To the editor:
A reliable, simple and safe sampling method applicable to a wide-age-range is required for community-based SARS-CoV-2 surveillance. Nasopharyngeal/oropharyngeal specimens require trained-personnel to perform and trigger sneezing, and coughing, which pose a risk of disease transmission.1 Self-administered sampling methods2, 3, 4 have been assessed in adult in-patients, but these methods do not translate well to the community setting, e.g., saliva collection is difficult in young children and the elderly, and variably reduces test sensitivity.5 A recent study published in the Journal of Infection reviewed the methodologies used in the estimation of diagnostic accuracy of SARS-CoV-2 real-time reverse transcription polymerase chain reaction (RT-PCR) and other nucleic acid amplification tests for COVID-19 and pointed out the importance in employing standardized guidelines for study designs and statistical methods.6
Here, we compared different sample collection methods and introduced nasal strip as a sensitive and low-risk collection method and assessed its application in both paediatric and adult subjects at the Prince of Wales Hospital, Hong Kong. Thirty-eight asymptomatic and symptomatic subjects hospitalized with COVID-19 were recruited prospectively by convenience sampling. The disease status was confirmed by two RT-PCR tests targeting different regions of the RdRp gene performed by the local hospital and Public Health Laboratory Service. Twenty infected adults (range: 22–74 years old) and eighteen children/adolescents (range: 6–17 years old) were recruited of whom ten were asymptomatic. Adult subjects or guardians of participants below 18 years old provided informed consent (see the Methods section in the Supplmentary Appendix).
We obtained nasal epithelial lining fluid (NELF) by nasal strip (n = 43), to compare against pooled nasopharyngeal and throat swabs (NPSTS) (n = 21) or deep throat saliva (DTS) (n = 22) collected within 24 h of the nasal strip. 13 paired nasal swabs were also collected right before the collection of nasal strip to evaluate their SARS-CoV-2 detection performance. All samples were subjected to viral RNA quantitation by real-time PCR targeting the nucleoprotein gene.7
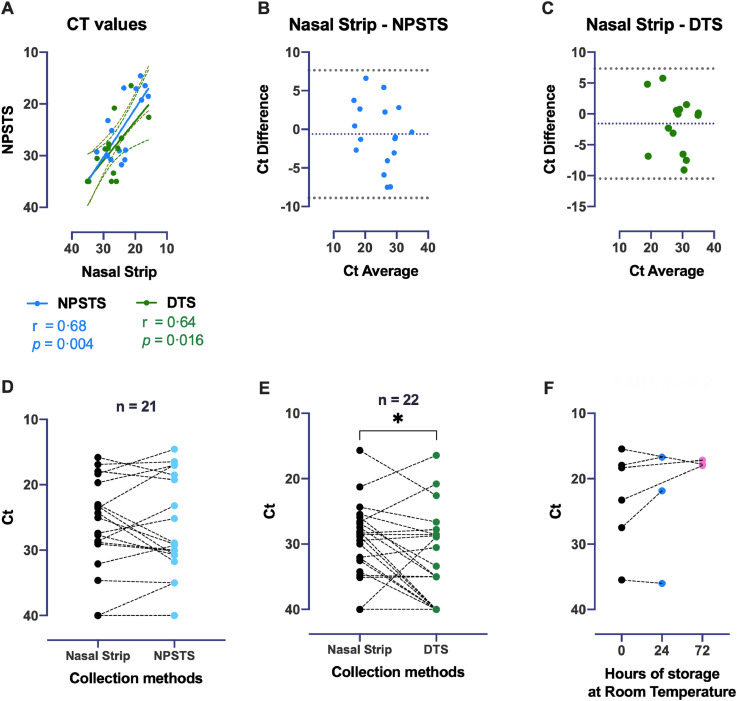
Spearman's test demonstrated significant correlation between nasal strip and NPSTS (p = 0.0003) and between nasal strip and DTS (p = 0.01) (Fig. 1 A). The agreement between nasal strip samples and NPSTS was 94.44% (17/18) and 100% (3/3) for NPSTS positive and negative samples (Table 1 , Fig. 1B). In contrast, the agreement between nasal strip specimens and DTS was 93.33% (14/15) and 14.29% (1/7) for DTS positive and negative samples, respectively (Table 1, Fig. 1C). Eight discrepant samples were identified (Table S1, Figure S1) of which seven were DTS specimens. Nasal strip outperformed DTS on six occasions, where negative result was reported in the latter. Four of these DTS specimens were collected from paediatric patients (Patients 1 to 4). Nasal strip samples were tested negative on two occasions when the reference test revealed Ct values of 35 and 28.92 (Figure S1). Wilcoxon signed rank test revealed that nasal strip and NPSTS gave similar Ct values (Fig. 1D, p = 0.76) while a lower Ct was detected in nasal strip compared to paired DTS (Fig. 1E, p = 0.016).
