Abstract
Purpose
To elucidate the genetic cause of intellectual deficiency and/or congenital malformations in two parental reciprocal translocation carriers and provide appropriate strategies of assisted reproductive therapy (ART).
Materials and methods
Two similar couples having a child with global developmental delay/intellectual disability symptoms attended the Reproductive and Genetic Hospital of CITIC-Xiangya (Changsha, China) in 2017 and 2019, respectively, in order to determine the cause(s) of the conditions affecting their child and to seek ART to have a healthy baby. Both of the healthy couples were not of consanguineous marriage, denied exposure to toxicants, and had no adverse life history. This study was approved by the Institutional Ethics Committee of the Reproductive & Genetic Hospital of CITIC-Xiangya, and written informed consent was obtained from the parents. Genetic diagnoses were performed by karyotype analysis, breakpoint mapping analysis of chromosomal translocation(s), single-nucleotide polymorphism (SNP) microarray analysis, and whole-exome sequencing (WES) for the two children and different appropriate reproductive strategies were performed in the two families.
Results
Karyotype analysis revealed that both patients carried parental reciprocal translocations [46,XY,t(7;16)(p13;q24)pat and 46,XY,t(13;17)(q12.3;p11.2)pat, respectively]. Follow-up breakpoint mapping analysis showed no interruption of associated genes, and SNP microarray analysis identified no significant copy number variations (CNVs) in the two patients. Moreover, WES results revealed that patients 1 and 2 harbored candidate compound heterozygous mutations of MCOLN1 [c.195G>C (p.K65N) and c.1061G>A (p.W354*)] and MCPH1 [c.877A>G (p.S293G) and c.1869_1870delAT (p.C624*)], respectively, that were inherited from their parents and not previously reported. Furthermore, the parents of patient 1 obtained 10 embryos during ART cycle, and an embryo of normal karyotype and non-carrier of observed MCOLN1 mutations according to preimplantation genetic testing for structural rearrangement and monogenic defect was successfully transferred, resulting in the birth of a healthy boy. The parents of patient 2 chose to undergo ART with donor sperm to reduce the risk of recurrence.
Conclusions
Systematic genetic diagnosis of two carriers of inherited chromosomal translocations accompanied by clinical phenotypes revealed their cause of disease, which was critical for genetic counseling and further ART for these families.
Electronic supplementary material
The online version of this article (10.1007/s10815-020-01986-1) contains supplementary material, which is available to authorized users.
Keywords: Reciprocal translocation, Developmental delay, Intellectual disability, Chromosome microdissection, Whole-exome sequencing, Assisted reproductive therapy
Introduction
With the development of assisted reproductive technology (ART), particularly preimplantation genetic testing (PGT) identifying possible genetic defects (chromosomal abnormality or monogenetic disease), the risk of an affected child is reducing. The accurate identification of potential genetic issues is a critical aspect of its use for families seeking assisted reproduction.
Reciprocal translocation is among the most common structural chromosomal rearrangements, with an incidence of approximately 0.1% in newborns [1, 2]. Reciprocal translocation involves breakage of two chromosomes and exchange of their distal chromosomal segments, resulting in two derivative chromosomes. Most balanced reciprocal translocation carriers are phenotypically normal, whereas due to quadrivalent formations during meiosis, they are at an increased fertility risk, including recurrent miscarriage, infertility, or having children with a variety of symptoms associated with genomic imbalances [3–5], while minority of offspring with parental balanced reciprocal translocation exhibited abnormal symptoms, which creates a challenge for genetic counseling. The cause of the phenotype(s) is critical not only for personal treatment but also for assessment of reproductive risk in the parents and in making ART strategy.
Global developmental delay (GDD) is defined as a significant delay in two or more developmental domains of gross/fine motor, cognitive, social/personal, or speech/language activities associated with daily life, with this mainly affecting ~ 1% to 3% of the population of children aged ≤ 5 years. Intellectual disability (ID) that involves problems with mental retardation or learning disability affects ~ 2% of children and young adults [6, 7], and the concurrence of GDD and ID is common in the pediatric population and introduces mental and economic burdens to their families [8, 9]. GDD/ID is a clinically and genetically heterogeneous group of diseases, with patients usually presenting a variety of symptoms, including combined congenital malformations, autism spectrum disorders, and/or seizure disorders [10, 11] that are difficult to clinically distinguish and a challenge for genetic counseling. Genetic causes are the primary factor related to these diseases and include chromosomal abnormalities, copy number variations (CNVs), and gene mutations [6, 12–14].
In this study, we performed systematic genetic diagnoses of two carriers of paternal reciprocal translocation and those exhibiting abnormal phenotypes in order to determine the underlying genetic causes, and corresponding ART strategies were made in the two families.
Materials and methods
Patients
Two similar couples having a child with GDD/ID symptoms attended the Reproductive and Genetic Hospital of CITIC-Xiangya (Changsha, China) in 2017 and 2019, respectively, in order to determine the cause(s) of the conditions affecting their child and to seek ART to have a healthy baby. Both of the healthy couples were not of consanguineous marriage, denied exposure to toxicants, and had no adverse life history. This study was approved by the Institutional Ethics Committee of the Reproductive & Genetic Hospital of CITIC-Xiangya, and written informed consent was obtained from the parents.
