Keywords: collecting duct, genome editing, kidney, phosphoproteomics, phosphorylation, protein kinases
Abstract
Vasopressin regulates osmotic water transport in the renal collecting duct by protein kinase A (PKA)-mediated control of the water channel aquaporin-2 (AQP2). Collecting duct principal cells express two seemingly redundant PKA catalytic subunits, PKA catalytic α (PKA-Cα) and PKA catalytic β (PKA-Cβ). To identify the roles of these two protein kinases, we carried out deep phosphoproteomic analysis in cultured mpkCCD cells in which either PKA-Cα or PKA-Cβ was deleted using CRISPR-Cas9-based genome editing. Controls were cells carried through the genome editing procedure but without deletion of PKA. TMT mass tagging was used for protein mass spectrometric quantification. Of the 4,635 phosphopeptides that were quantified, 67 phosphopeptides were significantly altered in abundance with PKA-Cα deletion, whereas 21 phosphopeptides were significantly altered in abundance with PKA-Cβ deletion. However, only four sites were changed in both. The target proteins identified in PKA-Cα-null cells were largely associated with cell membranes and membrane vesicles, whereas target proteins in PKA-Cβ-null cells were largely associated with the actin cytoskeleton and cell junctions. In contrast, in vitro incubation of mpkCCD proteins with recombinant PKA-Cα and PKA-Cβ resulted in virtually identical phosphorylation changes. In addition, analysis of total protein abundances in in vivo samples showed that PKA-Cα deletion resulted in a near disappearance of AQP2 protein, whereas PKA-Cβ deletion did not decrease AQP2 abundance. We conclude that PKA-Cα and PKA-Cβ serve substantially different regulatory functions in renal collecting duct cells and that differences in phosphorylation targets may be due to differences in protein interactions, e.g., mediated by A-kinase anchor proteins, C-kinase anchoring proteins, or PDZ binding.
INTRODUCTION
The actions of vasopressin in collecting duct principal cells (24) are mediated by a G-protein-coupled receptor, vasopressin receptor 2 (gene symbol: Avpr2), which couples to the heterotrimeric G protein stimulatory α-subunit (Gsα) with activation of adenylyl cyclase 6 (23). Increased intracellular cAMP levels result in physiological effects in collecting duct principal cells, largely through activation of protein kinase A (PKA) (5, 6, 9, 12, 13, 18, 20–22). Two of the most important physiological end effects are 1) membrane trafficking changes that increase the abundance of the water channel protein aquaporin-2 (AQP2) in the plasma membrane (19) and 2) increased transcription of the Aqp2 gene (8, 15, 25), both of which contribute to vasopressin-induced increases in osmotic water transport across the collecting duct epithelium. PKA is a widely studied protein that has been viewed by most investigators as a single entity, although its catalytic subunits are coded in mammalian genomes by two separate genes, PKA catalytic α (PKA-Cα; gene symbol: Prkaca) and PKA catalytic β (PKA-Cβ; gene symbol: Prkacb). At the amino acid level, the two are 91% identical and the catalytic domains are very similar. In native collecting duct cells and mpkCCD cells, PKA-Cα and PKA-Cβ are expressed at comparable levels (1, 10). (A third entity, PKA catalytic γ, is not widely expressed and will not be considered in this paper.) We have recently succeeded in using CRISPR-Cas9 to create disruptive mutations in both PKA catalytic genes [PKA double knockout (PKA dKO)] in vasopressin-responsive kidney epithelial cells (mpkCCD cells) (10). We then used phosphoproteomics to identify a large number of novel PKA targets as well as many secondary changes in phosphorylation due to loss of PKA-mediated regulation of other kinases and phosphatases (10). Whether the two PKA catalytic proteins have redundant regulatory functions, as is implicitly assumed in many studies involving PKA-mediated regulation, has not been tested. Here, we carried out mass spectrometry-based quantitative proteomics and phosphoproteomics in both PKA-Cα and PKA-Cβ single knockouts (KOs) in mpkCCD collecting duct cells to address this issue. Thus, the goal of the present study was to identify the relative roles of PKA-Cα and PKA-Cβ in the signaling processes triggered by vasopressin.
METHODS
Cell culture.
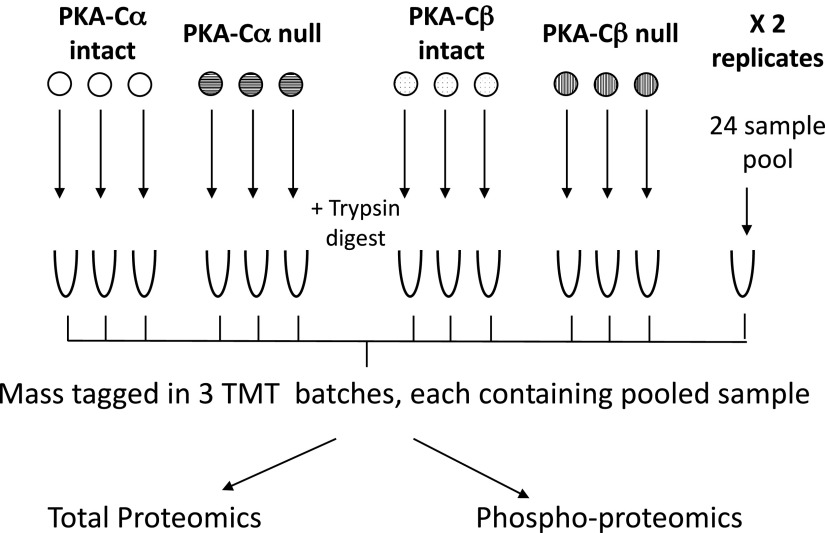
The present study used immortalized mpkCCD cells in which either Prkaca or Prkacb gene expression was deleted (“PKA-Cα-null” and “PKA-Cβ-null”) by introducing mutations using CRISPR-Cas9 (10). Control cell lines (“PKA intact”) were carried through the CRISPR-Cas9 protocol but did not show deletion of either PKA catalytic gene. We used three independent PKA-Cα-null lines and three independent PKA-intact controls. Similarly, we used three independent PKA-Cβ-null lines and three different independent PKA-intact controls, giving three biological replicates for each kinase. Experiments with these lines were performed in duplicate (i.e., 2 technical replicates each), giving 24 samples in total. The overall scheme is shown in Fig. 1. Cells were cultured as previously described (14). Briefly, cells were initially maintained in complete medium, DMEM-F-12 containing 2% serum and other supplements (5 μg/mL insulin, 50 nM dexamethasone, 1 nM triiodothyronine, 10 ng/mL epidermal growth factor, 60 nM sodium selenite, and 5 μg/mL transferrin, all from Sigma). Cells were seeded on permeable membrane supports (Transwell) and grown in complete medium containing 0.1 nM 1-desamino-8-d-arginine-vasopressin (dDAVP; basal side only) for 4 days, during which time cells formed confluent monolayers. The medium was then changed to simple medium (DMEM-F-12 with dexamethasone, sodium selenite, and transferrin and no serum) with 0.1 nM dDAVP and maintained for 3 days, at which time cells were harvested for proteomic analysis. Because dDAVP was present in the culture medium under all conditions, it was not an experimental variable in the present study.
Fig. 1.
Experimental method. Cells were grown on permeable supports with three biological replicates as indicated. There were two technical replicates each, resulting in a total of 24 samples. Proteins were isolated, trypsinized, and labeled using isotopic tags (TMT-11 plex). The 24 samples were combined into 3 batches of 8 each with a shared sample pooled from all 24 samples. The combined samples were processed for proteomic and phosphoproteomic analysis as described in methods.
Sample preparation for total and phospho-proteomics.
Cells were washed three times with ice-cold PBS and lysed with TEAB buffer (ThermoFisher) with SDS (1%) containing protease and phosphatase inhibitors (Halt, ThermoFisher). Membranes were scraped, and samples were homogenized using a QIAshredder (Qiagen). Protein concentrations were measured using the Pierce BCA Protein Assay Kit. Protein lysates were reduced with 20 mM DTT for 1 h at 25°C and then alkylated with 40 mM iodoacetamide for 1 h at 25°C in the dark. Proteins were acetone precipitated before digestion with recombinant trypsin [1:50 (wt/wt), ThermoFisher] overnight at 37°C. The resulting peptides were quantified using the Pierce Quantitative Colorimetric Peptide Assay. For each replicate, 250 μg of peptide were labeled using the TMT11Plex Mass Tag Labeling Kit (lot no. UE283355, Thermo Scientific) following the manufacturer’s instructions, and they were combined as shown in Fig. 1. A total of 3 labeling batches using the same TMT11 Plex Mass Tag kit were run to quantify all 24 samples. Each batch included a common pooled sample containing a mixture of all 24 experimental samples. Samples were combined and desalted using hydrophilic-lipophilic-balanced extraction cartridges (Oasis) and then fractionated into 12 fractions using high pH reverse-phase chromatography (Agilent 1200 HPLC System). After an aliquot was taken from each fraction for total proteomics, phosphopeptides were sequentially enriched following the Sequential Enrichment from Metal Oxide Affinity Chromatography protocol (Thermo Scientific) using High-Select TiO2 and then High-Select Fe-NTA Phosphopeptide enrichment kits (Thermo Scientific). Samples were then vacuum dried and stored at −80°C until analysis.
Dried peptides were resuspended with 0.1% formic acid and 2% acetonitrile in LC-MS grade water (J.T. Baker) before mass spectrometry analysis. Peptides (total and phospho-) were analyzed using a Dionex UltiMate 3000 Nano LC system connected to an Orbitrap Fusion Lumos mass spectrometer equipped with an EASY-Spray ion source (ThermoFisher Scientific). Peptides were introduced into a peptide nanotrap at a flow rate of 5 μL/min. The trapped peptides were fractionated with a reverse-phase EASY-Spray PepMap column (C18, 75 μm × 50 cm) using a linear gradient of 4−32% acetonitrile in 0.1% formic acid (120 min at 0.3 μL/min). The Thermo Scientific Synchronous Precursor Selection-MS3 (SPS-MS3) workflow (17) was selected on the mass spectrometer for TMT quantification. The main settings for MS3 were as follows: HCD activation, 65% normalized collision energy, 2 m/z isolated window, 50,000 Orbitrap resolution, AGC target of 15,000, and 120-ms maximum injection time.
Mass spectrometry data processing.
Raw mass spectra were searched against the Mus musculus UniProtKB (27) reference proteome (proteome ID: UP000000589, release 2019_06, plus contaminant database) using MaxQuant (3) 1.6.7.0. Reporter ion MS3 with TMT10plex was specified as the labeling type, and lot-specific TMT isotopic impurity correction factors were used as recommended in the TMT product data sheets. Carbamidomethyl (C) was configured as fixed modifications. Variable modifications included phospho (STY), oxidation (M), and acetyl (Protein-N-terminal). The false discovery rate was controlled at 1% (target-decoy). “Trypsin/P” was set as the digestion enzyme with up to two missed cleavages allowed. Other parameters were set to the defaults. We used the “proteinGroups.txt” output file as the input data for total proteome analyses. Both “Phospho (STY)Sites.txt” and “evidence.txt” output files were used for phosphoproteome analyses. For each TMT batch, corrected reporter ion intensities were first normalized to make the total sum intensities for each channel equal. The average of the common pooled channel from three TMT batches was then used to normalize the batch effects.
