Abstract
Background
Idiopathic pulmonary fibrosis (IPF) is the most common of short telomere phenotypes. Familial clustering of IPF is common, but the genetic basis remains unknown in more than one-half of cases. We identified a 65-year-old man with familial IPF, short telomere length, and low telomerase RNA levels. He was diagnosed with a short telomere syndrome after developing hematologic complications post-lung transplantation, but no mutations were identified in a clinical testing pipeline.
Research Question
What is the molecular basis underlying the familial IPF and low telomerase RNA levels in this patient?
Study Design and Methods
We analyzed whole-genome sequence data and performed functional molecular studies on cells derived from the patient and his family.
Results
We identified a previously unreported synonymous variant c.942G>A p.K314K in DKC1, the gene encoding the dyskerin ribonucleoprotein, which is required for telomerase RNA biogenesis. The mutation created a competing de novo exonic splicing enhancer, and the misspliced product was degraded by nonsense-mediated decay causing an overall dyskerin deficiency in mutation carriers. In silico tools identified other rare silent DKC1 variants that warrant functional evaluation if found in patients with short telomere-mediated disease.
Interpretation
Our data point to silent mutation in telomere maintenance genes as a mechanism of familial pulmonary fibrosis. In contrast to DKC1 missense mutations, which primarily manifest in children as dyskeratosis congenita, hypomorphic mutations affecting dyskerin levels likely have a predilection to presenting in adults as pulmonary fibrosis.
Key Words: bone marrow failure, lung transplantation, telomerase
Abbreviations: IPF, idiopathic pulmonary fibrosis; LCL, lymphoblastoid cell line; MAF, minor allele frequency; NMD, nonsense-mediated decay; PCR, polymerase chain reaction; RT-PCR, reverse transcription-polymerase chain reaction
FOR EDITORIAL COMMENT, SEE PAGE 2249
Idiopathic pulmonary fibrosis (IPF) is a progressive, age-related disease. Several lines of evidence have linked short telomere length to genetic susceptibility to IPF.1,2 Approximately 30% to 35% of patients with familial IPF carry germline mutations in a telomerase or other telomere maintenance genes.3, 4, 5, 6, 7, 8, 9, 10 Moreover, at least one-half of patients with IPF, both with isolated and familial cases, have short telomere lengths that fall in the lowest decile of the population.2,11 The short telomere length in these patients is genetically determined, systemic, and has been documented in multiple leukocyte lineages, alveolar epithelial cells, and lung fibroblasts.2,12 At least 10% of patients with IPF show features of extrapulmonary telomere-mediated disease.2,13 These include varying degrees of bone marrow failure, immunodeficiency, and liver disease including hepatopulmonary syndrome.13, 14, 15, 16 In the setting of lung transplantation, where immunosuppressive regimens challenge bone marrow and immune reserves, cytopenias and infectious complications, especially cytomegalovirus, are common and can unmask the diagnosis of telomere-mediated IPF.14,17,18 The diagnosis of telomere-mediated IPF, either clinically or as identified by combining telomere length and DNA sequencing, has thus been increasingly appreciated to have implications for patient care in lung transplantation as well as other settings.19
Among the known familial pulmonary fibrosis genes, the most commonly mutated group affects the integrity, biogenesis, and catalytic functions of the RNA component of the telomerase enzyme. Telomerase RNA (TR, also known as TERC) is an essential core component of the telomerase enzyme; it provides the template for telomere repeat addition to the reverse transcriptase TERT. In addition to the TR gene itself, four of the eight known telomere-related IPF genes play a role in TR biogenesis. DKC1, which encodes dyskerin, is required for TR posttranscriptional stability,20 while NAF1 encodes a ribonucleoprotein that is required for TR trafficking and eventual assembly with TERT.9,21 Mutations in ZCCHC8 and PARN disrupt TR 3′-end maturation and processing of the nascent and adenylated TR, respectively.10,22 We previously identified a number of families with low TR levels who have no obvious mutations in the known genes,9 suggesting there are yet undiscovered mechanisms that compromise TR integrity. Here we show that a “silent” mutation in DKC1, which does not affect its protein sequence, manifests as familial IPF by compromising dyskerin levels through missplicing.
Materials and Methods
Subjects and Study Approval
Subjects were recruited as part of the Johns Hopkins Telomere Syndrome Registry as previously described.15,23 They were assigned the short telomere syndrome diagnosis if they had short telomere length with classic features of a familial short telomere syndrome (eg, history of aplastic anemia and IPF), or had classic short telomere syndrome features with abnormally low telomerase RNA levels even in the absence of a family history.24 The study was approved by the Johns Hopkins Medicine Institutional Review Board, and all the subjects gave written informed consent. Lymphoblastoid cell lines (LCLs) were generated by Epstein-Barr virus transduction as previously described.25
Telomere Length Measurement
Telomere length was measured in peripheral blood leukocytes by flow cytometry and fluorescence in situ hybridization (flowFISH) as previously described.26,27
Quantitative Reverse Transcription-Polymerase Chain Reaction
Quantitative reverse transcription-polymerase chain reaction (qRT-PCR) was performed by the SYBR Green method, random hexamer-primed cDNA, and TR-specific primers with standards as previously detailed.9 DKC1 (RefSeq ID: NM_001363) cDNA was amplified with this primer pair: exon 10-F (5′-atatgaggacggcattgagg) and exon 11-R (5′-ccatggtcgcaggtagagat). Both TR and DKC1 levels were normalized relative to ARF3. Reactions were performed in triplicate, and results were replicated at least twice for all data shown, each time from an independent RNA isolation.
