Abstract
Background
Metformin is a widely prescribed antidiabetic BCS Class III drug (low permeability) that depends on active transport for its absorption and disposition. It is recommended by the US Food and Drug Administration as a clinical substrate of organic cation transporter 2/multidrug and toxin extrusion protein for drug–drug interaction studies. Cimetidine is a potent organic cation transporter 2/multidrug and toxin extrusion protein inhibitor.
Objective
The objective of this study was to provide mechanistic whole-body physiologically based pharmacokinetic models of metformin and cimetidine, built and evaluated to describe the metformin-SLC22A2 808G>T drug–gene interaction, the cimetidine-metformin drug–drug interaction, and the impact of renal impairment on metformin exposure.
Methods
Physiologically based pharmacokinetic models were developed in PK-Sim® (version 8.0). Thirty-nine clinical studies (dosing range 0.001–2550 mg), providing metformin plasma and urine data, positron emission tomography measurements of tissue concentrations, studies in organic cation transporter 2 polymorphic volunteers, drug–drug interaction studies with cimetidine, and data from patients in different stages of chronic kidney disease, were used to develop the metformin model. Twenty-seven clinical studies (dosing range 100–800 mg), reporting cimetidine plasma and urine concentrations, were used for the cimetidine model development.
Results
The established physiologically based pharmacokinetic models adequately describe the available clinical data, including the investigated drug–gene interaction, drug–drug interaction, and drug–drug–gene interaction studies, as well as the metformin exposure during renal impairment. All modeled drug–drug interaction area under the curve and maximum concentration ratios are within 1.5-fold of the observed ratios. The clinical data of renally impaired patients shows the expected increase in metformin exposure with declining kidney function, but also indicates counter-regulatory mechanisms in severe renal disease; these mechanisms were implemented into the model based on findings in preclinical species.
Conclusions
Whole-body physiologically based pharmacokinetic models of metformin and cimetidine were built and qualified for the prediction of metformin pharmacokinetics during drug–gene interaction, drug–drug interaction, and different stages of renal disease. The model files will be freely available in the Open Systems Pharmacology model repository. Current guidelines for metformin treatment of renally impaired patients should be reviewed to avoid overdosing in CKD3 and to allow metformin therapy of CKD4 patients.
Electronic supplementary material
The online version of this article (10.1007/s40262-020-00896-w) contains supplementary material, which is available to authorized users.
Key Points
| A whole-body physiologically based pharmacokinetic model of metformin, the fourth most commonly prescribed drug in the USA, has been carefully developed and evaluated to describe the metformin concentrations in blood, kidney, and urine. In addition, a whole-body physiologically based pharmacokinetic model of cimetidine, a potent multidrug and toxin extrusion protein 1 inhibitor used in drug–drug interaction studies, has been established. |
| These models have been applied to describe and predict the metformin-SLC22A2 808G>T drug–gene interaction, the cimetidine-metformin drug–drug interaction, and a combined drug–drug–gene interaction study, in which different SLC22A2 genotypes were additionally challenged with cimetidine co-administration. |
| Furthermore, the pathophysiological changes during renal impairment have been assessed and implemented to describe the increased metformin exposure of patients with different stages of chronic kidney disease. For severe chronic kidney disease, this analysis indicates an induction of organic cation transporter 2 and multidrug and toxin extrusion protein 1, possibly as an adaptation to progressing uremia/hyperuricemia. The final pathophysiologically based pharmacokinetic model was applied to generate metformin dosing recommendations for CKD3A-4 patients. |
Introduction
Metformin is an oral antidiabetic that reduces blood glucose levels. It is the first-line therapy for type 2 diabetes mellitus (T2DM) and the fourth most commonly prescribed outpatient medication in the USA, with almost 80 million prescriptions in 2017 [1].
Metformin is a BCS Class III drug of high solubility and very low permeability, positively charged at physiological pH and depends on active transport to cross biological membranes. The metformin rate of absorption is slower than its rate of elimination [2] and the absorption is restricted to the upper intestine [3], leading to incomplete absorption of metformin, an oral bioavailability of 50–60%, and the excretion of approximately 30% of an oral dose, unabsorbed, with the feces [2, 4]. Furthermore, the absorption of metformin is saturable, with higher doses showing decreased dose-normalized plasma concentrations and a decreased fraction excreted to urine [4, 5]. Following its absorption, metformin is not bound to plasma proteins [2, 4, 6], not metabolized [2, 6], and not secreted to bile [2, 4, 7], but excreted unchanged with the urine by passive glomerular filtration and active renal secretion through the sequential action of organic cation transporter 2 (OCT2) and multidrug and toxin extrusion protein 1 (MATE1). Although there are early reports of MATE2-K expression in the human kidney [8, 9], a recent quantitative study found only negligible amounts of MATE2-K compared to MATE1 [10]. Renal clearance is approximately 500 mL/min [11] with a strong correlation between the renal clearances of metformin and creatinine [4]. Patients with renal impairment show a marked increase in metformin exposure, with three- to ten-fold higher plasma trough concentrations in chronic kidney disease (CKD) stages 3A-5 [12]. As a consequence, metformin is contraindicated in patients with a glomerular filtration rate (GFR) < 30 mL/min (i.e., CKD stages 4 and 5) [13, 14], depriving these patients of metformin as a treatment option.
The impact of genetic polymorphisms on the absorption and disposition of metformin (drug–gene interactions or “DGIs”) has been investigated in a multitude of clinical trials, yielding to some extent contradictory results. The transporters of primary interest in these studies were the plasma membrane monoamine transporter (PMAT), OCT1, OCT2, and MATE1, where variations in OCT2 seem to have the largest impact on the plasma concentrations of metformin [15–19]. The most common polymorphism in the gene encoding for OCT2 is the SLC22A2 808G>T single-nucleotide polymorphism [20], which results in an amino acid exchange from alanine to serine (A270S) and presumably increased function, leading to decreased exposure with ~ 13–20% decreased maximum concentration (Cmax) [17, 18, 21].
A third factor that impacts metformin exposure is drug–drug interactions (DDIs). Metformin displays a list of 333 DDIs, with 13 major and 293 moderate interactions [22]. Even though some of these occur on the pharmacodynamic level, pharmacokinetic DDIs are clinically relevant and may call for an adjustment of the co-administration regimen. As metformin is exclusively eliminated by glomerular filtration and secretion through the renal organic cation transport system, co-treatment with a potent inhibitor of this transport pathway, such as cimetidine, decreases the renal clearance of metformin and increases metformin exposure (+ 50% area under the curve [AUC]) [21, 23]. Metformin is recommended by the US Food and Drug Administration as an OCT2/MATE victim drug for clinical DDI studies [24].
The aim of this study was to build and evaluate a whole-body physiologically based pharmacokinetic (PBPK) model of metformin, applicable (1) to describe the impact of the metformin-SLC22A2 808G>T DGI on metformin exposure, (2) to dynamically model the cimetidine-metformin DDI, and (3) to analyze the impact of renal impairment on metformin exposure and generate dose recommendations for different stages of CKD. The newly developed and thoroughly evaluated metformin and cimetidine models will be freely available in the Open Systems Pharmacology PBPK model repository (https://www.open-systems-pharmacology.org), and the Electronic Supplementary Material (ESM) to this article is compiled to serve as a comprehensive and transparent documentation and reference.
