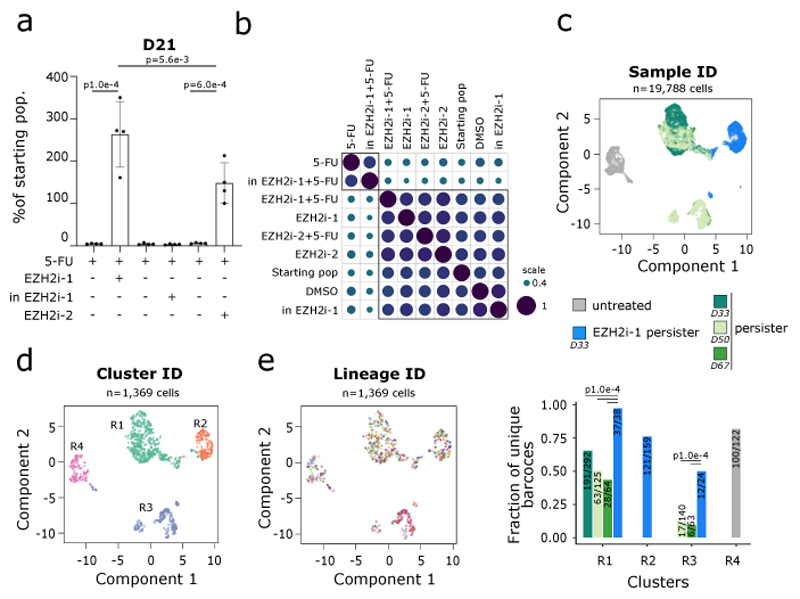
To further test if H3K27me3 was a lock to the emergence of persister cells under chemotherapy exposure, we next depleted H3K27me3 from epigenomes prior to chemotherapy treatment in three TNBC cell lines (MDA-MB-468, BT20 and HCC38, Extended Data Fig. 8). Using EZH2i-1, in addition to an inactive isomer (UNC240033) and a second EZH2i (GSK12634 referred to as EZH2i-2), we showed that erasing H3K27me3 – without perturbing EZH2 protein levels – increased the number of persister cells with either EZH2 inhibitor, while the inactive isomer had no effect (Fig. 4a, Extended Data Fig. 8). By comparing bulk lineage barcode frequencies across conditions, we further demonstrated that EZH2 inhibitors were actually affecting cell fate under chemotherapy exposure, by rescuing the biased lineage frequency induced by 5-FU treatment (Fig. 4b, Extended Data Fig. 8c). We observed that co-treatment with EZH2i-1 or EZH2i-2 increased correlation scores between lineage frequencies in 5-FU- or DMSO-treated populations, whereas co-treatment with the inactive isomer had no effect on lineage frequencies (Fig. 4b, Extended Data Fig. 8c). Performing combined single-cell transcriptomics and lineage tracing as in Figure 1, we then compared the fraction of unique barcodes across expression clusters under 5-FU exposure with or without EZH2i-1, termed “EZH2i-1 persister” and “persister” respectively (Fig. 4c-e). We observed that co-treating cells with EZH2i-1 significantly increased the fraction of unique lineage barcodes in CDH2+ persister populations, identified in Figure 1 (R3 cluster, P < 10-4 and over 2 fold-increase, Fig. 4e). Altogether, our results showed that H3K27me3 depletion with EZH2 inhibitors rescued the biased lineage frequency observed under chemotherapy treatment, and enabled a wider variety of cells to switch to the CDH2+ drug-tolerant state. Overall, depleting H3K27me3 from untreated cells not only launched a persister-like expression program, but it also enhanced the potential of each cancer cell to tolerate chemotherapy.
Fig. 4. EZH2 inhibition rescues cell fate bias upon chemotherapy exposure.
All the experiments were performed in MDA-MB-468 cells. a. Histogram representing the number of cells after treatment with 5-FU alone or 5-FU and EZH2i over 21 days, relative to the number of cells at D0. Cells were pre-treated with EZH2i-1, inactive EZH2i-1 or EZH2i-2 for 10 days prior to chemotherapy treatment. (n = 3, Mean ± s.d., ANOVA test). b. Clustering of samples according to lineage barcode frequencies, detected by bulk analysis, using Spearman correlation score. c. UMAP representation of scRNA-seq datasets, colored according to the sample of origin. Cells were treated with DMSO (untreated) or with 5-FU alone (persister) or with 5-FU and EZH2i-1 (EZH2i-1 persister). d. UMAP representation of scRNA-seq datasets, barcoded cells were selected and colored according to hierarchical clusters e. (Left) UMAP representation of scRNA-seq datasets, barcoded cells were selected and colored according to lineage barcode. (Right) Histogram of the lineage barcodes diversity detected in the scRNA-seq data within hierarchical clusters, and across samples. Colors correspond to sample ID as in 4c. Ratio of unique barcodes/total barcodes and P value are indicated (two-sided Fisher’s test).
