Abstract
The current pandemic of the 2019 novel coronavirus (COVID-19) caused by SARS-CoV-2 (severe acute respiratory syndrome coronavirus-2) has raised significant public health concern. Rapid, affordable, and accurate diagnostics of SARS-CoV-2 is essential for early treatment and control of the disease spread. In the past few years, CRISPR technology has shown great potential for highly sensitive and specific molecular diagnostics. Amid the ongoing COVID-19 pandemic, there is an increasing interest in implementing CRISPR-based diagnostic principles to develop fast and precise methods for detecting SARS-CoV-2. In this work, we reviewed and summarized these CRISPR-based diagnostic systems as well as their characteristics and challenges. We also provided future perspectives of CRISPR-based sensing towards point-of-care molecular diagnosis applications.
Keywords: COVID-19, SARS-CoV-2, CRISPR, Cas12, Cas13, Point-of-care diagnosis
1. Introduction
Coronaviruses are enveloped positive-sense RNA viruses, which are commonly associated with acute respiratory infections in humans (Phan 2020). In late December 2019, several local health facilities reported patients with pneumonia of unknown causes in Wuhan, Hubei Province, China (Zhu et al., 2020). The causative pathogen has been identified as a novel enveloped RNA betacoronavirus (Wang et al., 2020d). Given the similarity to the previously isolated severe acute respiratory syndrome coronavirus (SARS-CoV), the new virus has been named SARS-CoV-2 (Castagnoli et al., 2020). On January 9th, 2021, there are a total of 87,589,206 confirmed cases and 1,906,606 deaths of SARS-CoV-2 reported globally by the World Health Organization (WHO) (WHO 2020). In this pandemic of SARS-CoV-2, reliable, early, and accurate diagnosis is crucial to provide timely medical support to the infected individual and facilitate the government agencies to prevent its spread to other individuals and saves lives. The current diagnostic of coronavirus includes detection of virus by genomic techniques using either reverse-transcription-quantitative PCR (RT-qPCR)-based method or sequencing (Corman et al., 2020; Li et al., 2020; Lu et al., 2020).
In the past few years, the emergence and development of clustered regularly interspaced short palindromic repeats (CRISPR) and CRISPR-associated Cas proteins opened a new avenue of research towards molecular diagnostics (Doudna and Charpentier 2014). The discovery of CRISPR took place in 1987 when Ishino et al. discovered an unusual repetitive DNA sequence in Escherichia coli (Barrangou and Marraffini, 2014; Ishino et al., 1987). Subsequent discoveries have unveiled the mechanisms of the CRISPR-Cas system, setting up a new era of CRISPR-Cas mediated adaptive immunity (Barrangou et al., 2007; Deveau et al., 2010; Horvath and Barrangou 2010; Murugan et al., 2017). The revolutionary application of CRISPR-Cas9 technology in the field of gene editing (Cong et al., 2013) paved the way for the subsequent applications in other CRISPR-Cas systems for basic sciences and clinical medicine (Ishino et al., 2018). In 2016, the CRISPR-Cas9 based diagnostics was first utilized to detect the Zike virus (Pardee et al., 2016), and in 2017 for detecting Staphylococcus aureus (Guk et al., 2017).
Later, the discovery of RNA-guided, RNA-targeting CRISPR effector Cas13 (Gootenberg et al., 2017) and subsequently founded Cas12 (Gootenberg et al., 2018) and Cas14 (Harrington et al., 2018) set up a stage of CRISPR-Cas12, 13 or 14-based nucleic acid detection (Karvelis et al., 2019; Kellner et al., 2019; Li et al., 2018b; Myhrvold et al., 2018). The mechanism of Cas12, 13, or 14 systems, which are programmed by guide RNA, allows for single-base specificity compared to the amplification-based molecular diagnostics techniques such as PCR (Gootenberg et al., 2017). The general CRISPR-based nucleic acid detection methods were reviewed in several previous studies (Anson et al., 2020; Ibrahim et al., 2020; van Dongen et al., 2020; Wang et al., 2020c). Due to the growing interest in developing CRISPR-based systems for sensitive and specific diagnostics of SARS-CoV-2 amid the ongoing COVID-19 pandemic, in this work, we specifically focused on the recent progress of applying Cas12 and Cas13 for SARS-CoV-2 detection and reviewed their assay, readout methods, and the system integration. We also discussed the challenges and future perspectives of CRISPR-based SARS-CoV-2 diagnosis.
2. Sample-to-answer workflow in CRISPR-based SARS-CoV-2 detection
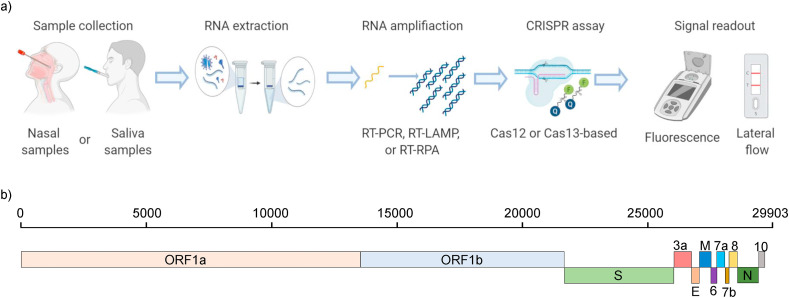
Fig. 1a illustrates the general steps for the sample-to-answer workflow of CRISPR-based diagnostics of SARS-CoV-2. In the first step, the samples are collected from the patients. Table 1 summarized the typical SARS-CoV-2 clinical samples with their collection materials, storage temperature, and viral load ranges. The most general forms of the SARS-CoV-2 samples are nasopharyngeal and oropharyngeal swabs (NPS and OPS) (Lin et al., 2020; Yang et al., 2020).
Fig. 1.
a) Workflow of CRISPR-based SARS-CoV-2 diagnostic schemes from sample to answer. b) Schematic presentation of the SARS-CoV-2 genome organization (Kim et al., 2020a).
Table 1.
SARS-CoV-2 clinical samples collection, storage, and typical viral load.
| Sample Type | Collection materials | Storage temperature until testing | Recommended temperature for shipment according | Viral load (Copies/mL) | Reference |
|---|---|---|---|---|---|
| Nasopharyngeal swabs | Dacron or polyester flocked swabs | 2–8 °C | 2–8 °C if ≤ 5 days −70 °C (dry ice) if > 5 days | 104–107 | (Fajnzylber et al., 2020; Wang et al., 2020d) |
| Oropharyngeal swabs | Dacron or polyester flocked swabs | 2–8 °C | 2–8 °C if ≤ 5 days −70 °C (dry ice) if > 5 days | 104–107 | (Fajnzylber et al., 2020; Wang et al., 2020d) |
| Saliva | Sterile container | 2–8 °C | 2–8 °C if ≤ 3 days −70 °C (dry ice) if > 3 days | 104–107 | (Azzi et al., 2020; Williams et al., 2020a) |
| Sputum | Sterile container | 2–8 °C | 2–8 °C if ≤ 2 days −70 °C (dry ice) if > 2 days | 104–109 | Fajnzylber et al. (2020) |
| Urine | Urine collection container | 2–8 °C | 2–8 °C if ≤ 5 days −70 °C (dry ice) if > 5 days | 102–103 | (Fajnzylber et al., 2020; Kim et al., 2020b) |
| Stool | Stool container | 2–8 °C | 2–8 °C if ≤ 5 days −70 °C (dry ice) if > 5 days | – | Kim et al. (2020b) |
A healthcare provider can collect them by utilizing dacron or polyester flocked swabs. Recently, saliva samples have been introduced as an alternative specimen source for detection of SARS-CoV-2 (McCormick-Baw et al., 2020; Williams et al., 2020b). Since it does not require specialized consumables, it causes less patient discomfort and reduces health worker exposure risk. In respect to storage temperature, all sample types can be stored at 2–8 °C for a short term (less than 2–5 days); however, specimens should be stored at −70 °C for longer storages and shipments. Pan et al. utilized quantitative RT-PCR assays to test different specimen types from 82 infected individuals (Pan et al., 2020). They discovered that samples obtained from sputum showed higher viral loads than OPS samples, followed by stool samples.
