Abstract
Direction selectivity is a fundamental feature in the visual system. In the retina, direction selectivity is independently computed by ON and OFF circuits. However, the advantages of extracting directional information from these two independent circuits are unclear. To gain insights, we examined the ON–OFF direction-selective ganglion cells (DSGCs), which recombine signals from both circuits. Specifically, we investigated the dendritic architecture of these neurons with the premise that asymmetries in architecture will provide insights into function. Scrutinizing the dendrites of dye-filled ON–OFF DSGCs reveals that the OFF arbors of these neurons are substantially denser. The increase in density can be primarily attributed to the higher branching seen in OFF arbors. Further, analysis of ON–OFF DSGCs in a previously published serial block-face electron microscopy dataset revealed that the denser OFF arbors packed more bipolar synapses per unit dendritic length. These asymmetries in the dendritic architecture suggest that the ON–OFF DSGC preferentially magnifies the synaptic drive of the OFF pathway, potentially allowing it to encode information distinct from the ON pathway.
Keywords: dendritic architecture, dendritic density, direction selectivity, morphology, ON–OFF, retinal ganglion cell, synaptic inputs
1. Introduction
Direction-selective retinal ganglion cells (DSGCs) respond maximally to motion in their “preferred” direction and minimally to movement in the opposite “null” direction. This computation is achieved by integrating inputs from presynaptic circuits, which are primarily composed of bipolar cells and starburst amacrine cells (reviewed by Vaney et al. 2012; Wei 2018; G. Schwartz 2021) (Figure 1A). There is a precise and detail-oriented synaptic wiring between these neurons that enables the DSGC to compute directional information across a variety of visual conditions (Briggman et al. 2011; Kim et al. 2014; Chen et al. 2016; Poleg-Polsky and Diamond 2016a, 2016b; Sethuramanujam et al. 2016, 2017).
Figure 1. Dendritic morphology of the ON and OFF arbors of ON–OFF DSGCs.
(A) Retinal cross-section illustrating the neuronal cell types involved in the direction-selective circuit. Photoreceptors stimulate bipolar cells (ON/OFF—BCs), which in turn activate downstream starburst amacrine cells (SACs) and direction-selective ganglion cells (DSGCs). DSGCs are of two types: ON DSGCs and ON–OFF DSGCs. ON DSGCs receive connections from ON bipolar cells and ON SACs, while ON–OFF DSGCs form connections with both ON and OFF bipolar cells and SACs. Importantly, the ON and OFF cells stratify at distinct strata of the inner plexiform layer (IPL). (B) Polar plot depicting the spike responses of an ON–OFF DSGC to spots of light moving in eight different directions. The ON and OFF responses are shown in red and blue, respectively. These responses were measured using loose patch recordings in the wholemount retina (see Methods). The total number of spikes generated by each direction is plotted. The radial line indicates the preferred direction of the DSGC, with the length of the line indicating the direction selectivity index (DSI) (see Methods). (C) Top panel shows an image of the dendrites of an ON–OFF DSGC filled with Alexa 488. The middle panel shows the rotated view of the same DSGC’s stratification in the inner plexiform layer. The green trace indicates the fluorescence intensity profile along the IPL depth. The minima between the two peaks was used to separate the ON and OFF arbors. The bottom panel shows the manual tracings of identified ON (red) and OFF (blue) dendrites. Scale bar represents 50 µm. (D, E) Similar to C, but show the ON (D) or OFF (E) dendrites of the DSGC separately.
DSGCs can be divided into subclasses based on the subtypes of connecting bipolar cells and starbursts. The ON DSGC responds only to increments in light intensity, as it synapses exclusively with ON bipolar cells and ON starbursts. In contrast, the ON–OFF DSGC synapses with both ON and OFF presynaptic neurons and consequently responds to both increments and decrements of light (Figure 1A) (Ariel and Daw 1982; Amthor et al. 1984). These ON and OFF neurons are housed in separate sublaminae of the inner plexiform layer (IPL), and the DSGC synapses with these neurons using distinct dendritic arbors (Famiglietti 1992; reviewed in Wei 2018) (Figure 1A). Importantly, both the ON and OFF circuits have the wherewithal to compute directional information independently (Cohen and Miller 1995; Kittila and Massey 1997; Vaney et al. 2012; Wei 2018) (Figure 1B). Therefore, in the ON–OFF DSGC, directional information is combined from two distinct direction-selective circuits.
Classically, it was believed that the ON and OFF pathways were mirror symmetric, differing solely in their preference for the polarity of light. However, studies in the past two decades clearly reveal that ON and OFF ganglion cells exhibit differences in other aspects, including kinetics and contrast sensitivity (Chichilnisky and Kalmar 2002; Zaghloul et al. 2003; Pandarinath et al. 2010; Soto et al. 2020), which can translate to encoding different aspects of the visual scene (Turner and Rieke 2016). It is becoming clear that these differences can be engendered by a number of presynaptic and postsynaptic changes in the ganglion cell circuitry (Dacey and Petersen 1992; Zaghloul et al. 2003; Sagdullaev et al. 2006; Zhang and Diamond 2009). In the case of DSGCs, presynaptic differences are observed in both starbursts and bipolar cells. For instance, ON starbursts preferentially receive glycinergic inhibition (Ishii and Kaneda 2014; Jain et al. 2022), while the OFF starburst cells receive stronger GABAergic inhibition (Chen et al. 2016). Further, the ON and OFF bipolar cells reveal differences in their feedback circuitry, which modulates the release of glutamate (Sethuramanujam et al. 2018; Huang et al. 2019; Hellmer et al. 2021).
Along with presynaptic differences, it is also possible that postsynaptic specializations in the ganglion cell process the information from the two pathways differently. For example, differences in the dendritic architecture (i.e., the dendritic arborization and synaptic patterns) can shape computations. Studies show that the dendritic architecture of distinct retinal ganglion cells determines a range of functions, starting from receptive fields (Yang and Masland 1994; Baden et al. 2016) to the cell’s ability to compute complex spatiotemporal features, including orientation selectivity (Nath and Schwartz 2016), high-definition detectors (Jacoby and Schwartz 2017) and direction selectivity (Trenholm et al. 2011; Matsumoto et al. 2019). Recently, it has been shown that small differences in the dendritic architecture can lead to distinct tuning curves in subtypes of ON DSGCs (Harris and Dunn 2023). However, given a lack of compelling data on the dendritic architecture of ON–OFF DSGCs, it is unclear whether the DSGC intrinsically processes ON and OFF information differently.
In this study, we examined the ON and OFF dendritic arbors of ON–OFF DSGCs by two-photon microscopy after filling individual cells in the wholemount retina with fluorescent dyes. Further, we also examined the synaptic input patterns in publicly available serial block-face electron microscopy (SBEM) datasets (Ding et al. 2016). Our results reveal that OFF arbors are more densely wired than ON arbors of the same DSGC. This increased density could be attributed to the higher branching observed in OFF arbors. Finally, the SBEM dataset reveals a higher density of bipolar synapses in OFF arbors compared to the ON arbors. These anatomical differences suggest that the DSGC inherently processes its ON and OFF signals differently.
