Summary
Gap junctions are present in most tissues and play essential roles in various biological processes. However, we know surprisingly little about the molecular mechanisms underlying gap junction formation. Here we uncover the essential role of a conserved EGF- and Laminin G- domain containing protein nlr-1/CASPR in the regulation of gap junction formation in multiple tissues across different developmental stages in C. elegans. NLR-1 is located in the gap junction perinexus, a region adjacent to but not overlapping with gap junctions, and forms puncta before the clusters of gap junction channels appear on the membrane. We show that NLR-1 can directly bind to actin to recruit F-actin networks at the gap junction formation plaque, and the formation of F-actin patches plays a critical role in the assembly of gap junction channels. Our findings demonstrate that nlr-1/CASPR acts as an early stage signal for gap junction formation through anchoring of F-actin networks.
Keywords: Gap junction, C. elegans, Contactin associated protein, F-actin
Graphical Abstract

In Brief/eTOC blurb:
Meng et al. reveal the role of the conserved EGF- and Laminin G- domain-containing protein nlr-1/CASPR in control of the formation of gap junctions in multiple tissues across different developmental stages, and show that NLR-1 forms gap junction through recruiting F-actin networks at the gap junction formation plaque
Introduction
Gap junctions are special junction structures that connect two cells to allow direct transfer of ions and small molecules, and they form plaques consisting of a few to thousands of units at appositional regions of cells (Connors and Long, 2004; Goodenough and Paul, 2009; Mese et al., 2007). Since the discovery of gap junctions in the 1950s, it has been a mystery how gap junctions are formed (Furshpan and Potter, 1959; Watanabe, 1958). The establishment of gap junctions requires the close contact of two cells, and adhesion molecules such as cadherins have been shown to be critical for bringing two cells together and triggering downstream signals during gap junction formation (Chakraborty et al., 2010; Govindarajan et al., 2010). Studies using transmission electron microscopy and electrophysiology document each step of gap junction formation and show that a “formation plaque” forms between two cells prior to the gap junction channels moving in (Hand and Gobel, 1972; Johnson et al., 1974; Johnson et al., 2012; Preus et al., 1981). Therefore, the study of gap junction formation can be divided into two parts: 1) how the formation plaque develops between two adjacent cells; and 2) how gap junction channels cluster at the formation plaque. Most studies to date have focused on the latter, dissecting the signals that regulate the clustering of gap junction channels at the formation plaque. They have revealed the essential roles of the MAGUK (membrane -associated guanylate kinase)-family member ZO-1 and posttranslational modifications of gap junction channels in clustering gap junction channels at the formation plaque, but these regulators are not involved in establishing the formation plaque (Berthoud et al., 2004; Falk et al., 2016; Falk et al., 2014; Flores et al., 2008; Giepmans and Moolenaar, 1998; Hunter et al., 2005; Marsh et al., 2017; Pogoda et al., 2016; Rhett et al., 2011; Segretain and Falk, 2004; Thevenin et al., 2013; Toyofuku et al., 1998). Although the C-terminus of connexins was shown to be involved in the establishment of the formation plaque and gap junctions (Johnson et al., 2012), little is known about the molecules that comprise the formation plaque, which makes it still very challenging to address how the formation plaque forms.
Gap junctions are found in most of, if not all, multicellular organisms, and are made up of connexins in vertebrates and innexins in invertebrates (Alexopoulos et al., 2004; Phelan, 2005). While they differ in their primary sequences, connexins and innexins have similar tertiary structures with intracellular amino- and carboxy- termini, two extracellular and one intracellular loops, and four transmembrane domains (Mese et al., 2007). Recent cryogenic electron microscopy (cryo-EM) studies of C. elegans INNEXIN-6 (INX-6) further highlighted the structural similarities between innexins and connexins and showed that the structures of transmembrane helices, extracellular loops and intracellular termini of INX-6 are remarkably similar to those of connexin26 (Cx26) (Maeda et al., 2009; Oshima et al., 2016b). The structural similarities between innexins and connexins are not only observed in monomeric channels but also in hexadecameric gap junctions, and both INX-6 and Cx26 gap junctions display a tsuzumi-shape (Maeda et al., 2009; Oshima et al., 2016b). Interestingly, it was reported that when a truncated INX-6 without its N-terminal was expressed in vitro and crystallized in lipid bilayers, a single INX-6 gap junction comprises 8 subunits in each hemichannel, which is different to the 6 subunits in connexin-based hemichannels (Maeda et al., 2009; Oshima et al., 2016a). However, it is still unclear whether this in vitro crystallized structure of the truncated INX-6 reflects functional gap junctions in vivo. Besides their structural similarity, innexin-based gap junctions also share many other similarities with connexin-based gap junctions: they both mediate sophisticated functions such as long-term potentiation and neuronal circuit modulation (Huang et al., 2018; Jin et al., 2016; Voelker et al., 2019; Welzel and Schuster, 2018); they are modulated by posttranslational modifications such as phosphorylation (Bauer et al., 2005; Calkins et al., 2015; Depriest et al., 2011; Welzel and Schuster, 2018); and their functions are regulated by intercellular binding proteins (Chen et al., 2007; Jang et al., 2017; Thevenin et al., 2013). A recent study shows that ectopically expressed vertebrate connexin36 (Cx36) in C. elegans AWC and AIY neurons can form functional Cx36-based gap junctions to affect neuronal function and animal behavior (Rabinowitch et al., 2014). This suggests even if connexins are not expressed in invertebrates, they may still possess certain conserved molecular mechanisms that could mediate the formation of connexin-based gap junctions. In the last two decades using electron microscopy, electrophysiology, and calcium imaging, studies from many labs have shown C. elegans gap junctions share many similarities with vertebrate ones, and C. elegans has become a powerful model to study the function and regulation of gap junctions in vivo (Bhattacharya et al., 2019; Chuang et al., 2007; Gao et al., 2018; Hall, 2017; Jang et al., 2017; Liu et al., 2011; Meng et al., 2016a; Meng et al., 2016b; Oshima et al., 2016b; Rabinowitch et al., 2014; Schumacher et al., 2012).
The cytoskeleton plays critical roles in many biological processes. In the case of gap junction formation, microtubules appear to be essential for the transportation of gap junction channels from the Golgi to the plasma membrane (Lauf et al., 2002) and the insertion of gap junction channels into the plaque region (Shaw et al., 2007). Microtubule capping proteins can bind with adhesion molecules such as cadherins to provide targeted delivery of gap junction cargoes (Butkevich et al., 2004). The interactions between gap junctions and microtubules are not only important for gap junction formation but also influence the properties of microtubules (Giepmans et al., 2001). However, microtubules are not always involved in gap junction formation, as studies show that the assembly of gap junctions in the preimplantation mouse conceptus is independent of microtubules (Kidder et al., 1987). F-actin has been shown to be associated with gap junctions through binding with cadherins (Bauer et al., 2006; Wei et al., 2005; Xu et al., 2001), but it is still unclear whether and how F-actin may affect gap junction formation.
Here we demonstrate that the conserved CASPR (Contactin-associated protein) family protein NLR-1 is required for gap junction formation and maintenance. We show that NLR-1 forms puncta at the gap junction perinexus, a region adjacent to but not overlapping with the gap junction plaque (Rhett et al., 2011), to anchor F-actin but not microtubules to trigger gap junction formation. Consistent with its role in gap junction formation, NLR-1 puncta appear prior to the clustering of gap junction channels. Knockdown of nlr-1 disrupts the organization of F-actin networks at the formation plaque, and as a consequence F-actin accompanied by gap junction channels randomly clusters along the membrane. These results support the essential role of NLR-1 and F-actin in gap junction formation and suggest that NLR-1 is involved in the establishmentof the formation plaque.
Results
Use of the C. elegans embryo as a model to study gap junction formation
To uncover the potential mechanisms of gap junction formation, we used the C. elegans embryo as a model, taking advantage of its limited cell number, large cell size and invariant gap junctions that are rapidly formed (Bossinger and Schierenberg, 1996; Sulston et al., 1983). To label gap junctions in embryos, we used a well-established inx-3/innexin::GFP transgene marker, which has been shown to label INX-3 containing gap junctions in vivo (Starich et al., 2003). We found that one of the earliest gap junction plaques occurred between EA and EP cells, two endodermal precursor cells, at the 16–24 cell stage of embryogenesis (Fig. 1A and 1B). EA and EP cells only form one gap junction plaque at the center of their adjoining membranes (Fig. 1B and 1C), making it a good model to study gap junction formation. There are three potential models for EA/EP gap junction formation (Fig. 1D): (1) The first model is through competition, in which gap junctions first randomly form between EA and EP cells, and then one of them becomes dominant through competition. As a result the rest of the gap junctions are eliminated, in a similar fashion as what has been described during the formation of neuromuscular junctions (Fields and Nelson, 1992; Mennerick and Zorumski, 2000; Santafe et al., 2004); (2) The second model is through fusion, where gap junctions randomly form along the EA/EP adjoining membranes and then fuse together to form the final gap junction plaque; (3) the third model is through growth, in which gap junctions are targeted to a specific region and grow into the final gap junction plaque. To examine how EA and EP cells build their gap junctions, we carried out live-cell imaging experiments. We found that EA and EP cells started to form gap junctions shortly after the cells are born, and the INX-3::GFP signal could only be observed at the center of their adjoining membranes, where the final gap junction plaque is localized, and not in any other regions of the membrane (Fig. 1E and 1F). We also noticed that the intensity of INX-3::GFP rapidly reached a plateau from its baseline (<40s), and INX-3::GFP puncta were stable at the center of the adjoining membranes throughout the lifetime of the EA and EP cells. A closer examination of the first group of gap junctions formed between EA and EP cells shows that clusters of gap junction channels emerge at the gap junction region from both the EA and EP cells at the same time and are subsequently assembled into gap junctions on the membrane (Fig. 1G). These results support that EA and EP cells form gap junctions through the growth model (Fig. 1D, model 3), meaning that gap junction channels are directly delivered to the region where gap junctions are formed and grow into the final gap junction plaque.
Figure 1. EA and EP cells form a single gap junction plaque at the center of the adjoining membranes.
