The soil bacterium Burkholderia thailandensis produces 2-alkyl-4(1H)-quinolones that are mostly methylated 4-hydroxyalkenylquinolines, a family of relatively unstudied metabolites similar to molecules also synthesized by Pseudomonas aeruginosa. Several of the methylated 4-hydroxyalkenylquinolines have antimicrobial activity against other species. We show that Bacillus subtilis strain 168 is particularly susceptible to N-oxidated methylalkenylquinolines (HMAQ-NOs). We confirmed that HMAQ-NO biosynthesis requires the previously unstudied protein HmqL. These results provide new information about the biology of 2-alkyl-4(1H)-quinolones, particularly the methylated 4-hydroxyalkenylquinolines, which are unique to B. thailandensis. This study also has importance for understanding B. thailandensis secondary metabolites and has implications for potential therapeutic development.
KEYWORDS: Burkholderia, cell-cell interaction, natural antimicrobial products, quinolones
ABSTRACT
The bacterium Burkholderia thailandensis produces an arsenal of secondary metabolites that have diverse structures and roles in the ecology of this soil-dwelling bacterium. In coculture experiments, B. thailandensis strain E264 secretes an antimicrobial that nearly eliminates another soil bacterium, Bacillus subtilis strain 168. To identify the antimicrobial, we used a transposon mutagenesis approach. This screen identified antimicrobial-defective mutants with insertions in the hmqA, hmqC, and hmqF genes involved in biosynthesis of a family of 2-alkyl-4(1H)-quinolones called 4-hydroxy-3-methyl-2-alkenylquinolines (HMAQs), which are closely related to the Pseudomonas aeruginosa 4-hydroxy-2-alkylquinolines (HAQs). Insertions also occurred in the previously uncharacterized gene BTH_II1576 (“hmqL”). The results confirm that BTH_II1576 is involved in generating N-oxide derivatives of HMAQs (HMAQ-NOs). Synthetic HMAQ-NO is active against B. subtilis 168, showing ∼50-fold more activity than HMAQ. Both the methyl group and the length of the carbon side chain account for the high activity of HMAQ-NO. The results provide new information on the biosynthesis and activities of HMAQs and reveal new insight into how these molecules might be important for the ecology of B. thailandensis.
IMPORTANCE The soil bacterium Burkholderia thailandensis produces 2-alkyl-4(1H)-quinolones that are mostly methylated 4-hydroxyalkenylquinolines, a family of relatively unstudied metabolites similar to molecules also synthesized by Pseudomonas aeruginosa. Several of the methylated 4-hydroxyalkenylquinolines have antimicrobial activity against other species. We show that Bacillus subtilis strain 168 is particularly susceptible to N-oxidated methylalkenylquinolines (HMAQ-NOs). We confirmed that HMAQ-NO biosynthesis requires the previously unstudied protein HmqL. These results provide new information about the biology of 2-alkyl-4(1H)-quinolones, particularly the methylated 4-hydroxyalkenylquinolines, which are unique to B. thailandensis. This study also has importance for understanding B. thailandensis secondary metabolites and has implications for potential therapeutic development.
INTRODUCTION
The saprophytic betaproteobacterium Burkholderia thailandensis is closely related to two pathogens, Burkholderia pseudomallei and Burkholderia mallei, which are the causative agents of melioidosis and glanders, respectively (1, 2). B. pseudomallei is also a saprophyte and causes respiratory or skin infections in humans following exposure to organisms in the environment, such as through skin contact with soil (3). B. mallei is a host-adapted pathogen and is spread to humans from horses and other ungulates, in which it is endemic in some regions (4). Because B. pseudomallei and B. mallei are tier 1 select agents and require handling under biosafety level 3 (BSL-3) laboratory conditions, B. thailandensis is often used as a surrogate to study the biology and virulence mechanisms of these pathogens (5). The development of versatile genetic techniques (6–9) and improvements in mouse models of melioidosis (10) have greatly improved the ability to study the biology of this relatively understudied group.
There has been much interest in elucidating the arsenal of small molecules produced by B. thailandensis, where there are at least 13 polyketide synthesis (PKS) gene clusters, many of which are conserved in B. mallei and/or B. pseudomallei. Although many of these metabolites have now been identified, only a few have been studied in much detail. One of the best studied is bactobolin (11, 12), which blocks translation by binding to a unique site in the 50S ribosomal subunit (13). Other PKS antibiotics are malleilactone (14, 15) and malleicyprol, a more toxic product of the malleilactone biosynthetic gene cluster (16), which contribute to virulence of B. pseudomallei (17). B. thailandensis also produces thailandenes, a group of polyenes with activity against Gram-positive bacteria (18). As with many bacterial natural products, malleilactone and thailandenes are not produced under standard laboratory conditions (14, 15, 18). Studies of these molecules were possible through genetic (14, 15) or chemical (19) elicitation of the gene clusters or through phenotype-based screening approaches (18).
Most of the PKS gene clusters are unique to this group of Burkholderia species. A few of them have analogous biosynthesis pathways in other Burkholderia species or even beyond the Burkholderia genus. For example, the hmqABCDEFG operon coding for enzymes responsible for the biosynthesis of a family of 4-hydroxy-2-alkylquinolines (HAQs) named 4-hydroxy-3-methyl-2-alkenylquinolines (HMAQs) are found in B. thailandensis, B. pseudomallei, and other members of the Burkholderia genus, such as Burkholderia ambifaria (20). The products made by the HmqABCDEFG enzymes have differing carbon chain lengths and saturation and presence of substitutions on the quinolone ring, such as methylation and oxidation. The relative abundances of these various congeners differ between species (21). The hmq operon is homologous to the pqs operon found in P. aeruginosa (Fig. 1) (21, 22). The molecules produced by Burkholderia also differ from those of P. aeruginosa in that most bear a methyl group at the 3′ position and possess an unsaturated aliphatic side chain, which are linked to the presence of the additional hmqG and hmqF genes, respectively (21). The main product of the P. aeruginosa pqs operon, 4-hydroxy-2-heptylquinoline (HHQ), is converted to 3,4-dihydroxy-2-heptylquinoline (Pseudomonas quinolone signal [PQS]) by the enzyme PqsH (23, 24). Both HAQs are involved in quorum sensing in P. aeruginosa and are detected by the MvfR (PqsR) regulator (25–27). No homologs of the pqsH and mvfR genes have been found in Burkholderia species (21).
FIG 1.
Biosynthesis of 4-hydroxy-2-alkylquinoline congeners. Burkholderia thailandensis uses the hmq gene products to synthesize HAQs including HMAQ and HMAQ-NO. In Pseudomonas aeruginosa, the analogous pqs gene products synthesize the related compounds HAQ, HAQ-NO, and PQS. Shown are the N-oxidated species referred to in the text, HQNO and HMAQ-NO-C9 with a double bond at the 1′-2′ position added by HmqF. The B. thailandensis compounds are methylated by HmqG, which does not have a homolog in P. aeruginosa. PqsH is needed for production of PQS, which is specific to P. aeruginosa.
We are interested in the small-molecule repertoire of B. thailandensis as an avenue to better understand the biology of this bacterium and make new discoveries in natural product biosynthesis. We observed that B. thailandensis culture fluid has significant antimicrobial activity that is not due to bactobolin, the only other known antimicrobial produced under these conditions. This bactobolin-independent activity was isolated to the hmq gene cluster using an approach involving transposon mutagenesis and screening for mutants exhibiting reduced antimicrobial activity. Purified and synthetic HAQ derivatives were assessed for the antimicrobial properties of several such biosynthetic products of the hmq genes, including HMAQ congeners and N-oxide derivatives (HMAQ-NOs) with various alkenyl side chain lengths. We also confirmed the involvement of hmqL in the biosynthesis of HMAQ-NO compounds. Our results provide new information on the biosynthesis and activities of the HMAQs produced by Burkholderia.
RESULTS
Antimicrobial activity of B. thailandensis bactobolin-null mutants.
Initial liquid coculture experiments with B. thailandensis strain E264 and B. subtilis strain 168 (hereinafter referred to as B. thailandensis or B. subtilis, respectively) showed that B. thailandensis has a strong growth advantage over B. subtilis. The growth advantage was so substantial that after overnight liquid coculture with B. thailandensis, B. subtilis decreased from a density of 106 cells per ml to below the limit of detection (<102 cells per ml). This result was not solely attributable to bactobolin, as a bactobolin-null mutant (BD20) also had the same growth advantage over B. subtilis (Fig. 2A). This observation led to the hypothesis that B. thailandensis has a previously uncharacterized antimicrobial activity against B. subtilis that is not mediated by bactobolin. To further explore this hypothesis, culture fluids of several B. thailandensis strains were harvested and tested for antimicrobial activity (Fig. 2B). As previously observed (11), filter-sterilized culture fluids of wild-type B. thailandensis saturating a paper filter disc placed on a lawn of B. subtilis caused a zone of growth inhibition around the filter disc, whereas there was no growth inhibition observed with the bactobolin-null BD20 strain (Fig. 2B, top panel). However, unprocessed culture fluid of both strains (wild type and BD20), which had not gone through the filter sterilization process, demonstrated antimicrobial activity (Fig. 2B, middle and bottom). This observation (i.e., that only unprocessed culture fluid had bactobolin-independent antimicrobial activity) could be explained by several possible hypotheses: first, that the filter sterilization process removes or inactivates antimicrobial activity, and second, that antimicrobial activity requires live cells. In support of the first hypothesis, the antimicrobial activity was observed in the absence of viable B. thailandensis bactobolin mutant cells; unprocessed B. thailandensis BD20 culture fluid had activity against B. subtilis when added directly to high-salt LB agar plates, which are conditions that do not allow B. thailandensis growth (Fig. 2B, bottom). Ethyl acetate extracts of B. thailandensis cultures also had activity against B. subtilis (Fig. S1 in the supplemental material). Together, these results suggest that B. thailandensis produces an antimicrobial other than bactobolin, which is eliminated by filter sterilization.
