Abstract
Natural compounds from diverse sources, including botanicals and commonly consumed foods and beverages, exert beneficial health effects via mechanisms that impact the epigenome and gene expression during disease pathogenesis. By targeting the so-called epigenetic ‘readers’, ‘writers’, and ‘erasers’, dietary phytochemicals can reverse abnormal epigenome signatures in cancer cells and preneoplastic stages. Thus, such agents provide avenues for cancer interception via prevention or treatment/therapeutic strategies. To date, much of the focus on dietary agents has been directed towards writers (e.g., histone acetyltransferases) and erasers (e.g., histone deacetylases), with less attention given to epigenetic readers (e.g., BRD proteins). The drug JQ1 was developed as a prototype epigenetic reader inhibitor, selectively targeting members of the bromodomain and extraterminal domain (BET) family, such as BRD4. Clinical trials with JQ1 as a single agent, or in combination with standard of care therapy, revealed antitumor efficacy but not without toxicity or resistance. In pursuit of second-generation epigenetic reader inhibitors, attention has shifted to natural sources, including dietary agents that might be repurposed as ‘JQ1-like’ bioactives. This review summarizes the current status of nascent research activity focused on natural compounds as inhibitors of BET and other epigenetic ‘reader’ proteins, with a perspective on future directions and opportunities.
Keywords: Bromodomains, Chromodomains, Cancer prevention, Epigenetics, Natural compounds
INTRODUCTION
Epigenetic ‘readers’, ‘writers’, and ‘erasers’ work in concert to coordinate the cellular epigenetic machinery and gene expression but are often deregulated in cancer and other pathologies [1]. Methylation of DNA, post-translational modifications (PTMs) to histones, chromatin-interacting proteins, and noncoding RNAs impart distinct molecular features that define transcriptionally ‘open’ euchromatin or ‘closed’ and repressive heterochromatin. The building blocks of chromatin are nucleosomes that consist of a coil of 147 bps of DNA wrapped around a core of eight, tightly packed histone proteins (dimers of histone H3, H4, H2A, and H2B). The N-terminal tails of histone proteins are subject to multiple PTMs that play a vital, dynamic role in regulating chromatin states and gene activity. For example, histone acetylation masks positively charged lysine residues in histone tails and lowers their affinity with negatively charged DNA, thereby opening transcription factor binding sites in gene promoters. Just as importantly, they also function as docking sites for reader proteins that recognize PTMs on histones, helping to recruit other factors that serve as transcriptional coactivators or corepressors [2].
Modifications at the level of DNA include cytosine methylation, hydroxymethylation, formylation, and carboxylation, whereas histone modifications are more numerous, with acetylation, methylation, and phosphorylation being the best studied to date. As noted above, these are dynamic alterations that, unlike the genetic changes in cancer, are potentially reversible by diet and lifestyle choices. Epigenetic writers modify DNA and histones by the addition of various chemical groups, whereas these can be reversed by epigenetic erasers, so that cells adapt and respond to environmental stimuli and endogenous signalling to regulate gene expression [1].
Examples of well-studied epigenetic writers are the DNA methyltransferases (DNMTs) that transfer a methyl group to 5’-cytosine residues, often present in CpG islands of gene promoters, facilitating gene repression, and histone acetyltransferases (HATs) that add acetyl groups to the ε-amino group of lysine residues in histones, promoting open chromatin and gene expression. Alternatively, histone methyltransferases (HMTs) can transfer a methyl group to arginine or lysine residues in histone tails, either activating or repressing gene expression depending on the number of methyl groups and the amino acid residue involved. These epigenetic marks can be removed by the erasing activities of Ten-eleven Translocation (TET) enzymes, histone deacetylases (HDACs) and histone demethylases.
Epigenetic readers are a group of diverse proteins with specialized docking domains that recognize and bind to different covalent modifications on DNA, histones, and non-histone proteins, laid down by the writers, and that assist other factors to mediate cellular and physiological outcomes [3]. Importantly, the epigenetic machinery responds to both internal and external/environmental stimuli to affect changes in gene expression but can be disrupted in cancer and other disease states. For this reason, much effort has been devoted to designing therapeutic agents that can target epigenetic readers, writers and erasers, several of which have been approved for clinical use. Examples include DNMT inhibitors (Azacytidine, Decitabine) for treating myelodysplastic syndromes and leukemias [4], and HDAC inhibitors (Vorinostat, Panobinostat) for targeting hematological malignancies [5]. Other drug candidates and lead compounds are in preclinical or clinical testing [1].
Natural compounds also have been explored in this regard, with several dietary phytochemicals known to reverse abnormal epigenetic signatures by interacting with HATs and HDACs. For example, curcumin found in the rhizome of Curcuma longa is a well-characterized HAT inhibitor [6], whereas the broccoli compound sulforaphane and the tea polyphenol epigallocatechin-3-gallate exert HDAC inhibitory activity [7,8]. An extensive literature exists on these and other natural dietary compounds, or their metabolites and other derivatives, influencing epigenetic factors that are altered in cancer [9-11]. A question that is often ignored concerns cause-and-effect, i.e., which epigenetic targets are critical for the anticancer mechanisms and which are simply bystanders. The answer, in part, relates to the concentrations of natural products employed in mechanistic studies vs. the doses achievable at target site(s) in vivo.
The majority of epigenetic drugs approved by the Food and Drug Administration, and those in the pipeline, target the writers and erasers, with many fewer developed specifically for inhibiting reader proteins. Readers of acetyl-lysine residues have garnered particular interest with the development of JQ1 as the first-in-class inhibitor of bromodomain and extraterminal domain (BET) family proteins, acting to downregulate MYC and other oncogenic targets for cancer prevention and therapy [12,13]. However, due to concerns over toxicity and resistance in some clinical trials with JQ1, attention has shifted to second-generation BET inhibitors, including natural compounds from botanical and food sources.
There are several reasons for studying botanicals and dietary factors as potential lead compounds for epigenetic reader inhibitors. First, the fundamental involvement of nutritional factors in epigenetic regulation is well established [14]; thus, identifying dietary agents as epigenetic reader inhibitors could provide new avenues for drug development, or inform on specific dietary recommendations. Second, food-derived compounds generally possess lower toxicity than standard therapies, while their metabolites and other derivatives represent untapped sources of novel epigenetic reader inhibitors [11,15]. Third, botanical and dietary compounds may give rise to beneficial synergistic outcomes when combined with new therapies, as illustrated by the HDAC inhibitors Vorinostat or sulforaphane with the BET inhibitor JQ1 [16,17].
Excellent reviews have focused on JQ1 and other synthetic agents, and this topic will not be covered here [18-21]. The current treatise considers natural compounds as inhibitors of epigenetic readers, an area that is much less well explored. After an overview of epigenetic readers, focusing mainly on acetyl- and methyl-readers, various natural compounds are introduced, ending with a summary and future perspectives.
EPIGENETIC READERS
Epigenetic readers can form higher-order multi-protein complexes that recognize and respond to the dynamic nature of PTMs on chromatin, arising from the crosstalk between the writers and the erasers. Readers are capable of initiating or silencing transcription, DNA repair, and other vital cellular processes via specialized domains that consist of a cavity or surface groove into which specific epigenetic marks can be accommodated. In the past 15 years or so, numerous such readers have been identified [22-33]. For example, the human proteome encodes over sixty acetyl reader bromodomains (BRDs) that are present in more than forty diverse proteins [34].
Readers are divided into families based on their ability to recognize specific modified amino acid residues, and also to distinguish different modification states of the same amino acid [22-24], such as histone H3 lysine 27 unmethylation (H3K27), monomethylation (H3K27me), dimethylation (H3K27me2), trimethylation (H3K27me3), and acetylation (H3K27ac). Specialized reader domains include the following: (1) BRDs that bind acetyl-lysine and target chromatin-modifying enzymes to specific sites; (2) chromatin organization modifier (chromodomain, CRD) moieties that interact with trimethylated lysines, resulting in either activation or silencing of gene expression; (3) malignant brain tumor (MBT) domains that recognize mono- and dimethylated lysines on histone H3 and histone H4 tails, thereby regulating transcription; (4) Tudor domains that can bind dimethylated and trimethylated lysines, as well as dimethylated arginines, and are involved in DNA repair and transcription; (5) proline–tryptophan–tryptophan–proline (PWWP) motifs, found in DNMTs that target trimethylated lysines; (6) plant homeodomains (PHDs) that are associated with multiple substrates; and (7) Yaf9, ENL, AF9, Taf1 and Sas5 (YEATS) domains that recognize crotonylated and acetylated lysine residues in active promoters and/or enhancers [3,25-27].
Examples of reader modules and the main histone PTMs recognized are provided in Table 1. The following sections will further discuss BRDs and CRDs which, to date, are the best characterized epigenetic readers.
Table 1.
Epigenetic reader modules and the histone marks recognized
| Modification | Residue | Family | Member |
|---|---|---|---|
| Acetylation | Lysine | BRD | BRD 2/3/4/T/7/9 |
| Methylation | Lysine | CRDs | MORF, MRG15 |
| Tudor domains | MBT, PHF1/19, TDRD7 | ||
| PWWP domains | BRPF1, NSD1-3 | ||
| Ankyrin repeats | GLP/G9a | ||
| Arginine | Tudor domains | WDR5, TDRD3, SMN1 | |
| Phosphorylation | Serine | 14-3-3 proteins BRCT domain | 14-3-3β/γ/η/ε/µ |
| Threonine | BIR domain | XRCC1, NBS1, BARD1 | |
| Tyrosine | PTB domain | ||
| Ubiquitination | Lysine | 53BP1 | |
| ADP-ribosylation | Glutamate | Macrodomains | RNF146 |
| Arginine | PBZ | APLF, CHFR | |
| Glutamate | WWE domain |
BRD, bromodomain; CRD, chromodomain; PWWP, Pro-Trp-Trp-Pro; MORF, monocytic leukemia zinc finger protein-related factor; MRG15, MORF4-related gene on chromosome 15; MBT, malignant brain tumor; PHF, plant homeodomain (PHD) finger; TDRD3/7, Tudor domain containing proteins 3 and 7; BRPF1, BRD and PHD finger-containing protein 1; NSD1-3, nuclear receptor binding SET domain proteins 1-3; GLP, G9a-like protein; WDR5, WD repeat domain 5; SMN1, survival of motor neuron 1; BRCT, BRCA1 C terminus; BIR, baculovirus IAP repeat; XRCC1, X-ray repair cross complementing 1; NBS1, Nijmegen breakage syndrome 1; BARD1, BRCA1 associated RING domain 1; PTB, phosphotyrosine-binding; 53BP1, tumor suppressor p53-binding protein 1; PBZ, PAR-binding zinc finger; WWE, Trp-Trp-Glu; RNF146, RING finger protein 146; APLF, aprataxin and PNKP like factor; CHFR, checkpoint with forkhead and RING finger domains (Adapted from Catia et al., 2019 [26]).