Methods
Software
Physiologically based pharmacokinetic models were developed using PK-Sim® and MoBi® modeling software (Open Systems Pharmacology Suite 8.0, https://www.open-systems-pharmacology.org). Published clinical study data were digitized with GetData Graph Digitizer 2.26.0.20 (© S. Fedorov). Model input parameter optimization (Levenberg–Marquardt algorithm, multiple starting values) and sensitivity analysis were performed in PK-Sim®. All pharmacokinetic parameters and model performance measures derived from simulated and/or observed data were calculated in R 3.6.1 (The R Foundation for Statistical Computing, Vienna, Austria). Plots were generated in R and RStudio 1.1.423 (RStudio, Inc., Boston, MA, USA).
Physiologically Based Pharmacokinetic Model Building
Physiologically based pharmacokinetic model building was started with an extensive literature search to collect physicochemical parameters, mechanistic information on absorption, distribution, metabolism, and excretion processes, as well as published clinical studies. The general procedure of PBPK model building, including parameter optimization and generation of virtual individuals and virtual populations, is described in the ESM.
Physiologically Based Pharmacokinetic Model Evaluation
Model performance was evaluated with multiple methods. First, predicted population plasma concentration–time profiles were compared with the data observed in the respective clinical studies. As the clinical data from literature is mostly reported as arithmetic means ± standard deviation, population prediction arithmetic means and 68% prediction intervals were plotted, which corresponds to the range of ± 1 standard deviation around the mean, if normal distribution is assumed. In addition, the predicted plasma concentration values of all studies were plotted against their corresponding observed values in goodness-of-fit plots.
Furthermore, model performance was evaluated by comparison of predicted to observed AUC and Cmax values. All AUC values were calculated from the time of drug administration to the time of the last concentration measurement (AUClast).
As quantitative measures of model performance, mean relative deviation (MRD) of all predicted plasma concentrations (Eq. 1) and geometric mean fold error (GMFE) of all predicted AUClast and Cmax values (Eq. 2) were calculated. MRD and GMFE values ≤ 2 characterize an adequate model performance.
| 1 |
where cpredicted,i is the predicted plasma concentration, cobserved,i is the corresponding observed plasma concentration, and k is the number of observed values.
| 2 |
where predicted PK parameteri is the predicted AUClast or Cmax value, observed PK parameteri is the corresponding observed AUClast or Cmax value, and m is the number of studies.
Finally, the physiological plausibility of the parameter estimates and the results of sensitivity analyses were assessed. A detailed description of the sensitivity calculation is given in the ESM.
Modeling the Impact of Polymorphism
The impact of genetic polymorphism on the pharmacokinetics of metformin was implemented by splitting the polymorphic transporter in question into two transporters with half of the initial reference concentration each, corresponding to the two homologous chromosomal alleles in diploid humans. Each “wild-type” allele present in the simulated population (one in heterozygous individuals and two in homozygous individuals) was modeled with the transport rate constant identified during the initial model development. Each “variant” allele was modeled with an adapted transport rate constant that was identified based on clinical studies of metformin in homozygous “variant” individuals.
Drug–Drug Interaction Modeling
For mechanistic DDI modeling, the type of interaction (competitive inhibition, mechanism-based inhibition, induction) and the interaction parameters were extracted from in-vitro literature. These parameters were incorporated into the perpetrator PBPK model, to dynamically describe the impact of the perpetrator on the victim drug. The mathematical implementation is shown in the ESM.
The DDI modeling performance was assessed by comparison of predicted vs observed victim drug plasma concentration–time profiles when administered alone and during co-administration. In addition, predicted DDI AUClast ratios (Eq. 3) and DDI Cmax ratios (Eq. 4) were evaluated.
| 3 |
| 4 |
As a quantitative measure of the prediction accuracy, GMFE values of the predicted DDI AUClast ratios and DDI Cmax ratios were calculated according to Eq. (2).
Modeling of Renal Impairment
To model the impact of renal impairment on the pharmacokinetics of metformin, a literature search was conducted to identify the pathophysiological changes that occur in conjunction with renal impairment, including their extent at the different stages of CKD. In a next step, these differences in anatomy and physiology were implemented to create renally impaired individuals and to describe the published clinical studies of metformin in patients with CKD.
Results
Metformin Physiologically Based Pharmacokinetic Model Building and Evaluation
A whole-body PBPK model of metformin has been successfully developed. Thirty-nine clinical studies of intravenous or oral administration covering a broad dosing range (0.001–2550 mg), 22 studies thereof with corresponding metformin fraction excreted to urine data, were utilized for PBPK model building and evaluation. In addition, human 11C-metformin tissue concentration positron emission tomography (PET) measurements in the kidneys, liver, skeletal muscle, and intestines were included. Clinical studies are listed in the ESM.
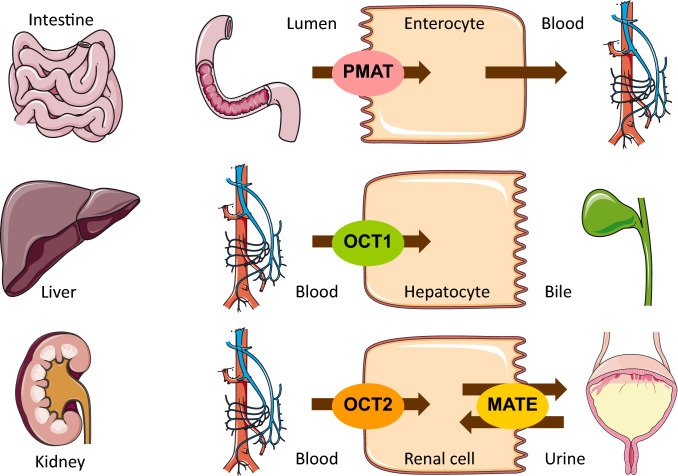
To describe the pharmacokinetics of metformin, active transport processes by PMAT, OCT1, OCT2, and MATE1 were implemented. These transporters were distributed and localized according to the current state of the literature, with their main sites of action illustrated in Fig. 1. PMAT was chosen to model the saturable absorption of metformin based on its good apparent affinity, high expression in the human small intestine, and localization at the luminal surface of enterocytes [25, 26], though the thiamine transporter 2 is also a likely candidate to contribute to the intestinal absorption of metformin [27, 28]. Renal excretion is modeled as passive glomerular filtration and active secretion through the sequential action of OCT2 and MATE1. Transporter distribution, localization, and transport directions are summarized in the system-dependent parameter table in the ESM; transport parameters are summarized in the metformin drug-dependent parameter table in the ESM.
Fig. 1.