2. Methods
2.1. Animals
Experiments were performed in adult C57BL/6J (RRID: IMSR_JAX:000664) mice or Hb9-EGFP (RRID: MGI_109160). All mice were between 2 and 16 months old in either sex. The animal protocols were made in accordance with the Canadian Council on Animal Care and approved by the University of Victoria’s Animal Care Committee.
2.2. Physiological Recordings
The physiological measurements were performed under conditions as described previously (Sethuramanujam et al. 2016, 2021). Mice were dark-adapted for ∼30–60 min before being anesthetized and decapitated. The retina was isolated in Ringer’s solution under infrared light. The isolated retina was then mounted on a 0.22-mm membrane filter (Millipore) with a precut window to allow light to reach the retina. In some experiments, the retina was mounted on glass coverslips coated with poly-l-lysine (Sigma). This whole mount preparation was viewed under an upright Olympus BX51 WI fluorescent microscope outfitted with a 40× water-immersion lens (Olympus Canada). The isolated retina was then perfused with warmed Ringer’s solution (35–37°C) containing 110 mM NaCl, 2.5 mM KCl, 1 mM CaCl2, 1.6 mM MgCl2, 10 mM dextrose, and 22 mM NaHCO3 that was bubbled with carbogen (95% O2:5% CO2). Perfusion rates were maintained at ∼3 mL/min. Unless otherwise noted, all reagents were purchased from Sigma-Aldrich Canada.
Electrophysiological measurements were made from ganglion cells in the mid-peripheral retina of all quadrants. Spike recordings were made with the cell-attached loose patch-clamp technique using 5–10 MΩ electrodes filled with Ringer’s solution. DSGCs were identified by their characteristic spike response to moving stimuli. Signals were acquired with the MultiClamp 700B amplifier and digitized at 100 kHz using the Digidata 1440A (Molecular devices). Spike activity was analyzed as described elsewhere (Trenholm et al. 2011). Neurons were considered direction selective when the direction selectivity index, calculated as a vector sum of the spike responses, was greater than 0.1 (Sethuramanujam et al. 2016). While the direction selectivity of the neurons was tested, the preferred direction of the neurons relative to the cardinal axes was not systematically tracked.
2.3. Imaging Dendritic Morphology
Once identified, DSGC dendritic morphologies were visualized by filling the cells with 100 µM Alexa-488 (Thermo Fisher). The dye’s fluorescence was imaged using two-photon excitation delivered using an Insight DeepSee+ laser (Spectra-Physics) tuned to 920 nm, guided by X/Y galvanometer mirrors (Cambridge Technology), and collected using photomultiplier tubes (Hama-matsu). The output voltage was digitized at 1 MHz (National Instruments) for image formation. A detailed description of the acquisition system is presented elsewhere (Jain et al. 2020). The image scans were acquired using custom software developed by Dr. Jamie Boyd (University of British Columbia) in the IgorPro environment. The X–Y pixel size of the acquired images ranged from 0.22 × 0.22 µm to 0.47 × 0.47 µm, and the resolution of the z-axis was 1 µm.
2.4. Morphological Analysis
The dendrites of ON and OFF arbors were traced manually using custom-written software in IgorPro (Sethuramanujam et al. 2021). ON and OFF arbors could be easily separated by the fluorescence intensity profile of filled dendrites along the IPL depth. Since the ON and OFF arbors of the DSGC stratify in narrow bands in the IPL, the intensity profile along the depth of the IPL can be visualized as two peaks (Figure 1C, middle panel). Along this profile, the minima—that is, the point that had the minimum fluorescence between the two peaks—was used to divide the ON and OFF arbors. The dendrites between the soma and the minima were considered ON dendrites, and dendrites beyond the minima were considered OFF dendrites. The average of the stack of images in each arbor was used for tracing dendrites. The total dendritic length was measured as the sum of the length of all the dendritic segments in each arbor. The dendritic area of each arbor was measured by calculating the area of a polygon connecting the endpoints of the dendrites. Sholl analysis was performed by constructing concentric rings with incremental radii of 10 µm centered at the soma. The number of intersections of the dendrites with each ring was considered for further analysis. Tortuosity was calculated as ΣLiTi/ΣLi, where Li is the length of individual dendritic segments, and Ti is the ratio of Li and the Euclidean distance between the segment’s endpoints. Finally, branch points were determined by manually counting the nodes in the arbor. All the above analyses were performed using custom code in IgorPro.
2.5. SBEM Analysis
An existing SBEM dataset was used for neuron reconstructions and synapse identification (Ding et al. 2016). The dimensions of the imaged volume were 65.9 × 211.2 × 262.9 µm3, and the voxel volume was 12 × 12 × 26 nm3. The volume consists of the IPL and the ganglion cell layer along with a fraction of the inner nuclear layer. Potential DSGCs and their synaptic partners were identified as described previously (Sethuramanujam et al. 2021). Potential ON–OFF DSGCs were first identified as ganglion cells with bi-stratified morphology in the IPL. Amacrine cell synapses to DSGC dendrites were identified by an electron-dense membrane thickening in presynaptic terminals filled with vesicles. A large fraction of these could be identified as starburst-mediated synapses based on their characteristic wraparound connections to DSGC dendrites. Bipolar synapses onto DSGCs were identified by their characteristic dyadic structure, where vesicles were concentrated on the presynaptic membrane at the junction abutting two postsynaptic neurons. The online platform “https://webknossos.org/” was used to trace the neuronal dendrites and annotate the synapses. The traces and annotations were imported in IgorPro for making figures.
2.6. Statistical Testing
Population data have been expressed as mean ± SEM and are indicated in the figure legends along with the number of samples. The Student’s t-test was used to compare values between ON and OFF arbors. In the case of Sholl analysis, the ON and OFF cumulative distributions were tested for significance using a Kolmogorov–Smirnov test. Differences were considered statistically significant when p ≤ 0.05.
3. Results
3.1. OFF Arbors Are Denser Than ON Arbors
We began by examining whether the ON and OFF dendritic arbors revealed gross morphological differences in ON–OFF DSGCs. Toward this endeavor, ON–OFF DSGCs were first identified in an isolated wholemount retina by their characteristic spike responses to the polarity of light stimuli and the direction of stimulus motion (Figure 1B). Once identified, these cells were injected with a fluorescent dye, and their dendritic tree was visualized using two-photon microscopy (see Methods) (Figure 1C–E, Top panels). Finally, the ON and OFF dendritic arbors were separated based on their distinct stratification pattern in the IPL and were manually traced for further analysis (Figure 1D,E) (see Methods). Using this method, we were able to image and reconstruct the ON and OFF dendrites of 44 cells (see Supporting Information), a subset of which are shown in Figure 2.
