Abstract
Coronavirus disease 2019 (COVID‐19), caused by the novel severe acute respiratory syndrome coronavirus 2 (SARS‐CoV‐2), has been a rising international cause of morbidity and mortality. Angiotensin‐converting enzyme 2 (ACE2) is identified as a key cell entry receptor for SARS‐CoV‐2 and suggested to be a limiting factor for viral entry at the initial infection stage. Recent studies have demonstrated that ACE2 expression is highly enriched in nasal epithelial cells and type II alveolar epithelial cells, highlighting the importance of respiratory tract as the primary target site of SARS‐CoV‐2. The expression of ACE2 in airway epithelial cells is tightly regulated by inflammatory milieu and environmental and internal stimuli. Very recently, ACE2 has been reported to have different expression levels in airways under distinct chronic inflammatory airway diseases, such as chronic obstructive pulmonary disease (COPD) and allergic asthma, which may associate with the COVID‐19 risk and affect the management of primary airway diseases. In this review, we focus on the cutting‐edge progress in distribution, expression, and regulation of ACE2 in respiratory system in physiological and pathological conditions, and their implication for the development of COVID‐19. We also discuss the management of airway diseases, including asthma, COPD, allergic rhinitis, and rhinosinusitis in the era of COVID‐19.
Keywords: airway disease, angiotensin‐converting enzyme 2, coronavirus disease 2019, management, severe acute respiratory syndrome coronavirus 2
1. INTRODUCTION
The current outbreak of coronavirus disease 2019 (COVID‐19), which was first reported in December 2019, spreads across the globe rapidly and has been declared as pandemic on 11 March 2020 by the World Health Organization. 1 , 2 , 3 , 4 COVID‐19 is caused by a novel severe acute respiratory syndrome coronavirus 2 (SARS‐CoV‐2) and associates with substantial morbidity and mortality. 2 , 3 , 4 , 5 , 6 , 7 , 8 As of 19 July 2020, more than 14.0 million cases have been confirmed and over 600,000 deaths. The clinical manifestations of COVID‐19 are heterogeneous, ranging from asymptomatic to severe disease and even death. 9 , 10 Currently, no specific antiviral drug proved by randomized controlled trials or vaccine is available for the prevention and treatment of COVID‐19. 8 , 10 , 11 A key challenge for clinicians is to identify risk and protective factors for COVID‐19 to improve outcomes. A number of epidemiological studies have identified older age, male gender, black and minority ethnicity, and various comorbidities including hypertension, diabetes, cardiovascular disease, and chronic obstructive pulmonary disease (COPD) as the risk factors for COVID‐19 (Table 1), associated with severe illness and death. 12 , 13 , 14 , 15 , 16 , 17 , 18 , 19 , 20 , 21 , 22 , 23 , 24 , 25 , 26 , 27 , 28 , 29 , 30 However, the mechanisms underpinning the contribution of these factors for the development of COVID‐19 are not fully understood, which may be associated with the capacity of SARS‐CoV‐2 to invade host cells and infect an individual person under different conditions.
TABLE 1.
Risk‐associated diseases or factors for COVID‐19
| Comorbidities | Hazard or odds ratios (95% confidence intervals) for infection or poor prognosis risk |
|---|---|
| Hypertension |
For composite end‐points (admission to intensive care unit, invasive ventilation, or death): 1.58 (1.07‐2.32) 13 ; For death in patients with severe illness: 1.77 (1.11‐2.84) 16 |
| Cardiovascular diseases | For death: 1.17 (1.12‐1.22) for patients with chronic heart disease 12 |
| Diabetes |
For death: 1.95 (1.83‐2.07) for patients with HbA1c ≥ 58 mmol/mol; 12 For composite end‐points (admission to intensive care unit, invasive ventilation, or death): 1.59 (1.03‐2.45) 13 |
| COPD |
For composite end‐points (admission to intensive care unit, invasive ventilation, or death): 2.68 (1.42‐5.06) 13 ; For severe illness: 5.69 (2.49‐13.00). 20 |
| Smoking |
For death: 1.19 (1.14‐1.24) for former smokers; 12 For death in critically ill patients: 5.21 (1.39‐19.52) 14 |
| Asthma |
For death: 1.13 (1.01‐1.26) for severe asthmatic patients with recent OCS use 12 ; For severe illness: 1.48 (1.15‐1.92) for non‐allergic asthmatic patients 27 |
| Malignancy |
For composite end‐points (admission to intensive care unit, invasive ventilation, or death): 3.50 (1.60‐7.64); 13 For death: 2.98 (1.76‐5.06) 15 |
| Obesity |
For severe illness: 3.40 (1.40‐2.86) for patients with BMI ≥ 28.0 22 For death: 1.40 (1.30‐1.52) for patients with BMI from 35.0 to 39.9 and 1.92 (1.72‐2.13) for patients with BMI ≥ 40.0 12 |
| Other factors | |
| Male |
For death: 1.59 (1.53‐1.65) for males 12 ; For death: 0.46 (0.35‐0.60) for females 15 ; For death in patients with severe illness: 1.72 (1.05‐2.82) for males. 16 |
| Advanced age |
For death: 2.40 (2.16‐2.66) for patients aged from 60 to 69 y, 6.08 (5.52‐6.69) for patients aged from 70 to 79 y, and 20.61 (18.72‐22.70) for those aged ≥80 y 12 ; For death: 5.06 (2.82‐9.06) for patients aged from 50 to 64 y, 9.97 (5.64‐17.63) for patients aged from 65 to 79 y, and 20.33 (10.76‐38.44) for those aged ≥80 y 15 ; For death in patients with severe illness: 1.72 (1.09‐2.73) for patients aged ≥65 y 16 |
| Blood group A |
For infection: 1.54 (1.12‐2.10) for subjects with blood group A 30 ; For infection: 0.65 (0.46‐0.93) for subjects with blood group O 30 |
| Black and minority ethnicity |
For death: 1.48 (1.30‐1.69) for Black patients 12 ; For death: 1.44 (1.32‐1.58) for South Asian patients 12 |
| Air pollution |
Incidence of COVID‐19 cases positively correlated with poor air quality (NO2, PM2.5, and PM10 levels) 25 ; Increase of 1 μg/m3 in PM2.5 associated with 8% increase in the COVID‐19 death rate 26 (preprint) |
Abbreviations: BMI, body mass index; COPD: chronic obstructive pulmonary disease; HbA1c, haemoglobin A1c; OCS, oral corticosteroids.
Severe acute respiratory syndrome coronavirus 2 belongs to the broad family of viruses known as coronaviruses and shares 79.6% sequence identity to SARS‐CoV, the virus accounting for SARS pandemic in 2003. 31 , 32 , 33 Similar to SARS‐CoV, SARS‐CoV‐2 has been discovered to enter cells via binding to angiotensin‐converting enzyme 2 (ACE2). 32 , 34 , 35 , 36 The SARS‐CoV‐2 spike protein demonstrates at least 10 times higher affinity in binding ACE2 than does SARS‐CoV. 37 High ACE2 expression on host cells increases the susceptibility to SARS‐CoV‐2, and blocking ACE2 signalling prevents the viral infection in vitro. 32 , 36 , 38 In humans, ACE2 expression has been found to be increased in the lung of smokers and patients with COPD, and those patients have been reported more likely to have severe COVID‐19, implicating a central role of ACE2 in COVID‐19 development. 13 , 19 , 39
Respiratory tract is continuously exposed to a multitude of pollutants and inhaled pathogens, making them a primary target of SARS‐CoV‐2. It is supported by the recent discovery of enriched ACE2 expression in nasal epithelial cells and type II alveolar (AT2) cells. 40 , 41 , 42 Airway diseases, including allergic rhinitis (AR), sinusitis, asthma, and COPD, are highly prevalent world‐wide, affecting up to 50% of the populations. 43 , 44 , 45 The current COVID‐19 pandemic makes the interaction between primary inflammatory airway diseases and COVID‐19 critical for mitigating COVID‐19 risk and management of primary airway diseases. Do patients with inflammatory airway diseases have an altered susceptibility to COVID‐19? Will those patients have a unique disease course if infected by SARS‐CoV‐2? Will COVID‐19 affect the management of the primary airway diseases? Understanding the expression and function of ACE2 in airways in physiological and pathological conditions may give valuable hints to answer these questions. In this review, we summarized cutting‐edge advances in the very fast‐moving field of ACE2 study and highlighted the changes in ACE2 expression and its implication for COVID‐19 risk and disease management in patients with common upper and lower inflammatory airway diseases, including AR, rhinosinusitis, asthma, and COPD.
