Abstract
Osteosarcoma is an aggressive malignancy with rapid development and poor prognosis. microRNA-19 (miR-19) plays an important role in several biological processes. Sprouty-related EVH1 domain protein 2 (SPRED2) is a suppressor of extracellular signal-regulated kinase/mitogen-activated protein kinase (ERK/MAPK) signaling to inhibit tumor development and progression by promoting autophagy. In this study, we investigated the roles of miR-19, SPRED2, and autophagy in osteosarcoma. We detected the expression of miR-19, SPRED2, epithelial–mesenchymal transition (EMT) markers, and autophagy-related proteins via quantitative real-time polymerase chain reaction or western blot. To evaluate the function of miR-19 and SPRED2, we used MTT and colony formation assays to detect cell proliferation, Transwell, and wound-healing assays to detect cell invasion and migration. Targetscan and luciferase reporter assays confirmed the relationship between SPRED2 and miR-19. The expression of miR-19 was significantly upregulated in osteosarcoma, while SPRED2 was downregulated. miR-19 inhibitor reduced cell proliferation, invasion, migration, and EMT, while its cell biological effects were partially reversed by addition of autophagy inhibitor 3-methyladenine (3-MA) or SPRED2 siRNA in osteosarcoma. SPRED2, a suppressor of ERK/MAPK pathway that is known to trigger autophagy, was identified as a direct target of miR-19. SPRED2 overexpression increased cell proliferation, invasion, migration, and EMT by promoting autophagy, and the effects could be inhibited by 3-MA. Collectively, these findings reveal an underlying mechanism for development of osteosarcoma. miR-19 was upregulated in osteosarcoma cells, and negatively regulated SPRED2, thus promoting the malignant transformation of osteosarcoma cells via inhibiting SPRED2-induced autophagy. Therefore, miR-19/SPRED2 may be a potential target for the treatment of osteosarcoma.
Keywords: osteosarcoma, miR-19, SPRED2, autophagy, malignant phenotypes
Introduction
Osteosarcoma accounts for about 60% of all bone tumors and is a highly graded primary tumor with the highest incidence in children and adolescents1,2. Despite improvements in diagnosis and treatment, and less exposure to social risks, there is still a lack of effective approaches for relapse prevention, and patients with unresectable, recurrent, or metastatic cancers still have poor prognosis3. Since many patients do not respond well to current therapy regimens, investigating the mechanism involved is helpful to improve effective treatments for osteosarcoma.
MicroRNAs (miRNAs) are small noncoding RNAs (10 to 22 nucleotides) by interaction with the 3′-untranslated region (UTR) of target genes to negatively regulate gene expression, leading to mRNA degradation or translational suppression4. The abnormal expression of miRNAs is involved in the pathogenesis of many diseases including cancer. According to the different target genes, miRNAs can be used as tumor suppressors or oncogenes5–7. Numerous studies have shown that miRNAs are involved in the development of osteosarcoma8,9. miR-19 is a key member miRNA of miR-17-92 cluster, and plays an oncogenic role, including promotion of tumor growth, progression, metastasis, and drug resistance through upregulating the expression of oncogenes or downregulating tumor suppressor genes, in multiple malignancies10–12. In addition, a recent study has shown an association of upregulation of miR-19 expression with osteosarcoma development13. However, the effect of microRNA-19 (miR-19) on the biological characteristics of osteosarcoma cells and the mechanism of action are still unclear. Further studies are needed to identify additional miR-19 target genes to elucidate the role and the mechanism of miR-19 in osteosarcoma.
Autophagy is a process of transferring cytoplasmic contents to lysosomes for degradation through double-membrane vesicles, which can lead to cell survival or death14. In particular, autophagy is closely related to apoptosis, a kind of programmed cell death, and the function of the autophagy is crucial to cell homeostasis15,16. In fact, disorders of the autophagy is associated with many kinds of cancer17. Beclin-1 acts as an autophagy-related protein to form autophagosomes through interaction with bcl-2 of anti-apoptotic protein. Microtubule-associated protein 1 light chain 3 (LC3-I) is also involved in autophagy through combining with phosphatidyl ethanolamine to form LC3-II, which is collected into the autophagosome membrane and degraded after the fusion of autophagosome with lysosome. Therefore, the LC3-II/LC3-I ratio is also an indicator of autophagy activity18. In terms of mechanisms, mammalian target of rapamycin (mTOR) is a serine/threonine kinase, which is regarded as the main negative regulator of autophagy19. mTOR signaling is downregulated in many diseases as it regulates cell proliferation. Subsequently, mTOR phosphorylation can be stimulated by the phosphoinositide-3 kinase-mediated activation of protein kinase B (PI3K/AKT), and extracellular signal-regulated kinase 1/2/mitogen-activated protein kinase (ERK1/2/MAPK) signaling20. PI3K/AKT and ERK1/2/MAPK signaling can trigger autophagy through activating mTOR pathway.