Metformin transporters. Main sites of action of the transporters that were implemented to model the absorption, distribution, and excretion of metformin. Several different studies report that they found no secretion of metformin to bile [2, 4, 7]. Drawings by Servier Medical Art, licensed under CC BY 3.0. MATE multidrug and toxin extrusion protein, OCT organic cation transporter, PMAT plasma membrane monoamine transporter
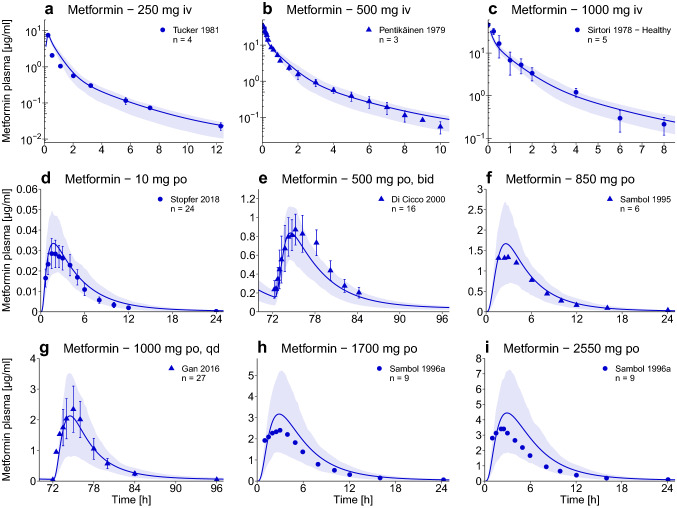
The good model performance is demonstrated in Fig. 2, using representative studies. Population predictions of all 39 clinical studies compared to observed data, shown in semi-logarithmic as well as linear plots, goodness-of-fit plots, and MRD values, are presented in the ESM. Predictions of metformin fraction excreted to urine are shown for all studies that provided observed data. For further evaluation of the model performance, predicted compared to observed AUClast and Cmax values, AUClast and Cmax GMFEs (1.20 and 1.24, respectively), and the results of the sensitivity analysis are documented in the ESM.
Fig. 2.
Metformin plasma concentrations. Population predictions of metformin plasma concentration–time profiles of representative intravenous (iv) and oral (po) studies, compared to observed data [2, 4–6, 39, 52–54]. Population prediction arithmetic means are shown as lines; the shaded areas illustrate the 68% population prediction intervals. Observed data are shown as dots (training dataset) or triangles (test dataset) ± standard deviation. Details on the study protocols, further studies, and quantitative model performance measures are provided in the ESM. bid twice daily, qd once daily
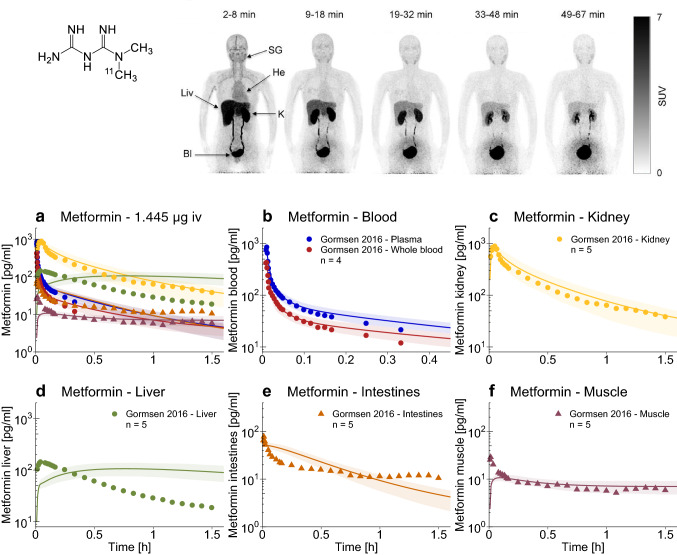
An important and novel feature of the presented model is the use of human 11C-metformin tissue concentration PET measurements for model development. As metformin is not metabolized, these PET images are unbiased by labeled metabolites. The unique transporter-controlled distribution of an intravenous 11C-metformin microdose over time [7] is shown in the upper part of Fig. 3. Noteworthy are the very high concentrations in the kidney, bladder, and liver, and the low permeation into other tissues. Population predictions of the quantified tissue concentrations are presented in the lower part of Fig. 3 and in the ESM. Metformin plasma, whole blood, kidney, and muscle concentrations are accurately described by the model, governed by the implemented transport processes and the low passive permeability of metformin.
Fig. 3.
Metformin tissue concentrations. Upper panel: pseudodynamic whole-body positron emission tomography imaging of a representative patient following an intravenous microdose of 11C-metformin [7]
© SNMMI. a–f Population predictions of metformin blood and tissue concentration–time profiles measured in the 11C-metformin positron emission tomography study, compared to observed data [7]. Population prediction arithmetic means are shown as lines; the shaded areas illustrate the 68% population prediction intervals. Observed data are shown as dots (training dataset) or triangles (test dataset). Linear plots are presented in the ESM. Bl bladder, He heart, iv intravenous, K kidney, Liv liver, SG submandibular gland, SUV standardized uptake value = concentration [kBq/mL]·body weight [g]/injected dose [kBq]
Impact of Organic Cation Transporter 2 Polymorphism
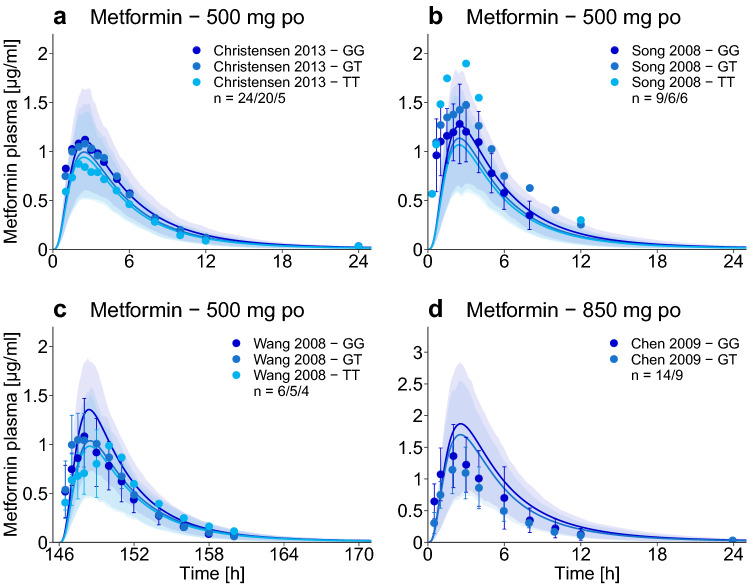
The impact of the SLC22A2 808G>T single-nucleotide polymorphism on metformin exposure (DGI) was implemented using the same Michaelis–Menten constant for both isoforms [17, 20, 29], but an increased transport rate constant for each minor OCT2 allele (808T) present in the simulated population. The 2.67-fold higher transport rate constant to describe the activity of the variant OCT2 (see metformin drug-dependent parameter table in the ESM) was optimized, based on the metformin plasma profiles of the homozygous 808TT populations studied by Christensen et al. and Wang et al. [18, 21]. The metformin plasma concentrations of all other 808TT and 808GT study populations were predicted and are presented in Fig. 4. Except for one study reporting a higher metformin exposure with the variant OCT2 [29], the DGI AUClast ratios for the hetero- and homozygous groups are well predicted, with 6/7 within two-fold of the observed ratios (see the ESM).