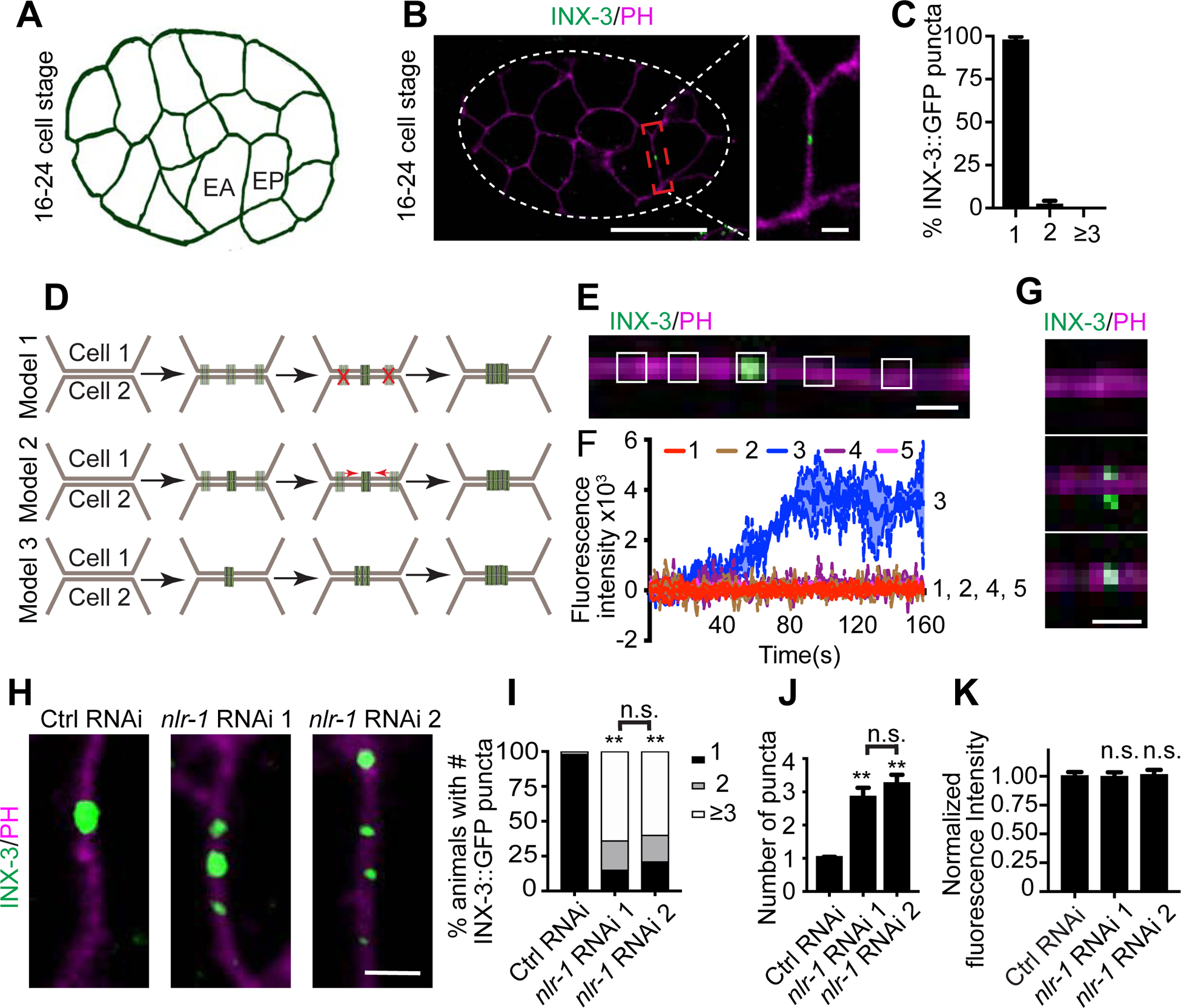
(A) A schematic of the 16–24 cell stage in C. elegans embryos. (B) Representative images of localization of INX-3::GFP at the 16–24 cell stage (Scale bar (left): 20 μm; Scale bar (right): 2 μm). Pleckstrin homology domain (PH domain) targeted mCherry was used to visualize the cell membrane. White contours outline the embryo. (C) Quantification of percentage of animals with 1, 2 or ≥3 INX-3::GFP puncta on the adjoining membranes of EA/EP cells. n≥ 85. (D) Potential models for EA/EP gap junction formation. (E) A representative image of INX-3::GFP localization on the adjoining EA/EP membranes with the white boxes showing the five regions of interest (ROI) for fluorescence intensity quantification (Scale bar: 1 μm). (F) Quantification of mean fluorescence intensity of the five ROI in 160 s. n=5 animals. (G) Representative images of INX-3::GFP localization on the adjoining EA/EP membranes during the early stages of gap junction formation (Scale bar: 1 μm). (H) Representative images of localization of INX-3::GFP on the adjoining EA/EP membranes in control and nlr-1 knockdown animals (Scale bar: 2 μm). (I) Quantification of percentage of animals with 1, 2 or ≥3 INX-3::GFP puncta on the adjoining membranes of EA/EP cells in control and nlr-1 knockdown animals. n≥85. Fisher’s exact test, **p < 0.005. (J, K) Quantification of average INX-3::GFP puncta number and normalized INX-3::GFP fluorescence intensity on the adjoining EA/EP membranes. n≥90. One-way ANOVA test; **p < 0.005, n.s: no significant difference.
With these findings we further asked what could be the molecular mechanisms underlying the formation of EA/EP gap junctions. Since gap junctions are formed on the membrane, it is reasonable to hypothesize that some membrane or extracellular membrane (ECM) proteins may play important roles in gap junction formation. As INX-3-containing gap junctions are essential for embryogenesis (Starich et al., 2003), any genes that are important for EA/EP gap junction formation are likely to be essential. Based on this, we carried out a RNAi-based genetic screen targeted on examining the function of the conserved and essential membrane/ECM proteins in EA/EP gap junction formation. After examining 103 genes (Supplemental Table 1), we found that RNAi-mediated knockdown of nlr-1 induced multiple clusters of INX-3::GFP puncta along the EA/EP adjoining membranes (Fig. 1H–1J and Supplemental Fig. 1A), while knockdown of any other genes tested did not affect EA/EP gap junctions. Despite the significant increase in the number of INX-3::GFP puncta, the total GFP fluorescence intensity on the adjoining EA/EP membrane and the overall expression pattern of inx-3 in nlr-1 knockdown animals were similar to those in control animals (Fig. 1K, and Supplemental Fig. 1B–1D, suggesting that nlr-1 regulates INX-3-containing gap junctions. We further confirmed these results by examining the effect of feeding animals with a second nlr-1 RNAi strain (Supplemental Fig. 1E) and showed that feeding both RNAi strains caused similar phenotypes (Fig. 1H–1K).
nlr-1 is required for gap junction formation and maintenance in embryos
nlr-1 is the C. elegans homolog of CNTNAP2 (Contactin-associated protein-like 2), a member of the CASPR (Contactin-associated protein) family proteins. CASPR family proteins have been shown to play important roles in neuronal development and to participate during the membrane localization of voltage-gated potassium channels (Poliak et al., 1999; Poliak et al., 2003; Traka et al., 2003). Besides nlr-1, C. elegans expresses another essential CASPR gene, itx-1 (Haklai-Topper et al., 2011), but knockdown of itx-1 did not cause any obvious defects in EA/EP gap junction formation (Supplemental Fig. 1F). CASPR4, a CASPR family protein, can regulate retinal ganglion development through interactions with Contactin 5 (Peng et al., 2017). We examined the only C. elegans Contactin homolog rig-6 (Schwarz et al., 2009) and did not observe any abnormalities in rig-6(lf) mutants (Supplemental Fig. 1G). To further understand the function of nlr-1 in EA/EP gap junction formation, we carried out live-cell imaging experiments. Different from what we observed in control animals, in which gap junction channels simultaneously cluster close to the gap junction region in both EA and EP cells and subsequently assemble into the formation plaque on the membrane (Fig. 2A, 2C, 2D), the gap junction channels in nlr-1 RNAi animals seem to randomly form clusters inside either the EA or EP cells (Fig. 2B, 2C, 2D). As a result, during the period of gap junction formation, multiple transient clusters of gap junction channels were observed in EA or EP cells in nlr-1 RNAi animals (Fig. 2D). To gain further insight into whether these transient puncta in nlr-1 RNAi animals were on the membrane, we quantified the crest distance between the fluorescence peaks of INX-3::GFP and PH::mCherry and found that in control animals the average crest distance between GFP and mCherry signals is close to zero, supporting the conclusion that INX-3::GFP puncta are on the membrane in control animals (Fig. 2E, Supplemental Fig. 2A). In contrast, the average crest distance between GFP and mCherry signals is about 300nm in nlr-1 RNAi animals (Fig. 2E, Supplemental Fig. 2A), suggesting that most of these transient puncta are not located in the membrane.
Figure 2. nlr-1 is required for INX-3 plaque formation and maintenance in embryos.
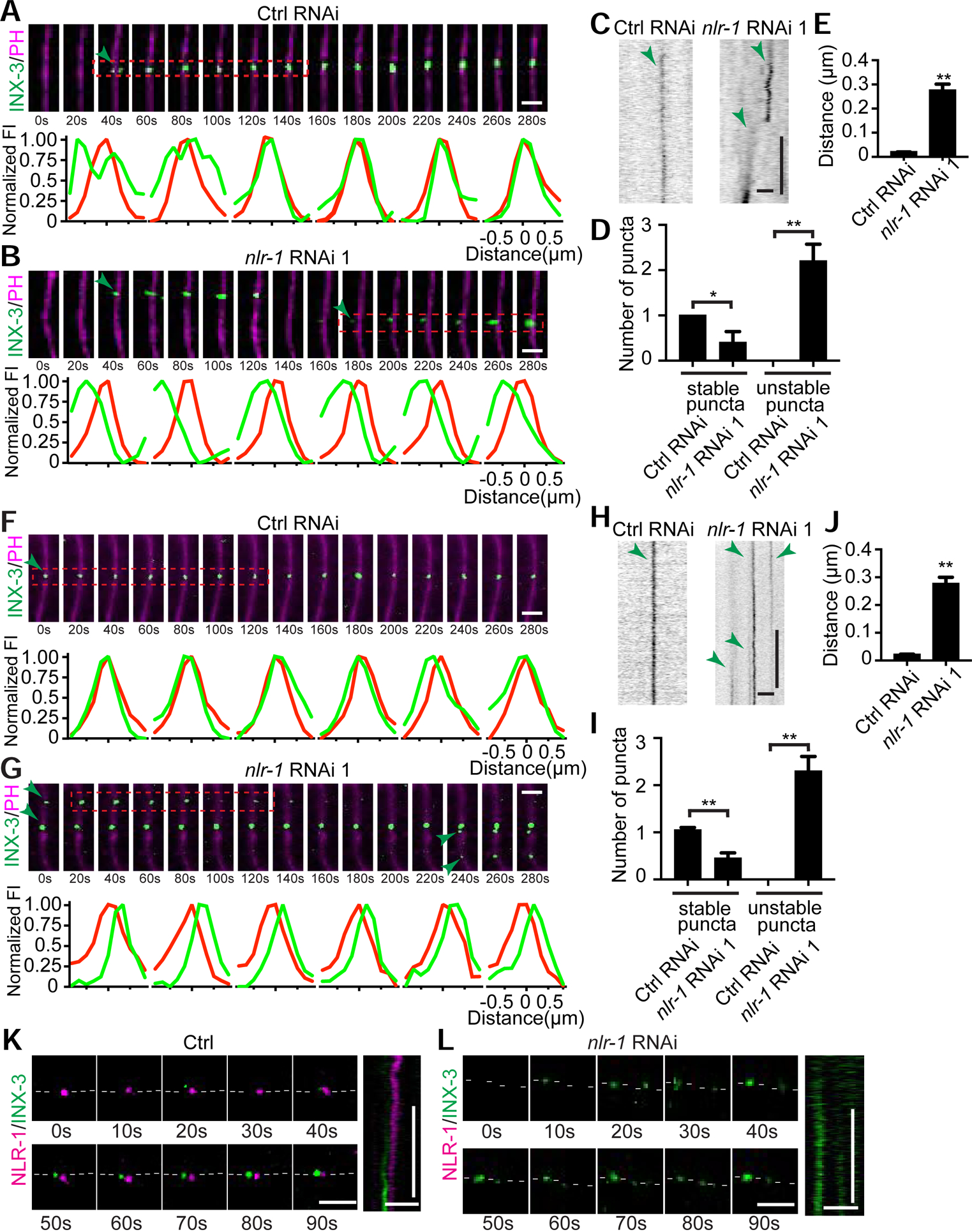
(A-E) nlr-1 is required for EA/EP gap junction formation. (A, B) Representative time-lapse images of INX-3::GFP puncta and profiles of GFP and mCherry fluorescence intensities at the regions indicated (red rectangle) during gap junction formation in control and nlr-1 knockdown animals (Scale bars: 2 μm). The interval between each image is 20s. Green arrows point to INX-3::GFP puncta. (C) Representative kymographs showing the dynamic of INX-3::GFP puncta formation (Scale bars (horizontal): 2 μm; Scale bars (vertical): 20 sec). Green arrows point to INX-3::GFP traces. (D) Quantification of stable and unstable puncta in control and nlr-1 knockdown animals during gap junction formation. (E) Quantification of average distance between the crests of the INX-3::GFP and PH::mCherry intensity profiles. Each point represents 5 experiments of at least 200 events. Student’s t-test, *p < 0.05; **p < 0.005. (F-J) nlr-1 is required for EA/EP gap junction maintenance. (F, G) Representative time-lapse images of INX-3::GFP puncta and GFP and mCherry fluorescence intensity profiles at the regions indicated (red rectangle) after gap junction formation in control and nlr-1 knockdown animals (Scale bars: 2 μm). The interval between each image is 20s. Green arrows point to INX-3::GFP puncta. (H) Representative kymographs showing the dynamic of INX-3::GFP puncta (Scale bars (horizontal): 2 μm; Scale bars (vertical): 20 sec). Green arrows point to INX-3::GFP traces. (I) Quantification of stable and unstable puncta in control and nlr-1 knockdown animals after gap junction formation. (J) Quantification of average distance between the crests of the INX-3::GFP and PH::mCherry intensity profiles. Each point represents 5 experiments of at least 200 events. Student’s t-test, **p < 0.005. (K, L) Representative time-lapse images (Scale bar: 2 μm) and kymographs (Horizontal scale bars: 2 μm; Vertical scale bars: 20 sec) of endogenous NLR-1 and INX-3 in control and nlr-1 knockdown animals during INX-3 plaque formation. White dash lines outline the plasma membrane.
