Abstract
Disrupted nucleocytoplasmic transport (NCT) has been implicated in neurodegenerative disease pathogenesis; however, the mechanisms by which disrupted NCT causes neurodegeneration remain unclear. In a Drosophila screen, we identified ref(2)P/p62, a key regulator of autophagy, as a potent suppressor of neurodegeneration caused by the GGGGCC hexanucleotide repeat expansion (G4C2 HRE) in C9orf72 that causes amyotrophic lateral sclerosis (ALS) and frontotemporal dementia (FTD). We found that p62 is increased and forms ubiquitinated aggregates due to decreased autophagic cargo degradation. Immunofluorescence and electron microscopy of Drosophila tissues demonstrate an accumulation of lysosome-like organelles that precedes neurodegeneration. These phenotypes are partially caused by cytoplasmic mislocalization of Mitf/TFEB, a key transcriptional regulator of autophagolysosomal function. Additionally, TFEB is mislocalized and downregulated in human cells expressing GGGGCC repeats and in C9-ALS patient motor cortex. Our data suggest that the C9orf72-HRE impairs Mitf/TFEB nuclear import, thereby disrupting autophagy and exacerbating proteostasis defects in C9-ALS/FTD.
Research organism: D. melanogaster
Introduction
A GGGGCC (G4C2) hexanucleotide repeat expansion (HRE) in chromosome nine open reading frame 72 (C9orf72) is the most common genetic cause of amyotrophic lateral sclerosis (ALS) and frontotemporal dementia (FTD), accounting for up to 40% of cases of familial ALS (DeJesus-Hernandez et al., 2011; ITALSGEN Consortium et al., 2011). ALS and/or FTD caused by mutations in C9orf72 (C9-ALS/FTD) is inherited in an autosomal dominant manner, suggesting that the HRE causes disease through gain-of-function or haploinsufficiency (DeJesus-Hernandez et al., 2011; Ling et al., 2013). Loss of C9orf72 function has been linked to disruption of autophagy and lysosome function, though neurodegeneration is not observed in C9orf72 knockout mice (Liu et al., 2016; Shi et al., 2018; Webster et al., 2016), suggesting that C9-ALS/FTD is primarily caused by toxicity of the HRE. Furthermore, expression of G4C2 repeats causes neurotoxicity in Drosophila and cell culture models of C9-ALS (Goodman et al., 2019a; Kramer et al., 2016; Tran et al., 2015). This toxicity has been proposed to occur through either G4C2 repeat RNA-mediated sequestration of RNA-binding proteins or translation of the G4C2 repeats into dipeptide-repeat proteins (DPRs) through non-canonical repeat-associated non-AUG (RAN) translation (Donnelly et al., 2013; Goodman et al., 2019a; Mori et al., 2013; Tran et al., 2015).
We previously conducted a Drosophila screen of candidate proteins that bound with moderate-to-high affinity to G4C2 RNA and identified modulation of the nucleocytoplasmic transport (NCT) pathway as a potent modifier of G4C2 toxicity in both fly and iPS neuron models of C9-ALS (Zhang et al., 2015a), a finding that has also been made by other groups (Freibaum et al., 2015; Jovičić et al., 2015). The mechanisms by which the G4C2 HRE disrupts NCT remain unclear, but potential mechanisms include G4C2 RNA binding to the master NCT regulator RanGAP (Zhang et al., 2015a), DPRs disrupting the nuclear pore complex (Boeynaems et al., 2016; Shi et al., 2017; Zhang et al., 2016), stress granules sequestering NCT factors (Zhang et al., 2018), or cytoplasmic TDP-43-dependent dysregulation of karyopherin-α (Chou et al., 2018; Gasset-Rosa et al., 2019; Solomon et al., 2018). Recently, a role for NCT disruption in Huntington’s disease and Alzheimer’s disease has been proposed, indicating that NCT disruption may be a common mechanism in several neurodegenerative diseases (Eftekharzadeh et al., 2018; Gasset-Rosa et al., 2017; Grima et al., 2017). However, the pathways affected by NCT disruption that cause neurodegeneration have not yet been elucidated.
In a Drosophila screen for modifiers of G4C2-mediated neurodegeneration (Zhang et al., 2015a), we identified refractory to sigma P (ref(2)P), the Drosophila homolog of p62/SQSTM1 (Sequestosome 1). p62/SQSTM1 functions in macroautophagy (hereafter termed autophagy), and mutations in p62/SQSTM1 are a rare genetic cause of ALS/FTD (Cirulli et al., 2015; Le Ber et al., 2013; Teyssou et al., 2013). Interestingly, many other genes implicated in ALS/FTD function in autophagy (Evans and Holzbaur, 2019; Lin et al., 2017; Ramesh and Pandey, 2017) such as tank-binding kinase 1 (TBK1), optineurin (OPTN1), ubiquilin 2 and 4 (UBQLN2 and 4), valosin-containing protein (VCP), charged multivesicular body protein 2B (CHMP2B), VAMP-associated protein B (VapB), and the C9orf72 protein itself (O'Rourke et al., 2015; Sellier et al., 2016; Sullivan et al., 2016; Ugolino et al., 2016; Webster et al., 2016; Yang et al., 2016). Organelles and protein aggregates are degraded via polyubiquitination and targeting to a newly forming autophagosome, followed by degradation upon fusion with the lysosome. Deletion of key autophagy genes in neurons is sufficient to cause neurodegeneration in mice (Hara et al., 2006; Komatsu et al., 2006).
Although autophagy and nucleocytoplasmic transport have both been implicated in neurodegeneration, it is unclear whether or how these two pathways interact in disease pathogenesis (Gao et al., 2017; Thomas et al., 2013). Here, we show that expression of expanded G4C2 repeats is sufficient to disrupt autophagy in Drosophila, leading to an accumulation of p62 and ubiquitinated protein aggregates. We find that autophagolysosomal defects are caused by loss of nuclear localization of the transcription factor Mitf (the Drosophila homolog of TFEB), which regulates transcription of genes involved in autophagolysosome biogenesis (Bouché et al., 2016; Palmieri et al., 2011; Sardiello et al., 2009; Zhang et al., 2015b). Furthermore, suppressing this NCT defect is sufficient to rescue Mitf nuclear localization, restoring autophagy and lysosome function and rescuing neurodegeneration. These findings suggest a pathogenic cascade in C9-ALS/FTD whereby NCT disruption causes a failure of autophagosome biogenesis and lysosome dysfunction that ultimately leads to neuronal death.
Results
Ref(2)P/p62 knockdown suppresses G4C2-mediated neurodegeneration
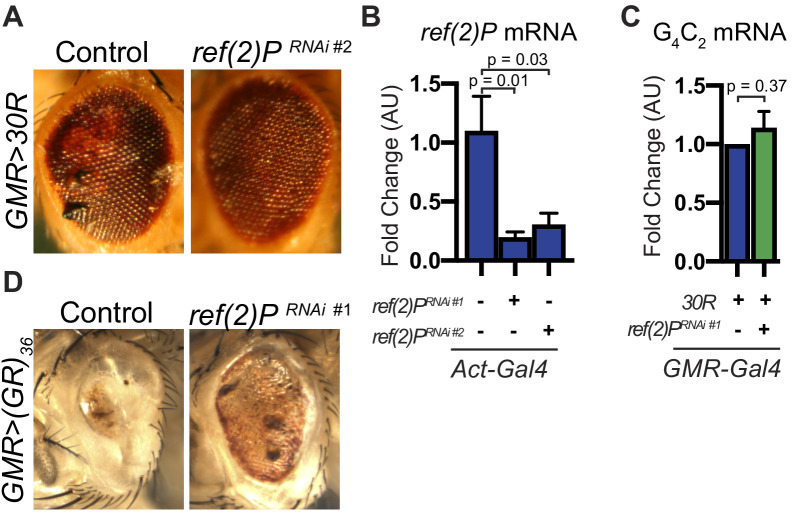
Expression of 30 G4C2 repeats (30R) in the eye using GMR-Gal4 results in progressive photoreceptor degeneration and visible ommatidial disruption by day 15 (Figure 1A; Xu et al., 2013; Zhang et al., 2015a). In a genetic modifier screen of over 800 RNAi lines, UAS-ref(2)PRNAi was among the strongest of 32 suppressors of G4C2-mediated eye degeneration (Zhang et al., 2015a; Figure 1A). ref(2)P is the Drosophila homolog of p62/SQSTM1, and this modifier is of particular interest because SQSTM1 mutations that cause loss of selective autophagy cause ALS/FTD (Cirulli et al., 2015; Goode et al., 2016; Le Ber et al., 2013), and p62 aggregates are pathological features of both familial and sporadic ALS (Al-Sarraj et al., 2011; Cooper-Knock et al., 2012). Knockdown of ref(2)P suppresses eye degeneration, whereas overexpression of ref(2)P enhances this phenotype (Figure 1A–B, Figure 1—figure supplement 1A). ref(2)P RNAi #1 expression reduced ref(2)P mRNA levels by ~80%, but did not alter G4C2 RNA levels in 30R expressing eyes (Figure 1—figure supplement 1B–C), suggesting that ref(2)P acts downstream of G4C2 transcription. Similarly, knockdown of ref(2)P also rescued eye degeneration in a second G4C2 model expressing 36 G4C2 repeats (36R) (Mizielinska et al., 2014; Figure 1C–D). We next assessed the ability of ref(2)P RNAi to rescue toxicity of G4C2 repeats in motor neurons using the 30R model and a new G4C2 model expressing 44R (Goodman et al., 2019b). As shown in Figure 1E–F, while expression of either 30R or 44R in motor neurons with vGlut-Gal4 leads to paralysis and lethality during pupal development, knockdown of ref(2)P partially rescues this phenotype, whereas overexpression of ref(2)P enhances the pupal lethality observed with 44R expression. These data suggest that ref(2)P is required for G4C2-mediated toxicity during Drosophila development.
Figure 1. Autophagy receptor Ref(2)P/p62 genetically suppresses G4C2-HRE-mediated degeneration.
(A) 15-day-old Drosophila eyes expressing GMR-Gal4 +/- UAS-30R (GMR>30R) with RNAi background (control), ref(2)P RNAi#1 or overexpression (OE) of ref(2)P. (B) Quantification of external eye degeneration in A by semi-quantitative scoring system. Data are reported as mean ± SEM. Kruskal-Wallis test, p<0.0001, followed by Dunn’s multiple comparisons, n > 15 adults. (C) 15-day-old Drosophila eyes expressing GMR-Gal4 +/- UAS-36R (GMR >36R) along with UAS-luciferase RNAi (control), UAS-ref(2)P RNAi #1, UAS-ref(2)P RNAi #2, or UAS-ref(2)P OE. (D) Quantification of external eye degeneration in C by semi-quantitative scoring system. Data are reported as mean ± SEM. Kruskal-Wallis test, p<0.0001, followed by Dunn’s multiple comparisons, n = 23, 62, 28, 10 adults respectively. (E) Percent of pupal eclosion of adult flies expressing the motor neuron driver vGlut-Gal4 +/- UAS-30R and RNAi background control or UAS-ref(2)P RNAi #1. Fisher’s exact test, n > 100 pupa. (F) Percent of pupal eclosion of adult flies expressing the motor neuron driver vGlut-Gal4 +/- UAS-44R along with UAS-luciferase RNAi, UAS-ref(2)P RNAi #1, or UAS-ref(2)P OE. Fisher’s exact test, n > 55 pupa. (G) Adult Drosophila expressing UAS-30R under the control of the inducible, pan-neuronal elavGS induced with 200 µM RU486 or vehicle alone and co-expressing control or UAS-ref(2)P RNAi #1. Data are reported as mean ± SEM. One-way ANOVA, ****p<0.0001, with Sidak’s multiple comparisons test, n = 9, 8, 8, 8 groups of 10 flies.
Figure 1—figure supplement 1. Ref(2)P/p62 genetically modifies G4C2-HRE.
To determine whether ref(2)P knockdown is able to suppress age-dependent neurodegeneration, we used a pan-neuronal, inducible ‘GeneSwitch’ driver (elavGS) in which 30R-expression leads to a marked reduction in climbing ability after 7 days (Figure 1G). This climbing defect is suppressed with coexpression of ref(2)P RNAi, suggesting that ref(2)P contributes to G4C2-mediated neurotoxicity in the adult nervous system. Since RAN-translation of arginine-containing DPRs have been implicated in G4C2-mediated toxicity in Drosophila (Kwon et al., 2014; Mizielinska et al., 2014), we next tested whether ref(2)P knockdown rescues poly-glycine-arginine (GR) repeat-mediated toxicity. As shown in Figure 1—figure supplement 1D, ref(2)P RNAi partially rescues the severe eye degeneration phenotype caused by poly(GR)36 expression. Together, these data indicate that ref(2)P, the Drosophila orthologue of p62/SQSTM1, modulates G4C2-mediated neurodegeneration.
G4C2 repeat expression impairs autophagic flux
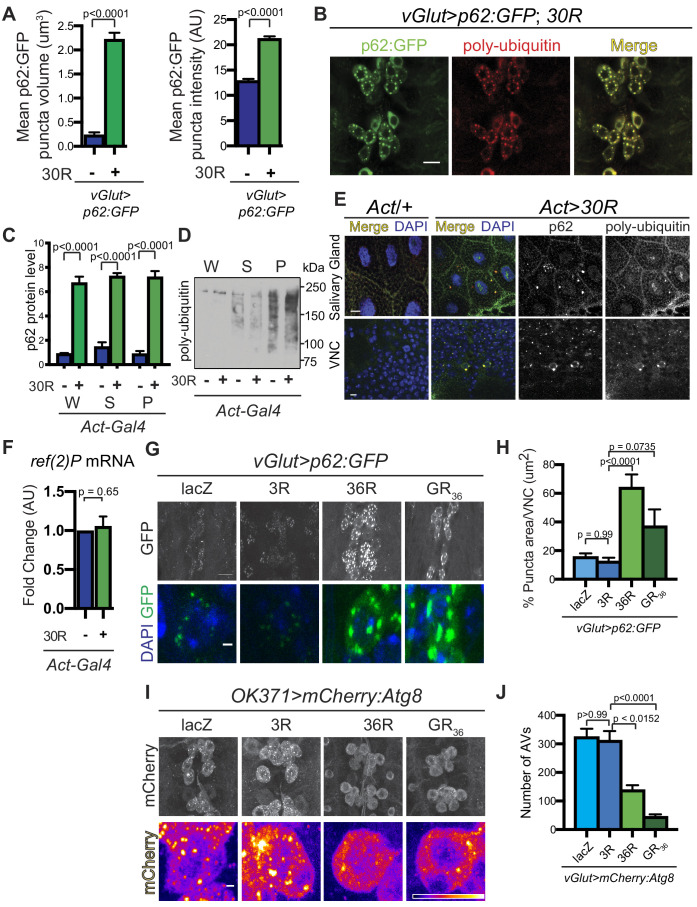
p62/SQSTM1-positive inclusions are a common pathologic feature seen in brains of C9-ALS/FTD patients where they colocalize with ubiquitin and DPRs (Al-Sarraj et al., 2011). We next investigated the localization of Ref(2)P protein (hereafter referred to as p62) in motor neurons. Expression of 30R leads to the formation of many large p62:GFP puncta in cell bodies compared to controls that strongly colocalize with poly-ubiquitinated proteins (Figure 2A–B, Figure 2—figure supplement 1A–B). Western blot analysis demonstrates that p62 and poly-ubiquitin are strongly upregulated in flies ubiquitously expressing 30R (Figure 2C, Figure 2—figure supplement 1C–D). Similarly, immunofluorescence staining with a p62 antibody shows endogenous p62 accumulations colocalizing with polyubiquitinated proteins in the ventral nerve cord and salivary gland of flies ubiquitously overexpressing 30R (Figure 2—figure supplement 1E). These data show that G4C2 repeat expression in fly models recapitulates the p62 accumulation with ubiquitinated protein aggregates seen in C9-ALS/FTD patient tissue and iPS neurons (Almeida et al., 2013; Mackenzie et al., 2014).
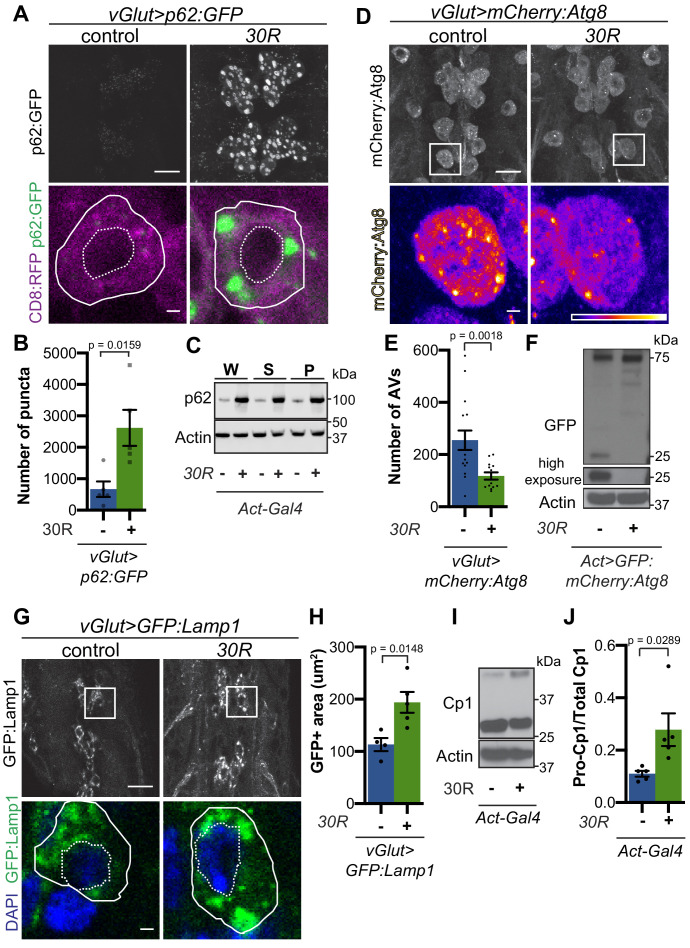
Figure 2. G4C2 repeat expression impairs autophagic flux.
(A) Drosophila motor neurons expressing UAS-p62:GFP +/- UAS-30R, showing multiple motor neuron cell bodies (top) or a representative cell co-expressing the membrane marker CD8:RFP (bottom). Plasma membrane outlined with solid white line; nucleus outlined with dotted line. Scale bar = 10 µm (top), 1 µm (bottom) (B) Quantification of number of p62:GFP puncta in Drosophila motor neuron cell bodies. Data are reported as mean ± SEM. Mann-Whitney test, n = 5 larvae per genotype. (C) Western blot of anti-p62 and anti-beta-actin showing the whole (W), supernatant (S) and pellet (P) fractions of lysates from Drosophila larvae ubiquitously expressing -/+ UAS-30R under the control of Act-Gal4. (D) Drosophila motor neurons expressing UAS-mCherry:Atg8 -/+ UAS-30R showing cell bodies (top) with an example single cell highlighting mCherry:Atg8-positive puncta (bottom). Scale bar = 10 µm (top), 1 µm (bottom). (E) Quantification of mCherry:Atg8-positive autophagic vesicles (AVs) in the ventral nerve cord of vGlut-Gal4/+ or vGlut >30R expressing flies. Data are reported as mean ± SEM. Mann-Whitney test, n = 16 and 13 larvae, respectively. (F) Western blot of anti-GFP and anti-beta-actin of lysates from whole Drosophila larvae ubiquitously expressing UAS-GFP:mCherry:Atg8 -/+ UAS-30R under the control of Act-Gal4 showing full length GFP:mCherry:Atg8 at 75 kDa and cleaved GFP at 25 kDa. (G) Drosophila motor neurons expressing UAS-GFP:Lamp1 (with N-terminal [luminal] GFP) -/+ UAS-30R under the control of vGlut-Gal4 in multiple cell bodies (top) or in a representative cell (bottom). Scale bar = 10 µm (top), 1 µm (bottom). (H) Quantification of GFP:Lamp1 positive area in G. Data are reported as mean ± SEM. Student’s t-test, n = 5 larvae. (I) Western of whole Act-Gal4 Drosophila larvae -/+ UAS-30R blotted for the lysosomal protease Cp1, showing pro- (inactive, upper band) and cleaved (active, lower band) Cp1. (J) Quantification of the ratio of pro-Cp1 to total Cp1 in I. Data are reported as mean ± SEM. Student’s t-test, n = 5 biological replicates.
Figure 2—figure supplement 1. p62:GFP accumulates in C9-ALS fly models and co-localizes with poly-ubiquitin.
Figure 2—figure supplement 2. Rescuing G4C2-mediated lysosome defects reduces neurodegeneration.
Increased p62 levels can be due to either increased transcription and/or translation or insufficient protein degradation (Korolchuk et al., 2010). Using qRT-PCR, we find that ref(2)P transcript levels are unchanged in G4C2 repeat-expressing larvae (Figure 2—figure supplement 1F), suggesting that G4C2 repeats cause p62 upregulation by inhibiting p62 degradation. Since p62 is degraded by autophagy and disrupted autophagic flux is known to cause p62 upregulation, we assessed autophagy in G4C2-repeat-expressing flies. We first co-expressed the tagged autophagosome marker mCherry:Autophagy-related 8 (Atg8, the fly orthologue of mammalian Microtubule-associated protein 1A/1B-light chain 3 (LC3)) with 30R in fly motor neurons and found a marked reduction in mCherry:Atg8 autophagic esicles (AVs) when compared to wild-type controls (Figure 2D–E). p62:GFP accumulation and loss of mCherry:Atg8 puncta were recapitulated in 36R and poly(GR)36 Drosophila models of C9-ALS/FTD (Figure 2—figure supplement 1G–J). Reduction of mCherry:Atg8-positive vesicles coupled with the accumulation of p62 and ubiquitin suggest that autophagic flux is impaired in these fly models of C9-ALS/FTD.
G4C2 repeat expression causes lysosome defects
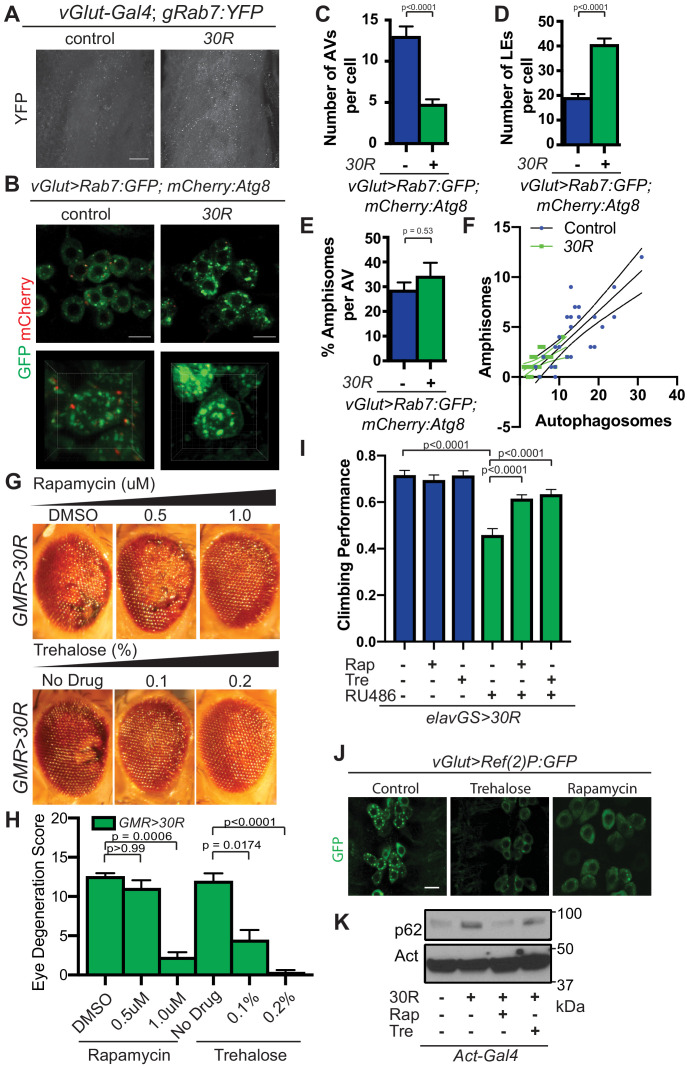
To further study lysosomal morphology and function, we expressed Lysosome- associated membrane protein 1 (Lamp1) with luminally-tagged GFP in our control and G4C2-expressing flies. Since GFP is largely quenched by the acidity of lysosomes in control animals (Pulipparacharuvil et al., 2005), the accumulation of GFP:Lamp-positive vesicles in 30R-expressing motor neurons suggests a defect in lysosomal acidity or targeting of GFP:Lamp to mature lysosomes (Figure 2G–H). Furthermore, we observe a marked increase in size and number of late endosomes and lysosomes using genomically tagged Ras-related GTP-binding protein 7, Rab7:YFP, throughout 30R-expressing motor neurons (Figure 2—figure supplement 2A) without alterations in early endosomes labeled with Rab5:YFP (data not shown). Together, these data demonstrate a marked expansion of the late endosome/lysosome compartment in G4C2-expressing neurons.