Karyotyping and fluorescence in situ hybridization analysis
Peripheral blood samples from the patients and their parents were collected, and lymphocytes were cultured for G-banding and fluorescence in situ hybridization (FISH) analyses. Metaphases were analyzed by GTG banding using trypsin and Giemsa staining, and karyotyping analysis was performed using standard methods according to the International System for Human Cytogenetic Nomenclature (2016, ISCN). FISH analysis was performed using commercially available sub-telomere-specific probes (Subtel 13q/Subtel 17p/CEP17; Abbott Laboratories, Chicago, IL, USA) according to manufacturer instructions.
Chromosome microdissection sequencing (MicroSeq)
Chromosome microdissection and follow-up next-generation sequencing (NGS) of junction-fragment DNA and precise mapping of the breakpoints were performed as previously described [15]. Briefly, eight copies of the region covering the breakpoints were dissected from the G-banding-metaphase spread of patient peripheral blood samples using glass needles. The dissected DNA fragments were amplified using the WGA4 kit (Sigma-Aldrich, St. Louis, MO, USA) according to manufacturer instructions. Amplified DNA products were evaluated by 2% agarose gel electrophoresis, followed by NGS using the Ion Xpress library kit (Life Technologies, Rockville, MD, USA) according to manufacturer instructions. The sequencing library for the amplified products was diluted to 2.5 pg/μL and amplified using the Ion PGM Template OT2 200 kit (Thermo Fisher Scientific, Waltham, MA, USA) for 5.5 h, with the Ion PGM sequencing 200 kit (v.2.0; Thermo Fisher Scientific) employed for sequencing. Sequence data were cleaned by removing the primer sequences and aligned to the reference genome (hg19) using Integrative Genomics Viewer (http://software.broadinstitute.org/software/igv/).
CNV analysis
SNP microarrays were used to analyze whole-genome CNVs. Genomic DNA was extracted from peripheral blood from the two patients using a QIAamp DNA midi kit (QIAGEN, Hilden, Germany) and used for testing according to the instructions of the CytoScan750K reagent kit (Affymetrix, Santa Clara, CA, USA). Chromosome Analysis Suite (Affymetrix) was used for image scanning and data analysis, and all CNVs were compared using the University of California–Santa Cruz Genome Browser (https://genome.ucsc.edu/; Feb. 2009; Human: GRCh37/hg19) and the DECIPHER database (https://decipher.sanger.ac.uk/).
Whole-exome sequencing
Genomic DNA extraction and subsequent whole-exome sequencing (WES) and data analyses for the two affected patients were performed by the Beijing Genome Institute (Shenzhen, China), as described previously [16]. All variants were classified into five categories—“benign,” “likely benign,” “uncertain significance,” “likely pathogenic,” and “pathogenic”—according to the American College of Medical Genetics and Genomics (ACMG) standards and guidelines for the interpretation of variations [17]. And candidate mutations were confirmed by Sanger sequencing of PCR-amplified products using specific primers.
Preimplantation genetic testing procedure
Different protocols for ovulation stimulation were administered to the females of the two couples, and subsequent oocyte retrieval, fertilization, culturing of the fertilized oocytes, and embryo scoring were described in previous reports [18]. Five to eight trophectoderm cells were obtained by laser-assisted blastocyst biopsy on the morning of day 6 after fertilization. Whole-genome amplification of the embryo biopsy samples was performed using a single-cell PicoPLEX WGA kit (Rubicon Genomics, Ann Arbor, MI, USA), and the products were analyzed by NGS for whole-genome copy numbers to identify balanced or unbalanced embryos [19]. For family 1, the informative SNPs (Table S1) flanking the breakpoint were amplified via PCR and PGT for MCOLN1 [c.195G>C (p.K65N) and c.1061G>A (p.W354*)] with short tandem repeat analysis performed on the balanced embryos.
Results
Clinical features
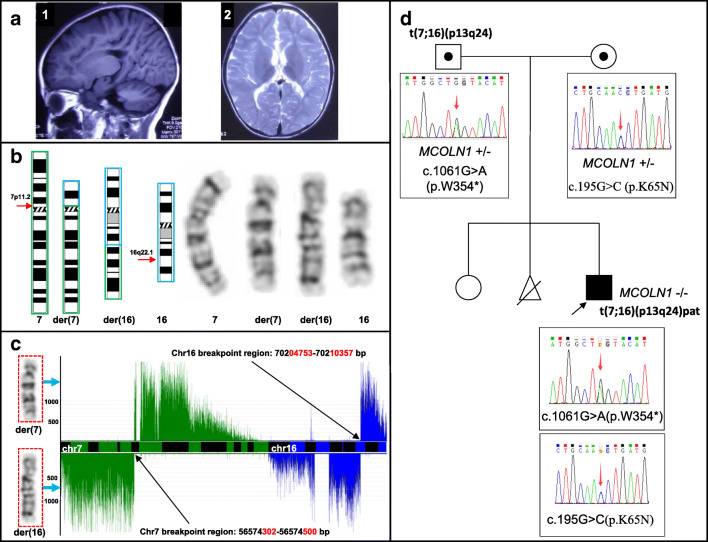
In family 1, the proband (patient 1) was a 7-year-old boy born from naturally conceived pregnancy at 36 weeks (birth weight: 4 kg). The main clinical phenotypes were cerebral palsy, mental retardation, developmental delay, mild corneal clouding, photophobia, and strabismus in the left eye. At birth, there were no obvious abnormalities. At 3 months, the patient could raise his head but could not turn over independently at 6 months. At 1 year old, the patient was diagnosed with developmental delay, including deficiencies in major and motor movements. The patient was able to sit at 1.5 years old and stand on tiptoes and exhibits a scissors gait with support at age 3 but remained unable to stand on his own. At 5 years old, the patient showed hip and knee flexion, as well as tension in multiple muscles, including bilateral adductors, hamstrings, and the Achilles tendons. At this time, the patient was clinically diagnosed with spastic cerebral palsy. At 7 years old, magnetic resonance imaging (MRI) revealed a decrease in white matter in the bilateral cerebral hemisphere, and cortical atrophy in the parietal lobe, hypoplastic corpus callosum, and broadening of the cerebral ventricles (Fig. 1A-1 and A-2). The mother had a history of adverse pregnancy involving an induced labor following fetal death in the uterus due to the umbilical cord around the neck.