Whole-Genome and Sanger Sequencing
Whole-genome sequencing was performed on blood-derived genomic DNA as previously described.10 Briefly, 150 paired-end sequencing was performed at Macrogen, using a HiSeq X platform (Illumina). Reads were aligned and called relative to GRCh37/hg19, using Isaac aligner (version 01.15.02.08) and variant caller (version 2.0.13), respectively. The mean depth of genome coverage for the proband was 88× with 98.5% covered at 30×. ANNOVAR-annotated VCF files were generated and filtered, using PhenoDB,28 on the basis of mutation frequency and predicted effect on protein sequence (ie, exonic or splicing). Nonsynonymous variants that were absent in dbSNP version 129 and version 131 and had a minor allele frequency (MAF) < 0.0001 in ESP650029 and 1000 Genomes30 were examined further if they additionally had an MAF < 0.005 in ExAC. Because these filtering criteria did not yield any obvious candidates, we manually queried BAM files of the five familial IPF TR-associated genes (Integrative Genomics Viewer): TR itself, DKC1, PARN, NAF1, and ZCCHC8 for other structural anomalies or synonymous variants. Segregation for the DKC1 candidate variant was performed by Sanger sequencing. Whole-exome sequence data were queried for silent mutations in other genes and analyzed as previously.5
In Silico Splice Prediction
To evaluate the impact of rare DKC1 synonymous variants on splicing, we focused on rare variants defined as having an MAF < 0.00002 in the Genome Aggregation Database (gnomAD).31 Splice predictions were performed with Human Splicing Finder (HSF version 3.1).32 Variants predicted to affect less than 25% of transcripts were considered likely benign in this context; the remaining variants were annotated as potentially pathogenic (e-Table 1). Variants with opposing effects or multiple predicted outcomes are also annotated in e-Table 1.
Immunoblot for Dyskerin
Immunoblotting was performed with the LI-COR system as described.9 Briefly, total protein from LCLs was isolated in RIPA buffer (Cell Signaling Technology) supplemented with cOmplete mini protease inhibitors (Roche), and lysates were quantified with a Pierce BCA protein assay kit (Thermo Fisher Scientific). Using the NuPAGE sodium dodecyl sulfate-polyacrylamide gel system, 15 μg was resolved (3%-8% Tris-acetate gel) with Tris-acetate running buffer (150 V for 1 h) and transferred, using the iBlot 2 dry blotting system (20 V for 7 min) (Thermo Fisher Scientific). After blocking (Odyssey blocking buffer, 2 h at room temperature), the following antibodies were used: dyskerin (rabbit, sc-48794, 1:500 [Santa Cruz Biotechnology], overnight at 4°C) and then actin (mouse, ab8226, 1:3,000 [Abcam], for 2 h). Membranes were incubated with anti-rabbit or anti-mouse IRDye secondary antibodies (IR680 or IR800, donkey, at 1:10,000), which were used for visualization on an Odyssey scanner, and quantification was done with ImageJ.33
Nonsense-Mediated Decay Analysis
We used cycloheximide to inhibit nonsense-mediated decay. Two million LCLs were incubated with cycloheximide (50 μg/mL; Cell Signaling) diluted in 2 mL of dimethyl sulfoxide-containing medium for 4 h. RNA was isolated and cDNA was prepared for DKC1 RT-PCR, using exon 9-F (5′-tggtgacaatgcatgatgtgc) and exon 10-R (5′-tgcttctcctttggtggtga). PCR products were resolved on a Novex 10% TBE (Tris-borate-EDTA) gel. DNA fragments were then gel isolated by the “crush and soak” method, using acrylamide gel elution buffer (0.5 M ammonium acetate, 1 mM EDTA, pH 8.0, and 0.1% [w/v] sodium dodecyl sulfate) at 55°C for 2 h. The DNA was then ethanol precipitated and resuspended. To obtain sufficient PCR product for sequencing, the eluted DNA was reamplified, as above, and then Sanger sequenced.
X-Inactivation/HUMARA Assay
X-inactivation analysis was performed using the HUMARA (human androgen receptor gene) assay.34,35 Briefly, genomic DNA was digested with a methylation-sensitive restriction enzyme, HpaII. PCR amplification was then performed with an FAM-labeled forward primer and an unlabeled reverse primer. Allele size was quantified on an ABI genome analyzer.