After sample collection, viral RNAs need to be extracted from the raw sample. RNA isolation procedures typically involve three general steps: lysis, separation of RNA from other macromolecules such as DNA, proteins, and lipids, followed by RNA elution. RNA-extraction kits are usually used to process patient samples (Eisen et al., 2020; Nalla et al., 2020). Viral RNA separation can be achieved by one of these three main methods (Ravi et al., 2020). Magnetic bead purification. In this method, magnetic beads are utilized to capture the viral RNA. An external magnetic field holds the beads in place during wash and collection. The magnetic format allows for rapid collection/concentration of the sample. However, the capture/release of particles can be laborious when performed manually. Spin column isolation. In this method, membranes usually fabricated by glass fiber, derivatized silica, or ion exchange membranes are used to trap viral RNAs. Afterward, a centrifugal force or vacuum is applied for wash and collection steps. This method is easy to perform and can be automated, but membranes can be clogged with particulate materials. Organic extraction. In this method, samples are homogenized in a phenol-containing solution and then centrifuged. During centrifugation, the upper aqueous phase contains the viral RNA. Therefore, the upper aqueous phase is recovered, and RNA is collected by alcohol precipitation and rehydration. This method is considered the best method for RNA extraction; however, this method is challenging to automate and laborious and manually intensive (Ravi et al., 2020).
After viral RNA isolation, an amplification step such as RT-PCR, RT-LAMP, or RT-RPA was often adopted to boost the limit of detection (LoD) (van Dongen et al., 2020). Afterward, a specific region of SARS-CoV-2 would be recognized by the Cas-CRISPR RNA (crRNA) complex. The genome size of the SARS-CoV-2 varies from 29.8 to 29.9-kilo nucleotides (knt). More than two-thirds of the genome comprises ORF1ab encoding orf1ab polyproteins; the other third consists of genes encoding structural proteins, including surface (S), envelope (E), membrane (M), and nucleocapsid N proteins (Khailany et al., 2020; Kim et al., 2020a) (Fig. 1b). The amplification primers and the crRNAs should be designed specifically to target the same SARS-CoV-2 region. For the signal readout, fluorescence (Li et al., 2018b) or colorimetric (Bai et al., 2019) based systems were two of the most common approaches.
3. Cas12-based SARS-Cov-2 detection
3.1. Cas12 system
Cas12 protein is one of the CRISPR family members, which can be programmed with a CRISPR RNA (crRNA) to specifically bind to the complementary single and double-stranded DNA targets (Aman et al., 2020; Pickar-Oliver and Gersbach 2019). Cas12 has emerged as an alternative for Cas9 due to its distinct features, such as the ability to target T-rich motifs and no need for trans-activating crRNA. To this date, various subtypes of Cas12 proteins have been discovered (Wang et al., 2020a). Table 2 summarizes various subtypes of Cas12 proteins and their characteristics. The size of Cas12 proteins is varying from 870 to 1228 amino acids (aa). Among them, Cas12a (cpf1) (Zetsche et al., 2015) and Cas12b (C2c1) (Liu et al., 2017) are the most well-known.
Table 2.
CRISPR-Cas12 subtypes.
| Cas12 subtype | Size (aa) | PAMa | protospacer (nt) | Cleaving target | Applicationb |
Refs | |
|---|---|---|---|---|---|---|---|
| Gene editing | Diagnostics | ||||||
| Cas12a (Cpf1) | 1228 | TTTV | 20–24 | dsDNA, ssDNA | Yes | Yes | (Chen et al., 2018; Li et al., 2018b) |
| Cas12b (C2c1) | 1129 | TTN | 20 | dsDNA, ssDNA | Yes | Yes | (Li et al., 2019a; Strecker et al., 2019) |
| Cas12c (C2c3) | 1302 | TG | – | dsDNA, ssDNA | Not yet | Not yet | (Harrington et al., 2020; Yan et al., 2019) |
| Cas12d (CasY) | ~1200 | TA | 17–19 | dsDNA | Not yet | Not yet | (Burstein et al., 2017; Harrington et al., 2020) |
| Cas12e (CasX) | ~980 | TTCN | 20 | dsDNA | Yes | Not yet | (Burstein et al., 2017; Yang and Patel 2019) |
| Cas12g1 | 767 | No PAM requirement | – | dsDNA, ssDNA | Not yet | Not yet | Yan et al. (2019) |
| Cas12h1 | 870 | RTR | – | dsDNA, ssDNA | Not yet | Not yet | Yan et al. (2019) |
| Cas12i1 | 1093 | TTN | – | dsDNA, ssDNA | Not yet | Not yet | Yan et al. (2019) |
N = any base; R = A/G; V = A/C/G.
This column presents the application of the Cas12 subtypes to this date (January 2021).
It was recently discovered that Cas12 could indiscriminately cleave the non-target surrounding DNAs (See cleaving targets in Table 2) once activated by a target DNA molecule matching its spacer sequence (Chen et al., 2018). This trans-cleavage activity of Cas12 makes it a powerful tool for detecting a specific target DNA in a mixture (Chen et al., 2018; Gootenberg et al. 2017, 2018; He et al., 2020; Li et al. 2018a, 2019b; Qin et al., 2019; Wang et al., 2019). So far, only Cas12a and Cas12b has been utilized for diagnostic applications, while other subtypes remain to be explored (Table 2). In the following, we primarily focus on Cas12a assay and its crRNA design rules due to its dominance in the applications.
3.2. Cas12a assay workflow
Fig. 2a illustrates the typical steps for the Cas12a-based detection of SARS-CoV-2. There are three main steps: reverse transcription (RT) amplification, Cas12a assay, and signal readout. After viral RNA extraction from raw samples, reverse transcription of the target RNAs and amplification process could be made in one or two steps. In the Cas12a assay, the target amplicons will be introduced to the Cas12a/crRNA complex (a.k.a. non-activated RNP). Upon the specific RNA-guided target binding, the Cas12a would perform collateral cleavage on the surrounding non-target reporters. The reporters are usually fluorophore quencher (FQ)-labeled single-stranded DNA or fluorophore biotin (FB)-labeled single-stranded DNA. After the trans-cleavage of the non-targeted ssDNA reporter, the signal readout can be performed as a fluorescence-based reaction or a single-plex colorimetric lateral flow reaction.
Fig. 2.
a) Typical steps for Cas12-based assay of SARS-CoV-2. b) Schematic of SARS-CoV-2 DNA amplicons detection with the Cas12a/crRNA complex. The target site and crRNA sequence are obtained from Dina et al. study (Ding et al., 2020).
3.3. Cas12a crRNA design rules
Cas12a crRNA is a single, 40–44 base, guide RNA, comprising a 20 nt constant stem-loop region (loop domain) and a 20–24 nt target-specific region (protospacer domain). The loop domain is specific to the Cas12a proteins, and the protospacer domain should be design based on the target sequence. crRNAs used with Cas12a identify locations in the target region with the protospacer adjacent motif (PAM) sequence, TTTV, where V is A, C, or G. Fig. 2b illustrates one example of SARS-CoV-2 recognition with Cas12a/crRNA complex (Ding et al., 2020). crRNA binds to the DNA strand opposite to the PAM sequence. Hence, the protospacer domain should be an exact 20–24 base after the PAM region. The crRNA design rules for the other types of Cas12 proteins are similar to Cas12a; however, the PAM region and protospacer length should be altered (Table 2).
3.4. Benchmarking Cas12-based SARS-CoV-2 detection
Table 3 presents an overview of the most current Cas12-based sensing methods for SARS-CoV-2. We organized this table into three sections: assay, signal readout, and performance.
Table 3.
Summary of different Cas12 based methods for SARS-CoV-2 diagnosis.
| Assay |
Signal readout | Performance |
Refs | |||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Sample source | Sample Prep | Effector | Targeted Gene Region | Pre-Amplification | Number of steps | Bulky instrumentation required |
Assay reaction time* | LoD | ||
| Synthetic gene fragments And nasopharyngeal swab |
Yes | Cas12a | N gene and E gene | RT-LAMP | Two-pot | Colorimetric | No | 30–40 min | 10 copies/μL | Broughton et al. |
| Synthetic gene fragments | No | Cas12a | ORF1ab | RT-RPA | Two-pot | Colorimetric and fluorescence | For colorimetric (No) for fluorescence detection (Yes) | 60 min | 10 copies/μL | Lucia et al. |
| Synthetic gene fragments | No | Cas12a | N gene | RT-RPA | One-pot | Fluorescence | Yes | 40 min | 10 copies/μL | Ding et al. |
| Synthetic gene fragments and nasopharyngeal swab | Yes | Cas12a and Cas12b | N gene and E gene | RT-LAMP | One- pot and two-pot | Fluorescence | Yes | 40 min | 10 copies/μL | Ali et al. |
| Synthetic gene fragments and clinical swab samples | Yes | Cas12b | N gene | RT-RAA | One pot | Colorimetric and fluorescence | For colorimetric (No) for fluorescence (Yes) | 60 min | 10 copies/μL | Guo et al. |
* Without the sample preparation time.