Though accumulation of p62 and ubiquitinated proteins could be caused by a failure of autophagic vesicles to fuse with the degradative endolysosomal compartment, we did not detect a decrease in mCherry:Atg8+, Rab7:GFP+ amphisomes in G4C2-expressing motor neuron cell bodies (Figure 2—figure supplement 2B–F). To assess autophagolysosomal function after fusion, we performed a GFP liberation assay on larvae expressing GFP:mCherry:Atg8 (Klionsky et al., 2016; Mauvezin et al., 2014). GFP is degraded more slowly than the rest of the mCherry:Atg8 protein, leaving a population of free GFP in functioning lysosomes. Free lysosomal GFP is not observed in G4C2-expressing larvae, suggesting an impairment in GFP:mCherry:Atg8 degradation by the lysosome (Figure 2F). To directly probe lysosome enzymatic activity, we performed Western analysis of Drosophila cathepsin Cp1. Whereas pro-Cp1 is normally cleaved to its mature form by acid hydrolases in lysosomes (Kinser and Dolph, 2012), larvae ubiquitously expressing 30R show an increase in the ratio of pro-Cp1 to Cp1, indicating a decrease in pro-Cp1 cleavage efficiency (Figure 2I–J). Together, these data suggest that lysosomes are expanded and dysfunctional in G4C2 repeat-expressing animals.
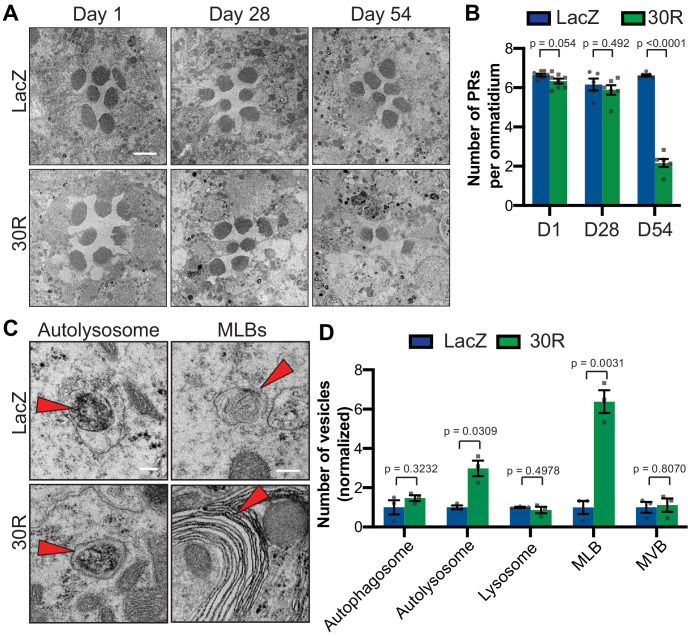
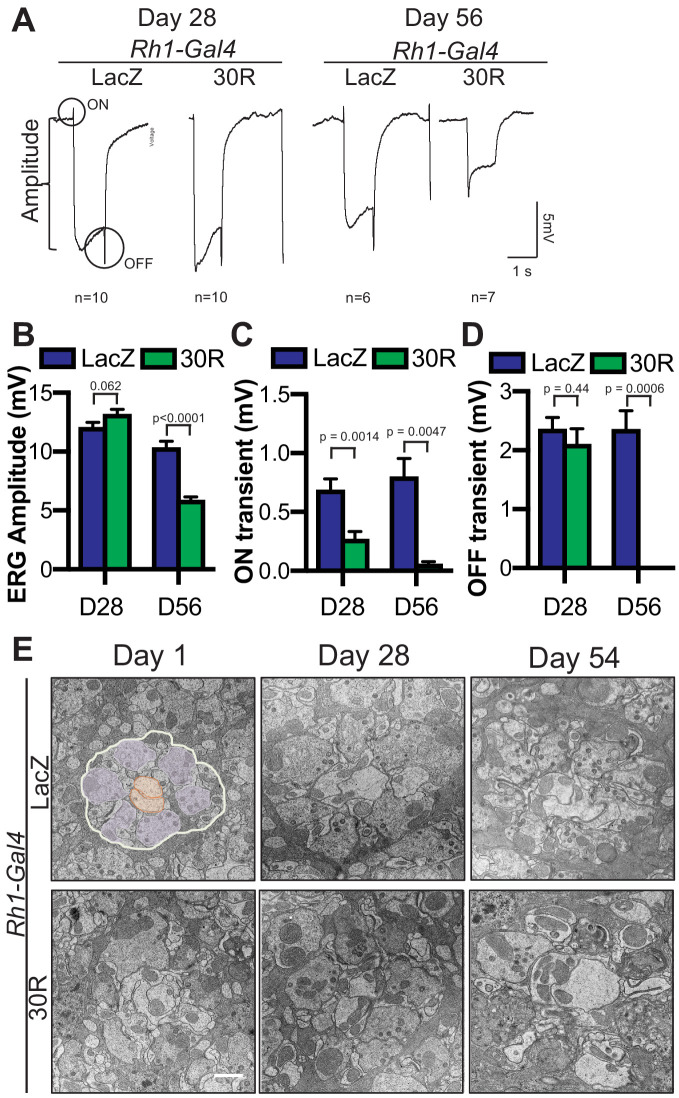
To investigate whether the autophagic pathway defects precede neurodegeneration in G4C2 repeat-expressing neurons, we performed transmission electron microscopy (TEM) on Drosophila eyes. As GMR-Gal4 is expressed throughout the development of the eye, we chose to perform electroretinograms (ERGs) of fly eyes selectively expressing 30R in photoreceptor neurons (PRs) using Rh1-Gal4, which turns on during adulthood. Rh1 >30R PRs show only a mild reduction of ON transient amplitude at 28 days, but a complete loss of ON and OFF transients and a decrease in ERG amplitude by 56 days (Figure 3—figure supplement 1A–D), indicating a slow and progressive loss of synaptic transmission and impaired phototransduction respectively. These changes also correspond to a marked loss of photoreceptors and synaptic terminals by 54 days which are not observed at 28 days (Figure 3A–B; Figure 3—figure supplement 1E). We therefore examined autophagic structures by TEM at 28 days, prior to cell loss. Strikingly, we observe a marked increase in the size and number of multilamellar bodies (MLBs) (Figure 3C–D). MLBs are commonly observed in lysosomal storage diseases and result from a deficiency of lysosomal hydrolases and accumulations of lysosomal lipids and membranes (Hariri et al., 2000; Weaver et al., 2002). Though we did not detect an alteration in the number of autophagosomes, lysosomes, or multivesicular bodies, we did see a significant increase in the number of autolysosomes (Figure 3C–D). These data suggest that autophagolysosomal function is disrupted in G4C2-expressing photoreceptor neurons at early stages of degeneration.
Figure 3. Autophagolysosomal defects precede neurodegeneration in photoreceptor neurons.
(A) Transmission electron microscopy (TEM) of rhabdomeres (cell bodies) in Rhodopsin1-Gal4 (Rh1-Gal4) driving UAS-LacZ (control) or UAS-30R at Day 1, Day 28, and Day 54 after eclosion. Scale bar = 2 µm. (B) Quantification of number of healthy (not split) photoreceptors (PRs) per ommatidium in A. Data are reported as mean ± SEM. Student’s t-test, n = 8, 8, 6, 6, 6, and 6 flies, respectively. (C) TEM images at 28 days of Drosophila eyes (rhabdomeres) -/+ 30R repeats expressed by Rh1-Gal4 showing representative autolysosomes and multilamellar bodies (MLBs), marked with red arrows. Scale bar = 200 nm. (D) Quantification of different vesicle types (autophagosomes, autolysosomes, lysosomes, MLBs, and multivesicular bodies (MVBs)) shown in TEM of rhabdomeres with Rh1-Gal4 driving UAS-LacZ or UAS-30R (as in C) normalized to LacZ (control). Data are reported as mean ± SEM. Student’s t-test, n = 3 adults per genotype.
Figure 3—figure supplement 1. Progressive synapse degeneration in G4C2-expressing photoreceptor neurons.
Given the impairment in autophagic flux, we hypothesized that genetic or pharmacologic manipulations that accelerate autophagy may suppress neurodegenerative phenotypes, whereas those that further impede autophagy would enhance the phenotypes. Indeed, in a candidate-based screen, activation of early steps in the autophagic pathway (e.g. by Atg1 overexpression) suppresses eye degeneration and blocking autophagosome/lysosome fusion (e.g. by Snap29 knockdown) enhances eye degeneration (Supplementary file 1). Similarly, pharmacologic activation of autophagy via inhibition of mTor with rapamycin or mTor-independent activation via trehalose (Sarkar et al., 2007) rescues neurodegenerative phenotypes and p62 accumulation (Figure 2—figure supplement 2G–K). Together, these data show that promoting autophagy or lysosomal fusion are potent suppressors of G4C2-mediated neurodegeneration.
Nucleocytoplasmic transport impairment disrupts autophagic flux
A diverse array of cellular pathways including autophagy, RNA homeostasis, and NCT are implicated in the pathogenesis of ALS and FTD (Balendra and Isaacs, 2018; Evans and Holzbaur, 2019; Gao et al., 2017; Lin et al., 2017; Ling et al., 2013; Ramesh and Pandey, 2017). However, the sequence of events in the pathogenic cascade remains unknown. Cytoplasmic protein aggregates or RNA stress granule formation is sufficient to disrupt nucleocytoplasmic transport (Woerner et al., 2016; Zhang et al., 2018). We therefore tested whether defects in autophagy are upstream, downstream, or in parallel with defects in NCT.
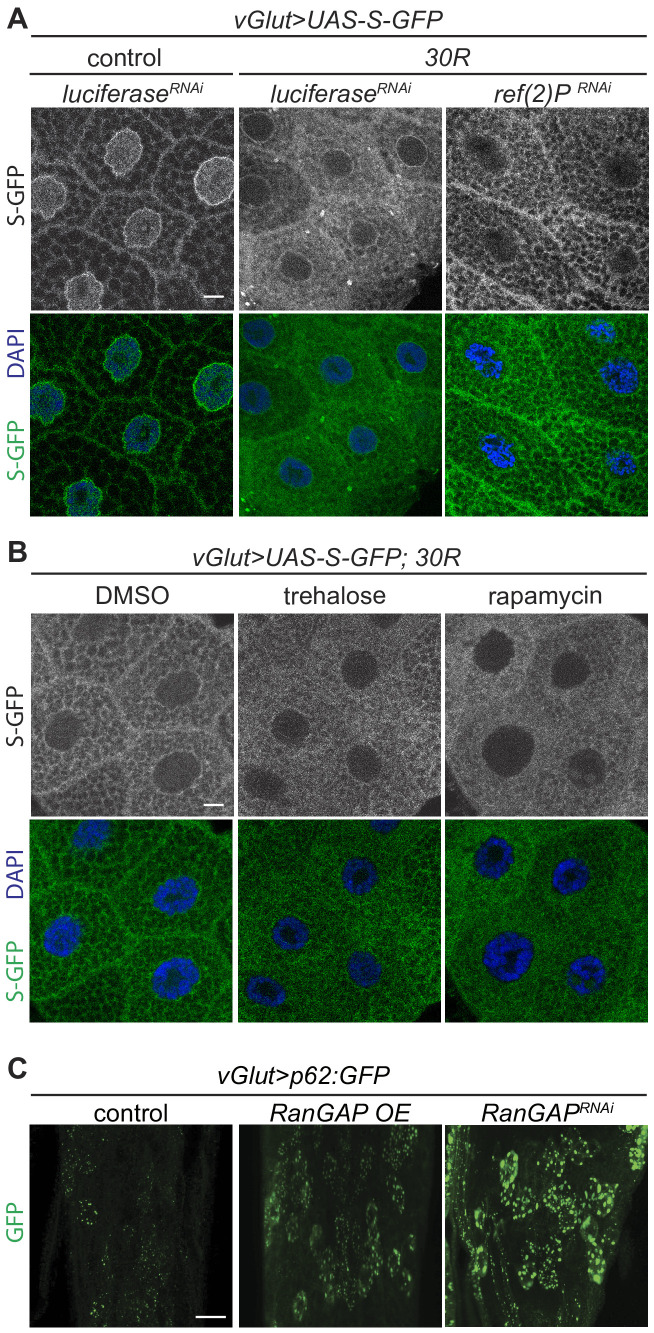
We first tested whether knockdown of ref(2)P rescues the mislocalization of the NCT reporter shuttle-GFP (S-GFP) containing both a nuclear localization sequence (NLS) and nuclear export sequence (NES). G4C2 repeat expression causes mislocalization of S-GFP to the cytoplasm (Zhang et al., 2015a), but knockdown of ref(2)P does not restore nuclear localization (Figure 5—figure supplement 1A). Similarly, stimulation of autophagy with rapamycin or trehalose fails to rescue S-GFP mislocalization in G4C2 expressing salivary glands (Figure 5—figure supplement 1B). Stimulating autophagy does not rescue NCT defects although it can rescue neurodegeneration, suggesting that autophagy defects are either independent of or downstream of NCT defects. Indeed, RanGAP knockdown increases the number and size of p62:GFP puncta, similar to the effects of overexpressing the G4C2 repeats (Figure 5—figure supplement 1C), suggesting that NCT disruption is sufficient to disrupt autophagic flux in Drosophila motor neurons.
Mitf is mislocalized and inactivated in Drosophila models of C9-ALS/FTD
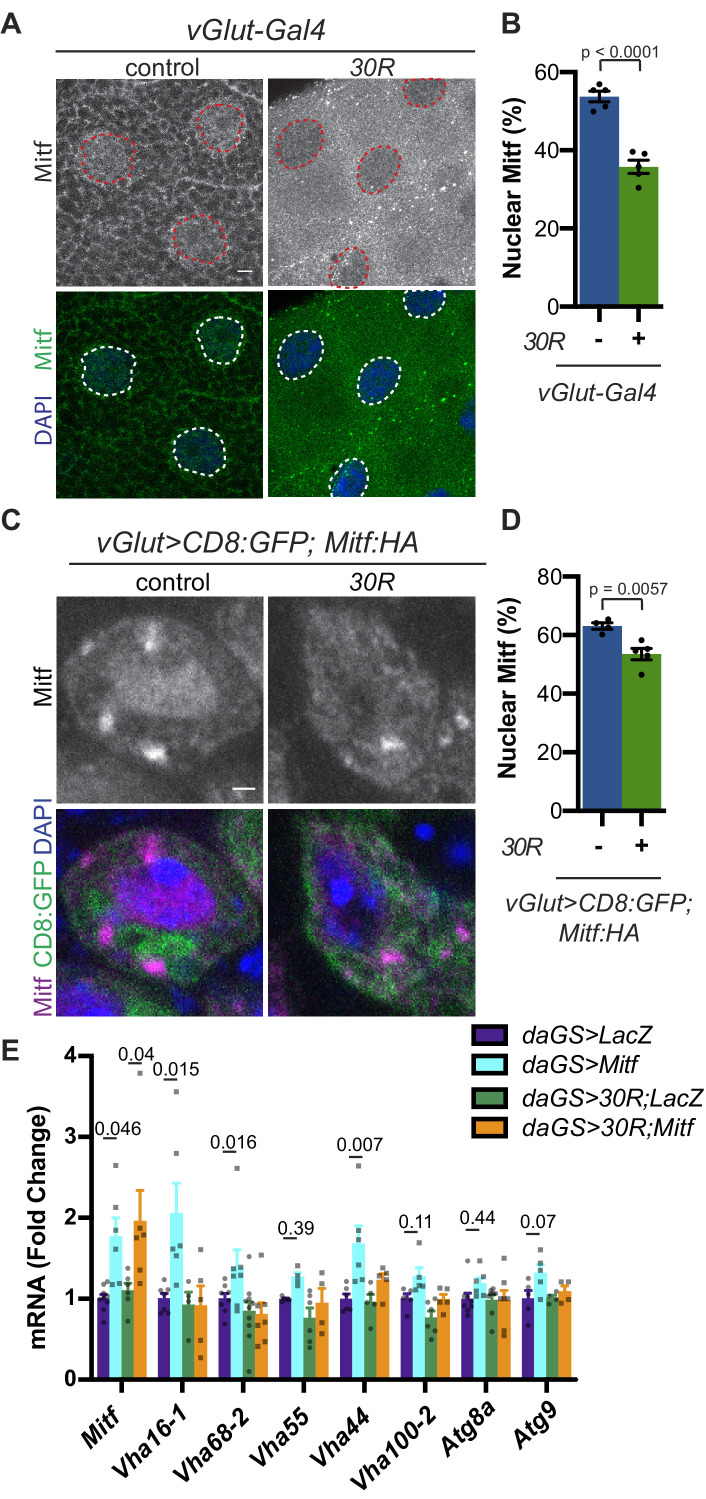
Because we observed a reduction in autophagosomes and expansion of lysosome-related organelles, we hypothesized that transcription factors regulating autophagolysosomal function may be mislocalized to the cytoplasm due to disrupted nuclear import. The MiT/TFE family of transcription factors (TFEB, TFE3, MITF, and TFEC) regulates multiple steps of autophagy from autophagosome biogenesis through lysosome acidification via a network of genes called the Coordinated Lysosome Expression And Regulation (CLEAR) network (Settembre et al., 2011). These transcription factors are regulated by localization between the cytoplasm and nucleus (Li et al., 2018). In Drosophila, this conserved transcription factor family is represented by a single homolog called Mitf (Bouché et al., 2016; Zhang et al., 2015b). Mitf knockdown in the nervous system causes lysosomal defects similar to those observed in G4C2-expressing flies (Bouché et al., 2016; Hallsson et al., 2004; Sardiello et al., 2009; Song et al., 2013). Additionally, TFEB levels are reduced in superoxide dismutase 1 (SOD1) mutant cell culture and mouse ALS models (Chen et al., 2015) as well as in ALS and Alzheimer’s patient brain tissue (Wang et al., 2016). Therefore, we hypothesized that impaired Mitf nucleocytoplasmic transport might underlie the autophagolysosomal phenotypes in fly models of C9-ALS. Indeed, both salivary gland cells and motor neurons expressing 30R show a reduction in percent nuclear Mitf (Figure 4A–D). To assess whether disrupted Mitf NCT alters CLEAR gene expression in adult heads, we expressed 30R using a ubiquitous inducible driver, daughterless-GeneSwitch (daGS). In control flies, a mild (~1.75 fold) overexpression of Mitf mRNA resulted in a significant upregulation of 3 of the 7 Mitf targets tested (the vesicular ATPase (v-ATPase) subunits Vha16-1, Vha68-2, and Vha44) and a trend towards upregulation of 4 others (Figure 4E). Importantly, co-expression of 30R with daGS >Mitf led to a similar ~2 fold increase in Mitf transcripts but did not induce Mitf target genes (Figure 4E). This lack of Mitf target induction in 30R flies suggests that decreased nuclear import of Mitf suppresses the ability of 30R-expressing flies to upregulate CLEAR genes in order to maintain or induce autophagic flux.
Figure 4. Mitf/TFEB is mislocalized from the nucleus and inactivated.
(A) Drosophila larval salivary glands -/+ UAS-30R under the control of vGlut-Gal4 stained with anti-Mitf and DAPI. Dotted lines outline nuclei. Scale bar = 10 µm. (B) Quantification of percent (%) nuclear Mitf (nuclear Mitf fluorescence/total fluorescence) in A. Data are reported as mean ± SEM. Student’s t-test, n = 5 larvae per genotype. (C) Drosophila motor neurons (MNs) expressing UAS-Mitf-HA and UAS-CD8:GFP -/+ UAS-30R under the control of vGlut-Gal4 stained with anti-HA, anti-GFP (membrane), and DAPI to show nuclear localization. Scale bar = 1 µm. (D) Quantification of percent (%) nuclear Mitf in C. Data are reported as mean ± SEM. Student’s t-test, n = 4 and 5 larvae, respectively, with at least 10 motor neurons per larva. (E) Quantitative RT-PCR to assess transcript levels of Mitf and seven target genes from lysates of Drosophila heads expressing control (UAS-LacZ) or UAS-30R driven by daGS in control conditions or with overexpression of Mitf. Data are reported as mean ± SEM. One-way ANOVA, p<0.0001, with Sidak’s multiple comparisons test, n > 4 biological replicates of 30 heads per genotype.
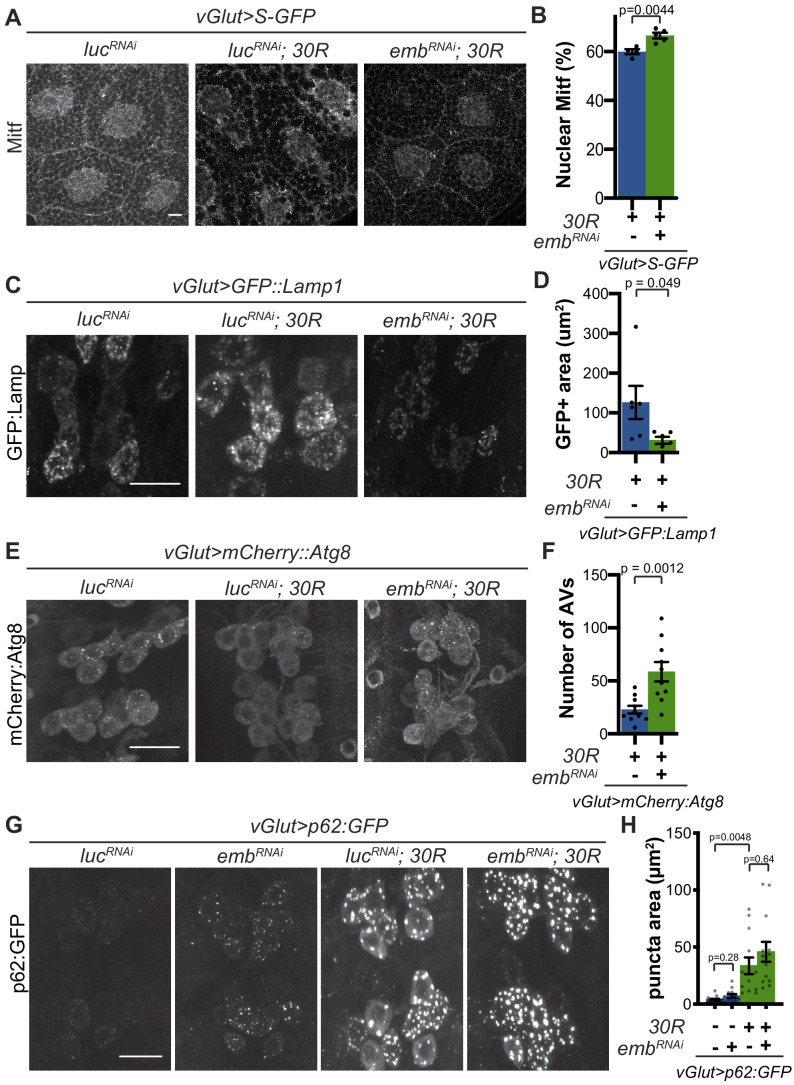
We next examined whether rescue of nucleocytoplasmic transport defects in 30R-expressing animals can rescue Mitf nuclear import and autophagolysosomal defects. Exportin-1 has recently been demonstrated to regulate Mitf/TFEB nuclear export (Li et al., 2018; Silvestrini et al., 2018). Knockdown of exportin-1 (Drosophila emb) rescues G4C2-mediated cytoplasmic Mitf mislocalization in the salivary gland (Figure 5A–B) and GFP:Lamp accumulation in motor neurons (Figure 5C–D). Importantly, emb knockdown increases the total number of autophagosomes in G4C2-expressing motor neuron cell bodies by ~3 fold (Figure 5E–F), suggesting that nuclear retention of Mitf rescues autophagolysosomal defects. However, emb knockdown caused a slight elevation of p62:GFP puncta intensity in controls and did not rescue the accumulations of p62:GFP in 30R-expressing motor neurons (Figure 5G–H). Together, these data indicate that autophagolysosomal dysfunction in 30R-expressing animals occurs downstream of nucleocytoplasmic transport disruption, whereas inhibition of nuclear export is not sufficient to rescue p62 accumulation.
Figure 5. Modulation of nucleocytoplasmic transport rescues autophagolysosome dysfunction.