Results
Patient With Classic Short Telomere Syndrome Features and Familial Pulmonary Fibrosis
The index patient was a 65-year-old man with familial IPF; he was diagnosed with IPF at age 58 and had a history of multiple nonmelanoma skin cancers and antecedent bone marrow failure associated with mild thrombocytopenia (Fig 1A). At the time of diagnosis, he reported having two male relatives with pulmonary fibrosis (Fig 1A). The patient underwent single lung transplantation; his posttransplant course was complicated by cytopenias, a persistent transfusion requirement, invasive fungal infection, and a need for renal replacement therapy. His immediate posttransplant course was reported in a series that described the recurrent posttransplant complications in patients with short telomere IPF.17 Subsequent to that report, he underwent a second lung transplantation, but continued to suffer from recurrent infections and died of sepsis 23 months after the initial transplant. On the basis of his presentation and family history, the diagnosis of a short telomere syndrome was recognized. Peripheral blood testing confirmed that his telomere length fell below the first age-adjusted percentile (Fig 1B), but no mutations were identified during genetic evaluation in a clinical testing pipeline.
Figure 1.
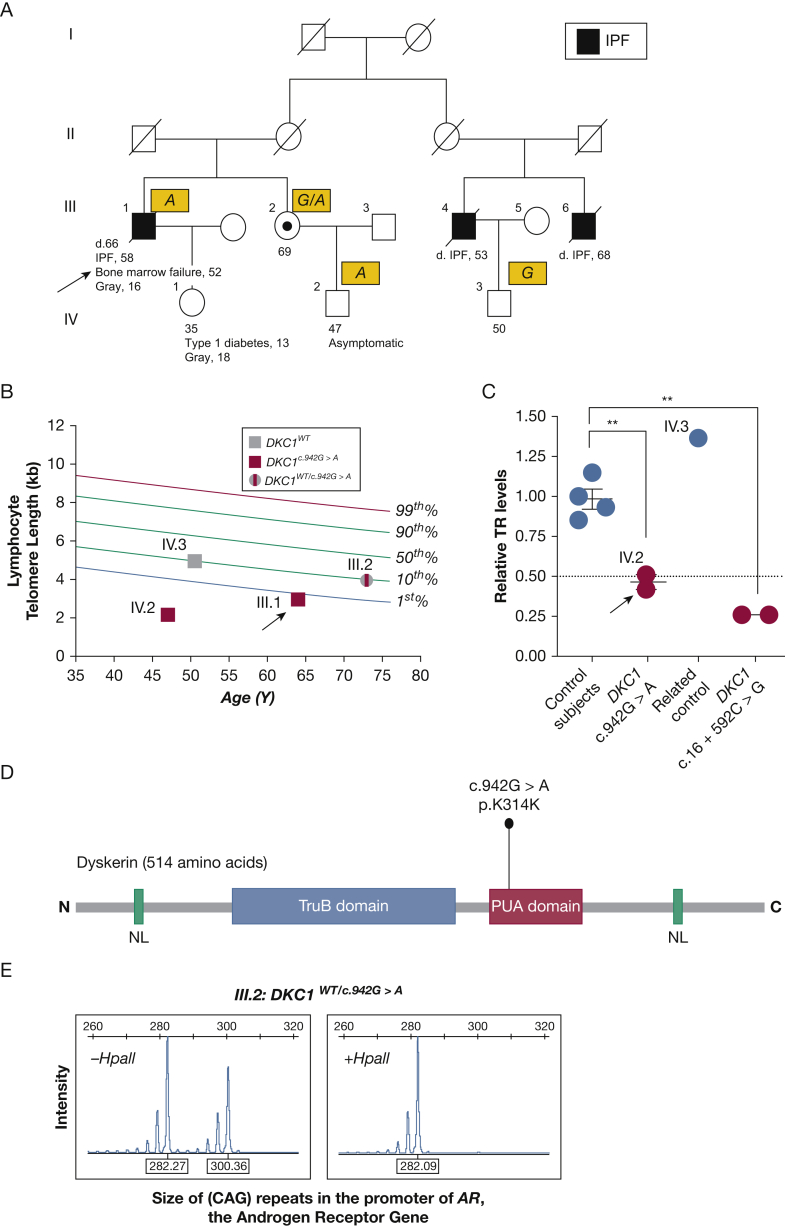
Rare synonymous germline mutation in DKC1 segregates with idiopathic pulmonary fibrosis (IPF), short telomere length, and low telomerase RNA (TR) levels. A, IPF proband (arrow) and family pedigree, with numbers below indicating age at assessment or diagnosis. “d.” refers to age at death from IPF. The yellow boxes contain the genotype at position 153,999,060 on chromosome X in the GRCh37/hg19 genome assembly. Telogram shows age-adjusted lymphocyte telomere length by flow cytometry and fluorescence in situ hybridization (flowFISH) in the proband and family [arrow and pedigree identifiers as in (A)]. C, Relative TR levels measured by quantitative real-time PCR in lymphoblastoid cell lines [arrow and pedigree identifiers as in (A)]. Each point refers to a control- or patient-derived sample as labeled. Error bars represent SEM and asterisks (∗∗) refer to P < .01 (two-sided, Student t-test). D, Scheme of dyskerin protein with known domains including the nuclear localization signal. The putative position of the DKC1 variant, if it were successfully translated, is near the end of the protein. E, Pherogram shows skewed X-chromosome inactivation pattern in blood-derived genomic DNA of proband’s sister, using HUMARA. It shows the size of CAG repeats in the promoter of the androgen receptor (AR) gene. Allele size with and without digestion with methylation-sensitive restriction endonuclease HpaII is shown. Only one allele is detectable after digestion, consistent with 100% skewing. HUMARA = human androgen receptor gene; NL = nuclear localization; WT = wild type.
