Abstract
The muscle protein titin plays a crucial role in passive elasticity and the disordered PEVK region within titin is central to that function. The PEVK region is so named due to its high proline, glutamate, valine and lysine content and the high charge density in this region results in a lack of organized structure within this domain. The PEVK region is highly extensible but the molecular interactions that contribute to the elastic nature of the PEVK still remain poorly described. The PEVK region is formed by two unique sequence motifs. The PPAK motif is a 26 to 28 amino acid sequence that contains a mixture of charged and hydrophobic residues and is the primary building block for the PEVK region. Poly-E sequence motifs vary in length and contain clusters of 3–4 glutamic acids distributed throughout the motif. In this study, we derived two 28-residue peptides from the human titin protein sequence and measured their structural characteristics over a range of pHs. Our results demonstrate that the poly-E peptide undergoes a shift from a more rigid and elongated state to a more collapsed state as pH decreases with the midpoint of this transition being at pH ~5.5. Interestingly, a similar conformational shift is not observed in the PPAK peptide. These results suggest that the poly-E motif might provide a nucleating site for the PEVK when the muscle is not in an extended state.
Keywords: Titin, PEVK, Circular dichroism, Intrinsically disordered proteins, Secondary structure, Peptide charge repulsions
Highlights
-
•
Poly-E peptides have a more extended conformation at pH 7.0 than PPAK peptides.
-
•
Poly-E peptides assume a more relaxed conformation below pH 6.
-
•
PPAK peptides do not show any conformational sensitivity to changes in pH.
1. Introduction
Titin is a giant elastic protein found in both striated and cardiac muscle that spans half of a sarcomere, from Z disc to M line [1]. It is known that titin plays a role in passive elasticity, sarcomere assembly and it acts as a sensor of sarcomeric stress and strain, but there are still gaps in the understanding of how titin performs these various functions [2,3]. The majority of titin's elastic function occurs within the I-band region of the protein [4]. This region is composed of two major structural motifs; two regions of repeated immunoglobulin (Ig) domains and a disordered region called the PEVK domain that sits between the two Ig regions [5,6]. PEVK derives its name from the fact that nearly 75% of the amino acids are either proline (P), glutamic acid (E), valine (V) or lysine (K) [7].
Two different sequence motifs have been identified within the PEVK region. The majority of the PEVK region is made up of PPAK motif segments, which are 26–28 residues in length composed of a mixture of hydrophobic and charged amino acids, mainly lysine [8]. PPAK motifs generally exist in clusters of 2–12 repeated sequence blocks [7]. Separating the PPAK clusters are poly-E motifs, which consists of clusters of glutamate residues. These regions contain approximately 45% glutamic acid, in clusters of 3–4 consecutive glutamic acid residues that are separated by 2–3 hydrophobic residues [7,8]. This results in a high negative charge density within poly-E segments.
One of titin's primary functions is to generate passive tension during muscle elongation, which helps maintain uniform sarcomere lengths during stretching and also to restore sarcomere length during muscle relaxation [14]. During an eccentric stretch, the regions of consecutive Ig domains extend into a linear arrangement under small passive forces and short sarcomere lengths [9]. At moderate to long sarcomere lengths, passive force is predominately controlled by the extensible PEVK region [9,10]. While there is a basic understanding of how titin regulates passive tension, a clear molecular level understanding of how PEVK generates the observed changes in force during eccentric contractions is still lacking.
Previous work has shown that the PEVK region is composed predominately of random coil sequence with a small amount of polyproline II (PII) type helices [11]. The most common way to model the elastic function of the PEVK region is as an entropic spring, where conformational entropy drives the observed passive force [12]. However, there is evidence that the PEVK region exhibits enthalpic characteristics at higher forces and titin-based myofibril stiffness depends on sarcomere ionic strength [9]. These observations suggest that PEVK might not act as a pure entropic spring [9,13] and highlight the need for a clearer understanding of the relationship between PEVK's function and any transient structural elements that PEVK might form.
Previous work has shown that the charged nature of PEVK is important for its function, but these studies were conducted at physiological pH (pH~7.2–7.4). Metabolically active muscle induced decreased pH within the muscle and the impact of these changes on the PEVK region has not been explored [14]. We hypothesized that the poly-E regions might undergo pH-dependent conformational changes due to the presence of clustered glutamate residues within the region. To test this hypothesis, we synthesized 28-amino acid peptides using PPAK and poly-E sequences from the human PEVK and explored their structural characteristics as a function of pH. Using a combination of CD, fluorescence and size exclusion chromatography, our results demonstrate that the poly-E peptide undergoes a conformational change around pH 5.5 while the PPAK peptide does not exhibit a similar pH dependency. These results highlight the potential impact of shifts in pH on PEVK function.