FIG 2.

Sensitivity of Bacillus subtilis strain 168 to a substance produced by Burkholderia thailandensis. (A) For liquid coculture growth, B. subtilis strain 168 was combined in a 1:1 ratio with either Burkholderia thailandensis E264 (WT) or bactobolin-deficient B. thailandensis (Bacto−, strain BD20) in LB broth and grown for 24 h at 37°C prior to plating to determine surviving CFU as described in Materials and Methods. Data are representative of three biological replicates. Error bars show standard deviations. (B) Growth inhibition of B. subtilis strain 168 following treatment with cultures or culture fluid from B. thailandensis after 18 h of growth on plates. B. thailandensis wild type (E264) or the bactobolin-defective mutant (Bacto−, strain BD20) was applied to a lawn of freshly plated B. subtilis 168, and plates were incubated at 30°C prior to imaging. Top, B. thailandensis culture fluid was filtered and used to saturate paper diffusion discs applied to the lawn of B. subtilis 168. A zone of clearing around a diffusion disc indicates the region where B. subtilis growth was inhibited. Results are similar to those previously reported (11). Middle, unfiltered B. thailandensis fluid (10 μl) was spotted directly onto B. subtilis 168. Bottom, unfiltered B. thailandensis fluid was spotted onto a lawn of B. subtilis 168, as in the experiment whose results are shown in the middle panel, but on medium containing 5% NaCl, which inhibits B. thailandensis growth.
Isolation and identification of antimicrobial-deficient transposon mutants.
To identify the genes required for the observed antimicrobial activity, we used a mutagenesis and screening approach. First, we randomly mutagenized the B. thailandensis bactobolin-null mutant BD20 with a transposon containing the trimethoprim resistance gene dhfR (Tn5::dhfR). Next, we screened the mutants (∼10,000) using a high-throughput method to assess antimicrobial activity (for experiment overview, see Fig. S2). Briefly, we added B. subtilis cells to cooled molten agar and mixed gently before pouring into plates. After the medium solidified, single isolated colonies (i.e., transposon mutants) were patched onto the plates. The next day, plates were assessed for zones of inhibition. B. thailandensis patches demonstrating reduced zones of inhibition compared with the B. thailandensis bactobolin-defective parent were reisolated for further study. We initially identified 60 antimicrobial-defective candidates. Of those, 9 were confirmed to have reduced antimicrobial activity against B. subtilis (Fig. 3A) with no observable growth defects (Table S1). These were mutants 7, 9, 14, 27, 31, 32, 56, 63, and 68 (Table 1).
FIG 3.
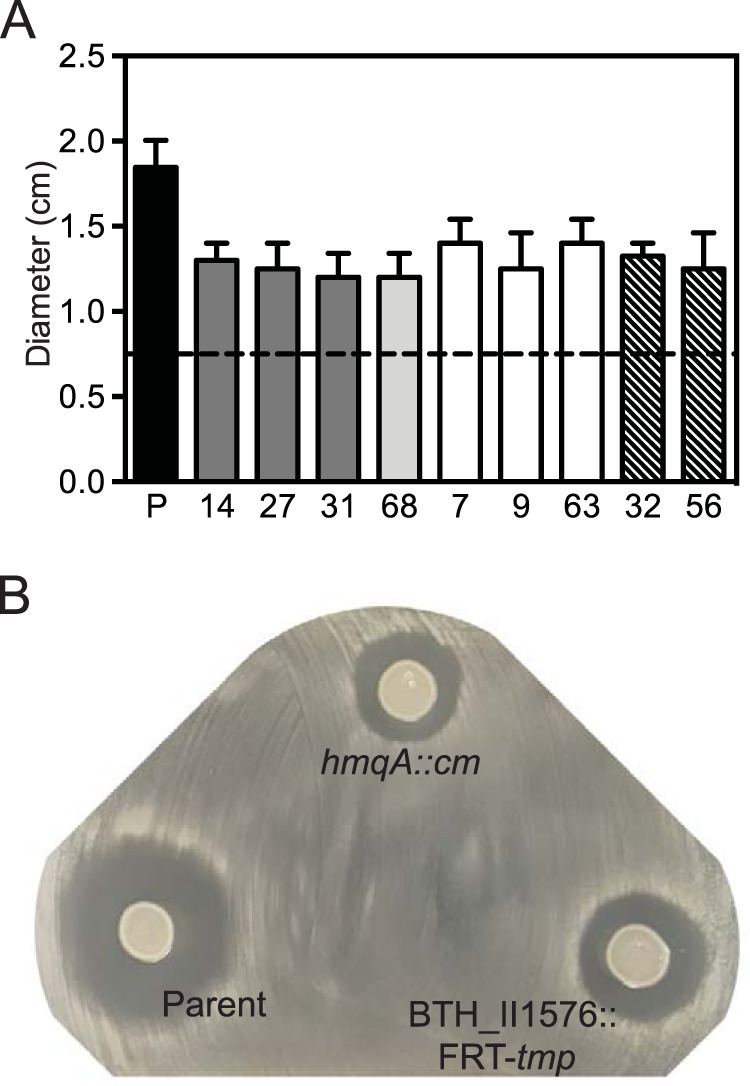
B. thailandensis transposon mutants with reduced effects on Bacillus subtilis strain 168. (A) Amounts of 5 μl of unfiltered fluid from B. thailandensis stationary-phase cultures were spotted onto lawns of freshly plated B. subtilis strain 168 and incubated overnight at 30°C. Results are shown as the diameters of the zones of inhibition. The black dashed line indicates the diameter of the spot of B. thailandensis culture. Transposon mutant numbers correspond with those in Table 1 and are shaded according to the gene mutation, as follows: dark gray, hmqA disruptions; light gray, hmqC disruption; white, hmqF disruptions; hatched, BTH_II1576 disruptions. P (parent), B. thailandensis bactobolin-deficient mutant BD20 used for transposon mutagenesis. Data are the averages of two biological replicates. Error bars show standard deviations. (B) Images of B. subtilis 168 lawns spotted with 5 μl unfiltered fluid from cultures of the B. thailandensis bactobolin-deficient strain BD20 (Parent) or BD20 with a disruption in hmqA (hmqA::cm) or BTH_II1576 (BTH_II1576::FRT-tmp) introduced by homologous recombination.
TABLE 1.
Locations of transposon insertions
| Mutant | Locus | Gene | Predicted gene function | Insertion location (gene length)a |
|---|---|---|---|---|
| 14 | BTH_II1935 | hmqA | 2-Aminobenzoate-CoA ligase | 1231 (1,626) |
| 27 | BTH_II1935 | hmqA | 2-Aminobenzoate-CoA ligase | 1513 (1,626) |
| 31 | BTH_II1935 | hmqA | 2-Aminobenzoate-CoA ligase | 1478 (1,626) |
| 68 | BTH_II1933 | hmqC | Unknown | 783 (1,107) |
| 7 | BTH_II1930 | hmqF | Polyketide synthase | 163 (3,972) |
| 9 | BTH_II1930 | hmqF | Polyketide synthase | 2524 (3,972) |
| 63 | BTH_II1930 | hmqF | Polyketide synthase | 2872 (3,972) |
| 32 | BTH_II1576 | “hmqL”b | Putative monooxygenase | 226 (1,167) |
| 56 | BTH_II1576 | “hmqL” | Putative monooxygenase | 998 (1,167) |
The number of the nucleotide just prior to the transposon insertion relative to the predicted translation start site is given, followed by the total nucleotides in the gene in parentheses.
hmqL, proposed name from reference 22.
The locations of the transposon insertions were identified using whole-genome resequencing of five of the mutants and PCR amplification of suspected target gene loci of the remaining four (see Materials and Methods). Of the nine mutants isolated, seven had insertions in the hmqABCDEFG operon (BTH_II1929 to -1935) (Table 1). The other two mutants had disruptions in a previously unstudied gene, BTH_II1576, which is predicted to encode a monooxygenase. To verify that the hmq locus and BTH_II1576 contribute to the antimicrobial defects observed for the transposon mutants, we disrupted hmqA or BTH_II1576 in the bactobolin-defective BD20 strain using homologous recombination. Both gene disruptions caused defects in B. subtilis growth inhibition similar to those observed with the transposon mutants (Fig. 3B), supporting the idea that the hmq genes and BTH_II1576 are important for the bactobolin-independent antimicrobial activity of B. thailandensis.
Identification and activities of hmq gene products.
Both the pqs and hmq gene products use anthranilic acid and fatty acid precursors to generate HMAQs and HAQs through the pathway illustrated in Fig. 1. The result of biosynthesis includes molecules with unsaturated or saturated side chains and N-oxide derivatives. The most abundant HMAQ in B. thailandensis E264 cultures is a congener with an unsaturated C9 side chain, 4-hydroxy-3-methyl-2-nonenylquinoline, referred to as HMNQ or HMAQ-C9:2′ (21). To test whether production of HMAQ-C9:2′ is absent in our transposon mutants, we measured this HMAQ in culture fluid using a liquid chromatography-tandem mass spectrometry (LC-MS/MS) method to find a product with the expected m/z of 284. Consistent with previous results (21, 28), transposon mutants with insertions in hmqA and hmqF had no HMAQ-C9:2′ (<0.05 μg/ml, the limit of detection). We also detected no HMAQ-C9:2′ in hmqC mutants, consistent with the proposed role of HmqC in HMAQ biosynthesis (Fig. 1). The BD20 parent strain and BTH_II1576 transposon mutants both readily produced this HMAQ congener (measured at 5 to 8 μg/ml).
We tested the activity of HMAQ-C9:2′ directly against B. subtilis using a standard MIC assay. The MIC of purified HMAQ-C9:2′ against B. subtilis was 50 μg/ml (Table 2). HMAQ-C9:2′ also inhibited the growth of Staphylococcus aureus (MIC of 25 μg/ml). We did not detect any antimicrobial activity of HMAQ-C9:2′ against Escherichia coli or Pseudomonas aeruginosa (MIC of >200 μg/ml). Of note, the concentration of HMAQ-C9:2′ in B. thailandensis cultures (5 to 8 μg/ml) is ∼5-fold lower than needed to inhibit B. subtilis growth (50 μg/ml), suggesting that HMAQ-C9:2′ alone is not sufficient for the observed effects on B. subtilis in our coculture experiments. Instead, we hypothesized that the antimicrobial activity in cocultures involved another product of the hmq genes.