Figure 2. Reconstruction of ON–OFF DSGC dendrites.
(A) The dendritic trees of six ON–OFF DSGCs are shown here. ON dendrites are labeled in red; OFF dendrites are labeled in blue. The soma is represented by the shaded circle. Note that the thickness of the dendrites is not drawn to scale. (B) The ON dendrites of the corresponding cells shown in A. (C) The OFF dendrites of the corresponding cells shown in A and B. Scale bar represents 50 µm. The reconstructions of all the cells are shown in Supporting Information.
To test for morphological differences, we first evaluated the dendritic field area of the ON and OFF arbors. The dendritic field was estimated as the area of a polygon linking the terminal ends of the dendrites. This approach revealed that the area of the ON arbor was 23107 ± 1566 µm2, while the OFF arbor had a dendritic field of 24782 ± 1489 µm2 (n = 44 DSGCs; Figure 3A). These values are comparable to previous studies employing similar methods to measure dendritic field size (Sun et al. 2002; Rivlin-Etzion et al. 2011; Trenholm et al. 2011). On average, the dendritic field of the OFF arbor was slightly larger than the ON arbor by 7% (n = 44 cells; p = 0.015; paired t-test; Figure 3A).
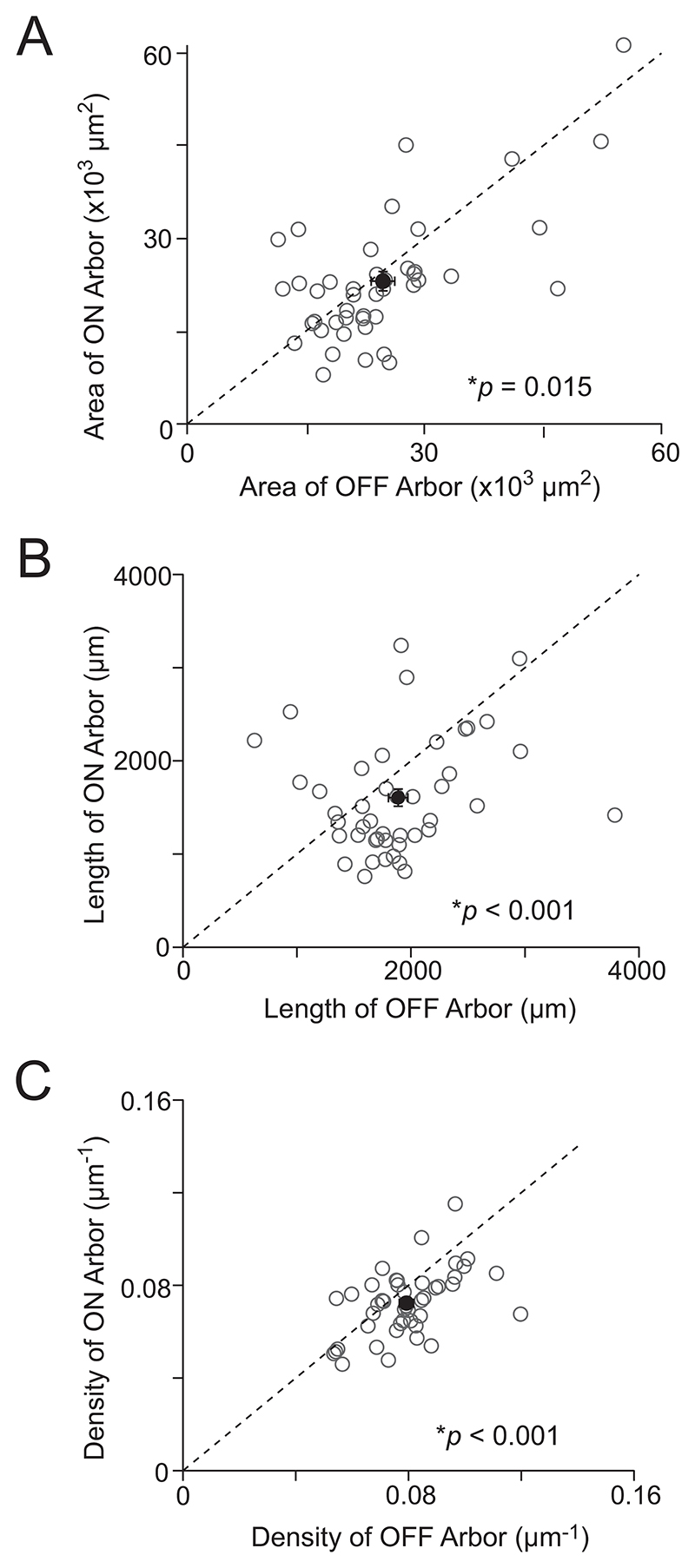
Figure 3. OFF arbor is denser in ON–OFF DSGCs.
(A) Scatter plots comparing the dendritic area of the ON and OFF arbors in DSGCs (n = 44). Open circles indicate individual cells, while the solid circle indicates the population mean. Data represented as mean ± SEM. (B, C) Similar to A, but for dendritic length and dendritic density. *indicates statistical significance when p is less than or equal to 0.05.
Next, we evaluated whether the arbors revealed any differences in the wiring density (dendritic length per unit area), as it has been suggested that the dendritic density may be a better indicator of function compared to the area (Bae et al. 2018). For this purpose, we first calculated the total length of dendrites in ON and OFF arbors. Our analysis revealed that the total dendritic length of the ON arbor was 1605 ± 92 µm, while the OFF arbor had a total dendritic length of 1888 ± 85 µm (n = 44 cells). Thus, the OFF arbor had 18% more wiring than the ON arbor of the same DSGC (p < 0.001; paired t-test; Figure 3B). Dividing the total wiring length by the dendritic area revealed that the OFF dendrites exhibited a 10% higher wiring density than the ON dendrites (0.0793 ± 0.002 µm−1 and 0.0723 ± 0.002 µm−1 for OFF and ON respectively; n = 44 cells; p < 0.001; paired t-test; Figure 3C). Hence, our analysis reveals that the OFF arbor is slightly larger but substantially denser when compared to its ON counterpart in ON–OFF DSGCs.
3.2. Increase in OFF Arbor Density Correlates With the Branching Pattern
Next, we examined the possible changes in the dendritic architecture that could lead to the observed increase in the density of the OFF arbor. We evaluated two mechanisms that would allow the OFF arbor to accommodate more dendrites per unit area: (a) tortuosity and (b) branching pattern.