Fig. 1.
Results of MRI and genetic analysis of patient 1. (A) Brain MRI revealed a decrease in white matter in the bilateral cerebral hemisphere, cortical atrophy in the parietal lobe, hypoplastic corpus callosum, broadening of the adjacent cerebral ventricles, and dull edges of the hypoplastic corpus callosum (A-1 and A-2). (B) Patient 1 carried 46,XY,t(7;16)(p13;q24). Red arrows indicate the translocation breakpoints. (C) MicroSeq analysis showed the breakpoint of Chr7 located in the region 56,574,302 bp to 56,574,500 bp and another one of Chr16 located in the region 70,204,753 bp to 70,210,357 bp. (D) WES analysis revealing that patient 1 carried compound heterozygous mutations of MCOLN1 [c.195G>C (p.K65N) and c.1061G>A (p.W354*)], which was confirmed by Sanger sequencing
In family 2, the proband (patient 2) was a 2-year-old boy born at 32 months (premature) and with a low birth weight (1.85 kg). Unlike patient 1, the boy was an offspring conceived by in vitro fertilization (IVF)/PGT using FISH. At 3 months, the patient had a fever and frequent convulsions accompanied by high levels of muscle tension. The patient was diagnosed with epilepsy according to an abnormal electroencephalogram and was administered antiepileptic drugs. At 2 years of age, he was able to turn over but could not raise his head, sit, or stand. Additionally, the patient exhibited severe visual impairment almost to the point of blindness but was able to hear. MRI results indicated very small brain tissue and revealed an occipital cistern arachnoid cyst characterized by softening in large areas of the brain. The parents had no corresponding clinical manifestations.
Results of cytogenetic analysis
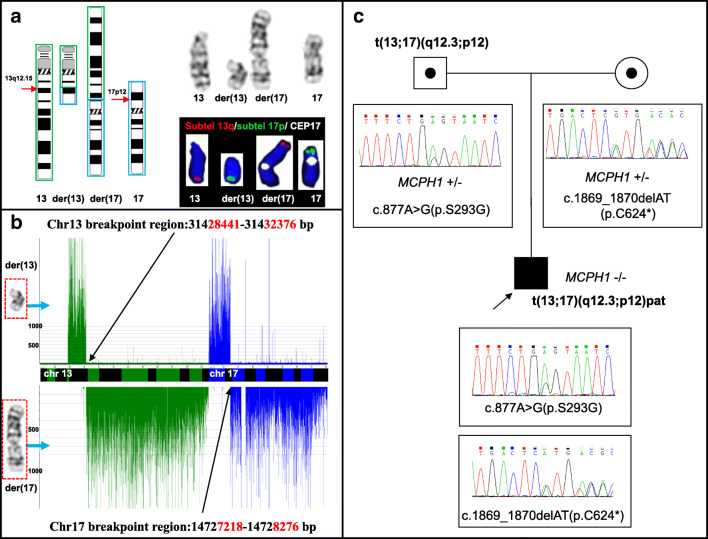
Karyotype G-banding coupled with FISH analysis indicated that the two patients inherited a paternal balanced translocation, with patient 1 carrying 46,XY,t(7;16)(p13;q24)pat (Fig. 1B), and patient 2 carrying 46,XY,t(13;17)(q12.3;p11.2)pat (Fig. 2A).
Fig. 2.
Results of genetic analysis of patient 2. (A) G-banding indicated that the translocation involved Chr13 and Chr17. Red arrows indicate the translocation breakpoints. (B) MicroSeq results indicating the breakpoint of Chr13 located in the region 31,428,441 bp to 31,432,376 bp and the other breakpoint of Chr17 located in the region 14,727,218 bp to 14,728,276 bp. (C) WES revealed that patient 2 carried compound heterozygous mutations of MCPH1 [c.877A>G (p.S293G) and c.1869_1870delAT (p.C624*)], which was confirmed by Sanger sequencing
Results of breakpoint mapping
The four junction fragments of derivative chromosomes in the two patients were successfully microdissected and amplified. The sequencing library for the amplified products included 128,665 reads to 1,003,450 reads, with an average coverage depth of the junction fragments ranging from 9.637 and 35.281 (Table S2). The der(16) breakpoint in patient 1 was located in the fourth intron of CLEC18C, whereas the other three breakpoints did not affect any known gene (Fig. 1C and Fig. 2B).