ACETYL READERS
First discovered in 1999 as domains that interacted with specific acetylated lysine residues to coordinate gene transcription, human BRDs are divided into eight sub-families [28,34]. Often linked by flexible sequences to other reader and catalytic domains within the same protein, BRDs facilitate diverse protein-protein interactions. For example, BRD and PHD finger-containing transcription factor (BPTF), a subunit of the imitation switch (ISWI) chromatin-remodelling complex, contains a PHD-BRD tandem module able to recognize both acetylated and methylated histone tails [29]. Other large chromatin-remodeling complexes also contain BRD modules that aid in either chromatin compaction or decompaction. Examples are bromodomain adjacent to zinc finger domain protein 2A (BAZ2A), a subunit of the nucleolar remodelling complex (NoRC) involved in noncoding RNA-dependent gene silencing [30], and Switch/sucrose non-fermenting (SWI/SNF)-related, matrix-associated, actin-dependent regulator of chromatin subfamily A members 2 and 4 (SMARCA2/4) found in SWI/SNF complexes, important in the control of cell differentiation and proliferation [31].
While interacting with histone acetylation marks, some BRD-containing proteins also engage as bi-functional ‘reader-writers’ in the coordinated (further) acetylation or methylation of nearby amino acid residues. Examples include the acetyltransferase p300/CBP-associated factor that acts as a transcriptional activator [32], and the methyltransferase mixed-lineage leukaemia protein, implicated in the expression of genes involved in self-renewal of haematopoietic stem cells, and enhanced by adjacent H3 acetylation [33].
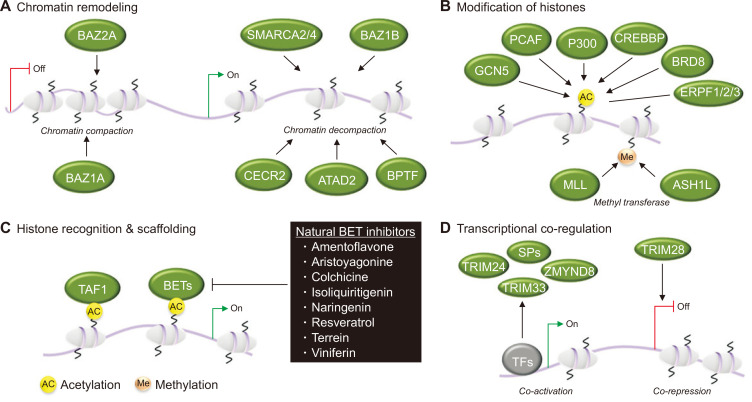
By recognizing acetylated histones, sub-families such as the BET members also act as scaffolds to recruit and segregate components of the transcriptional machinery that positively regulate growth-promoting genes [34]. Furthermore, BRD-containing proteins can affect gene expression by serving as transcriptional co-regulators. Examples include zinc-finger MYND domain-containing protein 8 (ZMYND8), which is a coactivator of estrogen receptor alpha (ERα) [35], or tripeptide motif-containing 24 (TRIM 24), which associates with nucleosomes through its BRD and PHD modules to activate androgen and retinoic acid receptors, hence aiding in gene regulation [36]. These various functional roles are illustrated in Figure 1.
Figure 1. Bromodomain (BRD)-containing proteins and gene regulation.
(A) BRD-containing proteins are frequently found in chromatin-remodeling complexes. For example, bromodomain adjacent to zinc finger domain (BAZ) proteins BAZ1A and BAZ2A promote chromatin compaction that leads to gene silencing. Other BRD-containing proteins promote chromatin decompaction, thus enhancing gene transcription. Examples include Switch/sucrose non-fermenting-related, matrix-associated, actin-dependent regulator of chromatin subfamily A members 2 and 4 (SMARCA2/4) and BAZ1B; cat eye syndrome critical region protein 2 (CECR2); BRD and PHD finger-containing transcription factor (BPTF); and ATPase family AAA domain-containing protein 2 (ATAD2). (B) BRD-containing proteins engage as bi-functional reader-writers in the acetylation and methylation of histones. They include acetyltransferases CREB-binding protein (CREBBP) and E1A-associated protein p300 (EP300), methyltransferases mixed-lineage leukaemia (MLL) and absent small and homeotic disks protein 1-like (ASH1L), or are part of larger histone-modifying complexes that include GCN5-like 2 (GCN5L2; also known as KAT2A), P300/CBP-associated factor (PCAF), BRD-containing protein 8 (BRD8) and the BRD and PHD finger-containing proteins (BRPFs), BRPF1, BRPF2 and BRPF3. (C) BRD-containing proteins can serve important roles in transcription by functioning as histone-recognizing scaffolds that promote the assembly of transcriptional complexes. For example, BRD and extraterminal (BET) proteins recruit components of the transcriptional machinery that positively regulate growth-promoting genes. Box: several natural compounds, including resveratrol and naringenin, are known BET inhibitors. Another BRD-containing protein, transcription initiation factor TFIID subunit 1 (TAF1), promotes transcription initiation. (D) BRD-containing speckled protein (SP) family members, zinc-finger MYND domain-containing protein 8 (ZMYND8), tripeptide motif-containing 24 (TRIM24) and TRIM33 can serve as transcriptional coregulators, while BRD-containing protein TRIM28 can act as a corepressor. TFs, transcription factors. Adapted from Takao and Panagis, 2017 [34], permission was obtained from the copyright holder.
Importantly, BRDs also recognize acetylation sites on nonhistone proteins with diverse cellular roles, including transcription factors such as GATA-binding factor 1 (GATA1) and the p65 subunit of NF-κB. In these examples, the presence of a BRD functionality helps to stabilize the complex of the transcription factor with the acetylated chromatin, thereby affecting transcriptional activation of target genes [37]. Other nonhistone proteins interacting with BRDs include DNA repair factors such as cell cycle and apoptosis regulator 2 (CCAR2) and p53 [17,34].
All BRDs are comprised of four α-helix bundles (Z, A, B, C) held together by ZA and BC loops. The ZA loop is relatively large, connecting αZ and αA, whereas the smaller BC loop connects αB and αC, combining to form a hydrophobic pocket with affinity for acetylated lysine residues on histone tails. The diversity of the ZA and BC loops provides for ligand specificity and selectivity towards specific acetylated histone sequences, thereby accounting for the eight BRD sub-families [38]. Furthermore, in BRD crystal structures, an array of five water molecules below the substrate binding pocket has been observed. These water molecules interact with inhibitors and ligands, and their locations provide important traits for drug design [39].
Although eight sub-families exist, this review focuses on BRD sub-families II and IV, due to the fact that studies reported to date on natural compounds, epigenetic readers and cancer have been directed towards these two families. Members of sub-family II garnered much interest after they were linked to the regulation of oncogenic MYC function. Thus, the BET family provides an attractive and ‘druggable’ target for cancer therapy [40], with JQ1 and several other small molecule inhibitors in the pipeline [19,41]. The first histone binding module inhibitor, JQ1, was designed to interact competitively with BRD4 at high specificity and potency [42]. This initial discovery paved the way for other reader inhibitor candidates, mainly focusing on the BET family. To date, JQ1 has been shown to inhibit pancreatic ductal adenocarcinoma, ovarian cancer, medulloblastoma, NUT midline carcinoma, and colorectal cancer, whereas I-BET151 was effective in mixed lineage leukemia [16,17,43-46]. These discoveries have opened doors to uncovering the druggable nature of other BRD sub-families [40].
For example, BRD family IV proteins implicated in cervical, bladder, uterine, endometrial and other malignancies [47-49] include BRD7, BRD9, ATPase family AAA domain-containing proteins 2 and 2b (ATAD2 and ATAD2b), and BRD and PHD finger-containing proteins 1 to 3 (BRPF1, BRPF2, and BRPF3), in which the PHD functionality also provides an additional mechanistic target [50]. Whereas BRPF1 is part of monocytic leukemia zinc finger protein (MOZ)/MOZ-related factor (MORF) HAT complexes, BRPF2 and BRPF3 are components of histone acetyltransferase binding to ORC1 (HBO1), which acetylates histone H4 at lysines 5, 8, and, 12 [51]. These complexes contain writer and reader domains that are implicated in disease pathogenesis, including acute myeloid leukemia for MOZ/MORF, and testicular tumors, mammary adenocarcinoma, and ovarian serous carcinoma for HBO1 [52,53]. The duality of ‘reader/writer’ or ‘reader/eraser’ functionalities increases the utility of protein complexes comprising epigenetic readers, for example, by docking onto existing histone acetylation marks with BRDs while simultaneously writing new methyl or acetyl groups on adjacent lysine/arginine residues [34].
Two of the family IV proteins, BRD7 and BRD9, are components of SWI/SNF chromatin remodeling complexes, and their BRDs are conserved with 71.8% identity and 88.7% similarity [50]. Downregulation at the mRNA level has been reported for BRD7 in colorectal and nasopharyngeal carcinomas [54,55], and for BRD9 in uveal melanoma [56]. At the protein level, high BRD7 is a favorable prognostic marker in renal cancer but an unfavorable one in melanoma, whereas BRD9 overexpression is a negative indicator in renal and liver cancer [57]. The BRD9, which recognizes histone H4K5acK8ac, has been shown to have an oncogenic role in various cancers, including ovarian cancer, squamous cell lung carcinoma and esophageal carcinoma [48]. Notably, BRD7 also regulates p53 and phosphoinositide 3-kinase (PI3K) activity through SWI/SNF complexes [58,59], whereas BRD9 has been implicated in the BET/BRD9 acetyl ‘switch’ on the Wnt coactivator CCAR2 [17]. To date, there is less research on the ligands of BRD7/BRD9 proteins compared with BET family members, but promising lead compounds do exist, including LP99 and I-BRD9 as selective inhibitors of BRD7 and BRD9 [60,61].
Another noteworthy candidate for cancer interception is ATAD2, containing an N-terminal AAA+ ATPase domain as well as a C-terminal BRD [38,62]. ATAD2 is a critical transcription factor in malignant cells and has been shown to upregulate antiapoptotic activity in prostate and breast cancer [63,64]. Due to its association with many cancer types, ATAD2 also has been identified as a potential therapeutic target [65]. Although ATAD2 and ATAD2b are highly conserved, with 94.4% overall similarity, little is known about ATAD2b functions in cancer. High ATAD2 and ATAD2b protein expression is an unfavourable prognostic marker for renal cancer [66].
METHYL READERS
CRDs consist of 40 to 50 amino acids that are found in 29 different proteins involved in chromatin remodeling, with critical roles in propagating and maintaining a repressed chromatin state. The domains typically consist of an N-terminal three-stranded β-sheet packed against a C-terminal α-helix, and the associated hydrophobic groove has affinity for specific methylated lysine residues on histone and nonhistone proteins [67]. Because they recognize various methylated substrates, they have been categorized as canonical or noncanonical according to mechanism of target recognition, context, and structure.
Those with high sequence homology to HP1 are referred to as canonical [67]. These domains recognize a conserved lysine-methylated motif, ARKS, which was originally discovered in the H3 tail at H3K9 and H3K27 sites. Canonical members in mammals include HP1 and polycomb CRD, CBX-like CRD in Chp1 protein [68], and those that reside in Pdd1p and Pdd3p proteins in free-living ciliates, Tetrahymena [69]. Noncanonical CRDs contain amino and carboxyl-terminal extensions and internal inserts that recognize targets along histone tails, such as methyl marks on histone H3K4, H3K36, and H4K20 [67]. They include CHD proteins that contain double CRDs, CRDs in the MORF-related gene family that are components of HAT and HDAC complexes, and CRDs in the MYST protein family of lysine acetyltransferases [70]. Tudor domains act as methyl-lysine and methyl-arginine readers [71], implicated in breast cancer [72], and a focus of druggable target modulation [26].
