Abstract
The potent DNA-binding compound triaminotriazine-acridine conjugate (Z1) functions by targeting T:T mismatches in CTG trinucleotide repeats (TNR) that are responsible for causing neurological diseases such as myotonic dystrophy type 1 (DM1), but its binding mechanism remains unclear. We solved a crystal structure of Z1 in complex with DNA containing three consecutive CTG repeats with three T:T mismatches. Crystallographic studies revealed that direct intercalation of two Z1 molecules at both ends of the CTG repeat induces thymine base flipping and DNA backbone deformation to form a four-way junction. The core of the complex unexpectedly adopts a U-shaped head-to-head topology to form a crossover of each chain at the junction site. The crossover junction is held together by two stacked G:C pairs at the central core that rotate with respect to each other in an X-shape to form two non-planar minor groove aligned G·C·G·C tetrads. Two stacked G:C pairs on both sides of the center core are involved in the formation of pseudo-continuous duplex DNA. Four metal-mediated base pairs are observed between the N7 atoms of G and CoII, an interaction that strongly preserves the central junction site. Beyond revealing a new type of ligand-induced, four-way junction, these observations enhance our understanding of the specific supramolecular chemistry of Z1 that is essential for the formation of a non-canonical DNA superstructure. The structural features described here serve as a foundation for the design of new sequence-specific ligands targeting mismatches in the repeat-associated structures.
Graphical Abstract

INTRODUCTION
Errors during DNA replication cause misincorporation of nucleotide bases in the DNA sequence leading to mispairing or mismatches between the bases.1–2 These mismatches are ubiquitous, and their insertion into the genome often have various physiological and pathological implications.3–4 The abnormal expansion of trinucleotide repeats (TNRs) often form non-canonical structures containing mismatches sandwiched between Watson-Crick-Franklin base pairs.5 Deficiency in the repair systems may lead to elevated rates of mismatches resulting in genetic instabilities, giving rise to neurological defects.6 Small molecules that recognize mismatches can be used as probes for the recognition mismatches to detect TNR related diseases that arise from defective DNA repair mechanisms.7
Myotonic dystrophy type 1 (DM1) is caused by the expansion of CTG trinucleotide repeats in the 3’-untranslated region of myotonic dystrophy protein kinase gene.8–9 The number of expanded CTG repeats correlates well with disease inheritability, age-of-onset and severity.10 Therefore, the CTG repeat length is often considered as a disease diagnostic marker and thus used for disease onset prediction.11–12 Acridine-based small molecules have been known since the 19th century to perform various therapeutic functions including acting as anti-parasitic, antiviral, and antibacterial agents.13 Moreover, acridine-based therapeutics were investigated for their antitumor activity and ability to bind to topoisomerase II.14 Triaminotriazine-acridine conjugate (Z1) (Figure 1a) was first developed using structure-based design to selectively recognize both d(CTG) and r(CUG) sites for the treatment of DM1.15 Z1 exhibited strong binding affinity to CTG repeat sequences, inhibiting transcription by binding to the mismatches in CTG repeats. The triaminotriazine unit was proposed to recognize and form a base triplet with T:T or to flip out one of the pyrimidine rings forming a Watson-Crick-Franklin-like base-pair with thymine while stacking on the acridine.16 However, the structural basis for the recognition of Z1 ligands to CTG repeats remains unclear.
Figure 1.
(a) Chemical structure of triaminotriazine acridine conjugate (ligand Z1). (b) Overall crystal structure of d[TT(CTG)3AA] complexed to Z1 in an asymmetric unit. Each DNA chain is shown in cartoon representation. Chain A is colored in green, chain B in cyan, chain C in pink and chain D in yellow. (c) Schematic representation of the overall crystal structure. (d) The core of the complex formed by a crossover of all four chains in G5-C6-T7-G8-C9 penta-sequence. (e) Two central stacked G:C pairings form two GC tetrads labeled as GC-I and GC-II. The additional stacked G:C pairing formed by adjacent G8pC9 bases of chains A and C are denoted as GC-III and GC-IV, respectively. The hydrogen bonding between the bases is shown with a black dotted line.
In the present study, the structure of Z1 bound to a palindromic duplex containing three consecutive CTG repeats has been determined. The DNA duplex is intercepted by two Z1s at both ends that induce significant structural distortions resulting in a new type of four-way junction featuring a double U-shaped conformation that assembles in a head-to-head manner. We also solved the crystal structure of Z1-d(TTCTGCTGCTGAA/TTCTGCAGCTGAA) to investigate the structural basis behind this interaction with a non-consecutive CTG motif. The unexpected features of the crystal structures with T:T mismatch specificity clearly demonstrates the ability of Z1 to induce a non-canonical DNA superstructure resembling a four-way junction. The structure allows the interactions between the Z1 and CTG repeat-associated DNA to be elucidated with atomic-level detail. These findings further enhance our knowledge about the adoption of higher-order non-canonical DNA conformations that depend on the nature of the interacting ligand and its selective supramolecular interactions.
RESULTS
Binding of Z1 forces deformation of the T:T mismatch DNA.