Following its formation, the gap junction plaque appears to be stable on the membrane, and no additional gap junctions were observed at other regions of the membrane throughout the lifetime of the EA and EP cells in control animals (Fig. 2F, 2H, 2I). Our data show that nlr-1 is also required for the maintenance of EA/EP gap junctions, as multiple transient INX-3::GFP puncta continually form along the EA/EP membrane in nlr-1 knockdown animals (Fig. 2G–2I). A close examination of the average crest distance between fluorescence peaks of INX-3::GFP and PH::mCherry in nlr-1 RNAi animals show most of these transient puncta are not on the membrane (Fig. 2J, Supplemental Fig. 2B). These results support the conclusion that nlr-1 is required for gap junction formation and maintenance. More specifically, nlr-1 is required for gap junction channels to assemble into the formation plaque on the membrane.
Given the importance of nlr-1 in gap junction formation and maintenance, we wanted to know where and when nlr-1 is expressed. To do so, we generated a knock-in strain with mNeonGreen fused to the C-terminus of NLR-1 and a second knock-in strain to label INX-3 with mKate at the C-terminus (Supplemental Fig. 2C). Both knock-in strains display similar behavior, brood size and gap junction morphology when compared with wild type, inx-3 and nlr-1 transgenic animals (Supplemental Fig. 2D and 2E), suggesting that knock-in of mNeonGreen and mKate did not affect the function of nlr-1 and inx-3. We found that both nlr-1 and inx-3 were expressed in the EA and EP cells (Fig. 2K, and Supplemental Fig. 2F). We then examined the dynamics of NLR-1 and INX-3 during gap junction formation and found that NLR-1 formed puncta at the gap junction perinexus and emerged prior to the formation of INX-3 puncta (Fig. 2K, and Supplemental Fig. 2H). After gap junction formation NLR-1 and INX-3 puncta are close to each other throughout the lifetime of the EA and EP cells (Supplemental Fig. 2G and 2H). Elimination of NLR-1 puncta caused multiple randomly-formed unstable INX-3 puncta to appear (Fig. 2L). Taken together, we believe that the localization of NLR-1 at the gap junction perinexus is important for its function in gap junction formation and maintenance.
nlr-1 is required for gap junction formation and function in pharyngeal muscles and neurons
To test whether nlr-1 is also important in other tissues, we examined gap junction formation in pharyngeal muscles, where both nlr-1 and inx-3 are expressed (Supplemental Fig. 3A) (Altun et al., 2009; Haklai-Topper et al., 2011). We focused our studies on gap junctions of pm5 muscle cells (Supplemental Fig. 3B) (Albertson and Thomson, 1976; Franks et al., 2006) and found that similar to what was found in EA/EP cells, NLR-1 localized at the gap junction perinexus (Fig. 3A–3C). INX-3::GFP forms about 20 puncta that evenly distribute along the adjoining membranes of the two pm5 muscle cells in control animals (Fig. 3D and 3E). In nlr-1 knockdown animals there were fewer INX-3::GFP puncta with increased average size (Fig. 3D–3F). As a result, the total GFP intensity along the adjoining membranes of the pm5 muscle cells did not change in nlr-1 RNAi animals when compared with control animals (Supplemental Fig. 3C). Further experiments also showed that knockdown of nlr-1 specifically in pharyngeal muscles was sufficient to generate similar phenotypes as those found in all-tissue knockdown animals, suggesting that nlr-1 functions cell autonomously to regulate gap junction formation in pharyngeal muscles (Supplemental Fig. 3D–3F).
Figure 3. nlr-1 regulates gap junction formation and function in muscles and neurons.
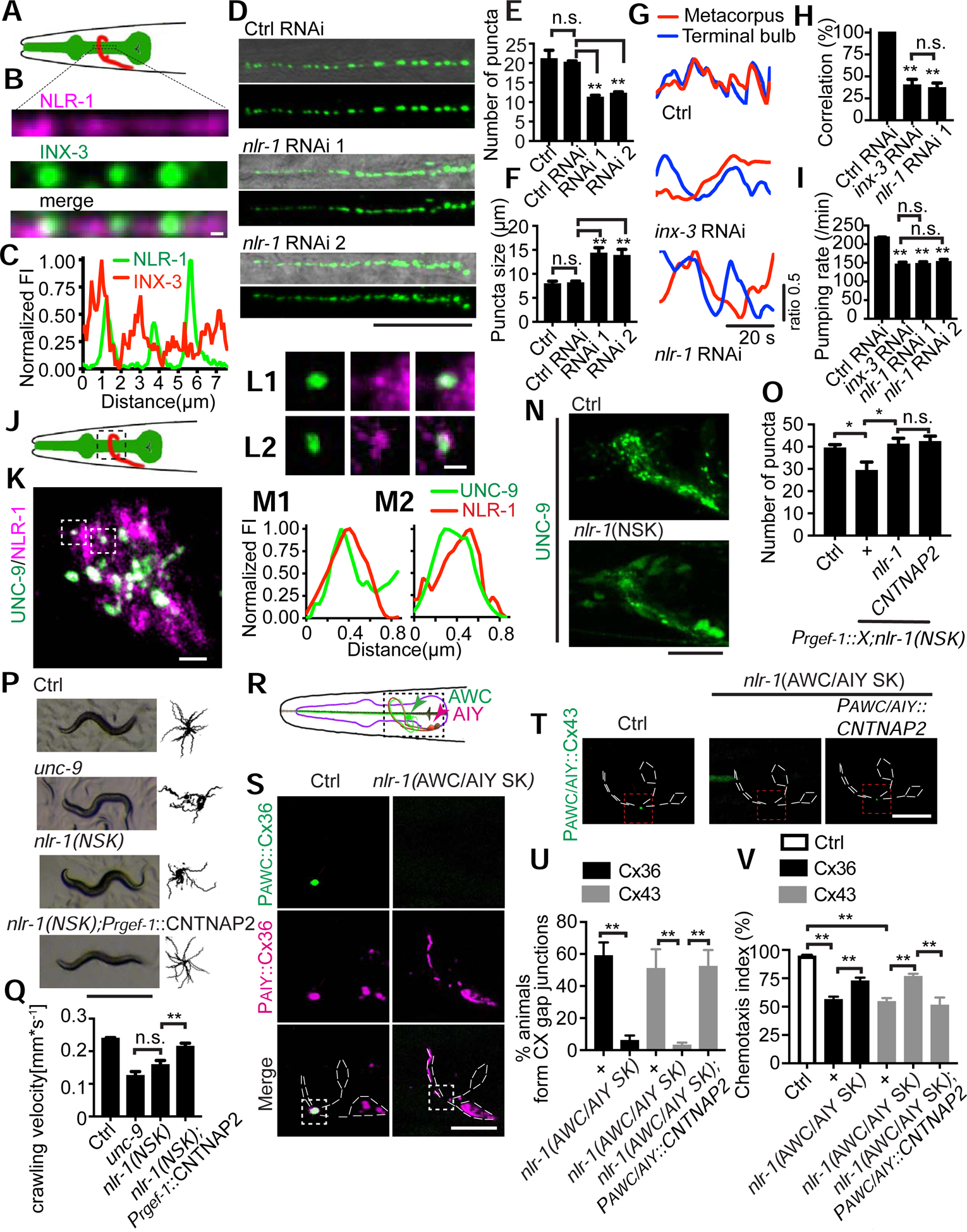
(A) A schematic of pharyngeal muscles (green). The black rectangles highlight the region of interest at pharyngeal pm5 muscles. (B, C) Representative images and fluorescence intensity profiles of NLR-1 and INX-3 localization along the adjoining membranes of pharyngeal pm5 muscles (Scale bar: 1 μm). (D) Representative images of INX-3::GFP puncta localization along the adjoining membranes of pm5 muscles. Images at the top show a merge of DIC and GFP images, and images at the bottom show GFP only (Scale bar: 10 μm). (E, F) Quantification of average number, average size of INX-3::GFP puncta in control, control RNAi and nlr-1 knockdown animals. One-way ANOVA test, n≥90. **p < 0.005, n.s: no significant difference. (G, H) Representative images and quantification of correlation between calcium transients in the metacorpus and terminal bulbs in control, inx-3 knockdown and nlr-1 knockdown animals. n≥60. One-way ANOVA test, **p < 0.005, n.s: no significant difference. (I) Quantification of pumping rate in control RNAi, inx-3 knockdown and nlr-1 knockdown animals. n≥30. One-way ANOVA test, **p < 0.005, n.s: no significant difference. (J) A schematic of the nerve ring (red). The black rectangles highlight the region of interest at the nerve ring. (K-M) Representative images of NLR-1 and UNC-9 localization (Scale bar: 2 μm in (K), Scale bar: 1 μm in (L)) and fluorescence intensity profiles of NLR-1 and UNC-9. (N) Representative images of UNC-9::GFP puncta localization in control and nlr-1 neuronal conditional knockout animals (Scale bar: 20 μm). (O) Quantification of UNC-9::GFP puncta localization in control, nlr-1(NSK) and transgenic rescue animals. Pregf-1 was used as a pan-neuronal promoter. n≥30. One-way ANOVA test, *p < 0.05, n.s: no significant difference. (P) Representative images and trajectories (20 animals) of 30 s locomotion are shown for wild type, unc-9(lf), nlr-1(NSK), and CNTNAP2 rescue animals (Scale bar: 0.5 mm). (Q) Quantification of crawling velocity of wild type, unc-9(lf), nlr-1(NSK) and CNTNAP2 rescue animals. n≥80. One-way ANOVA test, **p < 0.005, n.s: no significant difference. (R) A schematic of AWC and AIY neurons. The black rectangles highlight the region of interest. (S) Representative images of AWC::Cx36::GFP;AIY::Cx36::mCherry in control and nlr-1 AWC/AIY specific knockout (AWC/AIY SK) animals (Scale bar: 5 μm). The white dashed lines outline the morphologies of AWC and AIY neurons. White arrowheads point to the Cx36-based gap junctions, which are localized at the contact region of AWC and AIY and are labeled by Cx36 from both AWC (green) and AIY (pink). Podr-1 and Pttx-3 were used as AWC and AIY specific promoters respectively. (T) Representative images of AWC::msfGFP; AIY::Cx43::msfGFP in control, nlr-1(AWC/AIY SK) and CNANAP2 rescue animals (Scale bar: 5 μm). The white dashed lines outline the morphologies of AWC and AIY neurons. White arrowheads point to the Cx43-based gap junctions, which are localized to the contact region between AWC and AIY. Podr-1 and Pttx-3 were used as AWC and AIY specific promoters respectively. (U) Percentage of animals with AWC/AIY gap junctions in control and nlr-1(AWC/AIY SK) conditions. n≥30. Student’s t-test, **p < 0.005. (V) Benzaldehyde chemotaxis behavior of wild type, connexin transgenic, connexin transgenic in nlr-1(AWC/AIY SK), and CNANAP2 rescue animals. n≥300. Student’s t-test, **p < 0.005.