3.4.1. Assay
Sample Sources. Most works used synthetic gene fragments to validate their assay. Working with synthetic gene fragments skipped the RNA extraction in sample preparation. Broughton et al., (2020) were one of the few studies that worked with real raw samples. To extract RNAs from the swab samples, they used two different kits: Qiagen DSP Viral RNA Mini kit and the MagNA Pure 24 instrument. Effector. Both Cas12a and Cas12b have been implemented as the effector in the CRISPR assay. While it is clear that Cas12a dominates, Guo et al., (2020) established a SARS-CoV-2 detection protocol based on their previously reported Cas12b-mediated DNA detection strategy (Teng et al., 2019).
Target SARS-CoV-2 Gene Region. The majority of studies evaluated the sensitivity of their assay based on multiple regions of the SARS-CoV-2 genome (Fig. 1b). For instance, Ali et al., (2020) designed their primers and crRNAs targeting N and E genes of the SARS-CoV-2 genome. To test the performance of their system, they considered 21 positive samples with qPCR. While the N gene assay captured 18 out of 21 samples ( 86%), the E gene assay only detected 8 out of 21 positive samples ( 38%). Lucia et al. designed multiple primers targeting different regions of ORF1ab and achieved an LoD ~10 copies/μL (Lucia et al., 2020). In order to design proper primers, the length of primers and amplicons should be compatible with the amplification method. For instance, RPA primers are preferred to be longer (30–35 nt) than typical PCR primers (18–30 nt) (Euler et al., 2013). Pre-amplification. As previously demonstrated, it is challenging for the CRISPR to detect the targets in a practical time (less than 1 h) when the target DNA concentration is extremely low (lower than 10 nM) (Nouri et al., 2020; Teng et al., 2019). Therefore, most CRISPR-based detection systems utilized different amplification methods such as PCR (Lee et al., 2017; Li et al., 2018b), LAMP (Li et al., 2019a), and RPA (Gootenberg et al., 2018) to enhance their sensitivity. For the RNA targets such as SARS-CoV-2, an additional step of RT should be performed before the application process. So far, most studies utilized isothermal amplification methods such as RT-LAMP (Ali et al., 2020; Broughton et al., 2020), RT-RPA (Ding et al., 2020; Lucia et al., 2020), and RT-RAA (Guo et al., 2020) for their Cas12-based assays. Utilization of isothermal amplification methods eliminates the need for a thermal cycler and increases the suitability of the assay for point-of-care (POC) applications. Number of steps. These pre-amplification process can be separated from CRISPR assay into two pot reactions or be merged as a one-pot reaction. Operating the assay in two-step complicates the testing process, increase the liquid handling, and potentially increases the risk of contaminations due to amplification products transferring. However, adding all the components (amplification and CRISPR reagents) in one pot would decrease the efficiency due to the possible cross-reactions and digestion of the initial amplification products by Cas/crRNA complex (Ali et al., 2020). Therefore, to run the assay in one pot, the Cas12 can be separated from the rest of the components by adding the Cas12 protein in a droplet on the tube wall (Ali et al., 2020; Ding et al., 2020). After running the amplification process, the Cas12 protein can then be mixed with reaction to start the CRISPR assay.
3.4.2. Signal readout
Various readout mechanisms such as fluorescence (Gootenberg et al., 2017), colorimetric (Yuan et al., 2020b), electrochemical (Dai et al., 2019), and electronic (Nouri et al., 2020) have been introduced for CRISPR based assays. The first Cas12 or Cas13-based sensing systems were based on the detection of an increase in fluorescence signal upon target sequence recognition by the RNP complex (Gootenberg et al., 2018; Li et al., 2019a). However, fluorescence readers are normally bulky, expensive, and not suitable for POC applications. Several studies have attempted to reduce the size and cost of fluorometers (Katzmeier et al., 2019; Qin et al., 2019). On the other hand, colorimetric sensors are suitable for POC applications since they are user-friendly, cost-effective, and accessible (van Dongen et al., 2020). Lateral flow assay (LFA) is the most common colorimetric readout system (Shao et al., 2019; Yuan et al., 2020a). Many studies adapted the LFAs platforms as their reading system for CRISPR sensing methods (Bai et al., 2019; Chang et al., 2020). In addition, the electronic readout is a highly promising approach due to its integration potential (Bruch et al., 2019; Dai et al., 2019). While electronic sensors such as nanopores were used with dCas9 (Ashley et al., 2018; Yang et al., 2018) and Cas12a-based (Nouri et al., 2020) assays, its adoption for CRISPR-based SARS-CoV-2 detection has yet to be developed.
3.4.3. Performance
The assay time for the Emergency Use Authorization (EUA) CDC qRT-PCR assay for detecting the SARS-CoV-2 is around 2 h (without considering the sample preparation time) (Broughton et al., 2020). By comparison, all Cas12-based studies reported their turnaround time less than 1 h (without considering the sample preparation time). The LoD for the CDC assay tested by the California Department of Public Health was estimated as 1 copy/μL reaction (Broughton et al., 2020). On the other hand, all Cas12a-based assays demonstrated a consistent LoD of 10 copies/μL (Ali et al., 2020; Broughton et al., 2020; Ding et al., 2020; Guo et al., 2020; Lucia et al., 2020).
4. Cas13-based SARS-Cov-2 detection
4.1. Cas13 system
Cas13 protein is another member of the CRISPR family. Unlike the other CRISPR proteins, Cas13 targets RNAs instead of DNAs (Aman et al., 2020). Cas13 encompasses four divergent subtypes (Cas13a to Cas13d) (Cox et al., 2017). The characteristics of these subtypes are summarized in Table 4 . The Cas13 proteins range in size from 954 to 1389 aa. Cas13a (a.k.a. C2c2) is the first and most well-known Cas13 protein which has been implemented in most Cas13-based diagnostic applications.
Table 4.
CRISPR-Cas13 subtypes.
| Cas13 subtype | Size (aa) | PFSa | Spacer (nt) | Cleaving target | Applicationb |
Refs | |
|---|---|---|---|---|---|---|---|
| Gene editing | Diagnostics | ||||||
| Cas13a (C2c2) | 1389 | H | 22–28 | ssRNA | Yes | Yes | (Abudayyeh et al., 2016; Cox et al., 2017) |
| Cas13b (C2c6) | 1224 | No PFS requirement | 30 | ssRNA | Yes | Yes | (Gootenberg et al., 2018; Smargon et al., 2017) |
| Cas13c (C2c7) | 1121 | NAN/NNA | – | ssRNA | Yes | Not yet | (Cox et al., 2017; Shmakov et al., 2017) |
| Cas13d | 954 | No PFS requirement | 22 | ssRNA | Yes | Not yet | (Konermann et al., 2018; Yan et al., 2018) |
N = any base; H = A/C/T.
This column presents the application of the Cas13 subtypes to this date (January 2021).
Similar to Cas12, Cas13 exhibits cleavage activity when it is activated by ssRNA sequence bearing complementarity to its crRNA spacer. However, activated Cas13 cleaves all surrounding ssRNAs (Table 4) instead of DNAs. This property has been utilized in vitro for highly specific diagnostics (Abudayyeh et al., 2019; Gootenberg et al., 2017; Kellner et al., 2019). So far, Cas13a was the most commonly used in most of the Cas13-based diagnostics. In the following, we will focus on the Cas13a assay and its crRNA design rules.
4.2. Cas13a assay workflow
Fig. 3a presents the typical steps of SARS-CoV-2 via Cas13a-based sensors. The process is almost identical to the Cas12a-based method with some small variations. Since RNA targets would only activate Cas13a proteins, an additional T7 transcription is needed after amplification to convert the DNA amplicons to RNAs. Fortunately, this step can be merged into the amplification and Cas13a assay as a single pot reaction (Kellner et al., 2019). In addition, because activated Cas13a cleaves the ssRNA rather than ssDNAs, reporters should be designed using ssRNA in contrast to ssDNA reporters used in Cas12a systems.
Fig. 3.
a) Typical steps for Cas13-based assay of SARS-CoV-2. b) Schematic of SARS-CoV-2 RNA detection mechanism with the Cas13a/crRNA complex. The target site and crRNA sequence are obtained from Hou et al. study (Hou et al., 2020).
4.3. Cas13a crRNA design rules
Cas13a crRNA contains a single stem-loop specific to Cas13a and a protospacer domain-specific to the target. Cas13a requires a protospacer domain of at least 22 nt length to efficiently cleave ssRNAs (Abudayyeh et al., 2016). The optimal protospacer length observed for Cas13a is 28 nucleotides along (Khan et al., 2018). The stem-loop structure of the crRNA is also critical for ssRNA cleavage. Abudayyeh et al., (2016) showed that a stem-loop longer than 24 nt is required for Cas13a to mediate ssRNA cleavage. Abudayyeh et al. also analyzed the flanking regions of protospacers. They found that sequences starting with a G immediately after the 3′ end of the protospacer were less effective relative to all other nucleotides (A, U, or C). Therefore, the protospacer-flanking site (PFS) should be considered in crRNA designs for the Cas13a effector. Fig. 3b shows an example of SARS-CoV-2 recognition with the Cas13a/crRNA complex, targeting ORF1ab (Hou et al., 2020). The protospacer domain is 27 nt long, complementing the target site of the SARS-CoV-2. The loop domain is a 36 nt long stem-loop structure that was obtained from a design by Gootenberg et al., (2017). The other subtypes of Cas13a crRNA designs are similar to Cas13a; however, the PFS region and protospacer domain length are different (Table 4).
