Abstract
The chromosomal DNA of bacteria is folded into a compact body called the nucleoid, which is composed essentially of DNA (∼80%), RNA (∼10%), and a number of different proteins (∼10%). These nucleoid proteins act as regulators of gene expression and influence the organization of the nucleoid by bridging, bending, or wrapping the DNA. These so-called architectural properties of nucleoid proteins are still poorly understood. For example, the reason why certain proteins compact the DNA coil in certain environments but make the DNA more rigid instead in other environments is the subject of ongoing debates. Here, we address the question of the impact of the self-association of nucleoid proteins on their architectural properties and try to determine whether differences in self-association are sufficient to induce large changes in the organization of the DNA coil. More specifically, we developed two coarse-grained models of proteins, which interact identically with the DNA but self-associate differently by forming either clusters or filaments in the absence of the DNA. We showed through Brownian dynamics simulations that self-association of the proteins dramatically increases their ability to shape the DNA coil. Moreover, we observed that cluster-forming proteins significantly compact the DNA coil (similar to the DNA-bridging mode of H-NS proteins), whereas filament-forming proteins significantly increase the stiffness of the DNA chain instead (similar to the DNA-stiffening mode of H-NS proteins). This work consequently suggests that the knowledge of the DNA-binding properties of the proteins is in itself not sufficient to understand their architectural properties. Rather, their self-association properties must also be investigated in detail because they might actually drive the formation of different DNA-protein complexes.
Significance
Many nucleoid proteins have two interrelated functions: they act as regulators of gene expression and shape the nucleoid by bridging, bending, or wrapping the DNA. It is usually accepted that the way these proteins bind to the DNA dictates the way they shape the DNA coil. For example, proteins that bridge distal DNA segments are expected to compact the nucleoid. Through coarse-grained modeling and Brownian dynamics simulations, we identify here yet another key parameter and show that protein self-association impacts very profoundly their architectural properties. Two proteins that interact similarly with the DNA but oligomerize differently may have strikingly different architectural properties, with one protein compacting the DNA coil and the other one instead making the DNA molecule more rigid.
Introduction
Bacteria lack a nucleus, but their chromosomal DNA is nevertheless folded into a compact body called the nucleoid, which is markedly different from the rest of the cytoplasm. The nucleoid is composed essentially of DNA (∼80%), RNA (∼10%), and a number of different proteins (∼10%) (1,2). These proteins act as regulators of gene expression (3, 4, 5) and influence the organization of the nucleoid by bridging, bending, or wrapping the DNA (5, 6, 7, 8). There are at least 12 different species of nucleoid proteins (9), among which HU (10), IHF (11), H-NS (12), Fis (13), and Lrp (14) have been extensively studied. It has been shown that the abundance of many of the nucleoid proteins varies dramatically in response to changes in the growth rate of the cell (15). Their occupancy landscape in the nucleoid (16) and along the genome (17) has also been investigated.
The mechanisms by which nucleoid proteins shape the DNA are still poorly understood. This is due, in part, to the fact that architectural properties are specific to each protein. For example, H-NS, ParB, and SMC form bridges between two DNA segments, but these bridges are qualitatively different and affect chromosome organization and gene regulation in contrasting ways (18). Moreover, several proteins exhibit dual architectural properties depending on several factors, such as the concentration of proteins and the DNA binding sequence (8). For example, HU is essentially known for its DNA-bending capabilities and Lrp for its DNA-bridging capabilities, but both of them are also able to wrap the DNA (8). Finally, subtle variations of the cytosol may alter dramatically the architectural properties of certain proteins. For example, an increase in the concentration of divalent cations in the cytosol causes H-NS to switch from the DNA-stiffening mode (characterized by rigid DNA/H-NS complexes) to the DNA-bridging mode (characterized by more compact DNA coils) (19).