DKC1 c.942G>A Segregates With Low TR and Short Telomere Length
To narrow the mutant pathway in this patient, we quantified TR levels in his cells, using real-time PCR, and found he had abnormally low TR levels, near 50% of control levels. This abnormality was similar to patients with short telomere IPF and patients with emphysema we had studied, who were subsequently found to carry deleterious mutations in NAF1 and ZCCHC89,10 (Fig 1C). We thus performed whole-genome sequencing and focused our analysis on genes that affect the biogenesis, processing, and localization of TR, but we found no nonsynonymous variants that fulfilled the preset criteria. We then manually examined the whole-genome data for intronic, synonymous, structural, and regulatory mutations in the five TR-related IPF genes including TR itself. One candidate in the X-chromosome DKC1 gene stood out. It fell in exon 10: c.942G>A; this variant predicted a silent protein substitution, p.K314K (Fig 1D). It was a compelling candidate because it had not been reported in 141,456 individuals in gnomAD and was predicted to create a de novo exonic splicing enhancer in silico. Moreover, the patient’s sister, who was a heterozygous carrier of this mutation, showed 100% skewed X-inactivation (Fig 1E). To directly test the pathogenicity of this variant, we recruited additional family members. We identified one asymptomatic male relative who carried the same mutation; he also had abnormally low TR levels and short telomere length, below the first age-adjusted percentile (Figs 1A-1C). In contrast, another male relative who was a noncarrier of the mutation had intact TR levels and normal telomere length (Figs 1A-1C). These data supported the IPF inheritance in this family being X-linked, and this rare silent mutation being potentially pathogenic.
DKC1 Mutation Alters Splicing and Causes Nonsense-Mediated Decay
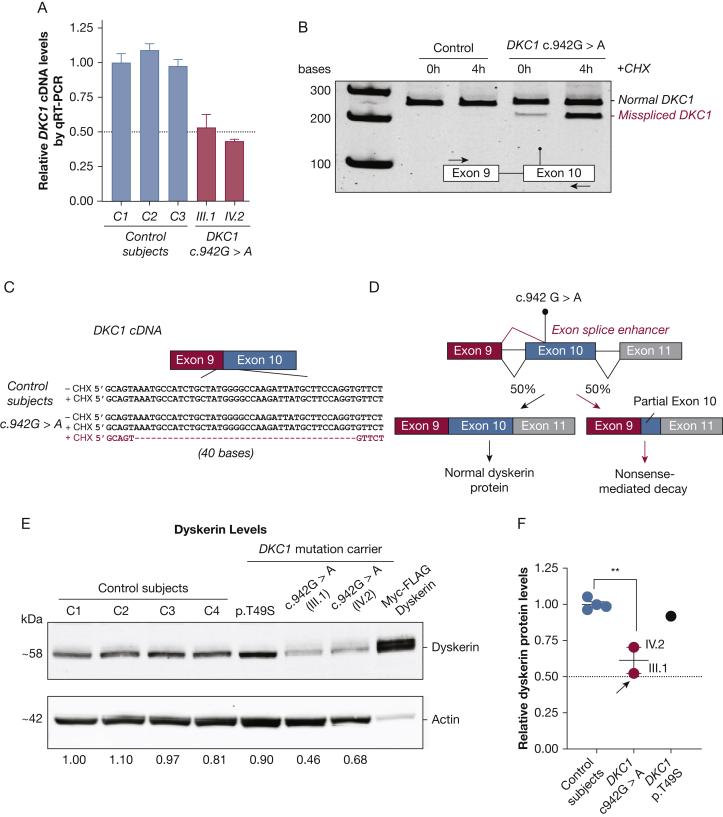
Some synonymous variants affect mRNA splicing,36 and we tested this hypothesis by performing RT-PCR on DKC1 using primers flanking the c.942G>A variant, but we could not detect an aberrant product in patient cells even though DKC1 mRNA levels were low (Fig 2A). Because in silico analysis predicted altered splicing, we tested whether the misspliced product may be degraded by nonsense-mediated decay (NMD). We exposed control- and proband-derived LCLs to cycloheximide, an inhibitor of NMD, and resolved the RT-PCR products. Under these conditions, we identified an aberrant product in the proband that was not seen in control cells (Fig 2B). It included a 40-base pair deletion in exon 10 (Figs 2C and 2D), consistent with c.942G>A creating a de novo exonic splicing enhancer. The new site competed with the canonical splice junction sequence at the native intron-exon boundary and led to an aberrant RNA that had similar abundance to the normally spliced product (Fig 2B). The aberrant product was predicted to encode a premature stop codon at amino acid 336, but the truncated product was not detected by immunoblot (Figs 2E and 2F). Indeed, immunoblotting for dyskerin showed that total levels were low in cells from mutation carriers, near 50% of healthy control cells, and in contrast to DKC1 missense mutation carrier, protein levels were intact (Figs 2E and 2F). These data establish that c.942G>A mutation impairs dyskerin abundance through NMD; the dyskerin insufficiency in turn leads to TR insufficiency.