In vitro phosphorylation experiments.
Three independent PKA dKO cell lines (10) were grown on permeable membrane supports as described above. Confluent monolayers were washed three times with ice-cold PBS and lysed with TEAB buffer (ThermoFisher) with SDS (1%) containing protease and phosphatase inhibitors. Membranes were scraped, samples were homogenized using a QIAshredder (Qiagen), and proteins were precipitated with acetone. The protein pellet was resuspended in TEAB buffer, and protein concentration was determined by the Pierce BCA Protein Assay Kit.
Equal quantities of proteins from the three PKA dKO cell lines were pooled together, and 500 μg pooled protein extract was mixed with either recombinant PKA-Cα (1.5 μg) or recombinant PKA-Cβ (1.5 μg) enzymes obtained from Genetex (PRKACA- GTX65206-pro and PRKACB-GTX65207-pro). Samples were incubated at 30°C for 24 h in buffer containing 50 mM Tris·HCl, 10 mM MgCl2, 0.1 mM EDTA, 2 mM DTT, and 250 μM ATP. Samples without any added kinases were used as controls. Following the in vitro kinase incubation, proteins underwent standard phosphoproteomic analysis as previously described.
Bioinformatics and data sharing.
Technical replicates were summarized by taking the median value. All analyses were performed using Perseus, Excel, and R software. Moderated P (Pmod) values were calculated using the empirical Bayes method, which integrates variance information from all peptides measured in the same LC-MS/MS run (11). Data have been deposited in PRIDE (as part of ProteomeXchange) with Accession No. PXD015050.
RESULTS
Previously, PKA-Cα-null and PKA-Cβ-null cells were characterized by immunoblot analysis, which showed that PKA-Cα and PKA-Cβ proteins were undetectable in the corresponding single KO lines (10). Both single KO lines grew well and formed confluent monolayered epithelial sheets when grown on permeable supports. Here, we used protein mass spectrometry to quantify changes in total protein abundances and changes in phosphorylation in PKA-Cα-null versus PKA-Cα-intact cells and PKA-Cβ-null versus PKA-Cβ-intact cells across the proteome.
Proteome-wide quantification of protein abundances.
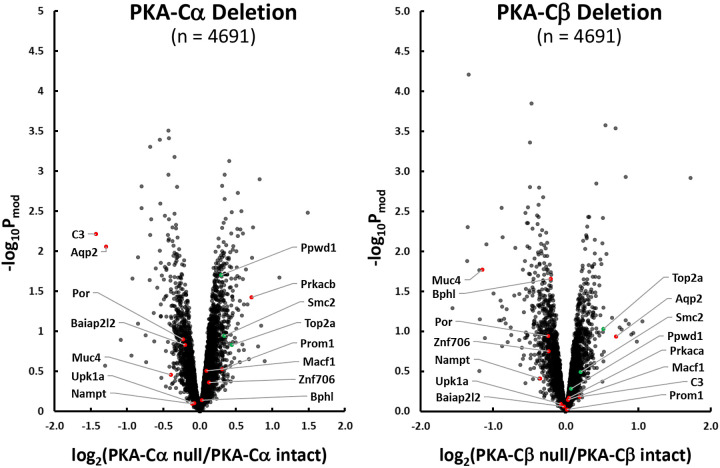
Quantitative data for the total abundance of all individual proteins are online at https://hpcwebapps.cit.nih.gov/ESBL/Database/PKA-singleKO-total/ and as Supplemental Data Set S1 in the Supplemental Material (provided online at https://hpcwebapps.cit.nih.gov/ESBL/Database/Supplemental_Data_PKA_sKO/). Figure 2 shows changes in protein abundances corresponding to 4,691 different genes in both PKA-Cα-null cells and PKA-Cβ-null cells versus their respective PKA-intact control cells. Each point shows values for a different protein. Proteins whose abundances were previously found to be significantly altered in PKA-Cα/PKA-Cβ dKO cells are marked by red or green points depending on the direction of change in dKO. Interestingly, Aqp2, which had previously been seen to be almost completely ablated in PKA dKO cells, was decreased in only PKA-Cα-null cells and not in PKA-Cβ-null cells. Two other proteins that were strongly decreased in abundance in PKA dKO experiments, namely, complement factor C3 (C3) and mucin-4 (Muc4) were differentially affected in the two single KO cells. C3 was selectively decreased in PKA-Cα-null cells, whereas mucin-4 was selectively decreased in PKA-Cβ-null cells. These data support the view that PKA-Cα and PKA-Cβ do not have the same physiological roles in collecting duct cells. Interestingly, PKA-Cβ was substantially increased in abundance in PKA-Cα-null cells, suggesting a compensatory response.
Fig. 2.
Effect of PKA-Cα (left) and PKA-Cβ (right) deletion on protein abundances in mouse mpkCCD cells. Red points indicate proteins decreased in PKA-Cα/PKA-Cβ double knockout cells (10). Green points indicate proteins increased in PKA-Cα/PKA-Cβ double knockout cells (10). Arrows to these red and green points show official gene symbols for the specific proteins. Pmod value, moderated P value.
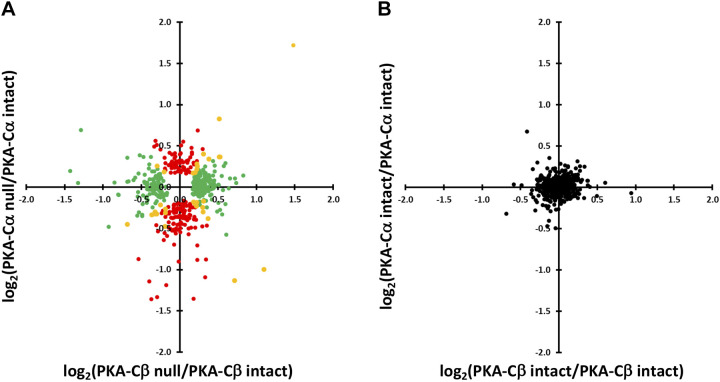
We plotted the changes in protein abundances in PKA-Cα-null cells versus those in PKA-Cβ-null cells for those proteins that changed significantly in either PKA-KO line (Fig. 3A). As can be seen, some proteins (yellow) underwent parallel changes in abundance in both single KO cell lines, whereas others underwent selective, significant changes in either PKA-Cα-null (red) or PKA-Cβ-null (green) cells. Control versus control comparisons for the same proteins are shown in Fig. 3B, illustrating the magnitude of changes expected from random variability in the mass spectrometric quantification. [The 95% confidence interval for log2(control/control) was 0.382.] A listing of the proteins that most convincingly changed [Pmod < 0.02 and |log(ratio)| > 0.5] in PKA-Cα-null cells is shown in Table 1, whereas changes in PKA-Cβ-null cells are shown in Table 2. Overall, we conclude that PKA-Cα and PKA-Cβ are not redundant with respect to regulation of protein abundances.
Fig. 3.
Comparison of effects of PKA-Cα and PKA-Cβ deletion on protein abundances. A: red points show proteins changed with PKA-Cα deletion but not PKA-Cβ deletion. Green points show proteins changed with PKA-Cβ deletion but not PKA-Cα deletion. Yellow points are changed in both, but not necessarily in the same direction. B: intrinsic variability of the data estimated by comparing values for PKA-intact controls versus other controls (see text).
Table 1.
Proteins with substantial changes in abundance in response to PKA-Cα deletion [Pmod < 0.02 and |log(ratio)| > 0.5]
| UniProt ID | Gene Symbol | Annotation | Log2 (PKA-Cα Null/PKA-Cα Intact) | Pmod Value | Log2 (PKA-Cβ Null/ PKA-Cβ Intact) | Pmod Value |
|---|---|---|---|---|---|---|
| P01027 | C3 | Complement C3 | −1.427 | 0.006 | 0.195 | 0.655 |
| P56402 | Aqp2 | Aquaporin-2 | −1.286 | 0.009 | 0.694 | 0.115 |
| Q9QYI5 | Dnajb2 | DnaJ homolog subfamily B member 2 | −0.850 | 0.012 | 0.056 | 0.848 |
| P97742 | Cpt1a | Carnitine O-palmitoyltransferase 1, liver isoform | −0.799 | 0.002 | −0.091 | 0.647 |
| Q8BH86 | Dglucy | d-glutamate cyclase, mitochondrial | −0.798 | 0.003 | −0.094 | 0.666 |
| P24472 | Gsta4 | Glutathione S-transferase A4 | −0.684 | 0.006 | −0.451 | 0.049 |
| Q9EPK8 | Trpv4 | Transient receptor potential channel subfamily V member 4 | −0.682 | 4.93 × 10−4 | 0.067 | 0.647 |
| Q9WTQ5 | Akap12 | A-kinase anchor protein 12 | −0.669 | 0.004 | 0.070 | 0.714 |
| Q99J39 | Mlycd | Malonyl-CoA decarboxylase, mitochondrial | −0.602 | 0.016 | 0.087 | 0.690 |
| Q8BLF1 | Nceh1 | Neutral cholesterol ester hydrolase 1 | −0.551 | 0.010 | −0.346 | 0.077 |
| P63011 | Rab3a | Ras-related protein Rab-3A | −0.550 | 4.05 × 10−4 | −0.115 | 0.324 |
| P26443 | Glud1 | Glutamate dehydrogenase 1, mitochondrial | −0.543 | 0.003 | −0.057 | 0.692 |
| G3X9Y6 | Akr1c19 | Aldo-keto reductase family 1, member C19 | −0.534 | 0.014 | −0.307 | 0.123 |
| Q8BG05 | Hnrnpa3 | Heterogeneous nuclear ribonucleoprotein A3 | 0.507 | 0.010 | 0.033 | 0.845 |
| D3YYU8 | Obsl1 | Obscurin-like protein 1 | 0.512 | 0.016 | 0.024 | 0.897 |
| P62482 | Kcnab2 | Voltage-gated potassium channel subunit β2 | 0.518 | 0.005 | 0.366 | 0.033 |
| Q8VI84 | Noc3l | Nucleolar complex protein 3 homolog | 0.524 | 0.002 | 0.369 | 0.016 |
| Q5DU09 | Znf652 | Zinc finger protein 652 | 0.527 | 0.006 | −0.275 | 0.108 |
| O88508 | Dnmt3a | DNA (cytosine-5)-methyltransferase 3A | 0.562 | 0.005 | −0.034 | 0.838 |
| O54786 | Dffa | DNA fragmentation factor subunit α | 0.576 | 0.003 | 0.255 | 0.127 |
| Q9D187 | Ciao2b | Cytosolic iron-sulfur assembly component 2B | 0.596 | 0.006 | −0.206 | 0.268 |
| Q61180 | Scnn1a | Amiloride-sensitive sodium channel subunit α | 0.637 | 0.017 | −0.016 | 0.945 |
| P0DOV2 | Ifi204 | Interferon-activable protein 204 | 0.722 | 0.012 | 0.108 | 0.664 |
| Q9JK42 | Pdk2 | Pyruvate dehydrogenase kinase isozyme 2 | 0.734 | 0.005 | 0.049 | 0.821 |
| P26645 | Marcks | Myristoylated alanine-rich C-kinase substrate | 1.486 | 0.003 | 1.718 | 0.001 |
| P05132 | Prkaca | cAMP-dependent protein kinase catalytic subunit α | 0.049 | 0.677 |
Pmod value, moderated P value.