Fig. 4.
Impact of organic cation transporter 2 (OCT2) polymorphism. Population predictions of metformin plasma concentration–time profiles in different SLC22A2 genotypes, compared to observed data [17, 18, 21, 29]. Population prediction arithmetic means are shown as dark blue (SLC22A2 808GG, reference genotype) or lighter blue (SLC22A2 808GT and 808TT, variant genotypes) lines. The shaded areas illustrate the respective 68% population prediction intervals. Observed data are shown as dots ± standard deviation. Details on the study protocols, semilogarithmic plots, and quantitative model performance measures are provided in the ESM. GG SLC22A2 808GG genotype, GT SLC22A2 808GT genotype, po oral, TT SLC22A2 808TT genotype
Drug–Drug Interaction Modeling with Cimetidine
In addition to the metformin model, a whole-body PBPK model of the potent MATE1 inhibitor cimetidine was developed. A detailed description of the cimetidine model development and evaluation is given in the ESM. The good model performance is demonstrated by population simulations compared to observed data, a goodness-of-fit plot, and MRD values. Furthermore, predicted AUClast and Cmax values are documented, which are in good agreement with the observed data with GMFEs of 1.14 and 1.17, respectively.
The cimetidine-metformin DDI was modeled as competitive inhibition of OCT1, OCT2, and MATE1 by cimetidine, using inhibition parameters from the literature [30]. However, cimetidine also is a BCS Class III drug (high solubility and low permeability) that is primarily excreted unchanged in the urine (renal clearance of approximately 400 mL/min [31]), indicating an important role of active transport in its distribution and excretion. As the only published information on cimetidine kidney concentrations is a postmortem tissue-to-serum partition coefficient of 14.9 [32], which is not applicable for parameter optimization, the cimetidine interaction parameters were fixed to literature values and one of the cimetidine-metformin DDI studies [33] was utilized to inform the intracellular kidney concentration in the cimetidine model parameter optimization. Population predictions of all clinical cimetidine-metformin DDI studies are presented in Fig. 5a–d. Predicted DDI AUClast and Cmax ratios are close to the observed values, with low GMFEs of 1.22 and 1.20, respectively (see the ESM).
Fig. 5.
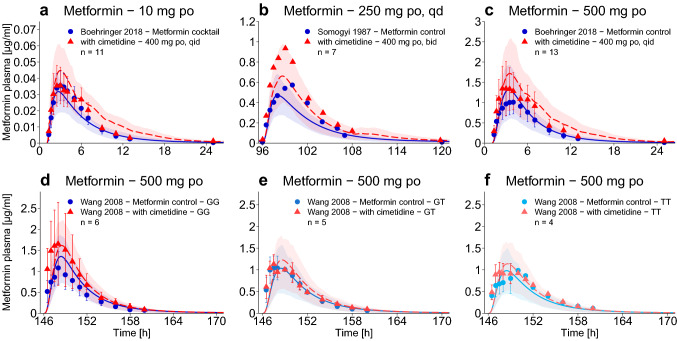
Impact of drug–drug interaction (DDI) and drug–drug–gene interaction (DDGI). Population predictions of metformin plasma concentration–time profiles before and during cimetidine co-treatment of different SLC22A2 genotypes, compared to observed data [21, 23, 33]. Population prediction arithmetic means are shown as solid blue (metformin only) or dashed red (DDI or DDGI) lines; the shaded areas illustrate the respective 68% population prediction intervals. Observed data are shown as dots (metformin only) or triangles (DDI or DDGI) ± standard deviation. Details on the study protocols, semilogarithmic plots, and quantitative DDI and DDGI prediction performance measures are provided in the ESM. bid twice daily, GG SLC22A2 808GG genotype, GT SLC22A2 808GT genotype, po oral, qd once daily, qid four times daily, TT SLC22A2 808TT genotype
In the OCT2 polymorphism study by Wang et al. [21], the different SLC22A2 genotypes were additionally challenged with cimetidine co-administration, to show the combined effects of SLC22A2 808G>T DGI (decreased metformin plasma concentrations) and cimetidine DDI (increased metformin plasma concentrations). The predictions of this drug–drug–gene interaction (DDGI) are presented in Fig. 5e, f. Comparison of the metformin exposure of the three different genotypes during cimetidine treatment (Fig. 5d–f, red triangles) shows that the impact of the polymorphism becomes more pronounced with inhibition of MATE1. Quantitative evaluation of all DDI and DDGI predictions with plots of predicted vs observed AUClast and Cmax ratios are presented in the ESM.
Modeling of Renal Impairment
The impact of renal impairment on metformin pharmacokinetics was modeled by implementation of pathophysiological changes for individuals with a GFR < 60 mL/min (CKD3A-CKD5). First, the actual individual GFR was used as reported. Second, renal secretion through OCT2 and MATE1 was decreased in proportion to the decrease in GFR, according to the “intact nephron hypothesis” [34–36]. Third, as metformin does not bind to plasma proteins, the levels of albumin and alpha-1-acid glycoprotein were not changed, but the hematocrit was gradually decreased with progressing stages of CKD [37]. As observed in previous PBPK analyses of drug pharmacokinetics during renal impairment [36, 38], these changes were not sufficient to describe the high metformin plasma concentrations in patients with CKD, suggesting the inhibition of further elimination pathways by uremic solutes that accumulate during renal impairment.
To incorporate this hypothesis by inhibition of basolateral OCT1 (liver uptake) and PMAT (skeletal muscle uptake), observed data of intravenously administered metformin in CKD3A-5 patients [6] were used to adjust the transport activities of OCT1 and PMAT for the different stages of CKD, yielding a linear correlation between transporter inhibition and GFR. This correlation was implemented and used to predict orally administered metformin in CKD. Furthermore, to capture the broader shape of the metformin plasma concentration–time profiles in patients with CKD, the permeability at the basolateral side of the small intestinal mucosa cells was decreased. This permeability was already adjusted in the model for healthy individuals, to release the metformin from the enterocytes into the blood. Considering the negligible passive permeability of metformin, there are probably transporters involved at this membrane barrier as well, but because of the current lack of information on the identity of such transporters, the local passive permeability was adjusted in the model. Decreasing the basolateral small intestinal permeability in CKD might well be a surrogate for the inhibition of these unknown transporters by accumulating uremic solutes, consistent with their inhibition of basolateral transporters of the liver.