Figure 2.
Synonymous DKC1 c.942G>A variant causes aberrant mRNA splicing and reduced dyskerin protein levels. A, Relative DKC1 mRNA levels measured by quantitative real-time polymerase chain reaction (qRT-PCR) in lymphoblastoid cell lines (LCLs), using primers in exons 10 and 11 downstream of the mutation. Error bars represent SEM and P values are two-sided, calculated by Student t-test. This experiment was independently replicated twice. B, DKC1 RT-PCR product resolved on a polyacrylamide gel after 0 or 4 h of cycloheximide treatment to inhibit nonsense-mediated decay in LCLs. The location of primers relative to the mutation (lollipop) is indicated in the embedded diagram. C, Sanger sequencing results of excised bands in (B) reveal an aberrant 40-base pair out-of-frame deletion in the proband. D, A diagram showing DKC1 c.942G>A impact on splicing. E, Immunoblot of dyskerin protein in LCLs from four healthy control subjects (C1 through C4) and a patient with a missense T49S DKC1 mutation with normal protein levels compared with the reduced dyskerin levels in the proband and the affected nephew (labeled with pedigree identifier from Fig 1A). Quantification of dyskerin levels relative to actin control is shown. Lane 8 contains protein extract from 293FT cells transfected with DKC1-MycFLAG tag to show specificity. F, Quantitation from three independently harvested protein lysates shows decreased dyskerin protein levels in male mutation carriers compared with control subjects and a T49S DKC1 mutant. Means were compared by Student t-test. Error bars represent SEM and P values are two-sided. cDNA = complementary DNA; CHX = cycloheximide.
Other Rare “Silent DKC1 Mutations” Are Candidates for Familial Short Telomere Syndromes
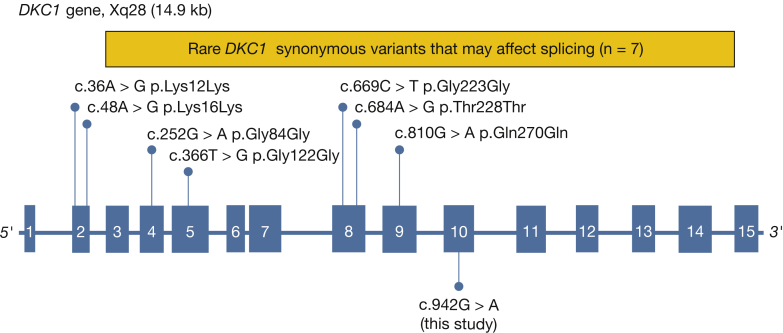
To test if other silent variants in telomere genes may explain the genetic basis in other uncharacterized families, we screened 26 unrelated short telomere syndrome probands for rare synonymous variants in TERT, RTEL1, and PARN in whole-exome and genome data but found no alterations. These data were consistent with this mechanism being rare and highlight the locus heterogeneity and diverse mechanisms that underlie the genetic basis of telomere-mediated diseases including IPF.10,37 Because the culprit variant we identified had been missed in a clinical pipeline, we queried in silico tools to test if other silent single-nucleotide variants may affect DKC1 splicing. Among the reported synonymous DKC1 variants in gnomAD at the time (141,456 individuals: 125,748 exomes and 15,708 genomes), 57 were rare and had a frequency similar to that of mutations seen in familial IPF, that is, less than 1 in 50,000 individuals (minor allele frequency < 0.00002 in male patients), and 35 of these had a single predicted interpretation of either “wild type” or “potentially pathogenic” (e-Table 1). Among these 35, seven variants were predicted to alter more than 25% of transcripts relative to wild-type splicing (Fig 3). Given the sensitivity of TR stability to dyskerin integrity,20 these variants, and undoubtedly others in DKC1, as well as other telomere-related genes, could be considered candidates for further functional and/or genetic characterization if identified in patients with short telomere syndrome features.
Figure 3.
Schematic map of disease-causing DKC1 splicing mutations and synonymous variants predicted to affect splicing. Scheme of genomic DKC1 locus showing the positions of potentially splice-altering synonymous variants identified from the Genome Aggregation Database (above the gene axis) at the time of query relative to the numbered exons. Variants shown fulfill the criteria of in silico prediction of > 25% from wild-type based on Human Splicing Finder (HSF) and are summarized from e-Table 1.