One advantage of using pm5 muscle cells to study gap junctions is that the function of these gap junctions is well-documented, where they are required for the synchronous muscular contraction between the pharyngeal metacorpus and terminal bulbs (Raizen and Avery, 1994; Shimozono et al., 2004; Simonsen et al., 2014). To test whether the morphological defects we observed in nlr-1 RNAi animals could cause any functional abnormalities, we used GCaMP6.0s as an indicator to monitor muscle contractions in the metacorpus and terminal bulbs. Consistent with previous studies, in control animals muscle contractions are tightly coupled between the metacorpus and terminal bulbs (Simonsen et al., 2014), and knockdown of inx-3 causes a significant decrease in the coupling between the metacorpus and terminal bulbs (Fig. 3G and 3H). We found that RNAi of nlr-1 lowered the coupling between the metacorpus and terminal bulb muscles from 100% to about 30%, which is similar to that in inx-3 knockdown animals (Fig. 3G and 3H). As a result, in nlr-1 or inx-3 RNAi animals, the GCaMP6 signal in the terminal bulb muscle is less than 30% of that in the metacorpus bulb muscle during metacorpus bulb contraction (Supplemental Fig. 3G). The pharyngeal pumping behavior is generated by the coordinated contraction of pharyngeal muscles and can be used as a behavioral readout for gap junction function in pharyngeal muscles (Simonsen et al., 2014). Consistent with our calcium imaging results, knockdown of nlr-1 in all tissues or specifically in pharyngeal muscles caused a significant decrease in pharyngeal pumping rate that is similar to what was observed in inx-3 RNAi animals (Fig. 3I and Supplemental Fig. 3H).
Since nlr-1 has strong expression in the nerve ring, the brain of C. elegans, we examined the function of nlr-1 in neuronal gap junction formation using a well-established neuronal gap junction marker UNC-9::GFP (Kawano et al., 2011; Liu et al., 2017) (Supplemental Fig. 3A). Consistent with what we found in embryos and pharyngeal muscles, NLR-1 localizes to the gap junction perinexus in the nerve ring (Fig. 3J–3M). As loss-of-function of nlr-1 causes lethality, and C. elegans neurons are resistant to feeding RNAi (Kamath et al., 2001), we generated an nlr-1 neuron-specific knockout (NSK) strain to examine its function in the nervous system (Golic and Lindquist, 1989; Sauer, 1987; Sauer and Henderson, 1988). We inserted two flippase recognition target (FRT) sites into noncoding regions of nlr-1, intron 13 and 3’UTR (Supplemental Fig. 3I), and expressed flippase (Flp) in the nervous system to generate a deletion that removes the transmembrane and the intracellular domains from the coding region. To ensure the efficiency of the knockout, we knocked in an mKate right after the C-terminus of NLR-1 and found that in the conditional knockout animals the neuronal expression of nlr-1 was abolished while nlr-1 had normal expression in other tissues (Supplemental Fig. 3J). As shown in Fig. 3N, 3O and Supplemental Fig. 3K, neuron-specific knockout (NSK) of nlr-1 caused decreases in both total number and average size of UNC-9::GFP puncta in the nerve ring region when compared with control animals. Consistent with this observation, we noticed that there are more neuronal cell bodies with visible UNC-9::GFP signal in nlr-1 NSK animals than in control animals (Fig. 3N). unc-9(lf) animals display the kinker uncoordinated (unc) phenotype, which has been shown to be associated with the gap junction function of unc-9 (Yeh et al., 2009). nlr-1(NSK) animals exhibit a similar kinker unc and decrease in locomotion as those in unc-9(lf) animals (Fig. 3P and 3Q). These data support the conclusion that nlr-1 regulates gap junction formation and function in neurons.
nlr-1 and its mammalian homolog CNTANAP2 can form connexin-based gap junctions in C. elegans
nlr-1 is a homolog of mammalian contactin associated proteins (CNTNAP), and human CNTNAP2 shares over 85% similarity with C. elegans NLR-1 (Supplemental Fig. 3L). To test whether CNTNAP2 is a functional homolog of nlr-1, and whether CNTNAP2 can regulate gap junction formation, we generated transgenes expressing the human CNTNAP2 in the nervous system of nlr-1(NSK) animals, and found that CNTNAP2 transgenes fully rescue both gap junction morphology and locomotion defects in nlr-1(NSK) animals (Fig. 3O–3Q). These results suggest nlr-1 and its mammalian homolog CNTNAP2 may play conserved functions in gap junction formation.
To further address the conserved role of nlr-1/CNTNAP2 in gap junctions, we examined whether they are involved in the formation of connexin-based gap junctions. Previous studies show that expression of vertebrate connexin 36 (Cx36) in C. elegans AIY and AWC neurons can form functional connexin-based gap junctions (Fig. 3R) (Rabinowitch et al., 2014). We examined whether nlr-1 was required for the formation of these Cx36-based gap junctions by specifically knocking out nlr-1 in AIY and AWC neurons, and found that Cx36 failed to form gap junctions in nlr-1 ((AWC/AIY specific knockout (AWC/AIY SK)) animals (Figure 3S and 3U). As a consequence the suppression of the response to benzaldehyde, a behavioral phenotype mediated by Cx36-based gap junctions (Rabinowitch et al., 2014), was largely released in nlr-1 (AWC/AIY SK) animals (Figure 3V). These results suggest that nlr-1 may play an important role in the formation of connexin-based gap junctions as well. To further support this conclusion, we tested another well-studied connexin – connexin 43 (Cx43). When expressed in AWC/AIY neurons, Cx43 formed similar gap junctions and caused the same degree of suppression in the response to benzaldehyde as that in Cx36 transgenes (Rabinowitch et al., 2014), and knockout of nlr-1 in AWC/AIY neurons similarly suppressed the formation of Cx43-based gap junctions and released the suppression of the response to benzaldehyde (Figure 3T–3V). Moreover, expression of human CNTNAP2 restored the formation of Cx43-based gap junctions and suppressed the response to benzaldehyde in nlr-1(AWC/AIY SK) animals (Figure 3T–3V). Taken together, we believe that nlr-1 and CNTNAP2 play a conserved role in gap junction formation and are involved in the formation of both innexin- and connexin- based gap junctions.
Disruption of F-actin causes defects in gap junction formation
In another RNAi screen targeted to examine the functions of cytoskeletal and motor proteins in gap junction formation, we found that knockdown of two key components of the WAVE regulatory complex, wve-1/WAVE and gex-3/NAP1 (Eden et al., 2002; Sawa et al., 2003; Withee et al., 2004), caused similar defects in EA/EP gap junction formation as those in nlr-1 knockdown animals, namely forming multiple random clusters of INX-3::GFP puncta outside the gap junction region (Fig. 4A–4C, Supplemental Fig. 4A). Given their phenotypic similarity, we further tested the genetic interactions between nlr-1 and the WAVE complex by examining the effect of knockdown of nlr-1 together with gex-3 or wve-1. As shown in Fig. 4A–4C, double RNAi knockdown animals did not show stronger phenotypes when compared with single RNAi knockdown animals. We also noticed that knockdown of gex-3 or wve-1 did not affect the localization or the size of NLR-1 puncta in embryos (Fig. 4D and Supplemental Fig. 4B). Since the WAVE complex plays essential roles in promoting actin nucleation and polymerization (Pantaloni et al., 2001; Pollard et al., 2000), these results suggest that either NLR-1 functions upstream of the WAVE complex in a linear signaling pathway, or both NLR-1 and the WAVE complex regulate gap junctions through F-actin. As NLR-1 does not have any potential WAVE complex binding sequence (Chen et al., 2014) and GEX-3 does not show obvious co-localization with NLR-1 or UNC-9 in neurons (Supplemental Fig. 4C and 4D), we tested whether nlr-1 regulates gap junction formation through F-actin. We first examined the effect of disassembling F-actin on gap junction formation by overexpressing a constitutively active form of actin-severing protein gsnl-1/gelsolin (P40) (Feldt et al., 2019; Meng et al., 2015). Due to the lethality caused by overexpression of gsnl-1(P40) in embryos, we carried out our studies in the nervous system and found that overexpression of gsnl-1(P40) in the nervous system caused similar gap junction formation defects as those in nlr-1(NSK) animals (Fig. 4E and 4F). More importantly, overexpression of gsnl-1(P40) did not further enhance gap junction phenotypes in nlr-1(NSK) animals, suggesting that nlr-1 may function through F-actin in the regulation of gap junction formation (Fig. 4E, 4F, and Supplemental Fig. 4E). Similar to what we observed in nlr-1(AWC/AIY SK) animals, overexpression of gsnl-1(P40) in AWC and AIY neurons also suppressed the formation of functional connexin-based gap junctions (Supplemental Fig. 4F–4H).