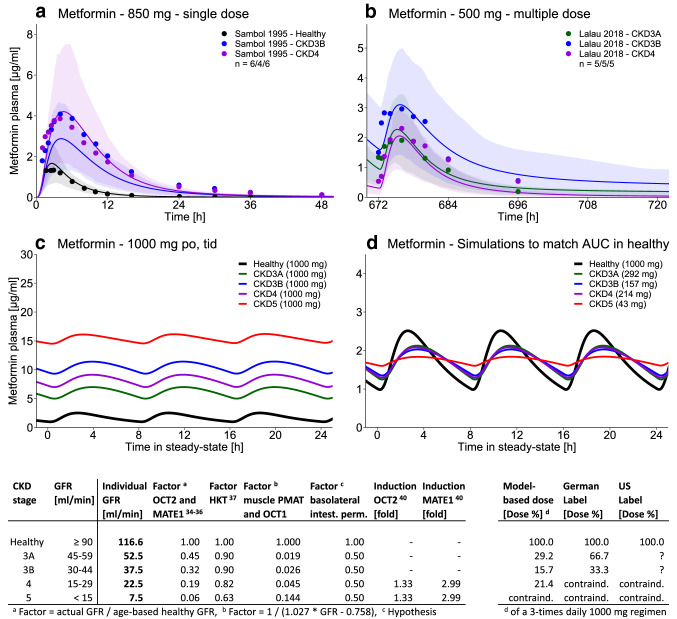
Finally, although one would expect a progressive or even exponential increase of metformin plasma concentrations with decreasing kidney function, no apparent difference in the exposure of CKD3B and CKD4 patients could be observed in the available clinical data [12, 39], indicating adaptive processes in severe renal disease. Therefore, induction of OCT2 and MATE1, as observed in hyperuricemic rats [40], was assumed and incorporated for CKD4-5 patients, greatly improving the predictions in severe renal disease (Fig. 6a, b). These changes in system-dependent parameters to model CKD3A-5 are summarized in Fig. 6 (left table). The model performance for all available clinical studies of metformin in renal disease is documented in the ESM.
Fig. 6.
Impact of renal impairment and model-based chronic kidney disease (CKD) dose recommendations. a, b Population predictions of metformin plasma concentration–time profiles in different stages of CKD, compared to observed data [12, 39]. Population prediction arithmetic means are shown as black (healthy) or colored (CKD3A-4) lines. The shaded areas illustrate the respective 68% population prediction intervals. Observed data are shown as dots in corresponding colors. c Simulations of metformin exposure in CKD3A-5 patients compared to healthy individuals using an oral (po) dose of 1000 mg, three times daily (tid). d Simulations of metformin exposure in CKD3A-5 patients to match the steady-state area under the curve (AUC) of 1000 mg po, tid in healthy individuals. The tables show the implementation of renal impairment (on the left) and the model-based dose recommendations compared to the guidance in the US and German labels (on the right). contraind. contraindicated, GFR glomerular filtration rate, HKT hematocrit, intest. perm. intestinal permeability, MATE multidrug and toxin extrusion protein, OCT organic cation transporter, PMAT plasma membrane monoamine transporter
The developed model of metformin in renal impairment was applied to generate dose recommendations for patients with CKD, that match the steady-state AUC of renally healthy individuals. Simulations of metformin plasma concentrations in CKD3A-5 patients compared to healthy volunteers, all treated with 1000 mg of metformin three times daily, are shown in Fig. 6c. Simulations of metformin in patients with CKD administered with the model-based dose recommendations are shown in Fig. 6d. In the table below, these recommendations are compared to the guidance in the US and German labels. While the US label provides no quantitative advice [13], the German label recommends a reduction to 67% of the dose for CKD3A and to 33% for CKD3B patients [14]. Based on the scarce clinical data of metformin exposure in renally impaired patients, the presented model suggests much lower doses of about 30% for CKD3A and of 20% for CKD3B and 4.
Discussion
A comprehensive whole-body PBPK model of metformin has been thoroughly built and evaluated, integrating the current knowledge on the mechanisms controlling the pharmacokinetics of this widely prescribed drug. The established model has been evaluated for prediction of the effects of the SLC22A2 808G>T polymorphism, the cimetidine-metformin DDI, and the impact of renal impairment, a frequent co-morbidity in patients with T2DM.
Several other PBPK models of metformin have been published previously [41–44], but our newly developed model is the first to integrate PET-measured human in-vivo metformin kidney concentrations, clinical data of microdose studies, and a mechanistic description of the saturable transporter-dependent absorption of metformin. The limitations of the presented model result from our lack of knowledge regarding the metformin pharmacokinetic processes in the liver and the expression levels of the different transporters throughout the body. As shown in Fig. 3d, the liver concentration–time profile following the intravenous 11C-metformin microdose is not adequately described. The uptake of metformin into the liver is modeled via OCT1, but, in accordance with the literature, no process for metformin metabolism or secretion to bile has been implemented. An unspecific hepatic metabolic clearance was tested, but did not improve the model (causing underestimation of the plasma concentrations in studies with therapeutic doses), supporting the idea that metformin is not metabolized. The plasma concentrations following the oral 11C-metformin microdose, and consequently also the measured tissue concentrations, are underpredicted for the 2 h of the oral PET study (see the ESM). However, the administered microdoses (1.445 µg intravenously and 0.856 µg orally) were more than 300,000 and 500,000 times below the lowest therapeutic dose of 500 mg. Given that the metformin pharmacokinetics are completely governed by saturable transport processes and that the plasma, whole blood, kidney, and muscle concentrations following the intravenous microdose are well described, this underprediction might be caused by a missing process for metformin absorption.
The effect of the SLC22A2 808G>T polymorphism is difficult to assess from the literature. In-vitro studies report a decreased metformin transport rate [29], equal activity [20], as well as increased transport velocity [17] for the variant OCT2 protein. In-vivo, two studies report decreased clearance by renal secretion in Korean and Chinese 808TT individuals [21, 29], whereof the Chinese study nevertheless shows a non-significantly lower metformin plasma Cmax for the 808TT group. However, two different studies report increased clearance by renal secretion in American and European individuals [17, 18], with corresponding decreases in metformin exposure in association with the minor allele. Given that OCT2 and MATE1 are working sequentially to transport metformin through the kidney, it is difficult to distinguish their impacts on renal secretion. Therefore, statements regarding the effect of polymorphisms or co-medications on OCT2 function should not be based on plasma concentrations or renal secretion alone, without concomitant assessment of MATE1 genotype/activity or kidney concentrations. This also holds true for the DDGI results by Wang et al. [21] (Fig. 5f), where the observed lack of cimetidine-metformin DDI in SLC22A2 808TT individuals is difficult to explain, because (1) this DDI is mainly caused by inhibition of MATE1, (2) the MATE1 genotypes were not analyzed in this study, and (3) so far there are no in-vitro results available on the impact of cimetidine on MATE1 variants. Another explanation for the weak effect of cimetidine in the SLC22A2 808TT group might be reduced transport of cimetidine by this OCT2 variant into the kidney and therefore less inhibition of MATE1, as previously proposed [30].
To model the renal impairment, renal secretion was decreased in proportion to the impaired GFR, based on the “intact nephron hypothesis”, which postulates that structurally damaged nephrons stop contributing to both passive renal filtration and active secretion, and that the remaining intact nephrons continue to function in glomerulo-tubular balance with appropriate adaptation to the patient’s needs [35]. This hypothesis has been successfully applied in previous PBPK analyses of renal impairment [36, 42, 45, 46]. The inhibition of liver drug uptake by uremic toxins in renal impairment has been postulated by Zhao et al. [38], based on the fact that the clearance of many nonrenally eliminated drugs is decreased in CKD, and based on their PBPK analysis of repaglinide in CKD4.