2. Materials and methods
All chemicals used in this study were purchased from standard chemical suppliers such as Fisher Scientific. Peptides used in this study were synthesized by AAPPtec and purified to >95% purity. The peptide sequences used in this study were:
PPAKVPEVPKKPVPEKKVPVPAPKKVEA (P1).
EVTEEPEEEPISEEEIPEEPPSIEEVEE (P2).
EVETEPEPEIESEIEPEPEPESEIEVEE (P3).
EVTPIPEEEPISPISIPEEPPSIPIVEE (P4).
EVTPIPPISPISEEEIPEEPPSIPIVEE (P5).
EVTPIPPISPISPISIPISPPSIPIVEE (P6).
The P1 and P2 peptides were based on exon sequences in the PEVK region of human titin. Peptides P3–P6 were derived using the P2 peptide sequence. Peptides were synthesized as both a tryptophan containing and tryptophan-free variants to aid with spectroscopic characterization of the structure. Tryptophan containing peptides will be referred to as P1+W, P2+W, P3+W, P4+W, P5+W and P6+W. Peptides were stored as a lyophilized powder and rehydrated for experiments.
2.1. Circular dichroism measurements
Tryptophan-free (∆Trp) peptides were reconstituted in milliQ water at a concentration of 600 μM and diluted to a final concentration of 0.1 mg/ml in 20 mM potassium phosphate at a range of pHs. Samples were equilibrated at 20 °C for an hour and spectra were measured using a JASCO J-810 circular dichroism spectropolarimeter equipped with a JASCO PFD-425S peltier controller. A quartz cuvette with a 1 mm pathlength was used and spectra were collected in triplicate using a 1 nm bandwidth and a 1 nm data pitch. Each spectrum was background subtracted and converted to molar ellipticity.
2.2. Fluorescence spectroscopy
Peptide samples were prepared using tryptophan containing peptides following the procedure as described above for the circular dichroism experiments. Fluorescence spectra were measured using a Molecular Devices SpectraMax M3 Multi-Mode Microplate Reader equipped with SoftMax Pro 7.0.3. Samples were excited at 280 nm and emission spectra were collected from 300 to 450 nm. The Center of Mass (CoM) of each spectrum was calculated using Equation (1), where I is intensity and is the wavenumber:
| (1) |
CoM was plotted as a function of pH.
2.3. Acrylamide quenching
Peptide samples were prepared using the tryptophan containing peptides as described above for the circular dichroism experiments. Samples were treated with acrylamide at final concentrations of 25, 50, 100, 150 and 200 mM at a range of pHs. Tryptophan emission spectra were collected as described above. Stern-Volmer plots were developed using Equation (2), where is tryptophan emission in the absence of acrylamide, is tryptophan emission in the presence of acrylamide, is the Stern-Volmer constant and is the acrylamide concentration.
| (2) |
2.4. Size Exclusion Chromatography
Peptide solutions were made by reconstituting tryptophan containing peptides to a final concentration of 1 mg/mL in a 20 mM potassium phosphate, 150 mM KCl buffer at a range of pHs. Samples were equilibrated at 20 °C for a minimum of an hour before separating the samples by Size Exclusion Chromatography-HPLC. The samples were separated on a TOSOH TSKgel G2500PWXL columns using a Waters 2695 Separation Module with Waters 2996 Photodiode Array Detector. The column was equilibrated with the appropriate buffer and all samples were run in triplicate. The point of maximum absorbance was used as the retention time for each sample.
3. Results
3.1. The poly-E peptide undergoes a pH dependent conformational change
Titin's PEVK region has been an area of interest because this region's important role in passive tension but the mechanism of how PEVK generates passive tension is poorly understood. The distribution of the two sequence motifs found in the PEVK region presumably plays a role in this function. While both sequence motifs have high charge densities, PPAK motifs have their charges distributed throughout the motif's sequence while poly-E motifs consist of clusters of glutamates separated by several hydrophobic residues [8]. Poly-E motifs also have a higher overall average charge density (45% glutamic acid for poly-E compared to 19% lysine for PPAK [8]). Based on the high charge density in this region, we hypothesized that the PEVK region might undergo pH-dependent conformational changes as the charged amino acids change between protonated and deprotonated states. This hypothesis was tested using two 28 amino acid peptides based on PPAK and poly-E sequences from human titin. It was expected that the poly-E peptides would be more likely to exhibit a pH-dependent conformational change since they have a higher charge density. The poly-E peptide sequence used in these studies was selected because its charge density is consistent with the average poly-E charge density. It should also be noted that poly-E regions are longer than the PPAK sequence, but we wanted to compare peptides of similar length and so a 28 amino acid section of poly-E motif sequence was used.