Figure 4. F-actin is required for gap junction formation.

(A, B) Representative images of localization of and percentage of animals with 1, 2 or ≥3 INX-3::GFP puncta on the adjoining EA/EP membranes in control, wve-1, gex-3, nlr-1, wve-1;nlr-1 and gex-3;nlr-1 knockdown animals (Scale bar: 2 μm). n≥80. Fisher’s exact test, **p < 0.005. (C) Quantification of average INX-3::GFP puncta number on the adjoining EA/EP membranes in control, gex-3, wve-1, nlr-1, gex-3;nlr-1 and wve-1;nlr-1 knockdown animals. n≥90. One-way ANOVA test, **p < 0.005. (D) Representative images of endogenous NLR-1 and INX-3 in control, gex-3 knockdown and wve-1 knockdown animals (Scale bar: 1 μm). (E, F) Representative images (E) and Quantification of puncta number (F) of UNC-9::GFP in control, nlr-1 (NSK), EX(gsnl-1(P40)) transgenes, and EX(gsnl-1(P40));nlr-1 (NSK) animals (Scale bar: 20 μm). n≥20. One-way ANOVA test, *p < 0.05.
As F-actin is a potential downstream target of nlr-1 in gap junction formation, we tested whether actin could directly bind to NLR-1 or gap junctions and found that actin could be co-immunoprecipitated with NLR-1, INX-3, and UNC-9, while tubulin did not show any interaction with NLR-1, INX-3, or UNC-9 (Fig. 5A–5D, and Supplemental Fig. 5A, 5B). NLR-1 also does not appear to directly bind to INX-3 or UNC-9 (Fig. 5E, 5F and Supplemental Fig. 5C). To address the functional significance of the interaction between NLR-1 and actin, we labeled F-actin using the calponin homology domain of F-actin binding protein utrophin (UtrCH) (Chia et al., 2014; Meng et al., 2015). We found that F-actin clustered at the center of the EA/EP membrane and partially co-localized with both NLR-1 and INX-3 in control embryos (Fig. 5G–5I). In contrast, in nlr-1 knockdown animals F-actin failed to concentrate at the gap junction formation plaque at the center of the EA/EP adjoining membrane and randomly formed patches accompanied by INX-3 along the membrane (Fig. 5H and 5I), supporting the conclusion that NLR-1 is required for the anchoring of F-actin at the formation plaque. Furthermore, we found that the intracellular domain of NLR-1 was required for its interaction with actin (Fig. 5C), and deletion of this domain from NLR-1 completely abolished its function in gap junction formation (Fig. 5J, 5K, and Supplemental Fig. 5D). These data support a model in which NLR-1 first appears at the gap junction perinexus and then recruits F-actin networks at the gap junction formation plaque, and subsequently gap junction channels interact with F-actin to be inserted into the gap junction region (Supplemental Fig. 5E).
Figure 5. NLR-1 interacts with actin to instruct gap junction formation.

(A-F) INX-3 and NLR-1 bind to actin but not tubulin. Proteins were collected from transgenes expressing GFP, INX-3::GFP (A, B); FLAG, NLR-1::FLAG, ΔNLR-1(C-terminal deletion)::FLAG (C); mNG, NLR-1::mNG (D); GFP;NLR-1::mNG, INX-3::GFP;NLR-1::mNG (E), and mNG;INX-3::GFP, INX-3::GFP;NLR-1::mNG (F) and then subjected to immunoprecipitation (IP) assays using specific antibody-coated beads as indicated. Western blots were done using antibodies presented at the right side of the images. (G-I) nlr-1 is required for F-actin to localize at the gap junction region. Representative images and fluorescence intensity profile along the EA/EP membrane of NLR-1::GFP and UtrCH::mCherry in control embryos (G), and INX-3::GFP and UtrCH::mCherry in control and nlr-1 knockdown animals (H) (Scale bar: 1 μm). (I) Percentage of UtrCh::mCherry fluorescence signal at the center of the EA/EP adjoining membrane. We evenly divided the EA/EP membrane into three sections, and the “center” was defined as the middle section. n≥20. Student’s t-test, **p < 0.005. (J, K) Representative images and quantification of UNC-9::GFP puncta in transgenes expressing either nlr-1::FLAG or Δ nlr-1(C-terminal deletion)::FLAG under the pan-neuronal Prgef-1 promoter in nlr-1 (NSK) animals (Scale bar: 20 μm). n≥20, One-way ANOVA test; *p < 0.05.
Discussion
In this study we used the C. elegans embryo as a model and carried out a RNAi-based screen to identify potential membrane or ECM proteins that are important for gap junction formation. We uncover the function of nlr-1/CASPR in regulating gap junction formation in embryonic cells, muscles and neurons, and show that NLR-1 likely functions through anchoring F-actin networks. The function of nlr-1 and its homologs appears to be conserved, as nlr-1 and CNTNAP2, the mammalian homolog of nlr-1, can regulate the formation of both innexin- and connexin- based gap junctions.
Use of invertebrates to study gap junctions
Invertebrate gap junctions are made up of innexins, and the first description of innexins was from the studies of Drosophila Pas/shaking-B and C. elegans unc-7 (Krishnan et al., 1993; Starich et al., 1993). With expression of 8 genes in Drosophila and 25 genes in C. elegans (Starich et al., 2001; Stebbings et al., 2002), innexins and innexin-based gap junctions are involved in most, it not all, aspects of animal biology: from germline maturation to embryogenesis; from neuronal development to learning and memory; from mechanosensation to locomotion; from food digestions to immune response; and from regeneration to degeneration (Guiza et al., 2018). The formation of innexin- and connexin- based gap junctions share the same three steps: 1) innexins/connexins are translated in the endoplasmic reticulum (ER), and then assembled at the Golgi apparatus; 2) after sorting into certain vesicles, hemichannels are transported to certain membrane or gap junction regions; and 3) hemichannels are inserted into a pre-defined gap junction formation plaque (Segretain and Falk, 2004). Mechanisms underlying many aspects of these steps remain unknown, including: how gap junction channels are oligomerized in the ER and then assembled at the Golgi apparatus; which vesicles and motor proteins are involved in the transportation of gap junction channels; how adjacent cells are specified to form gap junction formation plaques; and how transient gap junctions are eliminated. Compared with vertebrate systems, invertebrate model organisms have their unique advantages in addressing these questions, such as access to powerful genetic tools, amenability to live imaging at the whole animal level, and possessing robust gap-junction-associated behaviors. Fully understanding of the biology of innexin-based gap junctions will likely provide a framework for dissecting mechanisms involved in the regulation of connexin-based gap junctions. Innexin- and connexin- based gap junctions could share common regulators as shown in this study, that nlr-1/CNTNAP2 regulate the formation of innexin- and connexin- based gap junctions through F-actin.
Intracellular and intercellular signals for gap junction formation
Gap junctions are formed between two cells and are regulated by both intracellular and intercellular signals (Segretain and Falk, 2004). For the intracellular signals, post-translational modifications of gap junction channels play a key role in gap junction formation (Falk et al., 2016; Thevenin et al., 2013). Many gap junction channels have multiple phosphorylation sites, and phosphorylation at different residues has distinct functions, including affecting the forward trafficking of gap junction channels to the membrane, docking gap junction channels on the membrane, regulating the opening of gap junctions, and initiating endocytosis (Pogoda et al., 2016). Some gap junction channels, such as connexin43/Cx43, can also be mono-ubiquitinated or poly-ubiquitinated, and these modifications are suggested to be involved in regulating either trafficking or degradation of gap junction channels (Berthoud et al., 2004; Falk et al., 2014). Besides posttranslational modifications, a MAGUK (membrane-associated guanylate kinase) family member ZO-1 has been shown to be critical for the clustering and stabilization of gap junction channels at the gap junction plaque (Falk et al., 2016; Flores et al., 2008; Giepmans and Moolenaar, 1998; Marsh et al., 2017; Segretain and Falk, 2004; Toyofuku et al., 1998). ZO-1 is a plasma membrane-associated scaffolding protein with three PDZ and one Src homology (SH3) domains at the N-terminus and can directly bind to and co-localize with gap junctions (Thevenin et al., 2013). However, these regulatory mechanisms appear to be important only for transporting and stabilizing gap junction channels into the gap junction region, but play little or no role in the establishment of the formation plaque.
The intercellular signals between two gap junction partner cells are believed to be critical in defining the formation plaque (Segretain and Falk, 2004). NLR-1 is a conserved CASPR family membrane protein and has the potential to form transhomophilic interactions between two cells through its EGF-like and laminin-G domains (Zou et al., 2017). Therefore, it is possible that NLR-1 serves as a key intercellular signal to bridge two cell membranes and to define gap junction regions and participate in the establishment of the formation plaque. This is supported by our findings, including: 1) NLR-1 forms puncta at peri-gap junction regions prior to the clusters of gap junction channels; 2) Knockdown of nlr-1 causes random and unstable insertions of gap junction channels only on the membrane of one cell, and these gap junction channels are not coupled with the gap junction channels from the other cell; 3) NLR-1 is required for the clustering of F-actin networks at the gap junction region, which is important for gap junction formation. In conclusion, we believe that NLR-1 is expressed at the peri-gap junction zone and is critical for gap junction formation through the anchoring of F-actin to define or regulate the formation plaque of gap junctions.
STAR Methods
Resource availability
Lead Contact
Further information and requests for resources should be directed to and will be fulfilled by the lead contact, Dong Yan(dong.yan@duke.edu).
Materials Availability
C. elegans strains and plasmids generated in this study are available from the lead contact without restriction.
Data and Code Availability
This study did not generate any unique datasets or code.
Experimental Model and Subject Details
C. elegans genetics
C. elegans strains were maintained on nematode growth media (NGM) plates using E. coli OP50 as a food source. Animals were grown according to standard methods (Brenner, 1974) at 20°C unless otherwise stated. Wild type worms were of the Bristol N2 strain. All transgenes and strains are described in Supplemental Table II. lwIs33 [Pinx-3::inx-3::GFP] was used to visualize gap junctions. L4440 empty vector was used as the RNAi control, and E. coli strain HT115 (DE3) was used to express RNAi vectors.
Method Details
RNA interference
RNA interference (RNAi) experiments were carried out by feeding animals with E. coli strain HT115 (DE3) expressing double-stranded RNA against target genes for at least two generations. All RNAi clones were obtained either from the C. elegans Ahringer RNAi feeding library (Kamath and Ahringer, 2003) or made by inserting cDNA fragments into the L4440 vector. RNAi clones were first cultured overnight using LB broth containing ampicillin (100 μg/ml) at 37°C, and then concentrated bacteria was seeded on NGM plates containing 100μg/ml ampicillin and 5mM isopropyl 1-thio-β-D-galactopyranoside (IPTG) and grown overnight at 37°C. Animals were grown on RNAi plates for at least two generations. In all experiments, GFP RNAi or unc-22 RNAi were included as controls to account for RNAi efficiency. Tissue-specific RNAi was carried out as described (Calixto et al., 2010; Feinberg and Hunter, 2003). In brief, to knock down gene expression only in pharyngeal muscles, we expressed sid-1 (cDNA) under the control of the pharyngeal muscle specific Pmyo-2 promoter in a sid-1(pk3321) mutant background. sid-1 mutants are resistant to RNAi, and transgenic animals [yadEX1352; sid-1(pk3321)] are resistant to RNAi in all tissues except in the cells expressing sid-1 cDNA.