To understand the structural basis behind the preference of Z1 for CTG DNA, we solved the crystal structure of a Z1 in complex with the palindromic DNA sequence, d[(5-BrU)TCTGCTGCTGAA], denoted Z1-TTT, at 1.55 Å resolution (Figure 1b and S1). The asymmetric unit shows two perpendicular antiparallel duplexes with four DNA strands, labeled as chain A, chain B, chain C and chain D (Figure 1c). The detailed analysis of complex structure reveals that the intercalation of Z1 to the end of the CTG repeat duplex induces thymine (T4) base flipping that allows for DNA backbone deformation of the central 5’-GCTGC. The backbone bends over 90o and forms a non-canonical four-way junction. These findings are also consistent with the Job-type titration plot of Z1 binding to DNA which showed a maximum at c.a. 0.67 (mole fraction of Z1), identifying the 2:1 stoichiometry of the complex in solution (Figure S2).
The core of the complex adopts a U-shaped head-to-head topology to form a crossover of each chain at the junction G5-C6-T7-G8-C9 penta-sequence (Figure 1d). The U-shaped kink is associated with an abrupt change in backbone torsion angle α by c.a. 140° between bases T4 and G5 in each chain thereby forming an unusual structure upon Z1 binding (Table S1). The central crossover core is formed by two stacked G:C pairs (denoted as GC-I, and GC-II) between 5’-G5pC6 of A and C chains and the C6pG5– 5’ of B and D chains. The adjacent G8pC9 bases of chains A and C are hydrogen bonded with C9pG8 from the B and D chains, respectively, to form two additional stacked G:C pairs (denoted as GC-III and GC-IV) on both sides (Figure 1e). In addition, the central stacked G5:C6 pairs from A, C and B, D chains are sandwiched between the T7 of each chain to form stable and continuous stacking interactions. These stacking interactions are similar to a previously reported structure where quinoxaline ring of echinomycin stabilized the continuous CpG base pairs in DNA containing a T:T mismatch.17 The thymine (T4) is present on the head of the U-turn with a χ angle in the range of −121° to −134° adopts an anti-conformation. The two backbone dihedral parameters α and ζ between T4 and G5 residue steps undergo drastic transitions (−167° to 67° for α and 52° to −95° for ζ, respectively) (Table S2). This causes T4 to be flipped in the opposite orientation of the DNA backbone with a higher degree of distortion generating an anti-conformation. Interestingly, the flipped out thymine shows no disordered electron density map and is found to be stabilized by the stacking interactions of symmetry-related 3’- terminal adenine (A13) residue in the crystal packing with an e-motif as observed previously.18
Furthermore, two arms of U-shaped duplex stack on each other to form a pseudo-continuous right-handed duplex DNA with G8 and C9 of strands A and C chains base-paired with C9 and G8 of strands B and D, respectively. Z1 intercalates through the minor groove side into the 5’-C3pT4 step (Figure 2). The binding of Z1 causes remarkable distortion in the pseudo duplex including the widening of the major (22.5 Å) and minor grooves (19.6 Å) at the central 3’-GpT step compared to the groove widths of canonical duplexes. This forces the backbone to unwind to generate additional space. The 5’ thymine (T4) is flipped out of the helix upon Z1 binding and the complementary thymine (T10) is stabilized through direct hydrogen bonding with the triaminotriazine moiety of Z1.
Figure 2.
The pseudo DNA duplex of the Z1-TTT complex structure viewed from (a) front and (b) side view. A pseudo-continuous right-handed duplex DNA is formed by the complementary base pairing between A and C chains and B and D, respectively shown as cartoon representation. The Z1 intercalates through the minor groove side into the 5’-C3pT4 step shown in blue stick and surface.
Structural detail of central crossover junction.
The palindromic pentanucleotide 5’-GCTGC sequence in the crossover junction is held together by two stacked G:C pairs at the central core (GC-I, and GC-II) and two stacked G:C pairs on both sides of the intersection (GC-III and GC-IV). From the side view, two central stacked G:C pairs are rotated c.a. 40° with X-shape to form two non-planar minor groove aligned G·C·G·C tetrads with Watson-Crick-Franklin base pairing between the complementary residues and the N2 amino of guanines hydrogen bonded to the O2 of adjacent cytosines. Two stacked G:C pairs on the side rotate 50° with respect to the central stacked G:C pairs giving 2-fold symmetry and are interconnected by metal ion-mediated interactions (Figure S3). For the first time, four CoII ions were found to mediate the base pairing between N7 on guanine (G8) in chains A, B, C and D to N7 on guanine (G5) from the chains D, C, B and A, respectively, that further stabilize the crossover junction sites. The 2Fo-Fc electron density (contoured at 1.0 σ) clearly shows that each CoII ion has octahedral coordination between two guanosine nitrogen and four water molecules (Figure 3a). This type of four stacked G:C pair arrangement in the crossover junction core gives rise to a propeller-like structure. The current structure is mostly hydrated, and several water molecules mediate significant interactions to stabilize the residues in the central crossover junction (Figure 3b). Two water molecules W206 and W56 bridge the interactions between N3 on G5 and O4 on C6 in the central core (Figure 3c), whereas W18 and W19 mediate the hydrogen-bonding interaction between the side GC pairs with the central ones (Figure 3d and 3e).