TABLE 2.
Antimicrobial activities of hydroxylalkylquinoline analogs
| Quinolone familya | Carbon chain | [M + H]+ | MIC (μg/ml)b
|
|
|---|---|---|---|---|
| B. subtilis 168 | S. aureus Newman | |||
| HMAQ | C9:2′ | 284 | 50 | 25 |
| HMAQ-NO | C9:2′ | 300 | 0.75 | 25 |
| HAQ (HHQ) | C7 | 242 | >200 | >200 |
| HAQ-NO (HQNO) | C7 | 259 | 25 | 25 |
| HMAQ-NO | C8:2′ | 286 | 1.5 | 6.25 |
| HMAQ-NO | C7:2′ | 272 | 6.25 | 12.5 |
HMAQ with a C9 carbon chain (HMAQ-C9:2′) was purified as described in reference 21. HMAQ-NO congeners were synthesized as described in Materials and Methods and Piochon et al. (31). HAQ-NO with a C7 carbon chain (HQNO) and HAQ with a C7 carbon chain (HHQ) were purchased from commercial sources (Cayman Chemicals and Sigma Aldrich, respectively).
No activity of any of the compounds against Pseudomonas aeruginosa strain PA14 or Escherichia coli strain JM109 was observed up to 200 μg/ml. Results are the averages of three independent experiments. In all cases, the range was <5%.
Biosynthesis and antimicrobial activity of HMAQ-NO.
The protein product of BTH_II1576 shares 52% amino acid sequence identity to the P. aeruginosa PqsL protein involved in HAQ biosynthesis. PqsL synthesizes 2-hydroxylaminobenzoylacetate (2-HABA) from 2-aminobenzoylacetate (2-ABA) as a step in the pathway to make N-oxide derivatives (HAQ-NOs) (Fig. 1, left column) (29, 30). We hypothesized that BTH_II1576 is similarly involved in the biosynthesis of N-oxide HMAQ (HMAQ-NO) in B. thailandensis (Fig. 1, right column). To test this hypothesis, we used LC-MS/MS to measure HMAQ-NO in the BTH_II1576 transposon mutants. We measured HMAQ-NO with an unsaturated C9 or C7 side chain, which are two abundant HMAQ congeners in B. thailandensis E264 cultures. Both of the BTH_II1576 mutants and our constructed BD20 BTH_II1576 mutant had undetectable HMAQ-NO-C9 (<0.05 μg/ml), whereas the BD20 parent had measurable levels (1.5 ± 0.5 μg/ml [mean ± standard deviation]). We also expressed BTH_II1576 ectopically from an IPTG (isopropyl-β-d-thiogalactopyranoside)-inducible lac promoter in the neutral glmS1 site in the engineered BTH_II1576 mutant genome, and we compared HMAQ-NO-C9 and the antimicrobial activities in this strain with those of an empty lac promoter-containing mutant or BD20 parent (Fig. 4). IPTG induction of BTH_II1576 in the mutant restored the production of HMAQ-NO (Fig. 4A) and increased the zone of inhibition of B. subtilis in colony outgrowth experiments (Fig. 4B), supporting the idea that BTH_II1576 is important for each of these processes. Furthermore, BTH_II1576 induction significantly decreased HMAQs, supporting the idea that the product of BTH_II1576 uses HMAQ precursors as substrate to generate HMAQ-NO. Together, our results confirm that the BTH_II1576 product is analogous to PqsL in HAQ-NO biosynthesis and is appropriately named HmqL, as previously proposed (22).
FIG 4.

BTH_II1576 (hmqL) involvement in HMAQ-NO production and activity against B. subtilis strain 168. (A) HMAQ-NO (C9) was quantified in stationary-phase B. thailandensis strains using LC-MS/MS and methods described previously (21). Error bars show standard deviations. (B) Antimicrobial activity of unfiltered B. thailandensis fluid (5 μl) on a lawn of freshly plated B. subtilis on plates containing 1 mM IPTG. Strains tested were the B. thailandensis bactobolin-deficient BD20 (Parent), BD20 with the IPTG-inducible Plac expression cassette inserted into the neutral glmS1 site in the genome (BD20 Plac), the constructed BD20 BTH_II1576 (hmqL) mutant with the Plac cassette in glmS1 (hmqL::tmp/Plac), or the BD20 hmqL mutant with Plac-hmqL in glmS1 (hmqL::tmp/Plac-hmqL).
Because HmqL generates HMAQ-NO and is important for the antimicrobial activity observed in B. thailandensis cultures, we tested the hypothesis that HMAQ-NO has antimicrobial activity against B. subtilis. We assessed the sensitivity of B. subtilis to the most abundant HMAQ-NO produced by B. thailandensis (21), synthetic HMAQ-NO-C9:2′ (31) (Table 2). The MIC of HMAQ-NO-C9:2′ against B. subtilis was 0.75 μg/ml. This MIC is below the measured concentration of HMAQ-NO-C9:2′ in B. thailandensis cell cultures (1.5 ± 0.5 μg/ml), supporting the idea that HMAQ-NO is primarily responsible for the observed antimicrobial activity against B. subtilis in cocultures with B. thailandensis. Interestingly, there was no difference in activity between the C9 congeners of HMAQ-NO and HMAQ against S. aureus strain Newman (MIC 25 μg/ml). Differences in diffusion or target site availability could explain the differences in relative activities of these two molecules in each species.
Antimicrobial activities of structurally related 4-hydroxy-2-alkylquinolines.
We found it intriguing that the C7 HAQ (HHQ) and its N-oxide derivative HAQ-NO-C7 (HQNO) were much less active against B. subtilis than the respective C9 HMAQ and HMAQ-NO molecules (Table 2). The difference in activity could be due to the difference in alkyl chain lengths or saturation levels. Alternatively, the presence of the methyl group in HMAQs could also affect the activity. To address the first possibility, we tested synthetic HMAQ-NO congeners with a C7 and a C8 unsaturated alkyl side chain against B. subtilis. Our results showed that the C8 and C7 HMAQ-NO molecules were 2- and 8-fold less active, respectively, than the C9 congener against B. subtilis (Table 2). These results suggest the molecule with the longer carbon chain length has higher activity of HMAQ-NO against B. subtilis. The C7 HMAQ-NO was also more active than HQNO (HAQ-NO-C7) by about 2-fold against the S. aureus Newman strain and 4-fold against B. subtilis (Table 2). HQNO differs from C7 HMAQ-NO in that it is unmethylated and has a saturated side chain. Thus, either methylation or saturation of the side chain also plays a role in the activity.
HMAQ promotes competition in liquid cocultures.
The results of our transposon mutant analysis suggest that HMAQs, and in particular HMAQ-NO-C9, eliminate B. subtilis from liquid cocultures, providing an explanation for our initial observation. To test this hypothesis, we competed B. subtilis with a B. thailandensis bactobolin-deficient BD20 strain containing either a single hmqA or hmqL mutation or an hmqA-hmqL mutation in liquid coculture experiments. Singly disrupting hmqA nearly abolished B. thailandensis activity against B. subtilis (Fig. 5A). However, singly disrupting hmqL showed only a small and not statistically significant reduction of activity. A strain in which both hmqL and hmqA were disrupted showed defects in antimicrobial activity similar to that of the hmqA single mutant, supporting the idea that HmqA is upstream from HmqL in the biosynthetic pathway. These results show that the activity against B. subtilis in liquid cocultures is due to hmq biosynthetic products but not HMAQ-NO alone. The activity either requires HMAQ-NO together with other hmq biosynthetic products or is independent of HMAQ-NO.
FIG 5.

Involvement of BTH_II1576 (hmqL) in liquid cocultures and cell pellet fraction localization of HMAQ and HMAQ-NO molecules. (A) Results of cocultures of B. subtilis strain 168 combined in a 1:1 ratio with the bactobolin-deficient B. thailandensis (BD20) parent strain or the parent strain bearing a constructed disruption of hmqA, hmqL, or both in LB broth and grown for 24 h at 37°C. Surviving CFU were enumerated by serial dilution and plating on LB agar containing, for B. subtilis, 5% NaCl (nonpermissive for B. thailandensis growth), and for B. thailandensis, 100 μg/ml gentamicin (nonpermissive for B. subtilis). Data are representative of three biological replicates, and the error bars represent standard deviations. Statistical significance was determined by repeated measures one-way analysis of variance (ANOVA), followed by Tukey’s multiple-comparison tests, and the letters A and B represent statistically significantly different groups (P < 0.05). (B) C9 congeners of HMAQ and HMAQ-NO were quantified in unfiltered and filtered fluid from cell-free B. thailandensis cultures, as well as in pelleted cells, using LC-MS/MS and methods described previously (21). Error bars show standard deviations.
Our initial observations suggested that the antimicrobial in B. thailandensis cultures was sensitive to filtration; thus, we also sought to test the sensitivities of HMAQ and HMAQ-NO to filtration. We measured the concentrations of each of these molecules in unfiltered and filtered fluid from cell-free B. thailandensis cultures. We also determined the concentrations of these molecules in pelleted cells to determine if they are primarily associated with the cell, similar to HAQs in P. aeruginosa (26, 32). We found that the percentages of HMAQs and HMAQ-NOs in the cell fraction, which included molecules associated with the cell membrane, periplasm, and cytoplasm, were 91% ± 2% and 71% ± 3%, respectively. Thus, these molecules are highly cell associated. Furthermore, filtration further depletes the molecules remaining in culture fluid to nearly undetectable levels (Fig. 5B). These results are consistent with the idea that HMAQs and HMAQ-NOs are removed by separation of the cells and filtration of the remaining fluid, providing an explanation as to how the activity of these molecules has been missed in prior experiments.