Genetic alterations in CRD proteins during tumor development negatively impact their recognition of methylation marks on histone and non-histone proteins, leading to misinterpretation of chromatin states. For example, mutations in the CRD protein CDYL that normally recognizes histone H3K27me3 is found in cervical cancer and is associated with poor prognosis [73]. Mutations in CBX proteins responsible for recruiting PRC1 complexes to H3K27me3 sites are associated with invasiveness and epithelial-to-mesenchymal transition in many types of cancers [74-76]. Because of the pivotal role that methyl readers play in gene regulation, oncogenesis, and other pathologies, a number of small molecules have been developed as drug candidates [18,77,78].
Despite the advancements in targeting epigenetic readers, fuelled by studies with JQ1 and related drug candidates, toxicity remains a concern and an impediment to breakthroughs for the major malignancies in the population [1,79]. This has led to interest in broadening the scope of the research to seek novel lead compounds, including natural bioactives from botanical and dietary sources. Examples of such natural compounds are discussed in the next section.
NATURAL COMPOUNDS
Natural compounds from diverse sources, such as plants, microorganisms, or marine species, have beneficial effects against cancer and other pathologies due to a combination of anti-inflammatory, anti-oxidant, pro-apoptotic, anti-angiogenic and anti-metastatic effects [80-84]. Phytochemicals in fruits, vegetables, and other natural sources make them ideal anti-cancer candidates because of their wide availability, good tolerability, and relatively low cost.
Dietary components act on epigenetic regulation at the level of DNA methylation, histone modifications, chromatin function and noncoding RNAs [11,51,85-92]. Among the many phytochemicals implicated as epigenetic modifiers are curcumin, genistein, quercetin, resveratrol, sulforaphane, tea catechins, triterpenoids, organoselenium and organosulfur compounds, short-chain fatty acids, indoles, and β-carotene. Reports have focused on the inhibition of writers, mainly DNMTs and HATs, or erasers such as HDACs. Some of the natural compounds or their whole food sources have entered preclinical and clinical trials [11,85,87,91,93-98].
Natural compounds from marine sources also have been reported to alter the activity of DNMTs, HATs, HMTs, and HDACs [99,100]. However, none of these compounds have been identified as acting specifically on epigenetic readers. For example, curcumin inhibits p300/CPB [101], but this occurs via the acetyltransferase activity rather than the BRD function in the dual ‘reader-writer’ protein. To our knowledge, there are no rigorous reports on phytochemical-derived antagonists of CRD-containing proteins. Few synthetic methyl-reader probes have been developed compared to acetyl-reader inhibitors, restricting our understanding of CRDs and the best ways to approach them as druggable candidates. Thus, the current review focuses mainly on natural compounds as potential inhibitors of the BET family member BRD4, with one example targeting BRD9 (see Table 2).
Table 2.
Natural compounds targeting epigenetic ‘reader’ domains
| Name & structure | Origin | Target | Reference | |
|---|---|---|---|---|
Naringenin triacetate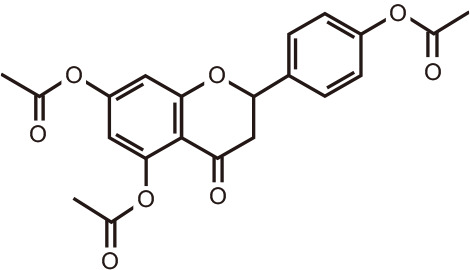
|
Citrus aurantium (bitter orange) | BRD4 | 104 | |
3-O-acetylpinobanksin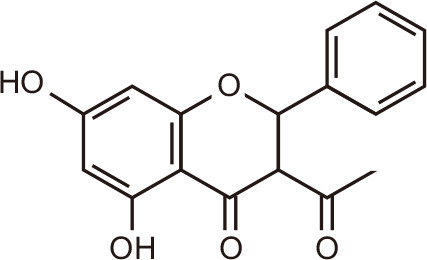
|
Black poplar tree | |||
Kaempferol tetracetate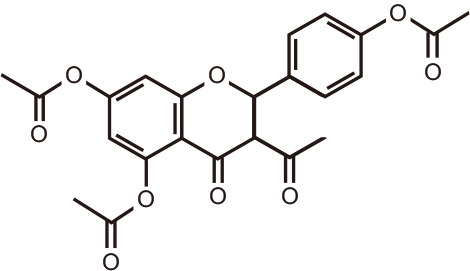
|
Cudrania tricuspidata (mandarin melonberry) | |||
Resveratrol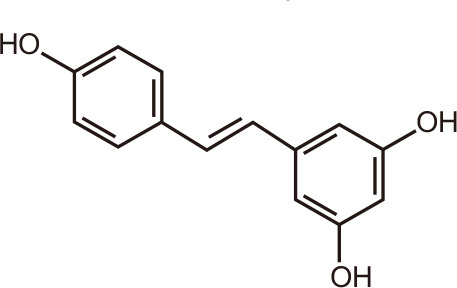
|
Peanuts, berries, grape skin, apples | BRD4 | 112 | |
Isoliquiritigenin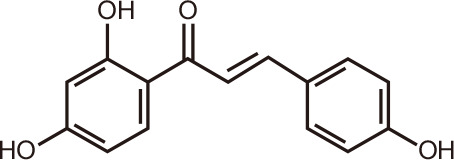
|
Glycyrrhiza glabra (licorice plant) | BRD4 | 113 | |
Amentoflavone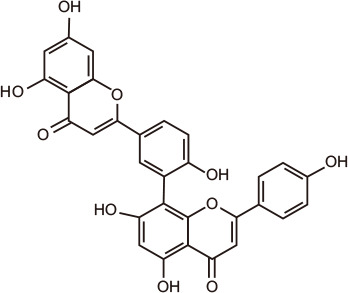
|
Ginkgo biloba, Hypericum perforatum (St. John’s Wort), Biophytum sensitivum, Nandina domestica (sacred bamboo) | BRD4 | 102 | |
Fisetin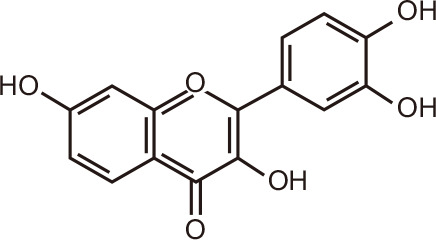
|
Cucumber, onion, strawberry, apple | |||
Aristoyagonine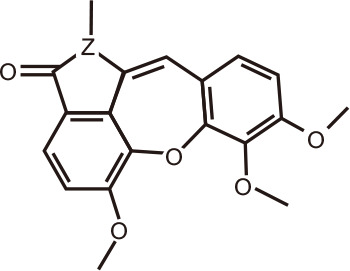
|
Sarcocapnos enneaphylla | BRD4 | 125 | |
α-Viniferin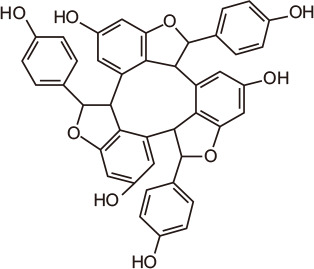
|
Caragana sinica, camphor tree, Astilbe grandis | BRD4 | 134 | |
Euscaphic acid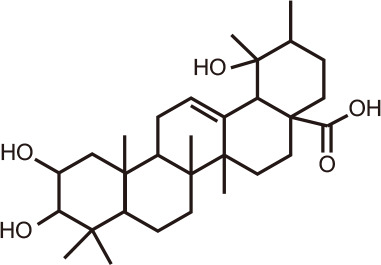
|
Plectranthus amboinicus (French thyme, Cuban oregano, Indian mint), Ziziphus jujuba (red date) | |||
Magnolol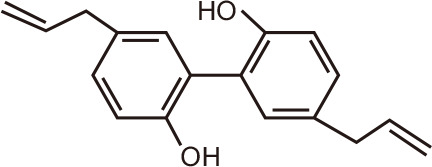
|
Magnolia tree | BRD9 | 135 | |
Colchicine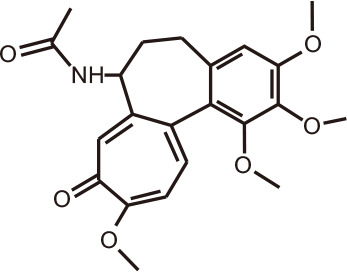
|
Colchiceine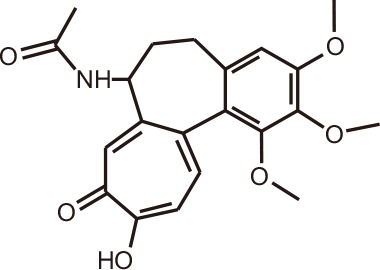
|
Colchicum speciosum (autumn crocus) | BRD4 | 126 |
Terrein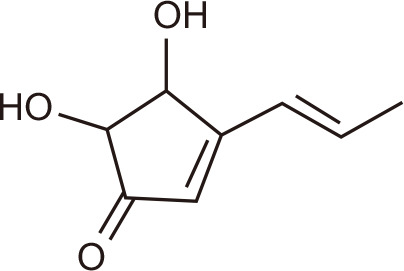
|
Mangrove-derived Streptomyces sp. SZ-A15 | BRD4 | 136 | |
4-oxo-Staurosporine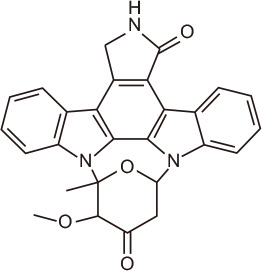
|
Marine-derived Streptomyces sp. NB-A22 | BRD4 | 130 | |
3,11α,12β,13β,6-pentahydroxy-11,12-dihydroperylen-6(13H)-one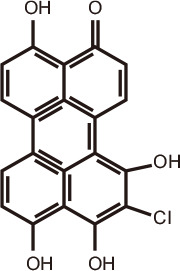
|
Marine fungi: Alternaria sp. NH-F6 | BRD4 | 137 | |
5-hydroxymethyl-furan-2-carboxylic acid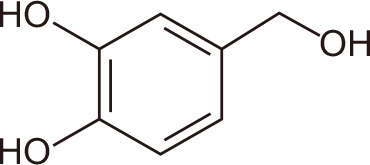
|
Marine fungi: Phoma sp. CZD-F11, Aspergillus sp. CZD-F18 | BRD4 | 138 | |
4-hydroxymethyl catechol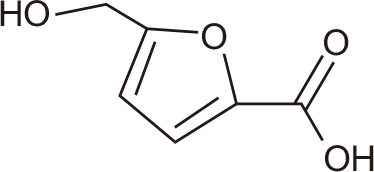
|
Marine fungi: Phoma sp. CZD-F11, Aspergillus sp. CZD-F18 | |||
12-hydroxy-verruculogen TR-2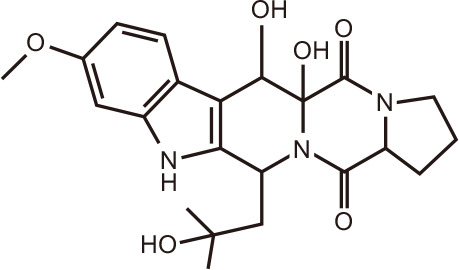
|
Fumitremorgin B
|
Penicillium sp. DT-F29 with Bacillus sp. B31 | BRD4 | 131 |
FLAVONOIDS
Through molecular docking and dynamic simulation in silico, flavonoids emerged as putative modulators of BRD proteins [102,103]. Natural compounds 3-O-acetylpinobanksin, naringenin triacetate, and kaempferol tetraacetate from poplar, bitter orange, and melonberry, respectively, interacted favorably with BET family members, in particular with the first BRD of BRD4, BRD4(1) [104]. The biflavonoid amentoflavone, found in Hypericum perforatum, Nandina domestica, Gingko biloba, and Biophytum sensitivum, was shown to dock into the N-acetyl-binding site of BRD4, making several contacts with the ZA channel and with non-canonical residues. Subsequent experiments confirmed the inhibition of BRD4 by amentoflavone in the micromolar range [102].