Protein assays
For expression studies in Supplemental Fig. 1D, we used an integrated transgene (lwIs33; cpIs55) expressing Pinx-3::inx3::GFP;Pmex-5::PH::mCherry. Proteins were extracted using SDS sample buffer containing 1mM DTT and freeze-thawed 20–50 times in dry ice/ethanol and a 37°C water bath. The samples were then denatured by heating to 95°C for 5 minutes.
For protein coimmunoprecipitation studies, worms were collected in 1xRIPA buffer containing protease inhibitor cocktail (Sigma-Aldrich). Total lysates were obtained by sonication with 15% amplified 5 second on/off pulse cycles, repeated 6 times. GFP-Trap (Chromo Tek) agarose beads, mNeonGreen-Trap magnetic beads (Chromo Tek) and Anti-FLAG M2 magnetic beads were used for immunoprecipitation.
The blots were probed with anti-GFP (1:2000, Abcam), anti-mNeongreen (1:1000, CST), anti-FLAG (1:5000, Sigma), anti-actin (1:5000, Sigma) and anti-tubulin (1:5000, Life Technologies) antibodies, then visualized with Amersham HRP-conjugated anti-rabbit or anti-mouse secondary antibodies at a 1:10000 dilution (GE Healthcare Life Sciences) using the SuperSignal West Femto kit (Pierce).
Fluorescence microscopy
Representative images were acquired with a Zeiss LSM700 confocal microscope using a Plan-Apochromat 60x/1.46 Plan-Apochromat objective. Embryos were immobilized on 1% agar slides with M9 buffer, and larva were immobilized on 1% agar slides with M9 buffer containing 1% 1-phenoxy-2-propanol. Live images were acquired using a 100x/1.4 Plan-Apochromat objective, an Andor EM-CCD camera (DU897), a spinning-disk confocal scan head (Yokogawa CSU-X1 Spinning Disk Unit), and the 488 nm (50 mW), 561 nm (50 mW) diode laser. Embryos were immobilized on 1% agar slides with M9 buffer.
In Fig. 1F, time zero was defined as 30 frames prior to the emergence of INX-3::GFP puncta. An adjacent area close to the ROI in each frame was used to subtract background signal. The fluorescence intensity relative to the initial intensity F0 were calculated as Fn - F0. Fn, fluorescence intensity at the nth frame after time zero; F0, fluorescence intensity at time zero.
Pharyngeal muscle calcium imaging
The calcium transients in the pharynx were measured by detecting changes in the fluorescence intensity of GCaMP 6.0s in L1 worms. Images were acquired with a Zeiss LSM700 confocal microscope using a 20x/0.8 Plan-Apochromat objective. Images were captured at 5 Hz. Animals were restricted from moving using 5% agar in M9 buffer with OP50 covered by a glass coverslip. The percent changes in fluorescence intensity relative to the initial intensity F0 were calculated as ΔF / F = (F – F0) / F0 × 100%. The average fluorescence intensity over 5 seconds in rest condition was taken as F0.
Measurement of pharyngeal pumping rates
Pharyngeal pumping rates were measured as described (Raizen et al., 2012). Briefly, worms were transferred to a fresh plate with OP50 or RNAi bacteria lawn and were videotaped through a stereomicroscope. Pumping rates were measured by counting the grinder movements and presented as pumps per minute.
CRISPR knock-in
yadck32 [nlr-1::mNeonGreen], yadck14 [inx-3::mKate] and yadck23 [nlr-1::frt::mKate::frt] were generated using Cas9-triggered homologous recombination with a self-excising selection cassette (SEC) (Dickinson and Goldstein, 2016). Homolog arms were amplified by PCR and then cloned into a backbone with fluorescence and SEC sequence using Gibson assembly (New England Biolabs). Synonymous substitutions were engineered in the guide RNA target sequence to disrupt the PAM. Guide RNA sequences were cloned into the Peft-3::Cas9+sgRNA expression vector pDD122 using standard mutagenesis protocols and co-injected into adult germlines with repair templates and extrachromosomal array markers (Dickinson et al., 2013). Candidate strains were selected by hygromycin B treatment, phenotypic identification (roller), and the absence of fluorescent extrachromosomal arrays. Primers for generating those strains are listed in Supplemental Table III.
Quantification and Statistical Analysis
Statistical Analysis
All data analysis was done using Graphpad Prism. For INX-3 or NLR-1 dynamic analyses, each point represents 5 experiments of at least 200 events. Quantification of percentage of animals with 1, 2 or ≥3 INX-3::GFP puncta on the adjoining membranes of EA/EP cells, at least 80 animals were analyzed. For quantification of average number and size of puncta, each data represents at least 3 experiments with more than 90 animals. For calcium imaging data, at least 60 animals were analyzed. For all behavior assays, at least 30 animals were analyzed. For fluorescence signal quantification, at least 20 animals were analyzed. For western blot, results were repeated at least 3 times. At detailed in the Figure legends, Student’s t-test, One-way ANOVA followed by Fisher’s LSD post-hoc test or Fisher’s exact test were conducted using GraphPad Prism 8. A p-value cutoff of 0.05 was considered statistically significant. The error bars in the figures indicate ± SEM.
Supplementary Material
Key Resource Table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| GFP-Trap Agarose | ChromoTek | # gtd-10, RRID: AB_2827592 |
| mNeonGreen-Trap Magnetic Agarose | ChromoTek | # ntma-10, RRID: AB_2827594 |
| Anti-FLAG M2 magnetic beads | Sigma-Aldrich | # M8823, RRID: AB_2637089 |
| Rabbit anti-GFP | Abcam | # ab183734, RRID: AB_2732027 |
| Rabbit anti-mNeongreen | CST | #53061 |
| Mouse Monolonal anti-FLAG M2 | Sigma-Aldrich | # F3165, RRID: AB_259529 |
| Rabbit Anti-actin | Sigma-Aldrich | # A2066, RRID: AB_476693 |
| Mouse anti-tubulin | Life Technologies | # 322600, RRID: AB_2533072 |
| Amersham HRP-conjugated anti-rabbit secondary antibodies | GE Healthcare Life Sciences | NA934, RRID: AB_772206 |
| Amersham HRP-conjugated anti-mouse secondary antibodies | GE Healthcare Life Sciences | NA931, RRID: AB_772210 |
| Chemicals, Peptides, and Recombinant Proteins | ||
| Benzaldehyde | Sigma-Aldrich | # B1334, CAS RN: 100-52-7 |
| Hygromycin | Sigma-Aldrich | # H3274, CAS RN: 31282-04-9 |
| Recombinant DNA | ||
| Pinx-3::Gcamp6.0 | this paper | PNYL963 |
| Pinx-3::mKate | this paper | PNYL964 |
| Pdpy-30::nlr-1 | this paper | PNYL992 |
| Pmyo-2::sid-1 | this paper | PNYL1199 |
| Punc-9::gfp::unc-9 | this paper | PNYL384 |
| Prgef-1::Flp::NLS | this paper | PNYL1198 |
| Prgef-1::nlr-1 | this paper | PNYL1167 |
| Prgef-1::CNTNAP2 | this paper | PNYL1341 |
| Podr-1::Flp::NLS | this paper | PNYL1329 |
| Pttx-3::Flp::NLS | this paper | PNYL1328 |
| Pttx-3::Cx43::msfgfp | this paper | PNYL1327 |
| Podr-1::Cx43::msfgfp | this paper | PNYL1323 |
| Prgef-1::gex-3::mCherry | this paper | PNYL1331 |
| Prgef-1::gsnl-1(P40) | this paper | PNYL1342 |
| Pdpy-30::gfp | this paper | PNYL1269 |
| Pdpy-30::3xflag | this paper | PNYL1343 |
| Pdpy-30::nlr-1::3xflag | this paper | PNYL1292 |
| Pinx-3::gfp::inx-3 | this paper | PNYL598 |
| Pdpy-30::mCherry::UtrCh | this paper | PNYL1277 |
| Pdpy-30:: △nlr-1::3xflag | this paper | PNYL1315 |
| Pdpy-30::mNeongreen | this paper | PNYL1263 |
| Pttx-3::gsnl-1(P40) | this paper | PNYL1344 |
| Podr-1::gsnl-1(P40) | this paper | PNYL1345 |
| Bacterial and Virus Strains | ||
| Escherichia coli OP50 | http://www.wormbook.org/chapters/www_strainmaintain/strainmaintain.html | N/A |
| Software and Algorithms | ||
| Graphpad Prism 8 | Graphpad Software | N/A |
| Fiji | imageJ | N/A |
All transgenes generated in this study are listed in the Supplemental Table 2.
All primers used this study are listed in the Supplemental Table 3
Highlights:
NLR-1 locates in the gap junction perinexus to regulate gap junction formation
NLR-1 can recruit F-actin networks at the gap junction formation plaque
The formation of F-actin patches plays a critical role in the assembly of gap junctions
nlr-1 and its mammalian homolog can form connexin-based gap junctions in C. elegans
Acknowledgements
We thank Dr. Jocelyn Shaw and Dr. Todd Starich for the EH489 strain [ncl-1(e1865) unc-36(e251)]; lwIs33 [inx-3::gfp; unc-36(+)], Dr. William Schafer and Denise Walker for Cx36 transgenes, and Bob Goldstein, Ari Pani, David Sherwood and Qiuyi Chi for sharing CRISPR protocol and plasmids. We also want to thank Albert Zhang, Monroe Monroe and other Yan lab members for comments on the manuscript. Caitlin Lopez, Robert Cline, Charlton Lu, Carson Herman, Kyle Ockerman, and many other undergraduate students in the lab were also involved in this project. Some strains were provided by the CGC, which is funded by the NIH Office of Research Infrastructure Programs (P40 OD010440). This project is supported by a NIH R01 grant NS105638 (to D.Y.). L.M. and D.Y. are also supported by a NIH R01 grant NS094171 (to D.Y.).
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Competing financial interests
The authors declare no competing financial interests.
Supplemental tables
Supplemental table 1, Genes tested for EA/EP gap junction formation, Related to Fig. 1.