HMAQ biosynthesis in B. ambifaria.
The Burkholderia ambifaria genome contains an hmq operon homologous to that of Burkholderia thailandensis (21). However, B. ambifaria does not produce HMAQ-NOs (21), presumably because it does not have a homolog of hmqL and pqsL. We predicted that introducing the B. thailandensis hmqL to B. ambifaria would enable production of HMAQ-NO. To test this prediction, we introduced the hmqL gene to B. ambifaria on plasmid pME6010 (33). Because HMAQ biosynthesis is less well characterized in this species, we used combined measurements of all three C7, C8, and C9 congeners of HMAQs for our analysis. We observed that B. ambifaria with pME6010 had no detectable HMAQ-NO, as previously reported (21). However, B. ambifaria with pME6010-hmqL produced measurable levels of HMAQ-NO (Fig. 6A), which is consistent with the idea that HmqL is the only missing enzyme and its presence permits the production of HMAQ-NO production in B. ambifaria. This strain also had 100-fold less HMAQ than the empty plasmid-only strain (Fig. 6A), suggesting strong competition for the HMAQ precursor, likely 2-aminobenzoyl(methyl)acetate (30) (the product of HmqADEG) (Fig. 3). We also tested whether the expression of HmqL caused B. ambifaria to inhibit B. subtilis growth. We spotted unfiltered culture fluid from B. ambifaria with pME6010 or pME6010-hmqL onto a lawn of B. subtilis. Only cultures of the strain expressing hmqL could inhibit B. subtilis growth (Fig. 6B). Together, the results provide further support for the idea that HmqL is crucial for production of the HMAQ-NO antimicrobials.
FIG 6.

Heterologous expression of hmqL in Burkholderia ambifaria. (A) HMAQ and HMAQ-NO in cultures of B. ambifaria HSJ1 cells containing either pME6010 or pMP6010-hmqL. Results are the average of three biological replicates and represent the sum of the C7, C8, and C9 congeners of each molecule. Error bars show standard deviations. (B) Antimicrobial activities of unfiltered fluid (5 μl) from cultures of Burkholderia ambifaria HSJ1 containing pME6010 or pME6010-hmqL spotted onto freshly spread lawns of B. subtilis on plates. Plates were imaged after 24 h of incubation at 37°C.
DISCUSSION
Investigation of the antimicrobial properties of HAQs dates back to 1945, when an “antibiotic metabolite” was described in P. aeruginosa (34). Although the biosynthesis steps and biology of the HAQs in P. aeruginosa have since been studied in detail, much less is known of those in B. thailandensis (21, 31, 35). The results of this study add new information to the known steps of biosynthesis of B. thailandensis HMAQs. Previous studies showed that enzymes analogous to PqsABCD in P. aeruginosa are involved in the synthesis of B. thailandensis HMAQ from anthranilate (Fig. 1, right column). In P. aeruginosa, the enzyme PqsL catalyzes an essential step in the synthesis of HAQ N-oxides (29, 30). B. thailandensis has no PqsH enzyme homolog and does not make 3-hydroxylated HAQs; a methyl is instead present as a substitution at that position. B. thailandensis is also missing a homolog of the HHQ/PQS receptor gene, mvfR. Our study validates the involvement of HmqL in N-oxide HMAQ biosynthesis and shows how the HMAQ family of molecules contribute to the arsenal of compounds used by B. thailandensis to compete with other species. The findings also provide new insight into the activities of specific B. thailandensis HMAQ family congeners against other bacteria.
Like many toxins, H(M)AQs have several known functions. In P. aeruginosa, where these molecules are best studied, the N-oxide congeners are potent antimicrobials that inhibit Gram-positive bacteria (24, 36, 37), and several of the HAQs are important for interspecies competition (38–40). Other B. thailandensis antimicrobials include bactobolin (12), malleilactone (14, 15), and thailandenes (18). This suite of diverse antimicrobials might be important for surviving competition with other microbes when space or other resources become limited. The loss of the hmq biosynthesis genes from the genome of the closely related host-adapted pathogen B. mallei supports a role of these genes in the saprophytic lifestyle of B. thailandensis. The current study demonstrates that the N-oxide HMAQs are important for antimicrobial activity toward other species under several laboratory coculture conditions, similar to P. aeruginosa HQNO. HQNO also has other known effects, such as enhancing biofilm formation (41, 42) or increasing resistance to antimicrobials (43, 44), and it remains to be seen if HMAQ-NO is similar in these other ways.
We find it interesting that the Burkholderia species do not have the enzyme responsible for generating PQS (PqsH) (Fig. 1). PQS has a variety of known functions, such as immune modulation (45), cell density-dependent gene regulation (24, 46), and iron sequestration (26). B. thailandensis might have lost the ability to synthesize PQS because these functions are not needed or because there is existing functional redundancy with other molecules or pathways. For example, the small molecule malleilactone might have similar biophysical properties and also sequester iron (14). It is also interesting that B. ambifaria lacks the HmqL enzyme responsible for generating N-oxide HMAQs, which are the most antimicrobial members of this family. The lack of PQS or any N-oxide analog in B. ambifaria strongly supports the idea that other products of this pathway have important functions that contribute to the survival of this species, although the biology of the other products of the Hmq system is not well understood.
A particularly interesting discovery in this work was that B. thailandensis HMAQ-NO-C9:2′ is much more active (33-fold) than P. aeruginosa HQNO (HAQ-NO-C7) against B. subtilis (Table 2). The heightened activity of HMAQ-NO compared with that of HQNO could be due to both side chain length and, possibly, methylation (or saturation). It remains to be seen whether the structural moieties important for this activity alter the target site of this molecule, the ability to penetrate B. subtilis cells, or some other aspect of this molecule. In addition to the N-oxide congeners, B. thailandensis produces a variety of HMAQs with side chains of various lengths and degrees of saturation (21, 26). Although these other molecules had less potent antimicrobial activities (Table 2), it is possible they contribute to competition in other ways. A previous study showed that different variations of HAQs used in combination can have synergistic antimicrobial effects on other bacteria by acting on distinctly different cellular targets (47). Thus, the diversity of H(M)AQs produced by B. thailandensis might serve to enhance activity against competitors or could be important for averting the development of antibiotic resistance in competitors.
Our findings that HMAQs and HMAQ-NOs are associated with the cell and are particularly potent toward B. subtilis explain how their antimicrobial activity was missed in prior studies (11, 48–50). P. aeruginosa HAQs are similarly cell associated (26). Given the antibacterial properties of HMAQ-NOs and HQNO, it is curious that these molecules are cell associated. It may be that their antimicrobial activity is important under conditions, such as biofilms, where close contact with other cells occurs, rendering it beneficial for the activity to remain associated with the cell surface. An alternative possibility is that their primary role involves another function that is needed at the cell surface. For example, the role of PQS in iron entrapment requires an interaction with the cell surface (26). It is also noteworthy that HMAQ-NOs, along with other antimicrobials produced by B. thailandensis, can be associated with outer membrane vesicles (OMVs) (51). OMVs are buds of the outer membrane that are released from the cell (52). B. thailandensis OMVs have antimicrobial properties (51), similar to OMVs of other Gram-negative bacteria (53). OMVs fuse with the cell membrane of a competitor to deliver toxins directly into the periplasm (53). Thus, antimicrobials associated with the cell membrane are likely to be packaged into OMVs, which can enhance their activity during competition.
MATERIALS AND METHODS
Bacterial culture conditions and reagents.
Bacteria were grown in lysogeny broth (LB) (10 g tryptone, 5 g yeast extract, and 5 g NaCl per liter), supplemented with 50 mM MOPS (morpholinepropanesulfonic acid) pH 7 in M9 minimal medium supplemented with 0.4% d-glucose and 10 mM para-chloro-phenylalanine (p-Cl-Phe; Sigma) for B. thailandensis counterselection during mutant construction or in DM medium (0.25× M63 salts, 1 mM MgSO4, 0.4% glycerol, 0.2% glucose, 1 μg/ml thiamine, and 40 μg/ml each of leucine, isoleucine, valine, tryptophan, glutamic acid, and glutamine) for transformation of PCR-generated products. For liquid cocultures, B. subtilis and B. thailandensis growth was at 37°C. For all other experiments, B. thailandensis growth was at 30°C, and all E. coli and B. ambifaria growth was at 37°C. 4-Hydroxy-2-heptylquinoline (HHQ) was purchased from Sigma (catalog number SML0747). 4-Hydroxy-2-heptylquinoline N-oxide (HQNO) was purchased from Cayman Chemicals (catalog number 15159). 4-Hydroxy-3-methyl-2-nonenylquinoline (HMNQ) was purified from B. thailandensis E264 cultures as described previously (21). The other hydroxyalkenylquinolines were synthesized as described below. For selection, trimethoprim was used at 100 μg/ml, gentamicin was used at 100 μg/ml, kanamycin was used at 500 μg/ml (B. thailandensis) or 50 μg/ml (E. coli), tetracycline was used at 225 μg/ml (B. ambifaria), and NaCl was used at 5% (for inhibiting B. thailandensis in coculture enumerations). IPTG (isopropyl β-d-1-thiogalactopyranoside) was added at 1 mM final concentration to cultures and plates, when appropriate. Genomic DNA, PCR and DNA fragments, and plasmid DNA were purified using a Puregene core A kit, plasmid purification miniprep kit, or PCR cleanup/gel extraction kits (Qiagen or IBI-MidSci) according to the manufacturer’s protocol.
Synthesis of N-oxides of hydroxyalkenylquinolines.