We initially tested our hypothesis by comparing retention time of the peak of maximum absorbance during size exclusion chromatography (SEC) at varying pHs. Both peptides were dissolved in mobile phase and separated on a SEC column equilibrated with a 20 mM phosphate buffer at the corresponding pH. We initially compared the conformation of the two peptides at pH 7.0 and noticed that the poly-E peptide exhibited a earlier retention time than the PPAK peptide. This suggests that the poly-E peptide exhibits a larger apparent hydrodynamic radius than PPAK even though the two peptides have the same number of amino acids, consistent with the model the poly-E has more elongated conformation (Fig. 1). Varying the pH of the mobile phase did not significantly alter the retention time of the PPAK peptide, indicating that pH does not influence the conformational state of the PPAK peptide over the pH range used in these experiments. In contrast, a shift to longer retention times was observed for the poly-E peptide as the pH of the mobile phase was decreased, suggesting that the poly-E peptide has a more compact conformation at lower pH. The observed shift in retention time begins around pH 6 and is believed to be induced by changes in the protonation of the glutamic acids in the peptide. The location of the transition was surprising since it is higher than the pKa for the side chain of free glutamic acid (pka = 4.0) [15], suggesting that either the high charge density or glutamate clustering is potentially influencing the pKa of the glutamate side chains.
Fig. 1.
Poly-E peptides exhibit pH-dependent changes in retention times. Peptides were separated on a TSK-GEL G2500PWXL column and the retention times were determined as a function of varying pH. The PPAK peptide (P1+W) shows no significant change in retention time. All poly-E peptides (P2+W – P6+W) have increased retention time at lower pHs. The P2+W and P3+W peptides show similar shifts, suggesting overall charge density is more important than clustering.
To determine which factor was more important, we repeated this experiment using a series of peptides derived from the poly-E peptide sequence. The P3+W peptide has an identical charge density as the P2+W peptide except the glutamates are evenly distributed throughout the peptide. Both P2+W and P3+W exhibited a similar pH sensitivity consistent with charge density and not clustering being the critical factor driving the magnitude of the conformational change. This is supported by the reduced pH sensitivity observed for the other modified peptides (P4+W – P6+W). It is interesting to note that all three peptides with reduced charge density had longer retention times in the physiological pH range and that there was a linear correlation between the magnitude of the change and the overall charge density of the peptide.
3.2. Poly-E peptide has a decreased fluorescence at lower pHs
The observed increased retention time for the poly-E peptide at lower pHs suggests that the peptide forms a more collapsed state, presumably due to decreased charge repulsion. If this model is accurate, we predicted that fluorescence intensity would decrease at lower pHs since a higher degree of conformational freedom would result in increased non-radiative decay of tryptophans in the excited state. Fluorescence spectra were measured for both the poly-E and PPAK peptides at a range of pHs and the emission intensity at 360 nm for each sample was compared to test this hypothesis. Emission intensity of P2+W decreased as pH decreases, with the largest changes occurring between pH 5 and 6 (Fig. 2), consistent with our SEC experiments. Each series of samples were normalized to facilitate comparison of the samples using the intensity at pH 3 as the reference. The intensity change exhibited a sigmoidal shape consistent with a transition between two unique conformational states (e.g. folded to unfolded states in classical globular protein unfolding). In this case, neither peptide is expected to exist in a folded conformation, but the observed transition is likely due to a change in most populated disordered conformation of the P2+W peptide. A change in fluorescence intensity was also observed for the PPAK peptide (P1+W) but the transition was linear and is consistent with the changes in fluorescence observed for free tryptophan. We therefore attribute the change in intensity in P1+W to pH-dependent shifts in tryptophan fluorescence rather than reflective of conformational changes in the peptides.
Fig. 2.
P2+W peptide decreases fluorescence intensity with decreased pH. Tryptophan fluorescence was measured for the P1+W and P2+W peptides at varying pH and the intensity at 360 nm was plotted as a function of pH. The intensity of the P2+W peptide exhibits a sigmoidal shape while the P1+W intensity is best fit as a line.