Fig. 1.
Correlation, agreement and comparison of the cycle threshold (Ct) values from nasal strip, NPSTS and DTS, and the stability of nasal strip sample at room temperature. The correlation coefficients of NPSTS and DTS are superimposed on the panel with trend lines estimated using simple linear regression (Panel A). Plot shows the available Ct values of 31 samples which had positive test results from both tests. Data on three samples with negative result in both nasal strip and NPSTS, one sample with negative result in nasal strip but a positive result in NPSTS (Ct value = 35), one sample with negative result in both nasal strip and DTS, one sample with negative result from nasal strip but positive result in DTS, and six samples with positive result in nasal strip but negative result DTS were excluded from the Spearman correlation analysis. Bland-Altman Plots indicate the agreement of nasal strip versus NPSTS (Panel B) and DTS (Panel C), respectively. The differences between the two measurements are plotted against their average Ct values. Almost all observations are located within 2 standard deviations of the mean difference, and no bias is shown. The plots show that the nasal strip gives consistent and comparable measurements versus the NPSTS and DTS. SARS-CoV-2 RNA load in nasal strip and NPSTS (n = 21) (Panel D) and DTS (n = 22) (Panel E). Samples were obtained from 36 in-patients who had a diagnosis of COVID‐19. Panel A shows SARS‐CoV‐2 RNA Ct in the nasal strip and NPSTS; panel B shows SARS‐CoV‐2 RNA Ct in the nasal strip and DTS. The lines indicate samples from the same patient obtained within 24 hours. Negative result is arbitrarily set as Ct = 40 and results were compared with the use of a Wilcoxon signed‐rank test (p < 0.05). Panel F shows the stability of nasal strip samples for the detection of SARS-CoV-2 (n = 6). Comparison of Ct upon 24 (blue) and 72 (pink) hours RT storage from nasal strips directly lysed after sample collection.
Table 1.
Number of tested samples and performance of the collection methods
| Nasal strip | |||||
|---|---|---|---|---|---|
| Reference | Positive | Negative | Total | Agreement (%) | |
| NPSTS (n = 21) | Positive | 17 | 1 | 18 | 94.44 |
| Negative | 0 | 3 | 3 | 100.00 | |
| Value (95% CI) | |||||
| Accuracy | 95.2 (76.18–99.88) | ||||
| DTS (n = 22) | Positive | 14 | 1 | 15 | 93.33 |
| Negative | 6 | 1 | 7 | 14.29 | |
| Value (95% CI) | |||||
| Accuracy | 68.18 (45.13–86.14) | ||||
No significant difference in the detection rate was observed between NPSTS (McNemar's test p = 1.000) or DTS (p = 0.13) for the qPCR test (NPSTS and DTS, p = 0.29). CT value equal or below 35 is defined as positive.
Of the 43 nasal strips collected, 13 were paired with a nasal swab sample obtained concurrently by a healthcare worker. A significant correlation was found between Ct values from the nasal strip and nasal swab specimens (r = 0.88, p = 0.0031, Figure S2A). Though nasal swab missed two positive cases detected by nasal strip and nasal strip missed one positive case detected by nasal swab, there was no significant difference detected between Ct values of the 13 paired samples (Figure S2B).
Finally, we collected nasal strip pairs from six patients to determine viral stability over time, viral RNA remained detectable after 24- and 72 h storage at room temperature (Fig. 1F).
The high correlation of nasal strip samples with the standard sampling methods is likely the result of steady NELF absorption with the strip in close contact with the nasal mucosa which reduces sample variability. This study also indicated the possible insensitivity of DTS, particularly in paediatric patients who are less able to provide DTS with consistent quality (Table S1) and how nasal strip would be a superior tool for surveillance of paediatric populations. Nasal strip is also a better collection method than NPSTS as it is less traumatic and irritating. The application of nasal strip reduces the risk of any sneezes and coughs and therefore lessens the risk of virus transmission. Nasal strip is a more comfortable and easier to apply sampling method than the other available standard sampling tools. Repeat nasal strip sampling as part of a community-based surveillance program is feasible in children and adults and likely to succeed as a result of its non-invasive nature (Video 1).
Compared with NPSTS, nasal strip sampling achieved an accuracy of 95.2% (Table 1). Nasal strip sample is comparable if not superior to other sampling methods reported in the literature, including self-administered tongue, lower- and mid-nasal specimens.4 Apart from its good accuracy, we assessed the validity of the nasal strip samples after prolonged room temperature storage so as to mimic the duration needed to post the specimens to the laboratory. This aspect was not assessed in previous studies, albeit an important criterion if a sampling method is adopted for community-based testing purposes. Our findings suggest that nasal strip would provide at least consistent qualitative results (positive or negative), as long as the Ct value is within the range of an inferred infectivity.8 This would be sufficient to identify potentially infectious individuals and susceptible contacts for further management and quarantine.