Results of CNV and WES analyses
The results of SNP microarray analysis indicated the presence of no pathological CNVs in genomic DNA of the two patients. According to WES, the average coverage depths of patients 1 and 2 were 270 ± 190- and 264 ± 191-fold, respectively, with the coverage areas > 20× at 99.6% and 99.3%, respectively. Following filtering and pedigree analysis, WES revealed a candidate pathogenic gene [MCOLN1 (NM_020533)] with novel compound heterozygous mutations [c.195G>C (p.K65N) (uncertain significance) and c.1061G>A (p.W354*) (likely pathogenic)] in patient 1. Sanger sequencing in-family analysis suggested that the two mutations were inherited from his mother and father (Fig. 1D). For patient 2, we identified a candidate causative gene [MCPH1 (NM_024596)] harboring a maternal mutation [c.877A>G (p.S293G) (uncertain significance)] and paternal mutation [c.1869_1870delAT (p.C624*) (likely pathogenic)] (Fig. 2C). These four mutations were previously unreported and all the allele frequencies were < 0.1% according to public databases.
PGT outcomes
The female of family 1 underwent one cycle of PGT, during which 17 oocytes were retrieved, and 15 oocytes were fertilized through intracytoplasmic sperm injection, resulting in 10 blastocysts obtained and eligible for biopsy (Table 1). The results of PGT revealed seven were unbalanced embryos and three were balanced embryos, of which two diagnosed as carriers of the 46,XY,t(7;16)(p13;q24) translocation and the MCOLN1 c.1061G>A mutation. Fortunately, the other embryo carried a normal karyotype and without observed MCOLN1 mutations (Table 1). This embryo was ultimately used for transfer, with prenatal diagnosis confirming PGT results, and a healthy baby boy was born.
Table 1.
The summary results of PGT in family 1
| Family | Father | Mother | TE biopsied cycle(s) | Embryo no. | PGT-SR | PGT-M | Embryo grade | Pregnancy outcome |
|---|---|---|---|---|---|---|---|---|
| Family 1 | 46,XY,t(7;16)(p13q24) /MCOLN1 c.1061G>A (p.W354*) | MCOLN1 c.195G>C (p.K65N) | 1 | 1 | 46,XY,t(7;16)(p13;q24)pat | c.1061G>A | 5AB | |
| 2 | 46,XY,dup(7)(p22.3-p11.2)(56.63Mb),del(16)(q22.2-q24.3)(18.37Mb) | ND | 5AB | |||||
| 3 | 46,XY,t(7;16)(p13;q24)pat | c.1061G>A | 5AB | |||||
| 4 | 46,XY,del(7)(p22.3-p11.2)(56.63Mb),dup(16)(q22.2-q24.3)(18.37Mb) | ND | 5AB | |||||
| 5 | 46,XY | Normal | 6AB | Deliver a healthy baby,46,XY | ||||
| 6 | 46,XX,dup(7)(q11.21-q36.3)(94.57Mb),del(16)(p13.3-q22.1)(67.81Mb) | ND | 6BB | |||||
| 7 | 46,XX,dup(7)(p22.3-p11.2)(56.63Mb),del(16)(q22.2-q24.3)(18.37Mb) | ND | 5BB | |||||
| 8 | 45,XX,-22 | ND | 5BB | |||||
| 9 | 46,XX,dup(7)(p22.3-p11.2)(56.63Mb),del(16)(q22.2-q24.3)(18.37Mb) | ND | 5BB | |||||
| 10 | 45,XX,-10,del(7)(q11.21-q36.3)(95.38Mb),dup(16)(p13.11-q22.1)(54.92Mb) | ND | 4BB |
PGT-SR, PGT for structural rearrangement; PGT-M, PGT for monogenic disease; ND, not detected
The couples of family 2 chose to undergo a routine IVF procedure with donor sperm because the defects of MCPH1 cannot explain all the clinical symptoms of patient 2. Yet, by this time, they are going through the process of IVF.
Discussion
Carriers of parental reciprocal translocation present congenital malformations, which makes genetic counseling and subsequent IVF strategies challenging. Most carriers of balanced reciprocal translocations exhibit no phenotypic effects; however, abnormal phenotypes associated with multiple congenital abnormalities in some cases [20] have been reported due to interruption of associated functional genes as a consequence of chromosomal breakpoints [21–25] or disturbances related to parental imprinting [26, 27]. And some studies revealed a correlation between diseases and genetic factors through breakpoint mapping [28–30]. Thus, here, we used a more cost-effective chromosomal microdissection approach combined with NGS technology to precisely locate chromosomal breakpoints. We found that the der(16) breakpoint in patient 1 interrupted the CLEC18C gene, whereas this gene was unrelated to the symptoms presented. Beyond that, we did not found interruptions of any related functional genes, and there was no loss or gain of genetic material in the breakpoint regions identified in the two patients (Figs. 1C and 2B).