We used an empirical approach to model the inhibition of liver and muscle uptake as a function of the degree of renal impairment. The inhibition of the basolateral intestinal permeability/transport in CKD is purely hypothetical, but was essential to describe the shape and elimination phase of the clinically observed data. The induction of OCT2 and MATE1 was demonstrated in hyperuricemic rats [40]. To confirm and refine these hypotheses, in-vitro studies of OCT1 and PMAT inhibition by uremic solutes are needed, to identify the toxins involved and to assess their inhibitory potential; the expression and role of transporters at the basolateral membrane of the intestinal mucosa has to be investigated; and the clinical relevance of OCT2 and MATE1 induction by uric acid in humans needs to be established.
Future applications include the modeling of further DGIs and DDIs, and ultimately, the individualized dose recommendation for real patients with multiple polymorphisms, co-medications, and co-morbidities. Although the effects of some of these interactions do not reach statistical significance in the blood, their impact on kidney or liver concentrations might well be substantial and of therapeutic relevance. The model application with the most immediate medical benefit is the generation of dose recommendations for renally impaired patients with T2DM. Chronic kidney disease is a frequent co-morbidity, but physicians are reluctant to prescribe metformin to patients with reduced renal function because of the contraindication given in most guidelines and fear of lactic acidosis caused by metformin accumulation [47]. These contraindications are based solely on the estimated GFR of the patient, even though for patients with stable renal disease, a dose adjustment based on renal function, with monitoring of the metformin plasma concentrations, would be perfectly feasible. A reduced dose of 500 mg metformin daily was reported to be safe for creatinine clearances as low as 20 mL/min [48] and in a group of CKD4 patients [12], which is in line with the presented model-based recommendation of 200 mg three times daily for CKD4 patients with T2DM.
Conclusions
Mechanistic whole-body PBPK models of metformin and cimetidine have been carefully developed and evaluated to integrate the current pharmacokinetic knowledge on these drugs and to describe the impact of the SLC22A2 808G>T polymorphism, the cimetidine-metformin DDI, and the pathophysiological changes during renal impairment on the exposure of metformin. Both models will be released open-source (https://www.open-systems-pharmacology.org) [49], to support metformin therapy, OCT2/MATE DDI studies during drug development, and to be used as input for pharmacodynamic glucose-homeostasis models [50, 51] and other PBPK/pharmacodynamic analyses. The presented analysis has generated insights into the pharmacokinetics during renal impairment, indicating that the kidneys of patients with severe renal disease might be able to adapt to uremia/hyperuricemia by induction of OCT2 and MATE1, as has been shown for hyperuricemic rats [40].
Electronic supplementary material
Below is the link to the electronic supplementary material.
Acknowledgements
Open Access funding provided by Projekt DEAL.
Compliance with Ethical Standards
Funding
This project has received funding from Boehringer Ingelheim Pharma GmbH & Co. KG and from the German Federal Ministry of Education and Research “NanoCare4.0 – Anwendungssichere Materialinnovationen” Program (BMBF Grant 03XP0196).
Conflict of Interest
Naoki Ishiguro, Thomas Ebner, Sabrina Wiebe, Fabian Müller, Peter Stopfer, and Valerie Nock are employees of Boehringer Ingelheim Pharma GmbH & Co. KG. Thorsten Lehr has received research grants from Boehringer Ingelheim Pharma GmbH & Co. KG and from the German Federal Ministry of Education and Research. Nina Hanke, Denise Türk, and Dominik Selzer have no conflicts of interest that are directly relevant to the content of this article.
Ethics Approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Consent to Participate
Informed consent was obtained from all individual participants included in the studies.
References
- 1.ClinCalc LLC. ClinCalc DrugStats database. 2019. Available from: https://clincalc.com/DrugStats/. Accessed 20 Nov 2019.
- 2.Pentikäinen PJ, Neuvonen PJ, Penttilä A. Pharmacokinetics of metformin after intravenous and oral administration to man. Eur J Clin Pharmacol. 1979;16:195–202. doi: 10.1007/bf00562061. [DOI] [PubMed] [Google Scholar]
- 3.Vidon N, Chaussade S, Noel M, Franchisseur C, Huchet B, Bernier JJ. Metformin in the digestive tract. Diabetes Res Clin Pract. 1988;4:223–229. doi: 10.1016/s0168-8227(88)80022-6. [DOI] [PubMed] [Google Scholar]
- 4.Tucker GT, Casey C, Phillips PJ, Connor H, Ward JD, Woods HF. Metformin kinetics in healthy subjects and in patients with diabetes mellitus. Br J Clin Pharmacol. 1981;12:235–246. doi: 10.1111/j.1365-2125.1981.tb01206.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Sambol NC, Chiang J, O’Conner M, Liu CY, Lin ET, Goodman AM, et al. Pharmacokinetics and pharmacodynamics of metformin in healthy subjects and patients with noninsulin-dependent diabetes mellitus. J Clin Pharmacol. 1996;36:1012–1021. doi: 10.1177/009127009603601105. [DOI] [PubMed] [Google Scholar]
- 6.Sirtori CR, Franceschini G, Galli-Kienle M, Cighetti G, Galli G, Bondioli A, et al. Disposition of metformin (N, N-dimethylbiguanide) in man. Clin Pharmacol Ther. 1978;24:683–693. doi: 10.1002/cpt1978246683. [DOI] [PubMed] [Google Scholar]
- 7.Gormsen LC, Sundelin EI, Jensen JB, Vendelbo MH, Jakobsen S, Munk OL, et al. In vivo imaging of human 11C-metformin in peripheral organs: dosimetry, biodistribution, and kinetic analyses. J Nucl Med. 2016;57:1920–1926. doi: 10.2967/jnumed.116.177774. [DOI] [PubMed] [Google Scholar]
- 8.Otsuka M, Matsumoto T, Morimoto R, Arioka S, Omote H, Moriyama Y. A human transporter protein that mediates the final excretion step for toxic organic cations. Proc Natl Acad Sci USA. 2005;102:17923–17928. doi: 10.1073/pnas.0506483102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Masuda S, Terada T, Yonezawa A, Tanihara Y, Kishimoto K, Katsura T, et al. Identification and functional characterization of a new human kidney-specific H+/organic cation antiporter, kidney-specific multidrug and toxin extrusion 2. J Am Soc Nephrol. 2006;17:2127–2135. doi: 10.1681/ASN.2006030205. [DOI] [PubMed] [Google Scholar]
- 10.Prasad B, Johnson K, Billington S, Lee C, Chung GW, Brown CDA, et al. Abundance of drug transporters in the human kidney cortex as quantified by quantitative targeted proteomics. Drug Metab Dispos. 2016;44:1920–1924. doi: 10.1124/dmd.116.072066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Graham GG, Punt J, Arora M, Day RO, Doogue MP, Duong JK, et al. Clinical pharmacokinetics of metformin. Clin Pharmacokinet. 2011;50:81–98. doi: 10.2165/11534750-000000000-00000. [DOI] [PubMed] [Google Scholar]
- 12.Lalau J-D, Kajbaf F, Bennis Y, Hurtel-Lemaire A-S, Belpaire F, De Broe ME. Metformin treatment in patients with type 2 diabetes and chronic kidney disease stages 3A, 3B, or 4. Diabetes Care. 2018;41:547–553. doi: 10.2337/dc17-2231. [DOI] [PubMed] [Google Scholar]
- 13.Bristol-Myers Squibb Company. Prescribing information: Glucophage® (metformin hydrochloride) tablets. 2017. https://www.accessdata.fda.gov/drugsatfda_docs/label/2017/020357s037s039,021202s021s023lbl.pdf. Accessed 20 Nov 2019.