Table 2.
Proteins with substantial changes in abundance in response to PKA-Cβ deletion [Pmod < 0.02 and |log(ratio)| > 0.5]
| UniProt ID | Gene Symbol | Annotation | Log2 (PKA-Cα Null/ PKA-Cα Intact) | Pmod Value | Log2 (PKA-Cβ Null/ PKA-Cβ Intact) | Pmod Value |
|---|---|---|---|---|---|---|
| Q8VCT4 | Ces1d | Carboxylesterase 1D | −0.370 | 0.440 | −1.358 | 0.013 |
| Q8CEZ4 | Mab21l4 | Protein mab-21-like 4 | 0.180 | 0.652 | −1.351 | 0.005 |
| P30115 | Gsta3 | Glutathione S-transferase A3 | −0.294 | 0.202 | −1.334 | 6.23 × 10−5 |
| P08074 | Cbr2 | Carbonyl reductase [NADPH] 2 | −0.178 | 0.683 | −1.186 | 0.017 |
| Q8JZM8 | Muc4 | Mucin-4 | −0.396 | 0.353 | −1.144 | 0.017 |
| P00329 | Adh1 | Alcohol dehydrogenase 1 | 0.334 | 0.348 | −1.094 | 0.008 |
| Q7TPW1 | Nexn | Nexilin | 0.228 | 0.349 | −0.881 | 0.003 |
| Q9D939 | Sult1c2 | Sulfotransferase 1C2 | −0.535 | 0.065 | −0.869 | 0.007 |
| Q62469 | Itga2 | Integrin α2 | −0.074 | 0.743 | −0.696 | 0.009 |
| Q99LB7 | Sardh | Sarcosine dehydrogenase, mitochondrial | −0.163 | 0.457 | −0.591 | 0.017 |
| Q9WUU7 | Ctsz | Cathepsin Z | −0.053 | 0.786 | −0.562 | 0.013 |
| Q8R3G9 | Tspan8 | Tetraspanin-8 | −0.199 | 0.212 | −0.557 | 0.003 |
| P22935 | Crabp2 | Cellular retinoic acid-binding protein 2 | −0.282 | 0.088 | −0.534 | 0.004 |
| P63082 | Atp6v0c | V-type H+-ATPase 16-kDa proteolipid subunit | −0.101 | 0.556 | −0.526 | 0.009 |
| Q80V42 | Cpm | Carboxypeptidase M | −0.077 | 0.607 | −0.516 | 0.004 |
| P12265 | Gusb | β-Glucuronidase | −0.032 | 0.819 | −0.516 | 0.003 |
| P17563 | Selenbp1 | Methanethiol oxidase | −0.122 | 0.525 | −0.506 | 0.019 |
| Q5FWI3 | Cemip2 | Cell surface hyaluronidase | 0.020 | 0.899 | 0.508 | 0.008 |
| P55264 | Adk | Adenosine kinase | −0.297 | 0.059 | 0.513 | 0.004 |
| Q99PT3 | Ino80b | INO80 complex subunit B | 0.048 | 0.658 | 0.551 | 2.66 × 10−4 |
| Q8K0C4 | Cyp51a1 | Lanosterol 14-α demethylase | −0.313 | 0.117 | 0.563 | 0.011 |
| P28661 | Septin4 | Septin-4 | 0.235 | 0.106 | 0.690 | 2.90 × 10−4 |
| Q80Z25 | Ofd1 | Oral-facial-digital syndrome 1 protein homolog | 0.518 | 0.020 | 0.826 | 0.001 |
| P26645 | Marcks | Myristoylated alanine-rich C-kinase substrate | 1.486 | 0.003 | 1.718 | 0.001 |
| P68181 | Prkacb | cAMP-dependent protein kinase catalytic subunit β | 0.717 | 0.038 |
Pmod value, moderated P value.
Proteome-wide quantification of protein phosphorylation.
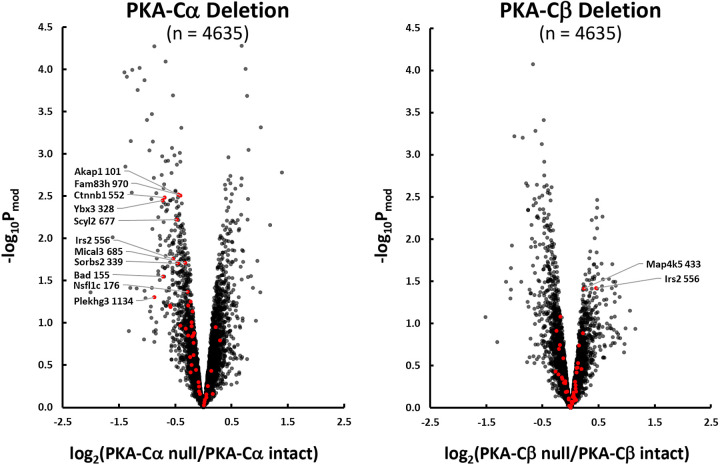
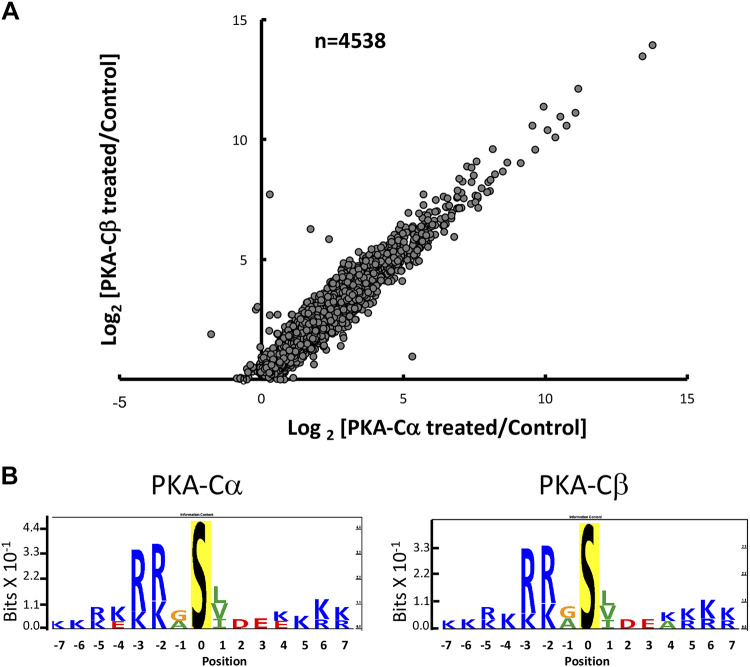
Quantitative data are provided at https://hpcwebapps.cit.nih.gov/ESBL/Database/PKA-singleKO-phospho/ and as Supplemental Data Set S2 for all phosphopeptides. Figure 4 shows changes in phosphopeptide abundances (n = 4635) in both PKA-Cα-null cells and PKA-Cβ-null cells versus their respective PKA-intact (control) cells. Each point shows values for a different phosphopeptide. Phosphopeptides corresponding to previously identified PKA target sites (10) are indicated in red. Most of the PKA sites were found to be decreased in PKA-Cα-null but not PKA-Cβ-null cells relative to controls. A listing of the phosphopeptides most convincingly changed [Pmod < 0.01 and |log(ratio)| > 0.5] in PKA-Cα-null cells is shown in Table 3, whereas changes in PKA-Cβ-null cells are shown in Table 4. Overall, we conclude that PKA-Cα and PKA-Cβ have a substantially different set of phosphorylation targets. This difference could be due either to a difference in target sequence preference or a difference in PKA-Cα and PKA-Cβ interactomes. Figure 5 shows the predicted target sequence preference logos for decreased single-site phosphopeptides in PKA-Cα- and PKA-Cβ-null cells, respectively. Only the PKA-Cα logo was consistent with the familiar (R/K)-(R/K)-X-p(S/T) motif generally accepted to be characteristic of PKA. Among the 82 phosphosites that were decreased in PKA-Cβ-null cells, only four possessed the (R/K)-(R/K)-X-p(S/T) motif. The logos raise the hypothesis that the two PKA catalytic subunits may have different target sequence preferences.
Fig. 4.
Effect of PKA-Cα (left) and PKA-Cβ (right) deletion on phosphopeptide abundances in mouse mpkCCD cells. Red points indicate phosphorylation sites altered in PKA-Cα/PKA-Cβ double knockout cells (10). Arrows to these red points show official gene symbol and amino acid number of phosphorylated site for those sites above the moderated P (Pmod) threshold (−log Pmod > 1.3).
Table 3.