(A) Drosophila larval salivary glands stained with anti-Mitf and DAPI expressing +/- UAS-30R, UAS-shuttle-GFP (S-GFP, not shown), and either control RNAi (UAS-lucRNAi) or exportin RNAi (UAS-embRNAi ) under the control of vGlut-Gal4. Scale bar = 10 µm (B) Quantification of percent (%) nuclear Mitf in A. Data are reported as mean ± SEM. Student’s t-test, n = 4 and 5 larvae, respectively. (C) Drosophila motor neurons expressing UAS-GFP:Lamp1 (N-terminal, luminal GFP) -/+ UAS-30R and UAS-lucRNAi or exportin RNAi (UAS-embRNAi ). Scale bar = 10 µm. (D) Quantification of C. Student’s t-test, n = 6 larvae. (E) Drosophila motor neurons expressing UAS-mCherry:Atg8 +/- UAS-30R and either control RNAi (UAS-lucRNAi) or exportin RNAi (UAS-embRNAi ). Scale bar = 10 µm. (F) Quantification of E. Data are reported as mean ± SEM. Mann-Whitney test, n = 10 larvae. (G) Drosophila motor neurons expressing UAS-p62:GFP -/+ UAS-30R and either control RNAi (lucRNAi) or exportin RNAi (embRNAi ) under the control of vGlut-Gal4. Scale bar = 10 µm. (H) Quantification of G. Data are reported as mean ± SEM. Brown-Forsythe and Welch ANOVA test, p<0.0001, followed by Dunnett’s T3 multiple comparisons, n = 12–14 larvae per genotype.
Figure 5—figure supplement 1. Nucleocytoplasmic transport disruption is upstream of autophagic defects.
Mitf rescues G4C2 repeat-mediated degeneration
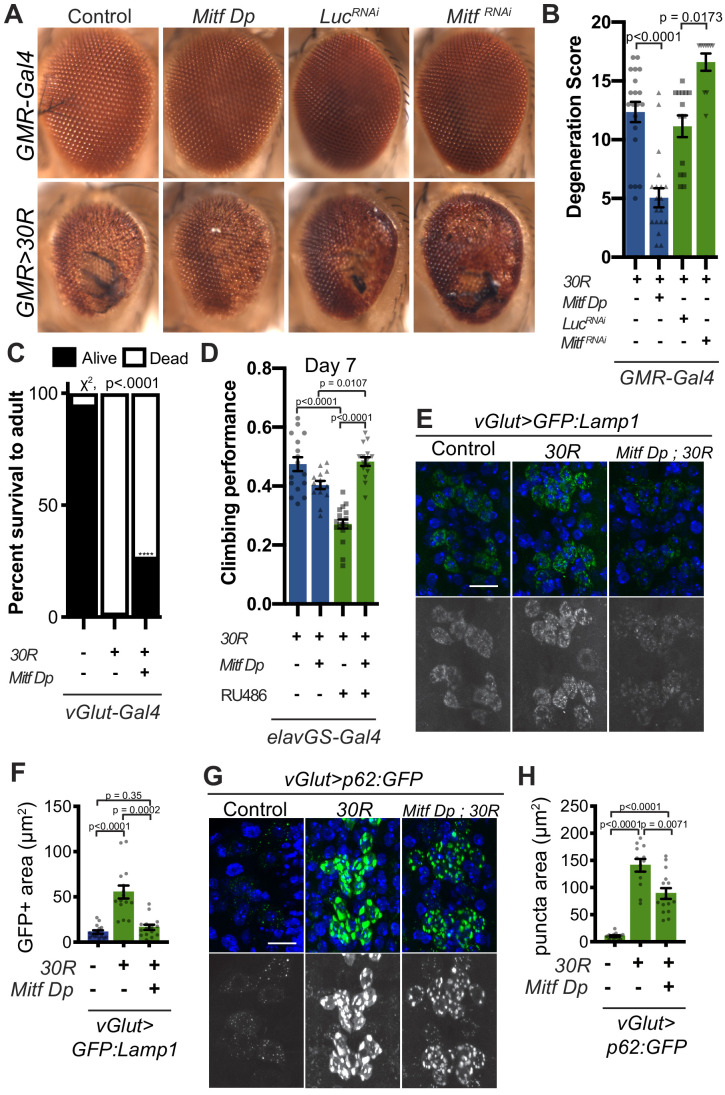
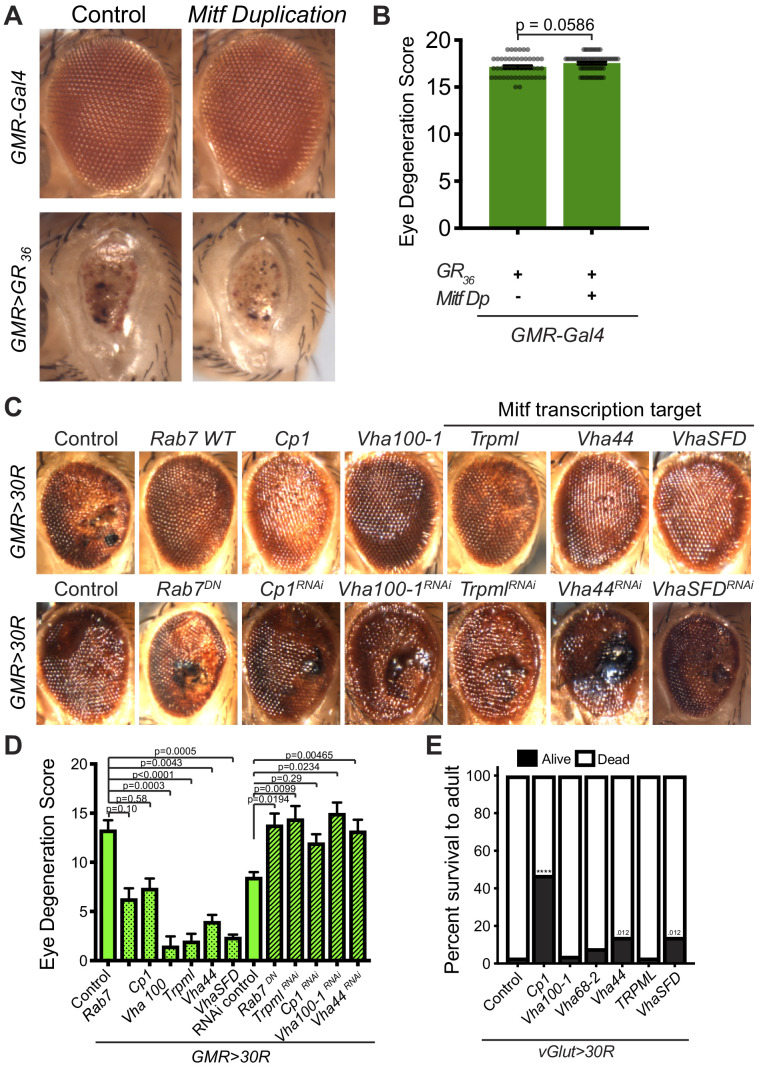
Since Mitf mislocalization contributes to autophagolysosome defects in a fly C9-ALS model, we hypothesized that increasing total levels of Mitf might compensate for impaired nuclear import. While high level Mitf overexpression is toxic in Drosophila (Hallsson et al., 2004), a genomic duplication construct containing the Mitf gene lacking the DNA repetitive intron 1 (Mitf Dp) (Zhang et al., 2015b), is sufficient to partially rescue 30R-mediated eye degeneration, while Mitf knockdown enhances eye degeneration (Figure 6A–B). Furthermore, pupal lethality caused by 30R expression in motor neurons and climbing impairment in elavGS >30R flies are also partially rescued by Mitf Dp (Figure 6C–D). In contrast, Mitf Dp did not rescue the severe rough eye phenotype observed with GMR-Gal4 overexpression of poly(GR)36 (Figure 6—figure supplement 1A–B), suggesting that Mitf Dp rescues toxicity caused by the G4C2 repeat RNA rather than the DPRs alone. To determine whether increased levels of Mitf rescue G4C2-mediated neurodegeneration through effects on the autophagolysosomal pathway, we examined GFP:Lamp and p62:GFP expression in 30R-expressing motor neurons. Indeed, Mitf Dp rescues increased GFP:Lamp1 expression (Figure 6E–F) and reduces p62:GFP accumulation in motor neurons of vGlut >30R larvae (Figure 6G–H). Thus, increasing Mitf levels in multiple neuronal subtypes in Drosophila suppresses G4C2-mediated neurotoxicity, consistent with our hypothesis that loss of nuclear Mitf is a key contributor to G4C2-mediated neurodegeneration.
Figure 6. Transcription factor Mitf/TFEB suppresses neurodegeneration caused by G4C2 expansion via lysosome activity.
(A) 15-day-old Drosophila eyes expressing UAS-30R under the control of GMR-Gal4, crossed to controls (w1118 or UAS-luciferase RNAi), genomic Mitf Duplication (Mitf Dp), or UAS-Mitf RNAi. (B) Quantification of external eye degeneration shown in A. Data are reported as mean ± SEM. Kruskal-Wallis test, p<0.0001, followed by Dunn’s multiple comparisons, n = 10–20 adults per genotype. (C) Percent of pupal eclosion in Drosophila expressing UAS-30R under the control of vGlut-Gal4 -/+ Mitf Dp compared to vGlut-Gal4/w1118 control. Fisher’s exact test, n = 133, 139, and 84 pupae, respectively. (D) Adult Drosophila expressing UAS-30R under the control of the inducible, pan-neuronal elavGS driver induced with 200 µM RU486 have decreased climbing ability at 7 days of age. Co-expressing Mitf Dp with UAS-30R rescues climbing ability. One-way ANOVA, p<0.0001, followed by Sidak’s multiple comparisons, n = 14–17 groups of 10 flies per genotype. (E) Representative images of motor neurons expressing UAS-GFP:Lamp1 for control (w1118), UAS-30R, or coexpressing Mitf Dp and UAS-30R. Scale bar = 10 µm (F) Quantification of the GFP positive (GFP+) area of GFP:Lamp1 in E. Data are reported as mean ± SEM. Brown-Forsythe and Welch ANOVA, p<0.0001, test followed by Dunnett’s T3 multiple comparisons, n = 15 per genotype. (G) Representative images of motor neurons coexpressing UAS-p62:GFP with no repeats (control, w1118), UAS-30R, and Mitf Dp with UAS-30R. Scale bar = 10 µm. (H) Quantification of p62:GFP GFP+ puncta area in F. Data are reported as mean ± SEM. Brown-Forsythe and Welch ANOVA test, p<0.0001, followed by Dunnett’s T3 multiple comparisons, n = 12–14 larvae per genotype.
Figure 6—figure supplement 1. Genetic increase of lysosome function rescues degeneration caused by G4C2 expression but not by poly-GR.
If the impaired lysosomal function we observe in our Drosophila model is contributing to neurodegeneration downstream of NCT defects, we would predict that genetic upregulation of key regulators of lysosome function may suppress degenerative phenotypes. Indeed, overexpression of Rab7, the small GTPase required for fusion of autophagosomes with lysosomes, or Trpml, a lysosomal calcium channel, suppress eye degeneration (Figure 6—figure supplement 1C–D). Furthermore, overexpression of key lysosomal v-ATPase subunits whose expression is regulated by Mitf also suppresses neurodegeneration in the Drosophila eye, while RNAi-mediated knockdown enhances degeneration (Figure 6—figure supplement 1C–D). Interestingly, loss of the ALS-associated gene ubqn in Drosophila was also rescued by increase in key lysosomal v-ATPase subunits or by nanoparticle mediated lysosome acidification (Şentürk et al., 2019). Overexpression of these Mitf-regulated genes also showed partial rescue of pupal lethality in animals expressing 30R in motor neurons (Figure 6—figure supplement 1E). These findings suggest a model whereby downregulation or cytoplasmic retention of Mitf targets leads to lysosomal disruption in G4C2-repeat-expressing flies.
Nuclear TFEB is reduced in human cells and motor cortex with GGGGCC repeat expansions
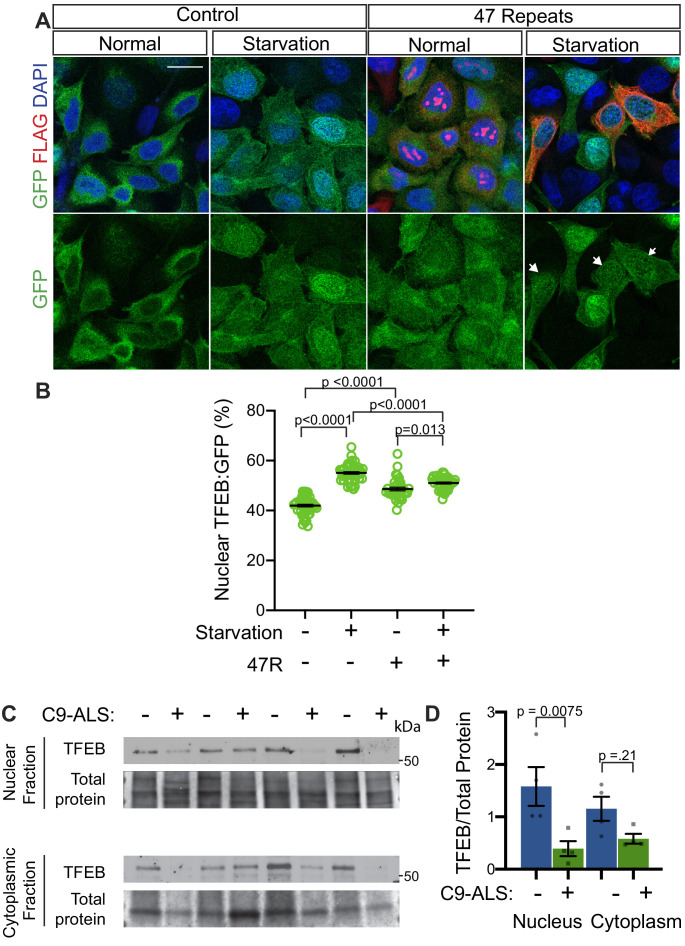
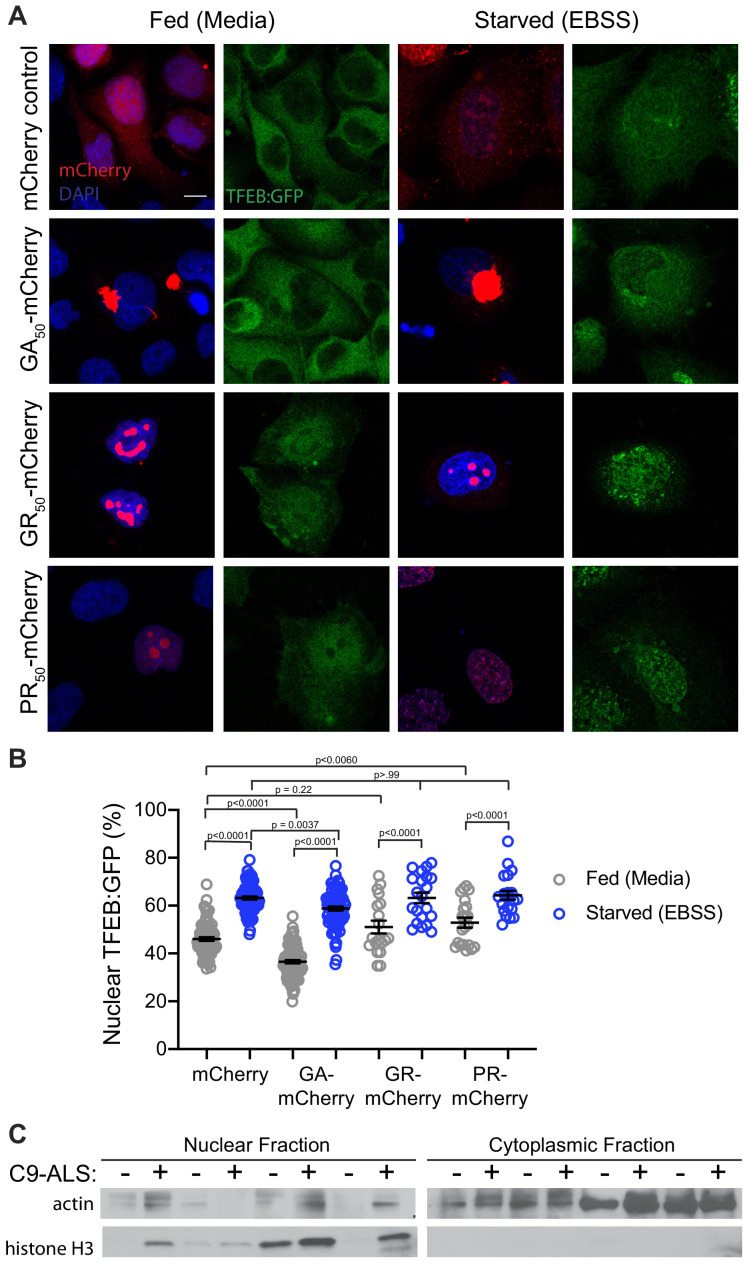
In humans, TFEB is the homolog of Drosophila Mitf that is best characterized for its role in autophagy and has been implicated in neurodegenerative disease (Cortes and La Spada, 2019; Martini-Stoica et al., 2016). Interestingly, a previous study showed nuclear TFEB was selectively depleted in the motor cortex of a sample of five ALS patients compared to five controls (Wang et al., 2016). To test the relevance of our findings in Drosophila models to human disease, we next examined whether G4C2 repeat expression impairs nuclear import of TFEB in HeLa cells stably expressing TFEB:GFP (Roczniak-Ferguson et al., 2012) using a 47-repeat (47R) G4C2 construct that expresses tagged DPRs (see Materials and methods). In control cells, TFEB:GFP is predominantly localized to the cytoplasm, whereas induction of autophagy by 3 hr starvation leads to robust nuclear translocation of TFEB (Figure 7A–B). In contrast, while 47R-expressing cells have a mild basal elevation of nuclear TFEB, the nuclear translocation of TFEB in response to starvation is significantly impaired relative to control cells (Figure 7A–B). We then tested the effect of expression of DPRs produced by alternate codons (i.e. in the absence of G4C2 repeats): poly-glycine-alanine (poly-GA50), poly-glycine-arginine (poly-GR50), and poly-proline-arginine (poly-PR50) (Figure 7—figure supplement 1A–B). While poly-GA50 causes a mild decrease in TFEB nuclear translocation, expression of poly-GR50 or poly-PR50 does not disrupt TFEB:GFP nuclear translocation. These data suggest that human cells expressing an expanded G4C2 repeat, but not DPRs, are unable to efficiently import TFEB into the nucleus in response to stimuli.
Figure 7. Nuclear TFEB is reduced in human cells expressing GGGGCC repeats and in C9-ALS human motor cortex.
(A) HeLa cells stably expressing TFEB:GFP transfected with 0R (Control) or a 47R construct (Flag tag in frame with poly-GR) in normal media (DMEM) or starved (3 hr in EBSS) conditions. White arrowheads indicate transfected cells in the 47R starved group. (B) Quantification of cells from A showing the percent (%) nuclear TFEB:GFP (nuclear/total) for each group. Data are presented as mean + SEM. One-way ANOVA, p<0.0001, with Sidak’s multiple comparisons, n = 47, 47, 35, and 38 cells. (C) Western blot for TFEB of human motor cortex samples fractionated into cytoplasmic and nuclear samples from postmortem control and C9-ALS patient brains. (D) Quantification of TFEB levels against total protein loading (Faststain) in control and C9-ALS patients. Data reported are mean ± SEM. One-way ANOVA, p=0.0142, with Sidak’s multiple comparisons, n = 4.
Figure 7—figure supplement 1. DPRs affect TFEB import in HeLa Cells.
To further investigate the relevance of loss of TFEB nuclear import to C9-ALS patients, we obtained human motor cortex samples from four non-neurological controls and four C9-ALS patients (Supplementary file 2). These samples were fractionated into cytoplasmic and nuclear-enriched fractions and assayed for TFEB using Western analysis. TFEB is reduced by an average of 76% in the nuclear fraction and by about 50% in the cytoplasm in C9-ALS compared to controls (Figure 7C–D, Figure 7—figure supplement 1C). These data suggest that TFEB protein is downregulated in C9-ALS/FTD motor cortex, but the greatest depletion occurs in the nucleus. Therefore, we propose a model whereby disruption of protein nuclear import by the C9orf72-HRE results in a failure of Mitf/TFEB to translocate to the nucleus to regulate the autophagic response to protein stress (Figure 8).
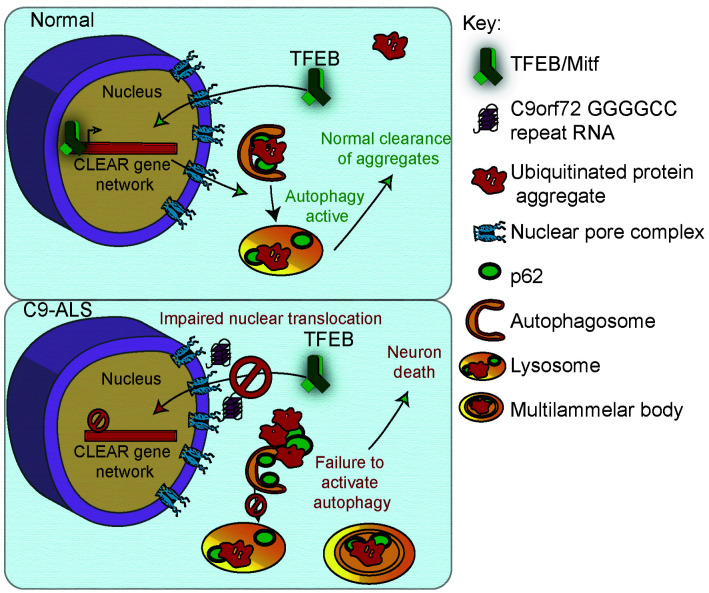
Figure 8. A proposed model of GGGGCC repeat expansion pathogenesis.
G4C2 repeat expansion causes nucleocytoplasmic transport disruption through multiple proposed mechanisms including G4C2 RNA binding of RanGAP and stress granule recruitment of nucleocytoplasmic transport machinery. Transport disruption leads to a blockage in the translocation of autophagy-mediating transcription factors such as Mitf/TFEB to the nucleus in response to proteotoxic stress. Failure to induce autophagic flux leads to autophagy pathway disruption such as the accumulation of large, non-degradative lysosomes and MLBs. Loss of autophagic flux leads to accumulation of Ref(2)P/ p62 and ubiquitinated protein aggregates, leading to chronic protein stress signaling and eventually neuronal cell death.
Discussion
Our work has revealed that the ALS-associated G4C2 hexanucleotide repeat is sufficient to disrupt multiple aspects of autophagy. In Drosophila, G4C2 repeats cause loss of autophagosomes and disrupt lysosomal structure and function. This accumulation of autolysosomes and lysosome-related organelles (MLBs) has been observed in lysosomal storage disorders and has been reported in spinal cord tissue from sporadic ALS patients (Bharadwaj et al., 2016; Parkinson-Lawrence et al., 2010; Sasaki, 2011). Regulation of protein and lipid homeostasis by the lysosome may be particularly important in neurons since they are post-mitotic and have high energy demands (Fraldi et al., 2016). Loss of function of C9orf72 also disrupts autophagy and lysosomal function in multiple cell types (Farg et al., 2014; Ji et al., 2017; O'Rourke et al., 2015; Sellier et al., 2016; Shi et al., 2018; Sullivan et al., 2016; Ugolino et al., 2016; Webster et al., 2016; Yang et al., 2016; Zhu et al., 2020), suggesting a mechanism whereby G4C2 repeats may have synergistically detrimental effects with haploinsufficient C9orf72 in C9-ALS/FTD patients. Additionally, multiple forms of familial ALS are caused by mutations in genes in autophagy and lysosome function (Evans and Holzbaur, 2019; Lin et al., 2017; Ramesh and Pandey, 2017), and upregulation of lysosome function has been proposed to be beneficial in multiple preclinical models of ALS (Donde et al., 2020; Mao et al., 2019; Şentürk et al., 2019; Shi et al., 2018). Thus, our findings suggest that, as has been shown in other forms of ALS, neurotoxicity of G4C2 repeats in C9 ALS-FTD is at least partially caused by disrupted autophagolysosomal function.
The finding that ref(2)P knockdown prevents or delays G4C2-mediated neurodegeneration is surprising, as p62/SQSTM1 is thought to link toxic ubiquitinated aggregates to LC3 to remove aggregates via selective autophagy (Cipolat Mis et al., 2016; Levine and Kroemer, 2008; Saitoh et al., 2015). However, other studies have also suggested that p62 may contribute to (rather than ameliorate) toxicity of ubiquitinated proteins. For example, Atg7-/- mice display severe defects in autophagy and accumulation of p62-positive protein aggregates in the liver and brain, and knockout of p62 in these mice prevents the formation of ubiquitinated aggregates and rescues liver dysfunction via suppression of chronic oxidative stress signaling (Komatsu et al., 2007). Additionally, Ataxia Telangiectasia Mutated-mediated DNA double stranded break repair is impaired in cultured neurons expressing the C9orf72-HRE, and this phenotype is rescued by p62 knockdown (Walker et al., 2017). These findings suggest that increases in p62 may contribute to DNA damage previously described in C9-ALS. Further, p62 is found to co-localize with DPRs in C9-ALS patients (Al-Sarraj et al., 2011; Mackenzie et al., 2014; Mori et al., 2013) and may promote protein aggregation. We hypothesize that p62-positive aggregate or oligomer formation in C9-patients contributes to neurotoxicity by activating downstream signaling pathways that are alleviated by autophagy-mediated clearance.