Tortuosity is a measure of the curvature/bendiness of the dendrites. This was calculated as the ratio of the total length of the dendrite to the shortest distance between the endpoints. A tortuosity of 1 implies a perfectly straight dendrite, whereas a number closer to 1.57 suggests that the dendrite is arched like a semicircle. In our analysis, the tortuosity of the arbor was estimated as a weighted average of the tortuosity of individual dendritic segments (see Methods), where a segment was defined as the length of the dendrite enclosed by either a branch point or the dendritic tip. This allowed us to dissociate the actual curvature of dendrites from the curvature induced by branching per se. The calculated tortuosity of the dendrites in the ON arbor was 1.154 ± 0.011, and that of the OFF arbor was 1.154 ± 0.008 (n = 44 cells; Figure 4A). For comparison, the tortuosity of the dendritic arbor in ON starburst amacrine cells was 1.101 ± 0.007 (n = 5 cells). Hence, the dendrites in DSGCs were inherently more curved (p < 0.001, unpaired t-test) compared to starbursts, concurring with many previous studies which termed it as “loopy” (Oyster et al. 1993; Sun et al. 2002; Rivlin-Etzion et al. 2011). But, importantly, the tortuosity of dendrites in ON and OFF arbors was not significantly different from each other (n = 44 cells; p = 0.818; paired t-test).
Figure 4. OFF arbor has higher branching.
(A) Scatter plots comparing the tortuosity of the ON and OFF arbors in DSGCs. Open circles indicate individual cells, while the solid circle indicates the population mean (n = 44). Data represented as mean ± SEM. (B) Similar to A, but for the number of branch points. *indicates statistical significance when p is less than or equal to 0.05.
Next, we investigated whether the branching pattern could explain the increased dendritic density. Toward this endeavor, we manually identified the number of branch points/nodes in the ON and OFF dendritic trees. We excluded branch points if one of the two offshoots was less than 2 microns in length. This approach revealed that the ON arbor had an average of 33 ± 1.9 branch points, while the OFF arbor had 38 ± 1.5 (n = 44 cells; p < 0.001; paired t-test; Figure 4B). Hence, the OFF arbors had 15% more branch points than the ON arbors. These observations suggest that the increased density in OFF arbors is due to the branching pattern and not due to the tortuosity of the dendrites.
3.3. ON and OFF Arbors Have Similar Substructures
The higher branching pattern observed in OFF dendrites could be uniformly distributed across the arbor or localized to distinct substructures. Given that these spatial patterns have distinct implications for neuronal function (see Discussion), we investigated the patterns using Sholl analysis (Figure 5). In this analysis, the number of dendrites intersecting with concentric rings centered at the soma was measured (Figure 5A). We hypothesized that an increase in branching at specific subregions of the dendritic field would be reflected in the distribution of dendritic intersections around the soma. Our analysis revealed that the number of intersections was higher in the OFF arbors compared to the ON arbors; however, in the case of both arbors, the number of intersections peaked at 50–60 µm (Figure 5B). This conformed with the observation that the OFF arbor had more dendrites, but importantly, it also suggested that the spatial pattern of branching was similar. To probe this further, we compared the cumulative distribution function of the intersections with increasing distance. As expected, there were more intersections in the OFF dendritic field compared to the ON dendritic field (Figure 5C). Yet, the normalized cumulative distribution functions were not significantly different from each other (Kolmogorov–Smirnov test, p = 0.77) (Figure 5D). This suggested that the basic dendritic pattern in ON and OFF arbors was similar, and the density increase in OFF arbors could be accounted for by a scaling factor. Hence, our analysis revealed that the increased branching in the OFF arbor was uniformly distributed in space and not localized to distinct subregions.
Figure 5. ON and OFF arbors have similar substructure.
(A) Illustration of Sholl analysis showing the intersections of dendrites of an OFF arbor (black) with a series of concentric circles (gray). The radius of the circles is 10 µm apart. (B) Distribution of the number of dendritic intersections of ON (black) and OFF (gray) arbors with increasing radial distance from the soma. At peak, the number of intersections in the OFF arbor was significantly greater than the ON arbor (n = 44, p < 0.01; paired t-test). Data represented as mean ± SEM. (C) Cumulative frequency distribution of the data shown in B. Data represented as mean ± SEM. (D) Similar to C, but the values are normalized. Data represented as mean ± SEM. *indicates statistical significance when p is less than or equal to 0.05.
3.4. OFF Dendrites Exhibit a Higher Density of Bipolar Synaptic Connections
Given that the OFF dendrites are denser, we proceeded to examine whether this directly engenders an increase in the synaptic density of the OFF arbors. To assess the synaptic density, we resorted to analyzing a publicly available SBEM dataset (Ding et al. 2016). In this dataset, we examined previously identified ON–OFF DSGCs (Sethuramanujam et al. 2021). Briefly, ON–OFF DSGCs were identified based on their bi-stratification in the IPL (Figure 6A) and their synaptic connections with starburst amacrine cells (Figure 6D). Here, we examined the synaptic connectivity onto the dendrites of three ON–OFF DSGCs.
Figure 6. Anatomical reconstruction of bipolar and starburst inputs on ON–OFF DSGC dendrites.
(A) The side-view of dendrites of two ON–OFF DSGCs (gray) reconstructed from a publicly available SBEM volume. Synaptic connections were identified in short 50 µm segments in the ON and OFF dendrites, indicated in green and pink, respectively. The open square indicates the soma. Two of the three DSGCs that were traced are shown here. (B) Top view of the same cells shown in A. (C) A 3D reconstruction of the synaptic connections onto a small 13-µm dendritic section of a DSGC (green) shown in A and B. A bipolar cell terminal is shown in blue, while two putative starburst terminals are shown in red. The black arrows indicate the synaptic sites. (D) An SBEM cross-section showing one of the starburst-DSGC synapses. The green color highlights the DSGC dendrite, and red indicates the presynaptic starburst terminal. The red arrow indicates the location of the synapse. (E) An SBEM cross-section showing a bipolar terminal (blue) forming a dyad with the DSGC dendrite (pink) and a second unidentified post-synaptic partner (yellow). The small red arrow shows the site of the ribbon synapse.
Given that this exercise aimed to evaluate the density of connections, we focused on identifying all the synaptic connections in dendritic segments about 50 microns in length. We selected six such dendritic segments in the ON and OFF arbors. To facilitate comparisons, we selected dendritic segments that were largely unbranched and perpendicular to the depth of the IPL (Figure 6A,B). Next, connections from inhibitory amacrine cells were identified by presynaptic terminals loaded with vesicles, which formed electron-dense structures abutting the DSGC’s dendrite (Figure 6D). Of these connections, starbursts could be identified as the presynaptic terminals that wrap around the DSGC dendrite, a structure termed as a “wraparound” (Figure 6C,D) (Briggman et al. 2011). A significant fraction of the amacrine cell synapses (9%) did not make wraparounds and were not considered for further analysis. Excitatory connections from bipolar cells were identified by the characteristic dyad formation, which shows vesicles concentrated at the junction of two postsynaptic dendrites (Figure 6E). It is important to note that DSGCs also receive cholinergic excitation from starburst amacrine cells. However, given the difficulty in identifying these connections in SBEM, we have not analyzed the cholinergic inputs (Sethuramanujam et al. 2021).