Phosphopeptides most convincingly changed in PKA-Cα-null cells versus PKA-Cα-intact [Pmod < 0.01 and |log(ratio)| > 0.5]
| UniProt ID | Gene Symbol† | Amino Acid Number | Annotation | Sequence | Log2 (PKA-Cα Null/ PKA-Cα Intact) | Pmod Value | Log2 (PKA-Cβ Null/ PKA-Cβ Intact) | Pmod Value |
|---|---|---|---|---|---|---|---|---|
| P16254 | Srp14 | 44 | Signal recognition particle 14-kDa protein | KS*SVEGLEPAENK | −1.41 | 1.09 × 10−4 | 0.18 | 0.472 |
| P31324 | Prkar2b | 112 | cAMP-dependent protein kinase type II-β regulatory subunit | RAS*VCAEAYNPDEEEDDAESR | −1.38 | 0.001 | −0.29 | 0.402 |
| P05213 | Tuba1b; Tuba4a; Tuba1a; Tuba3a; Tuba1c |
158 | Tubulin α1B chain; tubulin α4A chain; tubulin α1A chain; tubulin α1C chain |
LS*VDYGK | −1.36 | 1.21 × 10−4 | 0.28 | 0.266 |
| E9PVZ8 | Golgb1 | 2655 | Golgi autoantigen, golgin subfamily b, macrogolgin 1 | KVS*EIEDQLK | −1.29 | 0.001 | 0.20 | 0.484 |
| Q68FG2 | Sptbn2 | 2254 | Spectrin β-chain | RGS*LGFYK | −1.28 | 0.003 | 0.20 | 0.576 |
| P16254 | Srp14 | 44 | Signal recognition particle 14-kDa protein | KS*SVEGLEPAENK | −1.27 | 1.02 × 10−4 | 0.25 | 0.276 |
| P14152 | Mdh1 | 241 | Malate dehydrogenase, cytoplasmic | KLS*SAMSAAK | −1.17 | 1.75 × 10−4 | 0.07 | 0.768 |
| Q9ES28 | Arhgef7 | 830 | Rho guanine nucleotide exchange factor 7 | S*LEEEQR | −1.13 | 9.54 × 10−5 | 0.18 | 0.371 |
| Q9DBC7 | Prkar1a | 83 | cAMP-dependent protein kinase type I-α regulatory subunit | EDEIS*PPPPNPVVK | −1.05 | 1.34 × 10−4 | −0.09 | 0.642 |
| G3X9K3 | Arfgef1 | 1076 | Brefeldin A-inhibited guanine nucleotide-exchange protein 1 | EGS*LTGTK | −0.96 | 0.001 | 0.09 | 0.688 |
| Q9CZX9 | Emc4 | 36 | Endoplasmic reticulum membrane protein complex subunit 4 | SDRGS*GQGDSLYPVGYLDK | −0.93 | 0.003 | 0.18 | 0.485 |
| Q8CI52 | Gramd1c | 532 | Protein aster-C | SS*TDLGFEAK | −0.92 | 3.38 × 10−4 | −0.04 | 0.832 |
| Q91XA2 | Golm1 | 299 | Golgi membrane protein 1 | PEEDS*QYPER | −0.91 | 0.008 | 0.08 | 0.794 |
| Q9DBC7 | Prkar1a | 83 | cAMP-dependent protein kinase type I-α regulatory subunit | TDSREDEIS*PPPPNPVVK | −0.90 | 0.001 | −0.11 | 0.599 |
| Q8CHT1 | Ngef | 84 | Ephexin-1 | RAS*DQADLPK | −0.88 | 0.002 | 0.32 | 0.179 |
| Q8CI52 | Gramd1c | 531 | Protein aster-C | S*STDLGFEAK | −0.87 | 5.31 × 10−5 | 0.08 | 0.558 |
| P43277 | Hist1h1c; Hist1h1d |
37 | Histone H1.2; histone H1.3 | KAS*GPPVSELITK | −0.84 | 0.003 | 0.26 | 0.263 |
| Q8C341 | Suco | 1069 | SUN domain-containing ossification factor | RTS*FPLIR | −0.80 | 0.006 | 0.13 | 0.597 |
| Q8K4L3 | Svil | 857 | Supervillin | KLS*VDNNTSATDYK | −0.77 | 0.009 | −0.03 | 0.918 |
| Q67FY2 | Bcl9l | 118 | B cell CLL/lymphoma 9-like protein | SVS*VDSGEQR | −0.75 | 0.002 | −0.86 | 0.001 |
| F6ZDS4 | Tpr | 2204 | Nucleoprotein TPR | TVPS*TPTLVVPHR | −0.74 | 0.004 | −0.47 | 0.047 |
| Q8C079 | Strip1 | 59 | Striatin-interacting protein 1 | KDS*EGYSESPDLEFEYADTDK | −0.73 | 0.001 | 0.08 | 0.642 |
| Q9JKB3 | Ybx3 | 328 | Y-box-binding protein 3 | S*RPLNAVSQDGK | −0.72 | 0.004 | 0.09 | 0.644 |
| Q6NZF1 | Zc3h11a | 740 | Zinc finger CCCH domain-containing protein 11A | RLS*SASTGKPPLSVEDDFEK | −0.71 | 0.006 | −1.00 | 0.001 |
| O35609 | Scamp3 | 78 | Secretory carrier-associated membrane protein 3 | KLS*PTEPR | −0.71 | 0.004 | 0.18 | 0.389 |
| P15066 | Jund; Jun | 100 | Transcription factor jun-D | LAS*PELER | −0.69 | 0.002 | 0.07 | 0.720 |
| Q02248 | Ctnnb1 | 552 | Catenin β1 | RTS*M^GGTQQQFVEGVR | −0.69 | 0.003 | −0.15 | 0.455 |
| Q8BUH8 | Senp7 | 12 | Sentrin-specific protease 7 | RAS*SEIVTEGK | −0.67 | 8.08 × 10−5 | −0.67 | 8.49 × 10−5 |
| Q6PGG2 | Gmip | 260 | GEM-interacting protein | S*REEAQAK | −0.67 | 0.001 | 0.33 | 0.057 |
| Q04899 | Cdk18 | 75 | Cyclin-dependent kinase 18 | RLS*LPMDIR | −0.66 | 0.005 | −0.40 | 0.057 |
| P20357 | Map2 | 1635 | Microtubule-associated protein 2 | S*GILVPSEK | −0.65 | 0.005 | −0.66 | 0.004 |
| Q61165 | Slc9a1 | 707 | Sodium/hydrogen exchanger 1 | IGS*DPLAYEPK | −0.63 | 0.004 | 0.17 | 0.366 |
| Q8K3X4 | Irf2bpl; Irf2bp2 | 13 | Probable E3 ubiquitin-protein ligase IRF2BPL; interferon regulatory factor 2-binding protein 2 | RQS*CYLCDLPR | −0.63 | 0.001 | 0.33 | 0.047 |
| P70698 | Ctps1 | 575 | CTP synthase 1 | SGSSS*PDSEITELK | −0.61 | 0.009 | 0.36 | 0.089 |
| A0A087WQ44 | Srcap | 2771 | Snf2-related CREBBP activator protein | TS*ADVEIR | −0.59 | 0.004 | −0.49 | 0.011 |
| P81122 | Irs2 | 362 | Insulin receptor substrate 2 | TAS*EGDGGAAGGAGTAGGR | −0.58 | 0.004 | 0.19 | 0.249 |
| Q02248 | Ctnnb1 | 551 | Catenin β1 | RT*SM^GGTQQQFVEGVR | −0.57 | 0.008 | −0.06 | 0.754 |
| F7BJB9 | Morc3 | 563 | MORC family CW-type zinc finger protein 3 | RLS*NPPVENSSYK | −0.57 | 0.002 | −0.29 | 0.062 |
| Q811L6 | Mast4; Mast1; Mast2 | 1436 | Microtubule-associated serine/threonine-protein kinase 4; microtubule-associated serine/threonine-protein kinase 2 | SAEPPRS*PLLK | −0.56 | 0.001 | −0.41 | 0.007 |
| Q8BG05 | Hnrnpa3 | 361 | Heterogeneous nuclear ribonucleoprotein A3 | SSGSPY*GGGYGSGGGSGGYGSR | −0.54 | 0.003 | −0.23 | 0.143 |
| P35569 | Irs1 | 3 | Insulin receptor substrate 1 | AS*PPDTDGFSDVR | −0.54 | 2.04 × 10−4 | −0.09 | 0.418 |
| P97310 | Mcm2 | 21 | DNA replication licensing factor MCM2 | RIS*DPLTSSPGR | −0.51 | 0.001 | −0.11 | 0.378 |
| Q9QXZ0 | Macf1 | 3889 | Microtubule-actin cross-linking factor 1 | QGS*FSEDVISHK | −0.51 | 0.008 | 0.23 | 0.177 |
| Q8K310 | Matr3 | 188 | Matrin-3 | RDS*FDDR | −0.50 | 0.003 | 0.11 | 0.423 |
| Q9DBR1 | Xrn2 | 473 | 5′-3′ exoribonuclease 2 | NSSPS*ISPNTSFASDGSPSPLGGIK | 0.50 | 0.003 | −0.41 | 0.010 |
| E1U8D0 | Soga1 | 1300 | Protein SOGA1 | APS*PTTAAGEESCK | 0.52 | 0.006 | 0.11 | 0.494 |
| Q8BTI8 | Srrm2 | 1338 | Serine/arginine repetitive matrix protein 2 | S*SSELSPEVVEK | 0.53 | 0.006 | 0.10 | 0.561 |
| Q8BIA4 | Fbxw8 | 86 | F-box/WD repeat-containing protein 8 | SRS*PPDRDATEPEPLVDQLIR | 0.57 | 0.003 | −0.24 | 0.126 |
| Q99K30 | Eps8l2 | 217 | Epidermal growth factor receptor kinase substrate 8-like protein 2 | QPGDS*PQAK | 0.57 | 0.007 | −0.02 | 0.893 |
| P63058 | Thra | 12 | Thyroid hormone receptor α | VECGS*DPEENSAR | 0.61 | 0.006 | −0.24 | 0.208 |
| Q9CQT2 | Rbm7 | 108 | RNA-binding protein 7 | SGSSHASQDASVSYPQHHVGNLS*PTSTSPNSYER | 0.63 | 0.008 | −0.10 | 0.608 |
| Q9WTQ5 | Akap12 | 584 | A-kinase anchor protein 12 | GPSEAPQEAEAEEGATS*DGEKKR | 0.64 | 0.008 | −0.02 | 0.908 |
| Q9QXM1 | Jmy | 704 | Junction-mediating and -regulatory protein | STAS*PVPCEEQCHSLPTVLQGQEK | 0.67 | 0.005 | −0.29 | 0.166 |
| P39053 | Dnm1 | 774; 777 | Dynamin-1 | RS*PTS*SPTPQR | 0.67 | 0.002 | −0.31 | 0.105 |
| Q9Z1T6 | Pikfyve | 1753 | 1-Phosphatidylinositol 3-phosphate 5-kinase | GTAGKS*PDLSSQK | 0.68 | 5.23 × 10−5 | −0.25 | 0.039 |
| Q8BTI8 | Srrm2 | 1343 | Serine/arginine repetitive matrix protein 2 | SSSELS*PEVVEK | 0.68 | 0.002 | 0.13 | 0.472 |
| Q8C078 | Camkk2 | 91 | Calcium/calmodulin-dependent protein kinase kinase 2 | DQPPEADGQELPLEASDPESRS*PLSGR | 0.75 | 9.91 × 10−5 | −0.14 | 0.317 |
| P58802 | Tbc1d10a | 407 | TBC1 domain family member 10A | AILDAEPGPRPALQPS*PSIR | 0.78 | 2.06 × 10−4 | −0.34 | 0.039 |
| P43274 | Hist1h1c; Hist1h1t; Hist1h1e; Hist1h1b; Hist1h1d |
102 | Histone H1.2; histone H1.4; histone H1.5; histone H1.3 |
GTGAS*GSFK | 0.79 | 0.006 | 0.10 | 0.663 |
| Q9ESE1 | Lrba | 979 | Lipopolysaccharide-responsive and beige-like anchor protein | DS*PISPHFTR | 0.80 | 0.003 | −0.10 | 0.659 |
| Q6PDN3 | Mylk | 355 | Myosin light chain kinase, smooth muscle | VPAIGSFS*PGEDRK | 0.81 | 0.001 | −0.21 | 0.261 |
| P56402 | Aqp2 | 256; 261 | Aquaporin-2 | QS*VELHS*PQSLPR | 1.02 | 4.83 × 10−4 | 0.44 | 0.064 |
| P40645 | Sox6 | 454 | Transcription factor SOX-6 | TS*PVNLPNK | 1.19 | 0.007 | −0.10 | 0.783 |
| P40645 | Sox6 | 439 | Transcription factor SOX-6 | S*PTSPTQNLFPASK | 1.40 | 0.002 | 0.13 | 0.708 |
PKA-Cβ-null data for the same peptides are given for comparison. Pmod value, moderated P value.