HMAQ-NOs were synthesized as previously described (31) from corresponding HMAQs in which the quinolone scaffold was built via the Conrad-Limpach approach (54). Briefly, aniline was condensed with diethyl 2-methyl-3-oxosuccinate and the resulting diester was cyclized under acidic conditions. Reduction of the quinolone ester followed by halogen substitution led to 2-chloromethyl-3-methylquinolin-4(1H)-ones, which were subjected to Suzuki-Miyaura cross-coupling (55) with commercially available alkenylboronic acid pinacol esters to provide HMAQs. Then, they were converted into the corresponding ethyl carbonates, oxidized with meta-chloroperbenzoic acid (mCPBA), and deprotected to yield HMAQ-NOs (56). The structures of HMAQ-NOs were confirmed by high-resolution mass spectrometry, as well as 1-dimensional (1-D) and 2-dimensional (2-D) nuclear magnetic resonance (NMR) analysis. Structural data are available in Piochon et al. (31).
Bacterial strains and genetic manipulations.
All bacterial strains, plasmids, and primers used in this study are listed in Tables 3 and 4. We used the wild type and mutant derivatives of B. thailandensis strain E264 (5). We used B. subtilis strain 168 (53), S. aureus strain Newman (57), P. aeruginosa strain PA14 (58), B. ambifaria strain HSJ1 (21), and E. coli strain DH5α for genetic manipulations (Invitrogen). The B. thailandensis bactobolin-defective mutant BD20 has a deletion of the bactobolin biosynthesis gene btaK as described previously (11). The B. thailandensis hmqA mutant was constructed by allelic exchange using methods described previously (6) and plasmid pMCG19. pMCG19 was constructed by first amplifying hmqA from the B. thailandensis E264 genome using primers hmqAfor and hmqArev containing HindIII and KpnI cleavage sites, respectively. The PCR product was digested with HindIII and KpnI and ligated to HindIII-KpnI-cut pEX18Tp-PheS (9). The chloramphenicol resistance cassette was amplified from pACYC184 (59) using primers CmFPstI and CmRPstI, each containing the PstI cleavage site, and ligated to the PstI site inside the hmqA gene in pEX18Tp-PheS-hmqA to make pMCG19.
TABLE 3.
Bacterial strains and plasmids used in this study
| Strain or plasmid | Relevant properties | Reference or source |
|---|---|---|
| Burkholderia thailandensis strains | ||
| E264 | Wild type | 5 |
| BD20 | E264 with deletion of btaK | 11 |
| JRK100 | E264 hmqA::cm | This study |
| JRK101 | E264 BTH_II1576::FRT-tmp | This study |
| JRK102 | BD20 hmqA::cm | This study |
| JRK103 | BD20 BTH_II1576::FRT-tmp | This study |
| JRK104 | BD20 glmS1 attn7::Plac; Kmr | This study |
| JRK105 | JRK103 glmS1 attn7::Plac; Kmr | This study |
| JRK106 | JRK103 glmS1 attn7::Plac-hmqL; Kmr | This study |
| Escherichia coli strains | ||
| DH5α | F− ϕ80lacZ ΔM15 Δ(lacZYA-argF)U169 hsdR17(rK− mK+) recA1 endA1 phoA supE44 thi-1 gyrA96 relA1 λ− | Invitrogen |
| JM109 | (traD36 proAB+ lacIq lacZ M15) endA1 recA1 hsdR17(rK− mK+) mcrA supE44 λ− gyrA96 relA1 (lac proAB) | 67 |
| Other strains | ||
| Bacillus subtilis 168 | Wild type | 68 |
| Staphylococcus aureus Newman | Wild type | 57 |
| Pseudomonas aeruginosa PA14 | Wild type | 58 |
| Burkholderia ambifaria HSJ1 | Wild type | 21 |
| Plasmids | ||
| pJRC125 | Suicide plasmid; Tpr | 6 |
| pMCG19 | pEX18Tp-PheS with an hmqA disrupted by a chloramphenicol resistance cassette (hmqA::cm) | This study |
| pTNS2 | Tn7 transposase-expressing helper plasmid; Ampr | 69 |
| pUC18miniTn7T-Plac-malR | Mobilizable mini-Tn7 vector with the lac promoter (Plac) for IPTG-inducible malR expression (used to construct pUC18-mini-Tn7T-Plac-malR); Kmr, Apr | 61 |
| pUC18miniTn7T-Plac-hmqL | pUC18miniTn7T-LAC-Km containing the BTH_II1576 gene (hmqL); Kmr, Apr | This study |
| pME6010 | pVS1-p15A shuttle vector; Tcr | 33 |
| pME6010-hmqL | pME6010 with the B. thailandensis BTH_II1576 (hmqL) gene | This study |
TABLE 4.
Primers used in this study
| Purpose, primer | Sequence (5′ to 3′) |
|---|---|
| Tn5 mutant identification | |
| hmqA F-1 | GATCTGCCATTGCTTTCCGCAACACG |
| hmqA R-1 | TCAGGCCGCTTGCACGTCG |
| hmqF F-1 | GCTGCATCTGAAGAGCATGGAGC |
| hmqF R-1 | CGTGCTCTCTTCGTGATATCCCATCC |
| hmqC F-1 | TCGGCAATGTGCGAAGCAAGGTC |
| hmqC R-1 | GAGCGGATTGTCGGCAACGAC |
| hmqL-Tn5-Tp-F2 | CGTCATGCCCAATGTGCGCTTG |
| hmqL-Tn5-Tp-R2 | GTTGGTTGACGACTGCGCGAAC |
| Constructing pUC18miniTn7T-Plac-hmqL (BTH_II1576) | |
| hmqL-ORF-F-SacI | ATATTAGAGCTCATGAAAAACAACCAAGTCGATG |
| hmqL-ORF-R-HindIII | ATATTAAAGCTTATTCCCGCTTCGTCCGCCAGC |
| Constructing pMCG19 (hmqA::cm) | |
| hmqAfor | ACGAAGCTTCATCTCTTGCCGCAGCTTGAA |
| hmqArev | ACGGTACCGATCATCAGCCTCGGCTACAC |
| CmFPstI | AAAACTGCAGGTGACGGAAGATCACTTCGCA |
| CmRPstI | AAAACTGCAGGCGTTTAAGGTCAACAATAACTGC |
| Constructing pME6010 hmqL | |
| hmqL-F | CCGAGATCTACCCAATTCATAGACCAGCGTTGC |
| hmqL-R | CCGGGTACCTCATGATGCGTACCTCCGTCGATT |
| Constructing the hmqL::dhfR mutant by natural transformation | |
| hmqL-Tn-for2 | CGTCATGCCCAATGTGCGCTTG |
| hmqL-Tn-rev2 | GTTGGTTGACGACTGCGCGAAC |
B. thailandensis BTH_II1576 (hmqL) mutants were made by transforming a PCR-amplified BTH_II1576::dhfR allele from transposon mutant 56 into the genome of strain BD20 using PCR transformation with a modified protocol similar to that of Thongdee et al. (60). Briefly, shaking B. thailandensis cultures were grown at 37°C to an optical density at 600 nm (OD600) of 0.5, concentrated 20-fold, and distributed to five aliquots of 50 μl. Each aliquot was mixed with 5 μl of gel-extracted hmqL::dhfR PCR product (amplified using hmqL-Tn-for2 and hmqL-Tn-rev2 primers). The cell-DNA mixture was spotted onto solid DM medium (DM liquid medium with 1.5% agar) and incubated at 37°C for 48 h. The DM plate growth was scraped up and collected, washed twice with DM, suspended in 200 μl DM, and spread onto LB agar containing trimethoprim. Mutant strains were verified by PCR amplifying the mutated region and sequencing the PCR product.
For ectopic expression of hmqL in B. thailandensis, this gene was placed under the control of the IPTG-inducible lac promoter in pUC18miniTn7T-LAC-Km (61). To construct this plasmid, we amplified hmqL from the B. thailandensis E264 genome using primers hmqL-ORF-F-SacI and hmqL-ORF-R-HindIII that incorporated the SacI and HindIII restriction enzyme sites, respectively, into the product. The amplicon was cut with SacI and HindIII and ligated to SacI- and HindIII-digested pUC18miniTn7T-Kan-Plac-malR (61) to make pUC18miniTn7T-Plac-hmqL (entirely removing the malR gene). This plasmid was used to transform competent B. thailandensis cells with the helper plasmid pTNS2 as described previously (62). We used PCR to verify insertion of the Plac-hmqL cassette into the attn7 site near glmS1.
We used plasmid pME6010 (33) for expressing the hmqL gene from B. thailandensis in B. ambifaria. The hmqL (BTH_II1576) gene was amplified from the B. thailandensis E264 genome using primers hmqL-F and hmqL-R that incorporated the BglII and KpnI sites into the amplicon. The product was cut with BglII and KpnI and ligated to BglII- and KpnI-digested pME6010 to make pMCG17. B. ambifaria strains with pME6010 plasmids were constructed by electroporation as previously described for B. thailandensis (6).
Liquid cocultures.
Logarithmic-phase overnight starter cultures (OD600 between 0.5 and 1.5) of B. subtilis and B. thailandensis were diluted to an OD600 of 0.05 and combined at a starting ratio of 1:1 in a 10-ml volume of LB in 125-ml baffled flasks. The flasks were incubated with shaking at 250 rpm at 37°C for 24 h before serially diluting and plating on LB agar plates containing gentamicin (to inhibit B. subtilis) or 5% NaCl (to inhibit B. thailandensis) and IPTG as appropriate to enumerate bacterial CFU.
Antimicrobial activity assays.
Antimicrobial activities of B. thailandensis culture fluid were assayed using disc diffusion (for filtered fluid) or outgrowth diffusion (for unclarified fluid) methods. For both methods, inocula for each of the B. thailandensis strains and B. subtilis were prepared by suspending a colony from an LB agar plate into LB broth and growing overnight at 30°C with shaking. B. subtilis overnight culture (100 μl) diluted 1:100 was spread onto LB agar plates and allowed to dry. Filter discs were placed on the B. subtilis lawns and saturated with B. thailandensis cultures that were either centrifuged and filter sterilized through a 0.2-μm membrane (for disc diffusion) or spotted directly onto the B. subtilis lawns (for outgrowth diffusion). The plates were incubated at 30°C for 24 h before observing zones of clearing of the B. subtilis lawns. The outgrowth assays were also conducted similarly on LB agar plates containing 5% NaCl, which inhibits growth of the B. thailandensis strains.