In this exercise, we annotated a grand total of 372 synapses over 623 microns of DSGC dendrites (ON and OFF combined). Of these 372 synapses, there were 280 starburst synapses and 66 bipolar synapses (Figure 7). Hence, there were about 4.5 starburst synapses/10 microns of DSGC dendrites, a number concurring with the high density of starburst connections predicted by previous studies (Sigal et al. 2015; Jain et al. 2020). Further, in the same set of dendrites, the density of excitatory bipolar connections was 1.1 synapses/10 microns. Consequently, inhibitory connections from starburst amacrine cells outnumber the bipolar connections by fourfold.
Figure 7. Density of synaptic inputs to ON and OFF dendrites of ON–OFF DSGCs.
(A) Skeletons of ON dendritic segments in ON–OFF DSGCs (each approximately 50 µm), with the location of the bipolar synapses shown in solid black circles. Six dendritic segments were reconstructed from three DSGCs (two segments each). Note that the same segments were analyzed for the bipolar and starburst synapses but were oriented differently to display the synaptic distribution clearly (D, E). (B) Same as A, but for the OFF dendrites. (C) A plot comparing the number of bipolar synapses made by ON and OFF dendrites. The solid black circle represents mean ± SEM (* indicates p = 0.004; unpaired t-tests), while individual dendrites are shown as hollow gray circles. (D–F) Similar to A–C, but for starburst synapses.
Finally, we investigated whether there were any differences in the starburst/bipolar connections made with ON and OFF dendrites of ON–OFF DSGCs. The ON dendrites of the DSGCs received 0.9 ± 0.2 bipolar synapses per 10 microns of dendrite, while the OFF dendritic segments received 1.3 ± 0.2 bipolar synapses (Figure 7A–C). Hence, the linear density of bipolar synapses to the OFF dendrites was 44% higher than the ON dendrites (n = 6 dendrites; p = 0.004; unpaired t-test; Figure 7C). In comparison, the DSGC’s ON dendrites received 4.7 ± 1.3 starburst synapses per 10 microns, and the OFF dendrites received 4.3 ± 0.8 starburst synapses (Figure 7D–F) (n = 6 dendrites; p = 0.506; unpaired t-test). Therefore, the density of inhibitory starburst connections did not reveal significant differences between the ON and OFF dendrites. Overall, these results show a clear and novel distinction in the pattern of synaptic inputs to the ON and OFF arbors of ON–OFF DSGCs.
4. Discussion
4.1. Denser Dendritic Architecture of OFF Arbors in ON–OFF DSGCs
The ON–OFF DSGCs have one of the better-characterized morphologies among retinal ganglion cells with their bi-stratified dendrites in the ON and OFF sublamina of the IPL (Figure 1A). In general, the ON–OFF DSGC’s dendrites are known for their relatively dense dendritic arborization and loopy nature, which crisscross between the two sublaminae in the IPL (Amthor et al. 1984, 1989; Oyster et al. 1993; Weng et al. 2005; Trenholm et al. 2011). The uniqueness of these features has allowed studies to distinguish DSGCs from other retinal ganglion cells (Sun et al. 2002; Rivlin-Etzion et al. 2011; Sethuramanujam et al. 2018).
While the general arborization in ON–OFF DSGCs is well characterized, it is unclear whether the ON and OFF arbors of these cells are distinct from each other. Classically, dendritic morphology has been studied using small sample sizes, and such studies have reported no significant differences between the ON and OFF arbors in both rabbit (Amthor et al. 1989; Oyster et al. 1993; Vaney 1994) and mouse retina (Rivlin-Etzion et al. 2011; Peng et al. 2017). Here, using a large sample size, we find that the OFF arbors have significantly higher dendritic area, length, and density compared to the corresponding ON arbors in murine ON–OFF DSGCs. It is possible that previous studies have focused on specific DSGC subpopulations, which do not exhibit differences in their ON and OFF arbors (Rivlin-Etzion et al. 2011; Peng et al. 2017). Given that the preferred direction of the DSGCs in this study was not tracked relative to the cardinal axes, it is not possible to draw a comparison in this regard. Nevertheless, the experiments presented here reveal novel distinctions in the morphologies of the ON and OFF arbors in murine ON–OFF DSGCs. While previous studies have documented differences in the dendritic morphologies of ON and OFF retinal ganglion cells belonging to the same class (Wässle, Boycott 1981; Wässle, Peichl 1981; Dacey and Petersen 1992; Goetz et al. 2022), this is the first instance showing differences in the ON and OFF branching patterns within the same ganglion cell.
The SBEM analysis presented here reveals that the synaptic inputs to ON and OFF arbors are different. Specifically, the density of excitatory bipolar inputs was higher in the OFF arbor, while the inhibitory starburst inputs were not significantly different between the arbors (Figure 7). Combining this information with the total dendritic wiring indicates that OFF arbors, on average, have ∼250 bipolar synapses and ∼800 starburst synapses. In contrast, the average ON arbor has ∼150 bipolar synapses and ∼750 starburst synapses. This large difference in the excitatory synapses could be distributed uniformly across the dendritic field or localized to specific regions within the dendritic field, a feature seen in some ganglion cells (G. W. Schwartz et al. 2012). However, two pieces of evidence suggest uniform distribution. First, the increase in dendritic density was scalable throughout the dendritic field (Figure 5), indicating that the ON and OFF arbors did not differ in their substructure. Second, despite using dendritic segments at varied radial distances from the soma (ON: 36–90 µm; OFF: 28–105 µm), the linear synaptic density was largely similar (Figure 7C). Hence, the data presented here support the notion of a uniform increase in the density of OFF bipolar synapses across the dendritic field.
The distinct architecture of ON and OFF arbors in DSGCs is somewhat surprising, given that they both arise from the same soma. It is possible that distinct patterns in the ON and OFF arbors require specific molecular mechanisms (Peng et al. 2017) that determine the branching pattern. Alternately, it is also possible that a spatial constraint in the ON layer—as it is shared between 7 types of DSGCs (Vaney et al. 2012; Wei 2018)—compared to the OFF layer results in a sparser ON arbor. This could be evaluated in future studies using mutant mice, like satb1 (Peng et al. 2017), which exhibit altered ON and OFF dendritic arbors in these DSGCs.