Fisetin, which is present in foods such as cucumber, onion, strawberry and apple, also docked favorably with BET family members in silico, but was a less potent inhibitor than amentoflavone of BRD4 [102]. Both compounds possess NF-κB modulatory activity and alter cell proliferation and apoptosis in cancer cells [105,106]. Fisetin also inhibits DNMT1 [107] and p300/CPB [108], a HAT that serves as a co-activator of NF-κB, explaining in part the reported NF-κB modulatory activity.
Resveratrol, a flavanol found naturally in peanuts, mulberries, raspberries, blueberries and grapes, inhibits class I, II, and IV HDACs in a dose-dependent manner [109], but has opposite effects on class III HDACs, serving as an allosteric sirtuin activating compound [110]. Thus, resveratrol regulates several signal transduction pathways involved in cancer, aging, inflammation, and neurodegeneration. In particular, resveratrol induces both intrinsic and extrinsic apoptotic pathways and interferes with different stages of cancer development [111]. Recent reports indicated that resveratrol might act as a pan-BET inhibitor [112]. Using isothermal titration calorimetry, a dissociation constant (Kd) of 6.6 µM towards BRD4(1) was found, corroborated by molecular docking studies in silico that showed favorable acetyl-lysine binding site interactions of resveratrol. Differential scanning calorimetry revealed 2°C to 3°C melting temperature shifts (ΔTm) of BET BRDs in the presence of 100 µM resveratrol. These findings suggested that resveratrol might exert anticancer activities, and other beneficial health effects, in part, by inhibiting cellular BRD functions.
An additional natural polyphenolic compound identified as a BRD4 inhibitor is the chalcone isoliquiritigenin (ISL), present in licorice, from the root of the herbaceous perennial legume Glycyrrhiza glabra. Crystallographic screening revealed that ISL interacts with BRD4 in a novel enthalpy driven mode [113]. Optimization and stabilization of the hydrogen bond network of the conserved water molecules contributed to favorable binding enthalpy, which makes ISL a novel lead compound for new BRD4 inhibitors. The fact that it binds competitively to BRD4 could explain, in part, the anticancer activity previously observed for ISL in various cancer cell lines [114].
Quercetin is another naturally occurring polyphenolic compound found in fruits and vegetables that has been linked to BRD inhibition, albeit indirectly, by promoting the antitumor effects of BET inhibitors [115]. Specifically, when pancreatic and thyroid cancer cells were co-treated with JQ1 and quercetin, there was reduced cell proliferation and enhanced apoptosis compared with either compound alone. Interestingly, quercetin targeted hnRNPA1, a nuclear protein involved in the regulation of mRNA translation of anti-apoptotic proteins. Indeed, hnRNPA1 knockdown enhanced the effects of BET inhibition on proliferation, apoptosis, and cell survival. Hence, quercetin might exert its effects on JQ1-mediated apoptosis in part by suppression of hnRNPA1. In a xenograft model of pancreatic cancer, quercetin downregulated hnRNPA1 in vivo and augmented the activity of BET inhibitors in suppressing tumor growth. Because hnRNPA1 and the BET protein BRD4 are co-expressed in human thyroid and pancreatic tumors, the authors suggested pursuing this combination therapy in advanced cancer patients [115].
These studies provided proof-of-principle that natural compounds in combination with synthetic BET inhibitors might help to reduce the unwanted side effects of BRD therapy by lowering the dose required for effective treatment.
VITAMINS
Another combination study of note showed that vitamin C, present ubiquitously in many fruits and vegetables, enhanced the sensitivity of melanoma cells to BET inhibitors, including JQ1. Vitamin C promotes DNA demethylation via the TET-mediated hydroxymethylation pathway, which leads to downregulation of HAT1. This also determines loss in acetylation of histone H4, leading to disruption of chromatin interactions involving BRD4 [116]. In ascorbate-deficient Gulo-/- mice, a lack of vitamin C attenuated the antitumor activity of JQ1 in melanoma xenografts, whereas vitamin C supplementation enhanced the tumor inhibition.
The SWI/SNF chromatin remodeling complex PBAF contains PB1 (BAF180), a protein with six BRDs that is critical for ligand-dependent activation of nuclear hormone receptors, including vitamin D receptor (VDR)-dependent transcription in response to vitamin D [117]. Recent studies showed that alternative recognition of an acetylated lysine in VDR by BRD7 and BRD9 directs association with PBAF and BAF chromatin remodeling complexes [118].
The B vitamins include thiamine, riboflavin, niacin, pantothenic acid, pyridoxine, biotin, folate, and various cobalamins. Folic acid has been investigated extensively in terms of epigenetic mechanisms regulating DNA and histone methylation, via the one-carbon pool and S-adenosyl methionine [119]. Recent studies in chronic myeloid leukemia cells showed that a fraction of cellular methylenetetrahydrofolate dehydrogenase 1 resides in the nucleus and is recruited to distinct genomic loci by interaction with BRD4 [120]. A well-known target of JQ1, MYC, also has been linked mechanistically to methylenetetrahydrofolate dehydrogenase-cylcohydrolase 2 in acute myeloid leukemia [121].
All-trans retinoic acid, which is synthesized from vitamin A as a physiological ligand for the retinoic acid receptor, combined effectively with JQ1 to exert potent anticancer activity in leukemia cells via c-MYC downregulation [122]. Retinoids and rexinoids have been widely studied in cancer prevention and therapy, and the associated acquired or intrinsic resistance mechanisms might be circumvented by combined BRD inhibition [123,124].
ALKALOIDS
Aristoyagonine is found in the extracts of the flowering plant Sarcocapnos enneaphylla, which belongs to the Papaveraceae family. It was the first natural compound to be reported as a BRD inhibitor, after screening assays on 8,000 compounds in the Korea Chemical Bank [125]. Aristoyagonine (5,7,8-trimethoxy-1-methylbenzo[6,7]oxepino[4,3,2-cd]isoindol-2(1H)-one) possesses a unique scaffold and belongs to the aristocularine alkaloid family. The IC50 in an alpha-screen was 3.27 µM, and 1.03 µM in an ELISA. Aristoyagonine had cytotoxic activity against I-BET-762-sensitive and I-BET-762-resistant cancer cells by downregulating c-MYC expression, and reduced tumor growth in a Ty82-xenograft mouse model [125]. This seminal report laid a foundation for the development of new BRD synthetic inhibitors based on the scaffold of aristoyagonine.
High-throughput virtual screening of more than 7 million small molecules, aimed at discovering novel inhibitors of BRD4, prioritized seven compounds for further validation by isothermal titration calorimetry [126]. Among the lead compounds was the alkaloid colchicine and its analogs, which showed binding affinity towards BRD4(1) in the range Kd = 20-46 µM. Co-crystallization structures had IC50 values of 13-17 µM, with the natural compounds occupying the acetyl-lysine recognition pocket of BRD4. These alkaloids are found in the herbaceous perennial flowering plant Colchicum speciosum, belonging to the family of Colchicaceae, more commonly known as the autumn crocus. Colchicine was extracted for centuries to ameliorate gout, before the advent of anticancer therapeutics targeting microtubules [127].
Indolocarbazoles are a class of natural compounds that have gained interest as anticancer agents through various mechanisms, including topoisomerase I-DNA complex stabilization and protein kinase inhibition [128,129]. These compounds isolated from Streptomyces strain NB-A22 showed the potential to act as BRD inhibitors. Ethyl acetate extracts yielded several indolocarbazole alkaloids, including staurosporine, which were evaluated for cytotoxic activity in PC3 prostate cancer cells and BRD4 inhibitory activity using the time-resolved fluorescence energy transfer (TR-FRET) assay. The results showed that the most efficient alkaloid was 4-oxo-staurosporine, with 60% inhibition at 10 µg/mL [130].
Two 2,5-diketopiperazine derivatives, namely, 12-hydroxyverruculogen TR-2 and Fumitremorgin B, were among several compounds isolated and characterized from co-cultures of Penicillium sp. DT-F29 with Bacillus sp. B31 [131]. The BRD4-inhibitory activity assay for these two compounds at 20 µM was 72.7% and 80.4%, respectively, equivalent to the positive control JQ1 (85.7%).
TERPENES AND STILBENES
Astilbe grandis Stapf ex E.H. Wilson, an herbaceous perennial plant grown in Korea and China, has been used in traditional medicine to treat wounds, bites and burns [132]. Extracts from this plant exhibited protective effects in UVB-treated keratinocytes via protein tyrosine phosphatase 1B [133]. Screening of compounds from Astilbe sp. for BRD4 inhibitory activity using a TR-FRET assay identified a new monoterpene, (S)-3-(2-hydroxyethyl)-5-(2-methylprop-1-en-1-yl)furan-2(5H)-one, as well as other natural compounds such as caffeic acid, mandelic acid, sonchifolinin B, α-viniferin, euscaphic acid, cianidanol, β-sitosterol, and stigmasterol.
The stilbene trimer α-viniferin and the triterpenoid euscaphic acid, also known as tormentic acid, exhibited IC50 values towards BRD4 of 13.2 µM and 17.39 µM, respectively. With inhibitory activity against HCC827 and A549 lung cancer cells and HeLa cervical carcinoma cells [134], these natural products warrant further attention as lead compounds for new BRD therapeutics.
LIGNANS
A docking study that aimed to identify anticancer molecular targets for compounds in the bark of Magnolia officinalis and Magnolia grandiflora prioritized BRD9, along with tankyrase-2 and casein kinase 2, as top hits among > 300 proteins screened [135]. The lignan magnolol and its structural derivatives fit the binding pockets of all three proteins as potential competitive inhibitors. However, to date, no reports have appeared that formally identified natural compounds as bona fide inhibitors of BRD9.