- 14.Heumann Pharma GmbH & Co. Generica KG. Prescribing information: metformin Heumann. 2017. Available from: https://www.heumann.de/fileadmin/user_upload/produkte/infos/Fachinformation-Metformin-Heumann.pdf. Accessed 20 Nov 2019.
- 15.Christensen MMH, Brasch-Andersen C, Green H, Nielsen F, Damkier P, Beck-Nielsen H, et al. The pharmacogenetics of metformin and its impact on plasma metformin steady-state levels and glycosylated hemoglobin A1c. Pharmacogenet Genomics. 2011;21:837–850. doi: 10.1097/FPC.0b013e32834c0010. [DOI] [PubMed] [Google Scholar]
- 16.Duong JK, Kumar SS, Kirkpatrick CM, Greenup LC, Arora M, Lee TC, et al. Population pharmacokinetics of metformin in healthy subjects and patients with type 2 diabetes mellitus: simulation of doses according to renal function. Clin Pharmacokinet. 2013;52:373–384. doi: 10.1007/s40262-013-0046-9. [DOI] [PubMed] [Google Scholar]
- 17.Chen Y, Li S, Brown C, Cheatham S, Castro RA, Leabman MK, et al. Effect of genetic variation in the organic cation transporter 2 on the renal elimination of metformin. Pharmacogenet Genomics. 2009;19:497–504. doi: 10.1097/FPC.0b013e32832cc7e9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Christensen MMH, Pedersen RS, Stage TB, Brasch-Andersen C, Nielsen F, Damkier P, et al. A gene-gene interaction between polymorphisms in the OCT2 and MATE1 genes influences the renal clearance of metformin. Pharmacogenet Genomics. 2013;23:526–534. doi: 10.1097/FPC.0b013e328364a57d. [DOI] [PubMed] [Google Scholar]
- 19.Stocker SL, Morrissey KM, Yee SW, Castro RA, Xu L, Dahlin A, et al. The effect of novel promoter variants in MATE1 and MATE2 on the pharmacokinetics and pharmacodynamics of metformin. Clin Pharmacol Ther. 2013;93:186–194. doi: 10.1038/clpt.2012.210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Zolk O, Solbach TF, König J, Fromm MF. Functional characterization of the human organic cation transporter 2 variant p270Ala%3eSer. Drug Metab Dispos. 2009;37:1312–1318. doi: 10.1124/dmd.108.023762. [DOI] [PubMed] [Google Scholar]
- 21.Wang Z-J, Yin OQP, Tomlinson B, Chow MSS. OCT2 polymorphisms and in-vivo renal functional consequence: studies with metformin and cimetidine. Pharmacogenet Genomics. 2008;18:637–645. doi: 10.1097/FPC.0b013e328302cd41. [DOI] [PubMed] [Google Scholar]
- 22.Drugs.com. Drug interactions checker. 2019. Available from: https://www.drugs.com/drug-interactions/metformin.html. Accessed 20 Nov 2019.
- 23.Somogyi A, Stockley C, Keal J, Rolan P, Bochner F. Reduction of metformin renal tubular secretion by cimetidine in man. Br J Clin Pharmacol. 1987;23:545–551. doi: 10.1111/j.1365-2125.1987.tb03090.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.US Food and Drug Administration. Drug development and drug interactions: table of substrates, inhibitors and inducers. 2019. Available from: https://www.fda.gov/drugs/drug-interactions-labeling/drug-development-and-drug-interactions-table-substrates-inhibitors-and-inducers. Accessed 15 Nov 2019.
- 25.Zhou M, Xia L, Wang J. Metformin transport by a newly cloned proton-stimulated organic cation transporter (plasma membrane monoamine transporter) expressed in human intestine. Drug Metab Dispos. 2007;35:1956–1962. doi: 10.1124/dmd.107.015495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Han TK, Proctor WR, Costales CL, Cai H, Everett RS, Thakker DR. Four cation-selective transporters contribute to apical uptake and accumulation of metformin in Caco-2 cell monolayers. J Pharmacol Exp Ther. 2015;352:519–528. doi: 10.1124/jpet.114.220350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Liang X, Chien H-C, Yee SW, Giacomini MM, Chen EC, Piao M, et al. Metformin is a substrate and inhibitor of the human thiamine transporter, THTR-2 (SLC19A3) Mol Pharm. 2015;12:4301–4310. doi: 10.1021/acs.molpharmaceut.5b00501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Liang X, Giacomini KM. Transporters involved in metformin pharmacokinetics and treatment response. J Pharm Sci. 2017;106:2245–2250. doi: 10.1016/j.xphs.2017.04.078. [DOI] [PubMed] [Google Scholar]
- 29.Song IS, Shin HJ, Shim EJ, Jung IS, Kim WY, Shon JH, et al. Genetic variants of the organic cation transporter 2 influence the disposition of metformin. Clin Pharmacol Ther. 2008;84:559–562. doi: 10.1038/clpt.2008.61. [DOI] [PubMed] [Google Scholar]
- 30.Ito S, Kusuhara H, Yokochi M, Toyoshima J, Inoue K, Yuasa H, et al. Competitive inhibition of the luminal efflux by multidrug and toxin extrusions, but not basolateral uptake by organic cation transporter 2, is the likely mechanism underlying the pharmacokinetic drug-drug interactions caused by cimetidine in the kidney. J Pharmacol Exp Ther. 2012;340:393–403. doi: 10.1124/jpet.111.184986. [DOI] [PubMed] [Google Scholar]
- 31.Somogyi A, Gugler R. Clinical pharmacokinetics of cimetidine. Clin Pharmacokinet. 1983;8:463–495. doi: 10.2165/00003088-198308060-00001. [DOI] [PubMed] [Google Scholar]
- 32.Schentag JJ, Cerra FB, Calleri GM, Leising ME, French MA, Bernhard H. Age, disease, and cimetidine disposition in healthy subjects and chronically ill patients. Clin Pharmacol Ther. 1981;29:737–743. doi: 10.1038/clpt.1981.104. [DOI] [PubMed] [Google Scholar]
- 33.Boehringer Ingelheim Pharma GmbH & Co. KG. The effect of potent inhibitors of drug transporters (verapamil, rifampin, cimetidine, probenecid) on pharmacokinetics of a transporter probe drug cocktail consisting of digoxin, furosemide, metformin and rosuvastatin. EudraCT 2017-001549-29. 2018. Available from: https://clinicaltrials.gov/ct2/show/record/NCT03307252. Accessed 20 Nov 2019.