Preventing H3K27me3 demethylation inhibits chemotolerance
Finally, we tested our ability to inhibit the emergence of persister cells by preventing the depletion of H3K27me3 under chemotherapy exposure using a KDM6A/B inhibitor (KDM6A/Bi – GSK-J435) simultaneously to chemotherapy. We tested in vitro and in vivo the ability of GSK-J4 to reduce the pool of persister cells upon chemotherapy exposure, reasoning that a reduced number of persister cells would increase the delay to recurrence. In vitro, in opposition to EZH2i, co-treatment with KDM6A/Bi led to a decrease in the number of persisters at day 21 and further abolished the growth of persister cells under 5-FU at day 60, whereas it had no effect on untreated cancer cells (Fig. 5a, Extended Data Fig. 9a-c). Interestingly, administrating KDM6i once persister cells have emerged, rather than in combination to chemotherapy, is inefficient (Fig. 5b, Extended Data Fig. 9d), demonstrating that the switch from untreated to drug-tolerant state, rather than the drug-tolerant state itself, was sensitive to KDM6i. These results were confirmed in two additional TNBC cell lines, BT20 and HCC38 (Extended Data Fig. 9e-f). In vivo, our objective was to test the potential of KDM6i to limit the emergence of persister cells, when administered simultaneously to chemotherapy. We compared the disease-free survival time of mice treated with Capecitabine alone (n = 25) or in combination with KDM6i (n = 25). The administration of KDM6i did not inhibit tumor progression in absence of chemotherapy nor was toxic for the mice (Fig. 5c). However, administered in combination to chemotherapy, it significantly increased the delay to recurrence (Fig. 5d, P = 4.0 × 10-2) in comparison to chemotherapy alone. Our in vitro and in vivo results together suggest that a fraction of cancer cells could need to actively demethylate H3K27me3 to tolerate chemotherapy. These results are consistent with a mechanism where persister genes would be repressed by H3K27me3 in untreated cells, and primed with stochastic H3K4me3 in a subset of cells, with the loss of H3K27me3 unlocking the transition to tolerance.
Fig. 5. Simultaneous KDM6i and chemotherapy treatment inhibits chemotolerance in vitro and delays recurrence in vivo.
a. Colony forming assay at day 60 for 5-FU treated MDA-MB-468 cells in combination with DMSO or indicated concentrations of the KDM6i GSK-J4. b. Colony forming assay at day 60 for 5-FU treated MDA-MB-468 cells in combination or not with 1 µM of GSK-J4 or its inactive isomer GSK-J5, either simultaneously - added at D0 - or added at day 39 of chemotherapy treatment. c. Relative tumor volumes for n = 60 mice treated with either DMSO, GSK-J4, Capecitabine or a combination of Capecitabine and GSK-J4. Dashed line indicates the threshold to detect recurrent tumors, RTV = 3. d. Kaplan-Meier plot of the overall disease-free survival probability since pathologic complete response pCR to initial treatment (tumor volume < 20 mm3), number of mice treated and P value are indicated (log-rank test).
Discussion
Our study shows that H3K27me3 landscapes are determinants of cell fate upon chemotherapy exposure in TNBC. We demonstrate that, prior to treatment, cells display bivalent chromatin landscapes priming the persister expression program with H3K4me3 and H3K27me3. In other words, genes are ready to be activated with H3K4me3, but are repressed with H3K27me3, which remains the lock to the activation of the persister expression program. Using EZH2 inhibitors and lineage tracing strategies, we further demonstrate that, depleting H3K27me3 from the epigenome rescues the cell fate bias normally observed upon chemotherapy insult; cells have an equal probability of surviving initial chemotherapy insult. Persister cells could be cells without H3K27me3 or the one releasing the H3K27me3 lock, or a mixture of both phenomena as shown here: co-treating cells with a H3K27me3 demethylase inhibitor together with 5-FU, we reduced, but not totally abrogated the number of persister cells. We propose that combining chemotherapy with histone demethylase inhibitors at the onset of chemotherapy exposure can delay recurrence by decreasing the pool of persister cells. Several studies had started to interrogate which epigenetic modifiers could regulate expression programs of persister or resistant cells10,34,36,37. Here, we show that even prior to treatment, the epigenome is already a key player, with a priming of the persister program. Our findings highlight how chromatin landscapes can shape the potential of cancer cells for chemotolerance.
Our finding that depletion of H3K27me3 with EZH2 inhibitors – at the onset of treatment – enhances chemotolerance leaves open questions regarding the role of H3K27me3 landscapes in cancer evolution, and the usage of EZH2i in cancer treatment. Depending on the context and the timing of administration, EZH2i might have drastically different effects, and such subtleties should be carefully considered before therapy combination. In contrast to our observations at the onset of treatment, TNBC cancer cells with long-term acquired chemo-resistance have been shown to display DNA hypomethylation and large H3K27me3 domains over transposable elements, hence a vulnerability to EZH2i38. EZH2i were also recently shown to lead to MHC Class I upregulation in cancer cells, thereby showing beneficial immunotherapeutic effects39,40. If such cell plasticity represents a therapeutic opportunity, our results also show that EZH2i could also lead, in some contexts, to the activation of a set of genes driving drug-persistence.
Isolated examples of bivalent promoters had been found in tumor cells41,42. Here we exhaustively map bivalent promoters genome-wide, revealing epigenomic priming of mammary stem cell genes and signaling pathways of known resistance pathways in TNBC43, including Hedgehog, WNT, TGF-β, and ATP-binding cassette drug transporters pathways. Such epigenomic priming is reminiscent of developmental bivalency priming mechanisms44 found in stem cells prior to differentiation and appears key for the rapid activation of the genes upon therapeutic stress. It remains to be understood how only a minority of bivalent genes are targeted by gene reactivation upon chemotherapy exposure – which could be associated to the nature of the treatment itself – and whether such priming mechanisms could be shared across cancer types. Determining the precise mechanisms that target H3K27me3 and H3K4me3 writers and readers to the TSSs of cancer-type-specific bivalent genes under therapeutic stress will be instrumental to identify co-factors that might serve as therapeutic targets to further limit the emergence of persister cells.
Methods
Our research complies with all relevant ethical regulations. Mouse care and housing were in accordance with institutional guidelines and the rules of the French Ethics Committee (project authorization no. 02163.02). Patient samples were studied after approval by the Institutional Review Committee of Institut Curie.