Through the development of coarse-grained models and Brownian dynamics simulations, we recently showed that the switch of H-NS proteins from the DNA-stiffening to the DNA-bridging mode may be due to the fact that an increase in the concentration of multivalent cations provokes an increase in the screening of electrostatic charges along the DNA backbone, which leads in turn to a decrease in the strength of DNA-protein interactions compared with protein-protein interactions (20). As a consequence, for concentrations of multivalent cations smaller than a certain threshold, proteins form filaments that stretch along the DNA molecule. In contrast, for larger concentrations of multivalent cations, proteins form clusters that connect genomically distant DNA sites (20). We argued that these two types of DNA-protein complexes may correspond to the DNA-stiffening and DNA-bridging modes of H-NS, respectively. Unfortunately, the model was not precise enough for protein filaments to increase the effective stiffness of the DNA chain and for protein clusters to significantly reduce the radius of the DNA coil. Moreover, this first study left an important question unanswered, namely, to what extent do the self-association properties of proteins influence their nucleoid architectural properties? In the model proposed in (20), proteins self-associate in the form of three-dimensional clusters, and the final conformation of DNA-protein complexes is actually driven by the relative strength of DNA-protein interactions compared with protein-protein ones. The question we address in this work is different in the sense that we consider two proteins that interact identically with the DNA chain but self-associate differently, and we want to determine whether such a difference in self-association properties is sufficient to induce large changes in the organization of the DNA coil. In addition to its obvious relevance in prokaryotes, this question may also be of fundamental importance in eukaryotes because it has recently been shown that slightly different variants of histone proteins form central tetramers with rather different properties, which may potentially influence drastically nucleosome assembly and disassembly (21). To answer this question, we developed two new, to our knowledge, coarse-grained models of proteins that interact identically with the DNA chain but self-associate differently. In the absence of DNA, model I proteins spontaneously form clusters, whereas model II proteins form filaments. We showed through Brownian dynamics simulations that self-association of the protein chains dramatically increases their ability to shape the DNA coil. Moreover, we observed that model I proteins significantly compact the DNA coil (similar to the DNA-bridging mode of H-NS), whereas model II proteins instead significantly increase the stiffness of the DNA chain (similar to the DNA-stiffening mode of H-NS). This work consequently suggests that the knowledge of the DNA-binding properties of given proteins is in itself not sufficient to understand their architectural properties. Rather, their self-association properties must also be investigated in detail because they might actually drive the formation of different DNA-protein complexes.
Methods
Model
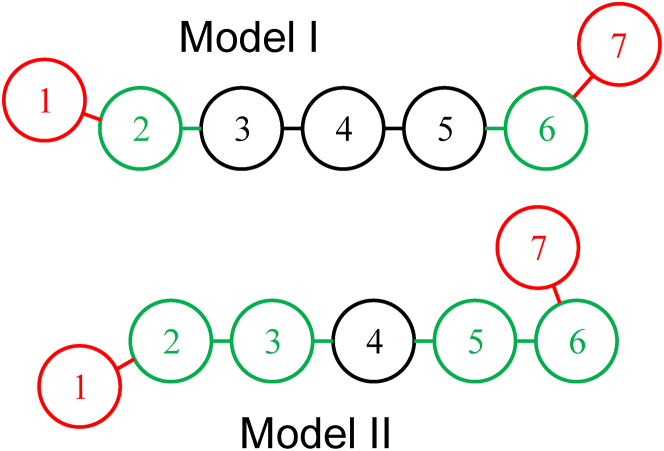
The two coarse-grained bead-and-spring models developed in the course of this study are described in detail in Model and Simulations in the Supporting Materials and Methods. In brief, they consist of a long DNA chain and 200 short protein chains enclosed in a confinement sphere. Each chain is composed of beads of equal size connected by springs. Concentrations of nucleotides and proteins are of the same order of magnitude as in vivo ones. For DNA, each bead represents 7.5 basepairs (bp), and the chain contains 2880 beads, equivalent to 21,600 bp, as in (20,22,23). Each protein chain contains seven beads with index m (1 ≤ m ≤ 7), where terminal beads m = 1 and 7 represent the two DNA-binding sites of each protein, whereas beads m = 2 and 6 (for model I) or m = 2, 3, 5 and 6 (for model II) represent the isomerization sites of the protein (Fig. 1). The overall potential energy of the system (Eq. S24) is the sum of four terms, which describe the internal energy of the DNA chain (Eq. S1), the internal energy of protein chains (Eq. S8), DNA-protein interactions (Eq. S14), and protein-protein interactions (Eq. S18). The first three terms are common to model I and model II. In particular, for both models, the two terminal beads of each protein chain (m = 1 and m = 7) can rotate without energy penalty around beads m = 2 and m = 6, respectively (Eq. S10), and can bind to the DNA chain with a maximal binding energy of −7.8 kBT (Eqs. S15 and S16; Fig. S2 a). Because of the free rotation of terminal beads, protein chains are significantly less rigid than the DNA chain, as is usually the case in vivo. Moreover, the DNA-protein binding energy is comparable to experimentally determined values for complexes of DNA and H-NS (∼−11.0 kBT) (24). Model I and II protein chains differ only in their isomerization properties. Indeed, for model I, beads m = 2 and m = 6 of one protein chain may bind to beads m = 2 and m = 6 of other protein chains, whereas for model II, beads m = 2 and m = 6 of one protein chain may bind to beads m = 3 and m = 5 of other protein chains (Eqs. S19, S21, and S23; Figs. 1 and S2 b). As a result, model I protein chains spontaneously form clusters, whereas model II protein chains form filaments. The binding interaction between two protein isomerization beads is modeled by a Lennard-Jones 3-6 potential of depth εLJ (Eq. S19). The isomerization binding energy for model I is −εLJ, whereas it varies with a slope close to −2εLJ for model II (Fig. S3). For comparison, remember that the experimentally determined value of the enthalpy change upon forming a complex between two H-NS dimers is −10.2 kBT (25).