While many groups have reported nucleocytoplasmic transport dysfunction in ALS, it has remained unclear how NCT disruption causes ALS. Stress granules can recruit nuclear pore proteins to the cytoplasm and cause nucleocytoplasmic transport defects, suggesting that the disruptions in phase separation of RNA-binding proteins may lie upstream of nucleocytoplasmic transport defects (Zhang et al., 2018). Recently, Ortega et al. discovered that hyperactivity of nonsense-mediated decay may lie downstream of nucleocytoplasmic transport, indicating that multiple proteostasis pathways may be disrupted (Ortega et al., 2020). Additionally, selective autophagy is required for nuclear pore turnover (Lee et al., 2020), implying that autophagy defects may contribute to the cytoplasmic nuclear pore pathology found in C9-ALS patients and animal models. Our data show that in Drosophila, HeLa cells, and human tissue, nucleocytoplasmic transport defects lead to an inability to activate TFEB translocation to the nucleus, causing widespread autophagy defects and accumulation of protein aggregates (Figure 8). Interestingly, genetic inhibition of nuclear export or increase in Mitf expression are able to strongly rescue autophagosome and lysosome phenotypes and neurodegeneration, but do not result in complete clearance of p62 accumulations (Figures 5–6). Additional studies will be needed to better understand the relationship between p62 accumulation, autophagy, nucleocytoplasmic transport, and neurodegeneration. Overall, these findings place nucleocytoplasmic transport defects in ALS upstream of proteostasis defects.
Importantly, TFEB has been previously proposed as a therapeutic target in ALS and other neurodegenerative disease (Cortes and La Spada, 2019). Upregulation of TFEB signaling helps clear multiple types of proteotoxic aggregates found in Alzheimer’s disease, Parkinson’s disease, Huntington’s disease, ALS and FTD (Decressac et al., 2013; Parr et al., 2012; Polito et al., 2014; Torra et al., 2018; Vodicka et al., 2016). Our study suggests that modulation of TFEB nucleocytoplasmic transport may be an additional therapeutic target, and that targeting both nucleocytoplasmic transport and autophagy may act synergistically in ALS and FTD.
Materials and methods
Drosophila genetics
Drosophila were raised on standard cornmeal-molasses food at 25°C. For eye degeneration, GMR-GAL4, UAS-30R/CyO, twi-GAL4, UAS-GFP were crossed to UAS-modifier lines or background controls and GMR-GAL4, UAS-30R/UAS-modifier or GMR-GAL4, UAS-30R/+ were selected (where UAS-modifier can be on any chromosome) from the offspring and aged at 25°C for 15 days. Eye degeneration is quantified using a previously described method (Ritson et al., 2010). Briefly, points were added if there was complete loss of interommatidial bristles, necrotic patches, retinal collapse, loss of ommatidial structure, and/or depigmentation of the eye. Eye images were obtained using a Nikon SMZ 1500 Microscope and Infinity 3 Luminera Camera with Image Pro Insight 9.1 software.
For pupal survival assay, either three males from vGlut-Gal4 or vGlut-Gal4; UAS-30R/TM6G80(Tb) were crossed to 5–6 female flies containing UAS-modifier lines or background controls. Parental adult crosses were transferred to fresh vials every 2–3 days. After 15 days, non-tubby pupated flies that were (either vGlut-Gal4/UAS-modifier, vGlut/+; UAS-30R, or vGlut-Gal4/UAS-modifier; UAS-30R) were scored as either eclosed (empty pupal case) or non-eclosed (typically a fully developed pharate adult fly unable to eclose from pupal case due to paralysis).
For the climbing assay, UAS-30R; elavGS were crossed to experimental or genetic background controls. Adults were transferred 3–5 days after eclosion to vials containing 200 μM RU486 food or ethanol vehicle alone and transferred to new vials every 2–3 days. After aging 7–10 days, groups of 10 flies were placed into empty food vials and were tapped to the bottom and then locomotor function assessed by their negative geotaxis (flies reflexively crawl against gravity) response as measured by ability to climb 8 cm in 10 seconds. Each cohort of 10 flies was tested 10 times to obtain an average. N represents individual cohorts of 10 flies.
Drosophila drug feeding
Cornmeal-molasses-yeast fly food was melted and then cooled for 5 min before being mixed with concentrations of mifepristone (RU486), rapamycin, or trehalose and cooled to room temperature. Ethanol or DMSO was used as a vehicle control. Parent flies were crossed on normal food, and then they were transferred to food containing drug every 2–3 days such that their offspring would develop in food containing drug or adult offspring were transferred to drug food once eclosed as noted. Wandering third-instar larvae were selected for immunostaining or western blot analysis. Adult flies were aged on the drug-containing food for 15 days before analyzing their eye morphology or assessed for climbing ability on the day noted.
Quantitative RT–PCR
For each genotype, mRNA was collected from 5 flies or 30 heads using the TRIzol reagent following the manufacturer’s protocol. Reverse transcription was performed using SuperScript III First-Strand synthesis kit following the manufacturer’s protocol. Quantitative PCR was performed using SYBR Green PCR system on a 7900HT fast Real-Time PCR system (Applied Biosystem). The primers for G4C2 repeats were designed to amplify a 3’ region immediately after the repeats in the UAS construct.
Immunofluorescence staining and imaging
For Drosophila ventral nerve cords, wandering third-instar larvae were dissected in HL3 (Stewart et al., 1994) using a standard larval fillet dissection then fixed in 4% paraformaldehyde (or Bouin's fixative for UAS-mCherry:Atg8 experiments) (Sigma) for 20 min, followed by wash and penetration with PBS 0.1% Triton X-100 (PBX) for 3 × 20 min washes. The tissues were blocked for 1 hr at room temperature in PBS with 5% normal goat serum (NGS) and 0.1% PBX, then stained with primary antibodies at 4C overnight (16 hr). Tissues were washed three times for 20 min each with 0.1% PBX. Secondary antibodies (Goat antibodies conjugated to Alexa Fluor 568, 488, 633) diluted in 0.1% PBX 5% NGS and incubated for 2 hr and then washed three times for 20 min each with 0.1% PBX. During one wash, DAPI was added to the prep at a final concentration of 1 µg/mL. Larvae were mounted in Fluoromount-G (Invitrogen).
Drosophila salivary glands were dissected using a standard protocol and stained as above excepting for stronger solubilization with 0.3% PBX. Fixed cells or tissues were analyzed under an LSM780 or LSM800 confocal microscope (Carl Zeiss) with their accompanying software using Plan Apochromat 63 ×, NA 1.4 DIC or Plan Apochromat 40×, 1.3 Oil DIC objectives (Carl Zeiss) at room temperature. Images were captured by an AxioCam HRc camera (Carl Zeiss) and were processed using ImageJ/Fiji. To quantify fluorescent intensities, after opening the images in ImageJ/Fiji, certain areas/bands were circled and the intensities were measured. Puncta were counted using the Analyze Particles function in Image J using the same thresholding across experiments. Images are representative and experiments were repeated two to five times.
Western blotting
Tissues or cells were homogenized and/or lysed in RIPA buffer (50 mM Tris-HCl pH 7.4, 150 mM NaCl, 0.1% SDS, 0.5% sodium deoxycholate, and 1% Triton X-100) supplemented with protease inhibitor cocktail (Complete, Roche) using microcentrifuge pestles, and then were incubated in RIPA buffer on ice for 20 min. Samples were spun down at 100 g for 5 min to remove carcass and unbroken cells. For protein quantification, solution was diluted and measured by BCA assay (Thermo Fischer Scientific).
For nucleocytoplasmic fractionation of autopsy tissue, fractionation was performed with the NE-PER Nuclear and Cytoplasmic Extraction Kit according to the manufacturer’s protocol. For detection of proteins in the whole fraction, Drosophila larvae were solubilized in 8M urea. For the soluble and pelleted fraction, larvae were first solubilized in RIPA buffer as described above. The samples were spun down at 15000 rpm for 20 min and the soluble supernatant was set aside. Freshly prepared 8M urea buffer (Sigma) was added to the pellet and dissolved through vortexing. Samples were spun again at 15000 rpm for 20 min and urea-soluble pellet fraction was collected. A small amount of sample buffer dye was added and urea-buffered protein samples were run immediately on SDS-PAGE without heating. For immunoblot, 10–50 µg of total protein sample was mixed with 4x Laemmli buffer (Bio-Rad) and heated at 98°C for 10 min. The protein samples were run on 4–15% SDS Mini-PROTEAN TGX Precast Gels (Bio-Rad) and transferred to nitrocellulose membrane. TBST (50 mM Tris-HCl pH 7.4, 1% Triton X-100) with 5% non-fat milk (Bio-Rad) was used for blocking.
Electroretinogram (ERG) Assay
For ERG recordings, Rh1-GAL4/UAS-LacZ and Rh1-GAL4/UAS-30R flies were aged at 25°C in 12 hr light/12 hr dark cycle. ERG recordings were performed as described (Şentürk et al., 2019). In brief, adult flies were immobilized on a glass slide by glue. A reference electrode was inserted in the thorax and a recording electrode was placed on the eye surface. Flies were maintained in the darkness for at least 2 min prior to 1 s flashes of white light pulses (LED source with daylight filter), during which retinal responses were recorded and analyzed using WinWCP (University of Strathclyde, Glasgow, Scotland) software. At least five flies were examined for each genotype and timepoint.
Transmission Electron Microscopy (TEM)
Rh1-GAL4/UAS-lacZ and Rh1-GAL4/UAS-30R flies were aged at 25°C in 12 hr light/12 hr dark cycle. Retinae of adult flies were processed for TEM imaging as previously described (Chouhan et al., 2016). Three flies were examined for each genotype and timepoint.
Plasmids Source and Construction
pSF-CAG-Amp (0G504) was purchased from Oxford Genetics. We generated a mammalian expression plasmid pSF-(G4C2)47-VFH (V5-Flag-His), which can express 47 G4C2 repeats with three different tags to monitor expression of DPRs (polyGP-V5, polyGA-His, and polyGR-Flag). pEGFP-(GA,GR, or PR)50 was obtained from Davide Trotti (Wen et al., 2014), and the GFP cDNA sequence was replaced with mCherry by digesting with BamHI and XhoI.
TFEB:GFP HeLa cell culture, transfection, and immunofluorescence analysis
HeLa cell line with stable expressing TFEB:GFP was a gift from Dr. Shawn Ferguson at Yale University. Hela cells were grown in DMEM media (Invitrogen) supplemented with 10% fetal bovine serum (Hyclone Laboratories Inc). The cell line was authenticated by observing nuclear translocation of TFEB:GFP in the presence of starvation (Figure 7). Absence of mycoplasma contamination was confirmed by staining with DAPI. Transfection was performed using Lipofectamine 2000 (Invitrogen) according to the manufacturer’s instructions. Briefly, 1–2 µg of cDNA was diluted into 100 µl of Opti-MEM I Medium (Invitrogen) and mixed gently. Lipofectamine 2000 mixture was prepared by diluting 2–4 µl of Lipofectamine 2000 in 100 µl of Opti-MEM I Medium. The ratio of DNA to Lipofectamine 2000 used for transfection was 1∶two as indicated in the manual. The DNA-Lipofectamine 2000 mixture was mixed gently and incubated for 20 min at room temperature. Cells were directly added to the 200 µl of DNA-Lipofectamine 2000 mixture. After 48 hr, transfected HeLa cells were treated with EBSS medium for 3 hr for starvation. HeLa cells were fixed with 4% PFA at room temperature for 15 min, washed three times with PBS, permeabilized for 10 min with 1% PBTX, washed another three times with PBS, and blocked for 1 hr at room temperature with 10% normal goat serum (Sigma) diluted in 0.1% PBTX. Cells were then incubated overnight at 4°C with primary antibody mouse anti-Flag antibody. After three washes in PBS (5 min each), cells were incubated for 1 hr at room temperature with secondary antibodies (goat anti-Alexa Fluor 568) diluted in the blocking solution. Cells were washed three times in PBS and mounted with Prolong Gold anti-fade reagent with DAPI (Cell Signaling).
Collection of human autopsied tissue
Human autopsied tissue used for these data are described in detail in Supplementary file 2. The use of human tissue and associated decedents’ demographic information was approved by the Johns Hopkins University Institutional Review Board and ethics committee (HIPAA Form five exemption, Application 11-02-10-01RD) and from the Ravitz Laboratory (UCSD) through the Target ALS Consortium.
Statistics
All quantitative data were derived from independent experiments. Each n value representing biological replicates is indicated in the figure legends. Statistical tests were performed in Prism version 8.3.1 or Microsoft Excel 16.34 and were performed as marked in the figure legends. All statistical tests were two-sided. Results were deemed significant when the P value = 0.05. No statistical methods were used to predetermine sample size. The investigators were not blinded during experiments.
Acknowledgements
This work was supported by NINDS R01NS082563 (TEL), R01NS094239 (TEL and JDR), F31 NS100401 (KMC), ALSA (TEL, KZ and JDR), and Target ALS (TEL, KZ, JDR, and HJB). KMC is a recipient of the PEO Scholar Award. HJB is an Investigator of the Howard Hughes Medical Institute. We thank Francesca Pignoni, Udai Pandey, Peng Jin, Adrian Isaacs, Eric Baehrecke, Helmut Kramer, Francesca Pignoni, Gábor Juhász, Patrick Dolph, L Miguel Martins, the Bloomington Drosophila Stock Center (NIH P40ODO18537) and Vienna Drosophila Research Center for Drosophila lines and/or antibodies and Shawn Ferguson and Davide Trotti for cell lines and constructs. The Johns Hopkins NINDS Multiphoton Imaging Core (NS050274) provided imaging equipment and expertise.
Appendix 1
Appendix 1—key resources table.
| Reagent type (species) or resource |
Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (D. melanogaster) | GMR-Gal4 | Bloomington Drosophila Stock Center | BDSC:1104 | w*; P{GAL4-ninaE.GMR}12 |
| Genetic reagent (D. melanogaster) | 30R | Peng Jin (Xu et al., 2013) | FlyBase: FBal0294759 |
w[1118];UAS-(G4C2)30 |
| Genetic reagent (D. melanogaster) | TRiP background control | Bloomington Drosophila Stock Center | BDSC: 36303 | y[1] v[1]; P{y[+t7.7]=CaryP}attP2 |
| Genetic reagent (D. melanogaster) | UAS-ref(2)PRNAi#1 | Bloomington Drosophila Stock Center | BDSC: 36111 | y[1] sc[*] v[1] sev[21]; P{y[+t7.7] v[+t1.8]=TRiP.HMS00551}attP2 |
| Genetic reagent (D. melanogaster) | UAS-ref(2)PRNAi #2 | Bloomington Drosophila Stock Center | BDSC: 33978 | y[1] sc[*] v[1] sev[21]; P{y[+t7.7] v[+t1.8]=TRiP.HMS00938}attP2 |
| Genetic reagent (D. melanogaster) | UAS-ref(2)P-HA | L.M. Martins (de Castro et al., 2013) | Flybase: FBtp0089618 | |
| Genetic reagent (D. melanogaster) | vGlut-Gal4 | Bloomington Drosophila Stock Center | Flybase: FBal0194519 | w[1118]; P{w[+mW.hs]=GawB}VGlut[OK371] |
| Genetic reagent (D. melanogaster) | elavGS | Adrian Isaacs | Flybase: FBtp0015149 | w[*]; P{elav-Switch.O} GSG301 |
| Genetic reagent (D. melanogaster) | UAS-poly(GR)36 | Adrian Isaacs (Mizielinska et al., 2014) | BDSC: 58692 | w[1118]; P{{y[+t7.7] w[+mC]=UAS poly-GR.PO-36}attP40 |
| Genetic reagent (D. melanogaster) | Act-Gal4 | Bloomington Drosophila Stock Center | Flybase: FBti0183703 | y[1] w[*]; P{Act5C-GAL4}17bFO1/TM6B, Tb1 |
| Genetic reagent (D. melanogaster) | UAS-ref(2)P: GFP | Thomas Neufeld (Chang and Neufeld, 2009) | Flybase: FBtp0041098 | |
| Genetic reagent (D. melanogaster) | UAS-mCherry-Atg8 | Bloomington Drosophila Stock Center | BDSC: 37749 | y[1] w[1118]; P{w[+mC]=UASp-GFP-mCherry-Atg8a}2 |
| Genetic reagent (D. melanogaster) | UAS-GFP:Lamp1 | Helmut Kramer (Pulipparacharuvil et al., 2005) | Flybase: FBtp0041063 | w[*]; P{w[+mC]=UAS-GFP-LAMP}2 |
| Genetic reagent (D. melanogaster) | UAS-3R; UAS-(G4C2)3 | Adrian Isaacs (Mizielinska et al., 2014) | BDSC: 58687 | w[1118]; P{{y[+t7.7] w[+mC]=UAS GGGGCC.3}attP40 |
| Genetic reagent (D. melanogaster) | UAS-36R; UAS-(G4C2)36 | Adrian Isaacs (Mizielinska et al., 2014) | BDSC: 58688 | w[1118]; P{{y[+t7.7] w[+mC]=UAS GGGGCC.36}attP40 |
| Genetic reagent (D. melanogaster) | UAS-44R; UAS-LDS(G4C2)44 | Nancy Bonini (Goodman et al., 2019b) |
BDSC: 84723 | w[1118]; P{w[+mC]=UAS-LDS-(G4C2)44.GR-GFP}9 |
| Genetic reagent (D. melanogaster) | UAS-LacZ | Bloomington Drosophila Stock Center | BDSC: 3956 | w[1118]; P{w[+mC]=UAS-lacZ.NZ}J312 |
| Genetic reagent (D. melanogaster) | Rh1-Gal4 | Bloomington Drosophila Stock Center | BDSC: 8961 | P{ry[+t7.2]=rh1 GAL4}3, ry[506] |
| Genetic reagent (D. melanogaster) | gRab7-YFP | Bloomington Drosophila Stock Center | BDSC: 62545 | w[1118]; TI{TI}Rab7[EYFP] |
| Genetic reagent (D. melanogaster) | UAS-Rab7-GFP | Bloomington Drosophila Stock Center | BDSC: 42706 | |
| Genetic reagent (D. melanogaster) | UAS-luciferaseRNAi | Bloomington Drosophila Stock Center | BDSC: 31603 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.JF01355}attP2 |
| Genetic reagent (D. melanogaster) | UAS-S-GFP; | Bloomington Drosophila Stock Center | BDSC: 7032 | w[1118]; P{w[+mC]=UAS-NLS-NES[+]-GFP}5A |
| Genetic reagent (D. melanogaster) | UAS-RanGAP | Zhang et al., 2015a | ||
| Genetic reagent (D. melanogaster) | UAS-RanGAPRNAi; | Bloomington Drosophila Stock Center | BDSC: 29565 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.JF03244}attP2/TM3, Sb[1] |
| Genetic reagent (D. melanogaster) | UAS-CD8:GFP | Bloomington Drosophila Stock Center | Flybase: FBti0012685 | y[1] w[*]; P{w[+mC]=UAS-mCD8::GFP.L}LL5 |
| Genetic reagent (D. melanogaster) | UAS-Mitf-HA | Francesca Pignoni (Zhang et al., 2015b) |
||
| Genetic reagent (D. melanogaster) | daGS | Bloomington Drosophila Stock Center | Flybase: FBtp0057039 |
w[*]; P{w[+mC]=da-GSGAL4.T} |
| Genetic reagent (D. melanogaster) | UAS-embargoed RNAi | Bloomington Drosophila Stock Center | BDSC: 31353 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.JF01311}attP2 |
| Genetic reagent (D. melanogaster) | Mitf duplication; MitfSI-RES | Francesca Pignoni (Zhang et al., 2015b) | Flybase: FBtp0115483 | |
| Genetic reagent (D. melanogaster) | UAS-Mitf RNAi | Bloomington Drosophila Stock Center | BDSC: 43998 | y[1] sc[*] v[1] sev[21]; P{y[+t7.7] v[+t1.8]=TRiP.HMS02712}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Rab7WT | Bloomington Drosophila Stock Center | BDSC: 23641 | y[1] w[*]; P{w[+mC]=UASp YFP.Rab7}21/SM5 |
| Genetic reagent (D. melanogaster) | UAS-Cp1EP | Bloomington Drosophila Stock Center | BDSC: 15957 | y[1] w[67c23]; P{w[+mC] y[+mDint2]=EPgy2}Cp1[EY05806] |
| Genetic reagent (D. melanogaster) | UAS-Vha100-1EP | Bloomington Drosophila Stock Center | BDSC: 63269 | w[1118]; P{w[+mC]=EP}Vha100-1[G4514]/TM6C, Sb[1] |
| Genetic reagent (D. melanogaster) | UAS-Trpml | Kartik Venkatachalam | Flybase: FBti0162438 | |