Figure 3.
(a) 2Fo-Fc map for the G5-CoII-G8 pair formed by each chain contoured at 1.0 σ. The coordinated metal ions (orange sphere) and water molecules (red sphere) appear clearly in the refined structure. Coordination and hydrogen bonds are shown by dashed lines. (b) 2Fo-Fc map for the water mediated stabilization of the central core residues. The water molecules are shown by red spheres and the hydrogen bonding is shown as dotted black lines. (c) The water W56 and W206 mediated stabilization of the central core residues. Enlarged view of W18 (d) and W19 (e) mediated hydrogen bonding between the G5 and G8 residues of each chain. The water molecules are shown by red spheres and the hydrogen bonding is shown as dotted black lines.
Structural details of the Z1 binding sites.
In the current complex structure, four Z1 molecules were observed in an asymmetric unit intercalated through the minor groove side at the end of each pseudo-duplex arm. The close-up view of the Z1 binding site is shown in Figure 4a, which showed that the acridine moiety stacks with the triaminotriazine. The stacked-intercalator inserts into the 5’-CpT/TpG-3’ step of the pseudo duplex where the acridine unit forms intramolecular stacking interactions with G11 base in the repeat sequence (Figure 4b). The intercalation of triaminotriazine causes the unpaired thymine (T4) to extrude out of the duplex and forms a complementary pairing to the other thymine (T10) through three hydrogen bond interactions. The flipping of thymine also causes the DNA backbone to bend c.a. 90° around the T4pG5 step that resulted in OP2 on guanine (G5) in such orientation that it forms a hydrogen-bonding interaction with one of the amino moieties of the triaminoazine ring (Figure 4a). In addition to these direct interactions between Z1 and DNA residues, three conserved water molecules were found to further stabilize the Z1-DNA complex (Figure 4c). LigPlot+ analysis19 showed that residue C3 and T4 of chain A, T10 and C11 of chain C, and G8 of chain D are in close van der Waals contact with the Z1 molecule in all four sites (Figure 4d). Altogether, these interactions contribute to the specific recognition of CTG repeats by Z1.
Figure 4.
Overall structure of Z1 binding site in the Z1-TTT complex. (a) Intercalation of Z1 through minor groove forms hydrogen bonding interactions with the T10 and G5 bases shown by black dotted lines. (b) The acridine moiety forms stacking interactions with the G11 base and the triaminoazine stacks on the G8 base. (c) 2Fo-Fc map for the three water mediated interactions between Z1 and DNA bases. The water molecules are shown by red spheres and the hydrogen bonding is shown as dotted black lines. (d) Hydrophobic interactions between Z1 and DNA bases as shown by the LIGPLOT+.
Comparison with the Z1–DNA complex with central Watson-Crick-Franklin base pair instead of T:T mismatch.
The stabilizing effects of Z1 binding to non-canonical duplex structures harboring one CTG repeat (AAT), two consecutive CTG repeats (ATT), two non-consecutive CTG repeats (TAT), and three consecutive CTG repeats (TTT) at a fixed Z1:DNA stoichiometry of 4:1 were tested using thermal duplex melting. We found that binding of Z1 leads to large stabilization effects on TTT (ΔTm = 16 °C) and TAT (ΔTm = 7.5 °C) oligonucleotides compared to AAT and ATT (Figure S4). Comparing the stabilization effects of Z1 on two CTG repeat DNAs, the ΔTm value of TAT increases significantly more than that of ATT (ΔTm = 4.7 °C) upon saturation with Z1 (10 μM), suggesting the binding of Z1 to neighboring CTG sites may cause steric hindrance and weaken the stabilizing effects of Z1 on DNA. These findings are consistent with the structural results showing that Z1 binds to TTT at both ends of DNA duplex and the central CTG is not involved in the interaction with Z1.
To understand whether the Z1 molecule binds at two CTG sites separated by at least one CTG site, the crystal structure of Z1 in complex with the palindromic DNA duplex harboring a central A:T base pair sequences, d(TTCTGCTGCTGAA/ TTCTGCAGCTGAA), denoted by Z1-TAT, at 1.65 Å resolution was solved for comparison (Figure S5). Each asymmetric unit of Z1-TAT complex contains two DNA duplexes bound by four Z1 molecules, in which one of the DNA duplex contains central A:T base pair and another possesses a central T:T base pair configuration (Figure S6). Upon superimposing the Z1-TTT and Z1-TAT complex structures shows an RMS deviation of 0.5 Å indicating both structures are identical globally with local differences (Figure S6a). Intriguingly, the Z1-TTT complex is more hydrated than that of Z1-TAT structure suggesting the importance of water in stabilizing the overall complexes (Figure S6b and S6c). Notable differences were found in the central T7-A7 base-pair geometries when comparing Z1-TTT to Z1-TAT. The T7:A7 base-pair shows less stacking interactions as the A7 was pushed outwards. The LigPlot+ analysis showed that the T7 of Z1-TTT complex has more van der Waals and water-mediated interactions with the flanking residues than A7 in the TAT complex (Figure S6d and S6e).