Patient samples and PDX models
Patient samples used in this study (n = 9) originated from patients treated at Institut Curie with residual triple-negative breast cancers post-neoadjuvant chemotherapy, who gave informed consent for the profiling. In this study, we used three xenograft models generated from three different residual triple-negative breast cancers post-neoadjuvant chemotherapy (HBCx95 called PDX_95, HBCx39 called PDX_39 and HBCx172 called PDX_172 in the manuscript, see Supplementary Table 5) established previously at Institut Curie with informed consent from the patient45,46. All female Swiss nude mice used in this study (6-7 weeks of age at the beginning of all experiments) were purchased from Charles River Laboratories and maintained under specific-pathogen-free conditions. Mouse care and housing were in accordance with institutional guidelines and the rules of the French Ethics Committee (project authorization no. 02163.02). The maximal tumor size permitted by the ethics committee is 2,000 mm3. For all mice, the maximal tumor size did not exceed 1,500 mm3.
Figure 1b: Five mice were not treated and kept as controls (termed “untreated”) and twenty-seven mice were treated orally with Capecitabine (Xeloda; Roche Laboratories) at a dose of 540 mg/kg, 5 d/week for 6 to 14 weeks. Relative tumor volumes (mm3) were measured as described previously17. Eight mice were sacrificed after the first round of chemotherapy to study “residual” tumors (2 mice) or “persister” (6 mice) human tumor cells. Seven mice with “recurrent” tumors (tumor volume between 200 and 600 mm3) were treated with a second round of Capecitabine to which they responded or not. “Resistant” refers to a tumor that maintains a constant volume under this second round of treatment.
Extended Data Figure 1f: Three mice were not treated and kept as controls (termed “untreated”) and fourteen mice were treated orally with Capecitabine at a dose of 540 mg/kg, 5 d/week for 7 weeks and sacrificed to study “persister” human tumor cells.
Extended Data Figure 1l: Six mice were not treated and kept as controls (termed “untreated”) and four mice were treated orally with Capecitabine for 7 weeks and sacrificed to study “persister” human tumor cells.
Figure 5c: Five mice were treated intraperitoneally with DMSO, five mice were treated intraperitoneally with GSK-J4 alone at a dose of 50 mg/kg, 5 d/week for 25 days. Twenty-five mice were treated orally with Capecitabine at a dose of 540 mg/kg, 5 d/week for 36 days and twenty-five mice were co-treated with Capecitabine and GSK-J4 for 36 days. Tumor volumes (mm3) were measured to follow recurrence. Figure 5d: Disease-free survival was defined as the number of days between the observation of a complete response (relative tumor volume RTV compared to volume at onset of treatment < 0.2) after the first round of Capecitabine treatment, and the appearance of a recurrent tumor (RTV > 3). Statistical analysis was performed using a log-rank test.
Before downstream analysis (scChIP-seq, scRNA-seq or sequential ChIP-seq), control and treated tumors were digested for 2 h at 37 °C with a cocktail of Collagenase I (Roche, Ref: 11088793001) and Hyaluronidase (Sigma-Aldrich, Ref: H3506). Cells were then individualized at 37 °C using a cocktail of 0.25% Trypsin-Versen (Thermo Fisher Scientific, Ref: 15040-033), Dispase II (Sigma-Aldrich, Ref: D4693) and DNase I (Roche, Ref: 11284932001) as described previously47. Then, eBioscience red blood cell lysis buffer (Thermo Fisher Scientific, Ref: 00-4333-57) was added to the cell suspension to remove red blood cells. To increase the viability of the final cell suspension, dead cells were removed using the Dead Cell Removal Kit (Miltenyi Biotec, Ref:130-090-101).
Cell lines, culture conditions and drug treatments
MDA-MB-468 cells (ATCC, HTB-132) were cultured in DMEM (Gibco-BRL, Ref: 11966025), supplemented with 10% heat-inactivated fetal calf serum (Gibco-BRL, Ref: 10270-106). HCC38 (ATCC, CRL-2314) and BT20 (ATCC, HTB-19) cell lines were cultured in RPMI 1640 (Gibco-BRL, Ref: 11875085), supplemented with 10% heat-inactivated fetal calf serum. All cell lines were cultured in a humidified 5% CO2 atmosphere at 37 °C, and were tested as mycoplasma negative. GSK-J4 (KDM6A/B inhibitor, Sigma, Ref: SML0701), GSK-J5 (GSK-J4 inactive isomer, Abcam, Ref: ab144397), UNC1999 (EZH2 inhibitor, Abcam, Ref: ab146152), UNC2400 (UNC1999 inactive isoform, Tocris, Ref: 4905) and GSK126 (EZH2 inhibitor, Sigma, Ref:) were used at indicated concentrations. Cells were treated with 5 μM of 5-FU (Sigma, Ref: F6627) alone or in combination with KDM6A/Bi or EZH2i for indicated days. For EZH2i, cells were pretreated with UNC1999, UNC2400 or GSK126 for 10 days before the addition of 5-FU for an additional 21 days (Fig. 4 and Extended Data Fig. 8).
Colony forming assay
TBNC cells were plated in 6 multi-well plates at a density of 200,000 cells per well and treated with the indicated drugs for 60 days (MDA-MB-468, Fig. 5a/b and Extended Data Fig. 9b) or 56 days (BT20) or 50 days (HCC38) (Extended Data Fig. 9). Cultures were incubated in humidified 37 °C incubators with an atmosphere of 5% CO2 in air, and treated plates were monitored for growth using a microscope. At the time of maximum foci formation, colony formation was evaluated after a staining with 0.5% Crystal Violet (Sigma, ref: C3886).
Cell proliferation, doubling time and IC50
MDA-MB-468, HCC38 and BT20 cells were stained with Trypan Blue (Invitrogen, Ref: T10282) exclusion test, and counted using a Countess automated cell counter (Invitrogen, Ref: C10228) at indicated time of treatment (Fig. 4a and Extended Data Fig. 8a/d/f).
