Abstract
Caulimoviridae is a family of non-enveloped reverse-transcribing plant viruses with non-covalently closed circular dsDNA genomes of 7.1–9.8 kbp in the order Ortervirales. They infect a wide range of monocots and dicots. Some viruses cause economically important diseases of tropical and subtropical crops. Transmission occurs through insect vectors (aphids, mealybugs, leafhoppers, lace bugs) and grafting. Activation of infectious endogenous viral elements occurs in Musa balbisiana, Petunia hybrida and Nicotiana edwardsonii. However, most endogenous caulimovirids are not infectious. This is a summary of the International Committee on Taxonomy of Viruses (ICTV) Report on the family Caulimoviridae, which is available at ictv.global/report/caulimoviridae.
Keywords: Caulimoviridae, ICTV Report, taxonomy
Virion
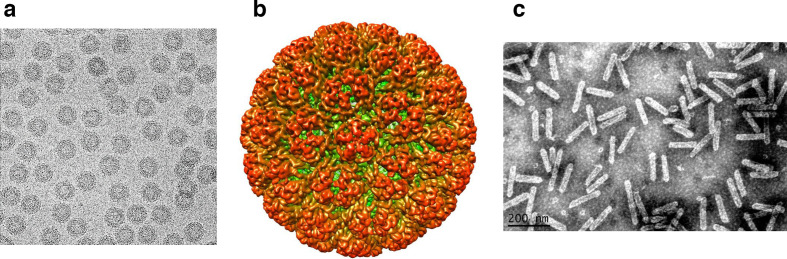
Virions are either isometric of 45–52 nm in diameter or, in the case of members of the genera Badnavirus and Tungrovirus, bacilliform particles of 30 nm × 60–900 nm (Table 1, Fig. 1). Virion sedimentation coefficient (S20,w) is 200–220 S; density in CsCl is 1.37 g cm−3. No envelope is present.
Table 1.
Characteristics of members of the family Caulimoviridae
|
Typical member: |
cauliflower mosaic virus-Cabb-S (V00141), species Cauliflower mosaic virus, genus Caulimovirus |
|---|---|
|
Virion |
Non-enveloped, isometric or bacilliform with a single-core capsid protein |
|
Genome |
7.1–9.8 kbp of non-covalently closed circular dsDNA with discontinuities in both genome strands at specific places |
|
Replication |
Cytoplasmic via reverse transcription of pregenomic RNA by viral reverse transcriptase. Terminally redundant pregenomic RNA is transcribed in the nucleus from repaired, covalently closed circular dsDNA by host DNA-directed RNA polymerase II |
|
Translation |
From capped and polyadenylated pregenomic RNA; in some viruses from subgenomic RNA and spliced versions of pregenomic RNA |
|
Host range |
Plants (monocots and dicots); some are transmitted by insects |
|
Taxonomy |
Realm Riboviria, kingdom Pararnavirae, phylum Artverviricota, class Revtraviricetes, order Ortervirales, multiple genera including >80 species |
Fig. 1.
Negative-contrast electron micrographs of virions of (a) cauliflower mosaic virus and (c) banana streak MY virus. (b) Tridimensional reconstruction of the cauliflower mosaic virus particle (images courtesy of Patrick Bron and Andrew D.W. Geering).
Genome
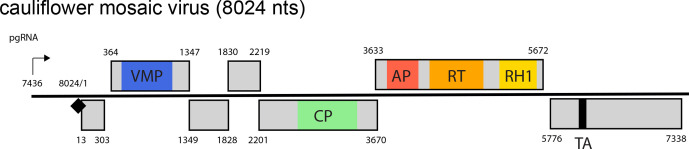
Virions contain a single molecule of non-covalently closed circular dsDNA of 7.1–9.8 kbp [1, 2] with discontinuities at specific sites in the negative-sense (one) and positive-sense strand (one to three). Genomes contain 1–8 ORFs encoding 5–6 conserved protein domains (Fig. 2), depending on the genus.
Fig. 2.
Caulimovirus genome linearised at the pregenomic RNA transcription start site (black arrow), numbered from the Met-tRNA primer binding site (black diamond). ORFs (light grey) include domains for the viral movement protein (VMP, blue), coat protein conserved C-terminus (CP, green), retropepsin (pepsin-like aspartic protease, AP, red), reverse transcriptase (RT, orange), RNase H1 (RH1, yellow), and translation transactivator (TA, black).