| Genetic reagent (D. melanogaster) | UAS-Vha44EP | Bloomington Drosophila Stock Center | BDSC: 20140 | y[1] w[67c23]; P{w[+mC] y[+mDint2]=EPgy2}Vha44[EY02202] |
| Genetic reagent (D. melanogaster) | UAS-VhaSFDEP | Bloomington Drosophila Stock Center | BDSC: 15758 | y[1] w[67c23]; P{w[+mC] y[+mDint2]=EPgy2}VhaSFD[EY04644]/CyO |
| Genetic reagent (D. melanogaster) | UAS-Rab7DN | Bloomington Drosophila Stock Center | BDSC: 9778 | y[1] w[*]; P{w[+mC]=UASp YFP.Rab7.T22N}06 |
| Genetic reagent (D. melanogaster) | UAS-Cp1RNAi | Bloomington Drosophila Stock Center | BDSC: 32932 | y[1] sc[*] v[1] sev[21]; P{y[+t7.7] v[+t1.8]=TRiP.HMS00725}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Vha100-1RNAi | Bloomington Drosophila Stock Center | BDSC: 26290 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.JF02059}attP2 |
| Genetic reagent (D. melanogaster) | UAS-TrpmlRNAi | Bloomington Drosophila Stock Center | BDSC: 31294 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.JF01239}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Vha44RNAi | Bloomington Drosophila Stock Center | BDSC: 33884 | y[1] sc[*] v[1] sev[21]; P{y[+t7.7] v[+t1.8]=TRiP.HMS00821}attP2 |
| Genetic reagent (D. melanogaster) | UAS-VhaSFDRNAi | Bloomington Drosophila Stock Center | BDSC: 40896 | y[1] sc[*] v[1] sev[21]; P{y[+t7.7] v[+t1.8]=TRiP.HMS02144}attP40 |
| Genetic reagent (D. melanogaster) | UAS-Atg6RNAi | Bloomington Drosophila Stock Center | BDSC: 35741 | y[1] sc[*] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMS01483}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Atg18aRNAi | Bloomington Drosophila Stock Center | BDSC: 34714 | y[1] sc[*] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMS01193}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Atg1 | Bloomington Drosophila Stock Center | BDSC: 51655 | y[1] w[*]; P{w[+mC]=UAS-Atg1.S}6B |
| Genetic reagent (D. melanogaster) | UAS-Atg7 | Bloomington Drosophila Stock Center | NA | w[1118]; P{w[+mC]=UAS-Atg7} |
| Genetic reagent (D. melanogaster) | UAS-Atg101RNAi | Bloomington Drosophila Stock Center | BDSC: 34360 | y[1] sc[*] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMS01349}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Atg8aRNAi | Bloomington Drosophila Stock Center | BDSC: 34340 | y[1] sc[*] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMS01328}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Atg5RNAi | Bloomington Drosophila Stock Center | BDSC: 27551 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.JF02703}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Atg5RNAi | Bloomington Drosophila Stock Center | BDSC: 34899 | y[1] sc[*] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMS01244}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Atg8aRNAi | Bloomington Drosophila Stock Center | BDSC: 28989 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.JF02895}attP2 e[*]/TM3, Sb[1] |
| Genetic reagent (D. melanogaster) | UAS-Atg14RNAi | Bloomington Drosophila Stock Center | BDSC: 55398 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMC04086}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Atg16RNAi | Bloomington Drosophila Stock Center | BDSC: 34358 | y[1] sc[*] v[1] sev[21]; P{y[+t7.7] v[+t1.8]=TRiP.HMS01347}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Atg16RNAi | Bloomington Drosophila Stock Center | BDSC: 58244 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMJ22265}attP40/CyO |
| Genetic reagent (D. melanogaster) | UAS-Atg17RNAi | Bloomington Drosophila Stock Center | BDSC: 36918 | y[1] sc[*] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMS01611}attP2/TM3, Sb[1] |
| Genetic reagent (D. melanogaster) | UAS-Atg6RNAi | Bloomington Drosophila Stock Center | BDSC: 28060 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.JF02897}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Atg8bRNAi | Bloomington Drosophila Stock Center | BDSC: 34900 | y[1] sc[*] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMS01245}attP2 |
| Genetic reagent (D. melanogaster) | UAS-bchsRNAi | Bloomington Drosophila Stock Center | BDSC: 42517 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMJ02083}attP40 |
| Genetic reagent (D. melanogaster) | UAS-Atg8bRNAi | Bloomington Drosophila Stock Center | BDSC: 27554 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.JF02706}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Atg9RNAi | Bloomington Drosophila Stock Center | BDSC: 28055 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.JF02891}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Atg18aRNAi | Bloomington Drosophila Stock Center | BDSC: 28061 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.JF02898}attP2 |
| Genetic reagent (D. melanogaster) | UAS-GyfRNAi | Bloomington Drosophila Stock Center | BDSC: 28896 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HM05106}attP2 |
| Genetic reagent (D. melanogaster) | Atg600096 | Bloomington Drosophila Stock Center | BDSC: 11487 | ry[506] P{ry[+t7.2]=PZ}Atg6[00096]/TM3, ry[RK] Sb[1] Ser[1] |
| Genetic reagent (D. melanogaster) | UAS-Atg4bRNAi | Bloomington Drosophila Stock Center | BDSC: 56046 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMS04249}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Atg17EP | Bloomington Drosophila Stock Center | BDSC: 15618 | y[1] w[67c23]; P{w[+mC] y[+mDint2]=EPgy2}Atg17[EY03045] |
| Genetic reagent (D. melanogaster) | bchs58 | Bloomington Drosophila Stock Center | BDSC: 9887 | y[1] w[*]; P{w[+mC]=EP}EP2299, bchs[58]/CyO |
| Genetic reagent (D. melanogaster) | UAS-Atg4aRNAi | Bloomington Drosophila Stock Center | BDSC: 35740 | y[1] sc[*] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMS01482}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Atg4aRNAi | Bloomington Drosophila Stock Center | BDSC: 44421 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.GLC01355}attP40 |
| Genetic reagent (D. melanogaster) | UAS-Atg10RNAi | Bloomington Drosophila Stock Center | BDSC: 40859 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMS02026}attP40 |
| Genetic reagent (D. melanogaster) | UAS-Atg16RNAi | Bloomington Drosophila Stock Center | BDSC: 34358 | y[1] sc[*] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMS01347}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Atg9RNAi | Bloomington Drosophila Stock Center | BDSC: 34901 | y[1] sc[*] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMS01246}attP2 |
| Genetic reagent (D. melanogaster) | UAS-ltRNAi | Bloomington Drosophila Stock Center | BDSC: 34871 | y[1] sc[*] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMS00190}attP2/TM3, Sb[1] |
| Genetic reagent (D. melanogaster) | UAS-Atg7RNAi | Bloomington Drosophila Stock Center | BDSC: 34369 | y[1] sc[*] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMS01358}attP2/TM3, Sb[1] |
| Genetic reagent (D. melanogaster) | Atg7d06996 | Bloomington Drosophila Stock Center | BDSC: 19257 | w[1118]; P{w[+mC]=XP}Atg7[d06996]/CyO |
| Genetic reagent (D. melanogaster) | UAS-Atg4aRNAi | Bloomington Drosophila Stock Center | BDSC: 28367 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.JF03003}attP2 |
| genetic reagent (D. melanogaster) | UAS-Atg8aRNAi | Bloomington Drosophila Stock Center | BDSC: 58309 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMJ22416}attP40 |
| Genetic reagent (D. melanogaster) | Bchs17 | Bloomington Drosophila Stock Center | BDSC: 9888 | y[1] w[*]; P{w[+mC]=EP}EP2299, bchs[17]/CyO |
| Genetic reagent (D. melanogaster) | UAS-Atg8aEP | Bloomington Drosophila Stock Center | BDSC: 10107 | w[1118] P{w[+mC]=EP}Atg8a[EP362] |
| Genetic reagent (D. melanogaster) | UAS-Atg2EP | Bloomington Drosophila Stock Center | BDSC: 17156 | w[1118]; P{w[+mC]=EP}Atg2[EP3697]/TM6B, Tb[1] |
| Genetic reagent (D. melanogaster) | UAS-Atg7RNAi | Bloomington Drosophila Stock Center | BDSC: 34369 | y[1] sc[*] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMS01358}attP2/TM3, Sb[1] |
| Genetic reagent (D. melanogaster) | UAS-Atg13RNAi | Bloomington Drosophila Stock Center | BDSC: 40861 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMS02028}attP40 |
| Genetic reagent (D. melanogaster) | UAS-Atg14RNAi | Bloomington Drosophila Stock Center | BDSC: 40858 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMS02025}attP40/CyO |
| Genetic reagent (D. melanogaster) | UAS-Atg3EP | Bloomington Drosophila Stock Center | BDSC: 16429 | y[1] w[67c23]; P{w[+mC] y[+mDint2]=EPgy2}Atg3[EY08396] |
| Genetic reagent (D. melanogaster) | UAS-Atg18bRNAi | Bloomington Drosophila Stock Center | BDSC: 34715 | y[1] sc[*] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMS01194}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Atg2RNAi | Bloomington Drosophila Stock Center | BDSC: 27706 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.JF02786}attP2 |
| Genetic reagent (D. melanogaster) | Snap29B6-21 | Bloomington Drosophila Stock Center | BDSC: 56818 | w[*]; P{ry[+t7.2]=neoFRT}42D Snap29[B6-21]/CyO, P{w[+mC]=GAL4 twi.G}2.2, P{w[+mC]=UAS-2xEGFP}AH2.2 |
| Genetic reagent (D. melanogaster) | UAS-Atg3RNAi | Bloomington Drosophila Stock Center | BDSC: 34359 | y[1] sc[*] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMS01348}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Atg2RNAi | Bloomington Drosophila Stock Center | BDSC: 34719 | y[1] sc[*] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMS01198}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Atg18bRNAi | Bloomington Drosophila Stock Center | BDSC: 34715 | y[1] sc[*] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.HMS01194}attP2 |
| Genetic reagent (D. melanogaster) | Atg4bP0997 | Bloomington Drosophila Stock Center | BDSC: 36340 | y[1] w[*]; P{w[+mC]=lacW}Atg4b[P0997] |
| Genetic reagent (D. melanogaster) | UAS-bchs-HA | Bloomington Drosophila Stock Center | BDSC: 51636 | y[1] w[*]; P{w[+mC]=UAS bchs.HA}32 |
| Cell line (Homo sapiens) | HeLa stably expressing TFEB:GFP | Shawn Ferguson (Roczniak-Ferguson et al., 2012) | ||
| Biological sample (Homo sapiens) | Control (non-neurological) and ALS postmortem motor cortex tissue | Ravitz laboratory (UCSD) through Target ALS Consortium; Brain Resource Center at JHMI | ||
| Antibody | Rabbit polyclonal anti-dsRed | Clontech | Cat#63249, RRID:AB_10013483 | 1:1000 for IF |
| Antibody | Mouse monoclonal anti- poly-ubiquitin | Enzo Life Sciences | Cat#BML-PW8805, RRID:AB_10541434 | 1:200 for IF |
| Antibody | Rabbit polyclonal anti-ref(2)P | Gabor Juhasz laboratory (Pircs et al., 2012) | 1:1000 for IF; 1:1000 for WB | |
| Antibody | Guinea pig polyclonal anti-Mitf | Francesca Pignoni laboratory (Zhang et al., 2015b) | 1:500 for IF | |
| Antibody | Rat monoclonal anti-HA | Roche | Cat# 11867423001, RRID:AB_390918 | 1:200 for IF |
| Antibody | Chicken polyclonal anti-GFP | Abcam | Cat# ab13970, RRID:AB_300798 | 1:1000 for IF; 1:1000 for WB |
| Antibody | Guinea pig polyclonal anti-Cp1 | Patrick Dolph laboratory (Kinser and Dolph, 2012) |
1:2500 for WB | |
| Antibody | Mouse monoclonal anti- beta actin (clone C4) | EMD Millipore | Cat# MAB1501, RRID:AB_2223041 | 1:1000 for WB |
| Antibody | Rabbit polyclonal anti- TFEB | Bethyl Biosciences | Cat# A303-673A, RRID:AB_11204751 | 1:2000 for WB |
| Antibody | Rabbit polyclonal anti-Histone H3 | Cell Signaling | Cat# 9715, RRID:AB_331563 | 1:1000 for WB |
| Antibody | Mouse monoclonal anti-FLAG | Sigma Aldrich | Cat# F3165, RRID:AB_259529 | 1:1000 for IF |
| Recombinant DNA reagent | pSF-CAG-Amp | Oxford Genetics | Cat# 0G504 | |
| Sequence-based reagent | Actin forward | Integrated DNA Technologies | q-RT-PCR primer | 5’- GCGCGGTTACTCTTTCACCA-3’ |
| Sequence-based reagent | Actin reverse | Integrated DNA Technologies | q-RT-PCR primer | 5’- ATGTCACGGACGATTTCACG-3’ |
| Sequence-based reagent | G4C2 repeats forward (UAS-30R) | Integrated DNA Technologies | q-RT-PCR primer | 5’- GGGATCTAGCCACCATGGAG-3’ |
| Sequence-based reagent | G4C2 repeats reverse (UAS-30R) | Integrated DNA Technologies | q-RT-PCR primer | 5’- TACCGTCGACTGCAGAGATTC-3’ |
| Sequence-based reagent | Mitf forward | Integrated DNA Technologies | q-RT-PCR primer | 5’-AGTATCGGAGTAGATGTGCCAC-3’ |
| Sequence-based reagent | Mitf reverse | Integrated DNA Technologies | q-RT-PCR primer | 5’- CGCTGAGATATTGCCTCACTTG-3’ |
| Sequence-based reagent | Vha16-1 forward | Integrated DNA Technologies | q-RT-PCR primer | 5’- TCTATGGCCCCTTCTTCGGA-3’ |
| Sequence-based reagent | Vha16-1 reverse | Integrated DNA Technologies | q-RT-PCR primer | 5’- AATGGCAATGATACCCGCCA-3’ |
| Sequence-based reagent | Vha68-2 forward | Integrated DNA Technologies | q-RT-PCR primer | 5’- CAAATATGGACGTGTCTTCGCT-3’ |
| Sequence-based reagent | Vha68-2 reverse | Integrated DNA Technologies | q-RT-PCR primer | 5’- CCGGATCTCCGACAGTTACG-3’ |
| Sequence-based reagent | Vha55 forward | Integrated DNA Technologies | q-RT-PCR primer | 5’- CGGGACTTTATCTCCCAGCC-3’ |
| Sequence-based reagent | Vha55 reverse | Integrated DNA Technologies | q-RT-PCR primer | 5’-TGACCTCATCGAGAATGACCAG-3’ |
| Sequence-based reagent | Vha44 forward | Integrated DNA Technologies | q-RT-PCR primer | 5’- TGGACTCGGAGTACCTGACC-3’ |
| Sequence-based reagent | Vha44 reverse | Integrated DNA Technologies | q-RT-PCR primer | 5’- CGTCACGTTGAACAGGCAGTA-3’ |
| Sequence-based reagent | Vha100-2 forward | Integrated DNA Technologies | q-RT-PCR primer | 5’- TGTTCCGTAGTGAGGAGATGG-3’ |
| Sequence-based reagent | Vha100-2 reverse | Integrated DNA Technologies | q-RT-PCR primer | 5’- TCACGTTCACATTCAAGTCGC-3’ |
| Sequence-based reagent | Atg8a forward | Integrated DNA Technologies | q-RT-PCR primer | 5’- GGTCAGTTCTACTTCCTCATTCG-3’ |
| Sequence-based reagent | Atg8a reverse | Integrated DNA Technologies | q-RT-PCR primer | 5’- GATGTTCCTGGTACAGGGAGC-3’ |
| Sequence-based reagent | Atg9 forward | Integrated DNA Technologies | q-RT-PCR primer | 5’- ACACGCCTCGAAACAGTGG-3’ |
| Sequence-based reagent | Atg9 reverse | Integrated DNA Technologies | q-RT-PCR primer | 5’- TCAAGGTCCTCGATGTGGTTC-3’ |
| Sequence-based reagent | ref(2)P forward | Integrated DNA Technologies | q-RT-PCR primer | 5' - ATGCCGGAGAAGCTGTTGAA - 3' |
| Sequence-based reagent | ref(2)P reverse | Integrated DNA Technologies | q-RT-PCR primer | 5' - ATCAGCGTCGATCCAGAAGG - 3' |
| Commercial assay or kit | SuperScript III First-Strand Synthesis System | Thermo Fischer Scientific | Cat #18080051 | |
| Commercial assay or kit | NE-PER Nuclear and Cytoplasmic Extraction Kit | Thermo Fischer Scientific | Cat #78833 | |
| Commercial assay or kit | BCA Assay | Thermo Fischer Scientific | Cat #23227 | |
| Commercial assay or kit | 4–15% Mini-PROTEAN TGX Precast Gel | Bio-Rad | Cat #4561083 | |
| Commercial assay or kit | One Shot TOP10 Chemically Competent E. coli | Thermo Fischer Scientific | Cat# C404006 | |
| Commercial assay or kit | Faststain | G-Biosciences | Cat #786–34 | |
| Commercial assay or kit | SYBR Select Master Mix | Thermo Fischer Scientific | Cat #4472908 | |
| Chemical compound, drug | Blotting Grade Blocker (nonfat dry milk) | Bio-Rad | Cat #1706404 | |
| Chemical compound, drug | Lipofectamine 2000 | Thermo Fischer Scientific | Cat #11668019 | |
| Chemical compound, drug | Mifepristone (RU486) | Millipore Sigma | Cat #M8046 | |
| Chemical compound, drug | Rapamycin | Selleckchem | Cat #S1039 | |
| Chemical compound, drug | D-(+)-Trehalose dihydrate | Millipore Sigma | Cat #T0167 | |
| Chemical compound, drug | TRIzol | Thermo Fischer Scientific | Cat #15596018 | |
| Chemical compound, drug | Protease Inhibitor Cocktail | Roche | Cat#11873580001 | |
| Software, algorithm | ImageJ | https://imagej.nih.gov/ij/ | ||
| Software, algorithm | GraphPad Prism 8 | https://www.graphpad.com/scientific-software/prism/ | ||
| Software, algorithm | IMARIS 9 | https://imaris.oxinst.com/ | ||
| Software, algorithm | Adobe Illustrator CC 2018 | https://www.adobe.com/products/illustrator | ||
| Software, algorithm | Image Pro Insight 9.1 | http://www.mediacy.com/imagepro | ||
| Software, algorithm | WinWCP | https://pureportal.strath.ac.uk/en/datasets/strathclyde-electrophysiology-software-winwcp-winedr |
Funding Statement
The funders had no role in study design, data collection and interpretation, or the decision to submit the work for publication.
Contributor Information
Thomas E Lloyd, Email: tlloyd4@jhmi.edu.
K VijayRaghavan, National Centre for Biological Sciences, Tata Institute of Fundamental Research, India.
Harry T Orr, University of Minnesota, United States.
Funding Information
This paper was supported by the following grants:
National Institute of Neurological Disorders and Stroke R01NS082563 to Thomas E Lloyd.
Amyotrophic Lateral Sclerosis Association 17-IIP-370 to Thomas E Lloyd.
National Institutes of Health P40OD018537 to Hugo J Bellen.
Howard Hughes Medical Institute to Hugo J Bellen.
National Institute of Neurological Disorders and Stroke R01NS094239 to Jeffrey D Rothstein, Thomas E Lloyd.
National Institute of Neurological Disorders and Stroke P30NS050274 to Thomas E Lloyd.
National Institute of Neurological Disorders and Stroke F31NS100401 to Kathleen M Cunningham.
ALSA to Jeffrey D Rothstein, Ke Zhang, Thomas E Lloyd.
Additional information
Competing interests
Reviewing editor, eLife.
No competing interests declared.
Author contributions
Conceptualization, Resources, Data curation, Formal analysis, Supervision, Funding acquisition, Validation, Investigation, Visualization, Methodology, Writing - original draft, Project administration, Writing - review and editing.
Investigation.
Formal analysis, Investigation, Visualization, Writing - review and editing.
Conceptualization, Investigation, Visualization.
Resources, Methodology.
Investigation, Visualization.
Investigation.
Investigation.
Investigation.
Investigation.
Supervision.
Conceptualization, Supervision, Funding acquisition, Investigation, Methodology, Writing - review and editing.
Conceptualization, Resources, Supervision, Investigation, Methodology, Writing - review and editing.
Conceptualization, Supervision, Funding acquisition, Investigation, Visualization, Methodology, Writing - original draft, Project administration, Writing - review and editing.
Ethics
Human subjects: The use of human tissue and associated decedents' demographic information was approved by the Johns Hopkins University Institutional Review Board and ethics committee (HIPAA Form 5 exemption, Application 11-02-10-01RD) and from the Ravitz Laboratory (UCSD) through the Target ALS Consortium.
Additional files
Flies expressing 30 G4C2 repeats in the eye under control of GMR-GAL4 were crossed to the indicated UAS line and scored for enhancement (<0) or suppression (>0) as described (Zhang et al., 2015a).
Motor cortex from postmortem brain autopsies from four C9-ALS patients and four non-neurological disease controls were used is this study (Figure 7). Patient ID, cause of death (diagnosis), gender, age at death, post-mortem interval (PMI) in hours, and presence of C9orf72 expanded hexanucleotide repeat are indicated.
Data availability
All data generated or analysed during this study are included in the manuscript and supporting files.