DISCUSSION
Because non-canonical structures of repetitive DNA sequences are hallmarks of many neurodegenerative diseases, small-molecule ligands targeting mismatch DNA can be a valuable tool in the diagnosis and treatment of neurological diseases.5, 20 Repetitive mismatches with polymer-like expansions and local instability can act as hotspots for small molecule recognition.21 Nakatani and coworkers showed that naphthyridine-containing compounds can recognize DNA mismatches in trinucleotide repeat DNA sequence with high affinity and specificity in vitro, and may induce repeat contractions in cells.22 Their findings led them to propose the use of naphthyridine analogs as a basis for the development of new types of therapeutic agents active against neurological disease.
Previous studies have shown that many DNA intercalators such as octahedral ruthenium complexes and actinomycin D can recognize DNA mismatches by extruding the mismatched base pairs and inducing a marked kink in the DNA structures.21, 23–24 Acridine-based compounds, including Z1, were rationally developed for targeting both d(CTG) and r(CUG) sites using structure-based design and for inhibiting both transcription of the expanded TNR of DM1 and inhibiting aberrant protein sequestration by the r(CUG) transcript.15, 25 The single stranded CTG repeat DNA can form stable, perpendicular, pseudo-duplexes when there are at least three repeating units. The present study demonstrates that the Z1 can tightly bind to T:T mismatches in CTG repeats and induce large scale structural deformation. The DNA backbone shows severe distortion in addition to thymine base flipping, resulting in a U-shaped structure (Figure 5a). Two U-shaped conformations form a marked crossover site of a -GCTGC- penta-sequence through the stacking, hydrogen bonding, and metal-mediated pairings to generate a four-way junction (4WJ).
Figure 5.
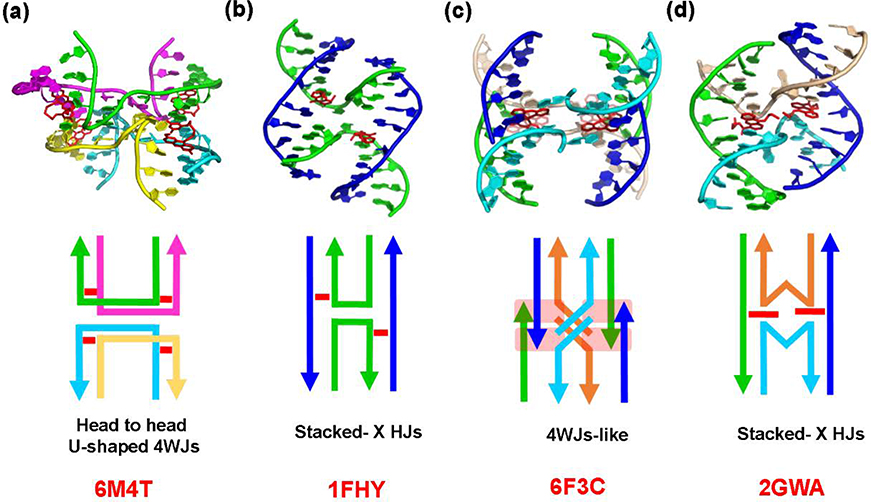
Comparison of the DNA topologies in different sequence context associated with ligand binding in four-way junction like structures. (a) Intercalation of Z1 to T:T mismatches in CTG repeat DNA resulted in U-shaped head to head four way junctions (4WJs) like topology. (b) hydroxymethyl-trimethylpsoralen (HMT) induces a Holliday junction (HJ) formation by cross-linking the thymine bases to cause stacked-X topology. (c) Binding of platinum complex [Pt(H2bapbpy)]-(PF6)2 to non-continuous pseudo duplexes causes cytosine base flipping to generate a pseudo-4WJ like crossover topology. (d) A bis(9-aminoacridine-4-carboxamide) ligand selectively recognizes the antiparallel stacked-X shaped Holliday junctions (HJ). The lower figure shows a schematic representation of each DNA topology due to different ligand binding. The respective PDB IDs for each structure are given in bold, red letters.