Multiple gene symbols are given when a peptide matches sequences in multiple proteins. *Phosphorylated amino acids.
Table 4.
Phosphopeptides changed most convincingly in PKA-Cβ -null cells versus PKA-Cβ intact [Pmod < 0.01 and |log(ratio)| > 0.5]
| UniProt ID | Gene Symbol | Amino Acid Number | Annotation | Sequence | Log2 (PKA-Cα Null/ PKA-Cα Intact) | Pmod Value | Log2 (PKA-Cβ Null/ PKA-Cβ Intact) | Pmod Value |
|---|---|---|---|---|---|---|---|---|
| Q6NZF1 | Zc3h11a | 740 | Zinc finger CCCH domain-containing protein 11A | RLS*SASTGKPPLSVEDDFEK | −0.71 | 0.006 | −1.00 | 0.001 |
| Q67FY2 | Bcl9l | 118 | B cell CLL/lymphoma 9-like protein | SVS*VDSGEQR | −0.75 | 0.002 | −0.86 | 0.001 |
| Q03173 | Enah | 719 | Protein enabled homolog | APST*STPEPTR | 0.00 | 0.988 | −0.79 | 0.002 |
| Q9QZQ1 | Mllt4 | 1201 | Afadin | ITSVS*TGNLCTEEQSPPPRPEAYPIPTQTYTR | −0.05 | 0.793 | −0.77 | 0.002 |
| O08796 | Eef2k | 66 | Eukaryotic elongation factor 2 kinase | T*ECGSTGSPASSFHFK | −0.13 | 0.552 | −0.75 | 0.004 |
| Q03173 | Enah | 720 | Protein enabled homolog | APSTS*TPEPTR | −0.11 | 0.633 | −0.75 | 0.005 |
| O55003 | Bnip3 | 60 | BCL2/adenovirus E1B 19 kDa protein-interacting protein 3 | SSHCDS*PPR | 0.27 | 0.222 | −0.70 | 0.007 |
| Q2VPU4 | Mlxip | 9 | MLX-interacting protein | AADVFM^CS*PR | −0.03 | 0.897 | −0.68 | 0.010 |
| Q8BUH8 | Senp7 | 12 | Sentrin-specific protease 7 | RAS*SEIVTEGK | −0.67 | 8.08 × 10−5 | −0.67 | 8.49 × 10−5 |
| P20357 | Map2 | 1635 | Microtubule-associated protein 2 | S*GILVPSEK | −0.65 | 0.005 | −0.66 | 0.004 |
| Q8VDZ4 | Zdhhc5 | 409 | Palmitoyltransferase ZDHHC5 | SEPSLEPESFRS*PTFGK | −0.27 | 0.144 | −0.64 | 0.003 |
| Q8CGF1 | Arhgap29 | 1241 | Rho GTPase-activating protein 29 | ESSEEPALPEGT*PTCQRPR | −0.04 | 0.743 | −0.63 | 0.001 |
| Q8VI36 | Pxn | 258 | Paxillin | IS*ASSATR | −0.01 | 0.966 | −0.62 | 0.004 |
| Q8C6B2 | Rtkn | 518 | Rhotekin | T*FSLDAAPADHSLGPSR | −0.01 | 0.962 | −0.60 | 0.005 |
| Q8C6B2 | Rtkn | 520 | Rhotekin | TFS*LDAAPADHSLGPSR | −0.13 | 0.423 | −0.60 | 0.002 |
| A1A535 | Veph1 | 422 | Ventricular zone-expressed PH domain-containing protein 1 | INAESNT*PGSGR | −0.34 | 0.070 | −0.59 | 0.005 |
| Q9Z2H5 | Epb41l1 | 540 | Band 4.1-like protein 1 | RLPS*SPASPSPK | −0.17 | 0.332 | −0.57 | 0.006 |
| P28661 | Sept4 | 68 | Septin-4 | PQS*PDLCDDDVEFR | −0.02 | 0.917 | −0.55 | 0.003 |
| Q8C5R2 | Proser2 | 223 | Proline and serine-rich protein 2 | LAGNEALSPTS*PSK | −0.26 | 0.119 | −0.55 | 0.004 |
| Q99MR1 | Gigyf1 | 227 | GRB10-interacting GYF protein 1 | ST*SPDGGPR | −0.05 | 0.640 | −0.51 | 0.001 |
| Q99MR1 | Gigyf1 | 226 | GRB10-interacting GYF protein 1 | S*TSPDGGPR | 0.05 | 0.723 | −0.50 | 0.003 |
| Q60864 | Stip1 | 481 | Stress-induced-phosphoprotein 1 | HDS*PEDVK | −0.12 | 0.498 | 0.57 | 0.005 |
PKA-Cα-null data for the same peptides are also given for comparison. Pmod value, moderated P value. *Phosphorylated amino acids.
Fig. 5.
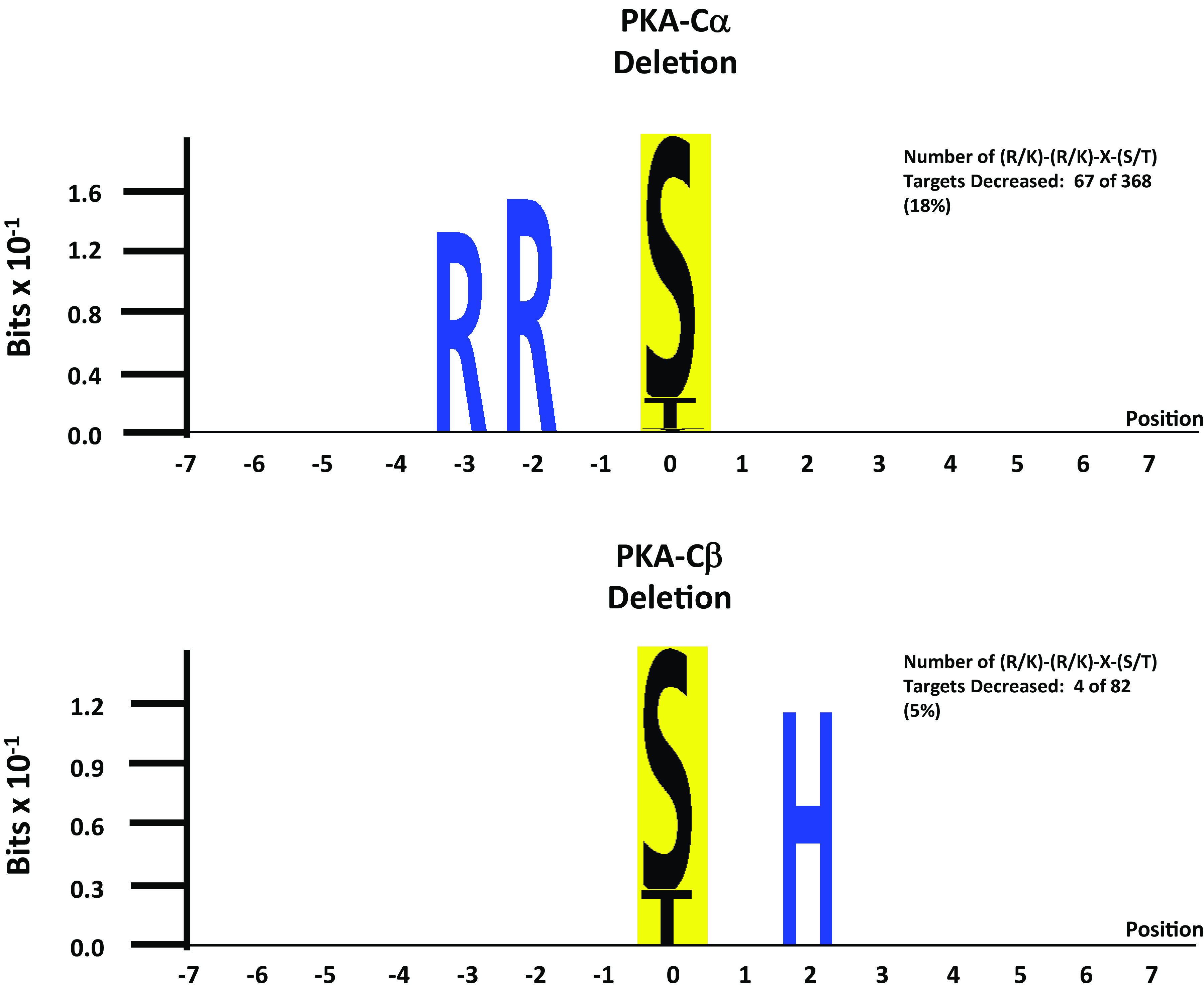
Sequence preference logos from sites decreased with PKA-Cα deletion (top) and PKA-Cβ deletion (bottom). Logos were generated using PTM-Logo with a background of all unchanged phosphopeptides.
In vitro phosphorylation by recombinant PKA-Cα versus PKA-Cβ.
PKA-Cα and PKA-Cβ may have different target preferences. To test this hypothesis directly, we carried out in vitro phosphorylation experiments using purified recombinant PKA-Cα and PKA-Cβ to phosphorylate protein extracts obtained from PKA dKO cells. The full data are shown in Supplemental Data Set S3. Figure 6A shows a comparison of the changes in phosphorylation by PKA-Cα versus PKA-Cβ, demonstrating that both PKA catalytic subunits phosphorylate virtually identical substrates. Figue 6B shows that the sequence preferences for PKA-Cα versus PKA-Cβ are virtually identical. This result rules out the hypothesis that the two PKA catalytic subunits have different substrate specificities. The remaining possibility, that the two catalytic subunits have different targets because they are propinquitous to a different set of proteins, remains as the most likely explanation for the differences in targets in the intact cells.
Fig. 6.
Results of in vitro phosphorylation experiments. Protein extracts from PKA double knockout cells (PKA-Cα and PKA-Cβ) were incubated with recombinant PKA-Cα or PKA-Cβ, and phosphorylation was quantified by mass spectrometry. A: there was a marked similarity between responses to the two recombinant kinase proteins. B: sequence preference logos derived from the analysis were almost identical.
Phosphoproteomics as a virtual proximity assay.
In addition to the target sequence, another factor important to the determination of targets for a protein kinase is colocalization because a kinase can only phosphorylate a protein with which it comes into physical contact. In this sense, phosphoproteomic analysis can be viewed as a kind of large-scale proximity assay, identifying kinase/target interactions. To evaluate whether differences in PKA-Cα and PKA-Cβ phosphorylation targets result in part from differences in PKA-Cα and PKA-Cβ localization, we identified Gene Ontology (GO) cellular component terms that are enriched in either set of phosphorylation targets in the intact cell experiments, relative to all proteins detected. Table 5 shows terms enriched in phosphoproteins with altered phosphorylation in PKA-Cα-null cells, whereas Table 6 shows terms enriched in phosphoproteins with altered phosphorylation in PKA-Cβ-null cells. Enriched terms in PKA-Cα-null cells were largely related to cell membranes and membrane vesicles, whereas enriched terms in PKA-Cβ-null cells were related to the actin cytoskeleton and cell junctions, suggesting that in vivo cellular subdomains of PKA-Cα and PKA-Cβ differ.