TERREIN
Mangroves comprising groups of trees and shrubs that grow in the coastal intertidal zone have been investigated as a source of novel bioactives, including the microbes that exist in a symbiotic relationship with host plants. One such microbe, Streptomyces sp. SZ-A15, is an actinomycete with potent antitumor activity in human PC3 prostate cancer cells. Ethyl acetate extracts of the fermented actinomycete yielded four new bisamides and two new rhamnosides, along with N1-acetyl-N7-phenylacetyl cadaverine, N-(2-hydroxy-3-phenylpropyl) acetamide, α-N-acetylanhydro-L-ornithine, and 2-cyclopenten-1-one,4,5-dihydroxy-3-propenyl-(8Cl). Only the latter compound, also known as terrein, had activity towards BRD4 in a TR-FRET assay, with 78.4% inhibition at 10 µM concentration [136].
MISCELLANEOUS COMPOUNDS
Ethyl acetate extracts of the marine fungus Alternaria sp. NH-F6 yielded 24 secondary metabolites, including perylenequinones, alternaric acid, butyrolactone, and cerebroside. After testing for BRD4 inhibitory activity using the TR-FRET assay, a perylenequinone compound was identified with 88% inhibition at a concentration of 10 µM [137]. Seven diphenyl ether analogs also were identified, including a new diphenyl ether from fermented extracts of Phoma sp. CZD-F11 and Aspergillus sp. CZD-F18. 5-Hydroxymethylfuran-2-carboxylic acid and 4-(hydroxymethyl) catechol exhibited BRD4-inhibitory activity of 78.5 and 76.4% at 10 µg/mL, respectively [138].
SUMMARY
Diverse natural dietary and non-dietary compounds from plants, microbes, and the marine environment provide a rich resource for identifying and characterizing the next-generation of epigenetic reader bioactives, acting at the level of chromatin and transcriptional coactivators/corepressors. Research to date has focused on BRD4 and other members of the BET family, seeking second-generation JQ1-like agents, with a few examining BRD9/7 inhibition. Magnolol potentially targeting BRD9 [134] provides proof-of-principle for natural compounds affecting non-BET acetyl readers, which is an exciting avenue for future growth, via the targeting of other sub-families. Studies reviewed herein have highlighted a number of different tools and approaches to defining new scaffolds from which to develop novel lead compounds as BRD inhibitors, mainly using in silico, in vitro, and cell-based assays.
Few if any of the reports have provided compelling evidence, especially in vivo, for BRD-specific inhibition by natural dietary compounds in the context of cancer treatment or prevention. Establishing specificity would entail knockdown or knockout strategies using small interference RNA or single-guide RNA (CRISPR/Cas 9) approaches, showing loss of the mechanistic target is pertinent for the BRD inhibitory functions. Given the ‘reader-writer’ and ‘reader-eraser’ duality aspects discussed above, it is possible to conceptualize how inhibition of certain readers might result in the erasure of abnormal epigenome signatures in cancer cells. However, what might be the consequences of BRD or CRD protein inhibition, and therefore impaired reader function, on the phenotype of normal cells? We might take lessons from the HDAC inhibitor field, in which normal cells exhibit homeostatic mechanisms that control or ‘buffer’ against extreme changes in histone acetylation, whereas cancer cells remain susceptible [11,17,91]. Parallel investigations in normal cells and tissues should be part of the standard operating procedure for future studies on epigenetic reader inhibition. This also will inform on toxicity issues, posed by drugs such as JQ1 in clinical trials, and avoidance strategies using natural compounds as reader inhibitors. These are key issues to be addressed in order to advance the field towards improved cancer interception via epigenetic combination agents [11,16].
A key question concerns the physiologically-relevant concentrations that can be achieved at the molecular target (e.g., BRD4 or BRD9) from dietary compounds, and the role of metabolism in generating intermediates with increased or decreased affinity for BRDs in situ [91]. Many of the docking studies in silico and bioassays in vitro focus on the parent compound, ignoring the likely scenario of extensive metabolism and excretion in vivo. A prime example is provided by the flavonoids and other polyphenols that interact with the gut microbiome, and undergo extensive phase I and phase II metabolism by host tissues [139-141].
Another important consideration relates to the potential synergistic (or antagonist) interactions with other natural compounds acting via alternative epigenetic mechanisms, for example involving lysine or arginine methyltransferases, HATs, HDACs, DNMTs, or TET enzymes. This exciting, nascent field is likely to move rapidly as new leads are pursued towards the prevention and interception of cancer, aging, and chronic degenerative diseases [142-145].
ACKNOWLEDGEMENTS
Original research from the laboratories of ED, PR and RHD that appeared in this review was supported in part by grants CA090890 and CA122959 from the National Cancer Institute, by the John S. Dunn Foundation, by AgriLife Research, and by a Chancellor’s Research Initiative from Texas A&M University. ED also acknowledges funding from the Polytechnic University of the Marche.
Footnotes
CONFLICTS OF INTEREST
No potential conflicts of interest were disclosed.
REFERENCES
- 1.Biswas S, Rao CM. Epigenetic tools (the writers, the readers and the erasers) and their implications in cancer therapy. Eur J Pharmacol. 2018;837:8–24. doi: 10.1016/j.ejphar.2018.08.021. [DOI] [PubMed] [Google Scholar]
- 2.Zhang G, Pradhan S. Mammalian epigenetic mechanisms. IUBMB Life. 2014;66:240–56. doi: 10.1002/iub.1264. [DOI] [PubMed] [Google Scholar]
- 3.Arrowsmith CH, Bountra C, Fish PV, Lee K, Schapira M. Epigenetic protein families: a new frontier for drug discovery. Nat Rev Drug Discov. 2012;11:384–400. doi: 10.1038/nrd3674. [DOI] [PubMed] [Google Scholar]
- 4.Contieri B, Duarte BKL, Lazarini M. Updates on DNA methylation modifiers in acute myeloid leukemia. Ann Hematol. 2020;99:693–701. doi: 10.1007/s00277-020-03938-2. [DOI] [PubMed] [Google Scholar]
- 5.Pinto V, Bergantim R, Caires HR, Seca H, Guimarães JE, Vasconcelos MH. Multiple myeloma: available therapies and causes of drug resistance. Cancers (Basel) 2020;12:407. doi: 10.3390/cancers12020407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Marcu MG, Jung YJ, Lee S, Chung EJ, Lee MJ, Trepel J, et al. Curcumin is an inhibitor of p300 histone acetylatransferase. Med Chem. 2006;2:169–74. doi: 10.2174/157340606776056133. [DOI] [PubMed] [Google Scholar]
- 7.Tortorella SM, Royce SG, Licciardi PV, Karagiannis TC. Dietary sulforaphane in cancer chemoprevention: the role of epigenetic regulation and HDAC inhibition. Antioxid Redox Signal. 2015;22:1382–424. doi: 10.1089/ars.2014.6097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Thakur VS, Gupta K, Gupta S. Green tea polyphenols increase p53 transcriptional activity and acetylation by suppressing class I histone deacetylases. Int J Oncol. 2012;41:353–61. doi: 10.3892/ijo.2012.1449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Yang AY, Kim H, Li W, Kong AN. Natural compound-derived epigenetic regulators targeting epigenetic readers, writers and erasers. Curr Top Med Chem. 2016;16:697–713. doi: 10.2174/1568026615666150826114359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Dashwood RH, Myzak MC, Ho E. Dietary HDAC inhibitors: time to rethink weak ligands in cancer chemoprevention? Carcinogenesis. 2006;27:344–9. doi: 10.1093/carcin/bgi253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Rajendran P, Ho E, Williams DE, Dashwood RH. Dietary phytochemicals, HDAC inhibition, and DNA damage/repair defects in cancer cells. Clin Epigenetics. 2011;3:4. doi: 10.1186/1868-7083-3-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Zaware N, Zhou MM. Bromodomain biology and drug discovery. Nat Struct Mol Biol. 2019;26:870–9. doi: 10.1038/s41594-019-0309-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Cochran AG, Conery AR, Sims RJ., 3rd Bromodomains: a new target class for drug development. Nat Rev Drug Discov. 2019;18:609–28. doi: 10.1038/s41573-019-0030-7. [DOI] [PubMed] [Google Scholar]
- 14.Stover PJ, James WPT, Krook A, Garza C. Emerging concepts on the role of epigenetics in the relationships between nutrition and health. J Intern Med. 2018;284:37–49. doi: 10.1111/joim.12768. [DOI] [PubMed] [Google Scholar]
- 15.Turrini E, Ferruzzi L, Fimognari C. Natural compounds to overcome cancer chemoresistance: toxicological and clinical issues. Expert Opin Drug Metab Toxicol. 2014;10:1677–90. doi: 10.1517/17425255.2014.972933. [DOI] [PubMed] [Google Scholar]
- 16.Mazur PK, Herner A, Mello SS, Wirth M, Hausmann S, Sánchez-Rivera FJ, et al. Combined inhibition of BET family proteins and histone deacetylases as a potential epigenetics-based therapy for pancreatic ductal adenocarcinoma. Nat Med. 2015;21:1163–71. doi: 10.1038/nm.3952. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Rajendran P, Johnson G, Li L, Chen YS, Dashwood M, Nguyen N, et al. Acetylation of CCAR2 establishes a BET/BRD9 acetyl switch in response to combined deacetylase and bromodomain inhibition. Cancer Res. 2019;79:918–27. doi: 10.1158/0008-5472.CAN-18-2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Arrowsmith CH, Schapira M. Targeting non-bromodomain chromatin readers. Nat Struct Mol Biol. 2019;26:863–9. doi: 10.1038/s41594-019-0290-2. [DOI] [PubMed] [Google Scholar]
- 19.Shi Y, Liu J, Zhao Y, Cao J, Li Y, Guo F. Bromodomain-containing protein 4: a druggable target. Curr Drug Targets. 2019;20:1517–36. doi: 10.2174/1574885514666190618113519. [DOI] [PubMed] [Google Scholar]
- 20.Wu Q, Heidenreich D, Zhou S, Ackloo S, Krämer A, Nakka K, et al. A chemical toolbox for the study of bromodomains and epigenetic signaling. Nat Commun. 2019;10:1915. doi: 10.1038/s41467-019-09672-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Waybright JM, James LI. Getting a handle on chemical probes of chomatin readers [published online ahead of print January 10, 2020] Future Med Chem. :doi: 10.4155/fmc-2019-0274. doi: 10.4155/fmc-2019-0274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Musselman CA, Lalonde ME, Côté J, Kutateladze TG. Perceiving the epigenetic landscape through histone readers. Nat Struct Mol Biol. 2012;19:1218–27. doi: 10.1038/nsmb.2436. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Dawson MA, Kouzarides T, Huntly BJ. Targeting epigenetic readers in cancer. N Engl J Med. 2012;367:647–57. doi: 10.1056/NEJMra1112635. [DOI] [PubMed] [Google Scholar]