The antimicrobial activities of purified, commercial, or synthesized hydroxyalkylquinoline compounds were assessed using a MIC assay according to a protocol modified from the 2020 guidelines of the Clinical and Laboratory Standards Institute (CLSI) (70). Inocula for each test organism were prepared by suspending a colony from an LB agar plate into tryptic soy broth (TSB) and growing for 3 to 5 h at 37°C with shaking, then adjusting the culture turbidity in TSB to an OD600 of 0.25, roughly the equivalent of a 1.0 McFarland standard (3 × 108 CFU per ml). These cell suspensions were used as inocula for microtiter MIC assays. A 2.5-μl inoculum, which corresponded to 1 × 106 cells, was added to a 100-μl well containing diluted cation-adjusted Mueller-Hinton II broth, and these were incubated with shaking for 24 h at 37°C. The MIC was defined as the lowest concentration of compound (μg/ml) with which bacterial growth in the well was not visible.
Transposon mutagenesis and screen.
Transposon mutagenesis was performed using the EZ-Tn5 <DHFR-1>Tnp Transposome kit (Epicentre) according to the manufacturer’s specifications. Briefly, electrocompetent cells of the B. thailandensis bactobolin-defective mutant BD20 were generated by growing cultures to mid-exponential phase (OD600 of 0.5 to 0.7), collecting them with centrifugation, washing the cell pellet three times in ice-cold 0.5 M sucrose (using 25% the volume of the original culture), and then resuspending the cell pellet in 100 μl ice-cold 0.5 M sucrose. Immediately, 1 μl transposome (the EZ-Tn5 transposon plus the needed transposase for transposition) was added to 50 μl electrocompetent cells in a 0.2-mm electroporation cuvette. This was electroporated with the Bio-Rad Gene Pulser II (using settings of 25 μF, 200 Ω, and 2.5 kV), and the cells were immediately recovered in 1 ml LB broth with shaking at 37°C for 1 h. At the end of the recovery, the culture was diluted 1:25, and 100-μl samples were plated on 20 LB plates with trimethoprim selection (100 μg/ml). The plates were incubated overnight at 37°C. The following day, single colonies were patched onto plates prepared with B. subtilis to screen for antimicrobial activity. Due to the scale required for the screen, we added B. subtilis directly to molten agar used to pour plates, as opposed to spreading B. subtilis lawns after pouring. To prepare the B. subtilis-agar medium, we added 1.43 ml of a stationary-phase B. subtilis culture (overnight growth) to 1 liter of cooled but molten LB agar medium (55 to 60°C), mixed gently, and poured. After a brief period to solidify and dry, plates were used to patch colonies isolated from the EZ-Tn5 <DHFR-1> transposon mutagenesis. Patched plates were incubated overnight at 30°C prior to identifying mutants defective for antimicrobial activity, as determined by reduced zones of B. subtilis growth inhibition compared with the growth of the B. thailandensis parent. Identified candidates were streaked for single B. thailandensis colonies on LB with gentamicin to prevent B. subtilis growth and retested in our assay to confirm the phenotype. Confirmed mutants with no apparent growth defects were subjected to whole-genome sequencing.
Identification of transposon insertion sites.
The transposon insertion locations of five transposon mutants (mutants 7, 14, 31, 32, and 56) were determined by whole-genome resequencing. DNA isolated from the transposon mutant strains was used to make sequencing libraries with 300-bp inserts. The libraries were sequenced on an Illumina MiSeq system using the NEBNext ultra II kit, generating approximately 1 million 200-bp paired-end reads per sample. The paired-end reads were assembled de novo into draft genomes using the SPAdes assembler with standard settings (63). For each de novo assembly, the contig with the Tn5 transposon sequence was located using a nucleotide search in the BLAST+ command line suite with individual blast databases for each transposon mutant (64). Clustal Omega was then used to precisely locate the sequence context of Tn5 insertion in each contig of interest (65). The genomic context for individual transposon insertions was then determined by blasting up- and downstream sequences against a database of all B. thailandensis E264 gene sequences to identify specific loci interrupted by Tn5 insertion. Finally, the raw reads were aligned to the B. thailandensis E264 ATCC 700388 reference genome (accession numbers NC_007650 and NC_007651, downloaded from https://www.burkholderia.com) using Strand NGS (Bangalore, India) software version 3.1.1 to confirm the insertion locus in each mutant. The remaining four transposon mutants (mutants 9, 27, 63, and 68) were assessed by PCR amplifying regions of the hmq locus (primers are given in Table S2 in the supplemental material). Mutations identified by either method were verified by Sanger sequencing of PCR-amplified products.
HMAQ and HMAQ-NO measurements from bacterial cultures.
To measure the production of HMAQ and HMAQ-NO in B. thailandensis cultures, samples were prepared by diluting stationary-phase B. thailandensis cultures to an OD600 of 0.05 into 5 ml of LB in 18-mm culture tubes and growing the cultures for 18 h with shaking at 250 rpm at 30°C. Where necessary, 1 mM IPTG was added to the LB at the beginning of the growth experiment. At 18 h, sample preparation and liquid chromatography-tandem mass spectrometry (LC-MS/MS) analyses were performed as described by Lépine et al. (66), with minor modifications. Briefly, for each sample, 300 μl of grown culture was mixed with 300 μl of high-performance liquid chromatography (HPLC)-grade methanol containing 4 ppm of 5,6,7,8-tetradeutero-4-hydroxy-2-heptylquinoline (HHQ-d4) as an internal standard, vortexed, and centrifuged for 5 min at maximum speed in a microcentrifuge. The supernatant-methanol solution was carefully recovered for analysis. Samples were analyzed by HPLC (Waters 2795; Waters, Mississauga, ON, Canada) equipped with a Kinetex (100- by 3.0-mm) 5-μm EVO C18 reverse-phase LC column (Phenomenex). The detector was a tandem quadrupole mass spectrometer (Quattro premier XE; Waters) equipped with a Z-spray interface using electrospray ionization in positive mode (ESI+). The capillary voltage was set at 3.0 kV, and the cone voltage at 21 V. The source temperature was kept at 120°C. Nitrogen was used as a nebulizing and drying gas at flow rates of 15 and 100 ml · min−1, respectively. Data were collected in scan mode from 130 to 530 Da. An acetonitrile-water gradient containing 1% acetic acid was used. The HPLC flow rate was 400 μl · min−1 split to 40 μl · min−1 by a Valco tee splitter. Quantification of all the HMAQs and HMAQ-NOs was performed using the response factor of 4-hydroxy-3-methyl-2-heptenylquinoline (HMAQ-C7) and using HHQ-d4 as an internal standard.
Data availability.
All strains, plasmids, and sequence data generated by these studies will be made available upon request (Chandler Laboratory [jrchandler@ku.edu], University of Kansas).
Supplementary Material
ACKNOWLEDGMENTS
This work was supported by the NIH through grant number R35GM133572, a pilot award from the COBRE Chemical Biology of Infectious Disease Program (grant number P20 GM113117) to J.R.C., and grant number R01 GM125714 to A.A.D. N.A.E. was supported by a K-INBRE fellowship (grant number P20 GM103418) and a KU undergraduate research award. K.L.A. was supported by award number ASFAHL19F0 from the Cystic Fibrosis Foundation. C.G. was supported by a Discovery grant from the Natural Sciences and Engineering Research Council of Canada (NSERC) under award number RGPIN-2016-04950, and E.D. was supported by a grant from the Canadian Institutes of Health Research (CIHR) under award number MOP-142466. C.G. holds a Fonds de recherche du Québec—Santé (FRQS) Research scholars junior 2 career award. E.D. holds the Canada Research Chair in Sociomicrobiology.
The KU sequencing facility is supported by P20 GM103418 and P20 GM103638.
Footnotes
Supplemental material is available online only.