Sprouty-related EVH1 domain (SPRED) proteins are membrane-associated suppressors of ERK/MAPK signaling and inhibit its activation through a series of mitogenic stimuli such as chemokines, growth factors, and cytokines21,22. SPREDs contain an N-terminal EVH1 domain, a central c-kit binding domain, and a C-terminal cysteine-rich Sprouty-related domain (SPR). Four mammalian SPRED isoforms have been studied. Recent studies have proved that Spred2 is regarded as a suppressor to impair tumor development and progression in many types of cancer, due to reduced ERK/adenosine monophosphate-activated protein kinase activity to trigger autophagy23. Sprouty2 is an effective suppressor of cell proliferation and migration in osteosarcoma cells, while Sprouty4 is not. However, the potential mechanism of SPRED2 in the occurrence and development of osteosarcoma has been poorly studied24. In this study, online software (http://www.targetscan.org) predicts that SPRED2 may be the target gene by miR-19 regulation. Therefore, we demonstrated the correction between miR-19, autophagy, and SPRED2 in osteosarcoma. The mechanism of miR-19 on the biological functions of osteosarcoma cells was investigated through autophagy inhibitor or knockdown SPRED2 expression in osteosarcoma cells. Rescue experiment proved that miR-19 promoted malignant transformation through targeting SPRED2 to impair the autophagy. The study provides experimental basis for miR-19/SPRED2 axis application in osteosarcoma therapy.
Materials and Methods
Tissue Specimens
Tissue samples were collected from the Second Affiliated Hospital of Guangzhou Medical University, Guangzhou, China. All the patients participating in this study signed informed consent. All experiments were performed in accordance with government policy and the Helsinki declaration.
Cell Culture, Reagents, and Antibodies
Mixed medium of 10% fetal bovine serum (FBS) (Dulbecco’s modified Eagle medium and F-12 mixed by 1:1) is used to culture normal human osteoblasts hFOB cells. MEM of 10% FBS is used to culture human osteosarcoma MG-63 and SOSP-9607cells. McCoy’s 5A medium 10% FBS is used to culture Saos-2 and U2OS cells. The above cells were incubated at 37°C in 5% CO2. The cell lines were all obtained from the Chinese Academy of Sciences Cell Bank (Beijing, China). The FBS and medium were purchased from Invitrogen (Carlsbad, CA, USA). 3-Methyladenine (3-MA) was purchased from Sigma (St Louis, MO, USA). SPRED2, Beclin-1, LC3, E-cadherin, N-cadherin, vimentin, and glyceraldehyde 3-phosphate dehydrogenase (GAPDH) antibodies were purchased from Abcam (Cambridge, MA, USA).
Cell Transfection
miR-19 inhibitor (5′-TCAGTTTTGCCCTATTTGCACA-3′), miR-19 mimics (5′-UGUGCAAAUCCAUGCAAAACUGA-3′), and SPRED2 siRNA (5′-AAGGACTTGGTCTACACCAAA-3′) were synthesized by GenePharma (Shanghai, China). Cells were transfected according to the manufacturer’s instructions of Lipofectamine 3000 (Invitrogen, Thermo Fisher Scientific, Inc., Waltham, MA, USA). SPRED2 was overexpressed using pLKO lentiviral vectors. After transfection of MG-63 and SOSP-9607 cells with the virus for 48 h, puromycin was added to the culture medium for screening stable clones.
Quantitative Real-Time Polymerase Chain Reaction
The mRNA expressions of SPRED2 and miR-19 were detected by quantitative real-time polymerase chain reaction (qRT-PCR). Extraction of total RNA from tissues and cell lines was done using Trizol reagent (Thermo Fisher Scientific). Reverse transcription of mRNA was performed using PrimeScript RT reagent kit (TaKara, Dalian, China). The qRT-PCR experiment accorded to instructions of an SYBR Premix Ex Taq Kit (TaKaRa) and was performed on an ABI 7500 (Applied Biosystem, TaKara, Dalian, China). Primers used in the qRT-PCR analysis were as follows: SPRED2 forward: 5′-GAAGGTACCCGGACAGGAAGATGAAGGG-3′, SPRED2 reverse: 5′-GAACTCGAGAGGGAGAAGGGAGGGAAACT-3′; miR-19 forward: 5′-CATAGTTGCACTACAAGAAG-3′, miR-19 reverse: 5′-GCACAACTACATTCTTCTTG-3′; GAPDH Forward: 5′-CACGGCAAATTCAACGGCACAGTCAAGG-3′, GAPDH reverse: 5′-GTTCACACCCATCACAAACATGG-3′; U6 forward: 5′-CTCGCTTCGGCAGCACA-3′, U6 reverse: 5′-AACGCTTCACGAATTTGCGT-3′. A ratio relative to the GAPDH and U6 was used as internal control.
Western Blot
Tissue and cells were lysed in RIPA Lysis Buffer, and using the BCA Protein Assay Kit (Beyotime Biotechnology, TaKara, Dalian, China) to measure the protein concentration followed by separation on sodium dodecyl sulfate-polyacrylamide gel electrophoresis for western blot analysis of N-cadherin, E-cadherin, vimentin, LC3I/II, BECLIN1, and SPRED2 expression using relevant primary antibodies and horseradish peroxidase-labeled secondary antibodies. The membrane was exposed and developed after addition of enhanced chemiluminescence reagent. GAPDH was used as a control.
MTT Assay
Cells were seeded in 96-well plates at a density of 2 × 104 cells per well. After 12, 24, 48, or 72 h, each well was added MTT solution (0.5 mg/ml in 20 µl phosphate-buffered saline) and incubated at 37°C for 4 h, then the absorbance at 570 nm was measured. Data came from three independent experiments, each with sextuplicate.