The structures of other DNA four-way junctions have been previously observed to form complexes with small ligands such as psoralen and platinum-based compounds, however, the U-shaped structure described here is unique (Figure 5). Thus, Eichman et al. reported a hydroxymethyl-trimethylpsoralen (HMT) that covalently binds to DNA, inducing Holliday junction (HJ) formation by cross-linking the thymine bases across complementary strands in the X-stacked structure (Figure 5b).26 Recently, a platinum complex [Pt(H2bapbpy)]-(PF6)2 (where H2bapbpy is N-(6-(6-(pyridin-2-ylamino)-pyridin-2-yl)-pyridin-2-yl)pyridin-2-amine) has been shown to bind to the short oligonucleotides that form non-continuous pseudo duplexes (Figure 5c).27 The interface between the platinum complex and the terminals of inter-duplexes induces cytosine base flipping to generate a pseudo-4WJ like crossover through stacking and hydrogen bonding as observed in the crystal packing of the complex. On the other hand, the ligands recognizing DNA superstructures can be an attractive strategy in drug design rather than targeting standard duplex DNA. For example, Brogden et al. reported 6-carbon-linked bis(9-aminoacridine-4-carboxamide) ligand that can selectively recognize the antiparallel HJ DNA (Figure 5d).28 The ligand binds across the center of the junction to form a special topology where a flipped adenine base is replaced by acridine intercalators on either side of the crossover. The detailed structural analysis of high-resolution ligand-DNA complexes can be useful to the design of new compounds with selective binding to higher-order DNA that self-assemble into superstructures such as 4WJs or HJs.
The design of new chemical compounds with distinct physical and supramolecular properties that have the ability to induce formation of different types of DNA structures is becoming an attractive area of research.29–30 There is increasing evidence that suggests the non-classical DNA structures also possess important roles in biology.31–32 Although at this time it is unfeasible to ascertain whether the current structure observed in the solid state also form in vivo. The structures reported in this study expands the diversity of unusual DNA motifs induced by ligand and thereof offer a new tool for rational design of controlled DNA superstructures or sensors. Nonetheless, we have identified a G-CoII-G mediated pairings that stabilize the crossover junction sites. Such metal ion mediated base pairs composed of natural nucleotides have been studied extensively for applications in the development of novel therapeutics and various DNA nanoarchitecture designs. For instance, a new type of non-helical DNA structure driven by a heavy metal ion has been identified that has a potential application in designing nanostructures such as DNA ‘tweezers’, ‘walkers’ or ‘gears’.33
Recently, Rousina-Webb et al. designed a metal ion selective DNA double crossover junction tile structure that has an application in designing robust, stable DNA-based material.34 Identifying the atomistic details of metal-mediated pairing will further the development of DNA nanotechnology. Inspired by the unexpected features of the current crystal structures, this work may also facilitate the development of more effective derivatives to target genomic mismatched bases. For example, modification of the triaminotriazine potion in the Z1 molecule may lead to derivatives with homo-purine mismatch specificities. In addition, the binding orientation of Z1 is such that the short linker is present in the minor groove, an appropriate position for specific interactions within the DNA duplex. The adjustment of the carbon chain length of the linker may affect the orientation of the acridine motif, whereas the modification of the acridine portion could affect the binding affinity to the DNA base pairs.
To summarize, recognition of T:T mismatches by Z1 results in the classic ligand binding-induced transformation, in this case triggering large-scale DNA deformations that ultimately lead to the formation of a non-canonical 4WJ. The ability to force DNA oligonucleotides to adopt a certain shape through sequence design and ligand binding to form supramolecular DNA may also have implications in the design of molecular devices such as ion sensors, nanowires and DNA magnets.34–35
EXPERIMENTAL SECTION
Drugs and oligonucleotides.
The triaminotriazine-acridine based ligand, Z1, was synthesized as described previously.15 Single stranded synthetic DNA oligonucleotide sequences were commercially synthesized by MDBio Inc. with purification performed by polyacrylamide gel electrophoresis (PAGE). Absorbance measurements were carried out in a quartz cuvette using a JASCO V-630 UV/VIS spectrophotometer. The concentrations of oligonucleotides were determined by Beer’s law (A = εbc, where A is the optical density at 260 nm, ε is the extinction coefficient, b is the cell path length (1 cm), and c is the DNA molar concentration). Oligomer extinction coefficients were estimated using tabulated values of monomer and dimer extinction coefficients with reasonable assumptions.36
Melting temperature.
The melting temperature (Tm) of the DNA oligonucleotides were analyzed using a JASCO UV/VIS spectrophotometer with 1 cm path length quartz cuvettes as previously described.17 Initially, the DNA oligomers (Table S3) were annealed by heating at 90 °C for 5 min to denature the strands and cooling on ice for 30 min. To ensure formation of the Z1-DNA complexes, 3 μM DNA oligomers were incubated with 12 μM Z1 in the presence of 12 μM CoII in MOPS buffer (20 mM MOPS, 300mM NaCl and 1 mM Na2EDTA, pH 6.5) at 4 °C overnight. The melting temperature curves were obtained by increasing the temperature from 4 °C to 95 °C at a rate of 1 °C/min and recorded every 0.5 min at 260 nm. The Tm values were determined from the observed curves using a polynomial curve fitting by Varian Cary Win UV Thermal application software (Ver. 3.0).
Job plot analysis.