REFERENCES
- 1.Cheng AC, Currie BJ. 2005. Melioidosis: epidemiology, pathophysiology, and management. Clin Microbiol Rev 18:383–416. doi: 10.1128/CMR.18.2.383-416.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Whitlock GC, Estes DM, Torres AG. 2007. Glanders: off to the races with Burkholderia mallei. FEMS Microbiol Lett 277:115–122. doi: 10.1111/j.1574-6968.2007.00949.x. [DOI] [PubMed] [Google Scholar]
- 3.Dance DA. 2000. Ecology of Burkholderia pseudomallei and the interactions between environmental Burkholderia spp. and human-animal hosts. Acta Trop 74:159–168. doi: 10.1016/s0001-706x(99)00066-2. [DOI] [PubMed] [Google Scholar]
- 4.Van Zandt KE, Greer MT, Gelhaus HC. 2013. Glanders: an overview of infection in humans. Orphanet J Rare Dis 8:131. doi: 10.1186/1750-1172-8-131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Brett PJ, DeShazer D, Woods DE. 1998. Burkholderia thailandensis sp. nov., a Burkholderia pseudomallei-like species. Int J Syst Bacteriol 48(Pt 1):317–320. doi: 10.1099/00207713-48-1-317. [DOI] [PubMed] [Google Scholar]
- 6.Chandler JR, Duerkop BA, Hinz A, West TE, Herman JP, Churchill ME, Skerrett SJ, Greenberg EP. 2009. Mutational analysis of Burkholderia thailandensis quorum sensing and self-aggregation. J Bacteriol 191:5901–5909. doi: 10.1128/JB.00591-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Choi KH, DeShazer D, Schweizer HP. 2006. mini-Tn7 insertion in bacteria with multiple glmS-linked attTn7 sites: example Burkholderia mallei ATCC 23344. Nat Protoc 1:162–169. doi: 10.1038/nprot.2006.25. [DOI] [PubMed] [Google Scholar]
- 8.Lopez CM, Rholl DA, Trunck LA, Schweizer HP. 2009. Versatile dual-technology system for markerless allele replacement in Burkholderia pseudomallei. Appl Environ Microbiol 75:6496–6503. doi: 10.1128/AEM.01669-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Barrett AR, Kang Y, Inamasu KS, Son MS, Vukovich JM, Hoang TT. 2008. Genetic tools for allelic replacement in Burkholderia species. Appl Environ Microbiol 74:4498–4508. doi: 10.1128/AEM.00531-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Lawrenz MB, Fodah RA, Gutierrez MG, Warawa J. 2014. Intubation-mediated intratracheal (IMIT) instillation: a noninvasive, lung-specific delivery system. J Vis Exp 2014:e52261. doi: 10.3791/52261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Duerkop BA, Varga J, Chandler JR, Peterson SB, Herman JP, Churchill ME, Parsek MR, Nierman WC, Greenberg EP. 2009. Quorum-sensing control of antibiotic synthesis in Burkholderia thailandensis. J Bacteriol 191:3909–3918. doi: 10.1128/JB.00200-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Seyedsayamdost MR, Chandler JR, Blodgett JA, Lima PS, Duerkop BA, Oinuma K, Greenberg EP, Clardy J. 2010. Quorum-sensing-regulated bactobolin production by Burkholderia thailandensis E264. Org Lett 12:716–719. doi: 10.1021/ol902751x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Amunts A, Fiedorczuk K, Truong TT, Chandler J, Peter Greenberg E, Ramakrishnan V. 2015. Bactobolin A binds to a site on the 70S ribosome distinct from previously seen antibiotics. J Mol Biol 427:753–755. doi: 10.1016/j.jmb.2014.12.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Biggins JB, Ternei MA, Brady SF. 2012. Malleilactone, a polyketide synthase-derived virulence factor encoded by the cryptic secondary metabolome of Burkholderia pseudomallei group pathogens. J Am Chem Soc 134:13192–13195. doi: 10.1021/ja3052156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Franke J, Ishida K, Hertweck C. 2012. Genomics-driven discovery of burkholderic acid, a noncanonical, cryptic polyketide from human pathogenic Burkholderia species. Angew Chem Int Ed Engl 51:11611–11615. doi: 10.1002/anie.201205566. [DOI] [PubMed] [Google Scholar]
- 16.Trottmann F, Franke J, Richter I, Ishida K, Cyrulies M, Dahse HM, Regestein L, Hertweck C. 2019. Cyclopropanol warhead in malleicyprol confers virulence of human- and animal-pathogenic Burkholderia species. Angew Chem Int Ed Engl 58:14129–14133. doi: 10.1002/anie.201907324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Moule MG, Spink N, Willcocks S, Lim J, Guerra-Assuncao JA, Cia F, Champion OL, Senior NJ, Atkins HS, Clark T, Bancroft GJ, Cuccui J, Wren BW. 2015. Characterization of new virulence factors involved in the intracellular growth and survival of Burkholderia pseudomallei. Infect Immun 84:701–710. doi: 10.1128/IAI.01102-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Park J-D, Moon K, Miller C, Rose J, Xu F, Ebmeier CC, Jacobsen JR, Mao D, Old WM, DeShazer D, Seyedsayamdost MR. 2020. Thailandenes, cryptic polyene natural products isolated from Burkholderia thailandensis using phenotype-guided transposon mutagenesis. ACS Chem Biol 15:1195–1203. doi: 10.1021/acschembio.9b00883. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Seyedsayamdost MR. 2014. High-throughput platform for the discovery of elicitors of silent bacterial gene clusters. Proc Natl Acad Sci U S A 111:7266–7271. doi: 10.1073/pnas.1400019111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Coulon PML, Groleau MC, Déziel E. 2019. Potential of the Burkholderia cepacia complex to produce 4-hydroxy-3-methyl-2-alkyquinolines. Front Cell Infect Microbiol 9:33. doi: 10.3389/fcimb.2019.00033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Vial L, Lépine F, Milot S, Groleau M-C, Dekimpe V, Woods DE, Déziel E. 2008. Burkholderia pseudomallei, B. thailandensis, and B. ambifaria produce 4-hydroxy-2-alkylquinoline analogues with a methyl group at the 3 position that is required for quorum-sensing regulation. J Bacteriol 190:5339–5352. doi: 10.1128/JB.00400-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Diggle SP, Winzer K, Chhabra SR, Worrall KE, Camara M, Williams P. 2003. The Pseudomonas aeruginosa quinolone signal molecule overcomes the cell density-dependency of the quorum sensing hierarchy, regulates rhl-dependent genes at the onset of stationary phase and can be produced in the absence of LasR. Mol Microbiol 50:29–43. doi: 10.1046/j.1365-2958.2003.03672.x. [DOI] [PubMed] [Google Scholar]
- 23.Gallagher LA, McKnight SL, Kuznetsova MS, Pesci EC, Manoil C. 2002. Functions required for extracellular quinolone signaling by Pseudomonas aeruginosa. J Bacteriol 184:6472–6480. doi: 10.1128/jb.184.23.6472-6480.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Déziel E, Lépine F, Milot S, He J, Mindrinos MN, Tompkins RG, Rahme LG. 2004. Analysis of Pseudomonas aeruginosa 4-hydroxy-2-alkylquinolines (HAQs) reveals a role for 4-hydroxy-2-heptylquinoline in cell-to-cell communication. Proc Natl Acad Sci U S A 101:1339–1344. doi: 10.1073/pnas.0307694100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Pesci EC, Milbank JB, Pearson JP, McKnight S, Kende AS, Greenberg EP, Iglewski BH. 1999. Quinolone signaling in the cell-to-cell communication system of Pseudomonas aeruginosa. Proc Natl Acad Sci U S A 96:11229–11234. doi: 10.1073/pnas.96.20.11229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Diggle SP, Matthijs S, Wright VJ, Fletcher MP, Chhabra SR, Lamont IL, Kong X, Hider RC, Cornelis P, Camara M, Williams P. 2007. The Pseudomonas aeruginosa 4-quinolone signal molecules HHQ and PQS play multifunctional roles in quorum sensing and iron entrapment. Chem Biol 14:87–96. doi: 10.1016/j.chembiol.2006.11.014. [DOI] [PubMed] [Google Scholar]
- 27.Xiao G, Déziel E, He J, Lépine F, Lesic B, Castonguay MH, Milot S, Tampakaki AP, Stachel SE, Rahme LG. 2006. MvfR, a key Pseudomonas aeruginosa pathogenicity LTTR-class regulatory protein, has dual ligands. Mol Microbiol 62:1689–1699. doi: 10.1111/j.1365-2958.2006.05462.x. [DOI] [PubMed] [Google Scholar]
- 28.Agarwal A, Kahyaoglu C, Hansen DB. 2012. Characterization of HmqF, a protein involved in the biosynthesis of unsaturated quinolones produced by Burkholderia thailandensis. Biochemistry 51:1648–1657. doi: 10.1021/bi201625w. [DOI] [PubMed] [Google Scholar]
- 29.Lépine F, Milot S, Déziel E, He J, Rahme LG. 2004. Electrospray/mass spectrometric identification and analysis of 4-hydroxy-2-alkylquinolines (HAQs) produced by Pseudomonas aeruginosa. J Am Soc Mass Spectrom 15:862–869. doi: 10.1016/j.jasms.2004.02.012. [DOI] [PubMed] [Google Scholar]
- 30.Drees SL, Ernst S, Belviso BD, Jagmann N, Hennecke U, Fetzner S. 2018. PqsL uses reduced flavin to produce 2-hydroxylaminobenzoylacetate, a preferred PqsBC substrate in alkyl quinolone biosynthesis in Pseudomonas aeruginosa. J Biol Chem 293:9345–9357. doi: 10.1074/jbc.RA117.000789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Piochon M, Coulon PML, Caulet A, Groleau M-C, Déziel E, Gauthier C. 2020. Synthesis and antimicrobial activity of Burkholderia-related 4-hydroxy-3-methyl-2-alkenylquinolones (HMAQs) and their N-oxide counterparts. J Nat Prod 83:2145–2154. doi: 10.1021/acs.jnatprod.0c00171. [DOI] [PubMed] [Google Scholar]
- 32.Lépine F, Déziel E, Milot S, Rahme LG. 2003. A stable isotope dilution assay for the quantification of the Pseudomonas quinolone signal in Pseudomonas aeruginosa cultures. Biochim Biophys Acta 1622:36–41. doi: 10.1016/s0304-4165(03)00103-x. [DOI] [PubMed] [Google Scholar]
- 33.Heeb S, Itoh Y, Nishijyo T, Schnider U, Keel C, Wade J, Walsh U, O’Gara F, Haas D. 2000. Small, stable shuttle vectors based on the minimal pVS1 replicon for use in gram-negative, plant-associated bacteria. Mol Plant Microbe Interact 13:232–237. doi: 10.1094/MPMI.2000.13.2.232. [DOI] [PubMed] [Google Scholar]
- 34.Hays EE, Wells IC, Katzman PA, Cain CK, Jacobs FA, Thayer SA, Doisy EA, Gaby WL, Roberts EC, Muir RD, Carroll CJ, Jones LR, Wade NJ. 1945. Antibiotic substances produced by Pseudomonas aeruginosa. Biol Chem 159:725–750. [Google Scholar]