Colony-Formation Assay
Cells were seeded in six-well plates at 2,000 cells per well. About 2 weeks, cell colonies were fixed with 4% paraformaldehyde, then stained with crystal violet.
Transwell Invasion Assays
For invasion assay, BD cell culture inserts (24-well insert, 8-µm pore size), which were coated Matrigel (BD Biosciences, Franklin Lakes, NJ, USA), were used following the manufacturer’s instructions. Two hundred microliters of cell suspension with serum-free medium was loaded into the upper chamber of the inserts, then 500 µl medium of 10% FBS was added to lower chambers. After incubation, cells on the upper layer of membrane were scraped by a cotton swab, then the membrane was fixed with 4% paraformaldehyde and stained with 0.5% crystal violet solution. Five random fields per filter were counted under a light microscope.
Dual-Luciferase Reporter Assay
SPRED2 3′-UTR was amplified using the following primers: forward: 5′-CCGCTCGAGTTGGACCAGGGCTTGCAA-3′, reverse: 5′-ATAAGAATGCGGCCGCGCAAGCCACCACTGTTTGAA-3′. The primers of the amplified SPRED2 mutant 3′-UTR were as follows: forward: 5′-GATTTGGGTTTCTGTGAGAGCTATC-3′, reverse: 5′-AGGCAAGAUGCUGGCAUAGCUG-3′. Cells cultured in 24-well plates were co-transfected with reporter plasmid and miR-19 mimics or negative control. After incubation, the firefly and Renilla luciferase activities were detected by the dual luciferase reporter assay system of Promega (Madison, WI, USA).
Statistical Analysis
Each experiment was performed at least three times. Statistical analyses were determined by Student’s t-test, and the differences between two groups or more than two groups were detected using analysis of variance. All data were expressed as mean ± standard deviation.
Results
1. miR-19 is highly expressed in osteosarcoma tissues and cells, while SPRED2 is poorly expressed
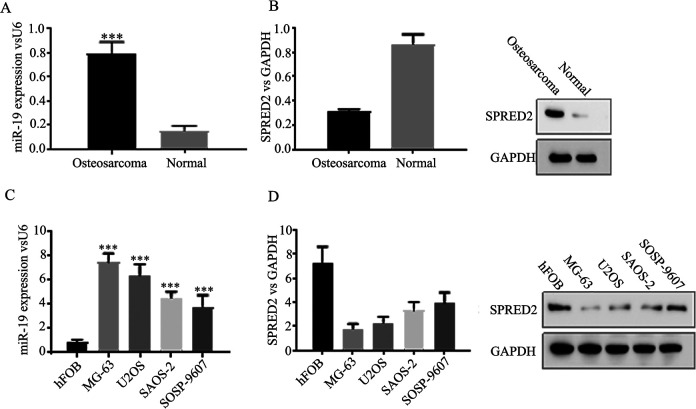
High expression of miR-19 is an important oncogenic event in many cancers, and miR-19 is the upstream miRNA of SPRED2 by target scanning. To investigate the relevance of miR-19 to SPRED2 expression, we used 30 osteosarcoma tissue and 30 normal specimens to analyze the expression of miR-19 and SPRED2. The expression of miR-19 in osteosarcoma tissue was higher than that in normal counterparts, whereas the mRNA and protein levels of SPRED2 in osteosarcoma were relatively lower than that in normal tissue (Fig. 1A, B). Furthermore, the expressions of miR-19 and SPRED2 were analyzed in human osteoblastic and osteosarcoma cell lines. As shown in Fig. 1C, D, in normal osteoblastic cell hFOB, miR-19 expression was relatively low, and on the contrary, SPRED2 expression was high. Meanwhile, we observed that the miR-19 expression was relatively high, which was negatively correlated with SPRED2 expression in osteosarcoma cell lines (Fig. 1C, D). Consistently, the high miR-19 expression and low SPRED2 expression were found in osteosarcoma, suggesting that SPRED2 may negatively correlate with miR-19 in osteosarcoma.
Figure 1.
The expression of miR-19 is upregulated, while SPRED2 is deregulated in osteosarcoma.
GAPDH: glyceraldehyde 3-phosphate dehydrogenase.