- 34.Bricker NS, Morrin PAF, Kime SW. The pathologic physiology of chronic Bright’s disease: an exposition of the “intact nephron hypothesis”. Am J Med. 1960;28:77–97. doi: 10.1016/0002-9343(60)90225-4. [DOI] [PubMed] [Google Scholar]
- 35.Bricker NS. On the meaning of the intact nephron hypothesis. Am J Med. 1969;46:1–11. doi: 10.1016/0002-9343(69)90053-9. [DOI] [PubMed] [Google Scholar]
- 36.Hsueh C-H, Hsu V, Zhao P, Zhang L, Giacomini KM, Huang S-M. PBPK Modeling of the effect of reduced kidney function on the pharmacokinetics of drugs excreted renally by organic anion transporters. Clin Pharmacol Ther. 2018;103:485–492. doi: 10.1002/cpt.750. [DOI] [PubMed] [Google Scholar]
- 37.Schmulenson E, Schlender J-F, Frechen S, Jaehde U. A physiologically-based pharmacokinetic modeling approach to assess the impact of chronic kidney disease. Annual Meeting of Population Approach Group Europe (PAGE); 2018; Montreux, Switzerland: abstract 8630. Available from: https://www.page-meeting.org/?abstract=8630. Accessed 30 Apr 2020.
- 38.Zhao P, Vieira MLT, Grillo JA, Song P, Wu T-C, Zheng JH, et al. Evaluation of exposure change of nonrenally eliminated drugs in patients with chronic kidney disease using physiologically based pharmacokinetic modeling and simulation. J Clin Pharmacol. 2012;52:91–108. doi: 10.1177/0091270011415528. [DOI] [PubMed] [Google Scholar]
- 39.Sambol NC, Chiang J, Lin ET, Goodman AM, Liu CY, Benet LZ, et al. Kidney function and age are both predictors of pharmacokinetics of metformin. J Clin Pharmacol. 1995;35:1094–1102. doi: 10.1002/j.1552-4604.1995.tb04033.x. [DOI] [PubMed] [Google Scholar]
- 40.Zhang G, Ma Y, Xi D, Rao Z, Sun X, Wu X. Effect of high uric acid on the disposition of metformin: in vivo and in vitro studies. Biopharm Drug Dispos. 2019;40:3–11. doi: 10.1002/bdd.2164. [DOI] [PubMed] [Google Scholar]
- 41.Xia B, Heimbach T, Gollen R, Nanavati C, He H. A simplified PBPK modeling approach for prediction of pharmacokinetics of four primarily renally excreted and CYP3A metabolized compounds during pregnancy. AAPS J. 2013;15:1012–1024. doi: 10.1208/s12248-013-9505-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Li J, Guo H-F, Liu C, Zhong Z, Liu L, Liu X-D. Prediction of drug disposition in diabetic patients by means of a physiologically based pharmacokinetic model. Clin Pharmacokinet. 2015;54:179–193. doi: 10.1007/s40262-014-0192-8. [DOI] [PubMed] [Google Scholar]
- 43.Burt HJ, Neuhoff S, Almond L, Gaohua L, Harwood MD, Jamei M, et al. Metformin and cimetidine: physiologically based pharmacokinetic modelling to investigate transporter mediated drug-drug interactions. Eur J Pharm Sci. 2016;88:70–82. doi: 10.1016/j.ejps.2016.03.020. [DOI] [PubMed] [Google Scholar]
- 44.Nishiyama K, Toshimoto K, Lee W, Ishiguro N, Bister B, Sugiyama Y. Physiologically-based pharmacokinetic modeling analysis for quantitative prediction of renal transporter-mediated interactions between metformin and cimetidine. CPT Pharmacometr Syst Pharmacol. 2019;8:396–406. doi: 10.1002/psp4.12398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Li G, Wang K, Chen R, Zhao H, Yang J, Zheng Q. Simulation of the pharmacokinetics of bisoprolol in healthy adults and patients with impaired renal function using whole-body physiologically based pharmacokinetic modeling. Acta Pharmacol Sin. 2012;33:1359–1371. doi: 10.1038/aps.2012.103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Posada MM, Bacon JA, Schneck KB, Tirona RG, Kim RB, Higgins JW, et al. Prediction of renal transporter mediated drug-drug interactions for pemetrexed using physiologically based pharmacokinetic modeling. Drug Metab Dispos. 2015;43:325–334. doi: 10.1124/dmd.114.059618. [DOI] [PubMed] [Google Scholar]
- 47.Kajbaf F, Arnouts P, de Broe M, Lalau J-D. Metformin therapy and kidney disease: a review of guidelines and proposals for metformin withdrawal around the world. Pharmacoepidemiol Drug Saf. 2013;22:1027–1035. doi: 10.1002/pds.3501. [DOI] [PubMed] [Google Scholar]
- 48.Duong JK, Roberts DM, Furlong TJ, Kumar SS, Greenfield JR, Kirkpatrick CM, et al. Metformin therapy in patients with chronic kidney disease. Diabetes Obes Metab. 2012;14:963–965. doi: 10.1111/j.1463-1326.2012.01617.x. [DOI] [PubMed] [Google Scholar]
- 49.Lippert J, Burghaus R, Edginton A, Frechen S, Karlsson M, Kovar A, et al. Open systems pharmacology community: an open access, open source, open science approach to modeling and simulation in pharmaceutical sciences. CPT Pharmacometrics Syst Pharmacol. 2019;8:878–882. doi: 10.1002/psp4.12473. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Schaller S, Willmann S, Lippert J, Schaupp L, Pieber TR, Schuppert A, et al. A generic integrated physiologically based whole-body model of the glucose-insulin-glucagon regulatory system. CPT Pharmacometrics Syst Pharmacol. 2013;2:e65. doi: 10.1038/psp.2013.40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Balazki P, Schaller S, Eissing T, Lehr T. A quantitative systems pharmacology kidney model of diabetes associated renal hyperfiltration and the effects of SGLT inhibitors. CPT Pharmacometrics Syst Pharmacol. 2018;7:788–797. doi: 10.1002/psp4.12359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Stopfer P, Giessmann T, Hohl K, Hutzel S, Schmidt S, Gansser D, et al. Optimization of a drug transporter probe cocktail: potential screening tool for transporter-mediated drug-drug interactions. Br J Clin Pharmacol. 2018;84:1941–1949. doi: 10.1111/bcp.13609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Di Cicco RA, Allen A, Carr A, Fowles S, Jorkasky DK, Freed MI. Rosiglitazone does not alter the pharmacokinetics of metformin. J Clin Pharmacol. 2000;40:1280–1285. [PubMed] [Google Scholar]
- 54.Gan L, Jiang X, Mendonza A, Swan T, Reynolds C, Nguyen J, et al. Pharmacokinetic drug-drug interaction assessment of LCZ696 (an angiotensin receptor neprilysin inhibitor) with omeprazole, metformin or levonorgestrel-ethinyl estradiol in healthy subjects. Clin Pharmacol Drug Dev. 2016;5:27–39. doi: 10.1002/cpdd.181. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.