Given that monogenic disorders are responsible for ~ 10% of GDD/ID diagnoses [31], we performed WES to investigate potential pathogenic mutations contributing to the patient phenotypes. We identified compound heterozygous mutations [c.195G>C (p.K65N)/c.1061G>A (p.W354*)] in MCOLN1 of patient 1 and [c.877A>G (p.S293G)/c.1869_1870delAT (p.C624*)] MCPH1 of patient 2. MCOLN1 contains 14 exons and encodes mucolipin-1, which comprises 580 amino acid residues and is a member of the transient receptor potential cation channel family [32]. In 2000, Bargal et al. [33] first identified MCOLN1 mutations causative of mucolipidosis IV (MLIV), a rare autosomal recessive disorder in which > 80% of diagnosed patients are Ashkenazi Jews. MLIV is mainly characterized by severe psychomotor retardation, corneal clouding, retinal dystrophy, optic atrophy, muscle tonus abnormalities, coarse facies, cognitive impairment, spasticity, or iron deficiency–related anemia [34–36]. In the present study, patient 1 mainly presented mental retardation, developmental retardation, corneal clouding, photophobia, and strabismus. Here, two novel MCOLN1 mutations were identified, and the c.195G>C (p.K65N) mutation was located in a highly conserved region between areas encoding the phosphoinositides domain (42–62 aa) [37] and the extracellular/lumenal pore loop domain (107–121 aa) [38]. Additionally, c.1061G>A (p.W354*) was a nonsense mutation resulting in a premature stop codon in exon 9 and resulting in a truncated protein containing only 354 amino acids and lacking the di-leucine internalization motif that mediates AP2-complex-dependent internalization [39]. These two mutations in MCOLN1 could explain the phenotype presented by patient 1. These two mutations were then used for follow-up PGT for monogenetic disease. Fortunately, the family obtained three embryos eligible for transfer, with the embryo showing a normal karyotype and no pathogenic mutation of MCOLN1, resulting in birth of a healthy baby boy (Table 1).
In patient 2, two candidate mutations in MCPH1 were identified, a gene associated with autosomal recessive primary microcephaly (MCPH). The MCPH1 gene plays an important role in mitotic spindle and centrosome formation, DNA damage repair, and transcription activation essential for normal brain development and maintenance of genome integrity [40–43]. Phenotypes associated with MCPH1 mutations include MCPH, autism spectrum disorder, and mental retardation [44–47]. Here, the candidate mutations of MCPH1 might explain the neurological abnormalities and microcephaly observed of patient 2 but not GDD. Due to the indeterminate genetic cause of the patient 2 phenotype, finally the parents chose to undergo IVF with donor sperm in order to reduce the risk of giving birth to a similarly afflicted child.
Here, we performed systematic genetic diagnoses of two children carrying balanced chromosomal translocation and presenting multiple phenotypes. Then, different IVF strategies were performed in the two families according to the different results of the genetic diagnoses, which reflected the significance of diagnosing genetic causes in the assisted reproductive process.
Electronic supplementary material
(DOCX 17 kb).
Authors’ contributions
Dehua Cheng and Shimin Yuan conducted the genetic studies, drafted the initial manuscript, and wrote the manuscript; Liang Hu participated in sequence alignment and performed the initial analyses; Keli Luo and Fei Gong participated in data collection; Duo Yi and Changfu Lu participated in data analysis and FISH experiment; Guangxiu Lu and Ge Lin conceived of the study; Yue-Qiu Tan conceived of the study, helped draft the initial manuscript, and wrote the manuscript. All authors read and approved the final manuscript.
Funding
This work was supported by grants from the National Key Research & Developmental Program of China (2018YFC1004900), the National Natural Science Foundation of China (81771645 and 81971447), Hunan Provincial Grant for Innovative Province Construction (2019SK4012), and Research Grant of CITIC-Xiangya (YNXM-201916).
Compliance with ethical standards
This study was approved by the Institutional Ethics Committee of the Reproductive & Genetic Hospital of CITIC-Xiangya, and written informed consent was obtained from the parents.
Conflict of interest
The authors declare that they have no conflict of interest.
Footnotes
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Dehua Cheng and Shimin Yuan contributed equally to this work.