As the acridine moiety of Z1 possesses fluorescent properties, Job-type titration spectra were used to monitor the interactions between CTG DNA and Z1. The binding of Z1 to CTG DNA causes enhanced fluorescence emission exhibiting a maximum near 525 nm and a fluorescence intensity enhancement near 420 nm in the excitation spectrum with 525 nm emission monitoring. Samples of varying molar ratios of Z1:DNA were prepared with the total concentration held constant at 2 μM and incubated at 25 °C for 24 h. All spectra were recorded at 25 °C in 20 mM MOPS (pH 6.5), 300 mM NaCl and 1 mM EDTA. The spectra were measured at 25 °C with a JASCO model FP-4500 spectrofluorimeter.37 Experiments were performed using a 3 × 3 mm quartz cell cuvette with a 1 cm path length. Plots of fluorescence intensity versus molar ratio were obtained to determine binding stoichiometry.
Crystallization.
To obtain the Z1-TTT crystals, 0.4 mM single-stranded d[BrUT(CTG)3AA] oligomer was pre-incubated with 0.7 mM ligand Z1 at 4 °C for 72 hours. Yellow-rhombic shaped crystals were grown after two weeks in a 3 μl drop containing 50 mM MES (pH = 6.5), 200 mM potassium chloride, 20 mM cobalt(II) chloride and 35% 2-methyl-2,4-pentanediol (MPD), equilibrated against 500 μl of 35% MPD using the sitting-drop vapor diffusion method. The Z1-TAT complex crystals were also obtained in similar conditions as described above where 0.2 mM d[TT(CTG)3AA] was mixed with an equimolar concentration of d(TTCTGCAGCTGAA) oligomer.
X-ray data collection, phasing, and structure refinement.
X-ray diffraction data of the Z1-TTT and Z1-TAT complexes were collected at beamlines TPS 05A and BL15A1 of the National Synchrotron Radiation Research Center (Hsinchu, Taiwan), respectively. The diffraction data for the Z1-TTT crystal in space group P21 with unit-cell parameters a = 57.2, b = 62.3, c = 29.7Å was collected at 100 K using the MX300HE detector at beamline 15A1. Single-wavelength anomalous diffraction (SAD) data was collected at a resolution of 2.1 Å from single peak wavelength using CoII as the anomalous scattering atom. The data with the anomalous signal was indexed, integrated, and scaled by HKL-2000, followed by CoII substructure localization using SHELX C/D/E.38 The resulting well-defined SAD electron density map was used to build initial models using the programs MIFit (Ver. 2010.10) and WinCoot (Ver. 0.8.4).39 The best built modeled structure of the Z1-TTT complex was used as a template to solve the phases of the Z1-TTT complex at a higher resolution of 1.55 Å. The structural refinements were performed using the PHENIX package (Ver. 1.14–3260).40 In addition, the diffraction data for the Z1-TAT crystal in space group P21 with unit-cell parameters a = 57.63, b = 62.09, c = 30.75 Å, was collected at 100 K on the MX300HS detector at beamline TPS05A. The phase for the Z1-TAT complex structure was determined by molecular replacement with Phaser MR in the PHENIX package suite (Ver. 1.14–3260) using the partial structure of the Z1-TTT complex as a template. The crystallographic and refinement statistics of these complexes are listed in (Table S4).
Supplementary Material
ACKNOWLEDGMENTS
The authors gratefully acknowledge support by the Ministry of Science and Technology, Taiwan (MOST 106-2628-M-005-001-MY3 to M.-H.H.) and the National Institutes of Health (R01AR069645 to S.C.Z.). L.D.H. is a member of the NIH Chemistry-Biology Interface Training Grant (NRSA 1-T-32-GM070421). The authors also thank the National Synchrotron Radiation Research Center (Taiwan) for X-ray diffraction data collection. We also thank Sarah E. Bonson for her useful suggestions on the manuscript.
Footnotes
Supporting Information
Job plot and melting temperatures for Z1-DNA complex and additional details of x-ray analysis including electron density maps, structural comparisons and data tables, and crystallographic and refinement statistics.