2. STRUCTURE AND FUNCTION OF ACE2
The 40 kb ACE2 gene located on chromosome Xp22 was first described in 2000. 46 , 47 The encoded human ACE2 protein is a type I transmembrane glycoprotein, which consists of 805 amino acids. 46 , 47 ACE2 orientates outside with the N‐terminus and anchors to the plasma membrane through a short intracellular C‐terminal tail. 46 , 47 , 48 The highly conserved catalytic site of ACE2 faces the extracellular space, where it can metabolize circulating peptides. 46 , 47 , 48 Despite 40% sequence identical of the active catalytic site between ACE and ACE2, they have distinct substrate specificity. 46 , 47 , 48 ACE2 functions as a carboxypeptidase, cleaving a single residue from angiotensin (Ang) II to generate Ang (1‐7) (Figure 1) and a single residue from Ang I to yield Ang (1‐9), whereas, as the peptidyl dipeptidase, ACE cleaves the decapeptide Ang I into an octapeptide Ang II. 49 Ang II is the key player in renin‐angiotensin system (RAS), a hormone system that maintains blood pressure homeostasis and fluid and salt balance. 50 Ang II mediates vasoconstriction and thus contributes to the overactivation of RAS, which is associated with a spectrum of diseases including hypertension, heart failure, and renal disease. 50 , 51 Therefore, ACE2 plays a crucial role in maintaining the balance of RAS by countering the activities of Ang II. Furthermore, ACE2 also can convert Ang A into alamandine, a protector in the cardiovascular system. 49 The membrane‐bound ACE2 can be cleaved by a disintegrin and metalloproteinase 17 to become the soluble form, which may function as a competitive interceptor of binding ligands of membrane ACE2. 52
FIGURE 1.
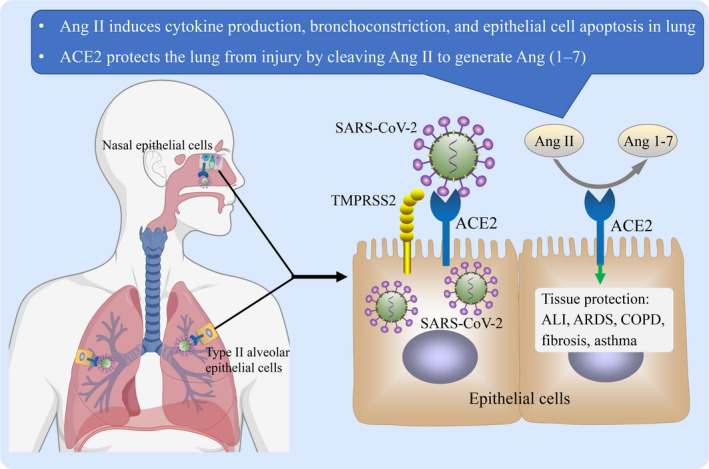
Expression and role of angiotensin‐converting enzyme 2 (ACE2) in airways. ACE2 is expressed in airways, with a particularly higher expression level in nasal epithelial cells and type II alveolar epithelial cells in the lung. Severe acute respiratory syndrome coronavirus 2 (SARS‐CoV‐2) binds to ACE2 expressed on human airway epithelial cells, and then, the serine protease TMPRSS2 cleaves and activates the spike protein of SARS‐CoV‐2, which ultimately facilities virus‐cell fusion and cell entry. As a peptidase, ACE2 catalyses and inactivates angiotensin (Ang) II and produces the vasodilator peptide Ang (1‐7). Ang II induces bronchoconstriction, vasoconstriction, fibroproliferation, cytokine expression, and cell apoptosis, thus promoting tissue injury. Accordingly, ACE2 has protective effects against tissue injury, including acute lung injury (ALI), acute respiratory distress syndrome (ARDS), fibrosis, chronic obstructive pulmonary disease (COPD), and allergic asthma
In lung, Ang II is able to induce bronchoconstriction, vasoconstriction, fibroproliferation, cytokine expression, and cell apoptosis, thus promoting tissue injury. 49 As a negative regulator of Ang II, ACE2 has been reported to protect the lung from injury (Figure 1). 49 In the acute lung injury/acute respiratory distress syndrome model, the lack of ACE2 expression in the lung resulted in exaggerated lung oedema, massive neutrophil accumulation, and deteriorated lung function. 53 Vice versa, treatment with recombinant ACE2 protein protected mice from severe acute lung injury. 53 Transforming growth factor (TGF)‐β1 is the most potent profibrotic cytokine and may act downstream of Ang II. 54 ACE2 overexpression significantly reduced TGF‐β1 levels and demonstrated a protective effect against the development of bleomycin‐induced fibrosis in murine models. 55 In the murine COPD model induced by cigarette smoking exposure, ACE2 overexpression in the lung significantly improved the lung function and pathological abnormalities. 56 In an ovalbumin‐challenged mouse model of asthma, Ang (1‐7) modulated ovalbumin‐induced increases in total cell counts, eosinophils, and lymphocytes, and decreased goblet cell hyper/metaplasia. 57 These results support an important role of Ang (1‐7) and ACE2 in reducing airway allergic inflammation (Figure 1). 57
Besides as a peptidase to catalyse Ang II cleavage in the physiological condition, ACE2 has been identified as a functional receptor for the SARS‐CoV and is important for the disease development after SARS‐CoV infection. 31 , 53 , 58 ACE2 purified from Vero E6 cells was able to bind the S1 domain of the SARS‐CoV spike protein in vitro. 31 ACE2 knockout mice showed reduced viral replication in the lungs in SARS‐CoV infection. 59 In contrast, transgenic mice expressing human ACE2 were highly susceptible to SARS‐CoV infection with more severe pulmonary lesions. 60 In addition, anti‐ACE2 antibody and soluble ACE2 treatment inhibited SARS‐CoV infection in mice. 58 Full‐length genome sequencing revealed that SARS‐CoV‐2 shares 79.6% sequence identity to SARS‐CoV. 32 SARS‐CoV‐2 has been found employing the same receptor as SARS‐CoV, ACE2, for host cell entry (Figure 1). 32 , 34 , 35 To invade the host cells, there is a need for another player, the transmembrane protease TMPRSS2, a cellular serine protease, which cleaves and activates the SARS‐CoV‐2 spike protein and facilitates human cell entry. 38 The spike protein of SARS‐CoV‐2 also contains a furin‐like cleavage site, 61 , 62 , 63 suggesting a potential role of furin and furin‐like proteases in SARS‐CoV‐2 cell entry. Wrapp et al described a 3.5‐angstrom‐resolution structure of the SARS‐CoV‐2 trimeric spike protein by cryo‐electron microscopy and found that SARS‐CoV‐2 spike protein binds to ACE2 at least 10 times more tightly than that of SARS‐CoV, 37 which may explain the higher transmission efficiency of SARS‐CoV‐2 than SARS‐CoV. Structurally, the receptor binding domain of the surface spike protein of SARS‐CoV‐2 is recognized by the extracellular peptidase domain of ACE2 mainly through the polar residues. 17 , 64
3. EXPRESSION OF ACE2 IN AIRWAYS
Airways consist of heterogeneous cell populations. The development of high‐throughput single‐cell RNA sequencing technique enables the robust profiling of the characteristics of specific cell types possible and offers an unbiased approach to study the expression of particular molecule in different cells. 65 , 66 By re‐analysing of deposited public available single‐cell RNA sequencing data sets, researches have conducted timely studies of the expression of ACE2 and other genes relevant to SARS‐CoV‐2 cell entry in airways.
3.1. Upper airways
Nasal cavity is the primary site of exposure to pollutants, airborne allergens, and inhaled pathogens and acts as the gateway to respiratory system. In patients with COVID‐19, nasal swabs have yielded higher viral loads than throat swabs, underscoring the nasal epithelium as a portal for initial infection and transmission of SARS‐CoV‐2. 32 Indeed, ACE2 and TMPRSS2 protein expressions have been detected in nasal epithelium by immunohistochemistry. 67 Recently, by analysing single‐cell RNA sequencing data of multiple tissues from healthy donors, Sungnak et al discovered an enriched ACE2 and TMPRSS2 mRNA expression in nasal epithelial cells, with the highest expression level observed in nasal secretory cells and ciliated cells (Figure 1). 40 Similarly, Ziegler et al observed significant enrichment of ACE2 mRNA expression in apical epithelial cells and, to a lesser extent, ciliated cells. 41 Consistently, the expression of ACE2 gene in goblet cells and ciliated cells was confirmed when analysing single‐cell RNA sequencing data derived from air‐liquid interface cultured nasal epithelial cells. 40 Notably, TMPRSS2 was found having a broader expression (28%) than ACE2 (4%) in nasal epithelial cells characterized by secretory phenotype, 41 suggesting that ACE2 may be the limiting factor for viral entry. In addition, TMPRSS2 is only expressed in a subset of ACE2‐positive cells. 40 Thus, SARS‐CoV‐2 may use alternative facilitating mechanism to entry host cells. A potential substitute for TMPRSS2 is cathepsin B, another protease, which was found to be expressed in more than 70% of ACE2‐positive cells. 40 ACE2 gene has been found to be coexpressed with genes involved in innate immunity in nasal epithelial cells, highlighting the important role of nasal epithelial cells in respiratory viral infection, spread and clearance. 40 , 41 In contrast, no significant ACE2 gene expression in immune cell populations, including T cells, dendritic cells, and mast cells, was discovered in nasal mucosal tissues. 40 , 41
Anosmia is typical of viral rhinitis, and smell and taste disorder has been reported as one of the common symptoms of COVID‐19 patients, 68 , 69 , 70 raising the possibility that SARS‐CoV‐2 may infect olfactory epithelium or sensory neurons. Indeed, high expression of ACE2 in epithelial cells of tongue has been reported based on the bulk and single‐cell RNA sequencing of oral tissues. 71 Expression of ACE2 and TMPRSS2 was also detected in human and murine olfactory epithelium. 72 , 73 , 74 Interestingly, by analysing a recently published single‐cell RNA sequencing data, Brann et al and Fodoulian et al found that sustentacular and olfactory stem cells, but not olfactory sensory neurons or neurons in olfactory bulb, expressed ACE2 and TMPRSS2 mRNA in humans (preprint). 73 , 74 In addition, enriched furin gene expression has been identified in olfactory epithelial gland cells, suggesting that other than TMPRSS2, furin may participate in helping SARS‐CoV‐2 cell entry. 41 , 73 The expression pattern of ACE2 protein in human olfactory epithelium was verified by immunohistochemical staining of biopsied tissues (preprint). 73 These data indicate that SARS‐CoV‐2 infection of support cells in olfactory epithelium and olfactory bulb may disrupt the structure of olfactory epithelium and impair the function of olfactory sensory neurons or bulb neurons, ultimately resulting the disturbance in smell perception in COVID‐19 patients.