Doubling time (Extended Data Fig. 2b) was calculated using this formula: DoublingTime = duration × log(2) / (log(Final Concentration) - log(Initial Concentration)).
For untreated condition and resistant condition, cell numbers were evaluated on cell population during 10 days (n = 3). For persister condition, cells were counted manually under the microscope at day 13 and day 30 of treatment. Doubling time of 5-FU dividing persister cells was studied from single cell to confluent colony by assaying cell number every 4 days during 27 days (n = 9 single cells).
MDA-MB-468 untreated and chemoresistant cells were plated in 96 multi-well plates at a density of 10,000 cells per well and treated with increased concentration of 5-FU (1 μM to 0.5 M) for 72 h. Cell cytotoxicity was assayed with XTT kit (Sigma, Ref: 11465015001) and IC50 was calculated as the concentration of 5-FU that is required to obtain 50% of cell viability (Extended Data Fig. 2b).
Cells were classified as ‘persister’, ‘growing persister’ or ‘resistant’ based on a combination of 2 biological markers : (i) doubling time under 5-FU, (ii) IC50 to 5-FU (Extended Data Fig. 2b, Fig. 1e):
-
-
‘persister’ correspond to non-dividing cells (infinite doubling time) or cells dividing with a doubling time significantly higher than resistant cells under 5-FU
-
-
‘resistant’ correspond to cells with a doubling time comparable to untreated cells and a significantly higher IC50 to 5-FU compared to untreated cells
It should be noted that due to low number of persister cells (0.01% of initial population), we could not measure the IC50 to 5-FU of these two states.
For Extended Data Figure 9d cells were treated with 5-FU at 5 µM and indicated concentrations of GSK-J4 or GSK-J5 (D-5 indicated 5 days pre-treatment with GSK-J4 or GSK-J5 before 5-FU treatment (D0), D0 indicated co-treatment 5-FU and GSK-J4 or GSK-J5, D10 and D30 indicated that treatment with GSK-J4 or GSK-J5 started 10 days or 30 days respectively after the onset of 5-FU treatment (D0). The number of persister cells were counted manually under the microscope at day 42 (n = 3).
The GraphPad PRISM 9 was used for statistics and the results represent the mean ± s.d. of three independent experiments. Statistical analysis was performed using the Bonferroni test for multiple comparisons between samples (Fig. 4a, Extended Data Fig. 8a/d/f, Extended Data Fig. 9b/d and Extended Data Fig. 2b-right) or one sided Wilcoxon rank test for the comparison between two conditions (Extended Data Fig. 2b-left).
Western blotting
In Extended Data Figure 8b/e/g, DMSO- and EZH2i-treated cells were lysed at 95 °C for 10 minutes in Laemmli buffer (50 mM Tris-HCl [pH 6.8], 2% SDS, 5% glycerol, 2 mM DTT, 2.5 mM EDTA, 2.5 mM EGTA, 4 mM Sodium Orthovanadate, 20 mM Sodium Fluoride, protease inhibitors, phosphatase inhibitors) and proteins concentrations were measured using a Pierce BCA protein Assay Kit (Thermo Fisher Scientific, Ref: 23225/23227). 10 µg of proteins were then separated on a 4-15% Mini-PROTEAN TGX Stain-Free Gel (Bio-Rad, Ref: 4568085) at 160V. After transfer, the membrane was blocked for 1 h at room temperature in PBS pH 7.4 containing 0.1% Tween-20 and 1% milk (Regilait). Incubation anti-H3K27me3 (1/2,000, Cell Signaling, Ref: 9733) or EZH2 (1/2,000, Cell Signaling, Ref: 5246) or Tubulin (1/1,000, Thermo Fisher Scientific, Ref: 31460) primary antibodies diluted in PBS pH 7.4, 0.1% Tween-20 were performed at 4 °C overnight. Following 2 h incubation at room temperature with an anti-rabbit or mouse peroxidase-conjugated secondary antibody (1/10,000, Thermo Fisher Scientific, Ref: 31460 or Ref: 31430) diluted in PBS pH 7.4, 0.1% Tween-20, antibody-specific labeling bands were revealed (Bio-Rad, ChemiDoc MP) using a SuperSignal West Pico PLUS Chemiluminescent Substrate (Thermo Fisher Scientific, Ref: 34579).
Lentivirus packaging and cell transduction
Lentivirus was produced by transfecting the barcode plasmids pRRL-CMV-GFP-BCv2AscI and p8.9-QV and pVSVG into HEK293T cells as previously described30. MDA-MB-468 cells from ATCC were infected at passage 11 with lentivirus produced from the barcode library (pRRL-CMV-GFP-BCv2AscI) which includes 18,206 different barcodes of 20 bp of a random stretch, at a low multiplicity of infection (MOI 0.1) to minimize the number of cells marked by multiple barcodes. Three weeks after transduction, cells were sorted for GFP expression to select cells with barcode insertion, and used for drug treatment.
Single-cell RNA-seq
For each single cell suspension (DMSO-D0-#1, 5-FU-D33-#1, 5-FU-D214-#1, 5-FU-D67-#2, 5-FU-D171-#2, 5-FU-D50-#3, 5-FU-D77-#3 and 5-FU-D202-#3) or PDX dissociated cells (PDX_95, PDX_39 or PDX_172, untreated and persister cells), approximately 3,000 cells were loaded on a Chromium Single Cell Controller Instrument (Chromium Single Cell 3'v3, 10X Genomics, Ref: PN-1000075) according to the manufacturer’s instructions. Samples and libraries were prepared according to the manufacturer's instructions. Libraries were sequenced on a NovaSeq 6000 (Illumina) in PE 28-8-91 with a coverage of 50,000 reads/cell.