REFERENCES
- (1).Li G-M Mechanisms and functions of DNA mismatch repair. Cell Res. 2008, 18, 85–98. [DOI] [PubMed] [Google Scholar]
- (2).Kunkel TA; Erie DA Eukaryotic Mismatch Repair in Relation to DNA Replication. Ann. Rev. Genetics 2015, 49, 291–313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (3).Li GM Mechanisms and functions of DNA mismatch repair. Cell Res. 2008, 18, 85–98. [DOI] [PubMed] [Google Scholar]
- (4).Rossetti G; Dans PD; Gomez-Pinto I; Ivani I; Gonzalez C; Orozco M The structural impact of DNA mismatches. Nucl. Acids Res. 2015, 43, 4309–4321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (5).Tseng W-H; Chang C.-k.; Wu P-C; Hu N-J; Lee G-H; Tzeng C-C; Neidle S; Hou M-H Induced-Fit Recognition of CCG Trinucleotide Repeats by a Nickel–Chromomycin Complex Resulting in Large-Scale DNA Deformation. Angew. Chem. Int. Ed. 2017, 56, 8761–8765. [DOI] [PubMed] [Google Scholar]
- (6).Subba Rao K Mechanisms of disease: DNA repair defects and neurological disease. Nat. Clin. Pract. Neurol. 2007, 3, 162–172. [DOI] [PubMed] [Google Scholar]
- (7).Satange R; Chang CK; Hou MH A survey of recent unusual high-resolution DNA structures provoked by mismatches, repeats and ligand binding. Nucl. Acids Res. 2018, 46, 6416–6434. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (8).Mirkin SM Expandable DNA repeats and human disease. Nature 2007, 447, 932–940. [DOI] [PubMed] [Google Scholar]
- (9).Thornton CA; Wang E; Carrell EM Myotonic dystrophy: approach to therapy. Curr. Opin. Genet. Dev. 2017, 44, 135–140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (10).Du J; Campau E; Soragni E; Jespersen C; Gottesfeld JM Length-dependent CTG·CAG triplet-repeat expansion in myotonic dystrophy patient-derived induced pluripotent stem cells. Human Molec. Genetics 2013, 22, 5276–5287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (11).Mor-Cohen R; Magal N; Gadoth N; Shohat T; Shohat M Correlation between the incidence of myotonic dystrophy in different groups in Israel and the number of CTG trinucleotide repeats in the myotonin gene. Am. J. Med. Genetics 1997, 71, 156–159. [DOI] [PubMed] [Google Scholar]
- (12).Budworth H; McMurray CT A brief history of triplet repeat diseases. Methods Mol. Biol. 2013, 1010, 3–17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (13).Chen R; Huo L; Jaiswal Y; Huang J; Zhong Z; Zhong J; Williams L; Xia X; Liang Y; Yan Z Design, Synthesis, Antimicrobial, and Anticancer Activities of Acridine Thiosemicarbazides Derivatives. Molecules 2019, 24, 2065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (14).Murphy MB; Mercer SL; Deweese JE, Chapter Five - Inhibitors and Poisons of Mammalian Type II Topoisomerases In Adv. Molec. Tox, Fishbein JC; Heilman JM, Eds. Elsevier: 2017; Vol. 11, pp 203–240. [Google Scholar]
- (15).Arambula JF; Ramisetty SR; Baranger AM; Zimmerman SC A simple ligand that selectively targets CUG trinucleotide repeats and inhibits MBNL protein binding. Proc. Natl. Acad. Sci. U.S.A. 2009, 106, 16068–16073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (16).Serrano JF; Lee J; Daniel Curet L; Hagler LD; Bonson SE; Schuster EJ; Zimmerman SC Development of novel macrocyclic small molecules that target CTG trinucleotide repeats. Bioorg. Med. Chem. 2019, 27, 2978–2984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (17).Wu P-C; Tzeng S-L; Chang C.-k.; Kao Y-F; Waring MJ; Hou M-H Cooperative recognition of T:T mismatch by echinomycin causes structural distortions in DNA duplex. Nucl. Acids Res. 2018, 46, 7396–7404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (18).Pan F; Zhang Y; Man VH; Roland C; Sagui C E-motif formed by extrahelical cytosine bases in DNA homoduplexes of trinucleotide and hexanucleotide repeats. Nucl. Acids Res. 2017, 46, 942–955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (19).Wallace AC; Laskowski RA; Thornton JM LIGPLOT: a program to generate schematic diagrams of protein-ligand interactions. Prot. Engin. 1995, 8, 127–134. [DOI] [PubMed] [Google Scholar]
- (20).Granzhan A; Kotera N; Teulade-Fichou MP Finding needles in a basestack: recognition of mismatched base pairs in DNA by small molecules. Chem. Soc. Rev. 2014, 43, 3630–3665. [DOI] [PubMed] [Google Scholar]