3.2. Lower airways
The lung is the most important target organ of SARS‐CoV‐2 infection. An analysis of single‐cell RNA sequencing data derived from normal human lung tissues revealed that ACE2 gene was expressed by 0.64% of cells in lung and 83% of ACE2‐expressing cells were AT2 cells and 5% were type I alveolar cells (AT1) (Figure 1). 75 Nevertheless, bronchial airway epithelial cells, fibroblasts, endothelial cells, and macrophages had low expression of ACE2. 75 Consistently, the expression of ACE2 in AT2, AT1, and ciliated cells was revealed by other studies in single‐cell level. 40 , 41 , 76 , 77 , 78 , 79 In line with the single‐cell RNA sequencing finding, by immunohistochemistry study, Bezara et al found that about 1% of surfactant protein C–positive AT2 cells were positive for ACE2 protein expression, whereas alveolar macrophages were negative for ACE2 staining (preprint). 80 Through investigating ACE2 protein expression in different anatomical regions, they found that apical ACE2 expression was rare and limited to ciliated cells in the trachea and bronchi (preprint). 80 In the submucosal glands of large airways, occasional serous cells and vessels near the acini were positive for ACE2, but ACE2 was regionally localized in ciliated cells in bronchioles (preprint). 80 Notably, regional distribution of ACE2 protein varied across different donors (preprint), 80 which possibly contributes to the difference in host susceptible to SARS‐CoV‐2 and variable clinical presentations of COVID‐19. Similar to the finding in nasal epithelial cells, gene ontology enrichment analysis showed that the ACE2‐expressing AT2 cells had high levels of multiple viral response‐related genes, supporting the role of these cells in viral infection and control. 41 In addition, based on single‐cell RNA sequencing data, TMPRSS2 expression has been identified in various cell types of the lung, and the abundance was lower in club cells, ciliated epithelial cells, and AT1 cells than in AT2 cells. 40 , 41 Ziegler et al reported 34.2% of AT2 cells expressing TMPRSS2, 1.4% expressing ACE2, and 0.8% co‐expressing TMPRSS2 and ACE2. 41 Colocalization of ACE2 and TMPRSS2 could be observed on the apical membrane of AT2 cells by immunofluorescence staining (preprint). 80 These findings again suggest that ACE2 is likely the limiting factor for cell entry of SARS‐CoV‐2 and apical ACE2 and TMPRSS2 colocalization facilitates the cell entry. 40 , 41
4. REGULATION OF ACE2 EXPRESSION AND ASSOCIATED FACTORS
Coronavirus disease 2019 outbreak stirs the great research interest in ACE2 expression regulation in airway epithelial cells. The transcriptional and functional characteristics of ACE2 may be shaped by environmental stimuli, inflammatory cytokines, and medical treatments in different disease states.
4.1. Interferon and viral infection
Antiviral interferons (IFNs) and subsequent IFN‐induced chemokines and cytokines produced by virus‐infected host cells are critical for effective antiviral response. 81 There are three types of IFNs: type I (IFN‐α and IFN‐β), type II (IFN‐γ), and type III (IFN‐λ), and each of them may have a non‐redundant role in host defence and immunopathology, particularly at epithelial barriers. 82 Recently, Ziegler et al identified that ACE2 represented an IFN‐stimulated gene in human airway epithelial cells. 41 They found that ACE2 gene expression was markedly up‐regulated by IFN‐α and to a lesser extent by IFN‐γ in primary human nasal basal epithelial cells (Figure 2). 41 They further found signal transducer and activator of transcription (STAT) 1, STAT3, interferon regulatory factor (IRF) 8, and IRF1 binding sites in the transcription start site of ACE2 in humans, supporting a role of IFNs in regulating ACE2 expression. 41 In vivo, Sajuthi et al found a positive correlation between ACE2 and the IFN response gene expression in nasal epithelial brushing cells collected from a large cohort of asthmatic and healthy children (preprint). 42 Upper airway viral infections are known to induce a robust IFN response. 83 In comparison with those from patients without viral infection, Ziegler et al found that ACE2 mRNA expression was up‐regulated in goblet cells and squamous cells in nasal wash samples from influenza virus–infected individuals. 41 Nevertheless, it should be noted that ACE2 mRNA expression was only slightly elevated in nasal basal epithelial cells following IFN‐α treatment in mouse. 41 Additionally, intranasal infection with murine gamma herpesvirus‐68 failed to change the ACE2 expression in viral‐transcript–positive cells or AT2 cells from the lungs of mice. 41 Therefore, a careful consideration of animal and cellular models will be required for assessing therapeutic interventions targeting the IFN system when studying ACE2‐associated biology.
FIGURE 2.
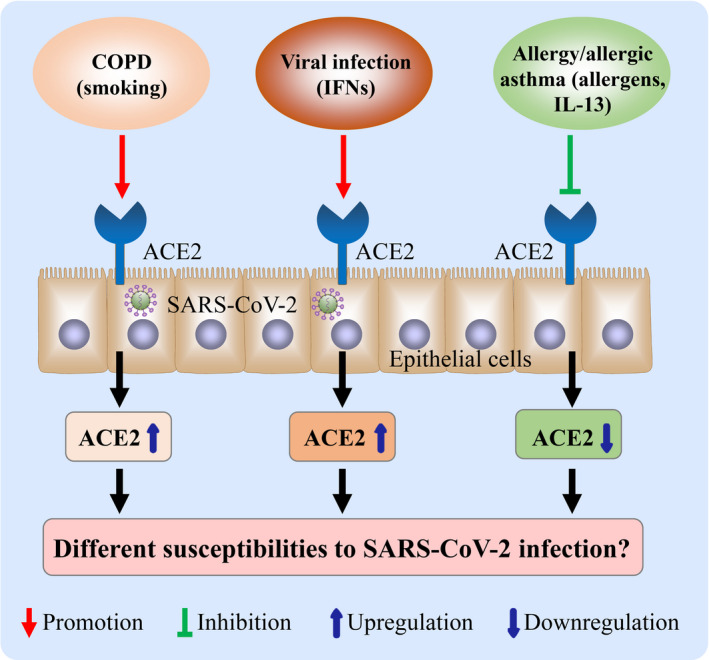
Regulation of angiotensin‐converting enzyme 2 (ACE2) expression in airway epithelial cells. Tobacco exposure promotes ACE2 expression in airway epithelial cells, which may associate with increased ACE2 expression in the lung of patients with chronic obstructive pulmonary disease (COPD). Interferon (IFN)‐α and IFN‐γ promote ACE2 expression in airway epithelial cells. Allergens and interleukin (IL)‐13 inhibit ACE2 expression in airway epithelial cells, may account for the lower ACE2 expression in patients with allergic asthma. Varied ACE2 in distinct chronic inflammatory airway diseases may contribute to different susceptibilities to severe acute respiratory syndrome coronavirus 2 (SARS‐CoV‐2) infection
4.2. Type 2 cytokine, allergen challenge, and eosinophil
Recently, several studies reported that IL‐13 down‐regulated ACE2 mRNA expression in human nasal and bronchial epithelial cells cultured with an air‐liquid interface method (Figure 2). 42 , 84 , 85 However, Ziegler et al showed that IL‐4 and IL‐13 had no effect on ACE2 mRNA expression in human nasal basal epithelial cells when cultured submerged. 41 It also should be noted that the data regarding the regulation of ACE2 expression at protein level are still lacking.