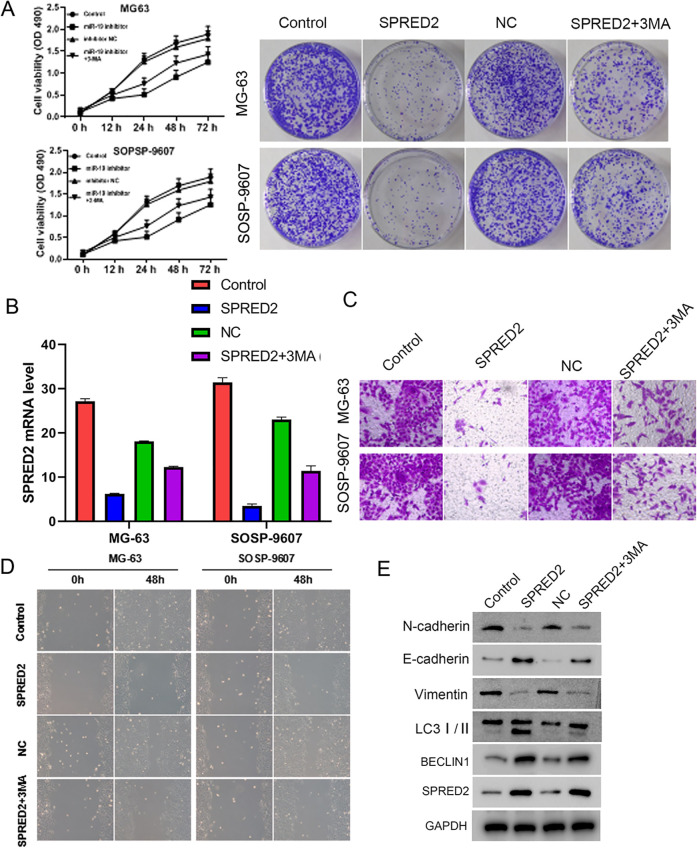
2. miR-19 inhibitor suppresses cell proliferation, invasion, migration, and epithelial–mesenchymal transition (EMT) via promoting autophagy in osteosarcoma cells
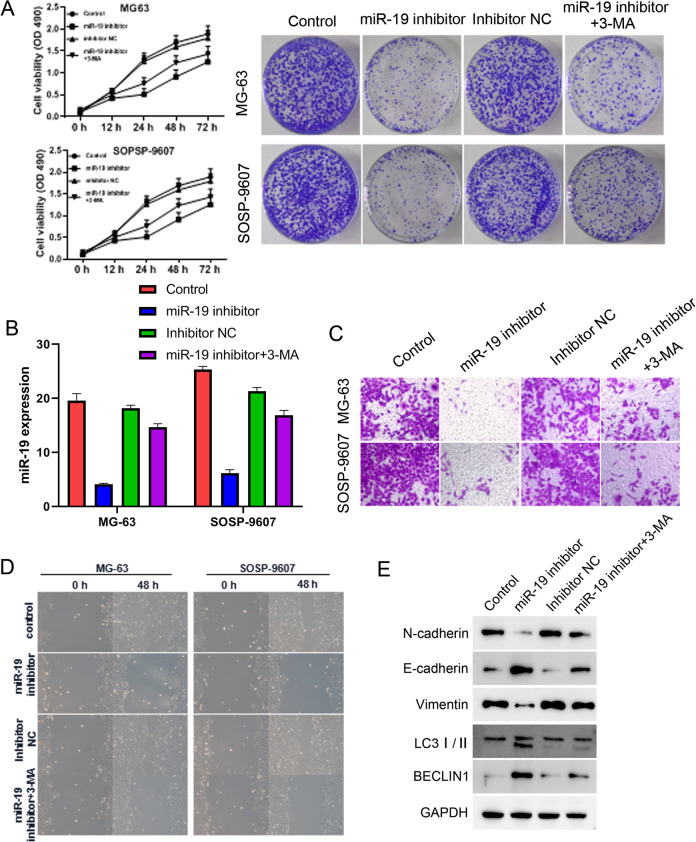
To examine the function of miR-19 in osteosarcoma and its relationship with autophagy, miR-19 inhibitor was used to treat MG-63 and SOSP-9607 cells with or without 3-MA, the inhibitor of autophagy25. Cell proliferation and colony formation were decreased after miR-19 inhibitor, whereas the inhibiting effects were partially reversed by 3-MA treatment (Fig. 2A). The miR-19 expression was reduced by miR-19 inhibitor treatment but not 3-MA, suggesting miR-19 was upstream of autophagy (Fig. 2B). In addition, Transwell and wound-healing assays were used to measure cell invasion and migration, respectively. After miR-19 inhibitor treatment, cell invasion and migration were reduced, while the cell treatment of 3-MA rescued the effect of miR-19 inhibitor compared with control (Fig. 2C, D). Moreover, the expressions of EMT and autophagy markers were assessed by western blot. miR-19 inhibitor downregulated N-cadherin and vimentin of mesenchymal-like marker, and upregulated E-cadherin of epithelial-like marker, LCII and BECLIN1 of autophagy markers, suggesting that it inhibits EMT and enhances the autophagy. Conversely, 3-MA weakened the suppression of EMT and the promotion of autophagy by miR-19 inhibitor (Fig. 2E). Taken together, these results suggest that miR-19 promotes cell proliferation, invasion, migration, and EMT by inhibiting cell autophagy in osteosarcoma.
Figure 2.
miR-19 inhibitor suppresses cell proliferation, invasion, migration, and epithelial–mesenchymal transition by enhancing autophagy in osteosarcoma cells.
GAPDH: glyceraldehyde 3-phosphate dehydrogenase.