References
- Al-Sarraj S, King A, Troakes C, Smith B, Maekawa S, Bodi I, Rogelj B, Al-Chalabi A, Hortobágyi T, Shaw CE. p62 positive, TDP-43 negative, neuronal cytoplasmic and intranuclear inclusions in the cerebellum and Hippocampus define the pathology of C9orf72-linked FTLD and MND/ALS. Acta Neuropathologica. 2011;122:691–702. doi: 10.1007/s00401-011-0911-2. [DOI] [PubMed] [Google Scholar]
- Almeida S, Gascon E, Tran H, Chou HJ, Gendron TF, Degroot S, Tapper AR, Sellier C, Charlet-Berguerand N, Karydas A, Seeley WW, Boxer AL, Petrucelli L, Miller BL, Gao FB. Modeling key pathological features of frontotemporal dementia with C9ORF72 repeat expansion in iPSC-derived human neurons. Acta Neuropathologica. 2013;126:385–399. doi: 10.1007/s00401-013-1149-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Balendra R, Isaacs AM. C9orf72-mediated ALS and FTD: multiple pathways to disease. Nature Reviews Neurology. 2018;14:544–558. doi: 10.1038/s41582-018-0047-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bharadwaj R, Cunningham KM, Zhang K, Lloyd TE. FIG4 regulates lysosome membrane homeostasis independent of phosphatase function. Human Molecular Genetics. 2016;25:681–692. doi: 10.1093/hmg/ddv505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boeynaems S, Bogaert E, Michiels E, Gijselinck I, Sieben A, Jovičić A, De Baets G, Scheveneels W, Steyaert J, Cuijt I, Verstrepen KJ, Callaerts P, Rousseau F, Schymkowitz J, Cruts M, Van Broeckhoven C, Van Damme P, Gitler AD, Robberecht W, Van Den Bosch L. Drosophila screen connects nuclear transport genes to DPR pathology in c9ALS/FTD. Scientific Reports. 2016;6:20877. doi: 10.1038/srep20877. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bouché V, Espinosa AP, Leone L, Sardiello M, Ballabio A, Botas J. Drosophila mitf regulates the V-ATPase and the lysosomal-autophagic pathway. Autophagy. 2016;12:484–498. doi: 10.1080/15548627.2015.1134081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chang YY, Neufeld TP. An Atg1/Atg13 complex with multiple roles in TOR-mediated autophagy regulation. Molecular Biology of the Cell. 2009;20:2004–2014. doi: 10.1091/mbc.e08-12-1250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Y, Liu H, Guan Y, Wang Q, Zhou F, Jie L, Ju J, Pu L, Du H, Wang X. The altered autophagy mediated by TFEB in animal and cell models of amyotrophic lateral sclerosis. American Journal of Translational Research. 2015;7:1574–1587. [PMC free article] [PubMed] [Google Scholar]
- Chou CC, Zhang Y, Umoh ME, Vaughan SW, Lorenzini I, Liu F, Sayegh M, Donlin-Asp PG, Chen YH, Duong DM, Seyfried NT, Powers MA, Kukar T, Hales CM, Gearing M, Cairns NJ, Boylan KB, Dickson DW, Rademakers R, Zhang YJ, Petrucelli L, Sattler R, Zarnescu DC, Glass JD, Rossoll W. TDP-43 pathology disrupts nuclear pore complexes and nucleocytoplasmic transport in ALS/FTD. Nature Neuroscience. 2018;21:228–239. doi: 10.1038/s41593-017-0047-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chouhan AK, Guo C, Hsieh YC, Ye H, Senturk M, Zuo Z, Li Y, Chatterjee S, Botas J, Jackson GR, Bellen HJ, Shulman JM. Uncoupling neuronal death and dysfunction in Drosophila models of neurodegenerative disease. Acta Neuropathologica Communications. 2016;4:62. doi: 10.1186/s40478-016-0333-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cipolat Mis MS, Brajkovic S, Frattini E, Di Fonzo A, Corti S. Autophagy in motor neuron disease: key pathogenetic mechanisms and therapeutic targets. Molecular and Cellular Neuroscience. 2016;72:84–90. doi: 10.1016/j.mcn.2016.01.012. [DOI] [PubMed] [Google Scholar]
- Cirulli ET, Lasseigne BN, Petrovski S, Sapp PC, Dion PA, Leblond CS, Couthouis J, Lu YF, Wang Q, Krueger BJ, Ren Z, Keebler J, Han Y, Levy SE, Boone BE, Wimbish JR, Waite LL, Jones AL, Carulli JP, Day-Williams AG, Staropoli JF, Xin WW, Chesi A, Raphael AR, McKenna-Yasek D, Cady J, Vianney de Jong JM, Kenna KP, Smith BN, Topp S, Miller J, Gkazi A, Al-Chalabi A, van den Berg LH, Veldink J, Silani V, Ticozzi N, Shaw CE, Baloh RH, Appel S, Simpson E, Lagier-Tourenne C, Pulst SM, Gibson S, Trojanowski JQ, Elman L, McCluskey L, Grossman M, Shneider NA, Chung WK, Ravits JM, Glass JD, Sims KB, Van Deerlin VM, Maniatis T, Hayes SD, Ordureau A, Swarup S, Landers J, Baas F, Allen AS, Bedlack RS, Harper JW, Gitler AD, Rouleau GA, Brown R, Harms MB, Cooper GM, Harris T, Myers RM, Goldstein DB, FALS Sequencing Consortium Exome sequencing in amyotrophic lateral sclerosis identifies risk genes and pathways. Science. 2015;347:1436–1441. doi: 10.1126/science.aaa3650. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cooper-Knock J, Hewitt C, Highley JR, Brockington A, Milano A, Man S, Martindale J, Hartley J, Walsh T, Gelsthorpe C, Baxter L, Forster G, Fox M, Bury J, Mok K, McDermott CJ, Traynor BJ, Kirby J, Wharton SB, Ince PG, Hardy J, Shaw PJ. Clinico-pathological features in amyotrophic lateral sclerosis with expansions in C9ORF72. Brain. 2012;135:751–764. doi: 10.1093/brain/awr365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cortes CJ, La Spada AR. TFEB dysregulation as a driver of autophagy dysfunction in neurodegenerative disease: molecular mechanisms, cellular processes, and emerging therapeutic opportunities. Neurobiology of Disease. 2019;122:83–93. doi: 10.1016/j.nbd.2018.05.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Castro IP, Costa AC, Celardo I, Tufi R, Dinsdale D, Loh SH, Martins LM. Drosophila ref(2)P is required for the parkin-mediated suppression of mitochondrial dysfunction in pink1 mutants. Cell Death & Disease. 2013;4:e873. doi: 10.1038/cddis.2013.394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Decressac M, Mattsson B, Weikop P, Lundblad M, Jakobsson J, Björklund A. TFEB-mediated autophagy rescues midbrain dopamine neurons from α-synuclein toxicity. PNAS. 2013;110:E1817–E1826. doi: 10.1073/pnas.1305623110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DeJesus-Hernandez M, Mackenzie IR, Boeve BF, Boxer AL, Baker M, Rutherford NJ, Nicholson AM, Finch NA, Flynn H, Adamson J, Kouri N, Wojtas A, Sengdy P, Hsiung GY, Karydas A, Seeley WW, Josephs KA, Coppola G, Geschwind DH, Wszolek ZK, Feldman H, Knopman DS, Petersen RC, Miller BL, Dickson DW, Boylan KB, Graff-Radford NR, Rademakers R. Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron. 2011;72:245–256. doi: 10.1016/j.neuron.2011.09.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Donde A, Sun M, Jeong YH, Wen X, Ling J, Lin S, Braunstein K, Nie S, Wang S, Chen L, Wong PC. Upregulation of ATG7 attenuates motor neuron dysfunction associated with depletion of TARDBP/TDP-43. Autophagy. 2020;16:672–682. doi: 10.1080/15548627.2019.1635379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Donnelly CJ, Zhang PW, Pham JT, Haeusler AR, Heusler AR, Mistry NA, Vidensky S, Daley EL, Poth EM, Hoover B, Fines DM, Maragakis N, Tienari PJ, Petrucelli L, Traynor BJ, Wang J, Rigo F, Bennett CF, Blackshaw S, Sattler R, Rothstein JD. RNA toxicity from the ALS/FTD C9ORF72 expansion is mitigated by antisense intervention. Neuron. 2013;80:415–428. doi: 10.1016/j.neuron.2013.10.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eftekharzadeh B, Daigle JG, Kapinos LE, Coyne A, Schiantarelli J, Carlomagno Y, Cook C, Miller SJ, Dujardin S, Amaral AS, Grima JC, Bennett RE, Tepper K, DeTure M, Vanderburg CR, Corjuc BT, DeVos SL, Gonzalez JA, Chew J, Vidensky S, Gage FH, Mertens J, Troncoso J, Mandelkow E, Salvatella X, Lim RYH, Petrucelli L, Wegmann S, Rothstein JD, Hyman BT. Tau protein disrupts nucleocytoplasmic transport in Alzheimer's Disease. Neuron. 2018;99:925–940. doi: 10.1016/j.neuron.2018.07.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Evans CS, Holzbaur ELF. Autophagy and mitophagy in ALS. Neurobiology of Disease. 2019;122:35–40. doi: 10.1016/j.nbd.2018.07.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Farg MA, Sundaramoorthy V, Sultana JM, Yang S, Atkinson RA, Levina V, Halloran MA, Gleeson PA, Blair IP, Soo KY, King AE, Atkin JD. C9ORF72, implicated in amytrophic lateral sclerosis and frontotemporal dementia, regulates endosomal trafficking. Human Molecular Genetics. 2014;23:3579–3595. doi: 10.1093/hmg/ddu068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fraldi A, Klein AD, Medina DL, Settembre C. Brain disorders due to lysosomal dysfunction. Annual Review of Neuroscience. 2016;39:277–295. doi: 10.1146/annurev-neuro-070815-014031. [DOI] [PubMed] [Google Scholar]
- Freibaum BD, Lu Y, Lopez-Gonzalez R, Kim NC, Almeida S, Lee KH, Badders N, Valentine M, Miller BL, Wong PC, Petrucelli L, Kim HJ, Gao FB, Taylor JP. GGGGCC repeat expansion in C9orf72 compromises nucleocytoplasmic transport. Nature. 2015;525:129–133. doi: 10.1038/nature14974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao FB, Almeida S, Lopez-Gonzalez R. Dysregulated molecular pathways in amyotrophic lateral sclerosis–frontotemporal dementia spectrum disorder. The EMBO Journal. 2017;36:2931–2950. doi: 10.15252/embj.201797568. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gasset-Rosa F, Chillon-Marinas C, Goginashvili A, Atwal RS, Artates JW, Tabet R, Wheeler VC, Bang AG, Cleveland DW, Lagier-Tourenne C. Polyglutamine-Expanded huntingtin exacerbates Age-Related disruption of nuclear integrity and nucleocytoplasmic transport. Neuron. 2017;94:48–57. doi: 10.1016/j.neuron.2017.03.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gasset-Rosa F, Lu S, Yu H, Chen C, Melamed Z, Guo L, Shorter J, Da Cruz S, Cleveland DW. Cytoplasmic TDP-43 De-mixing independent of stress granules drives inhibition of nuclear import, loss of nuclear TDP-43, and cell death. Neuron. 2019;102:339–357. doi: 10.1016/j.neuron.2019.02.038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goode A, Butler K, Long J, Cavey J, Scott D, Shaw B, Sollenberger J, Gell C, Johansen T, Oldham NJ, Searle MS, Layfield R. Defective recognition of LC3B by mutant SQSTM1/p62 implicates impairment of autophagy as a pathogenic mechanism in ALS-FTLD. Autophagy. 2016;12:1094–1104. doi: 10.1080/15548627.2016.1170257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goodman LD, Prudencio M, Kramer NJ, Martinez-Ramirez LF, Srinivasan AR, Lan M, Parisi MJ, Zhu Y, Chew J, Cook CN, Berson A, Gitler AD, Petrucelli L, Bonini NM. Toxic expanded GGGGCC repeat transcription is mediated by the PAF1 complex in C9orf72-associated FTD. Nature Neuroscience. 2019a;22:863–874. doi: 10.1038/s41593-019-0396-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goodman LD, Prudencio M, Srinivasan AR, Rifai OM, Lee VM, Petrucelli L, Bonini NM. eIF4B and eIF4H mediate GR production from expanded G4C2 in a Drosophila model for C9orf72-associated ALS. Acta Neuropathologica Communications. 2019b;7:62. doi: 10.1186/s40478-019-0711-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grima JC, Daigle JG, Arbez N, Cunningham KC, Zhang K, Ochaba J, Geater C, Morozko E, Stocksdale J, Glatzer JC, Pham JT, Ahmed I, Peng Q, Wadhwa H, Pletnikova O, Troncoso JC, Duan W, Snyder SH, Ranum LPW, Thompson LM, Lloyd TE, Ross CA, Rothstein JD. Mutant huntingtin disrupts the nuclear pore complex. Neuron. 2017;94:93–107. doi: 10.1016/j.neuron.2017.03.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hallsson JH, Haflidadóttir BS, Stivers C, Odenwald W, Arnheiter H, Pignoni F, Steingrímsson E. The basic helix-loop-helix leucine zipper transcription factor mitf is conserved in Drosophila and functions in eye development. Genetics. 2004;167:233–241. doi: 10.1534/genetics.167.1.233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hara T, Nakamura K, Matsui M, Yamamoto A, Nakahara Y, Suzuki-Migishima R, Yokoyama M, Mishima K, Saito I, Okano H, Mizushima N. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature. 2006;441:885–889. doi: 10.1038/nature04724. [DOI] [PubMed] [Google Scholar]
- Hariri M, Millane G, Guimond MP, Guay G, Dennis JW, Nabi IR. Biogenesis of multilamellar bodies via autophagy. Molecular Biology of the Cell. 2000;11:255–268. doi: 10.1091/mbc.11.1.255. [DOI] [PMC free article] [PubMed] [Google Scholar]
- ITALSGEN Consortium. Renton AE, Majounie E, Waite A, Simón-Sánchez J, Rollinson S, Gibbs JR, Schymick JC, Laaksovirta H, van Swieten JC, Myllykangas L, Kalimo H, Paetau A, Abramzon Y, Remes AM, Kaganovich A, Scholz SW, Duckworth J, Ding J, Harmer DW, Hernandez DG, Johnson JO, Mok K, Ryten M, Trabzuni D, Guerreiro RJ, Orrell RW, Neal J, Murray A, Pearson J, Jansen IE, Sondervan D, Seelaar H, Blake D, Young K, Halliwell N, Callister JB, Toulson G, Richardson A, Gerhard A, Snowden J, Mann D, Neary D, Nalls MA, Peuralinna T, Jansson L, Isoviita VM, Kaivorinne AL, Hölttä-Vuori M, Ikonen E, Sulkava R, Benatar M, Wuu J, Chiò A, Restagno G, Borghero G, Sabatelli M, Heckerman D, Rogaeva E, Zinman L, Rothstein JD, Sendtner M, Drepper C, Eichler EE, Alkan C, Abdullaev Z, Pack SD, Dutra A, Pak E, Hardy J, Singleton A, Williams NM, Heutink P, Pickering-Brown S, Morris HR, Tienari PJ, Traynor BJ. A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron. 2011;72:257–268. doi: 10.1016/j.neuron.2011.09.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ji YJ, Ugolino J, Brady NR, Hamacher-Brady A, Wang J. Systemic deregulation of autophagy upon loss of ALS- and FTD-linked C9orf72. Autophagy. 2017;13:1254–1255. doi: 10.1080/15548627.2017.1299312. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jovičić A, Mertens J, Boeynaems S, Bogaert E, Chai N, Yamada SB, Paul JW, Sun S, Herdy JR, Bieri G, Kramer NJ, Gage FH, Van Den Bosch L, Robberecht W, Gitler AD. Modifiers of C9orf72 dipeptide repeat toxicity connect nucleocytoplasmic transport defects to FTD/ALS. Nature Neuroscience. 2015;18:1226–1229. doi: 10.1038/nn.4085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kinser RD, Dolph PJ. Cathepsin proteases mediate photoreceptor cell degeneration in Drosophila. Neurobiology of Disease. 2012;46:655–662. doi: 10.1016/j.nbd.2012.03.004. [DOI] [PubMed] [Google Scholar]
- Klionsky DJ, Abdelmohsen K, Abe A, Abedin MJ, Abeliovich H, Acevedo Arozena A, Adachi H, Adams CM, Adams PD, Adeli K, Adhihetty PJ, Adler SG, Agam G, Agarwal R, Aghi MK, Agnello M, Agostinis P, Aguilar PV, Aguirre-Ghiso J, Airoldi EM, Ait-Si-Ali S, Akematsu T, Akporiaye ET, Al-Rubeai M, Albaiceta GM, Albanese C, Albani D, Albert ML, Aldudo J, Algül H, Alirezaei M, Alloza I, Almasan A, Almonte-Beceril M, Alnemri ES, Alonso C, Altan-Bonnet N, Altieri DC, Alvarez S, Alvarez-Erviti L, Alves S, Amadoro G, Amano A, Amantini C, Ambrosio S, Amelio I, Amer AO, Amessou M, Amon A, An Z, Anania FA, Andersen SU, Andley UP, Andreadi CK, Andrieu-Abadie N, Anel A, Ann DK, Anoopkumar-Dukie S, Antonioli M, Aoki H, Apostolova N, Aquila S, Aquilano K, Araki K, Arama E, Aranda A, Araya J, Arcaro A, Arias E, Arimoto H, Ariosa AR, Armstrong JL, Arnould T, Arsov I, Asanuma K, Askanas V, Asselin E, Atarashi R, Atherton SS, Atkin JD, Attardi LD, Auberger P, Auburger G, Aurelian L, Autelli R, Avagliano L, Avantaggiati ML, Avrahami L, Awale S, Azad N, Bachetti T, Backer JM, Bae DH, Bae JS, Bae ON, Bae SH, Baehrecke EH, Baek SH, Baghdiguian S, Bagniewska-Zadworna A, Bai H, Bai J, Bai XY, Bailly Y, Balaji KN, Balduini W, Ballabio A, Balzan R, Banerjee R, Bánhegyi G, Bao H, Barbeau B, Barrachina MD, Barreiro E, Bartel B, Bartolomé A, Bassham DC, Bassi MT, Bast RC, Basu A, Batista MT, Batoko H, Battino M, Bauckman K, Baumgarner BL, Bayer KU, Beale R, Beaulieu JF, Beck GR, Becker C, Beckham JD, Bédard PA, Bednarski PJ, Begley TJ, Behl C, Behrends C, Behrens GM, Behrns KE, Bejarano E, Belaid A, Belleudi F, Bénard G, Berchem G, Bergamaschi D, Bergami M, Berkhout B, Berliocchi L, Bernard A, Bernard M, Bernassola F, Bertolotti A, Bess AS, Besteiro S, Bettuzzi S, Bhalla S, Bhattacharyya S, Bhutia SK, Biagosch C, Bianchi MW, Biard-Piechaczyk M, Billes V, Bincoletto C, Bingol B, Bird SW, Bitoun M, Bjedov I, Blackstone C, Blanc L, Blanco GA, Blomhoff HK, Boada-Romero E, Böckler S, Boes M, Boesze-Battaglia K, Boise LH, Bolino A, Boman A, Bonaldo P, Bordi M, Bosch J, Botana LM, Botti J, Bou G, Bouché M, Bouchecareilh M, Boucher MJ, Boulton ME, Bouret SG, Boya P, Boyer-Guittaut M, Bozhkov PV, Brady N, Braga VM, Brancolini C, Braus GH, Bravo-San Pedro JM, Brennan LA, Bresnick EH, Brest P, Bridges D, Bringer MA, Brini M, Brito GC, Brodin B, Brookes PS, Brown EJ, Brown K, Broxmeyer HE, Bruhat A, Brum PC, Brumell JH, Brunetti-Pierri N, Bryson-Richardson RJ, Buch S, Buchan AM, Budak H, Bulavin DV, Bultman SJ, Bultynck G, Bumbasirevic V, Burelle Y, Burke RE, Burmeister M, Bütikofer P, Caberlotto L, Cadwell K, Cahova M, Cai D, Cai J, Cai Q, Calatayud S, Camougrand N, Campanella M, Campbell GR, Campbell M, Campello S, Candau R, Caniggia I, Cantoni L, Cao L, Caplan AB, Caraglia M, Cardinali C, Cardoso SM, Carew JS, Carleton LA, Carlin CR, Carloni S, Carlsson SR, Carmona-Gutierrez D, Carneiro LA, Carnevali O, Carra S, Carrier A, Carroll B, Casas C, Casas J, Cassinelli G, Castets P, Castro-Obregon S, Cavallini G, Ceccherini I, Cecconi F, Cederbaum AI, Ceña V, Cenci S, Cerella C, Cervia D, Cetrullo S, Chaachouay H, Chae HJ, Chagin AS, Chai CY, Chakrabarti G, Chamilos G, Chan EY, Chan MT, Chandra D, Chandra P, Chang CP, Chang RC, Chang TY, Chatham JC, Chatterjee S, Chauhan S, Che Y, Cheetham ME, Cheluvappa R, Chen CJ, Chen G, Chen GC, Chen G, Chen H, Chen JW, Chen JK, Chen M, Chen M, Chen P, Chen Q, Chen Q, Chen SD, Chen S, Chen SS, Chen W, Chen WJ, Chen WQ, Chen W, Chen X, Chen YH, Chen YG, Chen Y, Chen Y, Chen Y, Chen YJ, Chen YQ, Chen Y, Chen Z, Chen Z, Cheng A, Cheng CH, Cheng H, Cheong H, Cherry S, Chesney J, Cheung CH, Chevet E, Chi HC, Chi SG, Chiacchiera F, Chiang HL, Chiarelli R, Chiariello M, Chieppa M, Chin LS, Chiong M, Chiu GN, Cho DH, Cho SG, Cho WC, Cho YY, Cho YS, Choi AM, Choi EJ, Choi EK, Choi J, Choi ME, Choi SI, Chou TF, Chouaib S, Choubey D, Choubey V, Chow KC, Chowdhury K, Chu CT, Chuang TH, Chun T, Chung H, Chung T, Chung YL, Chwae YJ, Cianfanelli V, Ciarcia R, Ciechomska IA, Ciriolo MR, Cirone M, Claerhout S, Clague MJ, Clària J, Clarke PG, Clarke R, Clementi E, Cleyrat C, Cnop M, Coccia EM, Cocco T, Codogno P, Coers J, Cohen EE, Colecchia D, Coletto L, Coll NS, Colucci-Guyon E, Comincini S, Condello M, Cook KL, Coombs GH, Cooper CD, Cooper JM, Coppens I, Corasaniti MT, Corazzari M, Corbalan R, Corcelle-Termeau E, Cordero MD, Corral-Ramos C, Corti O, Cossarizza A, Costelli P, Costes S, Cotman SL, Coto-Montes A, Cottet S, Couve E, Covey LR, Cowart LA, Cox JS, Coxon FP, Coyne CB, Cragg MS, Craven RJ, Crepaldi T, Crespo JL, Criollo A, Crippa V, Cruz MT, Cuervo AM, Cuezva JM, Cui T, Cutillas PR, Czaja MJ, Czyzyk-Krzeska MF, Dagda RK, Dahmen U, Dai C, Dai W, Dai Y, Dalby KN, Dalla Valle L, Dalmasso G, D'Amelio M, Damme M, Darfeuille-Michaud A, Dargemont C, Darley-Usmar VM, Dasarathy S, Dasgupta B, Dash S, Dass CR, Davey HM, Davids LM, Dávila D, Davis RJ, Dawson TM, Dawson VL, Daza P, de Belleroche J, de Figueiredo P, de Figueiredo RC, de la Fuente J, De Martino L, De Matteis A, De Meyer GR, De Milito A, De Santi M, de Souza W, De Tata V, De Zio D, Debnath J, Dechant R, Decuypere JP, Deegan S, Dehay B, Del Bello B, Del Re DP, Delage-Mourroux R, Delbridge LM, Deldicque L, Delorme-Axford E, Deng Y, Dengjel J, Denizot M, Dent P, Der CJ, Deretic V, Derrien B, Deutsch E, Devarenne TP, Devenish RJ, Di Bartolomeo S, Di Daniele N, Di Domenico F, Di Nardo A, Di Paola S, Di Pietro A, Di Renzo L, DiAntonio A, Díaz-Araya G, Díaz-Laviada I, Diaz-Meco MT, Diaz-Nido J, Dickey CA, Dickson RC, Diederich M, Digard P, Dikic I, Dinesh-Kumar SP, Ding C, Ding WX, Ding Z, Dini L, Distler JH, Diwan A, Djavaheri-Mergny M, Dmytruk K, Dobson RC, Doetsch V, Dokladny K, Dokudovskaya S, Donadelli M, Dong XC, Dong X, Dong Z, Donohue TM, Doran KS, D'Orazi G, Dorn GW, Dosenko V, Dridi S, Drucker L, Du J, Du LL, Du L, du Toit A, Dua P, Duan L, Duann P, Dubey VK, Duchen MR, Duchosal MA, Duez H, Dugail I, Dumit VI, Duncan MC, Dunlop EA, Dunn WA, Dupont N, Dupuis L, Durán RV, Durcan TM, Duvezin-Caubet S, Duvvuri U, Eapen V, Ebrahimi-Fakhari D, Echard A, Eckhart L, Edelstein CL, Edinger AL, Eichinger L, Eisenberg T, Eisenberg-Lerner A, Eissa NT, El-Deiry WS, El-Khoury V, Elazar Z, Eldar-Finkelman H, Elliott CJ, Emanuele E, Emmenegger U, Engedal N, Engelbrecht AM, Engelender S, Enserink JM, Erdmann R, Erenpreisa J, Eri R, Eriksen JL, Erman A, Escalante R, Eskelinen EL, Espert L, Esteban-Martínez L, Evans TJ, Fabri M, Fabrias G, Fabrizi C, Facchiano A, Færgeman NJ, Faggioni A, Fairlie WD, Fan C, Fan D, Fan