To further validate this conclusion, we plotted the Center of Mass (CoM) for each spectrum since this reduces pipet errors and other factors that can influence the accuracy of fluorescence intensity. CoM exhibits a rapid red shift of both peptides between pH 3 and 4, in contrast to the observed shift in intensity over the same pH range (Fig. 3). To facilitate comparison between the samples, each data set was normalized (Fig. 3 Inset). A comparable shift is not observed in free tryptophan, which suggests that both peptides undergo a pH dependent conformational shift at low pH. Both peptides exhibit a red shift between pH 4 and pH 5 as well though, unlike between pH 3 and 4 where the shift is similar for both peptides, there is a significant shift for P2+W but only a small shift for P1+W and then the CoM doesn't undergo any additional significant shifts between pH 5 and pH 8. In contrast, P2+W continues to undergo red-shifts in the CoM through pH 6 and then plateaus through pH 8. These results suggest that both peptides undergo pH-dependent conformational shifts between pH 3 and 4. However, P2+W continues to shift conformation between pH 4 and 6 while P1+W does not undergo comparable shifts. This is consistent with P2+W undergoing sequence-dependent conformational shifts that P1+W is not capable of undergoing. Overall, these results support the model that the poly-E peptide undergoes a structural transition to a more flexible conformation at lower pHs.
Fig. 3.
P2+W peptide shows a red shift with decreased pH. P2+W exhibits CoM shifts between pH 3 and pH 6. In contrast, P1+W has a significant pH shift between pH 3 and 4, followed by a small shift between pH 4 and 5 and then no significant CoM after pH 5. Insert. Normalized Center of Mass curves showing magnitude of CoM shifts as a function of pH.
3.3. Acrylamide quenching of poly-E is increased at higher pHs
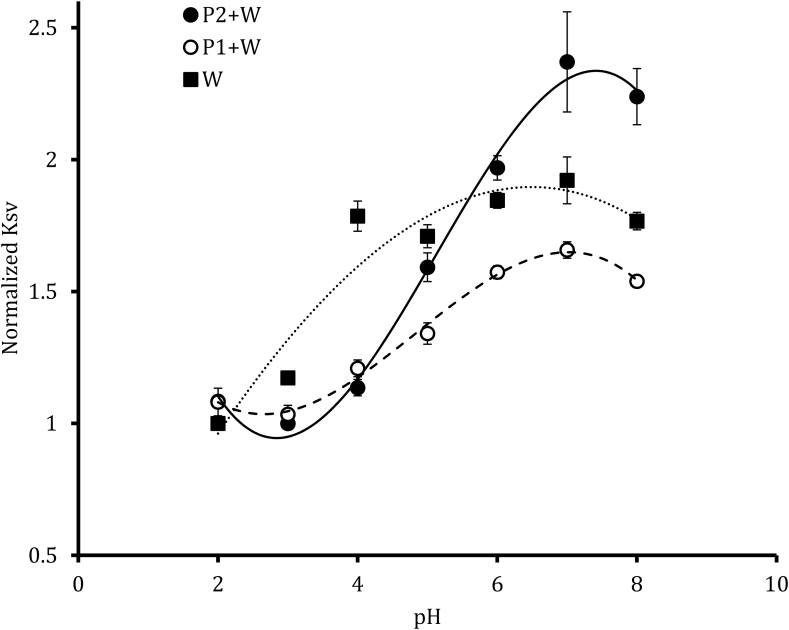
Our initial experiments suggested that the poly-E peptide has two conformational states; an extended conformation at higher pH and a more compact state at lower pH. Based on this model, we predicted that the tryptophan on the peptide would be more sensitive to quenching at higher pHs due to the extended conformation of the peptide. To test this model, we performed an acrylamide quenching experiment, where each peptide was mixed with a range of acrylamide concentrations (50–250 mM) over a range of pHs and the Stern-Volmer quenching constant was measured (Fig. 4). As can be seen, there is a shift in P1+W that is consistent with the shifts observed in free tryptophan. The P2+W peptide exhibits similar quenching at lower pH, consistent with the peptide being in a similar conformation as the P1+W peptide. However, at higher pHs, the P2+W peptide exhibits a higher degree of quenching, which is consistent with this peptide being in a more extended conformation, supporting our proposed model.
Fig. 4.
P2+W peptide undergoes more quenching at higher pHs. The values of P2+W shows a sigmoidal shift with pH where a major transition occurs between pH 4 and pH 7. The of free tryptophan amino acid remain constant between that region of pH.