4.4. Benchmarking Cas13-based SARS-CoV-2 detection
Table 5 gives an overview of the most current Cas13 sensing methods for SARS-CoV-2. In the following, we will benchmark their methods and performances.
Table 5.
Summary of different Cas13 based methods for SARS-CoV-2 diagnosis.
| Assay |
Signal readout | Performance |
Refs | |||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Sample source | Sample Prep |
Effector | Gene or region detected | Amplification Type | Number of steps | Bulky instrumentation required |
Assay reaction time* | LoD | ||
| Synthetic gene fragments | No | Cas13a | N gene | RT-PCR | Two step | Colorimetric | No | Not reported | 10 copies/μL | Rauch et al. |
| Synthetic gene fragments And saliva samples |
Yes | Cas13a | ORF1a | RT-RPA | One pot and two step | Colorimetric and fluorescence | For colorimetric (No) for fluorescence (Yes) | 1 h | 10 copies/μL | Arizti-Sanz et al. |
| Synthetic gene fragments And nasopharyngeal swab |
Yes | Cas13a | ORF1ab and N | RT-RPA | Two step | Fluorescence | yes | 40 min | 1 copies/μL | Hou et al. |
| Synthetic gene fragments | No | Cas13a | Not reported | RT-RPA | Two step | Colorimetric | No | 50 min | 10 copies/μL | Metsky et al. |
| Synthetic gene fragments and nasopharyngeal and throat swabs | Yes | Cas13a | Orf1ab, S, and N | RT-RPA | Two step | Colorimetric and fluorescence | For colorimetric (No) for fluorescence (Yes) |
35–70 min | 10 copies/μL | Patchsung et al. |
* Without the sample preparation time.
4.4.1. Assay
Sample Sources. Most reported Cas13-based work tested their system with real samples after validating their assays with synthetic genes. Arizti-Sanz et al., (2020) tested the saliva samples and successfully detected SARS-CoV-2 via Cas13a assay. Their RNA extraction was performed using MagMAX™ mirVana™ Total RNA isolation kit. In other studies (Hou et al., 2020; Patchsung et al., 2020), nasopharyngeal swabs were used. Effector. So far, most studies employed Cas13a as their assay effector for detecting SARS-CoV-2. Although other Cas13 family members have not yet been implemented for diagnostics, it is highly desirable to exploring them since they could relax the PFS sequence constraints when designing the specific target region. Target SARS-CoV-2 Gene Region. Analogous to Cas12 assays, most studies examined multiple genes as their targets to optimize the performance. Nevertheless, distinct genes were claimed by different studies as the best target to achieve higher sensitivities. For instance, Hou et al., (2020) reached the best overall performance of sensitivity and specificity by selecting their target from the ORF1ab region (after evaluating the performance of two sets of primers targeting N and Orf1ab genes). On the other hand, Patchsung et al., (2020) achieved an LoD of 10 copies/μL by targeting the S gene in comparison to LoD of 104 copies/μL for three sets of primers targeting N and Orf1ab genes. Pre-amplification. RT-RPA was selected as the amplification method for most Cas13-based methods for SARS-CoV-2 detection. Compared to LAMP, RPA appears to be an attractive technology for diagnostics as it is both faster and has a simpler primer design (Zou et al., 2020). RPA could also be performed at room temperature, which makes it ideal for POC applications (Crannell et al., 2014). Number of steps. In the Cas13a assay, the amplification process can be merged with T7 transcription and Cas13-based detection. Arizti-Sanz et al., (2020) compared the sensitivity of one and two steps assay. Initially, they observed that the sensitivity of the two steps assay was dramatically lower than one step (LoD of 106 copies/μL in respect to LoD of 10 copies/μL for two steps assay). This was due to the incompatibility of enzymatic reactions with the given conditions. Subsequently, after optimizing pHs, monovalent salt, magnesium, and primer concentrations, they achieved an identical LoD for two and one steps assays (10 copies/μL).
4.4.2. Signal readout
Similar to Cas12-based methods, fluorescence (Hou et al., 2020; Rauch et al., 2020), and colorimetric (Metsky et al., 2020) were the only chosen signal readout methods for Cas13-based methods for SARS-CoV-2 detection. Rauch et al., (2020) were one of the studies that explored the utilization of low-cost, portable, and field-ready instruments for their assay and signal readout system. For PCR amplification, they used miniPCR mini16, which was an affordable thermocycler and convenient for POC applications. Instead of using conventional fluorometers, they used a P51 cardboard fluorescence visualizer powered by a 9V battery for their readout system.
4.4.3. Performance
The performance of the Cas13-based assays was almost identical to Cas12-based ones. Most studies reporter their assay LoD as 10 copies/μL, and their turnaround time was less than 1 h (Table 5).
5. Summary and future perspective
CRISPR technology provides an appealing opportunity to facilitate superior alternatives or improvements by providing a rapid, in-field, sensitive, and specific assay for SARS-CoV-2 detection. So far, the majority of the CRISPR-Cas systems utilized Cas12 or Cas13 proteins as the CRISPR effectors in their assays. However, other types of CRISPR effectors such as Cas9 (Wang et al., 2020b) and dCas9 (Lee et al., 2018; Yang et al., 2018) have been implemented for other pathogen detection. Utilizing the other types of CRISPR proteins could expand the targeting range (Wang et al., 2020a) and provide new signal readout methods (Hajian et al., 2019; Koo et al., 2018). While CRISPR-based sensing showed great potential in specific, sensitive, and affordable nucleic acid detection in the last couple of years, this field is still in its beginning. Particularly, the development of ‘sample-in-answer-out’ CRISPR-based point of care devices is highly desirable. To this end, we need to address the following challenges.
Integrated sample preparation. According to guidelines by WHO, key features in accessing the practicality of a POC diagnostic device are affordable, sensitive, specific, user-friendly, rapid and robust, equipment-free, and deliverable (ASSURED). Based on these guidelines, the most critical lacking factor in CRISPR methods is the assay runtime (Anson et al., 2020). Sample preparation consists of a large part of the CRISPR-based diagnostic methods. In addition, the sample preparations are almost performed separately for all existing studies. This process would increase the complexity of the CRISPR-based test and reduce the user-friendliness of the assay. It is desirable to create a diagnostic test that combines all steps with only a single action required by the user. Fortunately, several studies have already looked into SARS-CoV-2 sample preparation to expedite and simplify the process (Marais et al., 2020; Rabe and Cepko 2020; Wozniak et al., 2020). In addition, some studies have also examined sample preparation-free assay by utilizing RT-PCR (Lübke et al., 2020) and RT-LAMP (Wei et al., 2020). Future works should focus on developing a fully streamlined process (raw sample-in-answer-out) by seamlessly combining the sample preparation with the CRISPR assay.
Limited target regions. For both Cas12 and Cas13 effectors, the target sequence is limited to specific regions due to the constraint of PAM or PFS. This could be problematic for short target sequences. Therefore, other Cas variations with altered PAM or PFS specificities should be explored and expanded. For example, Kleinstiver et al. engineered an enhanced Acidaminococcus sp. Cas12a variant (enAsCas12a) (Kleinstiver et al., 2019). The novel enAsCas12a has the ability to recognize VTTV/TTTT/TTCN/TATV PAMs. The discovery of the new Cas proteins and also utilizing the other types of Cas proteins such as Cas9 (van Dongen et al., 2020) and Cas14 (Harrington et al., 2018) could significantly enhance the flexibility of choosing and designing the specific target regions.
Multiplexing. Many applications require the detection of more than one target in a single reaction, so-called multiplexing. Multiplex detection from a single sample offers numerous advantages such as rapid turnaround times and low sample consumption (Dincer et al., 2017). However, multiplexed detection is challenging because of the interference between recognition molecules and various analytes and possible cross-reactions (Li et al., 2019c). One of the major issues of Cas12 and Cas13 systems for multiplexed sensing is the non-specific collateral cleavage of all ssDNA/ssRNA sequences, possibly destroying the other targets. Gootenberg et al. developed one of the first multiplexed Cas systems by using four different Cas effectors (PsmCas13b, LwaCas13a, CcaCas13b, and AsCas12a) to detect four different targets in a single reaction (Gootenberg et al., 2018). While this approach showed the possibility of multiplexed sensing in CRISPR systems, the scale-up of the multiplex sensing is limited by the amount of different Cas effectors. This can be a significant research topic for future CRISPR based nucleic acid detection.