3. miR-19 inhibitor reduces cell proliferation, invasion, migration, and EMT through upregulating SPRED2 expression in osteosarcoma cells
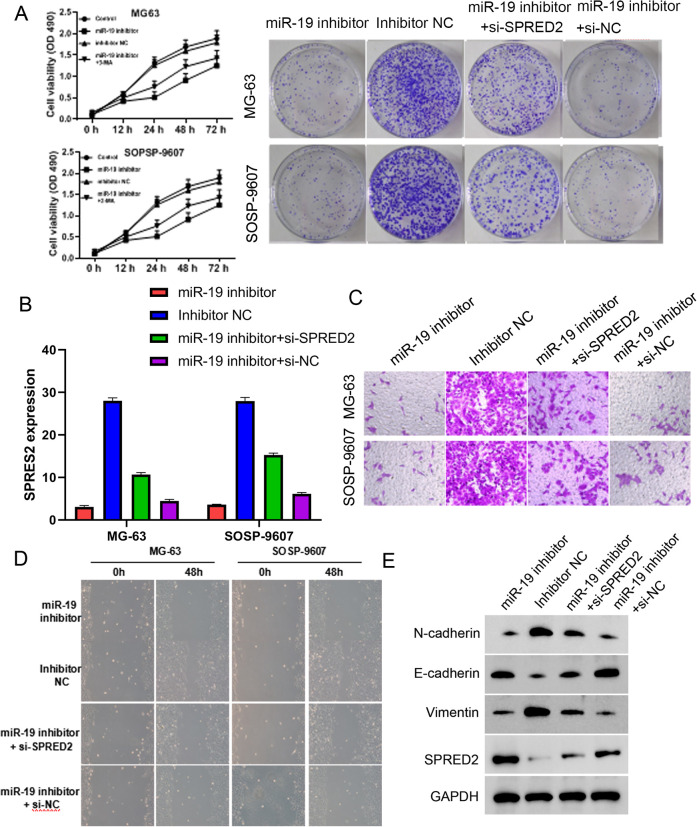
Having shown that miR-19 inhibitor enhanced LC3II and BECLIN1 protein level, we wanted to investigate whether the effects of miR-19 were regulated by SPRED2, an autophagy activator. First, miR-19 inhibitor was used to treat MG-63 and SOSP-9607 cells with or without the SPRED2 siRNA. After SPRES2 siRNA treatment, reduced cell proliferation and colony formation induced by miR-19 inhibitor were partially reversed (Fig. 3A). The SPRES2 expression was significantly upregulated by miR-19 inhibitor at mRNA and protein level, and SPRES2 siRNA downregulated the SPRES2 expression of increased by miR-19 inhibitor, suggesting SPRES2 was the potential target of miR-19 (Fig. 3B). Knockdown of SPRES2 partially rescued the decreased cell invasion and migration mediated by miR-19 inhibitor (Fig. 3C, D). In addition, the downregulation of N-cadherin and vimentin, and upregulation of E-cadherin and LC3II and BECLIN1 by miR-19 inhibitor were weakened by SPRES2 siRNA (Fig. 3E). Knockdown of SPRES2 partiality reverses the effect of miR-19 inhibitor, indicating that miR-19 promotes cell proliferation, invasion, migration, and EMT via targeting SPRES2 to inhibit cell autophagy in osteosarcoma.
Figure 3.
Effect of miR-19 inhibitor on cell proliferation, invasion, migration, epithelial–mesenchymal transition, and autophagy is reversed by knockdown SPRED2 in osteosarcoma cells.
4. SPERED2 is a target of miR-19
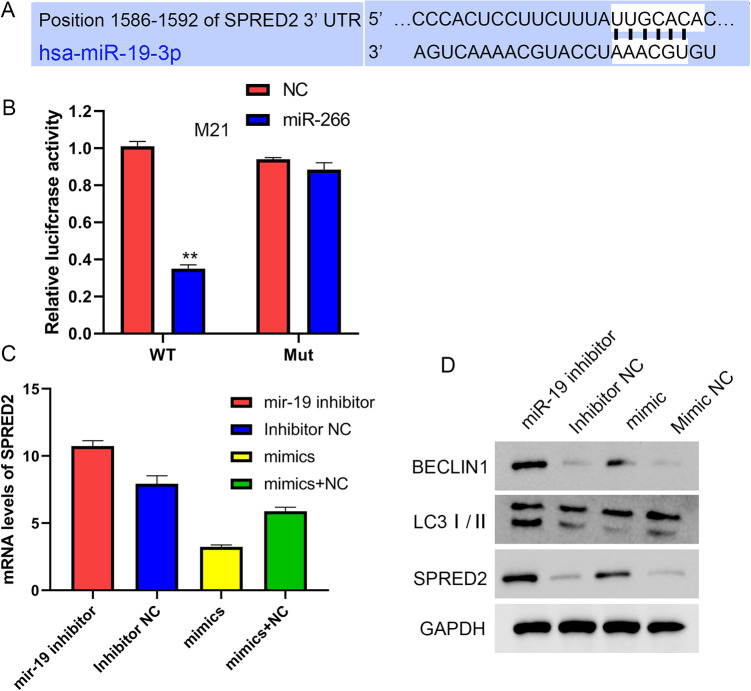
SPRED2 was predicted as a potential target for miR-19 via the TargetScan databases (Fig. 4A). To confirm whether SPRED2 is regulated by miR-19, SPRED2 wild-type 3′-UTR and mutations in the assumed miR-19 binding site of the SPRED2 3′-UTR were constructed into luciferase reporter vector, respectively, and the plasmids were transfected together with miR-19 mimics into the MG-63 cells. The luciferase activity assay showed that miR-19 mimics reduced the activity of the wild-type but not the mutant SPRED2 3′-UTR, indicating that miR-19 binds with the SPRED2 3′-UTR and reduces its promoter activity (Fig. 4A). To validate the ability of miR-19 to suppress the expression of SPRED2, miR-19 mimics and inhibitor treated the MG-63 cells. The results showed that miR-19 mimics significantly decreased the mRNA and protein levels of SPRED2, which were increased by miR-19 inhibitor along with elevated LC3II and BECLIN1 of autophagy marker expression (Fig. 4C, D). These results indicated that SPRED2/autophagy axis played an important role in the effects of miR-19 on osteosarcoma.