Table 5.
Gene Ontology cellular component terms enriched in the list of phosphoproteins with altered phosphorylation in PKA-Cα-null cells
| Term | Number | Fold Enrichment | Fisher Exact | Proteins |
|---|---|---|---|---|
| Membrane raft | 8 | 4.3 | 3.20×10−4 | EGFR, EFHD2, PRKAR2B, PRKAR1A, PIKFYVE, CXADR, CTNNB1, HDAC6 |
| Perinuclear region of cytoplasm | 12 | 2.3 | 3.90×10−3 | EGFR, PACS1, EPN3, PRKAR2B, FBXW8, PAK2, SCYL2, PIKFYVE, SPTBN2, RAB3IP, CTNNB1, HDAC6 |
| Membrane part | 26 | 1.6 | 6.40×10−3 | GPRC5C, ACSS2, CXADR, CHCHD6, CTNNB1, EFHD2, PRKAR2B, PIKFYVE, PRKAA1, SYNPO, SREBF1, EGFR, EPN3, OLFR120, KANSL1, SLC33A1, STIM2, CRB3, EMC4, BNIP3L, PRKAR1A, SPTBN2, GRAMD1C, DNM1, HDAC6, SUCO |
| Cytoplasmic vesicle | 11 | 2 | 1.90×10−2 | EGFR, PACS1, SREBF1, ARHGAP21, EPN3, SCYL2, RAN, PIKFYVE, SPTBN2, CXADR, DNM1 |
| Integral component of membrane | 16 | 1.6 | 3.60×10−2 | SREBF1, EGFR, OLFR120, GPRC5C, KANSL1, SLC33A1, STIM2, CRB3, ACSS2, CXADR, CHCHD6, EMC4, BNIP3L, PIKFYVE, GRAMD1C, SUCO |
| Membrane-bounded vesicle | 21 | 1.4 | 5.00×10−2 | SREBF1, PACS1, EGFR, SRP14, EPN3, GPRC5C, RAN, CRB3, CXADR, HNRNPA1, CTNNB1, ARHGAP21, FAM65A, PRKAR2B, RPL30, SCYL2, PIKFYVE, SPTBN2, TUBA1B, DNM1, MDH1 |
Table 6.
Gene Ontology cellular component terms enriched in the list of phosphoproteins with altered phosphorylation in PKA-Cβ-null cells
| Term | Number | Fold Enrichment | Fisher Exact | Proteins |
|---|---|---|---|---|
| Cytoskeleton | 23 | 1.7 | 2.70 × 10−3 | DPF2, SEPT4, ENAH, DYNC1LI2, PDLIM5, AKAP12, IGF2BP2, NEXN, MYO9A, PXN, SMC3, TNKS1BP1, EPB41L1, MACF1, MAP2, MAP4, RANBP2, CDC42EP4, CEP170B, UBXN6, AFAP1, SYNPO, SEPT9 |
| Adherens junction | 16 | 1.9 | 5.10 × 10−3 | EGFR, ENAH, PDLIM5, AKAP12, NEXN, CXADR, PXN, TNKS1BP1, EPB41L1, LIMD1, EEF1D, ERC1, EPS8L2, AFAP1, TJP2, SEPT9 |
| Cell junction | 19 | 1.7 | 1.30 × 10−2 | CLDN8, EGFR, ENAH, PDLIM5, AKAP12, CXADR, NEXN, PXN, TNKS1BP1, EPB41L1, LIMD1, EEF1D, ERC1, CDC42EP4, EPS8L2, AFAP1, TJP2, SYNPO, SEPT9 |
| Plasma membrane | 21 | 1.5 | 1.90 × 10−2 | CLDN8, EGFR, ENAH, IRS2, ZDHHC5, PDLIM5, LRBA, AKAP12, WWC1, ZBTB16, CXADR, PXN, AQP2, TNKS1BP1, EPB41L1, MACF1, MAP4, CDC42EP4, EPS8L2, TJP2, SYNPO |
| Actin cytoskeleton | 9 | 2.1 | 2.60 × 10−2 | ENAH, MACF1, PDLIM5, CDC42EP4, MYO9A, AFAP1, PXN, SEPT9, SYNPO |
PKA-Cα and PKA-Cβ regulate different protein kinases.
Many phosphorylation sites that changed in PKA-Cα-null or PKA-Cβ-null cells did not conform to the classic PKA target motif and presumably undergo changes in phosphorylation as a result of secondary effects on other protein kinases. Table 7 shows protein kinase catalytic proteins that underwent changes in phosphorylation in either PKA-Cα-null or PKA-Cβ-null cells. Many of the altered phosphorylation sites have known effects on enzyme activity, as indicated in the last column. The affected kinases, namely, calcium/calmodulin-dependent protein kinase kinase 2 (Camkk2), epidermal growth factor receptor kinase (Egfr), myosin light chain kinase (Mylk), p21-activated kinase (Pak2), AMP-activated protein kinase-α1 (Prkaa1), and salt-inducible kinase 2 (Sik2), all underwent changes in phosphorylation in response to PKA-Cα deletion, whereas only Egfr underwent a change in phosphorylation in response to PKA-Cβ deletion. This supports the conclusion that PKA-Cα or PKA-Cβ have different downstream signaling networks.
Table 7.
Protein kinases with altered phosphorylation in response to deletion of PKA-Cα or PKA-Cβ
| Gene Symbol | Annotation | Amino Acid Number | Centralized Sequence‡ | Log2 (PKA-Cα Null/ PKA-Cα Intact) | Pmod Value | Log2 (PKA-Cβ Null/ PKA-Cβ Intact) | Pmod Value | Kinase Class | Effect of Phosphorylation on Activity† |
|---|---|---|---|---|---|---|---|---|---|
| Camkk2 | Calcium/calmodulin-dependent protein kinase kinase 2 | 85 | QELPLEAS*DPESRSP | 0.41 | 0.005 | −0.17 | 0.169 | Other | Increases |
| Camkk2 | Calcium/calmodulin-dependent protein kinase kinase 2 | 91 | ASDPESRS*PLSGRKM | 0.75 | 9.91 × 10−5 | −0.14 | 0.317 | Other | NI |
| Cdk18 | Cyclin-dependent kinase 18 | 75 | EDLNKRLS*LPMDIRL | −0.61 | 0.012 | −0.39 | 0.086 | CMGC | NI |
| Cdk18 | Cyclin-dependent kinase 18 | 109 | TRMSRRAS*LSDIGFG | −0.65 | 0.042 | −0.18 | 0.543 | CMGC | NI |
| Eef2k | Eukaryotic elongation factor 2 kinase | 66 | YYSNLTKT*ECGSTGS | −0.13 | 0.552 | −0.75 | 0.004 | Atypical | NI |
| Egfr | Epidermal growth factor receptor | 695 | RELVEPLT*PSGEAPN | −0.55 | 0.024 | −0.63 | 0.012 | Tyr | Altered receptor internalization |
| Map4k5 | Mitogen-activated protein kinase kinase kinase kinase 5 | 335 | SRAERTAS*EINFDKL | −0.70 | 0.028 | −0.48 | 0.112 | STE | NI |
| Map4k5 | Mitogen-activated protein kinase kinase kinase kinase 5 | 400 | PPKPRVNT*YPEDSLP | −0.80 | 0.027 | −0.04 | 0.894 | STE | NI |
| Mast4 | Microtubule-associated serine/threonine-protein kinase 4 | 1382 | SPLARTPS*PTPQPTS | 0.04 | 0.885 | −0.55 | 0.047 | AGC | NI |
| Mast4 or Mast1 or Mast2 | Microtubule-associated serine/threonine-protein kinase 4; microtubule-associated serine/threonine-protein kinase 2 | 1436 | KSAEPPRS*PLLKRVQ | −0.559 | 0.001 | −0.411 | 0.007 | AGC | NI |
| Mylk | Myosin light chain kinase, smooth muscle | 355 | VPAIGSFS*PGEDRK | 0.81 | 0.001 | −0.21 | 0.261 | CAMK | Increases |
| Nrbp2 | Nuclear receptor-binding protein 2 | 20 | EREREDES*EDESDIL | −0.02 | 0.870 | −0.42 | 0.002 | Other | NI |
| Pak2 | Serine/threonine-protein kinase PAK 2 | 141 | TVKQKYLS*FTPPEKD | −2.00 | 0.043 | −1.30 | 0.167 | STE | Increases |
| Pak2 | Serine/threonine-protein kinase PAK 2 | 197 | TKSIYTRS*VIDPIPA | −1.30 | 0.039 | −0.21 | 0.719 | STE | Increases? |
| Prkaa1 | 5′-AMP-activated protein kinase catalytic subunit α1 | 496 | ATPQRSGS*ISNYRSC | −0.41 | 0.009 | −0.13 | 0.347 | CAMK | Decreases |
| Scyl2 | SCY1-like protein 2 | 677 | QGKQKRGS*LTLEEKQ | −0.47 | 0.006 | 0.09 | 0.541 | Other | NI |
| Sik2 | Serine/threonine-protein kinase SIK2 | 342 | ERLKSHRS*SFPVEQR | −0.42 | 0.013 | −0.14 | 0.333 | CAMK | Increases? |
| Slk | STE20-like serine/threonine-protein kinase | 777 | SKAKDSGS*VSLQETR | −0.21 | 0.293 | −0.43 | 0.041 | STE | NI |
| Wnk1 | Serine/threonine-protein kinase WNK1 | 165 | TSKDRPVS*QPSLVGS | 0.09 | 0.578 | 0.42 | 0.020 | Other | NI |
Pmod value, moderated P value; NI, no information available.
Phosphorylated amino acids. R or K in positions −3 and −2 are underlined.
Based on information at PhospoSitePlus (https://www.phosphosite.org).
Differential regulation of cAMP signaling proteins by PKA-Cα and PKA-Cβ.
One factor involved in the localization of PKA in the cell is its interaction with anchoring proteins such as A-kinase anchoring proteins (AKAPs). Table 8 shows altered phosphopeptides belonging to AKAP proteins or proteins with GO terms containing “cAMP” or “cyclic-AMP” to identify possible other AKAP interactors. Among the phosphoproteins in this list are Akap1, Akap12, cAMP phosphodiesterase 4C (Pde4c), cAMP phosphodiesterase 7A (Pde7a), and Prkaa1 as well as two PKA regulatory subunits, RIα (Prkar1a) and RIIβ (Prkar2b). All of these (except for one site in Akap12) underwent changes in response to PKA-Cα deletion but not PKA-Cβ deletion. Thus, the phosphorylation evidence suggests that PKA-Cα interacts more extensively with components of AKAP complexes than does PKA-Cβ, at least with regard to phosphorylation.