References
- 1.Hamerton JL, Canning N, Ray M, Smith S. A cytogenetic survey of 14,069 newborn infants. I. Incidence of chromosome abnormalities. Clin Genet. 1975;8(4):223–243. doi: 10.1111/j.1399-0004.1975.tb01498.x. [DOI] [PubMed] [Google Scholar]
- 2.Maeda T, Ohno M, Matsunobu A, Yoshihara K, Yabe N. A cytogenetic survey of 14,835 consecutive liveborns. Jinrui Idengaku Zasshi. 1991;36(1):117–129. doi: 10.1007/BF01876812. [DOI] [PubMed] [Google Scholar]
- 3.Pasinska M, Lazarczyk E, Julga K, Bartnik-Glaska M, Nowakowska B, Haus O. Multiple occurrence of psychomotor retardation and recurrent miscarriages in a family with a submicroscopic reciprocal translocation t(7;17)(p22;p13.2) BMC Med Genet. 2018;11(1):69. doi: 10.1186/s12920-018-0384-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Chen JK, Liu P, Hu LQ, Xie Q, Huang QF, Liu HL. A foetus with 18p11.32-q21.2 duplication and Xp22.33-p11.1 deletion derived from a maternal reciprocal translocation t(X;18)(q13;q21.3) Mol Cytogenet. 2018;11:37. doi: 10.1186/s13039-018-0381-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Paththinige CS, Sirisena ND, Kariyawasam U, Ediriweera RC, Kruszka P, Muenke M, et al. A child with multiple congenital anomalies due to partial trisomy 7q22.1 --> qter resulting from a maternally inherited balanced translocation: a case report and review of literature. BMC Med Genet. 2018;11(1):44. doi: 10.1186/s12920-018-0366-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Larizza L, Finelli P. Developmental disorders with intellectual disability driven by chromatin dysregulation: clinical overlaps and molecular mechanisms. Clin Genet. 2019;95(2):231–240. doi: 10.1111/cge.13365. [DOI] [PubMed] [Google Scholar]
- 7.Ropers HH. Genetics of early onset cognitive impairment. Annu Rev Genomics Hum Genet. 2010;11:161–187. doi: 10.1146/annurev-genom-082509-141640. [DOI] [PubMed] [Google Scholar]
- 8.Silibello G, Vizziello P, Gallucci M, Selicorni A, Milani D, Ajmone PF, Rigamonti C, de Stefano S, Bedeschi MF, Lalatta F. Daily life changes and adaptations investigated in 154 families with a child suffering from a rare disability at a public centre for rare diseases in Northern Italy. Ital J Pediatr. 2016;42(1):76. doi: 10.1186/s13052-016-0285-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.McKenzie J, McConkey R. Caring for adults with intellectual disability: the perspectives of family carers in South Africa. J Appl Res Intellect Disabil. 2016;29(6):531–541. doi: 10.1111/jar.12209. [DOI] [PubMed] [Google Scholar]
- 10.de Ligt J, Willemsen MH, van Bon BW, Kleefstra T, Yntema HG, Kroes T, et al. Diagnostic exome sequencing in persons with severe intellectual disability. N Engl J Med. 2012;367(20):1921–1929. doi: 10.1056/NEJMoa1206524. [DOI] [PubMed] [Google Scholar]
- 11.Topper S, Ober C, Das S. Exome sequencing and the genetics of intellectual disability. Clin Genet. 2011;80(2):117–126. doi: 10.1111/j.1399-0004.2011.01720.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.van Bokhoven H. Genetic and epigenetic networks in intellectual disabilities. Annu Rev Genet. 2011;45:81–104. doi: 10.1146/annurev-genet-110410-132512. [DOI] [PubMed] [Google Scholar]
- 13.Kessi M, Xiong J, Wu L, Yang L, He F, Chen C, Pang N, Duan H, Zhang W, Arafat A, Yin F, Peng J. Rare copy number variations and predictors in children with intellectual disability and epilepsy. Front Neurol. 2018;9:947. doi: 10.3389/fneur.2018.00947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Han JY, Jang JH, Park J, Lee IG. Targeted next-generation sequencing of Korean patients with developmental delay and/or intellectual disability. Front Pediatr. 2018;6:391. doi: 10.3389/fped.2018.00391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Hu L, Cheng D, Gong F, Lu C, Tan Y, Luo K, Wu X, He W, Xie P, Feng T, Yang K, Lu G, Lin G. Reciprocal translocation carrier diagnosis in preimplantation human embryos. EBioMedicine. 2016;14:139–147. doi: 10.1016/j.ebiom.2016.11.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Tan YQ, Tu C, Meng L, Yuan S, Sjaarda C, Luo A, du J, Li W, Gong F, Zhong C, Deng HX, Lu G, Liang P, Lin G. Loss-of-function mutations in TDRD7 lead to a rare novel syndrome combining congenital cataract and nonobstructive azoospermia in humans. Genet Med. 2019;21(5):1209–1217. doi: 10.1038/gim.2017.130. [DOI] [PubMed] [Google Scholar]
- 17.Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17(5):405–424. doi: 10.1038/gim.2015.30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Hu L, Wei Y, Luo K, Xie P, Gong F, Xiong B, Tan Y, Lu G, Lin G. Clinical outcomes in carriers of complex chromosomal rearrangements: a retrospective analysis of comprehensive chromosome screening results in seven cases. Fertil Steril. 2018;109(3):486–492. doi: 10.1016/j.fertnstert.2017.11.021. [DOI] [PubMed] [Google Scholar]