J, Fang S, Fanto M, Fanzani A, Farkas T, Faure M, Favier FB, Fearnhead H, Federici M, Fei E, Felizardo TC, Feng H, Feng Y, Feng Y, Ferguson TA, Fernández ÁF, Fernandez-Barrena MG, Fernandez-Checa JC, Fernández-López A, Fernandez-Zapico ME, Feron O, Ferraro E, Ferreira-Halder CV, Fesus L, Feuer R, Fiesel FC, Filippi-Chiela EC, Filomeni G, Fimia GM, Fingert JH, Finkbeiner S, Finkel T, Fiorito F, Fisher PB, Flajolet M, Flamigni F, Florey O, Florio S, Floto RA, Folini M, Follo C, Fon EA, Fornai F, Fortunato F, Fraldi A, Franco R, Francois A, François A, Frankel LB, Fraser ID, Frey N, Freyssenet DG, Frezza C, Friedman SL, Frigo DE, Fu D, Fuentes JM, Fueyo J, Fujitani Y, Fujiwara Y, Fujiya M, Fukuda M, Fulda S, Fusco C, Gabryel B, Gaestel M, Gailly P, Gajewska M, Galadari S, Galili G, Galindo I, Galindo MF, Galliciotti G, Galluzzi L, Galluzzi L, Galy V, Gammoh N, Gandy S, Ganesan AK, Ganesan S, Ganley IG, Gannagé M, Gao FB, Gao F, Gao JX, García Nannig L, García Véscovi E, Garcia-Macía M, Garcia-Ruiz C, Garg AD, Garg PK, Gargini R, Gassen NC, Gatica D, Gatti E, Gavard J, Gavathiotis E, Ge L, Ge P, Ge S, Gean PW, Gelmetti V, Genazzani AA, Geng J, Genschik P, Gerner L, Gestwicki JE, Gewirtz DA, Ghavami S, Ghigo E, Ghosh D, Giammarioli AM, Giampieri F, Giampietri C, Giatromanolaki A, Gibbings DJ, Gibellini L, Gibson SB, Ginet V, Giordano A, Giorgini F, Giovannetti E, Girardin SE, Gispert S, Giuliano S, Gladson CL, Glavic A, Gleave M, Godefroy N, Gogal RM, Gokulan K, Goldman GH, Goletti D, Goligorsky MS, Gomes AV, Gomes LC, Gomez H, Gomez-Manzano C, Gómez-Sánchez R, Gonçalves DA, Goncu E, Gong Q, Gongora C, Gonzalez CB, Gonzalez-Alegre P, Gonzalez-Cabo P, González-Polo RA, Goping IS, Gorbea C, Gorbunov NV, Goring DR, Gorman AM, Gorski SM, Goruppi S, Goto-Yamada S, Gotor C, Gottlieb RA, Gozes I, Gozuacik D, Graba Y, Graef M, Granato GE, Grant GD, Grant S, Gravina GL, Green DR, Greenhough A, Greenwood MT, Grimaldi B, Gros F, Grose C, Groulx JF, Gruber F, Grumati P, Grune T, Guan JL, Guan KL, Guerra B, Guillen C, Gulshan K, Gunst J, Guo C, Guo L, Guo M, Guo W, Guo XG, Gust AA, Gustafsson ÅB, Gutierrez E, Gutierrez MG, Gwak HS, Haas A, Haber JE, Hadano S, Hagedorn M, Hahn DR, Halayko AJ, Hamacher-Brady A, Hamada K, Hamai A, Hamann A, Hamasaki M, Hamer I, Hamid Q, Hammond EM, Han F, Han W, Handa JT, Hanover JA, Hansen M, Harada M, Harhaji-Trajkovic L, Harper JW, Harrath AH, Harris AL, Harris J, Hasler U, Hasselblatt P, Hasui K, Hawley RG, Hawley TS, He C, He CY, He F, He G, He RR, He XH, He YW, He YY, Heath JK, Hébert MJ, Heinzen RA, Helgason GV, Hensel M, Henske EP, Her C, Herman PK, Hernández A, Hernandez C, Hernández-Tiedra S, Hetz C, Hiesinger PR, Higaki K, Hilfiker S, Hill BG, Hill JA, Hill WD, Hino K, Hofius D, Hofman P, Höglinger GU, Höhfeld J, Holz MK, Hong Y, Hood DA, Hoozemans JJ, Hoppe T, Hsu C, Hsu CY, Hsu LC, Hu D, Hu G, Hu HM, Hu H, Hu MC, Hu YC, Hu ZW, Hua F, Hua Y, Huang C, Huang HL, Huang KH, Huang KY, Huang S, Huang S, Huang WP, Huang YR, Huang Y, Huang Y, Huber TB, Huebbe P, Huh WK, Hulmi JJ, Hur GM, Hurley JH, Husak Z, Hussain SN, Hussain S, Hwang JJ, Hwang S, Hwang TI, Ichihara A, Imai Y, Imbriano C, Inomata M, Into T, Iovane V, Iovanna JL, Iozzo RV, Ip NY, Irazoqui JE, Iribarren P, Isaka Y, Isakovic AJ, Ischiropoulos H, Isenberg JS, Ishaq M, Ishida H, Ishii I, Ishmael JE, Isidoro C, Isobe K, Isono E, Issazadeh-Navikas S, Itahana K, Itakura E, Ivanov AI, Iyer AK, Izquierdo JM, Izumi Y, Izzo V, Jäättelä M, Jaber N, Jackson DJ, Jackson WT, Jacob TG, Jacques TS, Jagannath C, Jain A, Jana NR, Jang BK, Jani A, Janji B, Jannig PR, Jansson PJ, Jean S, Jendrach M, Jeon JH, Jessen N, Jeung EB, Jia K, Jia L, Jiang H, Jiang H, Jiang L, Jiang T, Jiang X, Jiang X, Jiang X, Jiang Y, Jiang Y, Jiménez A, Jin C, Jin H, Jin L, Jin M, Jin S, Jinwal UK, Jo EK, Johansen T, Johnson DE, Johnson GV, Johnson JD, Jonasch E, Jones C, Joosten LA, Jordan J, Joseph AM, Joseph B, Joubert AM, Ju D, Ju J, Juan HF, Juenemann K, Juhász G, Jung HS, Jung JU, Jung YK, Jungbluth H, Justice MJ, Jutten B, Kaakoush NO, Kaarniranta K, Kaasik A, Kabuta T, Kaeffer B, Kågedal K, Kahana A, Kajimura S, Kakhlon O, Kalia M, Kalvakolanu DV, Kamada Y, Kambas K, Kaminskyy VO, Kampinga HH, Kandouz M, Kang C, Kang R, Kang TC, Kanki T, Kanneganti TD, Kanno H, Kanthasamy AG, Kantorow M, Kaparakis-Liaskos M, Kapuy O, Karantza V, Karim MR, Karmakar P, Kaser A, Kaushik S, Kawula T, Kaynar AM, Ke PY, Ke ZJ, Kehrl JH, Keller KE, Kemper JK, Kenworthy AK, Kepp O, Kern A, Kesari S, Kessel D, Ketteler R, Kettelhut IC, Khambu B, Khan MM, Khandelwal VK, Khare S, Kiang JG, Kiger AA, Kihara A, Kim AL, Kim CH, Kim DR, Kim DH, Kim EK, Kim HY, Kim HR, Kim JS, Kim JH, Kim JC, Kim JH, Kim KW, Kim MD, Kim MM, Kim PK, Kim SW, Kim SY, Kim YS, Kim Y, Kimchi A, Kimmelman AC, Kimura T, King JS, Kirkegaard K, Kirkin V, Kirshenbaum LA, Kishi S, Kitajima Y, Kitamoto K, Kitaoka Y, Kitazato K, Kley RA, Klimecki WT, Klinkenberg M, Klucken J, Knævelsrud H, Knecht E, Knuppertz L, Ko JL, Kobayashi S, Koch JC, Koechlin-Ramonatxo C, Koenig U, Koh YH, Köhler K, Kohlwein SD, Koike M, Komatsu M, Kominami E, Kong D, Kong HJ, Konstantakou EG, Kopp BT, Korcsmaros T, Korhonen L, Korolchuk VI, Koshkina NV, Kou Y, Koukourakis MI, Koumenis C, Kovács AL, Kovács T, Kovacs WJ, Koya D, Kraft C, Krainc D, Kramer H, Kravic-Stevovic T, Krek W, Kretz-Remy C, Krick R, Krishnamurthy M, Kriston-Vizi J, Kroemer G, Kruer MC, Kruger R, Ktistakis NT, Kuchitsu K, Kuhn C, Kumar AP, Kumar A, Kumar A, Kumar D, Kumar D, Kumar R, Kumar S, Kundu M, Kung HJ, Kuno A, Kuo SH, Kuret J, Kurz T, Kwok T, Kwon TK, Kwon YT, Kyrmizi I, La Spada AR, Lafont F, Lahm T, Lakkaraju A, Lam T, Lamark T, Lancel S, Landowski TH, Lane DJ, Lane JD, Lanzi C, Lapaquette P, Lapierre LR, Laporte J, Laukkarinen J, Laurie GW, Lavandero S, Lavie L, LaVoie MJ, Law BY, Law HK, Law KB, Layfield R, Lazo PA, Le Cam L, Le Roch KG, Le Stunff H, Leardkamolkarn V, Lecuit M, Lee BH, Lee CH, Lee EF, Lee GM, Lee HJ, Lee H, Lee JK, Lee J, Lee JH, Lee JH, Lee M, Lee MS, Lee PJ, Lee SW, Lee SJ, Lee SJ, Lee SY, Lee SH, Lee SS, Lee SJ, Lee S, Lee YR, Lee YJ, Lee YH, Leeuwenburgh C, Lefort S, Legouis R, Lei J, Lei QY, Leib DA, Leibowitz G, Lekli I, Lemaire SD, Lemasters JJ, Lemberg MK, Lemoine A, Leng S, Lenz G, Lenzi P, Lerman LO, Lettieri Barbato D, Leu JI, Leung HY, Levine B, Lewis PA, Lezoualc'h F, Li C, Li F, Li FJ, Li J, Li K, Li L, Li M, Li M, Li Q, Li R, Li S, Li W, Li W, Li X, Li Y, Lian J, Liang C, Liang Q, Liao Y, Liberal J, Liberski PP, Lie P, Lieberman AP, Lim HJ, Lim KL, Lim K, Lima RT, Lin CS, Lin CF, Lin F, Lin F, Lin FC, Lin K, Lin KH, Lin PH, Lin T, Lin WW, Lin YS, Lin Y, Linden R, Lindholm D, Lindqvist LM, Lingor P, Linkermann A, Liotta LA, Lipinski MM, Lira VA, Lisanti MP, Liton PB, Liu B, Liu C, Liu CF, Liu F, Liu HJ, Liu J, Liu JJ, Liu JL, Liu K, Liu L, Liu L, Liu Q, Liu RY, Liu S, Liu S, Liu W, Liu XD, Liu X, Liu XH, Liu X, Liu X, Liu X, Liu Y, Liu Y, Liu Z, Liu Z, Liuzzi JP, Lizard G, Ljujic M, Lodhi IJ, Logue SE, Lokeshwar BL, Long YC, Lonial S, Loos B, López-Otín C, López-Vicario C, Lorente M, Lorenzi PL, Lõrincz P, Los M, Lotze MT, Lovat PE, Lu B, Lu B, Lu J, Lu Q, Lu SM, Lu S, Lu Y, Luciano F, Luckhart S, Lucocq JM, Ludovico P, Lugea A, Lukacs NW, Lum JJ, Lund AH, Luo H, Luo J, Luo S, Luparello C, Lyons T, Ma J, Ma Y, Ma Y, Ma Z, Machado J, Machado-Santelli GM, Macian F, MacIntosh GC, MacKeigan JP, Macleod KF, MacMicking JD, MacMillan-Crow LA, Madeo F, Madesh M, Madrigal-Matute J, Maeda A, Maeda T, Maegawa G, Maellaro E, Maes H, Magariños M, Maiese K, Maiti TK, Maiuri L, Maiuri MC, Maki CG, Malli R, Malorni W, Maloyan A, Mami-Chouaib F, Man N, Mancias JD, Mandelkow EM, Mandell MA, Manfredi AA, Manié SN, Manzoni C, Mao K, Mao Z, Mao ZW, Marambaud P, Marconi AM, Marelja Z, Marfe G, Margeta M, Margittai E, Mari M, Mariani FV, Marin C, Marinelli S, Mariño G, Markovic I, Marquez R, Martelli AM, Martens S, Martin KR, Martin SJ, Martin S, Martin-Acebes MA, Martín-Sanz P, Martinand-Mari C, Martinet W, Martinez J, Martinez-Lopez N, Martinez-Outschoorn U, Martínez-Velázquez M, Martinez-Vicente M, Martins WK, Mashima H, Mastrianni JA, Matarese G, Matarrese P, Mateo R, Matoba S, Matsumoto N, Matsushita T, Matsuura A, Matsuzawa T, Mattson MP, Matus S, Maugeri N, Mauvezin C, Mayer A, Maysinger D, Mazzolini GD, McBrayer MK, McCall K, McCormick C, McInerney GM, McIver SC, McKenna S, McMahon JJ, McNeish IA, Mechta-Grigoriou F, Medema JP, Medina DL, Megyeri K, Mehrpour M, Mehta JL, Mei Y, Meier UC, Meijer AJ, Meléndez A, Melino G, Melino S, de Melo EJ, Mena MA, Meneghini MD, Menendez JA, Menezes R, Meng L, Meng LH, Meng S, Menghini R, Menko AS, Menna-Barreto RF, Menon MB, Meraz-Ríos MA, Merla G, Merlini L, Merlot AM, Meryk A, Meschini S, Meyer JN, Mi MT, Miao CY, Micale L, Michaeli S, Michiels C, Migliaccio AR, Mihailidou AS, Mijaljica D, Mikoshiba K, Milan E, Miller-Fleming L, Mills GB, Mills IG, Minakaki G, Minassian BA, Ming XF, Minibayeva F, Minina EA, Mintern JD, Minucci S, Miranda-Vizuete A, Mitchell CH, Miyamoto S, Miyazawa K, Mizushima N, Mnich K, Mograbi B, Mohseni S, Moita LF, Molinari M, Molinari M, Møller AB, Mollereau B, Mollinedo F, Mongillo M, Monick MM, Montagnaro S, Montell C, Moore DJ, Moore MN, Mora-Rodriguez R, Moreira PI, Morel E, Morelli MB, Moreno S, Morgan MJ, Moris A, Moriyasu Y, Morrison JL, Morrison LA, Morselli E, Moscat J, Moseley PL, Mostowy S, Motori E, Mottet D, Mottram JC, Moussa CE, Mpakou VE, Mukhtar H, Mulcahy Levy JM, Muller S, Muñoz-Moreno R, Muñoz-Pinedo C, Münz C, Murphy ME, Murray JT, Murthy A, Mysorekar IU, Nabi IR, Nabissi M, Nader GA, Nagahara Y, Nagai Y, Nagata K, Nagelkerke A, Nagy P, Naidu SR, Nair S, Nakano H, Nakatogawa H, Nanjundan M, Napolitano G, Naqvi NI, Nardacci R, Narendra DP, Narita M, Nascimbeni AC, Natarajan R, Navegantes LC, Nawrocki ST, Nazarko TY, Nazarko VY, Neill T, Neri LM, Netea MG, Netea-Maier RT, Neves BM, Ney PA, Nezis IP, Nguyen HT, Nguyen HP, Nicot AS, Nilsen H, Nilsson P, Nishimura M, Nishino I, Niso-Santano M, Niu H, Nixon RA, Njar VC, Noda T, Noegel AA, Nolte EM, Norberg E, Norga KK, Noureini SK, Notomi S, Notterpek L, Nowikovsky K, Nukina N, Nürnberger T, O'Donnell VB, O'Donovan T, O'Dwyer PJ, Oehme I, Oeste CL, Ogawa M, Ogretmen B, Ogura Y, Oh YJ, Ohmuraya M, Ohshima T, Ojha R, Okamoto K, Okazaki T, Oliver FJ, Ollinger K, Olsson S, Orban DP, Ordonez P, Orhon I, Orosz L, O'Rourke EJ, Orozco H, Ortega AL, Ortona E, Osellame LD, Oshima J, Oshima S, Osiewacz HD, Otomo T, Otsu K, Ou JH, Outeiro TF, Ouyang DY, Ouyang H, Overholtzer M, Ozbun MA, Ozdinler PH, Ozpolat B, Pacelli C, Paganetti P, Page G, Pages G, Pagnini U, Pajak B, Pak SC, Pakos-Zebrucka K, Pakpour N, Palková Z, Palladino F, Pallauf K, Pallet N, Palmieri M, Paludan SR, Palumbo C, Palumbo S, Pampliega O, Pan H, Pan W, Panaretakis T, Pandey A, Pantazopoulou A, Papackova Z, Papademetrio DL, Papassideri I, Papini A, Parajuli N, Pardo J, Parekh VV, Parenti G, Park JI, Park J, Park OK, Parker R, Parlato R, Parys JB, Parzych KR, Pasquet JM, Pasquier B, Pasumarthi KB, Patschan D, Patterson C, Pattingre S, Pattison S, Pause A, Pavenstädt H, Pavone F, Pedrozo Z, Peña FJ, Peñalva MA, Pende M, Peng J, Penna F, Penninger JM, Pensalfini A, Pepe S, Pereira GJ, Pereira PC, Pérez-de la Cruz V, Pérez-Pérez ME, Pérez-Rodríguez D, Pérez-Sala D, Perier C, Perl A, Perlmutter DH, Perrotta I, Pervaiz S, Pesonen M, Pessin JE, Peters GJ, Petersen M, Petrache I, Petrof BJ, Petrovski G, Phang JM, Piacentini M, Pierdominici M, Pierre P, Pierrefite-Carle V, Pietrocola F, Pimentel-Muiños FX, Pinar M, Pineda B, Pinkas-Kramarski R, Pinti M, Pinton P, Piperdi B, Piret JM, Platanias LC, Platta HW, Plowey ED, Pöggeler S, Poirot M, Polčic P, Poletti A, Poon AH, Popelka H, Popova B, Poprawa I, Poulose SM, Poulton J, Powers SK, Powers T, Pozuelo-Rubio M, Prak K, Prange R, Prescott M, Priault M, Prince S, Proia RL, Proikas-Cezanne T, Prokisch H, Promponas VJ, Przyklenk K, Puertollano R, Pugazhenthi S, Puglielli L, Pujol A, Puyal J, Pyeon D, Qi X, Qian WB, Qin ZH, Qiu Y, Qu Z, Quadrilatero J, Quinn F, Raben N, Rabinowich H, Radogna F, Ragusa MJ, Rahmani M, Raina K, Ramanadham S, Ramesh R, Rami A, Randall-Demllo S, Randow F, Rao H, Rao VA, Rasmussen BB, Rasse TM, Ratovitski EA, Rautou PE, Ray SK, Razani B, Reed BH, Reggiori F, Rehm M, Reichert AS, Rein T, Reiner DJ, Reits E, Ren J, Ren X, Renna M, Reusch JE, Revuelta JL, Reyes L, Rezaie AR, Richards RI, Richardson DR, Richetta C, Riehle MA, Rihn BH, Rikihisa Y, Riley BE, Rimbach G, Rippo MR, Ritis K, Rizzi F, Rizzo E, Roach PJ, Robbins J, Roberge M, Roca G, Roccheri MC, Rocha S, Rodrigues CM, Rodríguez CI, de Cordoba SR, Rodriguez-Muela N, Roelofs J, Rogov VV, Rohn TT, Rohrer B, Romanelli D, Romani L, Romano PS, Roncero MI, Rosa JL, Rosello A, Rosen KV, Rosenstiel P, Rost-Roszkowska M, Roth KA, Roué G, Rouis M, Rouschop KM, Ruan DT, Ruano D, Rubinsztein DC, Rucker EB, Rudich A, Rudolf E, Rudolf R, Ruegg MA, Ruiz-Roldan C, Ruparelia AA, Rusmini P, Russ DW, Russo GL, Russo G, Russo R, Rusten TE, Ryabovol V, Ryan KM, Ryter SW, Sabatini DM, Sacher M, Sachse C, Sack MN, Sadoshima J, Saftig P, Sagi-Eisenberg R, Sahni S, Saikumar P, Saito T, Saitoh T, Sakakura K, Sakoh-Nakatogawa M, Sakuraba Y, Salazar-Roa M, Salomoni P, Saluja AK, Salvaterra PM, Salvioli R, Samali A, Sanchez AM, Sánchez-Alcázar JA, Sanchez-Prieto R, Sandri M, Sanjuan MA, Santaguida S, Santambrogio L, Santoni G, Dos Santos CN, Saran S, Sardiello M, Sargent G, Sarkar P, Sarkar S, Sarrias MR, Sarwal MM, Sasakawa C, Sasaki M, Sass M, Sato K, Sato M, Satriano J, Savaraj N, Saveljeva S, Schaefer L, Schaible UE, Scharl M, Schatzl HM, Schekman R, Scheper W, Schiavi A, Schipper HM, Schmeisser H, Schmidt J, Schmitz I, Schneider BE, Schneider EM, Schneider JL, Schon EA, Schönenberger MJ, Schönthal AH, Schorderet DF, Schröder B, Schuck S, Schulze RJ, Schwarten M, Schwarz TL, Sciarretta S, Scotto K, Scovassi AI, Screaton RA, Screen M, Seca H, Sedej S, Segatori L, Segev N, Seglen PO, Seguí-Simarro JM, Segura-Aguilar J, Seki E, Sell C, Seiliez I, Semenkovich CF, Semenza GL, Sen U, Serra AL, Serrano-Puebla A, Sesaki H, Setoguchi T, Settembre C, Shacka JJ, Shajahan-Haq AN, Shapiro IM, Sharma S, She H, Shen CK, Shen CC, Shen HM, Shen S, Shen W, Sheng R, Sheng X, Sheng ZH, Shepherd TG, Shi J, Shi Q, Shi Q, Shi Y, Shibutani S, Shibuya K, Shidoji Y, Shieh JJ, Shih CM, Shimada Y, Shimizu S, Shin DW, Shinohara ML, Shintani M, Shintani T, Shioi T, Shirabe K, Shiri-Sverdlov R, Shirihai O, Shore GC, Shu CW, Shukla D, Sibirny AA, Sica V, Sigurdson CJ, Sigurdsson EM, Sijwali PS, Sikorska B, Silveira WA, Silvente-Poirot S, Silverman GA, Simak J, Simmet T, Simon AK, Simon HU, Simone C, Simons M, Simonsen A, Singh R, Singh SV, Singh SK, Sinha D, Sinha S, Sinicrope FA, Sirko A, Sirohi K, Sishi BJ, Sittler A, Siu PM, Sivridis E, Skwarska A, Slack R, Slaninová I, Slavov N, Smaili SS, Smalley KS, Smith DR, Soenen SJ, Soleimanpour SA, Solhaug A, Somasundaram K, Son JH, Sonawane A, Song C, Song F, Song HK, Song JX, Song W, Soo KY, Sood AK, Soong TW, Soontornniyomkij V, Sorice M, Sotgia F, Soto-Pantoja DR, Sotthibundhu A, Sousa MJ, Spaink HP, Span PN, Spang A, Sparks JD, Speck PG, Spector SA, Spies CD, Springer W, Clair DS, Stacchiotti A, Staels B, Stang MT, Starczynowski DT, Starokadomskyy P, Steegborn C, Steele JW, Stefanis L, Steffan J, Stellrecht CM, Stenmark H, Stepkowski TM, Stern ST, Stevens C, Stockwell BR, Stoka V, Storchova Z, Stork B, Stratoulias V, Stravopodis DJ, Strnad P, Strohecker AM, Ström AL, Stromhaug P, Stulik J, Su YX, Su Z, Subauste CS, Subramaniam S, Sue CM, Suh SW, Sui X, Sukseree S, Sulzer D, Sun FL, Sun J, Sun J, Sun SY, Sun Y, Sun Y, Sun Y, Sundaramoorthy V, Sung J, Suzuki H, Suzuki K, Suzuki N, Suzuki T, Suzuki YJ, Swanson MS, Swanton C, Swärd K, Swarup G, Sweeney ST, Sylvester PW, Szatmari Z, Szegezdi E, Szlosarek PW, Taegtmeyer H, Tafani M, Taillebourg E, Tait SW, Takacs-Vellai K, Takahashi Y, Takáts S, Takemura G, Takigawa N, Talbot NJ, Tamagno E, Tamburini J, Tan CP, Tan L, Tan ML, Tan M, Tan YJ, Tanaka K, Tanaka M, Tang D, Tang D, Tang G, Tanida I, Tanji K, Tannous BA, Tapia JA, Tasset-Cuevas I, Tatar M, Tavassoly I, Tavernarakis N, Taylor A, Taylor GS, Taylor GA, Taylor JP, Taylor MJ, Tchetina EV, Tee AR, Teixeira-Clerc F, Telang S, Tencomnao T, Teng BB, Teng RJ, Terro F, Tettamanti G, Theiss AL, Theron AE, Thomas KJ, Thomé MP, Thomes PG, Thorburn A, Thorner J, Thum T, Thumm M, Thurston TL, Tian L, Till A, Ting JP, Titorenko VI, Toker L, Toldo S, Tooze SA, Topisirovic I, Torgersen ML, Torosantucci L, Torriglia A, Torrisi MR, Tournier C, Towns R, Trajkovic V, Travassos LH, Triola G, Tripathi DN, Trisciuoglio D, Troncoso R, Trougakos IP, Truttmann AC, Tsai KJ, Tschan MP, Tseng YH, Tsukuba T, Tsung A, Tsvetkov AS, Tu S, Tuan HY, Tucci M, Tumbarello DA, Turk B, Turk V, Turner RF, Tveita AA, Tyagi SC, Ubukata M, Uchiyama Y, Udelnow A, Ueno T, Umekawa M, Umemiya-Shirafuji R, Underwood BR, Ungermann C, Ureshino RP, Ushioda R, Uversky VN, Uzcátegui NL, Vaccari T, Vaccaro MI, Váchová L, Vakifahmetoglu-Norberg H, Valdor R, Valente EM, Vallette F, Valverde AM, Van den Berghe G, Van Den Bosch L, van den Brink GR, van der Goot FG, van der Klei IJ, van der Laan LJ, van Doorn WG, van Egmond M, van Golen KL, Van Kaer L, van Lookeren Campagne M, Vandenabeele P, Vandenberghe W, Vanhorebeek I, Varela-Nieto I, Vasconcelos MH, Vasko R, Vavvas DG, Vega-Naredo I, Velasco G, Velentzas AD, Velentzas PD, Vellai T, Vellenga E, Vendelbo MH, Venkatachalam K, Ventura N, Ventura S, Veras PS, Verdier M, Vertessy BG, Viale A, Vidal M, Vieira HL, Vierstra RD, Vigneswaran N, Vij N, Vila M, Villar M, Villar VH, Villarroya J, Vindis C, Viola G, Viscomi MT, Vitale G, Vogl DT, Voitsekhovskaja OV, von Haefen C, von Schwarzenberg K, Voth DE, Vouret-Craviari V, Vuori K, Vyas JM, Waeber C, Walker CL, Walker MJ, Walter J, Wan L, Wan X, Wang B, Wang C, Wang CY, Wang C, Wang C, Wang C, Wang D, Wang F, Wang F, Wang G, Wang HJ, Wang H, Wang HG, Wang H, Wang HD, Wang J, Wang J, Wang M, Wang MQ, Wang PY, Wang P, Wang RC, Wang S, Wang TF, Wang X, Wang XJ, Wang XW, Wang X, Wang X, Wang Y, Wang Y, Wang Y, Wang YJ, Wang Y, Wang Y, Wang YT, Wang Y, Wang ZN, Wappner P, Ward C, Ward DM, Warnes G, Watada H, Watanabe Y, Watase K, Weaver TE, Weekes CD, Wei J, Weide T, Weihl CC, Weindl G, Weis SN, Wen L, Wen X, Wen Y, Westermann B, Weyand CM, White AR, White E, Whitton JL, Whitworth AJ, Wiels J, Wild F, Wildenberg ME, Wileman T, Wilkinson DS, Wilkinson S, Willbold D, Williams C, Williams K, Williamson PR, Winklhofer KF, Witkin SS, Wohlgemuth SE, Wollert T, Wolvetang EJ, Wong E, Wong GW, Wong RW, Wong VK, Woodcock EA, Wright KL, Wu C, Wu D, Wu GS, Wu J, Wu J, Wu M, Wu M, Wu S, Wu WK, Wu Y, Wu Z, Xavier CP, Xavier RJ, Xia GX, Xia T, Xia W, Xia Y, Xiao H, Xiao J, Xiao S, Xiao W, Xie CM, Xie Z, Xie Z, Xilouri M, Xiong Y, Xu C, Xu C, Xu F, Xu H, Xu H, Xu J, Xu J, Xu J, Xu L, Xu X, Xu Y, Xu Y, Xu ZX, Xu Z, Xue Y, Yamada T, Yamamoto A, Yamanaka K, Yamashina S, Yamashiro S, Yan B, Yan B, Yan X, Yan Z, Yanagi Y, Yang DS, Yang JM, Yang L, Yang M, Yang PM, Yang P, Yang Q, Yang W, Yang WY, Yang X, Yang Y, Yang Y, Yang Z, Yang Z, Yao MC, Yao PJ, Yao X, Yao Z, Yao Z, Yasui LS, Ye M, Yedvobnick B, Yeganeh B, Yeh ES, Yeyati PL, Yi F, Yi L, Yin XM, Yip CK, Yoo YM, Yoo YH, Yoon SY, Yoshida K, Yoshimori T, Young KH, Yu H, Yu JJ, Yu JT, Yu J, Yu L, Yu WH, Yu XF, Yu Z, Yuan J, Yuan ZM, Yue BY, Yue J, Yue Z, Zacks DN, Zacksenhaus E, Zaffaroni N, Zaglia T, Zakeri Z, Zecchini V, Zeng J, Zeng M, Zeng Q, Zervos AS, Zhang DD, Zhang F, Zhang G, Zhang GC, Zhang H, Zhang H, Zhang H, Zhang H, Zhang J, Zhang J, Zhang J, Zhang J, Zhang JP, Zhang L, Zhang L, Zhang L, Zhang L, Zhang MY, Zhang X, Zhang XD, Zhang Y, Zhang Y, Zhang Y, Zhang Y, Zhang Y, Zhao M, Zhao WL, Zhao X, Zhao YG, Zhao Y, Zhao Y, Zhao YX, Zhao Z, Zhao ZJ, Zheng D, Zheng XL, Zheng X, Zhivotovsky B, Zhong Q, Zhou GZ, Zhou G, Zhou H, Zhou SF, Zhou XJ, Zhu H, Zhu H, Zhu WG, Zhu W, Zhu XF, Zhu Y, Zhuang SM, Zhuang X, Ziparo E, Zois CE, Zoladek T, Zong WX, Zorzano A, Zughaier SM. Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition) Autophagy. 2016;12:1–222. doi: 10.1080/15548627.2015.1100356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Komatsu M, Waguri S, Chiba T, Murata S, Iwata J, Tanida I, Ueno T, Koike M, Uchiyama Y, Kominami E, Tanaka K. Loss of autophagy in the central nervous system causes neurodegeneration in mice. Nature. 2006;441:880–884. doi: 10.1038/nature04723. [DOI] [PubMed] [Google Scholar]