- 35.Diggle SP, Lumjiaktase P, Dipilato F, Winzer K, Kunakorn M, Barrett DA, Chhabra SR, Camara M, Williams P. 2006. Functional genetic analysis reveals a 2-alkyl-4-quinolone signaling system in the human pathogen Burkholderia pseudomallei and related bacteria. Chem Biol 13:701–710. doi: 10.1016/j.chembiol.2006.05.006. [DOI] [PubMed] [Google Scholar]
- 36.Machan ZA, Taylor GW, Pitt TL, Cole PJ, Wilson R. 1992. 2-Heptyl-4-hydroxyquinoline N-oxide, an antistaphylococcal agent produced by Pseudomonas aeruginosa. J Antimicrob Chemother 30:615–623. doi: 10.1093/jac/30.5.615. [DOI] [PubMed] [Google Scholar]
- 37.Smirnova IA, Hagerhall C, Konstantinov AA, Hederstedt L. 1995. HOQNO interaction with cytochrome b in succinate:menaquinone oxidoreductase from Bacillus subtilis. FEBS Lett 359:23–26. doi: 10.1016/0014-5793(94)01442-4. [DOI] [PubMed] [Google Scholar]
- 38.Nguyen AT, Jones JW, Ruge MA, Kane MA, Oglesby-Sherrouse AG. 2015. Iron depletion enhances production of antimicrobials by Pseudomonas aeruginosa. J Bacteriol 197:2265–2275. doi: 10.1128/JB.00072-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Mashburn LM, Jett AM, Akins DR, Whiteley M. 2005. Staphylococcus aureus serves as an iron source for Pseudomonas aeruginosa during in vivo coculture. J Bacteriol 187:554–566. doi: 10.1128/JB.187.2.554-566.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Korgaonkar A, Trivedi U, Rumbaugh KP, Whiteley M. 2013. Community surveillance enhances Pseudomonas aeruginosa virulence during polymicrobial infection. Proc Natl Acad Sci U S A 110:1059–1064. doi: 10.1073/pnas.1214550110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Mitchell G, Seguin DL, Asselin AE, Déziel E, Cantin AM, Frost EH, Michaud S, Malouin F. 2010. Staphylococcus aureus sigma B-dependent emergence of small-colony variants and biofilm production following exposure to Pseudomonas aeruginosa 4-hydroxy-2-heptylquinoline-N-oxide. BMC Microbiol 10:33. doi: 10.1186/1471-2180-10-33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Fugere A, Lalonde Seguin D, Mitchell G, Déziel E, Dekimpe V, Cantin AM, Frost E, Malouin F. 2014. Interspecific small molecule interactions between clinical isolates of Pseudomonas aeruginosa and Staphylococcus aureus from adult cystic fibrosis patients. PLoS One 9:e86705. doi: 10.1371/journal.pone.0086705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Hoffman LR, Déziel E, D’Argenio DA, Lepine F, Emerson J, McNamara S, Gibson RL, Ramsey BW, Miller SI. 2006. Selection for Staphylococcus aureus small-colony variants due to growth in the presence of Pseudomonas aeruginosa. Proc Natl Acad Sci U S A 103:19890–19895. doi: 10.1073/pnas.0606756104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Lightbown JW. 1954. An antagonist of streptomycin and dihydrostreptomycin produced by Pseudomonas aeruginosa. J Gen Microbiol 11:477–492. doi: 10.1099/00221287-11-3-477. [DOI] [PubMed] [Google Scholar]
- 45.Hooi DS, Bycroft BW, Chhabra SR, Williams P, Pritchard DI. 2004. Differential immune modulatory activity of Pseudomonas aeruginosa quorum-sensing signal molecules. Infect Immun 72:6463–6470. doi: 10.1128/IAI.72.11.6463-6470.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Bredenbruch F, Geffers R, Nimtz M, Buer J, Haussler S. 2006. The Pseudomonas aeruginosa quinolone signal (PQS) has an iron-chelating activity. Environ Microbiol 8:1318–1329. doi: 10.1111/j.1462-2920.2006.01025.x. [DOI] [PubMed] [Google Scholar]
- 47.Wu Y, Seyedsayamdost MR. 2017. Synergy and target promiscuity drive structural divergence in bacterial alkylquinolone biosynthesis. Cell Chem Biol 24:1437–1444.e3. doi: 10.1016/j.chembiol.2017.08.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Chandler JR, Heilmann S, Mittler JE, Greenberg EP. 2012. Acyl-homoserine lactone-dependent eavesdropping promotes competition in a laboratory co-culture model. ISME J 6:2219–2228. doi: 10.1038/ismej.2012.69. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Benomar S, Evans KC, Unckless RL, Chandler JR. 2019. Efflux pumps in Chromobacterium species increase antibiotic resistance and promote survival in a co-culture competition model. Appl Environ Microbiol 85:e00908-19. doi: 10.1128/AEM.00908-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Evans KC, Benomar S, Camuy-Velez LA, Nasseri EB, Wang X, Neuenswander B, Chandler JR. 2018. Quorum-sensing control of antibiotic resistance stabilizes cooperation in Chromobacterium violaceum. ISME J 12:1263–1272. doi: 10.1038/s41396-018-0047-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Wang Y, Hoffmann JP, Chou CW, Honer Zu Bentrup K, Fuselier JA, Bitoun JP, Wimley WC, Morici LA. 2020. Burkholderia thailandensis outer membrane vesicles exert antimicrobial activity against drug-resistant and competitor microbial species. J Microbiol 58:550–562. doi: 10.1007/s12275-020-0028-1. [DOI] [PubMed] [Google Scholar]
- 52.Beveridge TJ. 1999. Structures of gram-negative cell walls and their derived membrane vesicles. J Bacteriol 181:4725–4733. doi: 10.1128/JB.181.16.4725-4733.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Li Z, Clarke AJ, Beveridge TJ. 1998. Gram-negative bacteria produce membrane vesicles which are capable of killing other bacteria. J Bacteriol 180:5478–5483. doi: 10.1128/JB.180.20.5478-5483.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Conrad M, Limpach L. 1887. Syntheses of quinoline derivatives using acetoacetic ester. Rep German Chem Soc 20:944–948. (In German.) [Google Scholar]
- 55.Salvaggio F, Hodgkinson JT, Carro L, Geddis SM, Galloway WRJD, Welch M, Spring DR. 2016. The synthesis of quinolone natural products from Pseudonocardia sp. Eur J Org Chem 2016:434–437. doi: 10.1002/ejoc.201501400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Woschek A, Mahout M, Mereiter K, Hammerschmidt F. 2007. Synthesis of 2-heptyl-1-hydroxy-4(1H)-quinolone—unexpected rearrangement of 4-(alkoxycarbonyloxy)quinoline N-oxides to 1-(alkoxycarbonyloxy)-4(1H)-quinolones. Synthesis 2007:1517–1522. doi: 10.1055/s-2007-966020. [DOI] [Google Scholar]
- 57.Duthie ES, Lorenz LL. 1952. Staphylococcal coagulase; mode of action and antigenicity. J Gen Microbiol 6:95–107. doi: 10.1099/00221287-6-1-2-95. [DOI] [PubMed] [Google Scholar]
- 58.Rahme LG, Stevens EJ, Wolfort SF, Shao J, Tompkins RG, Ausubel FM. 1995. Common virulence factors for bacterial pathogenicity in plants and animals. Science 268:1899–1902. doi: 10.1126/science.7604262. [DOI] [PubMed] [Google Scholar]
- 59.Chang AC, Cohen SN. 1978. Construction and characterization of amplifiable multicopy DNA cloning vehicles derived from the P15A cryptic miniplasmid. J Bacteriol 134:1141–1156. doi: 10.1128/JB.134.3.1141-1156.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Thongdee M, Gallagher LA, Schell M, Dharakul T, Songsivilai S, Manoil C. 2008. Targeted mutagenesis of Burkholderia thailandensis and Burkholderia pseudomallei through natural transformation of PCR fragments. Appl Environ Microbiol 74:2985–2989. doi: 10.1128/AEM.00030-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Klaus JR, Deay J, Neuenswander B, Hursh W, Gao Z, Bouddhara T, Williams TD, Douglas J, Monize K, Martins P, Majerczyk C, Seyedsayamdost MR, Peterson BR, Rivera M, Chandler JR. 2018. Malleilactone is a Burkholderia pseudomallei virulence factor regulated by antibiotics and quorum sensing. J Bacteriol 200:e00008-18. doi: 10.1128/JB.00008-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Choi KH, Mima T, Casart Y, Rholl D, Kumar A, Beacham IR, Schweizer HP. 2008. Genetic tools for select-agent-compliant manipulation of Burkholderia pseudomallei. Appl Environ Microbiol 74:1064–1075. doi: 10.1128/AEM.02430-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Lesin VM, Nikolenko SI, Pham S, Prjibelski AD, Pyshkin AV, Sirotkin AV, Vyahhi N, Tesler G, Alekseyev MA, Pevzner PA. 2012. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477. doi: 10.1089/cmb.2012.0021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL. 2009. BLAST+: architecture and applications. BMC Bioinformatics 10:421. doi: 10.1186/1471-2105-10-421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Soding J, Thompson JD, Higgins DG. 2011. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7:539. doi: 10.1038/msb.2011.75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Lépine F, Milot S, Groleau MC, Déziel E. 2018. Liquid chromatography/mass spectrometry (LC/MS) for the detection and quantification of N-acyl-L-homoserine lactones (AHLs) and 4-hydroxy-2-alkylquinolines (HAQs). Methods Mol Biol 1673:49–59. doi: 10.1007/978-1-4939-7309-5_4. [DOI] [PubMed] [Google Scholar]
- 67.Yanisch-Perron C, Vieira J, Messing J. 1985. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene 33:103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]
- 68.Burkholder PR, Giles NH Jr. 1947. Induced biochemical mutations in Bacillus subtilis. Am J Bot 34:345–348. doi: 10.1002/j.1537-2197.1947.tb12999.x. [DOI] [PubMed] [Google Scholar]
- 69.Choi KH, Gaynor JB, White KG, Lopez C, Bosio CM, Karkhoff-Schweizer RR, Schweizer HP. 2005. A Tn7-based broad-range bacterial cloning and expression system. Nat Methods 2:443–448. doi: 10.1038/nmeth765. [DOI] [PubMed] [Google Scholar]
- 70.Clinical and Laboratory Standards Institute (CLSI). 2020. Performance standards for antimicrobial susceptibility testing, 30th ed. CLSI supplement M100. Clinical and Laboratory Standards Institute, Wayne, Pennsylvania. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All strains, plasmids, and sequence data generated by these studies will be made available upon request (Chandler Laboratory [jrchandler@ku.edu], University of Kansas).