Table 8.
AKAPs and cAMP-associated proteins with altered phosphorylation in either PKA-Cα or PKA-Cβ-null cells
| UniProt ID | Gene Symbol | Annotation | Amino Acid Number | Centralized Sequence† | Log2(PKA-Cα Null/ PKA-Cα Intact) | Pmod Value | Log2(PKA-Cβ Null/ PKA-Cβ Intact) | Pmod Value |
|---|---|---|---|---|---|---|---|---|
| O08715 | Akap1 | A-kinase anchor protein 1, mitochondrial | 101 | TRQVRRRS*ESSGNLP | −0.41 | 0.003 | 0.00 | 0.966 |
| Q9WTQ5 | Akap12 | A-kinase anchor protein 12 | 22 | PAESDTPS*ELELSGH | 0.55 | 0.035 | −0.09 | 0.716 |
| Q9WTQ5 | Akap12 | A-kinase anchor protein 12 | 583 | AEAEEGAT*SDGEKKR | 0.43 | 0.018 | 0.11 | 0.490 |
| Q9WTQ5 | Akap12 | A-kinase anchor protein 12 | 584 | EAEEGATS*DGEKKRE | 0.64 | 0.008 | −0.02 | 0.908 |
| Q9WTQ5 | Akap12 | A-kinase anchor protein 12 | 683 | KRARKASS*SDDEGGP | 0.51 | 0.011 | −0.11 | 0.514 |
| Q9WTQ5 | Akap12 | A-kinase anchor protein 12 | 684 | RARKASSS*DDEGGPR | 0.52 | 0.019 | −0.20 | 0.305 |
| Q9WTQ5 | Akap12 | A-kinase anchor protein 12 | 1292 | SNEEQSIS*PEKREMG | 0.40 | 0.061 | 0.49 | 0.026 |
| Q3UEI1 | Pde4c | cAMP-specific 3′,5′-cyclic phosphodiesterase 4C | 301 | ELRRSSHT*SLPTAAI | −0.43 | 0.034 | 0.26 | 0.173 |
| P70453 | Pde7a | High-affinity cAMP-specific 3′,5′-cyclic phosphodiesterase 7A | 58 | FETERRGS*HPYIDFR | −0.44 | 0.020 | −0.40 | 0.031 |
| Q5EG47 | Prkaa1 | 5′-AMP-activated protein kinase catalytic subunit α1 | 496 | ATPQRSGS*ISNYRSC | −0.41 | 0.009 | −0.13 | 0.347 |
| Q9DBC7 | Prkar1a | cAMP-dependent protein kinase type I-α regulatory subunit | 83 | DSREDEIS*PPPPNPV | −1.01 | 3.99 × 10−4 | 0.05 | 0.816 |
| P31324 | Prkar2b | cAMP-dependent protein kinase type II-β regulatory subunit | 112 | NRFTRRAS*VCAEAYN | −1.38 | 0.001 | −0.29 | 0.402 |
Pmod value, moderated P value.
Phosphorylated amino acids. R or K in positions −3 and −2 are underlined.
An important role of vasopressin signaling in renal collecting duct cells is transcriptional regulation, particularly regulation of transcription of the Aqp2 gene. Table 9 shows transcription factors with altered phosphorylation in PKA-Cα-null or PKA-Cβ-null cells. Among all transcription factors, 241 phosphopeptides were quantified. There were 14 phosphopeptides that were substantially changed in abundance in 13 different transcription factor proteins. Most of the altered sites had proline in position +1 relative to the phosphorylated S or T, signifying altered phosphorylation by protein kinases in the cyclin-dependent kinase or MAPK families. Only one transcription factor underwent differential phosphorylation at a site with the (R/K)-(R/K)-X-p(S/T) motif, namely, “cAMP-dependent transcription factor ATF-7,” which showed a marked decrease in phosphorylation in PKA-Cα-null but not PKA-Cβ-null cells. Atf7 is a b-ZIP transcription factor that is inactive as a homodimer but can transactivate genes when heterodimerized with members of the activator protein-1 family, including Jund (7), which underwent a decrease in phosphorylation. Beyond the transcription factors, various transcriptional coregulators may participate in vasopressin-mediated regulation of transcription. One such protein is β-catenin, which shows increased phosphorylation at S552, a PKA target site (10), in response to vasopressin. Interestingly, S552 of β-catenin showed a marked decrease in phosphorylation in PKA-Cα-null cells but not PKA-Cβ-null cells (Table 3). In general, as with other categories of proteins, changes in phosphorylation of transcription factors and their coregulators are different in PKA-Cα-null or PKA-Cβ-null cells, again supporting the conclusion that the two PKA catalytic subunits fulfill different regulatory functions.
Table 9.
Phosphopeptides from transcription factor proteins that underwent changes in abundance with PKA-Cα and/or PKA-Cβ deletion
| UniProt ID | Gene Symbol | Annotation | Amino Acid Number | Centralized Sequence | Log2 (PKA-Cα Null/ PKA-Cα Intact) | Pmod Value | Log2 (PKA-Cβ Null/ PKA-Cβ Intact) | Pmod Value | Transcription Factor Family |
|---|---|---|---|---|---|---|---|---|---|
| Q3US59 | Atf7 | cAMP-dependent transcription factor ATF-7 | 326 | TGGRRRRT*VDEDPDE | −1.19 | 0.039 | −0.60 | 0.266 | bZIP |
| Q61103 | Dpf2 | Zinc finger protein ubi-d4 | 142 | DPRVDDDS*LGEFPVS | −0.44 | 0.061 | −0.59 | 0.017 | zf-C2H2 |
| Q8K5C0 | Grhl2 | Grainyhead-like protein 2 homolog | 2 | ______MS*QESDNNK | 0.42 | 0.038 | −0.11 | 0.539 | CP2 |
| P15066 | Jund (or Jun) | Transcription factor jun-D (or jun) | 100 | LGLLKLAS*PELERLI | −0.69 | 0.002 | 0.07 | 0.720 | bZIP |
| Q2VPU4 | Mlxip | MLX-interacting protein | 9 | AADVFMCS*PRRPRSR | −0.03 | 0.897 | −0.68 | 0.010 | bHLH |
| P40645 | Sox6 | Transcription factor SOX-6 | 439 | KTAEPVKS*PTSPTQN | 1.40 | 0.002 | 0.13 | 0.708 | HMG |
| P40645 | Sox6 | Transcription factor SOX-6 | 454 | LFPASKTS*PVNLPNK | 1.19 | 0.007 | −0.10 | 0.783 | HMG |
| Q9WTN3 | Srebf1 | Sterol regulatory element-binding protein 1 | 96 | KVTPAPLS*PPPSAPA | −0.80 | 0.024 | 0.22 | 0.500 | bHLH |
| Q9JM73 | Srf | Serum response factor | 220 | TCLNSPDS*PPRSDPT | 0.41 | 0.007 | −0.05 | 0.710 | SRF |
| P63058 | Thra | Thyroid hormone receptor α | 12 | PSKVECGS*DPEENSA | 0.61 | 0.006 | −0.24 | 0.208 | TH |
| P62960 | Ybx1 | Nuclease-sensitive element-binding protein 1 | 174 | EKNEGSES*APEGQAQ | −0.47 | 0.020 | −0.05 | 0.792 | CSD |
| Q3UQ17 | Zbtb16 | MCG3834 | 307 | EESGEQLS*PPVEAGQ | −0.44 | 0.236 | −1.04 | 0.012 | ZBTB |
| Q8BIQ6 | Zfp947 | MCG23335 | 283 | FTQKSSLS*IHQMYHT | 0.50 | 0.044 | 0.07 | 0.768 | zf-C2H2 |
| O35615 | Zfpm1 | Zinc finger protein ZFPM1 | 681 | RGSEGSQS*PGSSVDD | 0.68 | 0.030 | −0.06 | 0.843 | zf-C2H2 |
Phosphorylated amino acids. R or K in positions −3 and −2 are underlined.
DISCUSSION
Based on the observations in this report, we conclude that PKA-Cα and PKA-Cβ do not have redundant regulatory functions. Indeed, while some overlap exists between the two in terms of phosphorylation targets, large differences were seen at a whole phosphoproteome level. PKA-Cα-null cells showed decreased phosphorylation dominated by sites with the classical PKA motif, viz. (R/K)-(R/K)-X-p(S/T), whereas only very few of the decreased phosphorylation sites in PKA-Cβ-null cells contained this motif. However, in vitro incubation with recombinant PKA-Cα and PKA-Cβ resulted in phosphorylation of virtually identical sites, with a predominance of the (R/K)-(R/K)-X-p(S/T) motif. A key question is “If the catalytic regions of PKA-Cα and PKA-Cβ have nearly identical in vitro target specificities, why is there such a difference in phosphorylation targets in the intact cells?”. The question was addressed by the bioinformatics analysis of phosphorylation targets, showing that completely different GO cellular component terms are associated with the two sets of phosphorylation targets. Specifically, PKA-Cα targets were largely related to cell membranes and membrane vesicles, whereas PKA-Cβ targets were related to the actin cytoskeleton and cell junctions, indicating that PKA-Cα and PKA-Cβ interact with different sets of proteins in cells. This difference could arise from differences in the noncatalytic regions of the two kinases that result in a different set of local interactions with potential substrates.
One possibility is that the two catalytic subunits could interact with different AKAPs and/or PKA regulatory subunits (16). Phosphoproteomic results point to a preferential association of PKA-Cα with Prkar2b, which showed a marked decrease at S112 in PKA-Cα null cells but not PKA-Cβ null cells (Table 8). The second possibility is that there could be differential interactions with so-called C-kinase anchoring proteins (26). The third possibility is that there may be a difference in protein-protein interactions with PDZ domain-containing proteins (2). PKA-Cα contains a class 1 COOH-terminal PDZ-ligand motif (–TxF), whereas PKA-Cβ ends with –CxF, which does not conform to a PDZ ligand motif. In a previous study (4), we showed that vasopressin treatment of native inner medullary collecting duct cells results in increased phosphorylation of several PDZ domain-containing proteins [connector enhancer of kinase suppressor of Ras 3 (Cnksr3), PDZ domain-containing protein GIPC1 (Gipc1), multiple-PDZ domain protein (Mpdz), and PDZ and LIM domain protein-5 (Pdlim5)]. That study showed that proteins with class I COOH-terminal three-amino acid motifs are more likely to show increases in phosphorylation in response to vasopressin than proteins without the motif.
In addition to phosphorylation differences, PKA-Cα and PKA-Cβ deletions resulted in many differences in what proteins underwent changes in total protein abundances. Thus, in terms of regulation of protein abundances, PKA-Cα and PKA-Cβ proteins appear to have nonredundant regulatory functions. For example, AQP2 was markedly decreased in PKA-Cα-null cells but not in PKA-Cβ-null cells, despite similar expression levels of the two (10). This result suggests that PKA-Cα is the predominant catalytic subunit responsible for the regulation of AQP2 abundance. Similarly, C3 was markedly decreased in PKA-Cα-null cells but not in PKA-Cβ-null cells. In contrast, the abundance of Muc4 was substantially decreased in PKA-Cβ-null cells but not in PKA-Cα-null cells. A prior study in mpkCCD cells has shown that, in PKA dKO cells, AQP2, C3, and Muc4 mRNA and protein levels were markedly decreased (10).