Figure 4.
SPRED2 is a target of miR-19.
GAPDH: glyceraldehyde 3-phosphate dehydrogenase.
5. SPRED2 impairs cell proliferation, invasion, migration, and EMT via triggering autophagy in osteosarcoma cells
To explore the effect of SPRED2 in osteosarcoma and the relationship with autophagy, SPRED2 stable overexpression in MG-63 and SOSP-9607 cell lines was established and also the treatment with or without the autophagy inhibitor 3-MA. Stable overexpression SPRED2 significantly reduced the cell proliferation and colony formation in MG-63 and SOSP-9607 cells, while the suppression was partially reversed after 3-MA treatment (Fig. 5A). SPRED2 mRNA and protein were significantly increased in SPRED2 overexpression cells, and 3-MA treatment did not affect SPRED2 expression (Fig. 5B). SPRED2 overexpression significantly decreased the number of invasion cells and migration distance, while 3-MA treatment significantly alleviated the suppression of overexpression SPRED2 on cell invasion and migration (Fig. 5C, D).
Figure 5.
Effect of SPRES2 on cell proliferation, invasion, migration, EMT, and autophagy is reversed by 3-MA treatment in osteosarcoma cells.
GAPDH: glyceraldehyde 3-phosphate dehydrogenase.
In addition, SPRED2 overexpression downregulated N-cadherin and vimentin, and upregulated E-cadherin, LC3II, and BECLIN1, suggesting that SPRED2 can reduce EMT and trigger autophagy. On the contrary, 3-MA weakened the suppression of EMT and the promotion of autophagy by SPRED2 (Fig. 5E). Collectively, the results suggest that SPRED2 decreases cell proliferation, invasion, migration, and EMT via activating cell autophagy.
Discussion
miR-19 is a member and the important oncogenic miRNA of miR-17-92 cluster, which is considered to be the first miRNA cluster with oncogenic potential26–28. So far, miR-19 has been proved to reduce apoptosis or chemosensitivity, and promote proliferation or migration in multiple cancers29,30. miR-19 enhances tumorigenesis by regulating target genes and related signaling pathways. In a previous study, high levels of miR-19 promoted cell proliferation or migration by triggering the extracellular signal–regulated kinase signaling pathway in osteosarcoma cells13. Multiple proteins including CtIP31, RhoB32, and p38α kinase33 could act as the target and negativity regulated by miR-19, indicating that in-depth mechanism research could help provide a novel multigene therapy for cancer treatment34.
In the study, we confirmed that miR-19 expression was high in osteosarcoma. Furthermore, miR-19 inhibitor could decrease proliferation and increase autophagy in osteosarcoma cells. These results also proved that miR-19 was a proto-oncogene miRNA related with cell autophagy. Autophagy plays an important role in maintaining cell homeostasis, while cancer cells have the ability to destroy intracellular homeostasis through disrupting the balance between cell apoptosis and proliferation and promote malignant growth15,16. However, autophagy has a dual function in development and progression of cancer and is a vital target in human disease prevention and therapy35. A previous study reported that autophagy inhibition diminishes the apoptosis of human osteosarcoma cells. In this study, 3-MA of autophagy inhibitor could rescue the biological effect of miR-19 inhibitor on MG-63 and SOSP-9607 cells by decreasing proliferation, invasion, migration, and impairing EMT, indicating that miR-19 promotes the malignant phenotypes of osteosarcoma cells via inhibiting autophagy.
SPRED2 is an important negative regulator of the ERK/MAPK pathway to trigger autophagy, and its expression is low in many cancers, resulting in overactivation of ERK/MAPK signaling pathway21,22. Many recent reports have demonstrated that overexpression SPRED2 increases the conversion of LC3 by interaction with the LC3, thereby triggering autophagy in HeLa and A549 cells36. Taken together, these observations indicate that SPRED2 acts as an activator of autophagy to induce tumor cell death. In this study, we confirmed SPRED2 was downregulated in osteosarcoma, suggesting that SPRED2 and miR-19 have opposite important functions in development of osteosarcoma. To further verify whether miR-19 promotes malignant transformation in osteosarcoma cells by regulating SPRED2, we transfected both miR-19 inhibitor and SPRED2 siRNA into osteosarcoma cells. The results showed that knockdown SPRED2 could partially rescue the biological effect of miR-19 inhibitor. Moreover, miR-19 inhibitor increased SPRED2 expression through mRNA upregulation, suggesting SPRED2 is a target gene of miR-19.
Luciferase reporter gene system was used to confirm whether SPRED2 is a target gene of miR-19. The results showed that miR-19 interacted with SPRED2 3′-UTR to regulate the expression of SPRED2. miR-19 mimics reduced the expression of SPRED2, LC3II, and BECLIN1, suggesting that SPRED2 involves in the regulation of autophagy in osteosarcoma like in other kinds of cancer. SPRED2 overexpression or cotreatment with 3-MA experiments indicates that upregulation of SPRED2 decreases malignant phenotypes of osteosarcoma by promoting autophagy.