Poly-E peptide shifts to a more pre-molten globule-like conformation at lower pH.
Circular Dichroism (CD) was used to further validate that the observed changes in P2 fluorescence intensity represented a conformational change in the peptide. Disordered peptides exist in an ensemble of states and their CD spectra are characterized by a minimum at 200 nm without many other structural features. It has been shown that plotting molar ellipticity at 200 nm (ME200) verse molar ellipticity at 222 nm (ME222) can provide some structural insights into IDPs. Coil-like conformations have a ME200 = −19,000 ± 2800 deg.cm2.dmol−1 and a ME222 = −1700 ± 700 deg.cm2.dmol−1 while pre-molten globule-like conformation have a ME200 = −10,700 ± 1300 deg.cm2.dmol−1 and a ME222 = −3900 ± 1100 deg.cm2.dmol−1 [16]. CD can provide insights into an IDP's conformational state it they resemble one of these two conformational states.
CD spectra for both the poly-E and PPAK peptides were collected from 250 to 190 nm at a range of pHs. There was a clear change in the molar ellipticity of the poly-E peptide at 200 nm as pH decreases (Fig. 5A). This transition begins at pH 6.0 (Fig. 5B), similar to what was observed in our other experiments, further supporting the model of a pH dependent conformational change. There was some variation in the molar ellipticity of the PPAK peptide at 200 nm over the range of pHs tested but the observed differences were not statistically significant and are believed to be a function of the dynamic nature of IDPs.
Fig. 5.
P2 peptide undergoes a pH dependent conformational change. A. CD spectra of P2 peptide from 250 nm to 190 nm. Spectra were collected between pH 2 and 9 at increments of 1 pH unit. The arrow represents the change in the minima at 200 nm with decreasing pH. B. Molar ellipticity at 200 nm for P1 and P2 peptides. P2 peptide decreases molar ellipticity between pH 4 and 6 while P1 peptide does not exhibit a similar conformational change.
The ME200 verse ME222 was plotted for the P1 and P2 peptides at each pH to determine if either peptide exhibited either pre-molten globule-like or coil-like characteristics (Fig. 6). The P1 peptide data was clustered together and did not demonstrate a pH dependence. The data also did not match either structural category, though the data more closely resembles a coil-like conformation than a pre-molten globule. A similar analysis of the poly-E spectra suggests that this peptide likely exists in a coil-like conformation and there is a clear shift in the coil-like nature of the peptide below pH 6. The coil-like conformational state is thought to contain ~30% PII in contrast to the pre-molten globule state, which is thought to contain ~10% PII [16]. The observed shift below pH 6 is in the direction of the pre-molten globule state, consistent with a decrease in PII character and a shift from a more elongated structure to a more collapsed structure observed in our other experiments.
Fig. 6.
P2 peptide exhibits coil-like structure that becomes more pre-molten globule like at lower pHs. The ME200 verse ME222 values for the P2 peptide are in the region associated with a coil-like structure while the P1 values are in a region of undefined structure on the plot. The P2 conformation exhibits a shift below pH 6 toward the region associated with a pre-molten globule conformation, potentially due to reduced PII helical content.
4. Discussion
The PEVK domain of the giant elastomeric protein titin is a primary contributor to passive force at moderate to long sarcomere lengths [5,9]. While it is fairly well established that this region plays a critical role in passive tension, there remain many unanswered questions regarding the mechanism of PEVK function. One of the unexplored aspects of this region is how pH impacts the structural characteristics of the PEVK region, which is the focus of this study. Metabolically active muscle experiences a decrease in pH due to the CO2 that is generated during aerobic respiration and the impact of pH shifts on the PEVK region is unknown. The highly charged nature of the PEVK region led us to hypothesize that decreased pH might result in changes in ionization state of the region that could alter conformational states.
In this study, we developed two 28-amino acid peptides based on the PPAK and poly-E motifs within the PEVK region and assessed the impact of decreased pH on the conformational flexibility of both peptides using a combination of SEC, fluorescence and circular dichroism. The PPAK peptide exhibited a minimal conformational change under the pH range that was tested. This is not very surprising since all the ionizable side chains in this sequence are lysine, which has a pKa of 10.5 in the free state and we only tested up to pH 9.0 since muscles do not generally experience alkaloidosis. In contrast, SEC showed that the poly-E peptide has an elongated conformation at physiological pH that switches into a more collapsed state at more acidic pHs. This result is supported by fluorescence, acrylamide quenching and circular dichroism data which all show similar conformational changes between pH 5 and pH 6. The shift observed in the CD experiments suggests that there could be a change in the PII content of the poly-E peptide, which would be consistent with the observed change in hydrodynamic radius in our SEC experiments. Taken together, these results show that the poly-E region can undergo pH-dependent conformational changes that could be physiologically significant.