Data resource.
In addition to the scientific findings highlighted above, this report provides added value in the form of two web resources that allow users to interrogate data from this paper. These resources have been included with other phosphoproteomic data on the Kidney Systems Biology Project website (https://hpcwebapps.cit.nih.gov/ESBL/Database/).
Limitations and future directions.
One limitation of this study is that the use of trypsin, in some cases, produces peptides that are too short to be quantified by mass spectrometry, causing certain important phosphorylation sites to be missed in the analysis. For example, the pKID domain of Creb1 contains a classic PKA phosphorylation site at S133 within the sequence LSRRPS*YRKILNDLS, which, when proteolyzed with trypsin, gives RPS*YR, which is too short to see by mass spectrometry. Future studies are needed using other proteases to allow more comprehensive analysis. Another limitation of the present study is that it was done exclusively in a cultured cell model of the collecting duct. Future studies are needed to assess the specific roles of the two PKA catalytic proteins in the native collecting duct.
GRANTS
This work was funded by National Heart, Lung, and Blood Institute (NHLBI) Projects ZIAHL001285 and ZIAHL006129 (to M.A.K.). K. Salhadar was a member of the Biomedical Engineering Student Internship Program (Robert Lutz, Director) supported by the National Institute for Biomedical Imaging and Bioengineering (June–August 2018). Mass spectrometry used the NHLBI Proteomics Core Facility (M. Gucek, Director).
DISCLOSURES
No conflicts of interest, financial or otherwise, are declared by the author(s).
AUTHOR CONTRIBUTIONS
V.R. and M.A.K. conceived and designed research; V.R., K.S. and C.Y. performed experiments; V.R., K.S., K.L., C.Y., and M.A.K. analyzed data; V.R., K.S., K.L., E.P., C.Y., and M.A.K. interpreted results of experiments; V.R., K.S., and M.A.K. prepared figures; V.R. and M.A.K. drafted manuscript; V.R., K.S., K.L., E.P., C.Y., and M.A.K. edited and revised manuscript; V.R., K.S., K.L., E.P., C.Y., and M.A.K. approved final version of manuscript.
ENDNOTES
Supplemental data are deposited at https://hpcwebapps.cit.nih.gov/ESBL/Database/Supplemental_Data_PKA_sKO/.
The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the data set Accession No. PXD015050 (https://www.ebi.ac.uk/pride/).
REFERENCES
- 1.Chen L, Lee JW, Chou CL, Nair AV, Battistone MA, Păunescu TG, Merkulova M, Breton S, Verlander JW, Wall SM, Brown D, Burg MB, Knepper MA. Transcriptomes of major renal collecting duct cell types in mouse identified by single-cell RNA-seq. Proc Natl Acad Sci USA 114: E9989–E9998, 2017. doi: 10.1073/pnas.1710964114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Colledge M, Dean RA, Scott GK, Langeberg LK, Huganir RL, Scott JD. Targeting of PKA to glutamate receptors through a MAGUK-AKAP complex. Neuron 27: 107–119, 2000. doi: 10.1016/S0896-6273(00)00013-1. [DOI] [PubMed] [Google Scholar]
- 3.Cox J, Mann M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat Biotechnol 26: 1367–1372, 2008. doi: 10.1038/nbt.1511. [DOI] [PubMed] [Google Scholar]
- 4.Deshpande V, Kao A, Raghuram V, Datta A, Chou CL, Knepper MA. Phosphoproteomic identification of vasopressin V2 receptor-dependent signaling in the renal collecting duct. Am J Physiol Renal Physiol 317: F789–F804, 2019. doi: 10.1152/ajprenal.00281.2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Fenton RA, Murali SK, Moeller HB. Advances in aquaporin-2 trafficking mechanisms and their implications for treatment of water balance disorders. Am J Physiol Cell Physiol 319: C1–C10, 2020. doi: 10.1152/ajpcell.00150.2020. [DOI] [PubMed] [Google Scholar]
- 6.Fushimi K, Uchida S, Hara Y, Hirata Y, Marumo F, Sasaki S. Cloning and expression of apical membrane water channel of rat kidney collecting tubule. Nature 361: 549–552, 1993. doi: 10.1038/361549a0. [DOI] [PubMed] [Google Scholar]
- 7.Hai T, Curran T. Cross-family dimerization of transcription factors Fos/Jun and ATF/CREB alters DNA binding specificity. Proc Natl Acad Sci USA 88: 3720–3724, 1991. doi: 10.1073/pnas.88.9.3720. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Hasler U, Mordasini D, Bens M, Bianchi M, Cluzeaud F, Rousselot M, Vandewalle A, Feraille E, Martin PY. Long term regulation of aquaporin-2 expression in vasopressin-responsive renal collecting duct principal cells. J Biol Chem 277: 10379–10386, 2002. doi: 10.1074/jbc.M111880200. [DOI] [PubMed] [Google Scholar]
- 9.Hasler U, Nielsen S, Féraille E, Martin PY. Posttranscriptional control of aquaporin-2 abundance by vasopressin in renal collecting duct principal cells. Am J Physiol Renal Physiol 290: F177–F187, 2006. doi: 10.1152/ajprenal.00056.2005. [DOI] [PubMed] [Google Scholar]
- 10.Isobe K, Jung HJ, Yang CR, Claxton J, Sandoval P, Burg MB, Raghuram V, Knepper MA. Systems-level identification of PKA-dependent signaling in epithelial cells. Proc Natl Acad Sci USA 114: E8875–E8884, 2017. doi: 10.1073/pnas.1709123114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Kammers K, Cole RN, Tiengwe C, Ruczinski I. Detecting significant changes in protein abundance. EuPA Open Proteom 7: 11–19, 2015. doi: 10.1016/j.euprot.2015.02.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Katsura T, Gustafson CE, Ausiello DA, Brown D. Protein kinase A phosphorylation is involved in regulated exocytosis of aquaporin-2 in transfected LLC-PK1 cells. Am J Physiol 272: F817–F822, 1997. doi: 10.1152/ajprenal.1997.272.6.F816. [DOI] [PubMed] [Google Scholar]
- 13.Klussmann E, Maric K, Wiesner B, Beyermann M, Rosenthal W. Protein kinase A anchoring proteins are required for vasopressin-mediated translocation of aquaporin-2 into cell membranes of renal principal cells. J Biol Chem 274: 4934–4938, 1999. doi: 10.1074/jbc.274.8.4934. [DOI] [PubMed] [Google Scholar]
- 14.Limbutara K, Kelleher A, Yang CR, Raghuram V, Knepper MA. Phosphorylation changes in response to kinase inhibitor H89 in PKA-null cells. Sci Rep 9: 2814, 2019. doi: 10.1038/s41598-019-39116-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Matsumura Y, Uchida S, Rai T, Sasaki S, Marumo F. Transcriptional regulation of aquaporin-2 water channel gene by cAMP. J Am Soc Nephrol 8: 861–867, 1997. [DOI] [PubMed] [Google Scholar]
- 16.Michel JJ, Scott JD. AKAP mediated signal transduction. Annu Rev Pharmacol Toxicol 42: 235–257, 2002. doi: 10.1146/annurev.pharmtox.42.083101.135801. [DOI] [PubMed] [Google Scholar]
- 17.Navarrete-Perea J, Yu Q, Gygi SP, Paulo JA. Streamlined tandem mass tag (SL-TMT) protocol: an efficient strategy for quantitative (phospho)proteome profiling using tandem mass tag-synchronous precursor selection-MS3. J Proteome Res 17: 2226–2236, 2018. doi: 10.1021/acs.jproteome.8b00217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Nedvetsky PI, Tabor V, Tamma G, Beulshausen S, Skroblin P, Kirschner A, Mutig K, Boltzen M, Petrucci O, Vossenkämper A, Wiesner B, Bachmann S, Rosenthal W, Klussmann E. Reciprocal regulation of aquaporin-2 abundance and degradation by protein kinase A and p38-MAP kinase. J Am Soc Nephrol 21: 1645–1656, 2010. doi: 10.1681/ASN.2009111190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Nielsen S, Chou CL, Marples D, Christensen EI, Kishore BK, Knepper MA. Vasopressin increases water permeability of kidney collecting duct by inducing translocation of aquaporin-CD water channels to plasma membrane. Proc Natl Acad Sci USA 92: 1013–1017, 1995. doi: 10.1073/pnas.92.4.1013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Nishimoto G, Zelenina M, Li D, Yasui M, Aperia A, Nielsen S, Nairn AC. Arginine vasopressin stimulates phosphorylation of aquaporin-2 in rat renal tissue. Am J Physiol Renal Physiol 276: F254–F259, 1999. doi: 10.1152/ajprenal.1999.276.2.F254. [DOI] [PubMed] [Google Scholar]
- 21.Olesen ET, Fenton RA. Aquaporin-2 membrane targeting: still a conundrum. Am J Physiol Renal Physiol 312: F744–F747, 2017. doi: 10.1152/ajprenal.00010.2017. [DOI] [PubMed] [Google Scholar]
- 22.Procino G, Carmosino M, Marin O, Brunati AM, Contri A, Pinna LA, Mannucci R, Nielsen S, Kwon TH, Svelto M, Valenti G. Ser-256 phosphorylation dynamics of aquaporin 2 during maturation from the ER to the vesicular compartment in renal cells. FASEB J 17: 1886–1888, 2003. doi: 10.1096/fj.02-0870fje. [DOI] [PubMed] [Google Scholar]
- 23.Roos KP, Strait KA, Raphael KL, Blount MA, Kohan DE. Collecting duct-specific knockout of adenylyl cyclase type VI causes a urinary concentration defect in mice. Am J Physiol Renal Physiol 302: F78–F84, 2012. doi: 10.1152/ajprenal.00397.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Salhadar K, Matthews A, Raghuram V, Limbutara K, Yang CR, Datta A, Chou CL, Knepper MA. Phosphoproteomic identification of vasopressin/cAMP/PKA-dependent signaling in kidney. Mol Pharmacol mol.120.119602, 2020. doi: 10.1124/mol.120.119602. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Sandoval PC, Claxton JS, Lee JW, Saeed F, Hoffert JD, Knepper MA. Systems-level analysis reveals selective regulation of Aqp2 gene expression by vasopressin. Sci Rep 6: 34863, 2016. doi: 10.1038/srep34863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Søberg K, Skålhegg BS. The molecular basis for specificity at the level of the protein kinase A catalytic subunit. Front Endocrinol (Lausanne) 9: 538, 2018. doi: 10.3389/fendo.2018.00538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.UniProt Consortium. Nucleic Acids Res 46: 2699, 2018. doi: 10.1093/nar/gky092. [DOI] [PMC free article] [PubMed] [Google Scholar]