Enhancing the CRISPR sensing performance. Nanomaterials and artificial intelligence have been introduced to enhance the performance of CRISPR sensing with respect to signal enhancement (Rabiee et al., 2020) and classification (Ibrahim et al., 2020). For instance, Bao et al. developed a simple visual detection system coupled with quantum dots as an ultra-brightness indicator and a CRISPR-Cas12a assay for isothermal viral DNA target sensing. They avoided bulky and complicated sensing instruments and utilized a handheld flashlight to distinguish between positive and negative samples. Furthermore, point of care sensing technology can be combined with artificial intelligence (AI) for improved data storage, classification, and sharing (Kaushik et al., 2020). For instance, Ibrahim et al. combined the internet of thing technology and machine learning with CRISPR sensing for wireless transmission of signals over the cloud to support decision making (Ibrahim et al., 2020). This opens up many possibilities for further enhancing the CRISPR sensing performance in the future.
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgments
This work was supported by the National Institutes of Health (R61AI147419), and National Science Foundation (1710831, 1902503, and 1912410). Any opinions, findings, and conclusions or recommendations expressed in this work are those of the authors and do not necessarily reflect the views of the National Science Foundation.
References
- Abudayyeh O.O., Gootenberg J.S., Kellner M.J., Zhang F. Nucleic acid detection of plant genes using CRISPR-Cas13. CRISPR J. 2019;2(3):165–171. doi: 10.1089/crispr.2019.0011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Abudayyeh O.O., Gootenberg J.S., Konermann S., Joung J., Slaymaker I.M., Cox D.B., Shmakov S., Makarova K.S., Semenova E., Minakhin L. C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science. 2016;353(6299):5573. doi: 10.1126/science.aaf5573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ali Z., Aman R., Mahas A., Rao G.S., Tehseen M., Marsic T., Salunke R., Subudhi A.K., Hala S.M., Hamdan S.M. iSCAN: an RT-LAMP-coupled CRISPR-Cas12 module for rapid, sensitive detection of SARS-CoV-2. Virus Res. 2020;288:198129. doi: 10.1016/j.virusres.2020.198129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aman R., Mahas A., Mahfouz M. Nucleic acid detection using CRISPR/cas biosensing technologies. ACS Synth. Biol. 2020;9(6):1226–1233. doi: 10.1021/acssynbio.9b00507. [DOI] [PubMed] [Google Scholar]
- Anson L., Carolyn L.R., Luke P.L. Critical review on where CRISPR meets molecular diagnostics. Prog. Biomed. Eng. 2020;3(1):012001. [Google Scholar]
- Arizti-Sanz J., Freije C.A., Stanton A.C., Boehm C.K., Petros B.A., Siddiqui S., Shaw B.M., Adams G., Kosoko-Thoroddsen T.-S.F., Kemball M.E. bioRxiv; 2020. Integrated Sample Inactivation, Amplification, and Cas13-Based Detection of SARS-CoV-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ashley J., Shukor Y., D'Aurelio R., Trinh L., Rodgers T.L., Temblay J., Pleasants M., Tothill I.E. Synthesis of MIP nanoparticles for α-casein detection using SPR as a milk allergen sensor. ACS Sens. 2018;3:418–424. doi: 10.1021/acssensors.7b00850. [DOI] [PubMed] [Google Scholar]
- Azzi L., Carcano G., Gianfagna F., Grossi P., Dalla Gasperina D., Genoni A., Fasano M., Sessa F., Tettamanti L., Carinci F. 2020. Saliva is a reliable tool to detect SARS-CoV-2. Journal of Infection. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bai J., Lin H., Li H., Zhou Y., Liu J., Zhong G., Wu L., Jiang W., Du H., Yang J. Cas12a-based on-site and rapid nucleic acid detection of African swine fever. Front. Microbiol. 2019;10:2830. doi: 10.3389/fmicb.2019.02830. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barrangou R., Fremaux C., Deveau H., Richards M., Boyaval P., Moineau S., Romero D.A., Horvath P. CRISPR provides acquired resistance against viruses in prokaryotes. Science. 2007;315(5819):1709–1712. doi: 10.1126/science.1138140. [DOI] [PubMed] [Google Scholar]
- Barrangou R., Marraffini L.A. CRISPR-Cas systems: prokaryotes upgrade to adaptive immunity. Mol. Cell. 2014;54(2):234–244. doi: 10.1016/j.molcel.2014.03.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Broughton J.P., Deng X., Yu G., Fasching C.L., Servellita V., Singh J., Miao X., Streithorst J.A., Granados A., Sotomayor-Gonzalez A., Zorn K., Gopez A., Hsu E., Gu W., Miller S., Pan C.-Y., Guevara H., Wadford D.A., Chen J.S., Chiu C.Y. CRISPR–Cas12-based detection of SARS-CoV-2. Nat. Biotechnol. 2020;38(7):870–874. doi: 10.1038/s41587-020-0513-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bruch R., Baaske J., Chatelle C., Meirich M., Madlener S., Weber W., Dincer C., Urban G.A. CRISPR/Cas13a-powered electrochemical microfluidic biosensor for nucleic acid amplification-free miRNA diagnostics. Adv. Mater. 2019;31(51):1905311. doi: 10.1002/adma.201905311. [DOI] [PubMed] [Google Scholar]
- Burstein D., Harrington L.B., Strutt S.C., Probst A.J., Anantharaman K., Thomas B.C., Doudna J.A., Banfield J.F. New CRISPR–Cas systems from uncultivated microbes. Nature. 2017;542(7640):237–241. doi: 10.1038/nature21059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Castagnoli R., Votto M., Licari A., Brambilla I., Bruno R., Perlini S., Rovida F., Baldanti F., Marseglia G.L. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection in children and adolescents: a systematic review. JAMA Pediatr. 2020;174(9):882–889. doi: 10.1001/jamapediatrics.2020.1467. [DOI] [PubMed] [Google Scholar]
- Chang Y., Deng Y., Li T., Wang J., Wang T., Tan F., Li X., Tian K. Visual detection of porcine reproductive and respiratory syndrome virus using CRISPR-Cas13a. Transb. Emerg. Dis. 2020;67(2):564–571. doi: 10.1111/tbed.13368. [DOI] [PubMed] [Google Scholar]
- Chen J.S., Ma E., Harrington L.B., Da Costa M., Tian X., Palefsky J.M., Doudna J.A. CRISPR-Cas12a target binding unleashes indiscriminate single-stranded DNase activity. Science. 2018;360(6387):436–439. doi: 10.1126/science.aar6245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cong L., Ran F.A., Cox D., Lin S., Barretto R., Habib N., Hsu P.D., Wu X., Jiang W., Marraffini L.A. Multiplex genome engineering using CRISPR/Cas systems. Science. 2013;339(6121):819–823. doi: 10.1126/science.1231143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Corman V.M., Landt O., Kaiser M., Molenkamp R., Meijer A., Chu D.K., Bleicker T., Brünink S., Schneider J., Schmidt M.L. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Euro Surveill. 2020;25(3):2000045. doi: 10.2807/1560-7917.ES.2020.25.3.2000045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cox D.B., Gootenberg J.S., Abudayyeh O.O., Franklin B., Kellner M.J., Joung J., Zhang F. RNA editing with CRISPR-Cas13. Science. 2017;358(6366):1019–1027. doi: 10.1126/science.aaq0180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crannell Z.A., Rohrman B., Richards-Kortum R. Equipment-free incubation of recombinase polymerase amplification reactions using body heat. PloS One. 2014;9(11):e112146. doi: 10.1371/journal.pone.0112146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dai Y., Somoza R.A., Wang L., Welter J.F., Li Y., Caplan A.I., Liu C.C. Exploring the trans-cleavage activity of CRISPR-cas12a (cpf1) for the development of a universal electrochemical biosensor. Angew. Chem. Int. Ed. 2019;58(48):17399–17405. doi: 10.1002/anie.201910772. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deveau H., Garneau J.E., Moineau S. CRISPR/Cas system and its role in phage-bacteria interactions. Annu. Rev. Microbiol. 2010;64:475–493. doi: 10.1146/annurev.micro.112408.134123. [DOI] [PubMed] [Google Scholar]