4.2. Functional Implications of Differences in the Dendritic Architecture of ON and OFF Arbors
It is well established that dendritic architecture is inextricably linked to function. For instance, in the human retina, studies indicate that ON midget and parasol cells exhibit 30%–50% larger dendritic fields compared to the corresponding OFF cells (Dacey and Petersen 1992). Not surprisingly, these ON cells with larger dendritic fields had larger receptive fields (Chichilnisky and Kalmar 2002; Soto et al. 2020). Along with the larger receptive fields, the ON cells also exhibited a higher signal-to-noise ratio and increased sensitivity to stimulus contrast. Both signal-to-noise ratio and increased sensitivity can be explained if the larger dendritic fields directly translate to more synaptic inputs (Demb et al. 2004). Higher synaptic input will allow even weak stimuli to activate sufficient independent inputs to average out the noise in the presynaptic circuitry and, at the same time, push the DSGC’s membrane to cross the spiking threshold. Indeed, it has been suggested that denser but smaller dendritic trees are also effective in increasing the number of synapses and, consequently, neuronal sensitivity (Balasubramanian and Sterling 2009). Our observations show that, in murine ON–OFF DSGCs, the OFF arbors are larger and denser, packing 18% more dendrites than the average ON arbor. This is further compounded by the increased synaptic density, leading to a 67% increase in the OFF excitatory inputs. Hence, under weak stimulus conditions, when the synaptic inputs to DSGCs are sparse and noisy, the increased synaptic density may facilitate the OFF arbors to reliably encode directional information. This hypothesis can be tested by subsequent studies evaluating the contrast sensitivity of the ON and OFF responses in these DSGCs using single-cell patch clamp recordings. However, it must be noted that the spike output of DSGCs will not only depend on the dendritic architecture but also on the relative activation of the presynaptic ON/OFF circuitry, which is dynamically modulated by the visual conditions (Pearson and Kerschensteiner 2015; Rosa et al. 2016). In order to isolate the effects of dendritic architecture, future studies have to directly activate the ON and OFF dendritic arbors using techniques like optogenetics/uncaging (Vlasits et al. 2016).
4.3. Differential Processing of ON and OFF Signals Likely Enhances the Computational Range of DSGCs
In the past decade, several studies have revealed that the convergence of distinct circuits allows DSGCs to compute direction under conditions that otherwise would not have been possible. For instance, in ON DSGCs, convergence of bipolar excitation and distinct glycinergic inhibitory inputs facilitate velocity tuning while maintaining direction computations (Sivyer et al. 2019; Summers and Feller 2022; Mani et al. 2023). Further, a convergence of distinct transient and sustained bipolar circuits facilitates computations particularly at slow speeds (Matsumoto et al. 2019). In the ON–OFF DSGC, it has been shown that distinct excitatory motifs facilitate computations at different contrast regimes (Poleg-Polsky and Diamond 2016a; Sethuramanujam et al. 2016, 2017). These motifs also help in encoding direction at different speeds (Hanson et al. 2019) and spatial scales (Sethuramanujam et al. 2018). In all these cases, the critical features are the differences in the processing mechanisms of presynaptic neurons. Our observations add another layer of complexity, showing that the ON–OFF DSGC intrinsically processes information from converging circuits differently. Specifically, our observations on the dendritic architecture of ON–OFF DSGCs predict that the OFF signals have a higher synaptic gain, which may allow the DSGC to encode information under weak/noisy conditions. Hence, our observations are in line with the growing evidence showing that distinct circuits converging at the level of DSGCs process information in complementary stimulus regimes and, consequently, enhance the range over which DSGCs can reliably encode information.
Supplementary Material
Supporting Information
Additional supporting information can be found online in the Supporting Information section.
Acknowledgments
We thank Dr. Gautam Awatramani for graciously allowing Dr. Santhosh Sethuramanujam to perform imaging experiments in his laboratory at the University of Victoria, Canada. We thank Prof. Malcolm Slaughter and Dr. Gautam Awatramani for commenting on the manuscript.
Funding
This work was supported by the DBT/Wellcome Trust India Alliance Fellowship (grant number: IA/I/21/2/505933), and a New Faculty Seed Grant (IP/22-23/0847/BT/NFSC/008986) from IIT Madras, awarded to Dr. Santhosh Sethuramanujam, a CSIR-HRDG fellowship awarded to Sheba Annie Philip, and a UGC Junior Research Fellowship awarded to Priyanka Parida.
Footnotes
Author Contributions
S.A.P., S.S., and N.P.S. designed the research. S.S. performed the imaging experiments. S.A.P. performed the analysis in Figures 1–5. S.V. helped in preliminary analysis. P.P. helped in the analysis of Figure 4B. N.P.S. performed the analysis in Figures 6 and 7. S.S. and S.A.P. wrote the article.
Conflicts of Interest
The authors declare no conflicts of interest.
Peer Review
The peer review history for this article is available at https://publons.com/publon/10.1002/cne.70023.
Data Availability Statement
The data presented in this study will be provided upon request by the corresponding author.
References
- Amthor FR, Oyster CW, Takahashi ES. Morphology of ON-OFF Direction-Selective Ganglion Cells in the Rabbit Retina. Brain Research. 1984;298:187–190. doi: 10.1016/0006-8993(84)91167-3. [DOI] [PubMed] [Google Scholar]
- Amthor FR, Takahashi ES, Oyster CW. Morphologies of Rabbit Retinal Ganglion Cells With Complex Receptive Fields. Journal of Comparative Neurology. 1989;280:97–121. doi: 10.1002/cne.902800108. [DOI] [PubMed] [Google Scholar]
- Ariel M, Daw NW. Pharmacological Analysis of Directionally Sensitive Rabbit Retinal Ganglion Cells. Journal of Physiology. 1982;324:161–185. doi: 10.1113/jphysiol.1982.sp014105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baden T, Berens P, Franke K, Román Rosón M, Bethge M, Euler T. The Functional Diversity of Retinal Ganglion Cells in the Mouse. Nature. 2016;529:345–350. doi: 10.1038/nature16468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bae JA, Mu S, Kim JS, et al. Digital Museum of Retinal Ganglion Cells With Dense Anatomy and Physiology. Cell. 2018;173:1293.e19–1306.e19. doi: 10.1016/j.cell.2018.04.040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Balasubramanian V, Sterling P. Receptive Fields and Functional Architecture in the Retina. Journal of Physiology. 2009;587:2753–2767. doi: 10.1113/jphysiol.2009.170704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Briggman KL, Helmstaedter M, Denk W. Wiring Specificity in the Direction-Selectivity Circuit of the Retina. Nature. 2011;471:183–188. doi: 10.1038/nature09818. [DOI] [PubMed] [Google Scholar]