Allergen challenge can initiate and perpetuate airway type 2 (T2) inflammation. 86 Jackson et al have found that allergic sensitization was inversely related to ACE2 mRNA expression in nasal epithelium in asthmatic children. 84 In addition, significant negative correlations between ACE2 mRNA expression and T2 biomarkers including the number of positive allergen‐specific immunoglobulin (Ig)E tests, total IgE, fractional exhaled nitric oxide, and nasal epithelial IL‐13 expression were found by them, although most correlation coefficients were low. 84 Furthermore, Jackson et al found that nasal cat allergen led to a significant reduction in ACE2 mRNA expression in nasal brush samples in adult AR patients allergic to cat. 84 Consistently, segmental allergen bronchoprovocation to dust mite, ragweed, or cat significantly reduced ACE2 mRNA expression in bronchial brushing cells in adult patients with mild asthma. 84 Therefore, respiratory allergen exposures and T2 inflammation decrease ACE2 expression in both upper and lower airways (Figure 2).
Although eosinophilia is a hallmark of allergic respiratory inflammation, 87 accumulating evidence indicates an antiviral activity of eosinophils. 87 Eosinopenia has been noted in patients with COVID‐19. 7 , 88 Patients with fewer peripheral blood eosinophils displayed worse radiographic aggravation and longer course of disease compared to those with normal eosinophil counts. 88 Peripheral blood eosinophil levels have been reported to be gradually increased in recovered patients before discharge, accompanied by the improved clinical status. 88 These data suggest that peripheral blood eosinophil count may be an indicator for diagnosis, clinical status monitor, and outcome prediction of COVID‐19. Furthermore, Camiolo et al observed an increased ACE2 gene expression in bronchial epithelial cells in asthmatic patients with low peripheral blood eosinophil counts compared with those with high eosinophil counts, 89 suggesting a negative effect of eosinophil on ACE2 expression.
Allergy is an immune response to allergen stimulation that is characterized by elevated T2 cytokines and eosinophilic inflammation. These above findings raise the possibility that allergy might be a protective factor for COVID‐19. In a single‐centre retrospective study with a small sample size (110 COVID‐19 patients) in China, Shi et al observed a lower rate of comorbid allergy in patients with COVID‐19 in comparison with the prevalence of allergic diseases in the general population. 90 When excluding patients with other underlying diseases and stratifying COVID‐19 patients into those with (n = 21) and without allergy history (n = 44), they found that patients with allergy demonstrated lower proportions of bilateral lung lesions on chest computed tomography scanning and severe illness and higher circulating total T cell counts than those without allergy. 90 However, the relationship between allergy and COVID‐19 obviously needs to be evaluated and validated in larger cohorts and those with different genetic and socio‐economic backgrounds.
4.3. Smoking
Several epidemiological studies reported that cigarette smoking exposure associated with increased risk for COVID‐19 and current and former smokers was likely to develop severe COVID‐19 than never smokers. 3 , 13 , 91 , 92 By comprehensively analysing transcriptomic and microarray data sets, Cai et al found a markedly higher ACE2 mRNA expression in ever smokers compared with never smokers in both airway epithelial cells and lung tissues in COPD and non‐COPD subjects. 19 Brake et al observed an enhanced ACE2 protein expression in resected lung tissues from smokers with COPD and smokers with normal lung function when compared to heathy non‐smoking individuals, with more prominent increase in smokers with COPD compared with smokers with normal lung function. 93 Leung et al found that current smokers had a significantly higher ACE2 mRNA expression in bronchial bushing cells than never smoker. 39 In addition, smoking status defined by never, former, and current smokers was significantly associated with ACE2 gene expression in large airway epithelium and lung tissues. 19 Therefore, these consistent results strongly support that tobacco exposure significant up‐regulates ACE2 gene expression in airways (Figure 2). However, further studies are needed to explore the mechanisms underlying tobacco‐induced up‐regulation of ACE2 pulmonary expression. In addition, electronic cigarette consumption has significantly increased, whether electronic cigarettes have effects on ACE2 expression remains to be determined.
4.4. Glucocorticoid and biologics
Intranasal and inhaled corticosteroids serve as the cornerstone for the treatment of inflammatory airway disorders such as AR, chronic rhinosinusitis (CRS), COPD, and asthma. 45 , 94 Specific concerns have been raised in relation to steroid therapy and possible risk of adverse outcomes in COVID‐19. Whether baseline corticosteroid treatment has an effect on ACE2 expression in airways and whether it even modifies the risk to SARS‐CoV‐2 infection remain open questions. Jackson et al reported that nasal corticosteroid use at the time of nasal sampling was not associated with alteration in ACE2 mRNA expression in nasal epithelial cells in asthmatic children. 84 We have recently found that dexamethasone treatment did not change ACE2 mRNA expression in human nasal epithelial cells in vitro (under review). However, Peters et al recently reported that use of inhaled corticosteroids was associated with lower expression of ACE2 in sputum cells, which comprised a mix of multiple cell types, including structural airway cells (eg epithelial cells and squamous cells) and immune cells (eg macrophages and granulocytes). 95
Additional questions of concern relate to the effect of biologics on COVID‐19 risk. Omalizumab (an anti‐IgE antibody), mepolizumab and reslizumab (blocking IL‐5), benralizumab (blocking the IL‐5 receptor), and dupilumab (blocking IL‐4Rα) reduce asthma‐related exacerbation and have been approved for the treatment of severe asthma. 96 In addition, dupilumab has recently been approved for the treatment of CRS with nasal polyps in the United States and Europe. 97 Currently, there is no evidence‐based study addressing the effect of biologics on the risk of COVID‐19 and ACE2 expression. However, given the finding of down‐regulation of ACE2 gene expression in airway epithelial cells by T2 cytokines, it is possible that reversal of T2 skewed allergic and eosinophilic inflammation by these biologics may potentially increase the risk of SARS‐CoV‐2 infection by up‐regulating ACE2 expression in airways. Very recently, Förster‐Ruhrmann et al reported a case of 53‐years‐old female patient with refractory CRS with nasal polyps and treated with dupilumab, and this patient presented a light course of COVID‐19. 98 Obviously, more studies are needed to clarify the impact of biologics on COVID‐19 in patients with chronic inflammatory airway diseases.
4.5. Other associated factors
A number of studies demonstrated a lower risk of SARS‐CoV‐2 infection in children compared with adults. 99 Older age has been independently associated with increased mortality of COVID‐19 after adjusting for comorbidities (Table 1). 12 , 14 In a cohort of 305 individuals aged 4–60 years, Bunyavanich et al found that ACE2 gene expression levels in nasal epithelial cells were the lowest in younger children and increased with age. 100 This age‐dependent ACE2 gene expression may account for the different risks and mortality of SARS‐CoV‐2 infection in different age groups.
Several studies have observed an increased susceptibility and mortality of SARS‐CoV‐2 infection in males compared with females (Table 1), 2 , 12 , 13 , 15 suggesting a gender‐related expression of ACE2. Indeed, as an X chromosome–encoded gene, renal ACE2 expression has been reported to be significantly down‐regulated by oestrogens. 101 In addition to the direct effect on ACE2 expression, sex hormone may also modify the immune response to viral infection. For examples, plasmacytoid dendritic cells from adult females produced more IFN‐α than those from adult males in response to virus infection. 102 In addition, emerging evidence has implicated an association between ethnicity and incidence or outcome of COVID‐19. By analysing large data sets (17 278 392 adults) in the UK, Williamson et al found that, after adjustment for other factors, Black and South Asian people were at a higher risk of death compared with people with White ethnicity. 12 In addition, Black and minority ethnicity people have been found to be associated with increased SARS‐CoV‐2 infection and poor clinical outcome. 23 , 24 The causes for ethnic disparities for COVID‐19 development are unclear and may be related to different socio‐economic and cultural background, lifestyle, genetic predisposition, and ACE2 expression.
5. AIRWAY DISEASE AND COVID‐19
5.1. Upper airway disease
AR and CRS represented the two most common upper airway diseases. Previous studies reported low incidence of AR in COVID‐19 patients, ranging from 0% to 1.8% in China. 7 , 103 However, those results generated were relied on the medical records, and may underestimate the incidences of upper airway comorbidities since they are usually not well considered under actual emergency situation. 7 , 103 Recently, we retrospectively studied a cohort of 1172 discharged COVID‐19 patients by integrating patients' medical records and telephone questionnaire and found that the prevalence of AR in COVID‐19 patients was 9.8%, which was comparable to that in general populations in Wuhan, China (9.7%) (preprint). 104 , 105 Additionally, we did not find an association between AR comorbidity and disease severity in COVID‐19 patients, suggesting that comorbidity of AR may not modify the disease development and expression of COVID‐19 (under review). In contrast to allergic asthma, 84 there was no difference in ACE2 mRNA and protein expressions between AR patients and healthy controls (under review). As for CRS, reduced ACE2 gene expression was found in nasal epithelial cells form nasal polyp tissues compared with that in nasal epithelial cells from inferior turbinate tissues. 41 Nevertheless, mounting evidence demonstrates a significant heterogeneity in inflammatory patterns (endotypes) of CRS. 106 Type 1 (T1), T2, and type 17 (T3) endotype and mixed endotype have been reported for CRS. 106 , 107 It would be interesting to investigate whether ACE2 expression will vary across different CRS endotypes. Very recently, Chhiba et al reported that COVID‐19 patients with rhinosinusitis showed a lower risk of hospitalization than those without rhinosinusitis. 108 However, Chhiba et al did not differentiate between the acute and CRS in their study. In addition, whether distinct CRS endotypes relate to different clinical expressions of COVID‐19 also remains to be studied.