Bulk lineage barcode library preparation and sequencing
Lineage barcodes are recovered by isolating genomic DNA from cells of interest (NucleoSpin Tissue, Mini kit for DNA from cells and tissue, Macherey Nagel, Ref: 740952.50). From the isolated genomic DNA, barcodes are amplified with three nested PCR steps as described in29 (see Supplementary Table 6 for primer sequence). In short, after a first specific PCR for the common region of the lineage barcodes, the amplified material was prepared for sequencing by addition of the Illumina sequencing adapters and indexing and purification. Sequencing was done in order to obtain 50 reads, on average, per barcoded cell.
Bulk ChIP-seq
ChIP experiments were performed as previously described15 on 3x106 MDA-MB-468 cells (DMSO-D67-#2, DMSO-D77-#3, DMSO-D113-#4, 5-FU-D67-#2, 5-FU-D77-#3, 5-FU-D113-#4) using an anti-H3K27me3 antibody (1/50, Cell Signaling Technology, Ref: 9733 - C36B11). Sequencing libraries were prepared using the NEBNext Ultra II DNA Library Prep Kit (NEB, Ref: E7645S) according to the manufacturer’s instructions. Libraries were sequenced on a NovaSeq 6000 (Illumina) in SE50 mode.
Single-cell ChIP-seq
Cells (DMSO-D60-#1, DMSO-D77-#3, DMSO-D131-#5, 5-FU-D33-#1, 5-FU-D67-#2, 5-FU-D171-#2, 5-FU-D147-#3, 5-FU-D131-#6) were labeled by 15 min incubation with 1 μM CFSE (CellTrace CFSE, Thermo Fisher Scientific, Ref: C34554). Cells were then resuspended in PBS supplemented with 30% Percoll, 0.1% Pluronic F68, 25 mM HEPES pH 7.4 and 50 mM NaCl. Cell encapsulation, bead encapsulation and 1:1 droplet fusion was performed as previously described15, see Supplementary Table 6 for the sequence of bead barcodes. Immunoprecipitation with H3K27me3 antibody (1/200, Cell signaling, Ref: 9733 - C36B11) or H3K4me3 antibody (1/200, Cell signaling, Ref: 9751-C42D8), DNA amplification and library were performed as in15. Libraries were sequenced on a NovaSeq 6000 (Illumina) in PE100, with 4 dark cycles on Read 2, with a coverage of 100,000 reads/cell.
Quantitative chromatin profiling with chromatin indexing
Chromatin isolation, indexing, immunoprecipitation and library preparation was adapted from48. Briefly, 50,000 MDA-MB-468 were lysed and digested with MNase for 20 min at 37 °C in the following buffer: 46 mM Tris-HCl pH 7.4, 0.154 M NaCl, 0.1% Triton, 0.1% NaDoc, 4.65 mM CaCl2, 0.47× Protease Inhibitor Cocktail (Roche, Ref: 11873580001) and 0.09 U/µl MNase (Thermo Scientific, Ref: EN0181). Fragmented nucleosomes were then ligated for at least 24 h at 16 °C to double-stranded barcoded adapters containing 8-bp barcodes to combine samples: Pac1-T7-Read2-8bpBarcode-linker-Pac1 (Extended Table 1). Next, 5 indexed chromatin samples (DMSO, 5-FU, UNC, 5-FU + UNC, GSK-J4) were pooled, each containing a different 8-bp barcode, to perform anti-H3K27me3 ChIP (1/50, Cell Signaling, Ref: 9733 - C36B11) on 250,000 cells in total in each pool. ChIP and DNA amplification was carried out as for scChIP-seq15 and a sequencing library was produced for both IP and input pools and sequenced on NovaSeq 6000 (Illumina) in PE100 mode.
Sequential ChIP-seq
Primary ChIP experiments were performed as described previously15 on 10 × 106 untreated MDA-MB-468, BT20 or HCC38 cells or untreated PDX_95, PDX_39 or PDX_172 tumor dissociated cells using the anti-H3K27me3 antibody (1/50, Cell Signaling, Ref: 9733 - C36B11 – MDA-MB-468) or anti-H3K4me3 antibody (1/50, Cell Signaling, Ref: 9751-C42D8 – MDA-MB-468-bis, BT20, HCC38 and PDX models). After washes, samples were eluted twice at 37 °C for 15 min under agitation in an elution buffer (50 mM Tris-HCl pH8, 5 mM EDTA, 20 mM DTT, 1% SDS) as in. Samples were diluted 10 times to decrease SDS and DTT concentration. 10% of the eluted chromatin was kept as primary ChIP. Secondary ChIP, re-ChIP, was performed overnight on the rest of the primary immuno-precipitated chromatin using an anti-H3K4me3 antibody (MDA-MB-468) or anti-H3K27me3 (MDA-MB-468-bis, BT20, HCC38 and PDX models) or using an anti-IgG antibody (1/250, Cell signaling, Ref: 3900 – all samples) as a control, to determine the background level of the re-ChIP experiment. After washes, samples were eluted twice at 65 °C for 15 min under agitation in 0.1 M NaHCO3 and 1% SDS as in49. After reverse crosslinking and DNA clean-up, 3 to 15 ng of immunoprecipitated DNA were used to prepare the sequencing libraries using the NEBNext Ultra II DNA Library Prep Kit (NEB, Ref: E7645S) according to the manufacturer’s instructions. Libraries were sequenced on a NovaSeq 6000 (Illumina) in SE100 mode. For MDA-MB-468, we verified that the two ways (H3K27me3→H3K4me3 and H3K4me3→H3K27me3) yielded similar results. We found a significant overlap of the 1,547 and 2,490 bivalent genes obtained with the two ways (P = 2.2 × 10-16, Extended Data Fig. 6g) and found that the enriched pathways were strongly correlated (Pearson’s r = 0.81, Extended Data Fig. 6h).