- Dincer C., Bruch R., Kling A., Dittrich P.S., Urban G.A. Multiplexed point-of-care testing–xPOCT. Trends Biotechnol. 2017;35(8):728–742. doi: 10.1016/j.tibtech.2017.03.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ding X., Yin K., Li Z., Lalla R.V., Ballesteros E., Sfeir M.M., Liu C. Ultrasensitive and visual detection of SARS-CoV-2 using all-in-one dual CRISPR-Cas12a assay. Nat. Commun. 2020;11(1):1–10. doi: 10.1038/s41467-020-18575-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Doudna J.A., Charpentier E. The new frontier of genome engineering with CRISPR-Cas9. Science. 2014;346(6213):1258096. doi: 10.1126/science.1258096. [DOI] [PubMed] [Google Scholar]
- Eisen A.K.A., Demoliner M., Gularte J.S., Hansen A.W., Schallenberger K., Mallmann L., Hermann B.S., Heldt F.H., de Almeida P.R., Fleck J.D. bioRxiv; Brazil: 2020. Comparison of Different Kits for SARS-CoV-2 RNA Extraction Marketed in. [DOI] [Google Scholar]
- Euler M., Wang Y., Heidenreich D., Patel P., Strohmeier O., Hakenberg S., Niedrig M., Hufert F.T., Weidmann M. Development of a panel of recombinase polymerase amplification assays for detection of biothreat agents. J. Clin. Microbiol. 2013;51(4):1110–1117. doi: 10.1128/JCM.02704-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fajnzylber J., Regan J., Coxen K., Corry H., Wong C., Rosenthal A., Worrall D., Giguel F., Piechocka-Trocha A., Atyeo C. SARS-CoV-2 viral load is associated with increased disease severity and mortality. Nat. Commun. 2020;11(1):1–9. doi: 10.1038/s41467-020-19057-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gootenberg J.S., Abudayyeh O.O., Kellner M.J., Joung J., Collins J.J., Zhang F. Multiplexed and portable nucleic acid detection platform with Cas13, Cas12a, and Csm6. Science. 2018;360(6387):439–444. doi: 10.1126/science.aaq0179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gootenberg J.S., Abudayyeh O.O., Lee J.W., Essletzbichler P., Dy A.J., Joung J., Verdine V., Donghia N., Daringer N.M., Freije C.A. Nucleic acid detection with CRISPR-Cas13a/C2c2. Science. 2017;356(6336):438–442. doi: 10.1126/science.aam9321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guk K., Keem J.O., Hwang S.G., Kim H., Kang T., Lim E.-K., Jung J. A facile, rapid and sensitive detection of MRSA using a CRISPR-mediated DNA FISH method, antibody-like dCas9/sgRNA complex. Biosens. Bioelectron. 2017;95:67–71. doi: 10.1016/j.bios.2017.04.016. [DOI] [PubMed] [Google Scholar]
- Guo L., Sun X., Wang X., Liang C., Jiang H., Gao Q., Dai M., Qu B., Fang S., Mao Y. SARS-CoV-2 detection with CRISPR diagnostics. Cell Discov. 2020;6(1):1–4. doi: 10.1038/s41421-020-0174-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hajian R., Balderston S., Tran T., DeBoer T., Etienne J., Sandhu M., Wauford N.A., Chung J.-Y., Nokes J., Athaiya M. Detection of unamplified target genes via CRISPR–Cas9 immobilized on a graphene field-effect transistor. Nat. Biomed. Eng. 2019;3(6):427–437. doi: 10.1038/s41551-019-0371-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harrington L.B., Burstein D., Chen J.S., Paez-Espino D., Ma E., Witte I.P., Cofsky J.C., Kyrpides N.C., Banfield J.F., Doudna J.A. Programmed DNA destruction by miniature CRISPR-Cas14 enzymes. Science. 2018;362(6416):839–842. doi: 10.1126/science.aav4294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harrington L.B., Ma E., Chen J.S., Witte I.P., Gertz D., Paez-Espino D., Al-Shayeb B., Kyrpides N.C., Burstein D., Banfield J.F. A scoutRNA is required for some type V CRISPR-cas systems. Mol. Cell. 2020;79(3):416–424. doi: 10.1016/j.molcel.2020.06.022. e415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- He Q., Yu D., Bao M., Korensky G., Chen J., Shin M., Kim J., Park M., Qin P., Du K. High-throughput and all-solution phase African Swine Fever Virus (ASFV) detection using CRISPR-Cas12a and fluorescence based point-of-care system. Biosens. Bioelectron. 2020;154:112068. doi: 10.1016/j.bios.2020.112068. [DOI] [PubMed] [Google Scholar]
- Horvath P., Barrangou R. CRISPR/Cas, the immune system of bacteria and archaea. Science. 2010;327(5962):167–170. doi: 10.1126/science.1179555. [DOI] [PubMed] [Google Scholar]
- Hou T., Zeng W., Yang M., Chen W., Ren L., Ai J., Wu J., Liao Y., Gou X., Li Y. medRxiv; 2020. Development and Evaluation of a CRISPR-Based Diagnostic for 2019-novel Coronavirus. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ibrahim A.U., Al-Turjman F., Sa’id Z., Ozsoz M. Futuristic CRISPR-based biosensing in the cloud and internet of things era: an overview. Multimed. Tool. Appl. 2020 doi: 10.1007/s11042-020-09010-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ishino Y., Krupovic M., Forterre P. History of CRISPR-cas from encounter with a mysterious repeated sequence to genome editing technology. J. Bacteriol. 2018;200(7) doi: 10.1128/JB.00580-17. e00580–00517. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ishino Y., Shinagawa H., Makino K., Amemura M., Nakata A. Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. J. Bacteriol. 1987;169(12):5429–5433. doi: 10.1128/jb.169.12.5429-5433.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karvelis T., Bigelyte G., Young J.K., Hou Z., Zedaveinyte R., Pociute K., Silanskas A., Venclovas Č., Siksnys V. 2019. PAM recognition by miniature CRISPR-Cas14 triggers programmable double-stranded DNA cleavage. bioRxiv, 654897. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Katzmeier F., Aufinger L., Dupin A., Quintero J., Lenz M., Bauer L., Klumpe S., Sherpa D., Dürr B., Honemann M. A low-cost fluorescence reader for in vitro transcription and nucleic acid detection with Cas13a. PloS One. 2019;14(12) doi: 10.1371/journal.pone.0220091. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaushik A.K., Dhau J.S., Gohel H., Mishra Y.K., Kateb B., Kim N.-Y., Goswami D.Y. Electrochemical SARS-CoV-2 sensing at point-of-care and artificial intelligence for intelligent COVID-19 management. ACS Appl. Bio Mater. 2020;3(11):7306–7325. doi: 10.1021/acsabm.0c01004. [DOI] [PubMed] [Google Scholar]
- Kellner M.J., Koob J.G., Gootenberg J.S., Abudayyeh O.O., Zhang F. SHERLOCK: nucleic acid detection with CRISPR nucleases. Nat. Protoc. 2019;14(10):2986–3012. doi: 10.1038/s41596-019-0210-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khailany R.A., Safdar M., Ozaslan M. Genomic characterization of a novel SARS-CoV-2. Gene Rep. 2020;19:100682. doi: 10.1016/j.genrep.2020.100682. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khan M.Z., Amin I., Hameed A., Mansoor S. CRISPR–Cas13a: prospects for plant virus resistance. Trends Biotechnol. 2018;36(12):1207–1210. doi: 10.1016/j.tibtech.2018.05.005. [DOI] [PubMed] [Google Scholar]
- Kim D., Lee J.-Y., Yang J.-S., Kim J.W., Kim V.N., Chang H. The architecture of SARS-CoV-2 transcriptome. Cell. 2020;181(4):914–921. doi: 10.1016/j.cell.2020.04.011. e910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim J.-M., Kim H.M., Lee E.J., Jo H.J., Yoon Y., Lee N.-J., Son J., Lee Y.-J., Kim M.S., Lee Y.-P. Detection and isolation of SARS-CoV-2 in serum, urine, and stool specimens of COVID-19 patients from the Republic of Korea. Osong Publ. Health Res. Perspect. 2020;11(3):112. doi: 10.24171/j.phrp.2020.11.3.02. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kleinstiver B.P., Sousa A.A., Walton R.T., Tak Y.E., Hsu J.Y., Clement K., Welch M.M., Horng J.E., Malagon-Lopez J., Scarfò I. Engineered CRISPR–Cas12a variants with increased activities and improved targeting ranges for gene, epigenetic and base editing. Nat. Biotechnol. 2019;37(3):276–282. doi: 10.1038/s41587-018-0011-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Konermann S., Lotfy P., Brideau N.J., Oki J., Shokhirev M.N., Hsu P.D. Transcriptome engineering with RNA-targeting type VI-D CRISPR effectors. Cell. 2018;173(3):665–676. doi: 10.1016/j.cell.2018.02.033. e614. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koo B., Kim D.-e., Kweon J., Jin C.E., Kim S.-H., Kim Y., Shin Y. CRISPR/dCas9-mediated biosensor for detection of tick-borne diseases. Sensor. Actuator. B Chem. 2018;273:316–321. doi: 10.1016/j.snb.2018.06.069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee H., Choi J., Jeong E., Baek S., Kim H.C., Chae J.-H., Koh Y., Seo S.W., Kim J.-S., Kim S.J. dCas9-mediated nanoelectrokinetic direct detection of target gene for liquid biopsy. Nano Lett. 2018;18(12):7642–7650. doi: 10.1021/acs.nanolett.8b03224. [DOI] [PubMed] [Google Scholar]