5.2. Lower airway disease
Although asthmatic patients are at risk of more severe outcome after common cold virus infection than people without asthma, 109 asthma has unexpectedly not been identified as a significant risk factor for severe COVID‐19. 5 , 7 , 110 Low prevalence of asthma (0%‐0.9%) was observed in patients with COVID‐19 in several studies in China. 13 , 16 , 91 , 111 In a US cohort including 1526 patients with COVID‐19, Chhiba et al recorded a relative high prevalence of asthma (14%); however, there was no significant difference in hospitalization rate or mortality between patients with and without asthma. 108 Notably, asthma is a heterogeneous disease with varying levels of severity and distinct endotypes. In a large cohort (17 278 392 adults) study in the UK, severe asthma (with oral steroid treatment) was identified as a risk factor associated with COVID‐19 death. 12 Allergic asthma is the most common type of asthma triggered by inhaled allergens. Jackson et al have recently reported reduced ACE2 gene expression in nasal and bronchial epithelial cells after allergen challenge in AR and allergic asthmatic patients (Figure 2). 84 In addition, Kimura et al found that ACE2 mRNA expression was reduced in bronchial epithelial cells in asthmatic patients with high T2 inflammation. 85 By analysing ACE2 gene expression in bronchial epithelial cells in asthmatic patients with different endotypes, Camiolo et al identified a positive correlation between ACE2 expression and scores of T1 gene expression and a negative correlation between ACE2 expression and scores of T2 gene expression. 89 These data suggest an association between asthma endotypes and COVID‐19, which was supported by the finding that T2‐low asthmatic patients demonstrated characteristics corresponding to risk factors for severe COVID‐19, including male sex and history of hypertension. 89 Taken together, although allergic asthma appears to be a protective factor for COVID‐19 (Figure 2), the associations between different phenotypic and endotypic asthma and COVID‐19 remain to be defined.
Among the airway inflammatory diseases, COPD is the most commonly reported comorbidity in COVID‐19 patients and the prevalence ranged from 1.5% to 5%. 13 , 91 , 112 Previous epidemiological studies have associated the COPD comorbidity with the severe illness and fatalities in COVID‐19 patients. 13 , 20 Guan et al reported that severe COVID‐19 cases had higher frequencies of COPD comorbidity than non‐severe cases. 13 Leung et al reported that ACE2 protein expression in the bronchial epithelial cells was significantly increased in COPD versus non‐COPD subjects. 39 Smoking is the primary aetiological factor for COPD, and evidence has shown increased ACE2 mRNA expression in smokers than in non‐smokers. 19 Interestingly, Leung et al found that ACE2 expression was still increased in bronchial epithelial cells in COPD than in non‐COPD subjects after adjusting the smoking status. 39 Therefore, it is likely that additional factors beyond smoking can modulate ACE2 expression in COPD patients.
6. IMPLICATION FOR TREATMENT
Current interim guidance from World Health Organization on clinical management of COVID‐19 advises against the administration of systemic corticosteroids. 113 For patients with primary chronic inflammatory airway diseases, several organizations suggest the patients with chronic airway diseases who are stable with inhaled or oral corticosteroid should continue their treatment. 113 , 114 , 115 , 116 , 117 The Allergic Rhinitis and its Impact of Asthma‐European Academy of Allergology and Clinical Immunology statement advises that for patients with COVID‐19 infection, intranasal corticosteroid treatment should be continued at the recommended dose for AR, since more sneezing after stopping means more spreading of the SARS‐CoV‐2. 114 The Global InNitiative for Asthma suggests that patients with asthma should not stop their prescribed inhaled corticosteroid during COVID‐19 pandemic, as stopping corticosteroids often leads to the potentially dangerous worsening of asthma. 115 Very recently, in a retrospective study of 1,526 COVID‐19 patients, Chhiba et al found that the ongoing use of intranasal corticosteroid was not associated with increased risk of COVID‐19–related hospitalization. 108 Both the American College of Allergy, Asthma & Immunology (ACAAI) and the American Academy of Allergy, Asthma & Immunology (AAAAI) also recommend patients with asthma to continue to use their maintenance medications even during the pandemic. 116 In addition, balancing the risk of losing disease control and the lack of evidence or expectation of increased infectivity or mortality, ACAAI and AAAAI recommend to continue administration of biologic agents. 116 The Global Initiative for Chronic Obstructive Lung Disease also advises that COPD patients should maintain their regular therapy, including corticosteroid administration, during the COVID‐19 pandemic. 117 Collectively, although glucocorticoids and biologics targeting T2 inflammation may modify the ACE2 expression in airways, there is no direct evidence that they will change the risk for COVID‐19. Considering the risk of exacerbation of primary airway diseases after stopping baseline treatment and subsequently increased potential risk to COVID‐19, the baseline treatments are suggested to be continued for patients with chronic inflammatory airway disorders.
Cardiovascular diseases including hypertension and heart failure are the leading causes of death globally. 118 As the first‐choice drugs in cardiovascular diseases, ACE inhibitors and angiotensin receptor blockers (ARBs) are thought to up‐regulate ACE2 expression, thus may increase the risk of COVID‐19. 118 Therefore, concerns have been raised regarding the use of ACE inhibitors and ARBs in the context of COVID‐19 pandemic. However, by analysing the gene expression of ACE2 and TMPRSS2 in 1,051 lung tissue samples, Milne et al found that ACE inhibitor use reduced ACE2 and TMPRSS2 gene expression, and ARBs use did not alter ACE2 and TMPRSS2 gene expression in the lung. 119 Several epidemiological studies demonstrated that the use of ACE inhibitors or ARBs was not associated with the increased risk or mortality of COVID‐19. 118 , 120 , 121 , 122 The American College of Cardiology, the American Heart Association, the Heart Failure Society of America, and the European Society of Cardiology suggest that ACE inhibitors and ARBs should not be discontinued for patients with cardiovascular diseases. 123 , 124
Antivirus drugs such as chloroquine, hydroxychloroquine, remdesivir, and lopinavir have been recommended for the treatment of COVID‐19, although none of them has been thoroughly proved by randomized controlled trials. 11 , 125 There are big controversies regarding their therapeutic effects on COVID‐19. Since ACE2 is likely an IFN‐stimulated gene and may be induced by viral infection, 41 antiviral treatment may reduce the ACE2 expression in airways in patients with COVID‐19. Of note, as the Ang II converting enzyme, ACE2 protects lung from a variety of injuries. 49 The approaches targeting ACE2 for treating COVID‐19 should be carefully considered in relation to the protective role of ACE2 in lung physiology and pathology.
In summary, enriched ACE2 expression in nasal epithelial cells and AT2 cells indicates that nose and lung are susceptible to the SARS‐CoV‐2 infection, thus representing the primary target organ of SARS‐CoV‐2. Expression of ACE2 in airways is tightly regulated by inflammatory milieu, such as IFNs and T2 cytokines, and environmental and internal stimuli, such as allergens and viruses. Altered ACE2 expression in chronic inflammatory airway diseases may modify COVID‐19 risk, which however needs to be proved by further epidemiological and functional studies. Potential approaches targeting ACE2 for COVID‐19 prevention and therapy should be developed under careful consideration regarding the multiple roles of ACE2 in lung physiology and pathology.
CONFLICT OF INTEREST
The authors declare no conflict of interests.
AUTHOR CONTRIBUTIONS
All authors participated in drafting and writing the manuscript and approved the manuscript.
Yao Y, Wang H, Liu Z. Expression of ACE2 in airways: Implication for COVID‐19 risk and disease management in patients with chronic inflammatory respiratory diseases. Clin Exp Allergy. 2020;50:1313–1324. 10.1111/cea.13746
Yin Yao and Hai Wang contributed equally to this work.
Funding information
This work was supported by the National Natural Science Foundation of China (NSFC) grants 81920108011 and 81630024.
DATA AVAILABILITY STATEMENT
Data sharing is not applicable to this article as no new data were created or analysed in this study.