- Chen Q, Pei Z, Koren D, Wei W. Stimulus-Dependent Recruitment of Lateral Inhibition Underlies Retinal Direction Selectivity. Elife. 2016;5:e21053. doi: 10.7554/eLife.21053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chichilnisky EJ, Kalmar RS. Functional Asymmetries in ON and OFF Ganglion Cells of Primate Retina. Journal of Neuroscience. 2002;22:2737–2747. doi: 10.1523/JNEUROSCI.22-07-02737.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cohen ED, Miller RF. Quinoxalines Block the Mechanism of Directional Selectivity in Ganglion Cells of the Rabbit Retina. Proceedings of the National Academy of Sciences of the United States of America. 1995;92:1127–1131. doi: 10.1073/pnas.92.4.1127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dacey DM, Petersen MR. Dendritic Field Size and Morphology of Midget and Parasol Ganglion Cells of the Human Retina. Proceedings of the National Academy of Sciences of the United States of America. 1992;89:9666–9670. doi: 10.1073/pnas.89.20.9666. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Demb JB, Sterling P, Freed MA. How Retinal Ganglion Cells Prevent Synaptic Noise From Reaching the Spike Output. Journal of Neurophysiology. 2004;92:2510–2519. doi: 10.1152/jn.00108.2004. [DOI] [PubMed] [Google Scholar]
- Ding H, Smith RG, Poleg-Polsky A, Diamond JS, Briggman KL. Species-Specific Wiring for Direction Selectivity in the Mammalian Retina. Nature. 2016;535:105–110. doi: 10.1038/nature18609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Famiglietti EV. Dendritic Co-Stratification of ON and ON-OFF Directionally Selective Ganglion Cells With Starburst Amacrine Cells in Rabbit Retina. Journal of Comparative Neurology. 1992;324:322–335. doi: 10.1002/cne.903240303. [DOI] [PubMed] [Google Scholar]
- Goetz J, Jessen ZF, Jacobi A, et al. Unified Classification of Mouse Retinal Ganglion Cells Using Function, Morphology, and Gene Expression. Cell Reports. 2022;40:111040. doi: 10.1016/j.celrep.2022.111040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hanson L, Sethuramanujam S, deRosenroll G, Jain V, Awatramani GB. Retinal Direction Selectivity in the Absence of Asymmetric Starburst Amacrine Cell Responses. Elife. 2019;8:e42392. doi: 10.7554/eLife.42392. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harris SC, Dunn FA. Asymmetric Retinal Direction Tuning Predicts Optokinetic Eye Movements Across Stimulus Conditions. Elife. 2023;12:e81780. doi: 10.7554/eLife.81780. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hellmer CB, Hall LM, Bohl JM, Sharpe ZJ, Smith RG, Ichinose T. Cholinergic Feedback to Bipolar Cells Contributes to Motion Detection in the Mouse Retina. Cell Reports. 2021;37:110106. doi: 10.1016/j.celrep.2021.110106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang X, Rangel M, Briggman KL, Wei W. Neural Mechanisms of Contextual Modulation in the Retinal Direction Selective Circuit. Nature Communications. 2019;10:2431. doi: 10.1038/s41467-019-10268-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ishii T, Kaneda M. ON-Pathway-Dominant Glycinergic Regulation of Cholinergic Amacrine Cells in the Mouse Retina. Journal of Physiology. 2014;592:4235–4245. doi: 10.1113/jphysiol.2014.271148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacoby J, Schwartz GW. Three Small-Receptive-Field Ganglion Cells in the Mouse Retina Are Distinctly Tuned to Size, Speed, and Object Motion. Journal of Neuroscience. 2017;37:610–625. doi: 10.1523/JNEUROSCI.2804-16.2016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jain V, Hanson L, Sethuramanujam S, et al. Gain Control by Sparse, Ultra-Slow Glycinergic Synapses. Cell Reports. 2022;38:110410. doi: 10.1016/j.celrep.2022.110410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jain V, Murphy-Baum BL, deRosenroll G, et al. The Functional Organization of Excitation and Inhibition in the Dendrites of Mouse Direction-Selective Ganglion Cells. Elife. 2020;9:e52949. doi: 10.7554/eLife.52949. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim JS, Greene MJ, Zlateski A, et al. Space–Time Wiring Specificity Supports Direction Selectivity in the Retina. Nature. 2014;509:331–336. doi: 10.1038/nature13240. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kittila CA, Massey SC. Pharmacology of Directionally Selective Ganglion Cells in the Rabbit Retina. Journal of Neurophysiology. 1997;77:675–689. doi: 10.1152/jn.1997.77.2.675. [DOI] [PubMed] [Google Scholar]
- Mani A, Yang X, Zhao TA, Leyrer ML, Schreck D, Berson DM. A Circuit Suppressing Retinal Drive to the Optokinetic System During Fast Image Motion. Nature Communications. 2023;14:5142. doi: 10.1038/s41467-023-40527-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsumoto A, Briggman KL, Yonehara K. Spatiotemporally Asymmetric Excitation Supports Mammalian Retinal Motion Sensitivity. Current Biology. 2019;29:3277.e5–3288.e5. doi: 10.1016/j.cub.2019.08.048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nath A, Schwartz GW. Cardinal Orientation Selectivity Is Represented by Two Distinct Ganglion Cell Types in Mouse Retina. Journal of Neuroscience. 2016;36:3208–3221. doi: 10.1523/JNEUROSCI.4554-15.2016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oyster CW, Amthor FR, Takahashi ES. Dendritic Architecture of ON-OFF Direction-Selective Ganglion Cells in the Rabbit Retina. Vision Research. 1993;33:579–608. doi: 10.1016/0042-6989(93)90181-u. [DOI] [PubMed] [Google Scholar]