Conclusion
Taken together, increased miR-19 and reduced SPRED2 expression are found in human osteosarcoma. miR-19 can promote malignant transformation of osteosarcoma cells via downregulating SPRED2 to inhibit autophagy. Therefore, miR-19/SPRED2 axis may offer a potential target for treating osteosarcoma.
Acknowledgments
We would like to acknowledge everyone for their helpful contributions to this paper.
Footnotes
Authors’ Contributions: Each author has made an important scientific contribution to the study and has assisted with the drafting or revising of the manuscript.
Availability of Data and Materials: The data are free to access and available upon request.
Consent to Publish: All of the authors have consented to publish this research.
Ethical Approval: The ethical approval was obtained from the Ethic Committee of The Second Affiliated Hospital of Guangzhou Medical University.
Statement of Human and Animal Rights: All experiments were performed in accordance with government policy and the Helsinki declaration.
Statement of Informed Consent: Written informed consent was obtained from all patients.
Declaration of Conflicting Interests: The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding: The author(s) received no financial support for the research, authorship, and/or publication of this article.
ORCID iD: Chuhai Xie  https://orcid.org/0000-0001-5991-2555
https://orcid.org/0000-0001-5991-2555
References
- 1. Isakoff MS, Bielack SS, Meltzer P, Gorlick R. Osteosarcoma: current treatment and a collaborative pathway to success. J Clin Oncol. 2015;33(27):3029–3035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Nikitovic D, Kavasi RM, Berdiaki A, Papachristou DJ, Tsiaoussis J, Spandidos DA, Tsatsakis AM, Tzanakakis GN. Parathyroid hormone/parathyroid hormone-related peptide regulate osteosarcoma cell functions: Focus on the extracellular matrix (Review). Oncol Rep. 2016;36(4):1787–1792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Otoukesh B, Boddouhi B, Moghtadaei M, Kaghazian P, Kaghazian M. Novel molecular insights and new therapeutic strategies in osteosarcoma. Cancer Cell Int. 2018;18:158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. O’Brien J, Hayder H, Zayed Y, Peng C. Overview of MicroRNA biogenesis, mechanisms of actions, and circulation. Front Endocrinol (Lausanne). 2018;9:402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Condrat CE, Thompson DC, Barbu MG, Bugnar OL, Boboc A, Cretoiu D, Suciu N, Cretoiu SM, Voinea SC. miRNAs as biomarkers in disease: latest findings regarding their role in diagnosis and prognosis. Cell. 2020;9(2):276. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Cui M, Wang H, Yao X, Zhang D, Xie Y, Cui R, Zhang X. Circulating microRNAs in cancer: potential and challenge. Front Genet. 2019;10:626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Wang H, Peng R, Wang J, Qin Z, Xue L. Circulating microRNAs as potential cancer biomarkers: the advantage and disadvantage. Clin Epigenetics. 2018;10:59. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Jacques C, Tesfaye R, Lavaud M, Georges S, Baud’huin M, Lamoureux F, Ory B. Implication of the p53-Related miR-34c, -125b, and -203 in the osteoblastic differentiation and the malignant transformation of bone sarcomas. Cells. 2020;9(4):810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Leichter AL, Sullivan MJ, Eccles MR, Chatterjee A. MicroRNA expression patterns and signalling pathways in the development and progression of childhood solid tumours. Mol Cancer. 2017;16(1):15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Jing ZF, Bi JB, Li ZL, Liu XK, Li J, Zhu YY, Zhang XT, Zhang Z, Li ZH, Kong CZ. miR-19 promotes the proliferation of clear cell renal cell carcinoma by targeting the FRK-PTEN axis. Onco Targets Ther. 2019;12:2713–2727. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Liu Y, Liu R, Yang F, Cheng R, Chen X, Cui S, Gu Y, Sun W, You C, Liu Z, Sun F, et al. miR-19a promotes colorectal cancer proliferation and migration by targeting TIA1. Mol Cancer. 2017;16(1):53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Niu SX, Ma X, Zhang Y, Liu YN, Chen XF, Gong HJ, Yao YX, Liu K, Zhang X. MicroRNA-19a and microRNA-19b promote the malignancy of clear cell renal cell carcinoma through targeting the tumor suppressor RhoB. Plos One. 2018;13(2):e0192790. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 13. Meng Q, Dai M, Nie X, Zhang W, Xu X, Li J, Mu H, Liu X, Qin L, Zhu X, Yan J, et al. MicroRNA-19 contributes to the malignant phenotypes of osteosarcoma in vitro by targeting Pax6. Tumour Biol. 2018;40(1):1010428317744704. [DOI] [PubMed] [Google Scholar]