- Komatsu M, Waguri S, Koike M, Sou YS, Ueno T, Hara T, Mizushima N, Iwata J, Ezaki J, Murata S, Hamazaki J, Nishito Y, Iemura S, Natsume T, Yanagawa T, Uwayama J, Warabi E, Yoshida H, Ishii T, Kobayashi A, Yamamoto M, Yue Z, Uchiyama Y, Kominami E, Tanaka K. Homeostatic levels of p62 control cytoplasmic inclusion body formation in autophagy-deficient mice. Cell. 2007;131:1149–1163. doi: 10.1016/j.cell.2007.10.035. [DOI] [PubMed] [Google Scholar]
- Korolchuk VI, Menzies FM, Rubinsztein DC. Mechanisms of cross-talk between the ubiquitin-proteasome and autophagy-lysosome systems. FEBS Letters. 2010;584:1393–1398. doi: 10.1016/j.febslet.2009.12.047. [DOI] [PubMed] [Google Scholar]
- Kramer NJ, Carlomagno Y, Zhang YJ, Almeida S, Cook CN, Gendron TF, Prudencio M, Van Blitterswijk M, Belzil V, Couthouis J, Paul JW, Goodman LD, Daughrity L, Chew J, Garrett A, Pregent L, Jansen-West K, Tabassian LJ, Rademakers R, Boylan K, Graff-Radford NR, Josephs KA, Parisi JE, Knopman DS, Petersen RC, Boeve BF, Deng N, Feng Y, Cheng TH, Dickson DW, Cohen SN, Bonini NM, Link CD, Gao FB, Petrucelli L, Gitler AD. Spt4 selectively regulates the expression of C9orf72 sense and antisense mutant transcripts. Science. 2016;353:708–712. doi: 10.1126/science.aaf7791. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kwon I, Xiang S, Kato M, Wu L, Theodoropoulos P, Wang T, Kim J, Yun J, Xie Y, McKnight SL. Poly-dipeptides encoded by the C9orf72 repeats bind nucleoli, impede RNA biogenesis, and kill cells. Science. 2014;345:1139–1145. doi: 10.1126/science.1254917. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Le Ber I, Camuzat A, Guerreiro R, Bouya-Ahmed K, Bras J, Nicolas G, Gabelle A, Didic M, De Septenville A, Millecamps S, Lenglet T, Latouche M, Kabashi E, Campion D, Hannequin D, Hardy J, Brice A, French Clinical and Genetic Research Network on FTD/FTD-ALS SQSTM1 mutations in french patients with frontotemporal dementia or frontotemporal dementia with amyotrophic lateral sclerosis. JAMA Neurology. 2013;70:1403–1410. doi: 10.1001/jamaneurol.2013.3849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee CW, Wilfling F, Ronchi P, Allegretti M, Mosalaganti S, Jentsch S, Beck M, Pfander B. Selective autophagy degrades nuclear pore complexes. Nature Cell Biology. 2020;22:159–166. doi: 10.1038/s41556-019-0459-2. [DOI] [PubMed] [Google Scholar]
- Levine B, Kroemer G. Autophagy in the pathogenesis of disease. Cell. 2008;132:27–42. doi: 10.1016/j.cell.2007.12.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li L, Friedrichsen HJ, Andrews S, Picaud S, Volpon L, Ngeow K, Berridge G, Fischer R, Borden KLB, Filippakopoulos P, Goding CR. A TFEB nuclear export signal integrates amino acid supply and glucose availability. Nature Communications. 2018;9:2685. doi: 10.1038/s41467-018-04849-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin G, Mao D, Bellen HJ. Amyotrophic lateral sclerosis pathogenesis converges on defects in protein homeostasis associated with TDP-43 mislocalization and Proteasome-Mediated degradation overload. Curr Top Dev Bio. 2017;121:111–171. doi: 10.1016/bs.ctdb.2016.07.004. [DOI] [PubMed] [Google Scholar]
- Ling SC, Polymenidou M, Cleveland DW. Converging mechanisms in ALS and FTD: disrupted RNA and protein homeostasis. Neuron. 2013;79:416–438. doi: 10.1016/j.neuron.2013.07.033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y, Pattamatta A, Zu T, Reid T, Bardhi O, Borchelt DR, Yachnis AT, Ranum LP. C9orf72 BAC mouse model with motor deficits and neurodegenerative features of ALS/FTD. Neuron. 2016;90:521–534. doi: 10.1016/j.neuron.2016.04.005. [DOI] [PubMed] [Google Scholar]
- Mackenzie IR, Frick P, Neumann M. The neuropathology associated with repeat expansions in the C9ORF72 gene. Acta Neuropathologica. 2014;127:347–357. doi: 10.1007/s00401-013-1232-4. [DOI] [PubMed] [Google Scholar]
- Mao D, Lin G, Tepe B, Zuo Z, Tan KL, Senturk M, Zhang S, Arenkiel BR, Sardiello M, Bellen HJ. VAMP associated proteins are required for autophagic and lysosomal degradation by promoting a PtdIns4P-mediated endosomal pathway. Autophagy. 2019;15:1214–1233. doi: 10.1080/15548627.2019.1580103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martini-Stoica H, Xu Y, Ballabio A, Zheng H. The Autophagy-Lysosomal pathway in neurodegeneration: a TFEB perspective. Trends in Neurosciences. 2016;39:221–234. doi: 10.1016/j.tins.2016.02.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mauvezin C, Ayala C, Braden CR, Kim J, Neufeld TP. Assays to monitor autophagy in Drosophila. Methods. 2014;68:134–139. doi: 10.1016/j.ymeth.2014.03.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mizielinska S, Gronke S, Niccoli T, Ridler CE, Clayton EL, Devoy A, Moens T, Norona FE, Woollacott IOC, Pietrzyk J, Cleverley K, Nicoll AJ, Pickering-Brown S, Dols J, Cabecinha M, Hendrich O, Fratta P, Fisher EMC, Partridge L, Isaacs AM. C9orf72 repeat expansions cause neurodegeneration in Drosophila through arginine-rich proteins. Science. 2014;345:1192–1194. doi: 10.1126/science.1256800. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mori K, Weng SM, Arzberger T, May S, Rentzsch K, Kremmer E, Schmid B, Kretzschmar HA, Cruts M, Van Broeckhoven C, Haass C, Edbauer D. The C9orf72 GGGGCC repeat is translated into aggregating dipeptide-repeat proteins in FTLD/ALS. Science. 2013;339:1335–1338. doi: 10.1126/science.1232927. [DOI] [PubMed] [Google Scholar]
- O'Rourke JG, Bogdanik L, Muhammad A, Gendron TF, Kim KJ, Austin A, Cady J, Liu EY, Zarrow J, Grant S, Ho R, Bell S, Carmona S, Simpkinson M, Lall D, Wu K, Daughrity L, Dickson DW, Harms MB, Petrucelli L, Lee EB, Lutz CM, Baloh RH. C9orf72 BAC transgenic mice display typical pathologic features of ALS/FTD. Neuron. 2015;88:892–901. doi: 10.1016/j.neuron.2015.10.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ortega JA, Daley EL, Kour S, Samani M, Tellez L, Smith HS, Hall EA, Esengul YT, Tsai Y-H, Gendron TF, Donnelly CJ, Siddique T, Savas JN, Pandey UB, Kiskinis E. Nucleocytoplasmic proteomic analysis uncovers eRF1 and Nonsense-Mediated decay as modifiers of ALS/FTD C9orf72 toxicity. Neuron. 2020;106:90–107. doi: 10.1016/j.neuron.2020.01.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Palmieri M, Impey S, Kang H, di Ronza A, Pelz C, Sardiello M, Ballabio A. Characterization of the CLEAR network reveals an integrated control of cellular clearance pathways. Human Molecular Genetics. 2011;20:3852–3866. doi: 10.1093/hmg/ddr306. [DOI] [PubMed] [Google Scholar]
- Parkinson-Lawrence EJ, Shandala T, Prodoehl M, Plew R, Borlace GN, Brooks DA. Lysosomal storage disease: revealing lysosomal function and physiology. Physiology. 2010;25:102–115. doi: 10.1152/physiol.00041.2009. [DOI] [PubMed] [Google Scholar]
- Parr C, Carzaniga R, Gentleman SM, Van Leuven F, Walter J, Sastre M. Glycogen synthase kinase 3 inhibition promotes lysosomal biogenesis and autophagic degradation of the amyloid-β precursor protein. Molecular and Cellular Biology. 2012;32:4410–4418. doi: 10.1128/MCB.00930-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pircs K, Nagy P, Varga A, Venkei Z, Erdi B, Hegedus K, Juhasz G. Advantages and limitations of different p62-based assays for estimating autophagic activity in Drosophila. PLOS ONE. 2012;7:e44214. doi: 10.1371/journal.pone.0044214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Polito VA, Li H, Martini-Stoica H, Wang B, Yang L, Xu Y, Swartzlander DB, Palmieri M, di Ronza A, Lee VM, Sardiello M, Ballabio A, Zheng H. Selective clearance of aberrant tau proteins and rescue of neurotoxicity by transcription factor EB. EMBO Molecular Medicine. 2014;6:1142–1160. doi: 10.15252/emmm.201303671. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pulipparacharuvil S, Akbar MA, Ray S, Sevrioukov EA, Haberman AS, Rohrer J, Krämer H. Drosophila Vps16A is required for trafficking to lysosomes and biogenesis of pigment granules. Journal of Cell Science. 2005;118:3663–3673. doi: 10.1242/jcs.02502. [DOI] [PubMed] [Google Scholar]
- Ramesh N, Pandey UB. Autophagy dysregulation in ALS: when protein aggregates get out of hand. Frontiers in Molecular Neuroscience. 2017;10:263. doi: 10.3389/fnmol.2017.00263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ritson GP, Custer SK, Freibaum BD, Guinto JB, Geffel D, Moore J, Tang W, Winton MJ, Neumann M, Trojanowski JQ, Lee VM, Forman MS, Taylor JP. TDP-43 mediates degeneration in a novel Drosophila model of disease caused by mutations in VCP/p97. Journal of Neuroscience. 2010;30:7729–7739. doi: 10.1523/JNEUROSCI.5894-09.2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roczniak-Ferguson A, Petit CS, Froehlich F, Qian S, Ky J, Angarola B, Walther TC, Ferguson SM. The transcription factor TFEB links mTORC1 signaling to transcriptional control of lysosome homeostasis. Science Signaling. 2012;5:ra42. doi: 10.1126/scisignal.2002790. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saitoh Y, Fujikake N, Okamoto Y, Popiel HA, Hatanaka Y, Ueyama M, Suzuki M, Gaumer S, Murata M, Wada K, Nagai Y. p62 plays a protective role in the autophagic degradation of polyglutamine protein oligomers in polyglutamine disease model flies. Journal of Biological Chemistry. 2015;290:1442–1453. doi: 10.1074/jbc.M114.590281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sardiello M, Palmieri M, di Ronza A, Medina DL, Valenza M, Gennarino VA, Di Malta C, Donaudy F, Embrione V, Polishchuk RS, Banfi S, Parenti G, Cattaneo E, Ballabio A. A gene network regulating lysosomal biogenesis and function. Science. 2009;325:473–477. doi: 10.1126/science.1174447. [DOI] [PubMed] [Google Scholar]
- Sarkar S, Davies JE, Huang Z, Tunnacliffe A, Rubinsztein DC. Trehalose, a novel mTOR-independent autophagy enhancer, accelerates the clearance of mutant huntingtin and α-Synuclein. Journal of Biological Chemistry. 2007;282:5641–5652. doi: 10.1074/jbc.M609532200. [DOI] [PubMed] [Google Scholar]
- Sasaki S. Autophagy in spinal cord motor neurons in sporadic amyotrophic lateral sclerosis. Journal of Neuropathology & Experimental Neurology. 2011;70:349–359. doi: 10.1097/NEN.0b013e3182160690. [DOI] [PubMed] [Google Scholar]
- Sellier C, Campanari ML, Julie Corbier C, Gaucherot A, Kolb-Cheynel I, Oulad-Abdelghani M, Ruffenach F, Page A, Ciura S, Kabashi E, Charlet-Berguerand N. Loss of C9ORF72 impairs autophagy and synergizes with polyQ Ataxin-2 to induce motor neuron dysfunction and cell death. The EMBO Journal. 2016;35:1276–1297. doi: 10.15252/embj.201593350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Şentürk M, Lin G, Zuo Z, Mao D, Watson E, Mikos AG, Bellen HJ. Ubiquilins regulate autophagic flux through mTOR signalling and lysosomal acidification. Nature Cell Biology. 2019;21:384–396. doi: 10.1038/s41556-019-0281-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Settembre C, Di Malta C, Polito VA, Garcia Arencibia M, Vetrini F, Erdin S, Erdin SU, Huynh T, Medina D, Colella P, Sardiello M, Rubinsztein DC, Ballabio A. TFEB links autophagy to lysosomal biogenesis. Science. 2011;332:1429–1433. doi: 10.1126/science.1204592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi KY, Mori E, Nizami ZF, Lin Y, Kato M, Xiang S, Wu LC, Ding M, Yu Y, Gall JG, McKnight SL. Toxic PRn poly-dipeptides encoded by the C9orf72 repeat expansion block nuclear import and export. PNAS. 2017;114:E1111–E1117. doi: 10.1073/pnas.1620293114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi Y, Lin S, Staats KA, Li Y, Chang WH, Hung ST, Hendricks E, Linares GR, Wang Y, Son EY, Wen X, Kisler K, Wilkinson B, Menendez L, Sugawara T, Woolwine P, Huang M, Cowan MJ, Ge B, Koutsodendris N, Sandor KP, Komberg J, Vangoor VR, Senthilkumar K, Hennes V, Seah C, Nelson AR, Cheng TY, Lee SJ, August PR, Chen JA, Wisniewski N, Hanson-Smith V, Belgard TG, Zhang A, Coba M, Grunseich C, Ward ME, van den Berg LH, Pasterkamp RJ, Trotti D, Zlokovic BV, Ichida JK. Haploinsufficiency leads to neurodegeneration in C9ORF72 ALS/FTD human induced motor neurons. Nature Medicine. 2018;24:313–325. doi: 10.1038/nm.4490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Silvestrini MJ, Johnson JR, Kumar AV, Thakurta TG, Blais K, Neill ZA, Marion SW, St Amand J, Reenan RA, Lapierre LR. Nuclear export inhibition enhances HLH-30/TFEB activity, autophagy, and lifespan. Cell Reports. 2018;23:1915–1921. doi: 10.1016/j.celrep.2018.04.063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Solomon DA, Stepto A, Au WH, Adachi Y, Diaper DC, Hall R, Rekhi A, Boudi A, Tziortzouda P, Lee YB, Smith B, Bridi JC, Spinelli G, Dearlove J, Humphrey DM, Gallo JM, Troakes C, Fanto M, Soller M, Rogelj B, Parsons RB, Shaw CE, Hortobágyi T, Hirth F. A feedback loop between dipeptide-repeat protein, TDP-43 and karyopherin-α mediates C9orf72-related neurodegeneration. Brain. 2018;141:2908–2924. doi: 10.1093/brain/awy241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Song W, Wang F, Savini M, Ake A, di Ronza A, Sardiello M, Segatori L. TFEB regulates lysosomal proteostasis. Human Molecular Genetics. 2013;22:1994–2009. doi: 10.1093/hmg/ddt052. [DOI] [PubMed] [Google Scholar]
- Stewart BA, Atwood HL, Renger JJ, Wang J, Wu C-F. Improved stability of Drosophila larval neuromuscular preparations in haemolymph-like physiological solutions. Journal of Comparative Physiology A. 1994;175:179–191. doi: 10.1007/BF00215114. [DOI] [PubMed] [Google Scholar]
- Sullivan PM, Zhou X, Robins AM, Paushter DH, Kim D, Smolka MB, Hu F. The ALS/FTLD associated protein C9orf72 associates with SMCR8 and WDR41 to regulate the autophagy-lysosome pathway. Acta Neuropathologica Communications. 2016;4:51. doi: 10.1186/s40478-016-0324-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Teyssou E, Takeda T, Lebon V, Boillée S, Doukouré B, Bataillon G, Sazdovitch V, Cazeneuve C, Meininger V, LeGuern E, Salachas F, Seilhean D, Millecamps S. Mutations in SQSTM1 encoding p62 in amyotrophic lateral sclerosis: genetics and neuropathology. Acta Neuropathologica. 2013;125:511–522. doi: 10.1007/s00401-013-1090-0. [DOI] [PubMed] [Google Scholar]
- Thomas M, Alegre-Abarrategui J, Wade-Martins R. RNA dysfunction and aggrephagy at the centre of an amyotrophic lateral sclerosis/frontotemporal dementia disease continuum. Brain. 2013;136:1345–1360. doi: 10.1093/brain/awt030. [DOI] [PubMed] [Google Scholar]
- Torra A, Parent A, Cuadros T, Rodríguez-Galván B, Ruiz-Bronchal E, Ballabio A, Bortolozzi A, Vila M, Bové J. Overexpression of TFEB drives a pleiotropic neurotrophic effect and prevents parkinson's Disease-Related Neurodegeneration. Molecular Therapy. 2018;26:1552–1567. doi: 10.1016/j.ymthe.2018.02.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tran H, Almeida S, Moore J, Gendron TF, Chalasani U, Lu Y, Du X, Nickerson JA, Petrucelli L, Weng Z, Gao FB. Differential toxicity of nuclear RNA foci versus dipeptide repeat proteins in a Drosophila Model of C9ORF72 FTD/ALS. Neuron. 2015;87:1207–1214. doi: 10.1016/j.neuron.2015.09.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ugolino J, Ji YJ, Conchina K, Chu J, Nirujogi RS, Pandey A, Brady NR, Hamacher-Brady A, Wang J. Loss of C9orf72 enhances autophagic activity via deregulated mTOR and TFEB signaling. PLOS Genetics. 2016;12:e1006443. doi: 10.1371/journal.pgen.1006443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vodicka P, Chase K, Iuliano M, Tousley A, Valentine DT, Sapp E, Kegel-Gleason KB, Sena-Esteves M, Aronin N, DiFiglia M. Autophagy activation by transcription factor EB (TFEB) in striatum of HDQ175/Q7 mice. Journal of Huntington's Disease. 2016;5:249–260. doi: 10.3233/JHD-160211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walker C, Herranz-Martin S, Karyka E, Liao C, Lewis K, Elsayed W, Lukashchuk V, Chiang SC, Ray S, Mulcahy PJ, Jurga M, Tsagakis I, Iannitti T, Chandran J, Coldicott I, De Vos KJ, Hassan MK, Higginbottom A, Shaw PJ, Hautbergue GM, Azzouz M, El-Khamisy SF. C9orf72 expansion disrupts ATM-mediated chromosomal break repair. Nature Neuroscience. 2017;20:1225–1235. doi: 10.1038/nn.4604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang H, Wang R, Xu S, Lakshmana MK. Transcription factor EB is selectively reduced in the nuclear fractions of Alzheimer’s and Amyotrophic Lateral Sclerosis Brains. Neuroscience Journal. 2016;2016:1–8. doi: 10.1155/2016/4732837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weaver TE, Na CL, Stahlman M. Biogenesis of lamellar bodies, lysosome-related organelles involved in storage and secretion of pulmonary surfactant. Seminars in Cell & Developmental Biology. 2002;13:263–270. doi: 10.1016/S1084952102000551. [DOI] [PubMed] [Google Scholar]
- Webster CP, Smith EF, Bauer CS, Moller A, Hautbergue GM, Ferraiuolo L, Myszczynska MA, Higginbottom A, Walsh MJ, Whitworth AJ, Kaspar BK, Meyer K, Shaw PJ, Grierson AJ, De Vos KJ. The C9orf72 protein interacts with Rab1a and the ULK1 complex to regulate initiation of autophagy. The EMBO Journal. 2016;35:1656–1676. doi: 10.15252/embj.201694401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wen X, Tan W, Westergard T, Krishnamurthy K, Markandaiah SS, Shi Y, Lin S, Shneider NA, Monaghan J, Pandey UB, Pasinelli P, Ichida JK, Trotti D. Antisense Proline-Arginine RAN dipeptides linked to C9ORF72-ALS/FTD form toxic nuclear aggregates that initiate in vitro and in vivo neuronal death. Neuron. 2014;84:1213–1225. doi: 10.1016/j.neuron.2014.12.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woerner AC, Frottin F, Hornburg D, Feng LR, Meissner F, Patra M, Tatzelt J, Mann M, Winklhofer KF, Hartl FU, Hipp MS. Cytoplasmic protein aggregates interfere with nucleocytoplasmic transport of protein and RNA. Science. 2016;351:173–176. doi: 10.1126/science.aad2033. [DOI] [PubMed] [Google Scholar]
- Xu Z, Poidevin M, Li X, Li Y, Shu L, Nelson DL, Li H, Hales CM, Gearing M, Wingo TS, Jin P. Expanded GGGGCC repeat RNA associated with amyotrophic lateral sclerosis and frontotemporal dementia causes neurodegeneration. PNAS. 2013;110:7778–7783. doi: 10.1073/pnas.1219643110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang M, Liang C, Swaminathan K, Herrlinger S, Lai F, Shiekhattar R, Chen JF. A C9ORF72/SMCR8-containing complex regulates ULK1 and plays a dual role in autophagy. Science Advances. 2016;2:e1601167. doi: 10.1126/sciadv.1601167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang K, Donnelly CJ, Haeusler AR, Grima JC, Machamer JB, Steinwald P, Daley EL, Miller SJ, Cunningham KM, Vidensky S, Gupta S, Thomas MA, Hong I, Chiu SL, Huganir RL, Ostrow LW, Matunis MJ, Wang J, Sattler R, Lloyd TE, Rothstein JD. The C9orf72 repeat expansion disrupts nucleocytoplasmic transport. Nature. 2015a;525:56–61. doi: 10.1038/nature14973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang T, Zhou Q, Ogmundsdottir MH, Möller K, Siddaway R, Larue L, Hsing M, Kong SW, Goding CR, Palsson A, Steingrimsson E, Pignoni F. Mitf is a master regulator of the v-ATPase, forming a control module for cellular homeostasis with v-ATPase and TORC1. Journal of Cell Science. 2015b;128:2938–2950. doi: 10.1242/jcs.173807. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang YJ, Gendron TF, Grima JC, Sasaguri H, Jansen-West K, Xu YF, Katzman RB, Gass J, Murray ME, Shinohara M, Lin WL, Garrett A, Stankowski JN, Daughrity L, Tong J, Perkerson EA, Yue M, Chew J, Castanedes-Casey M, Kurti A, Wang ZS, Liesinger AM, Baker JD, Jiang J, Lagier-Tourenne C, Edbauer D, Cleveland DW, Rademakers R, Boylan KB, Bu G, Link CD, Dickey CA, Rothstein JD, Dickson DW, Fryer JD, Petrucelli L. C9ORF72 poly(GA) aggregates sequester and impair HR23 and nucleocytoplasmic transport proteins. Nature Neuroscience. 2016;19:668–677. doi: 10.1038/nn.4272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang K, Daigle JG, Cunningham KM, Coyne AN, Ruan K, Grima JC, Bowen KE, Wadhwa H, Yang P, Rigo F, Taylor JP, Gitler AD, Rothstein JD, Lloyd TE. Stress granule assembly disrupts nucleocytoplasmic transport. Cell. 2018;173:958–971. doi: 10.1016/j.cell.2018.03.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu Q, Jiang J, Gendron TF, McAlonis-Downes M, Jiang L, Taylor A, Diaz Garcia S, Ghosh Dastidar S, Rodriguez MJ, King P, Zhang Y, La Spada AR, Xu H, Petrucelli L, Ravits J, Da Cruz S, Lagier-Tourenne C, Cleveland DW. Reduced C9ORF72 function exacerbates gain of toxicity from ALS/FTD-causing repeat expansion in C9orf72. Nature Neuroscience. 2020;23:615–624. doi: 10.1038/s41593-020-0619-5. [DOI] [PMC free article] [PubMed] [Google Scholar]