4.1. Impact of pH shifts on titin function
The cellular environment within the sarcomere is a complex ionic mixture that fluctuates in composition as a function of the activation state of the muscle [17]. One of the major changes is an increase in calcium concentration to open binding sites for the myosin heads on the thin filament [18]. The crossbridge cycling that occurs in active muscle results in a high level of metabolic activity and a subsequent decrease in pH [19]. The high charge density in the PEVK region makes this region especially susceptible to these ionic and pH fluctuations. Initial attempts to model force spectroscopy data collected on PEVK constructs used the wormlike chain model, which adequately models the behavior of the PEVK region at low to moderate extensions as an entropic spring [13]. However, this approach does not accurately model PEVK behavior at high force unless an enthalpic contribution is included to account for electrostatic interactions as part of the model [9].
Combining the results from this study with what is known about the spring-like characteristics of the PEVK region suggest an intriguing functional model. High force extensions of PEVK include an enthalpic contribution, which presumably comes from interactions between lysines in the PPAK motifs and glutamates in the poly-E regions. However, the data from this study suggests that if the high force stretch were to occur in a metabolically active muscle, where acidotic conditions might exist, that these electrostatic interactions might be disrupted due to protonation of the glutamic acid residues and the PEVK region might act purely as an entropic spring. It should be noted that the structural transition measured in this study occurs at the very edge of the pH range that has been measured in active muscle [19] and therefore would only occur under extreme conditions, assuming that the conformational shift occurs in the same pH range in a complete poly-E motif sequence. However, since the poly-E peptide used in this study is less than one-half of the actual motif it is derived from (28 amino acid verse 55 amino acids), it is possible that the full motif might exhibit a structural transition at a higher pH range than was observed in this study. A complete assessment of the relationship between charge density and sequence length will be necessary to understand which motifs are likely to exhibit pH sensitivity.
4.2. Role for the rigid nature of the poly-E sequence
A striking result from this study is the conformational difference between the PPAK and poly-E peptides at physiological pH. There is a significant difference in retention time between the two peptides, even though they have an identical number of amino acids, demonstrating that the poly-E motif has a more elongated conformation than the PPAK motif. The predominate motif within the PEVK region is the PPAK motif occupying 75% of PEVK's amino acid content with poly-E motifs occurring between groups of 2–12 PPAK motifs [8]. Unlike the PPAK motif which is 26–28 amino acids long, poly-E motifs range from 26 amino acids up to 134 amino acids and have a glutamate content as little as 32% and up to 48%, with the average glutamate content being approximately 45%. Because of the amino acid distribution, the overall charge of PEVK is slightly negative (23% acidic and 19%basic amino acid content [8]). The sequence variability in the PEVK region most likely has functional implications as poly-E regions with lower glutamate content likely have more conformational flexibility compared to regions with high glutamate content. There is significant alternative splicing in the PEVK region so a careful comparison of which exons are included in particular types of muscle types might provide interesting insights into the relationship between splice variants and function of particular muscle types. The cardiac N2B isoform has a shorter PEVK region than N2BA isoform or the N2A isoform in the soleus muscle (163 residues, 600–800 residues and 2174 residues respectively) [20]. The larger PEVK regions have a net acidic character due to the presence of the poly-E regions. For example, the PEVK region from soleus muscle has a predicted pI of 5.1 [21]. In constrast, the N2B PEVK region, which doesn't contain any poly-E segments, has a predicted pI of 9.7 [21] and therefore most likely to bind actin (see below).There is likely functional significance to the absence of poly-E regions in the N2B isoform since the ratio of N2B:N2BA changes during development and during heart failure. In the case of dilated cardiomyopathy, the N2B:N2BA ratio increases by 63%, resulting in increased passive stiffness [22,23].
The apparent rigid nature of the poly-E peptide may also have particular significance when it comes to the high passive tension measured in long, lengthening stretches. Studies on homopolypeptide chains of glutamates have found that the polyglutamic acid chains are mechanically resilient. For example, a 25 residue polyglutamate chain resists extension at forces up to 180 pN [24]. This is further supported by studies demonstrating that PEVK segments with a larger number of poly-E motifs are stiffer than with a lower number of poly-E motifs [6]. This further highlights the importance of considering motif content within the PEVK region in considering titin's contribution to force.