- (21).Satange R; Chuang C-Y; Neidle S; Hou M-H Polymorphic G:G mismatches act as hotspots for inducing right-handed Z DNA by DNA intercalation. Nucl. Acids Res. 2019, 47, 8899–8912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (22).Nakamori M; Panigrahi GB; Lanni S; Gall-Duncan T; Hayakawa H; Tanaka H; Luo J; Otabe T; Li J; Sakata A; Caron MC; Joshi N; Prasolava T; Chiang K; Masson JY; Wold MS; Wang X; Lee M; Huddleston J; Munson KM; Davidson S; Layeghifard M; Edward LM; Gallon R; Santibanez-Koref M; Murata A; Takahashi MP; Eichler EE; Shlien A; Nakatani K; Mochizuki H; Pearson CE A slipped-CAG DNA-binding small molecule induces trinucleotide-repeat contractions in vivo. Nature Genetics 2020, 52, 146–159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (23).Song H; Kaiser JT; Barton JK Crystal structure of Delta-[Ru(bpy)(2)dppz](2)(+) bound to mismatched DNA reveals side-by-side metalloinsertion and intercalation. Nature Chem. 2012, 4, 615–620. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (24).Lo YS; Tseng WH; Chuang CY; Hou MH The structural basis of actinomycin D-binding induces nucleotide flipping out, a sharp bend and a left-handed twist in CGG triplet repeats. Nucl. Acids Res. 2013, 41, 4284–4294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (25).Nguyen L; Luu LM; Peng S; Serrano JF; Chan HY; Zimmerman SC Rationally designed small molecules that target both the DNA and RNA causing myotonic dystrophy type 1. J. Am. Chem. Soc. 2015, 137, 14180–14189. [DOI] [PubMed] [Google Scholar]
- (26).Eichman BF; Mooers BHM; Alberti M; Hearst JE; Ho PS The crystal structures of psoralen cross-linked DNAs: drug-dependent formation of Holliday junctions11Edited by I. Tinoco. J. Molec. Biol. 2001, 308, 15–26. [DOI] [PubMed] [Google Scholar]
- (27).van Rixel VHS; Busemann A; Wissingh MF; Hopkins SL; Siewert B; van de Griend C; Siegler MA; Marzo T; Papi F; Ferraroni M; Gratteri P; Bazzicalupi C; Messori L; Bonnet S Induction of a Four-Way Junction Structure in the DNA Palindromic Hexanucleotide 5′-d(CGTACG)-3′ by a Mononuclear Platinum Complex. Angew. Chem. Int. Ed. 2019, 58, 9378–9382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (28).Brogden AL; Hopcroft NH; Searcey M; Cardin CJ Ligand bridging of the DNA Holliday junction: molecular recognition of a stacked-X four-way junction by a small molecule. Angew. Chem. Int. Ed. Engl. 2007, 46, 3850–3854. [DOI] [PubMed] [Google Scholar]
- (29).Takahara PM; Rosenzweig AC; Frederick CA; Lippard SJ Crystal structure of double-stranded DNA containing the major adduct of the anticancer drug cisplatin. Nature 1995, 377, 649–652. [DOI] [PubMed] [Google Scholar]
- (30).Song H; Kaiser JT; Barton JK Crystal structure of Δ-[Ru(bpy)2dppz]2+ bound to mismatched DNA reveals side-by-side metalloinsertion and intercalation. Nature Chem. 2012, 4, 615–620. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (31).Maizels N Dynamic roles for G4 DNA in the biology of eukaryotic cells. Nature Structural & Molec. Biol. 2006, 13, 1055–1059. [DOI] [PubMed] [Google Scholar]
- (32).Chu B; Zhang D; Hwang W; Paukstelis PJ Crystal Structure of a Tetrameric DNA Fold-Back Quadruplex. J. Am. Chem. Soc. 2018, 140, 16291–16298. [DOI] [PubMed] [Google Scholar]
- (33).Liu H; Shen F; Haruehanroengra P; Yao Q; Cheng Y; Chen Y; Yang C; Zhang J; Wu B; Luo Q; Cui R; Li J; Ma J; Sheng J; Gan J A DNA Structure Containing AgI-Mediated G:G and C:C Base Pairs. Angew. Chem. Int. Ed. 2017, 56, 9430–9434. [DOI] [PubMed] [Google Scholar]
- (34).Rousina-Webb A; Lachance-Brais C; Rizzuto FJ; Askari MS; Sleiman HF Transition-Metal-Functionalized DNA Double-Crossover Tiles: Enhanced Stability and Chirality Transfer to Metal Centers. Angew. Chem. Int. Ed. 2020, 59, 4091–4098. [DOI] [PubMed] [Google Scholar]
- (35).Hong F; Jiang S; Lan X; Narayanan RP; Šulc P; Zhang F; Liu Y; Yan H Layered-Crossover Tiles with Precisely Tunable Angles for 2D and 3D DNA Crystal Engineering. J. Am. Chem. Soc. 2018, 140, 14670–14676. [DOI] [PubMed] [Google Scholar]
- (36).Murphy JH; Trapane TL Concentration and extinction coefficient determination for oligonucleotides and analogs using a general phosphate analysis. Anal. Biochem. 1996, 240, 273–282. [DOI] [PubMed] [Google Scholar]
- (37).Hung H-C; Liu C-L; Hsu JTA; Horng J-T; Fang M-Y; Wu S-Y; Ueng S-H; Wang M-Y; Yaw C-W; Hou M-H Development of an Anti-Influenza Drug Screening Assay Targeting Nucleoproteins with Tryptophan Fluorescence Quenching. Anal. Chem. 2012, 84, 6391–6399. [DOI] [PubMed] [Google Scholar]
- (38).Sheldrick GM A short history of SHELX. Acta Crystallogr. Sect. A, Found. Crystallogr. 2008, 64, 112–122. [DOI] [PubMed] [Google Scholar]
- (39).Emsley P; Lohkamp B; Scott WG; Cowtan K Features and development of Coot. Acta Crystallogr. Sect. D, Biol. Crystallogr. 2010, 66, 486–501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (40).Liebschner D; Afonine PV; Baker ML; Bunkoczi G; Chen VB; Croll TI; Hintze B; Hung LW; Jain S; McCoy AJ; Moriarty NW; Oeffner RD; Poon BK; Prisant MG; Read RJ; Richardson JS; Richardson DC; Sammito MD; Sobolev OV; Stockwell DH; Terwilliger TC; Urzhumtsev AG; Videau LL; Williams CJ; Adams PD Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. Sect. D, Struct. Biol. 2019, 75, 861–877. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.