Replication
Following entry into the cell, the virion is targeted to the nucleus by a nuclear localization signal in the N-terminus of the capsid protein. Discontinuities in the genome are sealed to give supercoiled DNA, which associates with histone proteins to form mini-chromosomes in the nucleus. These are transcribed by host DNA-directed RNA polymerase II to give a greater-than-genome length transcript (35S or 34S RNA) that has a terminal redundancy of 35 to 270 nt. This transcript (pregenomic RNA) serves as a template for reverse transcription to give the negative-sense strand DNA and as a polycistronic mRNA for expression of at least some of the ORFs [3].
Unlike retroviruses, the episomal replication cycle does not involve an integration phase [4–6]. Negative-sense strand DNA synthesis is primed by host cytosolic tRNAmet. Synthesis of both strands is performed by the viral reverse transcriptase and RNase H1. RNase H1-resistant polypurine stretches serve as primer for positive-sense DNA synthesis. The site-specific discontinuities are at the priming sites for both negative- and positive-sense strand DNA synthesis and are made by the oncoming strand displacing the existing strand for a short distance and not ligating to form a closed circle [2].
Taxonomy
Current taxonomy: ictv.global/report/caulimoviridae. Members of the genera Badnavirus and Tungrovirus have bacilliform virions whereas members of the genera Caulimovirus, Cavemovirus, Petuvirus, Rosadnavirus, Solendovirus and Soymovirus have isometric virions. The number of ORFs ranges between one (petuviruses and vacciniviruses), three or more (badnaviruses), four (cavemoviruses, dioscoviruses, solendoviruses and tungroviruses), seven (caulimoviruses), seven or eight (soymoviruses) and eight (rosadnaviruses). Insect-mediated transmission has been reported for badnaviruses, caulimoviruses and tungroviruses. Infectious endogenous viral elements (EVEs) have been reported for several banana streak viruses (Badnavirus), petunia vein clearing virus (Petuvirus) and tobacco vein clearing virus (Solendovirus).
Resources
Current ICTV Report on the family Caulimoviridae: ictv.global/report/caulimoviridae
Funding information
Production of this summary, the online chapter, and associated resources was funded by a grant from the Wellcome Trust (WT108418AIA).
Acknowledgements
Members of the ICTV Report Consortium are Stuart G. Siddell, Andrew J. Davison, Elliot J. Lefkowitz, Sead Sabanadzovic, Peter Simmonds, Donald B. Smith, Richard J. Orton and F. Murilo Zerbini.
Conflicts of interest
The authors declare that there are no conflicts of interest
References
- 1.Bousalem M, Douzery EJP, Seal SE. Taxonomy, molecular phylogeny and evolution of plant reverse transcribing viruses (family Caulimoviridae) inferred from full-length genome and reverse transcriptase sequences. Arch Virol. 2008;153:1085–1102. doi: 10.1007/s00705-008-0095-9. [DOI] [PubMed] [Google Scholar]
- 2.Hohn T, Rothnie H. Plant pararetroviruses: replication and expression. Curr Opin Virol. 2013;3:621–628. doi: 10.1016/j.coviro.2013.08.013. [DOI] [PubMed] [Google Scholar]
- 3.Pooggin MM, Ryabova LA. Ribosome shunting, polycistronic translation, and evasion of antiviral defenses in plant pararetroviruses and beyond. Front Microbiol. 2018;9:644. doi: 10.3389/fmicb.2018.00644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Diop SI, Geering ADW, Alfama-Depauw F, Loaec M, Teycheney PY, et al. Tracheophyte genomes keep track of the deep evolution of the Caulimoviridae . Sci Rep. 2018;8:572. doi: 10.1038/s41598-017-16399-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Geering AD, Scharaschkin T, Teycheney PY. The classification and nomenclature of endogenous viruses of the family Caulimoviridae . Arch Virol. 2010;155:123–131. doi: 10.1007/s00705-009-0488-4. [DOI] [PubMed] [Google Scholar]
- 6.Geering AD, Maumus F, Copetti D, Choisne N, Zwickl DJ, et al. Endogenous florendoviruses are major components of plant genomes and hallmarks of virus evolution. Nat Commun. 2014;5:5269. doi: 10.1038/ncomms6269. [DOI] [PMC free article] [PubMed] [Google Scholar]