CUT&Tag on frozen tumor samples
CUT&Tag was performed as in16 with minor modifications on 50,000 to 100,000 nuclei with indicated antibody (1/50, Cell Signaling Antibodies : Anti-H3K27me3, Ref: 9733- C36B11, Anti-H3K4me3, Ref: 9751- C42D8)16,50. All washes were performed in a volume of 500 µl and all centrifugations were done using a swinging bucket centrifuge at 1,300g, 4 min, at 4 °C for nuclei preparation and 600g, 8 min, 4 °C for subsequent steps. Nuclei were extracted and permeabilized from 10-20 mg frozen tumor tissues by incubating samples 10 min on ice in 6 ml ice-cold NE1 buffer (20 mM HEPES pH7.2, 10 mM KCl, 0.5 mM spermidine, 20% glycerol, 1%BSA, 1% NP-40, 0.01% digitonin, 1× proteases inhibitor) after mechanical dissociation. Following antibody incubation and tagmentation, samples were incubated for 1 h at 55 °C with max speed agitation with 3 µl 10% SDS and 2.5 µl 20 mg/ml proteinase K. After DNA extraction (Qiagen, Ref: 139046 MaXtract High density), PCR amplification (with 17 cycles, 20 s at 63 °C combined annealing/extension step) of the sequencing libraries was performed and profiles were checked on the Agilent TapeStation using High-sensitivity D1000 reagents. CUT&Tag libraries were sequenced on a NovaSeq 6000 (Illumina) in PE50 mode.
Whole-exome sequencing
Genomic DNA from samples (DMSO-D0, DMSO-D147-#3, DMSO-D171-#5, DMSO-D131-#6, 5-FU-D67-#2, 5-FU-D153-#2, 5-FU-D50-#3, 5-FU-D147-#3, 5-FU-D171-#5 and 5-FU-D131-#6) were extracted with NucleoSpin Tissue, Mini kit for DNA from cells and tissue (Macherey Nagel, Ref: 740952.50) and sequenced on a NovaSeq 6000 (Illumina) with a 100× depth.
Extended Data
Extended Figure 1.
Extended Figure 2.
Extended Figure 3.
Extended Figure 4.
Extended Figure 5.
Extended Figure 6.
Extended Figure 7.
Extended Figure 8.
Extended Figure 9.
Supplementary Material
Acknowledgments
Single-cell experiments were performed with the Single-Cell platform of the Institut Curie. We thank Antonin Morillon for critical reading of the manuscript. This work was supported by the ATIP Avenir program, by Plan Cancer, by the SiRIC-Curie program SiRIC Grants #INCa-DGOS-4654 and #INCa-DGOS-Inserm_12554, and by a starting ERC grant from the H2020 program #948528-ChromTrace (to C.V.), and by the Fondation de France #00107944 (to J.M.). The work was supported by an ATIP-Avenir grant from CNRS and Bettencourt-Schueller Foundation, by the Labex CelTisPhyBio #ANR-11-LABX-0038 and by a starting ERC grant from the H2020 program #758170-Microbar (to L.P.). High-throughput sequencing was performed by the ICGex NGS platform of the Institut Curie supported by the grants Equipex #ANR-10-EQPX-03, by the France Génomique Consortium from the Agence Nationale de la Recherche #ANR-10-INBS-09-08 ("Investissements d’Avenir" program), by the ITMO-Cancer Aviesan - Plan Cancer III and by the SiRIC-Curie program SiRIC Grant #INCa-DGOS-4654.
Footnotes
Author contributions Statement
J.M., A.D., C.L., L.B., S.B.T., A.E. and A.T. performed experiments. A.D. and A.M.L. contributed equally to the paper. scChIP-seq experiments were conducted together with S.F. and K.G.. PDX experiments were performed by E.M., L.S. and A.D.. M.B. and S.B. performed sequencing. P.P. and C.V. performed omics data analysis. Lineage barcoding data were analyzed by A.M.L., C.V. and L.P.. Whole-exome sequencing data were analyzed together with E.L.. C.V., L.P. and J.M. conceived and designed experiments. C.V., J.M., P.P. and L.P. wrote the manuscript with input from all authors.
Competing Interests Statement
C.V. is founder and equity holder of One Biosciences. The remaining authors declare no competing interests.
Data availability
All sequencing files were deposited to GEO under the super series GSE164716.
Code availability
All statistical analyses were performed in R (v4.1) using custom R scripts. Codes for data analysis are available at the following repositories https://github.com/vallotlab/ChemoPersistance, release v1.0.0 doi 10.5281/zenodo.6010802 and https://github.com/TeamPerie/lentiviral_barcode_detection_in10X_data/.