- 14. Su Z, Yang Z, Xu Y, Chen Y, Yu Q. Apoptosis, autophagy, necroptosis, and cancer metastasis. Mol Cancer. 2015;14:48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. He C, Bassik MC, Moresi V, Sun K, Wei Y, Zou Z, An Z, Loh J, Fisher J, Sun Q, Korsmeyer S, et al. Exercise-induced BCL2-regulated autophagy is required for muscle glucose homeostasis. Nature. 2012;481(7382):511–515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Carroll B, Korolchuk VI, Sarkar S. Amino acids and autophagy: cross-talk and co-operation to control cellular homeostasis. Amino Acids. 2015;47(10):2065–2088. [DOI] [PubMed] [Google Scholar]
- 17. Yun CW, Lee SH. The roles of autophagy in cancer. Int J Mol Sci. 2018;19(11):3466. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Yoshii SR, Mizushima N. Monitoring and measuring autophagy. Int J Mol Sci. 2017;18(9):1865. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Kim YC, Guan KL. mTOR: a pharmacologic target for autophagy regulation. J Clin Invest. 2015;125(1):25–32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Alers S, Loffler AS, Wesselborg S, Stork B. Role of AMPK-mTOR-Ulk1/2 in the regulation of autophagy: cross talk, shortcuts, and feedbacks. Mol Cell Biol. 2012;32(1):2–11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Mason JM, Morrison DJ, Basson MA, Licht JD. Sprouty proteins: multifaceted negative-feedback regulators of receptor tyrosine kinase signaling. Trends Cell Biol. 2006;16(1):45–54. [DOI] [PubMed] [Google Scholar]
- 22. Masoumi-Moghaddam S, Amini A, Morris DL. The developing story of Sprouty and cancer. Cancer Metastasis Rev. 2014;33(2–3):695–720. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Kawazoe T, Taniguchi K. The Sprouty/Spred family as tumor suppressors: coming of age. Cancer Sci. 2019;110(5):1525–1535. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Rathmanner N, Haigl B, Vanas V, Doriguzzi A, Gsur A, Sutterluty-Fall H. Sprouty2 but not Sprouty4 is a potent inhibitor of cell proliferation and migration of osteosarcoma cells. FEBS Lett. 2013;587(16):2597–2605. [DOI] [PubMed] [Google Scholar]
- 25. Hendil KB, Lauridsen AM, Seglen PO. Both endocytic and endogenous protein degradation in fibroblasts is stimulated by serum/amino acid deprivation and inhibited by 3-methyladenine. Biochem J. 1990;272(3):577–581. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Wang W, Zhang A, Hao Y, Wang G, Jia Z. The emerging role of miR-19 in glioma. J Cell Mol Med. 2018;22(10):4611–4616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Olive V, Bennett MJ, Walker JC, Ma C, Jiang I, Cordon-Cardo C, Li QJ, Lowe SW, Hannon GJ, He L. miR-19 is a key oncogenic component of mir-17-92. Genes Dev. 2009;23(24):2839–2849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Donayo AO, Johnson RM, Tseng HW, Izreig S, Gariepy A, Mayya VK, Wu E, Alam R, Lussier C, Jones RG, Duchaine TF. Oncogenic biogenesis of pri-miR-17 approximately 92 reveals hierarchy and competition among polycistronic microRNAs. Mol Cell. 2019;75(2):340–356.e310. [DOI] [PubMed] [Google Scholar]
- 29. Murphy BL, Obad S, Bihannic L, Ayrault O, Zindy F, Kauppinen S, Roussel MF. Silencing of the miR-17∼92 cluster family inhibits medulloblastoma progression. Cancer Res. 2013;73(23):7068–7078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Han YC, Vidigal JA, Mu P, Yao E, Singh I, Gonzalez AJ, Concepcion CP, Bonetti C, Ogrodowski P, Carver B, Selleri L, et al. An allelic series of miR-17 approximately 92-mutant mice uncovers functional specialization and cooperation among members of a microRNA polycistron. Nat Genet. 2015;47(7):766–775. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Huhn D, Kousholt AN, Sorensen CS, Sartori AA. miR-19, a component of the oncogenic miR-17 approximately 92 cluster, targets the DNA-end resection factor CtIP. Oncogene. 2015;34(30):3977–3984. [DOI] [PubMed] [Google Scholar]
- 32. Glorian V, Maillot G, Poles S, Iacovoni JS, Favre G, Vagner S. HuR-dependent loading of miRNA RISC to the mRNA encoding the Ras-related small GTPase RhoB controls its translation during UV-induced apoptosis. Cell Death Differ. 2011;18(11):1692–1701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Guinot A, Oeztuerk-Winder F, Ventura JJ. miR-17-92/p38alpha dysregulation enhances Wnt signaling and selects Lgr6+ cancer stem-like cells during lung adenocarcinoma progression. Cancer Res. 2016;76(13):4012–4022. [DOI] [PubMed] [Google Scholar]
- 34. Trang P, Wiggins JF, Daige CL, Cho C, Omotola M, Brown D, Weidhaas JB, Bader AG, Slack FJ. Systemic delivery of tumor suppressor microRNA mimics using a neutral lipid emulsion inhibits lung tumors in mice. Mol Ther. 2011;19(6):1116–1122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Loos B, Klionsky DJ, Du Toit A, Hofmeyr JS. On the relevance of precision autophagy flux control in vivo - Points of departure for clinical translation. Autophagy. 2020;16(4):750–762. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Jiang K, Liu M, Lin G, Mao B, Cheng W, Liu H, Gal J, Zhu H, Yuan Z, Deng W, Liu Q, et al. Tumor suppressor Spred2 interaction with LC3 promotes autophagosome maturation and induces autophagy-dependent cell death. Oncotarget. 2016;7(18):25652–25667. [DOI] [PMC free article] [PubMed] [Google Scholar]