REFERENCES
- 1. Wang C, Horby PW, Hayden FG, Gao GF. A novel coronavirus outbreak of global health concern. Lancet. 2020;395:470‐473. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Huang C, Wang Y, Li X, et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet. 2020;395:497‐506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Guan WJ, Ni ZY, Hu Y, et al. Clinical characteristics of coronavirus disease 2019 in China. N Engl J Med. 2020;382:1708‐1720. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Goyal P, Choi JJ, Pinheiro LC, et al. Clinical characteristics of Covid‐19 in New York city. N Engl J Med. 2020;382:2372‐2374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Wu Z, McGoogan JM. Characteristics of and important lessons from the coronavirus disease 2019 (COVID‐19) outbreak in China. JAMA. 2020;323(13):1239. [DOI] [PubMed] [Google Scholar]
- 6. Dong X, Cao YY, Lu XX, et al. Eleven faces of coronavirus disease 2019. Allergy. 2020;75:1699‐1709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Zhang JJ, Dong X, Cao YY, et al. Clinical characteristics of 140 patients infected with SARS‐CoV‐2 in Wuhan, China. Allergy. 2020;75:1730‐1741. [DOI] [PubMed] [Google Scholar]
- 8. Azkur AK, Akdis M, Azkur D, et al. Immune response to SARS‐CoV‐2 and mechanisms of immunopathological changes in COVID‐19. Allergy. 2020;75:1564‐1581. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Sakurai A, Sasaki T, Kato S, et al. Natural history of asymptomatic SARS‐CoV‐2 infection. N Engl J Med. 2020;383(9):885–886. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Wiersinga WJ, Rhodes A, Cheng AC, Peacock SJ, Prescott HC. Pathophysiology, transmission, diagnosis, and treatment of coronavirus disease 2019 (COVID‐19): a review. JAMA. 2020;324(8):782. [DOI] [PubMed] [Google Scholar]
- 11. Sanders JM, Monogue ML, Jodlowski TZ, Cutrell JB. Pharmacologic treatments for coronavirus disease 2019 (COVID‐19): a review. JAMA. 2020. 10.1001/jama.2020.6019 [DOI] [PubMed] [Google Scholar]
- 12. Williamson EJ, Walker AJ, Bhaskaran K, et al. OpenSAFELY: factors associated with COVID‐19 death in 17 million patients. Nature. 2020. 10.1038/s41586-020-2521-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Guan WJ, Liang WH, Zhao Y, et al. Comorbidity and its impact on 1590 patients with Covid‐19 in China: a nationwide analysis. Eur Respir J. 2020;14(55):2000547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Zhang JJ, Cao YY, Tan G, et al. Clinical, radiological and laboratory characteristics and risk factors for severity and mortality of 289 hospitalized COVID‐19 patients. Allergy. 2020. 10.1111/all.14496 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Meng Y, Lu W, Guo E, et al. Cancer history is an independent risk factor for mortality in hospitalized COVID‐19 patients: a propensity score‐matched analysis. J Hematol Oncol. 2020;13:75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Li X, Xu S, Yu M, et al. Risk factors for severity and mortality in adult COVID‐19 inpatients in Wuhan. J Allergy Clin Immunol. 2020;146:110‐118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Shang J, Ye G, Shi K, et al. Structural basis of receptor recognition by SARS‐CoV‐2. Nature. 2020;581:221‐224. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Guo T, Fan Y, Chen M, et al. Cardiovascular implications of fatal outcomes of patients with coronavirus disease 2019 (COVID‐19). JAMA Cardiol. 2020;27(5):1‐8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Cai G, Bosse Y, Xiao F, Kheradmand F, Amos CI. Tobacco smoking increases the lung gene expression of ACE2, the receptor of SARS‐CoV‐2. Am J Respir Crit Care Med. 2020;201:1557‐1559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Lippi G, Henry BM. Chronic obstructive pulmonary disease is associated with severe coronavirus disease 2019 (COVID‐19). Respir Med. 2020;167:105941. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Sattar N, McInnes IB, McMurray JJV. Obesity is a risk factor for severe COVID‐19 infection: multiple potential mechanisms. Circulation. 2020;142:4‐6. [DOI] [PubMed] [Google Scholar]
- 22. Cai Q, Chen F, Wang T, et al. Obesity and COVID‐19 severity in a designated hospital in Shenzhen, China. Diabetes Care. 2020;43:1392‐1398. [DOI] [PubMed] [Google Scholar]
- 23. Khunti K, Singh AK, Pareek M, Hanif W. Is ethnicity linked to incidence or outcomes of covid‐19? BMJ. 2020;369:m1548. [DOI] [PubMed] [Google Scholar]
- 24. Webb Hooper M, Nápoles AM, Pérez‐Stable EJ. COVID‐19 and racial/ethnic disparities. JAMA. 2020;323:2466‐2467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Fattorini D, Regoli F. Role of the chronic air pollution levels in the Covid‐19 outbreak risk in Italy. Environ Pollut. 2020;264:114732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Wu X, Nethery RC, Sabath BM, et al. Exposure to air pollution and COVID‐19 mortality in the United States: a nationwide cross‐sectional study. MedRxiv; 2020. 10.1101/2020.04.05.20054502 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Zhu ZZ, Hasegawa K, Ma BS, et al. Association of asthma and its genetic predisposition with the risk of severe COVID‐19. J Allergy Clin Immunol. 2020;146:327‐329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Roberton T, Carter ED, Chou VB, et al. Early estimates of the indirect effects of the COVID‐19 pandemic on maternal and child mortality in low‐income and middle‐income countries: a modelling study. Lancet Glob Health. 2020;8:e901‐e908. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Lau FH, Majumder R, Torabi R, et al. Vitamin D insufficiency is prevalent in severe COVID‐19. BioRxiv. 2020. 10.1101/2020.04.24.20075838 [DOI] [Google Scholar]
- 30. Wu Y, Feng Z, Li P, Yu Q. Relationship between ABO blood group distribution and clinical characteristics in patients with COVID‐19. Clin Chim Acta. 2020;509:220‐223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Li WH, Moore MJ, Vasilieva N, et al. Angiotensin‐converting enzyme 2 is a functional receptor for the SARS coronavirus. Nature. 2003;426:450‐454. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Zhou P, Yang XL, Wang XG, et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020;579:270‐273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. de Wit E, van Doremalen N, Falzarano D, Munster VJ. SARS and MERS: recent insights into emerging coronaviruses. Nat Rev Microbiol. 2016;14:523‐534. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Letko M, Marzi A, Munster V. Functional assessment of cell entry and receptor usage for SARS‐CoV‐2 and other lineage B betacoronaviruses. Nat Microbiol. 2020;5:562‐569. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Wan Y, Shang J, Graham R, et al. Receptor recognition by the novel coronavirus from Wuhan: an analysis based on decade‐long structural studies of SARS coronavirus. J Virol. 2020;94. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Walls AC, Park YJ, Tortorici MA, Wall A, McGuire AT, Veesler D. Structure, function, and antigenicity of the SARS‐CoV‐2 spike glycoprotein. Cell. 2020;181:281‐292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Wrapp D, Wang NS, Corbett KS, et al. Cryo‐EM structure of the 2019‐nCoV spike in the prefusion conformation. Science. 2020;367:1260‐1263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Hoffmann M, Kleine‐Weber H, Schroeder S, et al. SARS‐CoV‐2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell. 2020;181:271‐280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Leung JM, Yang CX, Tam A, et al. ACE‐2 expression in the small airway epithelia of smokers and COPD patients: implications for COVID‐19. Eur Respir J. 2020;55:2000688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Sungnak W, Huang N, Becavin C, et al. SARS‐CoV‐2 entry factors are highly expressed in nasal epithelial cells together with innate immune genes. Nat Med. 2020;26:681‐687. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Ziegler C, Allon SJ, Nyquist SK, et al. SARS‐CoV‐2 receptor ACE2 is an interferon‐stimulated gene in human airway epithelial cells and is enriched in specific cell subsets across tissues. Cell. 2020;181:1016‐1035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Sajuthi SP, DeFord P, Jackson ND, et al. Type 2 and interferon inflammation strongly regulate SARS‐CoV‐2 related gene expression in the airway epithelium. BioRxiv; 2020. 10.1101/2020.04.09.034454 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Pawankar R. Allergic diseases and asthma: a global public health concern and a call to action. World Allergy Organ J. 2014;7:12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Celli BR, Wedzicha JA. Update on clinical aspects of chronic obstructive pulmonary disease. N Engl J Med. 2019;381:1257‐1266. [DOI] [PubMed] [Google Scholar]
- 45. Dutau G, Annesi‐Maesanno I, Klossek JM, et al. The allergic rhinitis and its impact on the asthma (ARIA 2008). Rev Fr Allergol. 2008;48:376‐379. [Google Scholar]