References
- 1.Vallette FM, et al. Dormant, quiescent, tolerant and persister cells: Four synonyms for the same target in cancer. Biochemical Pharmacology. 2019;162:169–176. doi: 10.1016/j.bcp.2018.11.004. [DOI] [PubMed] [Google Scholar]
- 2.Shen S, Vagner S, Robert C. Persistent Cancer Cells: The Deadly Survivors. Cell. 2020;183:860–874. doi: 10.1016/j.cell.2020.10.027. [DOI] [PubMed] [Google Scholar]
- 3.Ramirez M, et al. Diverse drug-resistance mechanisms can emerge from drug-tolerant cancer persister cells. Nat Commun. 2016;7:10690. doi: 10.1038/ncomms10690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Shaffer SM, et al. Rare cell variability and drug-induced reprogramming as a mode of cancer drug resistance. Nature. 2017;546:431–435. doi: 10.1038/nature22794. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Cortazar P, et al. Pathological complete response and long-term clinical benefit in breast cancer: the CTNeoBC pooled analysis. The Lancet. 2014;384:164–172. doi: 10.1016/S0140-6736(13)62422-8. [DOI] [PubMed] [Google Scholar]
- 6.Hata AN, et al. Tumor cells can follow distinct evolutionary paths to become resistant to epidermal growth factor receptor inhibition. Nat Med. 2016;22:262–269. doi: 10.1038/nm.4040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Kim C, et al. Chemoresistance Evolution in Triple-Negative Breast Cancer Delineated by Single-Cell Sequencing. Cell. 2018;173:879–893.:e13. doi: 10.1016/j.cell.2018.03.041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Echeverria GV, et al. Resistance to neoadjuvant chemotherapy in triple-negative breast cancer mediated by a reversible drug-tolerant state. Sci Transl Med. 2019;11:eaav0936. doi: 10.1126/scitranslmed.aav0936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Sharma SV, et al. A Chromatin-Mediated Reversible Drug-Tolerant State in Cancer Cell Subpopulations. Cell. 2010;141:69–80. doi: 10.1016/j.cell.2010.02.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Liau BB, et al. Adaptive Chromatin Remodeling Drives Glioblastoma Stem Cell Plasticity and Drug Tolerance. Cell Stem Cell. 2017;20:233–246.:e7. doi: 10.1016/j.stem.2016.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Rambow F, et al. Toward Minimal Residual Disease-Directed Therapy in Melanoma. Cell. 2018;174:843–855.:e19. doi: 10.1016/j.cell.2018.06.025. [DOI] [PubMed] [Google Scholar]
- 12.Nik-Zainal S, et al. Landscape of somatic mutations in 560 breast cancer whole-genome sequences. Nature. 2016;534:47–54. doi: 10.1038/nature17676. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Mazor T, et al. DNA Methylation and Somatic Mutations Converge on the Cell Cycle and Define Similar Evolutionary Histories in Brain Tumors. Cancer Cell. 2015;28:307–317. doi: 10.1016/j.ccell.2015.07.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Gaiti F, et al. Epigenetic evolution and lineage histories of chronic lymphocytic leukaemia. Nature. 2019;569:576–580. doi: 10.1038/s41586-019-1198-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Grosselin K, et al. High-throughput single-cell ChIP-seq identifies heterogeneity of chromatin states in breast cancer. Nat Genet. 2019;51:1060–1066. doi: 10.1038/s41588-019-0424-9. [DOI] [PubMed] [Google Scholar]
- 16.Kaya-Okur HS, et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat Commun. 2019;10:1930. doi: 10.1038/s41467-019-09982-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Marangoni E, et al. A New Model of Patient Tumor-Derived Breast Cancer Xenografts for Preclinical Assays. Clinical Cancer Research. 2007;13:3989–3998. doi: 10.1158/1078-0432.CCR-07-0078. [DOI] [PubMed] [Google Scholar]
- 18.Longley DB, Harkin DP, Johnston PG. 5-fluorouracil: mechanisms of action and clinical strategies. Nat Rev Cancer. 2003;3:330–338. doi: 10.1038/nrc1074. [DOI] [PubMed] [Google Scholar]
- 19.Chen Z, et al. TGF-β-induced transgelin promotes bladder cancer metastasis by regulating epithelial-mesenchymal transition and invadopodia formation. EBioMedicine. 2019;47:208–220. doi: 10.1016/j.ebiom.2019.08.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Ulanovskaya OA, Zuhl AM, Cravatt BF. NNMT promotes epigenetic remodeling in cancer by creating a metabolic methylation sink. Nat Chem Biol. 2013;9:300–306. doi: 10.1038/nchembio.1204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Shaul YD, et al. Dihydropyrimidine Accumulation Is Required for the Epithelial-Mesenchymal Transition. Cell. 2014;158:1094–1109. doi: 10.1016/j.cell.2014.07.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Liang L, Zeng M, Pan H, Liu H, He Y. Nicotinamide N-methyltransferase promotes epithelial-mesenchymal transition in gastric cancer cells by activating transforming growth factor-β1 expression. Oncol Lett. 2018 doi: 10.3892/ol.2018.7885. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 23.Oren Y, et al. Cycling cancer persister cells arise from lineages with distinct programs. Nature. 2021;596:576–582. doi: 10.1038/s41586-021-03796-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Fischer KR, et al. Epithelial-to-mesenchymal transition is not required for lung metastasis but contributes to chemoresistance. Nature. 2015;527:472–476. doi: 10.1038/nature15748. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Ryan S-L, et al. Targeting NF-κB-mediated inflammatory pathways in cisplatin-resistant NSCLC. Lung Cancer. 2019;135:217–227. doi: 10.1016/j.lungcan.2019.07.006. [DOI] [PubMed] [Google Scholar]
- 26.Zheng X, et al. Epithelial-to-mesenchymal transition is dispensable for metastasis but induces chemoresistance in pancreatic cancer. Nature. 2015;527:525–530. doi: 10.1038/nature16064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Zeng D, et al. Inhibition of Notch1 reverses EMT and chemoresistance to cisplatin via direct downregulation of MCAM in triple‐negative breast cancer cells. Int J Cancer. 2020;147:490–504. doi: 10.1002/ijc.32911. [DOI] [PubMed] [Google Scholar]