Discussion
We report, to our knowledge, a first case of familial IPF caused by a silent mutation in a telomerase-related gene. Our data further underscore the causal link between telomere dysfunction and genetic susceptibility to familial IPF, as well as the specificity of the clinical IPF-bone marrow failure co-occurrence for germline defects in telomere maintenance.13 Although the synonymous mutation we identified was not detected in a clinical pipeline, the short telomere syndrome diagnosis was initially confirmed by telomere length testing. The clinical presentation of this patient raises a contemporary question related to the indications for genetic and molecular testing in patients with IPF. The proband we studied had a history of thrombocytopenia predating his lung transplantation, but he was diagnosed in the post-lung transplantation period after developing a transfusion requirement. At the time of his presentation, clinical testing for telomere length and genetic studies were in their early stages of clinical use and the natural history of post-lung transplant complications had not yet been well-delineated. In recent years, limited bone marrow reserves and T cell immunodeficiency in patients with short telomere IPF have been well-documented in numerous settings.14,17,18 Our group has thus suggested that attenuating immunosuppression in the lung transplant setting may avert some complications without increasing the risk of graft rejection.14 Such an approach has however not yet been systematically studied. The case we present here raises a question as to the indications of telomere length and genetic testing in pre-transplant evaluations, in at least the subset of patients with IPF with overt short telomere syndrome features, given that it may inform the risk of complications and identify strategies to avert them.
Dyskerin is responsible for the stability and biogenesis of hundreds of box H/ACA motif-containing noncoding RNAs including TR.38 It is also an essential pseudouridylase catalyzing uridine to pseudouridine residues of ribosomal and other RNAs.38 Mutations in DKC1 were initially identified in male patients with the genodermatosis dyskeratosis congenita.39 Nearly all the mutations reported since are missense mutations and associated with a clinical syndrome of acquired mucocutaneous abnormalities in children that define dyskeratosis congenita.40,41 However, in the family we report here, there were no features of dyskeratosis congenita and the initial presentation in this family was pulmonary disease. It may therefore be that hypomorphic mutations that affect dyskerin levels but retain the dyskerin sequence are more likely to be associated with adult-onset disease. Indeed, dyskerin missense mutations that retain protein stability manifest in children,5,35 whereas patients with mutations that disrupt dyskerin levels, via intronic or, as we report here, exonic splicing mutations, are diagnosed in adulthood.35,36 Consistent with this framework, a missense mutation at the same residue, c.941A>G p.K314R, was previously reported in pediatric-onset severe disease.42 The heterogeneity of clinical presentations associated with DKC1 mutations may thus be explained in part by whether a mutation compromises the protein sequence or retains the sequence but affects abundance.
Conclusions
We report on a silent mutation that causes missplicing by creating a competing splice enhancer. In other Mendelian disorders, synonymous mutations have been shown to affect RNA structure, translation initiation, and the rate of translation.43 Our data point to silent mutations in telomere-related genes as a novel yet rare mechanism of familial IPF. They highlight the importance of considering this class of variants in the clinical diagnostic pipeline of patients thought to have short telomere syndromes.
Acknowledgments
Author contributions: V. G. identified the causal mutation, designed and performed the functional studies, and wrote the manuscript with M. A. S. E. S. identified the proband in a screen for patients with low TR levels and performed the initial whole-genome sequence analysis. S. A. analyzed clinical data. M. A. evaluated the family, oversaw the project, and wrote the manuscript with V. G.
Financial/nonfinancial disclosures: None declared.
Role of sponsors: The sponsor had no role in the design of the study, the collection and analysis of the data, or the preparation of the manuscript.
Other contributions: The authors are grateful to all the patients who participated in this research, and to their family members. The authors are grateful for helpful comments from Armanios laboratory members and for assistance from Nuria Amat with the X-inactivation experiment. The authors appreciate support from the Johns Hopkins Genetic Resource Core Facility team.
Additional information: The e-Table can be found in the Supplemental Materials section of the online article.
Footnotes
FUNDING/SUPPORT: The authors acknowledge support from NIH grants RO1 CA225027 and RO1 HL119476 and funding from the Williams, Commonwealth, and S&R Foundations. M. A. acknowledges an award in the name of Mrs. P. Godrej. V. G. was supported by NIH T32 HL007534 and F32 HL142207.