- 46. Donoghue M, Hsieh F, Baronas E, et al. A novel angiotensin‐converting enzyme‐related carboxypeptidase (ACE2) converts angiotensin I to angiotensin 1–9. Circ Res. 2000;87:E1‐E9. [DOI] [PubMed] [Google Scholar]
- 47. Tipnis SR, Hooper NM, Hyde R, Karran E, Christie G, Turner AJ. A human homolog of angiotensin‐converting enzyme. Cloning and functional expression as a captopril‐insensitive carboxypeptidase. J Biol Chem. 2000;275:33238‐33243. [DOI] [PubMed] [Google Scholar]
- 48. Turner AJ, Hooper NM. The angiotensin‐converting enzyme gene family: genomics and pharmacology. Trends Pharmacol Sci. 2002;23:177‐183. [DOI] [PubMed] [Google Scholar]
- 49. Hamming I, Cooper ME, Haagmans BL, et al. The emerging role of ACE2 in physiology and disease. J Pathol. 2007;212:1‐11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Patel S, Rauf A, Khan H, Abu‐Izneid T. Renin‐angiotensin‐aldosterone (RAAS): the ubiquitous system for homeostasis and pathologies. Biomed Pharmacother. 2017;94:317‐325. [DOI] [PubMed] [Google Scholar]
- 51. Chen L, Hao G. The role of angiotensin‐converting enzyme 2 in coronaviruses/influenza viruses and cardiovascular disease. Cardiovasc Res. 2020;116(12):1932–1936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Lambert DW, Yarski M, Warner FJ, et al. Tumor necrosis factor‐alpha convertase (ADAM17) mediates regulated ectodomain shedding of the severe‐acute respiratory syndrome‐coronavirus (SARS‐CoV) receptor, angiotensin‐converting enzyme‐2 (ACE2). J Biol Chem. 2005;280:30113‐30119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Imai Y, Kuba K, Rao S, et al. Angiotensin‐converting enzyme 2 protects from severe acute lung failure. Nature. 2005;436:112‐116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Jia H. Pulmonary Angiotensin‐converting enzyme 2 (ACE2) and inflammatory lung disease. Shock. 2016;46:239‐248. [DOI] [PubMed] [Google Scholar]
- 55. Shenoy V, Ferreira AJ, Qi YF, et al. The angiotensin‐converting enzyme 2/Angiogenesis‐(1–7)/Mas axis confers cardiopulmonary protection against lung fibrosis and pulmonary hypertension. Am J Resp Crit Care. 2010;182:1065‐1072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Xue T, Wei N, Xin Z, Qingyu X. Angiotensin‐converting enzyme‐2 overexpression attenuates inflammation in rat model of chronic obstructive pulmonary disease. Inhal Toxicol. 2014;26:14‐22. [DOI] [PubMed] [Google Scholar]
- 57. Magalhaes GS, Rodrigues‐Machado MG, Motta‐Santos D, et al. Angiotensin‐(1–7) attenuates airway remodelling and hyperresponsiveness in a model of chronic allergic lung inflammation. Br J Pharmacol. 2015;172:2330‐2342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Han DP, Penn‐Nicholson A, Cho MW. Identification of critical determinants on ACE2 for SARS‐CoV entry and development of a potent entry inhibitor. Virology. 2006;350:15‐25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Kuba K, Imai Y, Rao S, et al. A crucial role of angiotensin converting enzyme 2 (ACE2) in SARS coronavirus‐induced lung injury. Nat Med. 2005;11:875‐879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Yang XH, Deng W, Tong Z, et al. Mice transgenic for human angiotensin‐converting enzyme 2 provide a model for SARS coronavirus infection. Comp Med. 2007;57:450‐459. [PubMed] [Google Scholar]
- 61. Coutard B, Valle C, de Lamballerie X, Canard B, Seidah NG, Decroly E. The spike glycoprotein of the new coronavirus 2019‐nCoV contains a furin‐like cleavage site absent in CoV of the same clade. Antiviral Res. 2020;176:104742. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62. Hasan A, Paray BA, Hussain A, et al. A review on the cleavage priming of the spike protein on coronavirus by angiotensin‐converting enzyme‐2 and furin. J Biomol Struct Dyn. 2020;1–9. 10.1080/07391102.2020.1754293 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63. Hoffmann M, Kleine‐Weber H, Pohlmann S. A multibasic cleavage site in the spike protein of SARS‐CoV‐2 is essential for infection of human lung cells. Mol Cell. 2020;78:779‐784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64. Yan R, Zhang Y, Li Y, et al. Structural basis for the recognition of SARS‐CoV‐2 by full‐length human ACE2. Science. 2020;367:1444‐1448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65. Papalexi E, Satija R. Single‐cell RNA sequencing to explore immune cell heterogeneity. Nat Rev Immunol. 2018;18:35‐45. [DOI] [PubMed] [Google Scholar]
- 66. Yofe I, Dahan R, Amit I. Single‐cell genomic approaches for developing the next generation of immunotherapies. Nat Med. 2020;26:171‐177. [DOI] [PubMed] [Google Scholar]
- 67. Bertram S, Heurich A, Lavender H, et al. Influenza and SARS‐coronavirus activating proteases TMPRSS2 and HAT are expressed at multiple sites in human respiratory and gastrointestinal tracts. PLoS One. 2012;7:e35876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68. Yan CH, Faraji F, Prajapati DP, et al. Association of chemosensory dysfunction and Covid‐19 in patients presenting with influenza‐like symptoms. Int Forum Allergy Rhinol. 2020;10:806‐813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69. Lechien JR, Chiesa‐Estomba CM, De Siati DR, et al. Olfactory and gustatory dysfunctions as a clinical presentation of mild‐to‐moderate forms of the coronavirus disease (COVID‐19): a multicenter European study. Eur Arch Oto‐Rhino‐L. 2020;277:2251‐2261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70. Menni C, Valdes AM, Freidin MB, et al. Real‐time tracking of self‐reported symptoms to predict potential COVID‐19. Nat Med. 2020;26:1037‐1040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71. Xu H, Zhong L, Deng J, et al. High expression of ACE2 receptor of 2019‐nCoV on the epithelial cells of oral mucosa. Int J Oral Sci. 2020;12:8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72. Butowt R, Bilinska K. SARS‐CoV‐2: olfaction, brain infection, and the urgent need for clinical samples allowing earlier virus detection. ACS Chem Neurosci. 2020;11:1200‐1203. [DOI] [PubMed] [Google Scholar]
- 73. Brann DH, Tsukahara T, Weinreb C, et al. Non‐neural expression of SARS‐CoV‐2 entry genes in the olfactory epithelium suggests mechanisms underlying anosmia in COVID‐19 patients. BioRxiv. 2020. 10.1101/2020.03.25.009084 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74. Fodoulian L, Tuberosa J, Rossier D, et al. SARS‐CoV‐2 receptor and entry genes are expressed by sustentacular cells in the human olfactory neuroepithelium. BioRxiv. 2020. 10.1101/2020.03.31.013268 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75. Zhao Y, Zhao Z, Wang Y, et al. Single‐cell RNA expression profiling of ACE2, the receptor of SARS‐CoV‐2. Am J Respir Crit Care Med. 2020. 10.1164/rccm.202001-0179LE [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76. Lukassen S, Chua RL, Trefzer T, et al. SARS‐CoV‐2 receptor ACE2 and TMPRSS2 are primarily expressed in bronchial transient secretory cells. EMBO J. 2020;39:e105114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77. Qi F, Qian S, Zhang S, Zhang Z. Single cell RNA sequencing of 13 human tissues identify cell types and receptors of human coronaviruses. Biochem Biophys Res Commun. 2020;526:135‐140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78. Zou X, Chen K, Zou J, et al. Single‐cell RNA‐seq data analysis on the receptor ACE2 expression reveals the potential risk of different human organs vulnerable to 2019‐nCoV infection. Front Med. 2020;14:185‐192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79. Li G, He X, Zhang L, et al. Assessing ACE2 expression patterns in lung tissues in the pathogenesis of COVID‐19. J Autoimmun. 2020;112:102463. 10.1016/j.jaut.2020.102463 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80. Bezara MO, Thurman A, Pezzulo A, et al. Heterogeneous expression of the SARS‐Coronavirus‐2 receptor ACE2 in the human respiratory tract. BioRxiv. 2020. 10.1101/2020.04.22.056127 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81. Bailey CC, Zhong G, Huang IC, Farzan M. IFITM‐family proteins: the cell's first line of antiviral defense. Annu Rev Virol. 2014;1:261‐283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82. Broggi A, Granucci F, Zanoni I. Type III interferons: balancing tissue tolerance and resistance to pathogen invasion. J Exp Med. 2020;217:e20190295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83. Steuerman Y, Cohen M, Peshes‐Yaloz N, et al. Dissection of influenza infection in vivo by single‐cell RNA sequencing. Cell Syst. 2018;6:679‐691. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84. Jackson DJ, Busse WW, Bacharier LB, et al. Association of respiratory allergy, asthma and expression of the SARS‐CoV‐2 receptor, ACE2. J Allergy Clin Immunol. 2020;146:203‐206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85. Kimura H, Francisco D, Conway M, et al. Type 2 inflammation modulates ACE2 and TMPRSS2 in airway epithelial cells. J Allergy Clin Immun. 2020;146:80‐88. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86. Wheatley LM, Togias A. Allergic rhinitis. New Engl J Med. 2015;372:456‐463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87. Lindsley AW, Schwartz JT, Rothenberg ME. Eosinophil responses during COVID‐19 infections and coronavirus vaccination. J Allergy Clin Immunol. 