References
- Albertson DG, and Thomson JN (1976). The pharynx of Caenorhabditis elegans. Philos Trans R Soc Lond B Biol Sci 275, 299–325. [DOI] [PubMed] [Google Scholar]
- Alexopoulos H, Bottger A, Fischer S, Levin A, Wolf A, Fujisawa T, Hayakawa S, Gojobori T, Davies JA, David CN, et al. (2004). Evolution of gap junctions: the missing link? Current biology : CB 14, R879–880. [DOI] [PubMed] [Google Scholar]
- Altun ZF, Chen B, Wang ZW, and Hall DH (2009). High resolution map of Caenorhabditis elegans gap junction proteins. Dev Dyn 238, 1936–1950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bauer R, Loer B, Ostrowski K, Martini J, Weimbs A, Lechner H, and Hoch M (2005). Intercellular communication: the Drosophila innexin multiprotein family of gap junction proteins. Chemistry & biology 12, 515–526. [DOI] [PubMed] [Google Scholar]
- Bauer R, Weimbs A, Lechner H, and Hoch M (2006). DE-cadherin, a core component of the adherens junction complex modifies subcellular localization of the Drosophila gap junction protein innexin2. Cell communication & adhesion 13, 103–114. [DOI] [PubMed] [Google Scholar]
- Berthoud VM, Minogue PJ, Laing JG, and Beyer EC (2004). Pathways for degradation of connexins and gap junctions. Cardiovascular research 62, 256–267. [DOI] [PubMed] [Google Scholar]
- Bhattacharya A, Aghayeva U, Berghoff EG, and Hobert O (2019). Plasticity of the Electrical Connectome of C. elegans. Cell 176, 1174–1189 e1116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bossinger O, and Schierenberg E (1996). Cell-cell communication in nematode embryos: differences between Cephalobus spec. and Caenorhabditis elegans. Dev Genes Evol 206, 25–34. [DOI] [PubMed] [Google Scholar]
- Brenner S (1974). The genetics of Caenorhabditis elegans. Genetics 77, 71–94. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Butkevich E, Hulsmann S, Wenzel D, Shirao T, Duden R, and Majoul I (2004). Drebrin is a novel connexin-43 binding partner that links gap junctions to the submembrane cytoskeleton. Current biology : CB 14, 650–658. [DOI] [PubMed] [Google Scholar]
- Calixto A, Chelur D, Topalidou I, Chen X, and Chalfie M (2010). Enhanced neuronal RNAi in C. elegans using SID-1. Nat Methods 7, 554–559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Calkins TL, Woods-Acevedo MA, Hildebrandt O, and Piermarini PM (2015). The molecular and immunochemical expression of innexins in the yellow fever mosquito, Aedes aegypti: insights into putative life stage- and tissue-specific functions of gap junctions. Comparative biochemistry and physiology Part B, Biochemistry & molecular biology 183, 11–21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chakraborty S, Mitra S, Falk MM, Caplan SH, Wheelock MJ, Johnson KR, and Mehta PP (2010). E-cadherin differentially regulates the assembly of Connexin43 and Connexin32 into gap junctions in human squamous carcinoma cells. The Journal of biological chemistry 285, 10761–10776. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen B, Brinkmann K, Chen Z, Pak CW, Liao Y, Shi S, Henry L, Grishin NV, Bogdan S, and Rosen MK (2014). The WAVE regulatory complex links diverse receptors to the actin cytoskeleton. Cell 156, 195–207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen B, Liu Q, Ge Q, Xie J, and Wang ZW (2007). UNC-1 regulates gap junctions important to locomotion in C. elegans. Current biology : CB 17, 1334–1339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chia PH, Chen B, Li P, Rosen MK, and Shen K (2014). Local F-actin network links synapse formation and axon branching. Cell 156, 208–220. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chuang CF, Vanhoven MK, Fetter RD, Verselis VK, and Bargmann CI (2007). An innexin-dependent cell network establishes left-right neuronal asymmetry in C. elegans. Cell 129, 787–799. [DOI] [PubMed] [Google Scholar]
- Connors BW, and Long MA (2004). Electrical synapses in the mammalian brain. Annual review of neuroscience 27, 393–418. [DOI] [PubMed] [Google Scholar]
- Depriest A, Phelan P, and Martha Skerrett I (2011). Tryptophan scanning mutagenesis of the first transmembrane domain of the innexin Shaking-B(Lethal). Biophys J 101, 2408–2416. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dickinson DJ, and Goldstein B (2016). CRISPR-Based Methods for Caenorhabditis elegans Genome Engineering. Genetics 202, 885–901. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dickinson DJ, Ward JD, Reiner DJ, and Goldstein B (2013). Engineering the Caenorhabditis elegans genome using Cas9-triggered homologous recombination. Nat Methods 10, 1028–1034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eden S, Rohatgi R, Podtelejnikov AV, Mann M, and Kirschner MW (2002). Mechanism of regulation of WAVE1-induced actin nucleation by Rac1 and Nck. Nature 418, 790–793. [DOI] [PubMed] [Google Scholar]
- Falk MM, Bell CL, Kells Andrews RM, and Murray SA (2016). Molecular mechanisms regulating formation, trafficking and processing of annular gap junctions. BMC cell biology 17 Suppl 1, 22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Falk MM, Kells RM, and Berthoud VM (2014). Degradation of connexins and gap junctions. FEBS letters 588, 1221–1229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feinberg EH, and Hunter CP (2003). Transport of dsRNA into cells by the transmembrane protein SID-1. Science 301, 1545–1547. [DOI] [PubMed] [Google Scholar]
- Feldt J, Schicht M, Garreis F, Welss J, Schneider UW, and Paulsen F (2019). Structure, regulation and related diseases of the actin-binding protein gelsolin. Expert Rev Mol Med 20, e7. [DOI] [PubMed] [Google Scholar]
- Fields RD, and Nelson PG (1992). Activity-dependent development of the vertebrate nervous system. Int Rev Neurobiol 34, 133–214. [DOI] [PubMed] [Google Scholar]
- Flores CE, Li X, Bennett MV, Nagy JI, and Pereda AE (2008). Interaction between connexin35 and zonula occludens-1 and its potential role in the regulation of electrical synapses. Proceedings of the National Academy of Sciences of the United States of America 105, 12545–12550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Franks CJ, Holden-Dye L, Bull K, Luedtke S, and Walker RJ (2006). Anatomy, physiology and pharmacology of Caenorhabditis elegans pharynx: a model to define gene function in a simple neural system. Invert Neurosci 6, 105–122. [DOI] [PubMed] [Google Scholar]
- Furshpan EJ, and Potter DD (1959). Transmission at the giant motor synapses of the crayfish. J Physiol 145, 289–325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao S, Guan SA, Fouad AD, Meng J, Kawano T, Huang YC, Li Y, Alcaire S, Hung W, Lu Y, et al. (2018). Excitatory motor neurons are local oscillators for backward locomotion. eLife 7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giepmans BN, and Moolenaar WH (1998). The gap junction protein connexin43 interacts with the second PDZ domain of the zona occludens-1 protein. Current biology : CB 8, 931–934. [DOI] [PubMed] [Google Scholar]
- Giepmans BN, Verlaan I, Hengeveld T, Janssen H, Calafat J, Falk MM, and Moolenaar WH (2001). Gap junction protein connexin-43 interacts directly with microtubules. Current biology : CB 11, 1364–1368. [DOI] [PubMed] [Google Scholar]
- Golic KG, and Lindquist S (1989). The FLP recombinase of yeast catalyzes site-specific recombination in the Drosophila genome. Cell 59, 499–509. [DOI] [PubMed] [Google Scholar]
- Goodenough DA, and Paul DL (2009). Gap junctions. Cold Spring Harbor perspectives in biology 1, a002576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Govindarajan R, Chakraborty S, Johnson KE, Falk MM, Wheelock MJ, Johnson KR, and Mehta PP (2010). Assembly of connexin43 into gap junctions is regulated differentially by E-cadherin and N-cadherin in rat liver epithelial cells. Molecular biology of the cell 21, 4089–4107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guiza J, Barria I, Saez JC, and Vega JL (2018). Innexins: Expression, Regulation, and Functions. Front Physiol 9, 1414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haklai-Topper L, Soutschek J, Sabanay H, Scheel J, Hobert O, and Peles E (2011). The neurexin superfamily of Caenorhabditis elegans. Gene Expr Patterns 11, 144–150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hall DH (2017). Gap junctions in C. elegans: Their roles in behavior and development. Developmental neurobiology 77, 587–596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hand AR, and Gobel S (1972). The structural organization of the septate and gap junctions of Hydra. J Cell Biol 52, 397–408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang H, Hayden DJ, Zhu CT, Bennett HL, Venkatachalam V, Skuja LL, and Hart AC (2018). Gap Junctions and NCA Cation Channels Are Critical for Developmentally Timed Sleep and Arousal in Caenorhabditis elegans. Genetics 210, 1369–1381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hunter AW, Barker RJ, Zhu C, and Gourdie RG (2005). Zonula occludens-1 alters connexin43 gap junction size and organization by influencing channel accretion. Mol Biol Cell 16, 5686–5698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jang H, Levy S, Flavell SW, Mende F, Latham R, Zimmer M, and Bargmann CI (2017). Dissection of neuronal gap junction circuits that regulate social behavior in Caenorhabditis elegans. Proceedings of the National Academy of Sciences of the United States of America 114, E1263–E1272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jin X, Pokala N, and Bargmann CI (2016). Distinct Circuits for the Formation and Retrieval of an Imprinted Olfactory Memory. Cell 164, 632–643. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson R, Hammer M, Sheridan J, and Revel JP (1974). Gap junction formation between reaggregated Novikoff hepatoma cells. Proc Natl Acad Sci U S A 71, 4536–4540. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson RG, Reynhout JK, TenBroek EM, Quade BJ, Yasumura T, Davidson KG, Sheridan JD, and Rash JE (2012). Gap junction assembly: roles for the formation plaque and regulation by the C-terminus of connexin43. Mol Biol Cell 23, 71–86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kamath RS, and Ahringer J (2003). Genome-wide RNAi screening in Caenorhabditis elegans. Methods 30, 313–321. [DOI] [PubMed] [Google Scholar]
- Kamath RS, Martinez-Campos M, Zipperlen P, Fraser AG, and Ahringer J (2001). Effectiveness of specific RNA-mediated interference through ingested double-stranded RNA in Caenorhabditis elegans. Genome biology 2, RESEARCH0002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kawano T, Po MD, Gao S, Leung G, Ryu WS, and Zhen M (2011). An imbalancing act: gap junctions reduce the backward motor circuit activity to bias C. elegans for forward locomotion. Neuron 72, 572–586. [DOI] [PubMed] [Google Scholar]
- Kidder GM, Rains J, and McKeon J (1987). Gap junction assembly in the preimplantation mouse conceptus is independent of microtubules, microfilaments, cell flattening, and cytokinesis. Proceedings of the National Academy of Sciences of the United States of America 84, 3718–3722. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krishnan SN, Frei E, Swain GP, and Wyman RJ (1993). Passover: a gene required for synaptic connectivity in the giant fiber system of Drosophila. Cell 73, 967–977. [DOI] [PubMed] [Google Scholar]