Understanding PEVK stiffness is not as simple as just considering the distribution of motifs. Studies on the PEVK region have shown that the persistence length of the PEVK region can be reduced by calcium and by increased ionic strength [10]. Interestingly, while there is an enthalpic contribution to high force stretching of the PEVK region, suggesting a role for electrostatic interactions, there is no evidence for direct interaction between PPAK and poly-E motifs [7]. It is clear from the body of work on the PEVK region that there is a complex interplay between motif distribution and environmental conditions behind the PEVK region's role in both passive and active force and pH is another potential regulating factor that needs to be considered.
Function of Poly-E's rigid conformation on F-Actin and PEVK interactions.
It has been well-established that the PEVK region is capable of binding actin filaments at physiological ionic strengths [25].This interaction appears to be electrostatically driven since KCl concentrations above 150 mM abolish this interaction [25]. There is a high heterogeneous charge distribution on the surface of F-actin has a heterogeneous charge distribution along its surface though the overall net surface charge is negative [26]. The slight negative charge on F-actin would suggest that the PPAK motifs would be more likely to bind to thin filaments. Force spectroscopy experiments have shown that PEVK binds indiscriminately along the actin filament and that the load-rate dependence of the rupture force of the PPAK is mono-phasic but it is biphasic for poly-E [25]. Poly-E constructs are also known to have longer persistence lengths (0.9 nm for poly-E and 0.6 nm for PPAK) [25]. These results point to an important role for stiffness of the poly-E region in the PEVK-actin interaction, which would be altered with decreased pH.
Our results are consistent with these previous results and we propose that the extended conformation of the poly-E motifs may facilitate the PEVK/actin interaction in two ways. First, the extended conformation might ensure adequate interaction between the poly-E regions and positively charged regions on F-actin. Second, the extended conformation might help space PPAK motifs appropriately so that they can interaction with negative regions of F-actin. In both cases, the more relaxed conformation at lower pH's would impact binding of this region to actin filaments.
5. Conclusions
The results of this study demonstrate that the conformational state of the poly-E motif is sensitive to pH changes and that it assumes a more flexible collapsed conformation at lower pHs. A similar effect is not observed with the PPAK motif, suggesting that the two motifs play unique functional roles. These results highlight the complex nature of the PEVK region and introduce a new factor that could influence function of this important region of titin.
CRediT authorship contribution statement
Dassanayake Mudiyanselage Sudarshi Premawardhana: Conceptualization, Methodology, Investigation, Formal analysis, Writing - original draft, and, Reviewing and Editing. Fang Zhang: Methodology, Investigation. Jin Xu: Supervision. Matthew J. Gage: Project administration, Conceptualization, Methodology, Funding acquisition, Writing - original draft, and, Reviewing and Editing.
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgements
This work was supported in part by NIH Grant R15GM132840-01 and by the Keck Foundation.
References
- 1.Trombitás K. Titin extensibility in situ: entropic elasticity of permanently folded and permanently unfolded molecular segments. The Journal of cell biology. 1998;140(4):853–859. doi: 10.1083/jcb.140.4.853. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Martonfalvi Z. Low-force transitions in single titin molecules reflect a memory of contractile history. J. Cell Sci. 2014;127(Pt 4):858–870. doi: 10.1242/jcs.138461. [DOI] [PubMed] [Google Scholar]