- 24.Taverna SD, Li H, Ruthenburg AJ, Allis CD, Patel DJ. How chromatin-binding modules interpret histone modifications: lessons from professional pocket pickers. Nat Struct Mol Biol. 2007;14:1025–40. doi: 10.1038/nsmb1338. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Greschik H, Schüle R, Günther T. Selective targeting of epigenetic reader domains. Expert Opin Drug Discov. 2017;12:449–63. doi: 10.1080/17460441.2017.1303474. [DOI] [PubMed] [Google Scholar]
- 26.Mio C, Bulotta S, Russo D, Damante G. Reading cancer: chromatin readers as druggable targets for cancer treatment. Cancers (Basel) 2019;11:61. doi: 10.3390/cancers11010061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Zhao D, Li Y, Xiong X, Chen Z, Li H. YEATS domain-a histone acylation reader in health and disease. J Mol Biol. 2017;429:1994–2002. doi: 10.1016/j.jmb.2017.03.010. [DOI] [PubMed] [Google Scholar]
- 28.Filippakopoulos P, Picaud S, Mangos M, Keates T, Lambert JP, Barsyte-Lovejoy D, et al. Histone recognition and large-scale structural analysis of the human bromodomain family. Cell. 2012;149:214–31. doi: 10.1016/j.cell.2012.02.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Ruthenburg AJ, Li H, Milne TA, Dewell S, McGinty RK, Yuen M, et al. Recognition of a mononucleosomal histone modification pattern by BPTF via multivalent interactions. Cell. 2011;145:692–706. doi: 10.1016/j.cell.2011.03.053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Zhou Y, Schmitz KM, Mayer C, Yuan X, Akhtar A, Grummt I. Reversible acetylation of the chromatin remodelling complex NoRC is required for non-coding RNA-dependent silencing. Nat Cell Biol. 2009;11:1010–6. doi: 10.1038/ncb1914. [DOI] [PubMed] [Google Scholar]
- 31.Singh M, Popowicz GM, Krajewski M, Holak TA. Structural ramification for acetyl-lysine recognition by the bromodomain of human BRG1 protein, a central ATPase of the SWI/SNF remodeling complex. Chembiochem. 2007;8:1308–16. doi: 10.1002/cbic.200600562. [DOI] [PubMed] [Google Scholar]
- 32.Nagy Z, Tora L. Distinct GCN5/PCAF-containing complexes function as co-activators and are involved in transcription factor and global histone acetylation. Oncogene. 2007;26:5341–57. doi: 10.1038/sj.onc.1210604. [DOI] [PubMed] [Google Scholar]
- 33.Milne TA, Briggs SD, Brock HW, Martin ME, Gibbs D, Allis CD, et al. MLL targets SET domain methyltransferase activity to Hox gene promoters. Mol Cell. 2002;10:1107–17. doi: 10.1016/S1097-2765(02)00741-4. [DOI] [PubMed] [Google Scholar]
- 34.Fujisawa T, Filippakopoulos P. Functions of bromodomain-containing proteins and their roles in homeostasis and cancer. Nat Rev Mol Cell Biol. 2017;18:246–62. doi: 10.1038/nrm.2016.143. [DOI] [PubMed] [Google Scholar]
- 35.Malovannaya A, Lanz RB, Jung SY, Bulynko Y, Le NT, Chan DW, et al. Analysis of the human endogenous coregulator complexome. Cell. 2011;145:787–99. doi: 10.1016/j.cell.2011.05.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Tsai WW, Wang Z, Yiu TT, Akdemir KC, Xia W, Winter S, et al. TRIM24 links a non-canonical histone signature to breast cancer. Nature. 2010;468:927–32. doi: 10.1038/nature09542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Wang CY, Filippakopoulos P. Beating the odds: BETs in disease. Trends Biochem Sci. 2015;40:468–79. doi: 10.1016/j.tibs.2015.06.002. [DOI] [PubMed] [Google Scholar]
- 38.Zeng L, Zhou MM. Bromodomain: an acetyl-lysine binding domain. FEBS Lett. 2002;513:124–8. doi: 10.1016/S0014-5793(01)03309-9. [DOI] [PubMed] [Google Scholar]
- 39.Owen DJ, Ornaghi P, Yang JC, Lowe N, Evans PR, Ballario P, et al. The structural basis for the recognition of acetylated histone H4 by the bromodomain of histone acetyltransferase gcn5p. EMBO J. 2000;19:6141–9. doi: 10.1093/emboj/19.22.6141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Vidler LR, Brown N, Knapp S, Hoelder S. Druggability analysis and structural classification of bromodomain acetyl-lysine binding sites. J Med Chem. 2012;55:7346–59. doi: 10.1021/jm300346w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Yang CY, Qin C, Bai L, Wang S. Small-molecule PROTAC degraders of the Bromodomain and Extra Terminal (BET) proteins - a review. Drug Discov Today Technol. 2019;31:43–51. doi: 10.1016/j.ddtec.2019.04.001. [DOI] [PubMed] [Google Scholar]
- 42.Filippakopoulos P, Qi J, Picaud S, Shen Y, Smith WB, Fedorov O, et al. Selective inhibition of BET bromodomains. Nature. 2010;468:1067–73. doi: 10.1038/nature09504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Grayson AR, Walsh EM, Cameron MJ, Godec J, Ashworth T, Ambrose JM, et al. MYC, a downstream target of BRD-NUT, is necessary and sufficient for the blockade of differentiation in NUT midline carcinoma. Oncogene. 2014;33:1736–42. doi: 10.1038/onc.2013.126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Qiu H, Jackson AL, Kilgore JE, Zhong Y, Chan LL, Gehrig PA, et al. JQ1 suppresses tumor growth through downregulating LDHA in ovarian cancer. Oncotarget. 2015;6:6915–30. doi: 10.18632/oncotarget.3126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Venkataraman S, Alimova I, Balakrishnan I, Harris P, Birks DK, Griesinger A, et al. Inhibition of BRD4 attenuates tumor cell self-renewal and suppresses stem cell signaling in MYC driven medulloblastoma. Oncotarget. 2014;5:2355–71. doi: 10.18632/oncotarget.1659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Dawson MA, Prinjha RK, Dittmann A, Giotopoulos G, Bantscheff M, Chan WI, et al. Inhibition of BET recruitment to chromatin as an effective treatment for MLL-fusion leukaemia. Nature. 2011;478:529–33. doi: 10.1038/nature10509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Ciró M, Prosperini E, Quarto M, Grazini U, Walfridsson J, McBlane F, et al. ATAD2 is a novel cofactor for MYC, overexpressed and amplified in aggressive tumors. Cancer Res. 2009;69:8491–8. doi: 10.1158/0008-5472.CAN-09-2131. [DOI] [PubMed] [Google Scholar]
- 48.Sima X, He J, Peng J, Xu Y, Zhang F, Deng L. The genetic alteration spectrum of the SWI/SNF complex: the oncogenic roles of BRD9 and ACTL6A. PLoS One. 2019;14:e0222305. doi: 10.1371/journal.pone.0222305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Klauke K, Radulović V, Broekhuis M, Weersing E, Zwart E, Olthof S, et al. Polycomb Cbx family members mediate the balance between haematopoietic stem cell self-renewal and differentiation. Nat Cell Biol. 2013;15:353–62. doi: 10.1038/ncb2701. [DOI] [PubMed] [Google Scholar]
- 50.Lloyd JT, Glass KC. Biological function and histone recognition of family IV bromodomain-containing proteins. J Cell Physiol. 2018;233:1877–86. doi: 10.1002/jcp.26010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Li S, Kuo HD, Yin R, Wu R, Liu X, Wang L, et al. Epigenetics/epigenomics of triterpenoids in cancer prevention and in health. Biochem Pharmacol. 2020;175:113890. doi: 10.1016/j.bcp.2020.113890. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Iizuka M, Takahashi Y, Mizzen CA, Cook RG, Fujita M, Allis CD, et al. Histone acetyltransferase Hbo1: catalytic activity, cellular abundance, and links to primary cancers. Gene. 2009;436:108–14. doi: 10.1016/j.gene.2009.01.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.LaMorte VJ, Dyck JA, Ochs RL, Evans RM. Localization of nascent RNA and CREB binding protein with the PML-containing nuclear body. Proc Natl Acad Sci USA. 1998;95:4991–6. doi: 10.1073/pnas.95.9.4991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Kaeser MD, Aslanian A, Dong MQ, Yates JR, 3rd, Emerson BM. BRD7, a novel PBAF-specific SWI/SNF subunit, is required for target gene activation and repression in embryonic stem cells. J Biol Chem. 2008;283:32254–63. doi: 10.1074/jbc.M806061200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Wu WJ, Hu KS, Chen DL, Zeng ZL, Luo HY, Wang F, et al. Prognostic relevance of BRD7 expression in colorectal carcinoma. Eur J Clin Invest. 2013;43:131–40. doi: 10.1111/eci.12024. [DOI] [PubMed] [Google Scholar]
- 56.Inoue D, Chew GL, Liu B, Michel BC, Pangallo J, D'Avino AR, et al. Spliceosomal disruption of the non-canonical BAF complex in cancer. Nature. 2019;574:432–6. doi: 10.1038/s41586-019-1646-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.The Human Protein Atlas, author. [Accessed September 1, 2020];Searches for BRD9 and BRD7. https://www.proteinatlas.org .
- 58.Chiu YH, Lee JY, Cantley LC. BRD7, a tumor suppressor, interacts with p85α and regulates PI3K activity. Mol Cell. 2014;54:193–202. doi: 10.1016/j.molcel.2014.02.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Drost J, Mantovani F, Tocco F, Elkon R, Comel A, Holstege H, et al. BRD7 is a candidate tumour suppressor gene required for p53 function. Nat Cell Biol. 2010;12:380–9. doi: 10.1038/ncb2038. [DOI] [PubMed] [Google Scholar]
- 60.Clark PG, Vieira LC, Tallant C, Fedorov O, Singleton DC, Rogers CM, et al. LP99: discovery and synthesis of the first selective BRD7/9 bromodomain inhibitor. Angew Chem Int Ed Engl. 2015;54:6217–21. doi: 10.1002/anie.201501394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Karim RM, Chan A, Zhu JY, Schönbrunn E. Structural basis of inhibitor selectivity in the BRD7/9 subfamily of bromodomains. J Med Chem. 2020;63:3227–37. doi: 10.1021/acs.jmedchem.9b01980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Neuwald AF, Aravind L, Spouge JL, Koonin EV. AAA+: a class of chaperone-like ATPases associated with the assembly, operation, and disassembly of protein complexes. Genome Res. 1999;9:27–43. [PubMed] [Google Scholar]
- 63.Altintas DM, Shukla MS, Goutte-Gattat D, Angelov D, Rouault JP, Dimitrov S, et al. Direct cooperation between androgen receptor and E2F1 reveals a common regulation mechanism for androgen-responsive genes in prostate cells. Mol Endocrinol. 2012;26:1531–41. doi: 10.1210/me.2012-1016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Boussouar F, Jamshidikia M, Morozumi Y, Rousseaux S, Khochbin S. Malignant genome reprogramming by ATAD2. Biochim Biophys Acta. 2013;1829:1010–4. doi: 10.1016/j.bbagrm.2013.06.003. [DOI] [PubMed] [Google Scholar]
- 65.Hussain M, Zhou Y, Song Y, Hameed HMA, Jiang H, Tu Y, et al. ATAD2 in cancer: a pharmacologically challenging but tractable target. Expert Opin Ther Targets. 2018;22:85–96. doi: 10.1080/14728222.2018.1406921. [DOI] [PubMed] [Google Scholar]
- 66.The Human Protein Atlas, author. [Accessed September 1, 2020];ATAD2B. https://www.proteinatlas.org/ENSG00000119778-ATAD2B/pathology/renal+cancer .