- Lee S.H., Yu J., Hwang G.H., Kim S., Kim H., Ye S., Kim K., Park J., Park D., Cho Y.-K. CUT-PCR: CRISPR-mediated, ultrasensitive detection of target DNA using PCR. Oncogene. 2017;36(49):6823–6829. doi: 10.1038/onc.2017.281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li L., Li S., Wu N., Wu J., Wang G., Zhao G., Wang J. HOLMESv2: a CRISPR-Cas12b-assisted platform for nucleic acid detection and DNA methylation quantitation. ACS Synth. Biol. 2019;8(10):2228–2237. doi: 10.1021/acssynbio.9b00209. [DOI] [PubMed] [Google Scholar]
- Li Q., Guan X., Wu P., Wang X., Zhou L., Tong Y., Ren R., Leung K.S.M., Lau E.H.Y., Wong J.Y., Xing X., Xiang N., Wu Y., Li C., Chen Q., Li D., Liu T., Zhao J., Liu M., Tu W., Chen C., Jin L., Yang R., Wang Q., Zhou S., Wang R., Liu H., Luo Y., Liu Y., Shao G., Li H., Tao Z., Yang Y., Deng Z., Liu B., Ma Z., Zhang Y., Shi G., Lam T.T.Y., Wu J.T., Gao G.F., Cowling B.J., Yang B., Leung G.M., Feng Z. Early transmission dynamics in wuhan, China, of novel coronavirus–infected pneumonia. N. Engl. J. Med. 2020;382(13):1199–1207. doi: 10.1056/NEJMoa2001316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li S.-Y., Cheng Q.-X., Liu J.-K., Nie X.-Q., Zhao G.-P., Wang J. CRISPR-Cas12a has both cis-and trans-cleavage activities on single-stranded DNA. Cell Res. 2018;28(4):491–493. doi: 10.1038/s41422-018-0022-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li S.-Y., Cheng Q.-X., Wang J.-M., Li X.-Y., Zhang Z.-L., Gao S., Cao R.-B., Zhao G.-P., Wang J. CRISPR-Cas12a-assisted nucleic acid detection. Cell Discov. 2018;4(1):1–4. doi: 10.1038/s41421-018-0028-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Y., Li S., Wang J., Liu G. CRISPR/Cas systems towards next-generation biosensing. Trends Biotechnol. 2019;37(7):730–743. doi: 10.1016/j.tibtech.2018.12.005. [DOI] [PubMed] [Google Scholar]
- Li Y., Liu L., Liu G. CRISPR/Cas multiplexed biosensing: a challenge or an insurmountable obstacle? Trends Biotechnol. 2019;37(8):792–795. doi: 10.1016/j.tibtech.2019.04.012. [DOI] [PubMed] [Google Scholar]
- Liu L., Chen P., Wang M., Li X., Wang J., Yin M., Wang Y. C2c1-sgRNA complex structure reveals RNA-guided DNA cleavage mechanism. Mol. Cell. 2017;65(2):310–322. doi: 10.1016/j.molcel.2016.11.040. [DOI] [PubMed] [Google Scholar]
- Lu R., Zhao X., Li J., Niu P., Yang B., Wu H., Wang W., Song H., Huang B., Zhu N. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet. 2020;395(10224):565–574. doi: 10.1016/S0140-6736(20)30251-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lübke N., Senff T., Scherger S., Hauka S., Andrée M., Adams O., Timm J., Walker A. Extraction-free SARS-CoV-2 detection by rapid RT-qPCR universal for all primary respiratory materials. J. Clin. Virol. 2020;130:104579. doi: 10.1016/j.jcv.2020.104579. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lucia C., Federico P.-B., Alejandra G.C. bioRxiv; 2020. An Ultrasensitive, Rapid, and Portable Coronavirus SARS-CoV-2 Sequence Detection Method Based on CRISPR-Cas12. [DOI] [Google Scholar]
- Marais G., Naidoo M., Hsiao N.y., Valley-Omar Z., Smuts H., Hardie D. The implementation of a rapid sample preparation method for the detection of SARS-CoV-2 in a diagnostic laboratory in South Africa. PloS One. 2020;15(10) doi: 10.1371/journal.pone.0241029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McCormick-Baw C., Morgan K., Gaffney D., Cazares Y., Jaworski K., Byrd A., Molberg K., Cavuoti D. Saliva as an alternate specimen source for detection of SARS-CoV-2 in symptomatic patients using cepheid xpert xpress SARS-CoV-2. J. Clin. Microbiol. 2020;58(8) doi: 10.1128/JCM.01109-20. e01109–01120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Metsky H.C., Freije C.A., Kosoko-Thoroddsen T.-S.F., Sabeti P.C., Myhrvold C. bioRxiv; 2020. CRISPR-based COVID-19 Surveillance Using a Genomically-Comprehensive Machine Learning Approach. [DOI] [Google Scholar]
- Murugan K., Babu K., Sundaresan R., Rajan R., Sashital D.G. The revolution continues: newly discovered systems expand the CRISPR-Cas toolkit. Mol. Cell. 2017;68(1):15–25. doi: 10.1016/j.molcel.2017.09.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Myhrvold C., Freije C.A., Gootenberg J.S., Abudayyeh O.O., Metsky H.C., Durbin A.F., Kellner M.J., Tan A.L., Paul L.M., Parham L.A. Field-deployable viral diagnostics using CRISPR-Cas13. Science. 2018;360(6387):444–448. doi: 10.1126/science.aas8836. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nalla A.K., Casto A.M., Huang M.-L.W., Perchetti G.A., Sampoleo R., Shrestha L., Wei Y., Zhu H., Jerome K.R., Greninger A.L. Comparative performance of SARS-CoV-2 detection assays using seven different primer-probe sets and one assay kit. J. Clin. Microbiol. 2020;58(6) doi: 10.1128/JCM.00557-20. e00557–00520. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nouri R., Jiang Y., Lian X.L., Guan W. Sequence-specific recognition of HIV-1 DNA with solid-state CRISPR-cas12a-assisted nanopores (SCAN) ACS Sens. 2020;5(5):1273–1280. doi: 10.1021/acssensors.0c00497. [DOI] [PubMed] [Google Scholar]
- Pan Y., Zhang D., Yang P., Poon L.L., Wang Q. Viral load of SARS-CoV-2 in clinical samples. Lancet Infect. Dis. 2020;20(4):411–412. doi: 10.1016/S1473-3099(20)30113-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pardee K., Green A.A., Takahashi M.K., Braff D., Lambert G., Lee J.W., Ferrante T., Ma D., Donghia N., Fan M. Rapid, low-cost detection of Zika virus using programmable biomolecular components. Cell. 2016;165(5):1255–1266. doi: 10.1016/j.cell.2016.04.059. [DOI] [PubMed] [Google Scholar]
- Patchsung M., Jantarug K., Pattama A., Aphicho K., Suraritdechachai S., Meesawat P., Sappakhaw K., Leelahakorn N., Ruenkam T., Wongsatit T. Clinical validation of a Cas13-based assay for the detection of SARS-CoV-2 RNA. Nat. Biomed. Eng. 2020 doi: 10.1038/s41551-020-00603-x. [DOI] [PubMed] [Google Scholar]
- Phan T. Novel coronavirus: from discovery to clinical diagnostics. Infection. Gene. Evol. 2020;79:104211. doi: 10.1016/j.meegid.2020.104211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pickar-Oliver A., Gersbach C.A. The next generation of CRISPR–Cas technologies and applications. Nat. Rev. Mol. Cell Biol. 2019;20(8):490–507. doi: 10.1038/s41580-019-0131-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qin P., Park M., Alfson K.J., Tamhankar M., Carrion R., Patterson J.L., Griffiths A., He Q., Yildiz A., Mathies R. Rapid and fully microfluidic Ebola virus detection with CRISPR-Cas13a. ACS Sens. 2019;4(4):1048–1054. doi: 10.1021/acssensors.9b00239. [DOI] [PubMed] [Google Scholar]