Supplementary Data
References
- 1.Armanios M. Telomerase and idiopathic pulmonary fibrosis. Mutat Res. 2012;730(1-2):52–58. doi: 10.1016/j.mrfmmm.2011.10.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Alder J.K., Chen J.J., Lancaster L. Short telomeres are a risk factor for idiopathic pulmonary fibrosis. Proc Natl Acad Sci U S A. 2008;105(35):13051–13056. doi: 10.1073/pnas.0804280105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Armanios M.Y., Chen J.J., Cogan J.D. Telomerase mutations in families with idiopathic pulmonary fibrosis. N Engl J Med. 2007;356(13):1317–1326. doi: 10.1056/NEJMoa066157. [DOI] [PubMed] [Google Scholar]
- 4.Tsakiri K.D., Cronkhite J.T., Kuan P.J. Adult-onset pulmonary fibrosis caused by mutations in telomerase. Proc Natl Acad Sci U S A. 2007;104(18):7552–7557. doi: 10.1073/pnas.0701009104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Alder J.K., Parry E.M., Yegnasubramanian S. Telomere phenotypes in females with heterozygous mutations in the dyskeratosis congenita 1 (DKC1) gene. Hum Mutat. 2013;34(11):1481–1485. doi: 10.1002/humu.22397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Kropski J.A., Mitchell D.B., Markin C. A novel dyskerin (DKC1) mutation is associated with familial interstitial pneumonia. Chest. 2014;146(1):e1–e7. doi: 10.1378/chest.13-2224. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Cogan J.D., Kropski J.A., Zhao M. Rare variants in RTEL1 are associated with familial interstitial pneumonia. Am J Respir Crit Care Med. 2015;191(6):646–655. doi: 10.1164/rccm.201408-1510OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Stuart B.D., Choi J., Zaidi S. Exome sequencing links mutations in PARN and RTEL1 with familial pulmonary fibrosis and telomere shortening. Nat Genet. 2015;47(5):512–517. doi: 10.1038/ng.3278. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Stanley S.E., Gable D.L., Wagner C.L. Loss-of-function mutations in the RNA biogenesis factor NAF1 predispose to pulmonary fibrosis-emphysema. Sci Transl Med. 2016;8(351):351ra107. doi: 10.1126/scitranslmed.aaf7837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Gable D.L., Gaysinskaya V., Atik C.C. ZCCHC8, the nuclear exosome targeting component, is mutated in familial pulmonary fibrosis and is required for telomerase RNA maturation. Genes Dev. 2019;33(19-20):1381–1396. doi: 10.1101/gad.326785.119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Cronkhite J.T., Xing C., Raghu G. Telomere shortening in familial and sporadic pulmonary fibrosis. Am J Respir Crit Care Med. 2008;178(7):729–737. doi: 10.1164/rccm.200804-550OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Alvarez D., Cardenes N., Sellares J. IPF lung fibroblasts have a senescent phenotype. Am J Physiol Lung Cell Mol Physiol. 2017;313(6):L1164–L1173. doi: 10.1152/ajplung.00220.2017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Parry E.M., Alder J.K., Qi X., Chen J.J., Armanios M. Syndrome complex of bone marrow failure and pulmonary fibrosis predicts germline defects in telomerase. Blood. 2011;117(21):5607–5611. doi: 10.1182/blood-2010-11-322149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Wagner C.L., Hanumanthu V.S., Talbot C.C., Jr. Short telomere syndromes cause a primary T cell immunodeficiency. J Clin Invest. 2018;128(12):5222–5234. doi: 10.1172/JCI120216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Gorgy A.I., Jonassaint N.L., Stanley S.E. Hepatopulmonary syndrome is a frequent cause of dyspnea in the short telomere disorders. Chest. 2015;148(4):1019–1026. doi: 10.1378/chest.15-0825. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Diaz de Leon A., Cronkhite J.T., Katzenstein A.L. Telomere lengths, pulmonary fibrosis and telomerase (TERT) mutations. PLoS One. 2010;5(5) doi: 10.1371/journal.pone.0010680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Silhan L.L., Shah P.D., Chambers D.C. Lung transplantation in telomerase mutation carriers with pulmonary fibrosis. Eur Respir J. 2014;44(1):178–187. doi: 10.1183/09031936.00060014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Popescu I., Mannem H., Winters S.A. Impaired cytomegalovirus immunity in idiopathic pulmonary fibrosis lung transplant recipients with short telomeres. Am J Respir Crit Care Med. 2019;199(3):362–376. doi: 10.1164/rccm.201805-0825OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Merck S.J., Armanios M. Shall we call them “telomere-mediated”? Renaming the idiopathic after the cause is found. Eur Respir J. 2016;48(6):1556–1558. doi: 10.1183/13993003.02115-2016. [DOI] [PubMed] [Google Scholar]
- 20.Mitchell J.R., Wood E., Collins K. A telomerase component is defective in the human disease dyskeratosis congenita. Nature. 1999;402(6761):551–555. doi: 10.1038/990141. [DOI] [PubMed] [Google Scholar]