- 67.Blus BJ, Wiggins K, Khorasanizadeh S. Epigenetic virtues of chromodomains. Crit Rev Biochem Mol Biol. 2011;46:507–26. doi: 10.3109/10409238.2011.619164. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Zocco M, Marasovic M, Pisacane P, Bilokapic S, Halic M. The Chp1 chromodomain binds the H3K9me tail and the nucleosome core to assemble heterochromatin. Cell Discov. 2016;2:16004. doi: 10.1038/celldisc.2016.4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Taverna SD, Coyne RS, Allis CD. Methylation of histone h3 at lysine 9 targets programmed DNA elimination in tetrahymena. Cell. 2002;110:701–11. doi: 10.1016/S0092-8674(02)00941-8. [DOI] [PubMed] [Google Scholar]
- 70.Sapountzi V, Côté J. MYST-family histone acetyltransferases: beyond chromatin. Cell Mol Life Sci. 2011;68:1147–56. doi: 10.1007/s00018-010-0599-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Gayatri S, Bedford MT. Readers of histone methylarginine marks. Biochim Biophys Acta. 2014;1839:702–10. doi: 10.1016/j.bbagrm.2014.02.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Morettin A, Paris G, Bouzid Y, Baldwin RM, Falls TJ, Bell JC, et al. Tudor domain containing protein 3 promotes tumorigenesis and invasive capacity of breast cancer cells. Sci Rep. 2017;7:5153. doi: 10.1038/s41598-017-04955-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Mulligan P, Westbrook TF, Ottinger M, Pavlova N, Chang B, Macia E, et al. CDYL bridges REST and histone methyltransferases for gene repression and suppression of cellular transformation. Mol Cell. 2008;32:718–26. doi: 10.1016/j.molcel.2008.10.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Forzati F, Federico A, Pallante P, Fedele M, Fusco A. Tumor suppressor activity of CBX7 in lung carcinogenesis. Cell Cycle. 2012;11:1888–91. doi: 10.4161/cc.20022. [DOI] [PubMed] [Google Scholar]
- 75.Karamitopoulou E, Pallante P, Zlobec I, Tornillo L, Carafa V, Schaffner T, et al. Loss of the CBX7 protein expression correlates with a more aggressive phenotype in pancreatic cancer. Eur J Cancer. 2010;46:1438–44. doi: 10.1016/j.ejca.2010.01.033. [DOI] [PubMed] [Google Scholar]
- 76.Pallante P, Sepe R, Federico A, Forzati F, Bianco M, Fusco A. CBX7 modulates the expression of genes critical for cancer progression. PLoS One. 2014;9:e98295. doi: 10.1371/journal.pone.0098295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Ginder GD, Williams DC., Jr Readers of DNA methylation, the MBD family as potential therapeutic targets. Pharmacol Ther. 2018;184:98–111. doi: 10.1016/j.pharmthera.2017.11.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Teske KA, Hadden MK. Methyllysine binding domains: structural insight and small molecule probe development. Eur J Med Chem. 2017;136:14–35. doi: 10.1016/j.ejmech.2017.04.047. [DOI] [PubMed] [Google Scholar]
- 79.Pervaiz M, Mishra P, Günther S. Bromodomain drug discovery - the past, the present, and the future. Chem Rec. 2018;18:1808–17. doi: 10.1002/tcr.201800074. [DOI] [PubMed] [Google Scholar]
- 80.Burkard M, Leischner C, Lauer UM, Busch C, Venturelli S, Frank J. Dietary flavonoids and modulation of natural killer cells: implications in malignant and viral diseases. J Nutr Biochem. 2017;46:1–12. doi: 10.1016/j.jnutbio.2017.01.006. [DOI] [PubMed] [Google Scholar]
- 81.Khuda-Bukhsh AR, Das S, Saha SK. Molecular approaches toward targeted cancer prevention with some food plants and their products: inflammatory and other signal pathways. Nutr Cancer. 2014;66:194–205. doi: 10.1080/01635581.2014.864420. [DOI] [PubMed] [Google Scholar]
- 82.Surh YJ. Cancer chemoprevention with dietary phytochemicals. Nat Rev Cancer. 2003;3:768–80. doi: 10.1038/nrc1189. [DOI] [PubMed] [Google Scholar]
- 83.Lampe JW. Diet and cancer prevention research: from mechanism to implementation. J Cancer Prev. 2020;25:65–9. doi: 10.15430/JCP.2020.25.2.65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Ko EY, Moon A. Natural products for chemoprevention of breast cancer. J Cancer Prev. 2015;20:223–31. doi: 10.15430/JCP.2015.20.4.223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Parasramka MA, Dashwood WM, Wang R, Abdelli A, Bailey GS, Williams DE, et al. MicroRNA profiling of carcinogen-induced rat colon tumors and the influence of dietary spinach. Mol Nutr Food Res. 2012;56:1259–69. doi: 10.1002/mnfr.201200117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Parasramka MA, Ho E, Williams DE, Dashwood RH. MicroRNAs, diet, and cancer: new mechanistic insights on the epigenetic actions of phytochemicals. Mol Carcinog. 2012;51:213–30. doi: 10.1002/mc.20822. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Schnekenburger M, Dicato M, Diederich M. Plant-derived epigenetic modulators for cancer treatment and prevention. Biotechnol Adv. 2014;32:1123–32. doi: 10.1016/j.biotechadv.2014.03.009. [DOI] [PubMed] [Google Scholar]
- 88.Beaver LM, Yu TW, Sokolowski EI, Williams DE, Dashwood RH, Ho E. 3,3'-Diindolylmethane, but not indole-3-carbinol, inhibits histone deacetylase activity in prostate cancer cells. Toxicol Appl Pharmacol. 2012;263:345–51. doi: 10.1016/j.taap.2012.07.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Myzak MC, Dashwood WM, Orner GA, Ho E, Dashwood RH. Sulforaphane inhibits histone deacetylase in vivo and suppresses tumorigenesis in Apc-minus mice. FASEB J. 2006;20:506–8. doi: 10.1096/fj.05-4785fje. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Nian H, Delage B, Ho E, Dashwood RH. Modulation of histone deacetylase activity by dietary isothiocyanates and allyl sulfides: studies with sulforaphane and garlic organosulfur compounds. Environ Mol Mutagen. 2009;50:213–21. doi: 10.1002/em.20454. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Rajendran P, Williams DE, Ho E, Dashwood RH. Metabolism as a key to histone deacetylase inhibition. Crit Rev Biochem Mol Biol. 2011;46:181–99. doi: 10.3109/10409238.2011.557713. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Kim D, Kim Y, Kim Y. Effects of β-carotene on expression of selected microRNAs, histone acetylation, and DNA methylation in colon cancer stem cells. J Cancer Prev. 2019;24:224–32. doi: 10.15430/JCP.2019.24.4.224. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Alumkal JJ, Slottke R, Schwartzman J, Cherala G, Munar M, Graff JN, et al. A phase II study of sulforaphane-rich broccoli sprout extracts in men with recurrent prostate cancer. Invest New Drugs. 2015;33:480–9. doi: 10.1007/s10637-014-0189-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Atwell LL, Hsu A, Wong CP, Stevens JF, Bella D, Yu TW, et al. Absorption and chemopreventive targets of sulforaphane in humans following consumption of broccoli sprouts or a myrosinase-treated broccoli sprout extract. Mol Nutr Food Res. 2015;59:424–33. doi: 10.1002/mnfr.201400674. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Bultman SJ. Interplay between diet, gut microbiota, epigenetic events, and colorectal cancer. Mol Nutr Food Res. 2017;61:10.1002/mnfr.201500902. doi: 10.1002/mnfr.201500902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Jones PA, Issa JP, Baylin S. Targeting the cancer epigenome for therapy. Nat Rev Genet. 2016;17:630–41. doi: 10.1038/nrg.2016.93. [DOI] [PubMed] [Google Scholar]
- 97.Myzak MC, Tong P, Dashwood WM, Dashwood RH, Ho E. Sulforaphane retards the growth of human PC-3 xenografts and inhibits HDAC activity in human subjects. Exp Biol Med (Maywood) 2007;232:227–34. [PMC free article] [PubMed] [Google Scholar]
- 98.Sapienza C, Issa JP. Diet, nutrition, and cancer epigenetics. Annu Rev Nutr. 2016;36:665–81. doi: 10.1146/annurev-nutr-121415-112634. [DOI] [PubMed] [Google Scholar]
- 99.Schnekenburger M, Dicato M, Diederich M. Epigenetic modulators from "The Big Blue": a treasure to fight against cancer. Cancer Lett. 2014;351:182–97. doi: 10.1016/j.canlet.2014.06.005. [DOI] [PubMed] [Google Scholar]
- 100.Seidel C, Schnekenburger M, Dicato M, Diederich M. Histone deacetylase modulators provided by Mother Nature. Genes Nutr. 2012;7:357–67. doi: 10.1007/s12263-012-0283-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Balasubramanyam K, Varier RA, Altaf M, Swaminathan V, Siddappa NB, Ranga U, et al. Curcumin, a novel p300/CREB-binding protein-specific inhibitor of acetyltransferase, represses the acetylation of histone/nonhistone proteins and histone acetyltransferase-dependent chromatin transcription. J Biol Chem. 2004;279:51163–71. doi: 10.1074/jbc.M409024200. [DOI] [PubMed] [Google Scholar]
- 102.Prieto-Martínez FD, Medina-Franco JL. Flavonoids as putative epi-modulators: insight into their binding mode with BRD4 bromodomains using molecular docking and dynamics. Biomolecules. 2018;8:61. doi: 10.3390/biom8030061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Raj U, Kumar H, Varadwaj PK. Molecular docking and dynamics simulation study of flavonoids as BET bromodomain inhibitors. J Biomol Struct Dyn. 2017;35:2351–62. doi: 10.1080/07391102.2016.1217276. [DOI] [PubMed] [Google Scholar]
- 104.Dhananjayan K. Molecular docking study characterization of rare flavonoids at the Nac-binding site of the first bromodomain of BRD4 (BRD4 BD1) J Cancer Res. 2015;2015:762716. doi: 10.1155/2015/762716. [DOI] [Google Scholar]
- 105.Hsu FT, Chiang IT, Kuo YC, Hsia TC, Lin CC, Liu YC, et al. Amentoflavone effectively blocked the tumor progression of glioblastoma via suppression of ERK/NF- κ B signaling pathway. Am J Chin Med. 2019;47:913–31. doi: 10.1142/S0192415X19500484. [DOI] [PubMed] [Google Scholar]
- 106.Kashyap D, Sharma A, Sak K, Tuli HS, Buttar HS, Bishayee A. Fisetin: a bioactive phytochemical with potential for cancer prevention and pharmacotherapy. Life Sci. 2018;194:75–87. doi: 10.1016/j.lfs.2017.12.005. [DOI] [PubMed] [Google Scholar]