- 28.Godwin P, et al. Targeting Nuclear Factor-Kappa B to Overcome Resistance to Chemotherapy. Front Oncol. 2013;3 doi: 10.3389/fonc.2013.00120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Eisele AS, et al. Erythropoietin directly remodels the clonal composition of murine hematopoietic multipotent progenitor cells. Elife. 2022;11:e66922. doi: 10.7554/eLife.66922. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Christensen S, et al. 5-Fluorouracil treatment induces characteristic T>G mutations in human cancer. Nat Commun. 2019;10:4571. doi: 10.1038/s41467-019-12594-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Konze KD, et al. An Orally Bioavailable Chemical Probe of the Lysine Methyltransferases EZH2 and EZH1. ACS Chem Biol. 2013;8:1324–1334. doi: 10.1021/cb400133j. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Keenan AB, et al. ChEA3: transcription factor enrichment analysis by orthogonal omics integration. Nucleic Acids Research. 2019;47:W212–W224. doi: 10.1093/nar/gkz446. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Chagraoui H, et al. SCL/TAL1 cooperates with Polycomb RYBP-PRC1 to suppress alternative lineages in blood-fated cells. Nat Commun. 2018;9:5375. doi: 10.1038/s41467-018-07787-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Göllner S, et al. Loss of the histone methyltransferase EZH2 induces resistance to multiple drugs in acute myeloid leukemia. Nat Med. 2017;23:69–78. doi: 10.1038/nm.4247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Kruidenier L, et al. A selective jumonji H3K27 demethylase inhibitor modulates the proinflammatory macrophage response. Nature. 2012;488:404–408. doi: 10.1038/nature11262. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Vinogradova M, et al. An inhibitor of KDM5 demethylases reduces survival of drug-tolerant cancer cells. Nat Chem Biol. 2016;12:531–538. doi: 10.1038/nchembio.2085. [DOI] [PubMed] [Google Scholar]
- 37.Hinohara K, et al. KDM5 Histone Demethylase Activity Links Cellular Transcriptomic Heterogeneity to Therapeutic Resistance. Cancer Cell. 2018;34:939–953.:e9. doi: 10.1016/j.ccell.2018.10.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Deblois G, et al. Epigenetic Switch–Induced Viral Mimicry Evasion in Chemotherapy-Resistant Breast Cancer. Cancer Discov. 2020;10:1312–1329. doi: 10.1158/2159-8290.CD-19-1493. [DOI] [PubMed] [Google Scholar]
- 39.Burr ML, et al. An Evolutionarily Conserved Function of Polycomb Silences the MHC Class I Antigen Presentation Pathway and Enables Immune Evasion in Cancer. Cancer Cell. 2019;36:385–401.:e8. doi: 10.1016/j.ccell.2019.08.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Mahadevan NR, et al. Intrinsic immunogenicity of small cell lung carcinoma revealed by its cellular plasticity. Cancer Discov. 2021:candisc.0913.2020. doi: 10.1158/2159-8290.CD-20-0913. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Hahn MA, et al. Loss of the Polycomb Mark from Bivalent Promoters Leads to Activation of Cancer-Promoting Genes in Colorectal Tumors. Cancer Research. 2014;74:3617–3629. doi: 10.1158/0008-5472.CAN-13-3147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Chaffer CL, et al. Poised Chromatin at the ZEB1 Promoter Enables Breast Cancer Cell Plasticity and Enhances Tumorigenicity. Cell. 2013;154:61–74. doi: 10.1016/j.cell.2013.06.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Nedeljković M, Damjanović A. Mechanisms of Chemotherapy Resistance in Triple-Negative Breast Cancer—How We Can Rise to the Challenge. Cells. 2019;8:957. doi: 10.3390/cells8090957. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Bernstein BE, et al. A Bivalent Chromatin Structure Marks Key Developmental Genes in Embryonic Stem Cells. Cell. 2006;125:315–326. doi: 10.1016/j.cell.2006.02.041. [DOI] [PubMed] [Google Scholar]
- 45.Cottu P, et al. Acquired Resistance to Endocrine Treatments Is Associated with Tumor-Specific Molecular Changes in Patient-Derived Luminal Breast Cancer Xenografts. Clinical Cancer Research. 2014;20:4314–4325. doi: 10.1158/1078-0432.CCR-13-3230. [DOI] [PubMed] [Google Scholar]
- 46.Marangoni E, et al. Capecitabine Efficacy Is Correlated with TYMP and RB1 Expression in PDX Established from Triple-Negative Breast Cancers. Clin Cancer Res. 2018;24:2605–2615. doi: 10.1158/1078-0432.CCR-17-3490. [DOI] [PubMed] [Google Scholar]
- 47.Petit V, et al. Optimization of tumor xenograft dissociation for the profiling of cell surface markers and nutrient transporters. Lab Invest. 2013;93:611–621. doi: 10.1038/labinvest.2013.44. [DOI] [PubMed] [Google Scholar]
- 48.van Galen P, et al. A Multiplexed System for Quantitative Comparisons of Chromatin Landscapes. Mol Cell. 2016;61:170–180. doi: 10.1016/j.molcel.2015.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Desvoyes B, Sequeira-Mendes J, Vergara Z, Madeira S, Gutierrez C. In: Plant Chromatin Dynamics. Bemer M, Baroux C, editors. Vol. 1675. Springer; New York: 2018. Sequential ChIP Protocol for Profiling Bivalent Epigenetic Modifications (ReChIP) pp. 83–97. [DOI] [PubMed] [Google Scholar]
- 50.Bartosovic M. Single-cell CUT&Tag profiles histone modifications and transcription factors in complex tissues. Nature Biotechnology. 33 doi: 10.1038/s41587-021-00869-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All sequencing files were deposited to GEO under the super series GSE164716.
All statistical analyses were performed in R (v4.1) using custom R scripts. Codes for data analysis are available at the following repositories https://github.com/vallotlab/ChemoPersistance, release v1.0.0 doi 10.5281/zenodo.6010802 and https://github.com/TeamPerie/lentiviral_barcode_detection_in10X_data/.