- Lauf U, Giepmans BN, Lopez P, Braconnot S, Chen SC, and Falk MM (2002). Dynamic trafficking and delivery of connexons to the plasma membrane and accretion to gap junctions in living cells. Proceedings of the National Academy of Sciences of the United States of America 99, 10446–10451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu P, Chen B, Mailler R, and Wang ZW (2017). Antidromic-rectifying gap junctions amplify chemical transmission at functionally mixed electrical-chemical synapses. Nat Commun 8, 14818. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu P, Chen B, and Wang ZW (2011). Gap junctions synchronize action potentials and Ca2+ transients in Caenorhabditis elegans body wall muscle. The Journal of biological chemistry 286, 44285–44293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maeda S, Nakagawa S, Suga M, Yamashita E, Oshima A, Fujiyoshi Y, and Tsukihara T (2009). Structure of the connexin 26 gap junction channel at 3.5 A resolution. Nature 458, 597–602. [DOI] [PubMed] [Google Scholar]
- Marsh AJ, Michel JC, Adke AP, Heckman EL, and Miller AC (2017). Asymmetry of an Intracellular Scaffold at Vertebrate Electrical Synapses. Current biology : CB 27, 3561–3567 e3564. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meng L, Chen CH, and Yan D (2016a). Regulation of Gap Junction Dynamics by UNC-44/ankyrin and UNC-33/CRMP through VAB-8 in C. elegans Neurons. PLoS genetics 12, e1005948. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meng L, Mulcahy B, Cook SJ, Neubauer M, Wan A, Jin Y, and Yan D (2015). The Cell Death Pathway Regulates Synapse Elimination through Cleavage of Gelsolin in Caenorhabditis elegans Neurons. Cell reports 11, 1737–1748. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meng L, Zhang A, Jin Y, and Yan D (2016b). Regulation of neuronal axon specification by glia-neuron gap junctions in C. elegans. eLife 5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mennerick S, and Zorumski CF (2000). Neural activity and survival in the developing nervous system. Mol Neurobiol 22, 41–54. [DOI] [PubMed] [Google Scholar]
- Mese G, Richard G, and White TW (2007). Gap junctions: basic structure and function. The Journal of investigative dermatology 127, 2516–2524. [DOI] [PubMed] [Google Scholar]
- Oshima A, Matsuzawa T, Murata K, Tani K, and Fujiyoshi Y (2016a). Hexadecameric structure of an invertebrate gap junction channel. Journal of molecular biology 428, 1227–1236. [DOI] [PubMed] [Google Scholar]
- Oshima A, Tani K, and Fujiyoshi Y (2016b). Atomic structure of the innexin-6 gap junction channel determined by cryo-EM. Nature communications 7, 13681. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pantaloni D, Le Clainche C, and Carlier MF (2001). Mechanism of actin-based motility. Science 292, 1502–1506. [DOI] [PubMed] [Google Scholar]
- Peng YR, Tran NM, Krishnaswamy A, Kostadinov D, Martersteck EM, and Sanes JR (2017). Satb1 Regulates Contactin 5 to Pattern Dendrites of a Mammalian Retinal Ganglion Cell. Neuron 95, 869–883 e866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Phelan P (2005). Innexins: members of an evolutionarily conserved family of gap-junction proteins. Biochimica et biophysica acta 1711, 225–245. [DOI] [PubMed] [Google Scholar]
- Pogoda K, Kameritsch P, Retamal MA, and Vega JL (2016). Regulation of gap junction channels and hemichannels by phosphorylation and redox changes: a revision. BMC cell biology 17 Suppl 1, 11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poliak S, Gollan L, Martinez R, Custer A, Einheber S, Salzer JL, Trimmer JS, Shrager P, and Peles E (1999). Caspr2, a new member of the neurexin superfamily, is localized at the juxtaparanodes of myelinated axons and associates with K+ channels. Neuron 24, 1037–1047. [DOI] [PubMed] [Google Scholar]
- Poliak S, Salomon D, Elhanany H, Sabanay H, Kiernan B, Pevny L, Stewart CL, Xu X, Chiu SY, Shrager P, et al. (2003). Juxtaparanodal clustering of Shaker-like K+ channels in myelinated axons depends on Caspr2 and TAG-1. J Cell Biol 162, 1149–1160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pollard TD, Blanchoin L, and Mullins RD (2000). Molecular mechanisms controlling actin filament dynamics in nonmuscle cells. Annu Rev Biophys Biomol Struct 29, 545–576. [DOI] [PubMed] [Google Scholar]
- Preus D, Johnson R, Sheridan J, and Meyer R (1981). Analysis of gap junctions and formation plaques between reaggregating Novikoff hepatoma cells. J Ultrastruct Res 77, 263–276. [DOI] [PubMed] [Google Scholar]
- Rabinowitch I, Chatzigeorgiou M, Zhao B, Treinin M, and Schafer WR (2014). Rewiring neural circuits by the insertion of ectopic electrical synapses in transgenic C. elegans. Nature communications 5, 4442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raizen D, Song BM, Trojanowski N, and You YJ (2012). Methods for measuring pharyngeal behaviors. WormBook, 1–13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raizen DM, and Avery L (1994). Electrical activity and behavior in the pharynx of Caenorhabditis elegans. Neuron 12, 483–495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rhett JM, Jourdan J, and Gourdie RG (2011). Connexin 43 connexon to gap junction transition is regulated by zonula occludens-1. Molecular biology of the cell 22, 1516–1528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Santafe MM, Salon I, Garcia N, Lanuza MA, Uchitel OD, and Tomas J (2004). Muscarinic autoreceptors related with calcium channels in the strong and weak inputs at polyinnervated developing rat neuromuscular junctions. Neuroscience 123, 61–73. [DOI] [PubMed] [Google Scholar]
- Sauer B (1987). Functional expression of the cre-lox site-specific recombination system in the yeast Saccharomyces cerevisiae. Mol Cell Biol 7, 2087–2096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sauer B, and Henderson N (1988). Site-specific DNA recombination in mammalian cells by the Cre recombinase of bacteriophage P1. Proc Natl Acad Sci U S A 85, 5166–5170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sawa M, Suetsugu S, Sugimoto A, Miki H, Yamamoto M, and Takenawa T (2003). Essential role of the C. elegans Arp2/3 complex in cell migration during ventral enclosure. J Cell Sci 116, 1505–1518. [DOI] [PubMed] [Google Scholar]
- Schumacher JA, Hsieh YW, Chen S, Pirri JK, Alkema MJ, Li WH, Chang C, and Chuang CF (2012). Intercellular calcium signaling in a gap junction-coupled cell network establishes asymmetric neuronal fates in C. elegans. Development 139, 4191–4201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwarz V, Pan J, Voltmer-Irsch S, and Hutter H (2009). IgCAMs redundantly control axon navigation in Caenorhabditis elegans. Neural Dev 4, 13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Segretain D, and Falk MM (2004). Regulation of connexin biosynthesis, assembly, gap junction formation, and removal. Biochim Biophys Acta 1662, 3–21. [DOI] [PubMed] [Google Scholar]
- Shaw RM, Fay AJ, Puthenveedu MA, von Zastrow M, Jan YN, and Jan LY (2007). Microtubule plus-end-tracking proteins target gap junctions directly from the cell interior to adherens junctions. Cell 128, 547–560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shimozono S, Fukano T, Kimura KD, Mori I, Kirino Y, and Miyawaki A (2004). Slow Ca2+ dynamics in pharyngeal muscles in Caenorhabditis elegans during fast pumping. EMBO Rep 5, 521–526. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simonsen KT, Moerman DG, and Naus CC (2014). Gap junctions in C. elegans. Front Physiol 5, 40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Starich T, Sheehan M, Jadrich J, and Shaw J (2001). Innexins in C. elegans. Cell Commun Adhes 8, 311–314. [DOI] [PubMed] [Google Scholar]
- Starich TA, Herman RK, and Shaw JE (1993). Molecular and genetic analysis of unc-7, a Caenorhabditis elegans gene required for coordinated locomotion. Genetics 133, 527–541. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Starich TA, Miller A, Nguyen RL, Hall DH, and Shaw JE (2003). The Caenorhabditis elegans innexin INX-3 is localized to gap junctions and is essential for embryonic development. Developmental biology 256, 403–417. [DOI] [PubMed] [Google Scholar]
- Stebbings LA, Todman MG, Phillips R, Greer CE, Tam J, Phelan P, Jacobs K, Bacon JP, and Davies JA (2002). Gap junctions in Drosophila: developmental expression of the entire innexin gene family. Mech Dev 113, 197–205. [DOI] [PubMed] [Google Scholar]
- Sulston JE, Schierenberg E, White JG, and Thomson JN (1983). The embryonic cell lineage of the nematode Caenorhabditis elegans. Dev Biol 100, 64–119. [DOI] [PubMed] [Google Scholar]
- Thevenin AF, Kowal TJ, Fong JT, Kells RM, Fisher CG, and Falk MM (2013). Proteins and mechanisms regulating gap-junction assembly, internalization, and degradation. Physiology 28, 93–116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Toyofuku T, Yabuki M, Otsu K, Kuzuya T, Hori M, and Tada M (1998). Direct association of the gap junction protein connexin-43 with ZO-1 in cardiac myocytes. The Journal of biological chemistry 273, 12725–12731. [DOI] [PubMed] [Google Scholar]
- Traka M, Goutebroze L, Denisenko N, Bessa M, Nifli A, Havaki S, Iwakura Y, Fukamauchi F, Watanabe K, Soliven B, et al. (2003). Association of TAG-1 with Caspr2 is essential for the molecular organization of juxtaparanodal regions of myelinated fibers. J Cell Biol 162, 1161–1172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Voelker L, Upadhyaya B, Ferkey DM, Woldemariam S, L’Etoile ND, Rabinowitch I, and Bai J (2019). INX-18 and INX-19 play distinct roles in electrical synapses that modulate aversive behavior in Caenorhabditis elegans. PLoS genetics 15, e1008341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Watanabe A (1958). The interaction of electrical activity among neurons of lobster cardiac ganglion. Jpn J Physiol 8, 305–318. [DOI] [PubMed] [Google Scholar]
- Wei CJ, Francis R, Xu X, and Lo CW (2005). Connexin43 associated with an N-cadherin-containing multiprotein complex is required for gap junction formation in NIH3T3 cells. The Journal of biological chemistry 280, 19925–19936. [DOI] [PubMed] [Google Scholar]
- Welzel G, and Schuster S (2018). Long-term potentiation in an innexin-based electrical synapse. Scientific reports 8, 12579. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Withee J, Galligan B, Hawkins N, and Garriga G (2004). Caenorhabditis elegans WASP and Ena/VASP proteins play compensatory roles in morphogenesis and neuronal cell migration. Genetics 167, 1165–1176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu X, Li WE, Huang GY, Meyer R, Chen T, Luo Y, Thomas MP, Radice GL, and Lo CW (2001). N-cadherin and Cx43alpha1 gap junctions modulates mouse neural crest cell motility via distinct pathways. Cell communication & adhesion 8, 321–324. [DOI] [PubMed] [Google Scholar]
- Yeh E, Kawano T, Ng S, Fetter R, Hung W, Wang Y, and Zhen M (2009). Caenorhabditis elegans innexins regulate active zone differentiation. J Neurosci 29, 5207–5217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zou Y, Zhang WF, Liu HY, Li X, Zhang X, Ma XF, Sun Y, Jiang SY, Ma QH, and Xu DE (2017). Structure and function of the contactin-associated protein family in myelinated axons and their relationship with nerve diseases. Neural regeneration research 12, 1551–1558. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
This study did not generate any unique datasets or code.