- 107.Lee WJ, Shim JY, Zhu BT. Mechanisms for the inhibition of DNA methyltransferases by tea catechins and bioflavonoids. Mol Pharmacol. 2005;68:1018–30. doi: 10.1124/mol.104.008367. [DOI] [PubMed] [Google Scholar]
- 108.Kim HJ, Kim SH, Yun JM. Fisetin inhibits hyperglycemia-induced proinflammatory cytokine production by epigenetic mechanisms. Evid Based Complement Alternat Med. 2012;2012:639469. doi: 10.1155/2012/639469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Farhan M, Ullah MF, Faisal M, Farooqi AA, Sabitaliyevich UY, Biersack B, et al. Differential methylation and acetylation as the epigenetic basis of resveratrol's anticancer activity. Medicines (Basel) 2019;6:24. doi: 10.3390/medicines6010024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Schultz MB, Rinaldi C, Lu Y, Amorim JA, Sinclair DA. Molecular and cellular characterization of SIRT1 allosteric activators. Methods Mol Biol. 2019;1983:133–49. doi: 10.1007/978-1-4939-9434-2_8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Lin HY, Tang HY, Davis FB, Davis PJ. Resveratrol and apoptosis. Ann N Y Acad Sci. 2011;1215:79–88. doi: 10.1111/j.1749-6632.2010.05846.x. [DOI] [PubMed] [Google Scholar]
- 112.Dutra LA, Heidenreich D, Silva GDBD, Man Chin C, Knapp S, Santos JLD. Dietary compound resveratrol is a pan-BET bromodomain inhibitor. Nutrients. 2017;9:1172. doi: 10.3390/nu9111172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Yokoyama T, Matsumoto K, Ostermann A, Schrader TE, Nabeshima Y, Mizuguchi M. Structural and thermodynamic characterization of the binding of isoliquiritigenin to the first bromodomain of BRD4. FEBS J. 2019;286:1656–67. doi: 10.1111/febs.14736. [DOI] [PubMed] [Google Scholar]
- 114.Peng F, Du Q, Peng C, Wang N, Tang H, Xie X, et al. A review: the pharmacology of isoliquiritigenin. Phytother Res. 2015;29:969–77. doi: 10.1002/ptr.5348. [DOI] [PubMed] [Google Scholar]
- 115.Pham TND, Stempel S, Shields MA, Spaulding C, Kumar K, Bentrem DJ, et al. Quercetin enhances the anti-tumor effects of BET inhibitors by suppressing hnRNPA1. Int J Mol Sci. 2019;20:4293. doi: 10.3390/ijms20174293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Mustafi S, Camarena V, Volmar CH, Huff TC, Sant DW, Brothers SP, et al. Vitamin C sensitizes melanoma to BET inhibitors. Cancer Res. 2018;78:572–83. doi: 10.1158/0008-5472.CAN-17-2040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Muller S, Filippakopoulos P, Knapp S. Bromodomains as therapeutic targets. Expert Rev Mol Med. 2011;13:e29. doi: 10.1017/S1462399411001992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Wei Z, Yoshihara E, He N, Hah N, Fan W, Pinto AFM, et al. Vitamin D switches BAF complexes to protect β cells. Cell. 2018;173:1135–49.e15. doi: 10.1016/j.cell.2018.04.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Sanderson SM, Gao X, Dai Z, Locasale JW. Methionine metabolism in health and cancer: a nexus of diet and precision medicine. Nat Rev Cancer. 2019;19:625–37. doi: 10.1038/s41568-019-0187-8. [DOI] [PubMed] [Google Scholar]
- 120.Sdelci S, Rendeiro AF, Rathert P, You W, Lin JG, Ringler A, et al. MTHFD1 interaction with BRD4 links folate metabolism to transcriptional regulation. Nat Genet. 2019;51:990–8. doi: 10.1038/s41588-019-0413-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Pikman Y, Puissant A, Alexe G, Furman A, Chen LM, Frumm SM, et al. Targeting MTHFD2 in acute myeloid leukemia. J Exp Med. 2016;213:1285–306. doi: 10.1084/jem.20151574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Kang C, Kim CY, Kim HS, Park SP, Chung HM. The bromodomain inhibitor JQ1 enhances the responses to all-trans retinoic acid in HL-60 and MV4-11 leukemia cells. Int J Stem Cells. 2018;11:131–40. doi: 10.15283/ijsc18021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.de Thé H. Differentiation therapy revisited. Nat Rev Cancer. 2018;18:117–27. doi: 10.1038/nrc.2017.103. [DOI] [PubMed] [Google Scholar]
- 124.Uray IP, Dmitrovsky E, Brown PH. Retinoids and rexinoids in cancer prevention: from laboratory to clinic. Semin Oncol. 2016;43:49–64. doi: 10.1053/j.seminoncol.2015.09.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Kim YH, Kim M, Yoo M, Kim JE, Lee HK, Heo JN, et al. A natural compound, aristoyagonine, is identified as a potent bromodomain inhibitor by mid-throughput screening. Biochem Biophys Res Commun. 2018;503:882–7. doi: 10.1016/j.bbrc.2018.06.091. [DOI] [PubMed] [Google Scholar]
- 126.Lucas X, Wohlwend D, Hügle M, Schmidtkunz K, Gerhardt S, Schüle R, et al. 4-Acyl pyrroles: mimicking acetylated lysines in histone code reading. Angew Chem Int Ed Engl. 2013;52:14055–9. doi: 10.1002/anie.201307652. [DOI] [PubMed] [Google Scholar]
- 127.McLoughlin EC, O'Boyle NM. Colchicine-binding site inhibitors from chemistry to clinic: a review. Pharmaceuticals (Basel) 2020;13:8. doi: 10.3390/ph13010008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Pereira ER, Belin L, Sancelme M, Prudhomme M, Ollier M, Rapp M, et al. Structure-activity relationships in a series of substituted indolocarbazoles: topoisomerase I and protein kinase C inhibition and antitumoral and antimicrobial properties. J Med Chem. 1996;39:4471–7. doi: 10.1021/jm9603779. [DOI] [PubMed] [Google Scholar]
- 129.Salas JA, Méndez C. Indolocarbazole antitumour compounds by combinatorial biosynthesis. Curr Opin Chem Biol. 2009;13:152–60. doi: 10.1016/j.cbpa.2009.02.003. [DOI] [PubMed] [Google Scholar]
- 130.Cheng X, Zhou B, Liu H, Huo C, Ding W. One new indolocarbazole alkaloid from the Streptomyces sp. A22. Nat Prod Res. 2018;32:2583–8. doi: 10.1080/14786419.2018.1428595. [DOI] [PubMed] [Google Scholar]
- 131.Yu L, Ding W, Wang Q, Ma Z, Xu X, Zhao X, et al. Induction of cryptic bioactive 2,5-diketopiperazines in fungus Penicillium sp. DT-F29 by microbial co-culture. Tetrahedron. 2017;73:907–14. doi: 10.1016/j.tet.2016.12.077. [DOI] [Google Scholar]
- 132.Sumiyoshi M, Kimura Y. Enhancing effects of a chromone glycoside, eucryphin, isolated from Astilbe rhizomes on burn wound repair and its mechanism. Phytomedicine. 2010;17:820–9. doi: 10.1016/j.phymed.2010.01.010. [DOI] [PubMed] [Google Scholar]
- 133.Na M, Cui L, Min BS, Bae K, Yoo JK, Kim BY, et al. Protein tyrosine phosphatase 1B inhibitory activity of triterpenes isolated from Astilbe koreana. Bioorg Med Chem Lett. 2006;16:3273–6. doi: 10.1016/j.bmcl.2006.03.036. [DOI] [PubMed] [Google Scholar]
- 134.Shi Y, Zhang D, Li S, Xuan X, Zhang L, Li Y, et al. Inhibitors of BRD4 protein from the roots of Astilbe grandis stapf ex E.H. Wilson [published online ahead of print August 22, 2019] Nat Prod Res. :doi: 10.1080/14786419.2019.1655414. doi: 10.1080/14786419.2019.1655414. [DOI] [PubMed] [Google Scholar]
- 135.Di Micco S, Pulvirenti L, Bruno I, Terracciano S, Russo A, Vaccaro MC, et al. Identification by Inverse Virtual Screening of magnolol-based scaffold as new tankyrase-2 inhibitors. Bioorg Med Chem. 2018;26:3953–7. doi: 10.1016/j.bmc.2018.06.019. [DOI] [PubMed] [Google Scholar]
- 136.Ma Q, Ding W, Chen Z, Ma Z. Bisamides and rhamnosides from mangrove actinomycete Streptomyces sp. SZ-A15. Nat Prod Res. 2018;32:761–6. doi: 10.1080/14786419.2017.1315578. [DOI] [PubMed] [Google Scholar]
- 137.Ding H, Zhang D, Zhou B, Ma Z. Inhibitors of BRD4 protein from a marine-derived fungus Alternaria sp. NH-F6. Mar Drugs. 2017;15:76. doi: 10.3390/md15030076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Wu X, Chen Z, Ding W, Liu Y, Ma Z. Chemical constituents of the fermentative extracts of marine fungi Phoma sp. CZD-F11 and Aspergillus sp. CZD-F18 from Zhoushan Archipelago, China. Nat Prod Res. 2018;32:1562–6. doi: 10.1080/14786419.2017.1389929. [DOI] [PubMed] [Google Scholar]
- 139.Cassidy A, Minihane AM. The role of metabolism (and the microbiome) in defining the clinical efficacy of dietary flavonoids. Am J Clin Nutr. 2017;105:10–22. doi: 10.3945/ajcn.116.136051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Del Rio D, Rodriguez-Mateos A, Spencer JP, Tognolini M, Borges G, Crozier A. Dietary (poly)phenolics in human health: structures, bioavailability, and evidence of protective effects against chronic diseases. Antioxid Redox Signal. 2013;18:1818–92. doi: 10.1089/ars.2012.4581. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Manach C, Williamson G, Morand C, Scalbert A, Rémésy C. Bioavailability and bioefficacy of polyphenols in humans. I. Review of 97 bioavailability studies. Am J Clin Nutr. 2005;81(1 Suppl):230S–242S. doi: 10.1093/ajcn/81.1.230S. [DOI] [PubMed] [Google Scholar]
- 142.Albini A, Bassani B, Baci D, Dallaglio K, Gallazzi M, Corradino P, et al. Nutraceuticals and "repurposed" drugs of phytochemical origin in prevention and interception of chronic degenerative diseases and cancer. Curr Med Chem. 2019;26:973–87. doi: 10.2174/0929867324666170920144130. [DOI] [PubMed] [Google Scholar]
- 143.Lippman SM, Abate-Shen C, Colbert Maresso KL, Colditz GA, Dannenberg AJ, Davidson NE, et al. AACR white paper: shaping the future of cancer prevention - a roadmap for advancing science and public health. Cancer Prev Res (Phila) 2018;11:735–78. doi: 10.1158/1940-6207.CAPR-18-0421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144.Traka MH, Melchini A, Mithen RF. Sulforaphane and prostate cancer interception. Drug Discov Today. 2014;19:1488–92. doi: 10.1016/j.drudis.2014.07.007. [DOI] [PubMed] [Google Scholar]
- 145.Vasan N, Baselga J, Hyman DM. A view on drug resistance in cancer. Nature. 2019;575:299–309. doi: 10.1038/s41586-019-1730-1. [DOI] [PMC free article] [PubMed] [Google Scholar]