2020;146:1‐7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88. Xie G, Ding F, Han L, et al. The role of peripheral blood eosinophil counts in COVID‐19 patients. Allergy. 2020. 10.1111/all.14465 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89. Camiolo MJ, Gauthier M, Kaminski N, et al. Expression of SARS‐CoV‐2 receptor ACE2 and coincident host response signature varies by asthma inflammatory phenotype. J Allergy Clin Immunol. 2020;146(2):315–324.e7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90. Shi W, Gao Z, Ding Y, et al. Clinical characteristics of COVID‐19 patients combined with allergy. Allergy. 2020;75(9):2405–2408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91. Wang D, Hu B, Hu C, et al. Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus‐infected pneumonia in Wuhan, China. JAMA. 2020;323:1061‐1069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92. Zhao Q, Meng M, Kumar R, et al. The impact of COPD and smoking history on the severity of Covid‐19: a systemic review and meta‐analysis. J Med Virol. 2020;92(10):1915–1921. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93. Brake SJ, Barnsley K, Lu W, et al. Smoking upregulates angiotensin‐converting enzyme‐2 receptor: a potential adhesion site for novel coronavirus SARS‐CoV‐2 (Covid‐19). J Clin Med. 2020;9(3):841. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94. Boardman C, Chachi L, Gavrila A, et al. Mechanisms of glucocorticoid action and insensitivity in airways disease. Pulm Pharmacol Ther. 2014;29:129‐143. [DOI] [PubMed] [Google Scholar]
- 95. Peters MC, Sajuthi S, Deford P, et al. COVID‐19 related genes in sputum cells in asthma: relationship to demographic features and corticosteroids. Am J Respir Crit Care Med. 2020;202:83‐90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96. Akenroye AT, Wood R, Keet C. Asthma, biologics, corticosteroids, and COVID‐19. Ann Allergy Asthma Immunol. 2020;125:12‐13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97. Bachert C, Zhang N, Cavaliere C, Weiping W, Gevaert E, Krysko O. Biologics for chronic rhinosinusitis with nasal polyps. J Allergy Clin Immunol. 2020;145:725‐739. [DOI] [PubMed] [Google Scholar]
- 98. Förster‐Ruhrmann U, Szczepek AJ, Bachert C, Olze H. COVID‐19 infection in a patient with severe chronic rhinosinusitis with nasal polyps during therapy with dupilumab. J Allergy Clin Immun. 2020;146:218‐220. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99. Castagnoli R, Votto M, Licari A, et al. Severe acute respiratory syndrome coronavirus 2 (SARS‐CoV‐2) infection in children and adolescents: a systematic review. JAMA Pediatr. 2020;174(9):882. [DOI] [PubMed] [Google Scholar]
- 100. Bunyavanich S, Do A, Vicencio A. Nasal gene expression of angiotensin‐converting enzyme 2 in children and adults. JAMA. 2020;323:2427‐2429. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101. Liu J, Ji H, Zheng W, et al. Sex differences in renal angiotensin converting enzyme 2 (ACE2) activity are 17beta‐oestradiol‐dependent and sex chromosome‐independent. Biol Sex Differ. 2010;1:6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102. Seillet C, Laffont S, Tremollieres F, et al. The TLR‐mediated response of plasmacytoid dendritic cells is positively regulated by estradiol in vivo through cell‐intrinsic estrogen receptor alpha signaling. Blood. 2012;119:454‐464. [DOI] [PubMed] [Google Scholar]
- 103. Jian L, Yi W, Zhang N, et al. Perspective: COVID‐19, implications of nasal diseases and consequences for their management. J Allergy Clin Immunol. 2020;146:67‐69. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104. Wang XD, Zheng M, Lou HF, et al. An increased prevalence of self‐reported allergic rhinitis in major Chinese cities from 2005 to 2011. Allergy. 2016;71:1170‐1180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105. Song J, Deng Y‐K, Wang H, et al. Self‐reported taste and smell disorders in patients with COVID‐19: distinct features in China. BioRxiv. 2020. 10.1101/2020.06.12.20128298 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106. Wang X, Zhang N, Bo M, et al. Diversity of TH cytokine profiles in patients with chronic rhinosinusitis: a multicenter study in Europe, Asia, and Oceania. J Allergy Clin Immunol. 2016;138:1344‐1353. [DOI] [PubMed] [Google Scholar]
- 107. Zhang Y, Gevaert E, Lou H, et al. Chronic rhinosinusitis in Asia. J Allergy Clin Immunol. 2017;140:1230‐1239. [DOI] [PubMed] [Google Scholar]
- 108. Chhiba KD, Patel GB, Vu THT, et al. Prevalence and characterization of asthma in hospitalized and non‐hospitalized patients with COVID‐19. J Allergy Clin Immunol. 2020;146(2):307–314.e4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109. Corne JM, Marshall C, Smith S, et al. Frequency, severity, and duration of rhinovirus infections in asthmatic and non‐asthmatic individuals: a longitudinal cohort study. Lancet. 2002;359:831‐834. [DOI] [PubMed] [Google Scholar]
- 110. Johnston SL. Asthma and COVID‐19: is asthma a risk factor for severe outcomes? Allergy. 2020;75:1543‐1545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111. Yang X, Yu Y, Xu J, et al. Clinical course and outcomes of critically ill patients with SARS‐CoV‐2 pneumonia in Wuhan, China: a single‐centered, retrospective, observational study. Lancet Respir Med. 2020;8:475‐481. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112. Grasselli G, Zangrillo A, Zanella A, et al. Baseline characteristics and outcomes of 1591 patients infected with SARS‐CoV‐2 admitted to ICUs of the lombardy region, Italy. JAMA. 2020;323:1574‐1581. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113. World Health Organization (WHO) . WHO Clinical Management of Severe Acute Respiratory Infection When Novel Coronavirus (nCoV) Infection is Suspected. Geneva, Switzerland: World Health Organization. https://www.who.int/publications‐detail/clinical‐management‐of‐severe‐acute‐respiratory‐infection‐when‐novel‐coronavirus‐(ncov)‐infection‐is‐suspected. Accessed February 2, 2020. [Google Scholar]
- 114. Bousquet J, Akdis C, Jutel M, et al. Intranasal corticosteroids in allergic rhinitis in COVID‐19 infected patients: an ARIA‐EAACI statement. Allergy. 2020. 10.1111/all.14302 [DOI] [PubMed] [Google Scholar]
- 115. GINA recommendations for inhaled asthma controller medications. Global Initiative for Asthma, 19 March 2020. https://ginasthma.org/recommendations‐for‐inhaled‐asthma‐controller‐medications/. Accessed 19 March, 2020.
- 116. Shaker MS, Oppenheimer J, Grayson M, et al. COVID‐19: pandemic contingency planning for the allergy and immunology clinic. J Allergy Clin Immunol Pract. 2020;8:1477‐1488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117. The Global Initiative for Chronic Obstructive Lung Disease (GOLD) . GOLD COVID‐19 guidance, the Global Initiative for Chronic Obstructive Lung Disease, 23 March 2020. https://goldcopd.org/gold‐covid‐19‐guidance/. Accessed 23 March, 2020.
- 118. Mancia G, Rea F, Ludergnani M, et al. Renin–Angiotensin–Aldosterone system blockers and the risk of Covid‐19. N Engl J Med. 2020;382:2431‐2440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119. Milne S, Yang CX, Timens W, et al. SARS‐CoV‐2 receptor ACE2 gene expression and RAAS inhibitors. Lancet Resp Med. 2020;8:E50‐E51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120. Fosbøl EL, Butt JH, Østergaard L, et al. Association of angiotensin‐converting enzyme inhibitor or angiotensin receptor blocker use with COVID‐19 diagnosis and mortality. JAMA. 2020;324:168‐177.] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121. de Abajo FJ, Rodríguez‐Martín S, Lerma V, et al. Use of renin–angiotensin–aldosterone system inhibitors and risk of COVID‐19 requiring admission to hospital: a case‐population study. Lancet. 2020;395:1705‐1714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122. Reynolds HR, Adhikari S, Pulgarin C, et al. Renin–angiotensin–aldosterone system inhibitors and risk of Covid‐19. N Engl J Med. 2020;382:2441‐2448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123. American College of Cardiology . HFSA/ACC/AHA statement addresses concerns re: using RAAS antagonists in COVID‐19. https://www.acc.org/latest‐in‐cardiology/articles/2020/03/17/08/59/hfsa‐acc‐aha‐statement‐addresses‐concerns‐re‐using‐raas‐antagonists‐in‐covid‐19. Accessed 17 March, 2020.
- 124. European Society of Cardiology . Position statement of the ESC Council on Hypertension on ACE‐inhibitors and angiotensin receptor blockers. https://www.escardio.org/Councils/Council‐on‐Hypertension‐(CHT)/News/position‐statement‐of‐the‐esc‐council‐on‐hypertension‐on‐ace‐inhibitors‐and‐ang. Accessed 13 March, 2020.
- 125. Cao B, Wang Y, Wen D, et al. A trial of lopinavir‐ritonavir in adults hospitalized with severe Covid‐19. N Engl J Med. 2020;382:1787‐1799. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Data sharing is not applicable to this article as no new data were created or analysed in this study.