There are several limitations in this study. This prospective study presents the cross-sectional data performed in a single hospital. The clinical sample pairs (n = 6) that underwent 24- to 72 h room temperature storage remained stable in terms of viral detection. However, the involvement of protease and RNase activity of individual subjects and its contribution to sample stability has not been fully elucidated. The current method provides detection of SARS-CoV-2 at the gene level but no information was obtained regarding the infectious titer.
Conclusion
Our nasal strip collection method serves as an excellent sampling method with comparable performance with NPSTS, DTS and nasal swab specimens in identifying subjects infected with SARS-CoV-2. This reliable, non-invasive, self-administered method with its extended sample stability makes it uniquely suited for repeated sampling and large-scale community study, especially for paediatric population.
Acknowledgments
We would like to acknowledge Prof Aaron HP Ho, Prof Megan YP Ho and Miss Yuan-yuan Wei (Department of Biomedical Engineering, Faculty of Engineering, The Chinese University of Hong Kong) who tailor-cut the nasal strips for this study and Dr KP Tao and Ms Fiona Cheng (Department of Paediatrics, The Chinese University of Hong Kong) for his technical support in molecular biology and her assistance in preparing all the sampling kits, respectively. We would like to offer our special thanks to Mr. Ernie Lee, Mr. Johnson Chan and Mr. Reneson Chan in contributing consecutive NELF samples during the testing stage and all the subjects who agreed to participate in this study. We thank Prof Ellis KL Hon (Department of Paediatrics, The Chinese University of Hong Kong) for his continuous encouragement to the research team. This study is supported by Health and Medical Research Fund commissioned grants (Ref: COVID190112 to RWYC; COVID190103 to MHW; COVID190107 to PKSC); The Chinese Univeristy of Hong Kong Project Impact Enhancement Fund (Ref: PIEF/Ph2/COVID/06) to MHW and Hong Kong Institute of Allergy Research Grant 2020 to RWYC.
Footnotes
Supplementary material associated with this article can be found, in the online version, at doi:10.1016/j.jinf.2020.11.009.
Appendix. Supplementary materials
Video 1. Illustration of the self-administering procedures to collect the NELF sample using nasal strip in child. https://youtu.be/qCplX-ounPA.
References
- 1.U.S. Food & Drug Administration. Recommendations on Providing Clear Instructions to Patients Who Self-Collect an Anterior Nares (Nasal) Sample in a Health Care Setting for SARS-CoV-2 Testing - Letter to Health Care Providers. In: Administration USFD, ed. 10/07/2020 ed 2020.
- 2.Wyllie AL, Fournier J, Casanovas-Massana A. Saliva or nasopharyngeal swab specimens for detection of SARS-CoV-2. N Engl J Med. 2020;383:1283–1286. doi: 10.1056/NEJMc2016359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Pere H, Podglajen I, Wack M. Nasal swab sampling for SARS-CoV-2: a convenient alternative in times of nasopharyngeal swab shortage. J Clin Microbiol. 2020:58. doi: 10.1128/JCM.00721-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Tu YP, Jennings R, Hart B. Swabs collected by patients or health care workers for SARS-CoV-2 testing. N Engl J Med. 2020;383:494–496. doi: 10.1056/NEJMc2016321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Lai CKC, Chen Z, Lui G. Prospective study comparing deep-throat saliva with other respiratory tract specimens in the diagnosis of novel coronavirus disease (COVID-19) J Infect Dis. 2020 doi: 10.1093/infdis/jiaa487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Axell-House DB, Lavingia R, Rafferty M, Clark E, Amirian ES, Chiao EY. The estimation of diagnostic accuracy of tests for COVID-19: A scoping review. J Infect. 2020;81:681–697. doi: 10.1016/j.jinf.2020.08.043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Chu DKW, Pan Y, Cheng SMS. Molecular diagnosis of a novel coronavirus (2019-nCoV) causing an outbreak of Pneumonia. Clin Chem. 2020;66:549–555. doi: 10.1093/clinchem/hvaa029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Perera R, Tso E, Tsang OTY. SARS-CoV-2 virus culture and subgenomic RNA for respiratory specimens from patients with mild coronavirus disease. Emerg Infect Dis. 2020:26. doi: 10.3201/eid2611.203219. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Video 1. Illustration of the self-administering procedures to collect the NELF sample using nasal strip in child. https://youtu.be/qCplX-ounPA.