- Rabe B.A., Cepko C. SARS-CoV-2 detection using isothermal amplification and a rapid, inexpensive protocol for sample inactivation and purification. Proc. Natl. Acad. Sci. Unit. States Am. 2020;117(39):24450–24458. doi: 10.1073/pnas.2011221117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rabiee N., Bagherzadeh M., Ghadiri A.M., Salehi G., Fatahi Y., Dinarvand R. ZnAl nano layered double hydroxides for dual functional CRISPR/Cas9 delivery and enhanced green fluorescence protein biosensor. Sci. Rep. 2020;10(1):1–15. doi: 10.1038/s41598-020-77809-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rauch J.N., Valois E., Solley S.C., Braig F., Lach R.S., Baxter N.J., Kosik K.S., Arias C., Acosta-Alvear D., Wilson M.Z. bioRxiv; 2020. A Scalable, Easy-To-Deploy, Protocol for Cas13-Based Detection of SARS-CoV-2 Genetic Material. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ravi N., Cortade D.L., Ng E., Wang S.X. Diagnostics for SARS-CoV-2 detection: a comprehensive review of the FDA-EUA COVID-19 testing landscape. Biosens. Bioelectron. 2020;165:112454. doi: 10.1016/j.bios.2020.112454. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shao N., Han X., Song Y., Zhang P., Qin L. CRISPR-Cas12a coupled with platinum nanoreporter for visual quantification of SNVs on a volumetric bar-chart chip. Anal. Chem. 2019;91(19):12384–12391. doi: 10.1021/acs.analchem.9b02925. [DOI] [PubMed] [Google Scholar]
- Shmakov S., Smargon A., Scott D., Cox D., Pyzocha N., Yan W., Abudayyeh O.O., Gootenberg J.S., Makarova K.S., Wolf Y.I. Diversity and evolution of class 2 CRISPR–Cas systems. Nat. Rev. Microbiol. 2017;15(3):169–182. doi: 10.1038/nrmicro.2016.184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smargon A.A., Cox D.B., Pyzocha N.K., Zheng K., Slaymaker I.M., Gootenberg J.S., Abudayyeh O.A., Essletzbichler P., Shmakov S., Makarova K.S. Cas13b is a type VI-B CRISPR-associated RNA-guided RNase differentially regulated by accessory proteins Csx27 and Csx28. Mol. Cell. 2017;65(4):618–630. doi: 10.1016/j.molcel.2016.12.023. e617. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strecker J., Jones S., Koopal B., Schmid-Burgk J., Zetsche B., Gao L., Makarova K.S., Koonin E.V., Zhang F. Engineering of CRISPR-Cas12b for human genome editing. Nat. Commun. 2019;10(1):1–8. doi: 10.1038/s41467-018-08224-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Teng F., Guo L., Cui T., Wang X.-G., Xu K., Gao Q., Zhou Q., Li W. CDetection: CRISPR-Cas12b-based DNA detection with sub-attomolar sensitivity and single-base specificity. Genome Biol. 2019;20(1):1–7. doi: 10.1186/s13059-019-1742-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Dongen J.E., Berendsen J.T., Steenbergen R.D., Wolthuis R.M., Eijkel J.C., Segerink L.I. Point-of-care CRISPR/Cas nucleic acid detection: recent advances, challenges and opportunities. Biosens. Bioelectron. 2020;166:112445. doi: 10.1016/j.bios.2020.112445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang B., Wang R., Wang D., Wu J., Li J., Wang J., Liu H., Wang Y. Cas12aVDet: a CRISPR/Cas12a-based platform for rapid and visual nucleic acid detection. Anal. Chem. 2019;91(19):12156–12161. doi: 10.1021/acs.analchem.9b01526. [DOI] [PubMed] [Google Scholar]
- Wang J., Zhang C., Feng B. The rapidly advancing Class 2 CRISPR-Cas technologies: a customizable toolbox for molecular manipulations. J. Cell Mol. Med. 2020;24(6):3256–3270. doi: 10.1111/jcmm.15039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang L., Shen X., Wang T., Chen P., Qi N., Yin B.-C., Ye B.-C. A lateral flow strip combined with Cas9 nickase-triggered amplification reaction for dual food-borne pathogen detection. Biosens. Bioelectron. 2020;165:112364. doi: 10.1016/j.bios.2020.112364. [DOI] [PubMed] [Google Scholar]
- Wang M., Zhang R., Li J. CRISPR/cas systems redefine nucleic acid detection: principles and methods. Biosens. Bioelectron. 2020;165:112430. doi: 10.1016/j.bios.2020.112430. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang X., Tan L., Wang X., Liu W., Lu Y., Cheng L., Sun Z. Comparison of nasopharyngeal and oropharyngeal swabs for SARS-CoV-2 detection in 353 patients received tests with both specimens simultaneously. Int. J. Infect. Dis. 2020 doi: 10.1016/j.ijid.2020.04.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wei S., Kohl E., Djandji A., Morgan S., Whittier S., Mansukhani M., Yeh R., Alejaldre J.C., Fleck E., D'Alton M. medRxiv; 2020. Field-deployable, Rapid Diagnostic Testing of Saliva Samples for SARS-CoV-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Who 2020. https://covid19.who.int/ Available at: [Accessed January 9th]
- Williams E., Bond K., Zhang B., Putland M., Williamson D.A. Saliva as a non-invasive specimen for detection of SARS-CoV-2. J. Clin. Microbiol. 2020 doi: 10.1128/JCM.00776-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williams E., Bond K., Zhang B., Putland M., Williamson D.A. Saliva as a noninvasive specimen for detection of SARS-CoV-2. J. Clin. Microbiol. 2020;58(8) doi: 10.1128/JCM.00776-20. e00776–00720. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wozniak A., Cerda A., Ibarra-Henríquez C., Sebastian V., Armijo G., Lamig L., Miranda C., Lagos M., Solari S., Guzmán A.M., Quiroga T., Hitschfeld S., Riveras E., Ferrés M., Gutiérrez R.A., García P. A simple RNA preparation method for SARS-CoV-2 detection by RT-qPCR. Sci. Rep. 2020;10(1):16608. doi: 10.1038/s41598-020-73616-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yan W.X., Chong S., Zhang H., Makarova K.S., Koonin E.V., Cheng D.R., Scott D.A. Cas13d is a compact RNA-targeting type VI CRISPR effector positively modulated by a WYL-domain-containing accessory protein. Mol. Cell. 2018;70(2):327–339. doi: 10.1016/j.molcel.2018.02.028. e325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yan W.X., Hunnewell P., Alfonse L.E., Carte J.M., Keston-Smith E., Sothiselvam S., Garrity A.J., Chong S., Makarova K.S., Koonin E.V. Functionally diverse type V CRISPR-Cas systems. Science. 2019;363(6422):88–91. doi: 10.1126/science.aav7271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang H., Patel D.J. CasX: a new and small CRISPR gene-editing protein. Cell Res. 2019;29(5):345–346. doi: 10.1038/s41422-019-0165-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang W., Restrepo-Pérez L., Bengtson M., Heerema S.J., Birnie A., van der Torre J., Dekker C. Detection of CRISPR-dCas9 on DNA with solid-state nanopores. Nano Lett. 2018;18(10):6469–6474. doi: 10.1021/acs.nanolett.8b02968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yuan C., Tian T., Sun J., Hu M., Wang X., Xiong E., Cheng M., Bao Y., Lin W., Jiang J. Universal and naked-eye gene detection platform based on the clustered regularly interspaced short palindromic repeats/cas12a/13a system. Anal. Chem. 2020;92(5):4029–4037. doi: 10.1021/acs.analchem.9b05597. [DOI] [PubMed] [Google Scholar]
- Yuan T., Mukama O., Li Z., Chen W., Zhang Y., de Dieu Habimana J., Zhang Y., Zeng R., Nie C., He Z. A rapid and sensitive CRISPR/Cas12a based lateral flow biosensor for the detection of Epstein–Barr virus. Analyst. 2020;145(19):6388–6394. doi: 10.1039/d0an00663g. [DOI] [PubMed] [Google Scholar]
- Zetsche B., Gootenberg J.S., Abudayyeh O.O., Slaymaker I.M., Makarova K.S., Essletzbichler P., Volz S.E., Joung J., Van Der Oost J., Regev A. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell. 2015;163(3):759–771. doi: 10.1016/j.cell.2015.09.038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu N., Zhang D., Wang W., Li X., Yang B., Song J., Zhao X., Huang B., Shi W., Lu R., Niu P., Zhan F., Ma X., Wang D., Xu W., Wu G., Gao G.F., Tan W. A novel coronavirus from patients with pneumonia in China, 2019. N. Engl. J. Med. 2020;382(8):727–733. doi: 10.1056/NEJMoa2001017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zou Y., Mason M.G., Botella J.R. Evaluation and improvement of isothermal amplification methods for point-of-need plant disease diagnostics. PloS One. 2020;15(6) doi: 10.1371/journal.pone.0235216. [DOI] [PMC free article] [PubMed] [Google Scholar]