- 3.Nishikawa K.C. Is titin a 'winding filament'? A new twist on muscle contraction. Proc. Biol. Sci. 2012;279(1730):981–990. doi: 10.1098/rspb.2011.1304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Gao M., Wilmanns M., Schulten K. Steered molecular dynamics studies of titin I1 domain unfolding. Biophys. J. 2002;83(6):3435–3445. doi: 10.1016/S0006-3495(02)75343-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Lu H. Unfolding of titin immunoglobulin domains by steered molecular dynamics simulation. Biophys. J. 1998;75(2):662–671. doi: 10.1016/S0006-3495(98)77556-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Nagy A. Hierarchical extensibility in the PEVK domain of skeletal-muscle titin. Biophys. J. 2005;89(1):329–336. doi: 10.1529/biophysj.104.057737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Duan Y. Studies on titin PEVK peptides and their interaction. Arch. Biochem. Biophys. 2006;454(1):16–25. doi: 10.1016/j.abb.2006.07.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Greaser M., Greaser M. Identification of new repeating motifs in titin. Proteins. 2001;43:145–149. doi: 10.1002/1097-0134(20010501)43:2<145::aid-prot1026>3.0.co;2-b. [DOI] [PubMed] [Google Scholar]
- 9.Linke W.A. Nature of PEVK-titin elasticity in skeletal muscle. Proc. Natl. Acad. Sci. Unit. States Am. 1998;95(14):8052. doi: 10.1073/pnas.95.14.8052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Granzier Henk L., Labeit S. The giant protein titin. Circ. Res. 2004;94(3):284–295. doi: 10.1161/01.RES.0000117769.88862.F8. [DOI] [PubMed] [Google Scholar]
- 11.Ma K., Kan L.-s., Wang K. Polyproline II helix is a key structural motif of the elastic PEVK segment of titin. Biochemistry. 2001;40(12):3427–3438. doi: 10.1021/bi0022792. [DOI] [PubMed] [Google Scholar]
- 12.Linke W.A., Granzier H. A spring tale: new facts on titin elasticity. Biophys. J. 1998;75(6):2613–2614. doi: 10.1016/S0006-3495(98)77706-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Linke W.A. PEVK domain of titin: an entropic spring with actin-binding properties. J. Struct. Biol. 2002;137(1–2):194–205. doi: 10.1006/jsbi.2002.4468. [DOI] [PubMed] [Google Scholar]
- 14.Jarvis K. Acidosis affects muscle contraction by slowing the rates myosin attaches to and detaches from actin. J. Muscle Res. Cell Motil. 2018;39(3–4):135–147. doi: 10.1007/s10974-018-9499-7. [DOI] [PubMed] [Google Scholar]
- 15.Forsyth W.R., Antosiewicz J.M., Robertson A.D. Empirical relationships between protein structure and carboxyl pKa values in proteins. Proteins: Structure, Function, and Bioinformatics. 2002;48(2):388–403. doi: 10.1002/prot.10174. [DOI] [PubMed] [Google Scholar]
- 16.Uversky V.N. Natively unfolded proteins: a point where biology waits for physics. Protein Sci. 2002;11(4):739–756. doi: 10.1110/ps.4210102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Kowalchuk J.M. Factors influencing hydrogen ion concentration in muscle after intense exercise. J. Appl. Physiol. 1988;65(5):2080–2089. doi: 10.1152/jappl.1988.65.5.2080. [DOI] [PubMed] [Google Scholar]
- 18.McKenna M.J. The roles of ionic processes in muscular fatigue during intense exercise. Sports Med. 1992;13(2):134–145. doi: 10.2165/00007256-199213020-00009. [DOI] [PubMed] [Google Scholar]
- 19.Fitts R.H. The cross-bridge cycle and skeletal muscle fatigue. J. Appl. Physiol. 2008;104(2):551–558. doi: 10.1152/japplphysiol.01200.2007. [DOI] [PubMed] [Google Scholar]
- 20.Nagy A. Differential actin binding along the PEVK domain of skeletal muscle titin. J. Cell Sci. 2004;117(Pt 24):5781–5789. doi: 10.1242/jcs.01501. [DOI] [PubMed] [Google Scholar]
- 21.Yamasaki R. Titin-actin interaction in mouse myocardium: passive tension modulation and its regulation by calcium/S100A1. Biophys. J. 2001;81(4):2297–2313. doi: 10.1016/S0006-3495(01)75876-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Neagoe C. Titin isoform switch in ischemic human heart disease. Circulation. 2002;106(11):1333–1341. doi: 10.1161/01.cir.0000029803.93022.93. [DOI] [PubMed] [Google Scholar]
- 23.Makarenko I. Passive stiffness changes caused by upregulation of compliant titin isoforms in human dilated cardiomyopathy hearts. Circ. Res. 2004;95(7):708–716. doi: 10.1161/01.RES.0000143901.37063.2f. [DOI] [PubMed] [Google Scholar]
- 24.Dougan L. Single homopolypeptide chains collapse into mechanically rigid conformations. Proc. Natl. Acad. Sci. U. S. A. 2009;106(31):12605–12610. doi: 10.1073/pnas.0900678106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Bianco P. Interaction forces between F-actin and titin PEVK domain measured with optical tweezers. Biophys. J. 2007;93(6):2102–2109. doi: 10.1529/biophysj.107.106153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Angelini T.E. Counterions between charged polymers exhibit liquid-like organization and dynamics. Proc. Natl. Acad. Sci. U. S. A. 2006;103(21):7962–7967. doi: 10.1073/pnas.0601435103. [DOI] [PMC free article] [PubMed] [Google Scholar]