- 19.Tan Y, Yin X, Zhang S, Jiang H, Tan K, Li J, Xiong B, Gong F, Zhang C, Pan X, Chen F, Chen S, Gong C, Lu C, Luo K, Gu Y, Zhang X, Wang W, Xu X, Vajta G, Bolund L, Yang H, Lu G, du Y, Lin G. Clinical outcome of preimplantation genetic diagnosis and screening using next generation sequencing. Gigascience. 2014;3(1):30. doi: 10.1186/2047-217X-3-30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Schluth-Bolard C, Labalme A, Cordier MP, Till M, Nadeau G, Tevissen H, Lesca G, Boutry-Kryza N, Rossignol S, Rocas D, Dubruc E, Edery P, Sanlaville D. Breakpoint mapping by next generation sequencing reveals causative gene disruption in patients carrying apparently balanced chromosome rearrangements with intellectual deficiency and/or congenital malformations. J Med Genet. 2013;50(3):144–150. doi: 10.1136/jmedgenet-2012-101351. [DOI] [PubMed] [Google Scholar]
- 21.Moyses-Oliveira M, Di-Battista A, Zamariolli M, Meloni VA, Bragagnolo S, Christofolini DM, et al. Breakpoint mapping at nucleotide resolution in X-autosome balanced translocations associated with clinical phenotypes. Eur J Hum Genet. 2019;27(5):760–771. doi: 10.1038/s41431-019-0341-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Yauy K, Schneider A, Ng BL, Gaillard JB, Sati S, Coubes C, Wells C, Tournaire M, Guignard T, Bouret P, Geneviève D, Puechberty J, Pellestor F, Gatinois V. Disruption of chromatin organisation causes MEF2C gene overexpression in intellectual disability: a case report. BMC Med Genet. 2019;12(1):116. doi: 10.1186/s12920-019-0558-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Aristidou C, Theodosiou A, Bak M, Mehrjouy MM, Constantinou E, Alexandrou A, Papaevripidou I, Christophidou-Anastasiadou V, Skordis N, Kitsiou-Tzeli S, Tommerup N, Sismani C. Position effect, cryptic complexity, and direct gene disruption as disease mechanisms in de novo apparently balanced translocation cases. PLoS One. 2018;13(10):e0205298. doi: 10.1371/journal.pone.0205298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Dutta UR, Rao SN, Pidugu VK, Vineeth VS, Bhattacherjee A, Bhowmik AD, et al. Breakpoint mapping of a novel de novo translocation t(X;20)(q11.1;p13) by positional cloning and long read sequencing. Genomics. 2019;111(5):1108–1114. doi: 10.1016/j.ygeno.2018.07.005. [DOI] [PubMed] [Google Scholar]
- 25.Schluth-Bolard C, Diguet F, Chatron N, Rollat-Farnier PA, Bardel C, Afenjar A, Amblard F, Amiel J, Blesson S, Callier P, Capri Y, Collignon P, Cordier MP, Coubes C, Demeer B, Chaussenot A, Demurger F, Devillard F, Doco-Fenzy M, Dupont C, Dupont JM, Dupuis-Girod S, Faivre L, Gilbert-Dussardier B, Guerrot AM, Houlier M, Isidor B, Jaillard S, Joly-Hélas G, Kremer V, Lacombe D, le Caignec C, Lebbar A, Lebrun M, Lesca G, Lespinasse J, Levy J, Malan V, Mathieu-Dramard M, Masson J, Masurel-Paulet A, Mignot C, Missirian C, Morice-Picard F, Moutton S, Nadeau G, Pebrel-Richard C, Odent S, Paquis-Flucklinger V, Pasquier L, Philip N, Plutino M, Pons L, Portnoï MF, Prieur F, Puechberty J, Putoux A, Rio M, Rooryck-Thambo C, Rossi M, Sarret C, Satre V, Siffroi JP, Till M, Touraine R, Toutain A, Toutain J, Valence S, Verloes A, Whalen S, Edery P, Tabet AC, Sanlaville D. Whole genome paired-end sequencing elucidates functional and phenotypic consequences of balanced chromosomal rearrangement in patients with developmental disorders. J Med Genet. 2019;56(8):526–535. doi: 10.1136/jmedgenet-2018-105778. [DOI] [PubMed] [Google Scholar]
- 26.Dupont JM, Cuisset L, Cartigny M, Le Tessier D, Vasseur C, Rabineau D, et al. Familial reciprocal translocation t(7;16) associated with maternal uniparental disomy 7 in a Silver-Russell patient. Am J Med Genet. 2002;111(4):405–408. doi: 10.1002/ajmg.10570. [DOI] [PubMed] [Google Scholar]
- 27.Conroy JM, Grebe TA, Becker LA, Tsuchiya K, Nicholls RD, Buiting K, Horsthemke B, Cassidy SB, Schwartz S. Balanced translocation 46,XY,t(2;15)(q37.2;q11.2) associated with atypical Prader-Willi syndrome. Am J Hum Genet. 1997;61(2):388–394. doi: 10.1086/514852. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Fukushi D, Yamada K, Suzuki K, Inaba M, Nomura N, Suzuki Y, Katoh K, Mizuno S, Wakamatsu N. Clinical and genetic characterization of a patient with SOX5 haploinsufficiency caused by a de novo balanced reciprocal translocation. Gene. 2018;655:65–70. doi: 10.1016/j.gene.2018.02.049. [DOI] [PubMed] [Google Scholar]
- 29.Cosemans N, Vandenhove L, Maljaars J, Van Esch H, Devriendt K, Baldwin A, et al. ZNF462 and KLF12 are disrupted by a de novo translocation in a patient with syndromic intellectual disability and autism spectrum disorder. Eur J Med Genet. 2018;61(7):376–383. doi: 10.1016/j.ejmg.2018.02.002. [DOI] [PubMed] [Google Scholar]