- 21.Egan E.D., Collins K. Biogenesis of telomerase ribonucleoproteins. RNA. 2012;18(10):1747–1759. doi: 10.1261/rna.034629.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Moon D.H., Segal M., Boyraz B. Poly(A)-specific ribonuclease (PARN) mediates 3′-end maturation of the telomerase RNA component. Nat Genet. 2015;47(12):1482–1488. doi: 10.1038/ng.3423. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Jonassaint N.L., Guo N., Califano J.A., Montgomery E.A., Armanios M. The gastrointestinal manifestations of telomere-mediated disease. Aging Cell. 2013;12(2):319–323. doi: 10.1111/acel.12041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Schratz K.E., Haley L., Danoff S.K. Cancer spectrum and outcomes in the Mendelian short telomere syndromes. Blood. 2020;135(22):1946–1956. doi: 10.1182/blood.2019003264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Penno M.B., Pedrotti-Krueger M., Ray T. Cryopreservation of whole blood and isolated lymphocytes for B-cell immortalization. J Tissue Cult Methods. 1993;15:43–47. [Google Scholar]
- 26.Baerlocher G.M., Vulto I., de Jong G., Lansdorp P.M. Flow cytometry and FISH to measure the average length of telomeres (flow FISH) Nat Protoc. 2006;1(5):2365–2376. doi: 10.1038/nprot.2006.263. [DOI] [PubMed] [Google Scholar]
- 27.Alder J.K., Hanumanthu V.S., Strong M.A. Diagnostic utility of telomere length testing in a hospital-based setting. Proc Natl Acad Sci U S A. 2018;115(10):E2358–E2365. doi: 10.1073/pnas.1720427115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Hamosh A., Sobreira N., Hoover-Fong J. PhenoDB: a new web-based tool for the collection, storage, and analysis of phenotypic features. Hum Mutat. 2013;34(4):566–571. doi: 10.1002/humu.22283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.NHLBI GO Exome Sequencing Project (ESP) ESP6500, Exome Variant Server. https://evs.gs.washington.edu/EVS/
- 30.1000 Genomes Project Consortium. Auton A., Brooks L.D., Durbin R.M. A global reference for human genetic variation. Nature. 2015;526(7571):68–74. doi: 10.1038/nature15393. https://www.internationalgenome.org/1000-genomes-browsers/ [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Karczewski K.J., Francioli L.C., Tiao G. Genome Aggregation Database Consortium. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature. 2020;581(7809):434–443. doi: 10.1038/s41586-020-2308-7. https://gnomad.broadinstitute.org Genome Aggregation Database (gnomAD), version 2.1.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Desmet F.O., Hamroun D., Lalande M., Collod-Beroud G., Claustres M., Beroud C. Human Splicing Finder: an online bioinformatics tool to predict splicing signals. Nucleic Acids Res. 2009;37(9):e67. doi: 10.1093/nar/gkp215. https://www.genomnis.com HSF version 3.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Schneider C.A., Rasband W.S., Eliceiri K.W. NIH Image to ImageJ: 25 years of image analysis. Nat Methods. 2012;9(7):671–675. doi: 10.1038/nmeth.2089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Allen R.C., Zoghbi H.Y., Moseley A.B., Rosenblatt H.M., Belmont J.W. Methylation of HpaII and HhaI sites near the polymorphic CAG repeat in the human androgen-receptor gene correlates with X chromosome inactivation. Am J Hum Genet. 1992;51(6):1229–1239. [PMC free article] [PubMed] [Google Scholar]
- 35.Parry E.M., Alder J.K., Lee S.S. Decreased dyskerin levels as a mechanism of telomere shortening in X-linked dyskeratosis congenita. J Med Genet. 2011;48(5):327–333. doi: 10.1136/jmg.2010.085100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Lee M., Roos P., Sharma N. Systematic computational identification of variants that activate exonic and intronic cryptic splice sites. Am J Hum Genet. 2017;100(5):751–765. doi: 10.1016/j.ajhg.2017.04.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Alder J.K., Stanley S.E., Wagner C.L., Hamilton M., Hanumanthu V.S., Armanios M. Exome sequencing identifies mutant TINF2 in a family with pulmonary fibrosis. Chest. 2015;147(5):1361–1368. doi: 10.1378/chest.14-1947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Meier U.T. The many facets of H/ACA ribonucleoproteins. Chromosoma. 2005;114(1):1–14. doi: 10.1007/s00412-005-0333-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Heiss N.S., Knight S.W., Vulliamy T.J. X-linked dyskeratosis congenita is caused by mutations in a highly conserved gene with putative nucleolar functions. Nat Genet. 1998;19(1):32–38. doi: 10.1038/ng0598-32. [DOI] [PubMed] [Google Scholar]
- 40.Dokal I. Dyskeratosis congenita in all its forms. Br J Haematol. 2000;110(4):768–779. doi: 10.1046/j.1365-2141.2000.02109.x. [DOI] [PubMed] [Google Scholar]
- 41.Knight S.W., Heiss N.S., Vulliamy T.J. X-linked dyskeratosis congenita is predominantly caused by missense mutations in the DKC1 gene. Am J Hum Genet. 1999;65(1):50–58. doi: 10.1086/302446. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Vulliamy T.J., Marrone A., Knight S.W., Walne A., Mason P.J., Dokal I. Mutations in dyskeratosis congenita: their impact on telomere length and the diversity of clinical presentation. Blood. 2006;107(7):2680–2685. doi: 10.1182/blood-2005-07-2622. [DOI] [PubMed] [Google Scholar]
- 43.Sauna Z.E., Kimchi-Sarfaty C. Understanding the contribution of synonymous mutations to human disease. Nat Rev Genet. 2011;12(10):683–691. doi: 10.1038/nrg3051. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.