- Pandarinath C, Victor JD, Nirenberg S. Symmetry Breakdown in the ON and OFF Pathways of the Retina at Night: Functional Implications. Journal of Neuroscience. 2010;30:10006–10014. doi: 10.1523/JNEUROSCI.5616-09.2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pearson JT, Kerschensteiner D. Ambient Illumination Switches Contrast Preference of Specific Retinal Processing Streams. Journal of Neurophysiology. 2015;114:540–550. doi: 10.1152/jn.00360.2015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peng Y-R, Tran NM, Krishnaswamy A, Kostadinov D, Martersteck EM, Sanes JR. Satb1 Regulates Contactin 5 to Pattern Dendrites of a Mammalian Retinal Ganglion Cell. Neuron. 2017;95:869.e6–883.e6. doi: 10.1016/j.neuron.2017.07.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poleg-Polsky A, Diamond JS. NMDA Receptors Multiplicatively Scale Visual Signals and Enhance Directional Motion Discrimination in Retinal Ganglion Cells. Neuron. 2016a;89:1277–1290. doi: 10.1016/j.neuron.2016.02.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poleg-Polsky A, Diamond JS. Retinal Circuitry Balances Contrast Tuning of Excitation and Inhibition to Enable Reliable Computation of Direction Selectivity. Journal of Neuroscience. 2016b;36:5861–5876. doi: 10.1523/JNEUROSCI.4013-15.2016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rivlin-Etzion M, Zhou K, Wei W, et al. Transgenic Mice Reveal Unexpected Diversity of ON-OFF Direction-Selective Retinal Ganglion Cell Subtypes and Brain Structures Involved in Motion Processing. Journal of Neuroscience. 2011;31:8760–8769. doi: 10.1523/JNEUROSCI.0564-11.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rosa JM, Morrie RD, Baertsch HC, Feller MB. Contributions of Rod and Cone Pathways to Retinal Direction Selectivity Through Development. Journal of Neuroscience. 2016;36:9683–9695. doi: 10.1523/JNEUROSCI.3824-15.2016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sagdullaev BT, McCall MA, Lukasiewicz PD. Presynaptic Inhibition Modulates Spillover, Creating Distinct Dynamic Response Ranges of Sensory Output. Neuron. 2006;50:923–935. doi: 10.1016/j.neuron.2006.05.015. [DOI] [PubMed] [Google Scholar]
- Schwartz G. Retinal Computation. Academic Press; 2021. [Google Scholar]
- Schwartz GW, Okawa H, Dunn FA, et al. The Spatial Structure of a Nonlinear Receptive Field. Nature Neuroscience. 2012;15:1572–1580. doi: 10.1038/nn.3225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sethuramanujam S, Awatramani GB, Slaughter MM. Cholinergic Excitation Complements Glutamate in Coding Visual Information in Retinal Ganglion Cells. Journal of Physiology. 2018;596:3709–3724. doi: 10.1113/JP275073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sethuramanujam S, Matsumoto A, deRosenroll G, et al. Rapid Multi-Directed Cholinergic Transmission in the Central Nervous System. Nature Communications. 2021;12:1374. doi: 10.1038/s41467-021-21680-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sethuramanujam S, McLaughlin AJ, deRosenroll G, Hoggarth A, Schwab DJ, Awatramani GB. A Central Role for Mixed Acetylcholine/GABA Transmission in Direction Coding in the Retina. Neuron. 2016;90:1243–1256. doi: 10.1016/j.neuron.2016.04.041. [DOI] [PubMed] [Google Scholar]
- Sethuramanujam S, Yao X, deRosenroll G, Briggman KL, Field GD, Awatramani GB. ‘Silent’ NMDA Synapses Enhance Motion Sensitivity in a Mature Retinal Circuit. Neuron. 2017;96:1099.e3–1111.e3. doi: 10.1016/j.neuron.2017.09.058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sigal YM, Speer CM, Babcock HP, Zhuang X. Mapping Synaptic Input Fields of Neurons With Super-Resolution Imaging. Cell. 2015;163:493–505. doi: 10.1016/j.cell.2015.08.033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sivyer B, Tomlinson A, Taylor WR. Simulated Saccadic Stimuli Suppress ON-Type Direction-Selective Retinal Ganglion Cells via Glycinergic Inhibition. Journal of Neuroscience. 2019;39:4312–4322. doi: 10.1523/JNEUROSCI.3066-18.2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soto F, Hsiang J-C, Rajagopal R, et al. Efficient Coding by Midget and Parasol Ganglion Cells in the Human Retina. Neuron. 2020;107:656–666.:e5. doi: 10.1016/j.neuron.2020.05.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Summers MT, Feller MB. Distinct Inhibitory Pathways Control Velocity and Directional Tuning in the Mouse Retina. Current Biology. 2022;32:2130.e3–2143.e3. doi: 10.1016/j.cub.2022.03.054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun W, Li N, He S. Large-Scale Morphological Survey of Mouse Retinal Ganglion Cells. Journal of Comparative Neurology. 2002;451:115–126. doi: 10.1002/cne.10323. [DOI] [PubMed] [Google Scholar]
- Trenholm S, Johnson K, Li X, Smith RG, Awatramani GB. Parallel Mechanisms Encode Direction in the Retina. Neuron. 2011;71:683–694. doi: 10.1016/j.neuron.2011.06.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Turner MH, Rieke F. Synaptic Rectification Controls Nonlinear Spatial Integration of Natural Visual Inputs. Neuron. 2016;90:1257–1271. doi: 10.1016/j.neuron.2016.05.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vaney DI. Territorial Organization of Direction-Selective Ganglion Cells in Rabbit Retina. Journal of Neuroscience. 1994;14:6301–6316. doi: 10.1523/JNEUROSCI.14-11-06301.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vaney DI, Sivyer B, Taylor WR. Direction Selectivity in the Retina: Symmetry and Asymmetry in Structure and Function. Nature Reviews Neuroscience. 2012;13:194–208. doi: 10.1038/nrn3165. [DOI] [PubMed] [Google Scholar]
- Vlasits AL, Morrie RD, Tran-Van-Minh A, et al. A Role for Synaptic Input Distribution in a Dendritic Computation of Motion Direction in the Retina. Neuron. 2016;89:1317–1330. doi: 10.1016/j.neuron.2016.02.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wässle H, Boycott BB, Illing RB. Morphology and Mosaic of ON- and OFF-Beta Cells in the Cat Retina and some Functional Considerations. Proceedings of the Royal Society of London Series B: Biological Sciences. 1981;212:177–195. doi: 10.1098/rspb.1981.0033. [DOI] [PubMed] [Google Scholar]
- Wässle H, Peichl L, Boycott BB. Morphology and Topography of ON- and OFF-Alpha Cells in the Cat Retina. Proceedings of the Royal Society of London Series B: Biological Sciences. 1981;212:157–175. doi: 10.1098/rspb.1981.0032. [DOI] [PubMed] [Google Scholar]
- Wei W. Neural Mechanisms of Motion Processing in the Mammalian Retina. Annual Review of Vision Science. 2018;4:165–192. doi: 10.1146/annurev-vision-091517-034048. [DOI] [PubMed] [Google Scholar]
- Weng S, Sun W, He S. Identification of ON–OFF Direction-Selective Ganglion Cells in the Mouse Retina. Journal of Physiology. 2005;562:915–923. doi: 10.1113/jphysiol.2004.076695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang G, Masland RH. Receptive Fields and Dendritic Structure of Directionally Selective Retinal Ganglion Cells. Journal of Neuroscience. 1994;14:5267–5280. doi: 10.1523/JNEUROSCI.14-09-05267.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zaghloul KA, Boahen K, Demb JB. Different Circuits for ON and OFF Retinal Ganglion Cells Cause Different Contrast Sensitivities. Journal of Neuroscience. 2003;23:2645–2654. doi: 10.1523/JNEUROSCI.23-07-02645.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang J, Diamond JS. Subunit- and Pathway-Specific Localization of NMDA Receptors and Scaffolding Proteins at Ganglion Cell Synapses in Rat Retina. Journal of Neuroscience. 2009;29:4274–4286. doi: 10.1523/JNEUROSCI.5602-08.2009. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The data presented in this study will be provided upon request by the corresponding author.