- 30.Xu XJ, Lv F, Liu Y, Wang JY, Song YW, Asan, et al. A cryptic balanced translocation involving COL1A2 gene disruption cause a rare type of osteogenesis imperfecta. Clin Chim Acta. 2016;460:33–39. doi: 10.1016/j.cca.2016.06.011. [DOI] [PubMed] [Google Scholar]
- 31.Miclea D, Peca L, Cuzmici Z, Pop IV. Genetic testing in patients with global developmental delay / intellectual disabilities. A review. Clujul Med. 2015;88(3):288–292. doi: 10.15386/cjmed-461. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Sun M, Goldin E, Stahl S, Falardeau JL, Kennedy JC, Acierno JS, Jr, et al. Mucolipidosis type IV is caused by mutations in a gene encoding a novel transient receptor potential channel. Hum Mol Genet. 2000;9(17):2471–2478. doi: 10.1093/hmg/9.17.2471. [DOI] [PubMed] [Google Scholar]
- 33.Bargal R, Avidan N, Ben-Asher E, Olender Z, Zeigler M, Frumkin A, Raas-Rothschild A, Glusman G, Lancet D, Bach G. Identification of the gene causing mucolipidosis type IV. Nat Genet. 2000;26(1):118–123. doi: 10.1038/79095. [DOI] [PubMed] [Google Scholar]
- 34.Amir N, Zlotogora J, Bach G. Mucolipidosis type IV: clinical spectrum and natural history. Pediatrics. 1987;79(6):953–959. [PubMed] [Google Scholar]
- 35.Bach G. Mucolipidosis type IV. Mol Genet Metab. 2001;73(3):197–203. doi: 10.1006/mgme.2001.3195. [DOI] [PubMed] [Google Scholar]
- 36.Altarescu G, Sun M, Moore DF, Smith JA, Wiggs EA, Solomon BI, Patronas NJ, Frei KP, Gupta S, Kaneski CR, Quarrell OW, Slaugenhaupt SA, Goldin E, Schiffmann R. The neurogenetics of mucolipidosis type IV. Neurology. 2002;59(3):306–313. doi: 10.1212/wnl.59.3.306. [DOI] [PubMed] [Google Scholar]
- 37.Zhang X, Li X, Xu H. Phosphoinositide isoforms determine compartment-specific ion channel activity. Proc Natl Acad Sci U S A. 2012;109(28):11384–11389. doi: 10.1073/pnas.1202194109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Li M, Zhang WK, Benvin NM, Zhou X, Su D, Li H, Wang S, Michailidis IE, Tong L, Li X, Yang J. Structural basis of dual Ca(2+)/pH regulation of the endolysosomal TRPML1 channel. Nat Struct Mol Biol. 2017;24(3):205–213. doi: 10.1038/nsmb.3362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Vergarajauregui S, Puertollano R. Two di-leucine motifs regulate trafficking of mucolipin-1 to lysosomes. Traffic. 2006;7(3):337–353. doi: 10.1111/j.1600-0854.2006.00387.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Gruber R, Zhou Z, Sukchev M, Joerss T, Frappart PO, Wang ZQ. MCPH1 regulates the neuroprogenitor division mode by coupling the centrosomal cycle with mitotic entry through the Chk1-Cdc25 pathway. Nat Cell Biol. 2011;13(11):1325–1334. doi: 10.1038/ncb2342. [DOI] [PubMed] [Google Scholar]
- 41.Zhou ZW, Tapias A, Bruhn C, Gruber R, Sukchev M, Wang ZQ. DNA damage response in microcephaly development of MCPH1 mouse model. DNA Repair (Amst) 2013;12(8):645–655. doi: 10.1016/j.dnarep.2013.04.017. [DOI] [PubMed] [Google Scholar]
- 42.Yang SZ, Lin FT, Lin WC. MCPH1/BRIT1 cooperates with E2F1 in the activation of checkpoint, DNA repair and apoptosis. EMBO Rep. 2008;9(9):907–915. doi: 10.1038/embor.2008.128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Liang Y, Gao H, Lin SY, Peng G, Huang X, Zhang P, Goss JA, Brunicardi FC, Multani AS, Chang S, Li K. BRIT1/MCPH1 is essential for mitotic and meiotic recombination DNA repair and maintaining genomic stability in mice. PLoS Genet. 2010;6(1):e1000826. doi: 10.1371/journal.pgen.1000826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Trimborn M, Richter R, Sternberg N, Gavvovidis I, Schindler D, Jackson AP, Prott EC, Sperling K, Gillessen-Kaesbach G, Neitzel H. The first missense alteration in the MCPH1 gene causes autosomal recessive microcephaly with an extremely mild cellular and clinical phenotype. Hum Mutat. 2005;26(5):496. doi: 10.1002/humu.9382. [DOI] [PubMed] [Google Scholar]
- 45.Perche O, Menuet A, Marcos M, Liu L, Paris A, Utami KH, et al. Combined deletion of two Condensin II system genes (NCAPG2 and MCPH1) in a case of severe microcephaly and mental deficiency. Eur J Med Genet. 2013;56(11):635–641. doi: 10.1016/j.ejmg.2013.07.007. [DOI] [PubMed] [Google Scholar]
- 46.Wang JK, Li Y, Su B. A common SNP of MCPH1 is associated with cranial volume variation in Chinese population. Hum Mol Genet. 2008;17(9):1329–1335. doi: 10.1093/hmg/ddn021. [DOI] [PubMed] [Google Scholar]
- 47.Farooq M, Baig S, Tommerup N, Kjaer KW. Craniosynostosis-microcephaly with chromosomal breakage and other abnormalities is caused by a truncating MCPH1 mutation and is allelic to premature chromosomal condensation syndrome and primary autosomal recessive microcephaly type 1. Am J Med Genet A. 2010;152A(2):495–497. doi: 10.1002/ajmg.a.33234. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
(DOCX 17 kb).