Figure 1.
Diagrams of protein chains for models I and II. Index m is indicated for each bead. Red circles represent DNA-binding beads (index m = 1 and 7), which rotate freely around beads with index m = 2 and 6, respectively. Green circles represent isomerization beads (index m = 2 and 6 for model I and index m = 2, 3, 5, and 6 for model II). In model I, beads m = 2 and m = 6 of one protein chain may bind to beads m = 2 and m = 6 of other protein chains. In model II, beads m = 2 and m = 6 of one protein chain may bind to beads m = 3 and m = 5 of other protein chains. All other features are common to the two models. Note that the two chains shown in the figure have minimal internal energy. To see this figure in color, go online.
Simulations
The dynamics of the models was investigated by integrating numerically Langevin equations of motion with kinetic energy terms dropped and time steps of 1.0 ps. Temperature T was assumed to be 298 K throughout the study. The value of the Debye length used in the simulations (rD = 1.07 nm) corresponds to a concentration of monovalent salt of 100 mM, which is the value that is generally assumed for the cytoplasm of bacterial cells. After each integration step, the position of the center of the confining sphere was slightly adjusted so as to coincide with the center of mass of the DNA molecule so that compact DNA-protein complexes do not stick to the wall of the confinement sphere and results are affected as little as possible by the interactions with the wall (26). Simulations were run for both models and values of εLJ ranging from 4 kBT to 12 kBT to check the impact of self-association of the protein chains on the equilibrium properties of the system. The upper limit was fixed to 12 kBT because the probability for model II proteins to form clusters instead of filaments becomes non-negligible for this value of εLJ and increases rapidly for larger values.
Results
Self-association of model I and II protein chains
Model I and II protein chains interact identically with the DNA chain but self-associate differently. The goal of this work is to determine whether the difference in self-association might result in different architectural properties of the proteins, that is, in DNA-protein complexes with substantially different conformations. A preliminary step consists in characterizing in some detail the complexes that protein chains form spontaneously in the absence of the DNA chain. To this end, 200 protein chains were introduced at random nonoverlapping positions in the confinement sphere, and the system was allowed to equilibrate for values of εLJ (the depth of the Lennard-Jones 3-6 potential that governs protein-protein interactions) ranging from 4 kBT to 12 kBT. Typical equilibration times range from 1 to 50 ms, depending on the model and the value of εLJ.
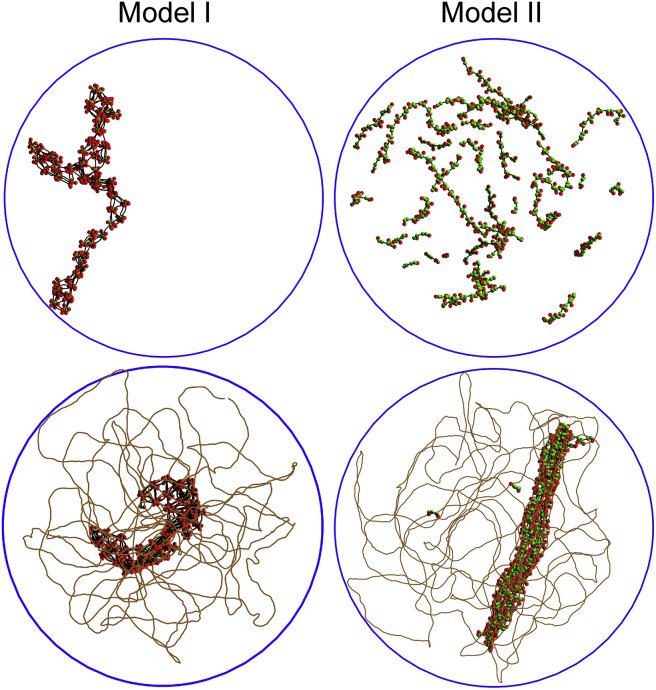
For the lowest values of εLJ, thermal noise is strong enough to prevent association of the protein chains, as was also the case for a previous model of H-NS-mediated compaction of bacterial DNA (27,28). In contrast, for larger values of εLJ, model I proteins form clusters, whereas model II proteins form filaments. Representative snapshots of equilibrated conformations are shown in the top row of Fig. 2. Evolution of protein complexes with increasing values of εLJ may be characterized by plotting q(s), the probability distribution for a protein chain to bind to s other protein chains. For this purpose, it was considered that two protein chains are bound if the interaction between at least two of their isomerization beads is attractive and of magnitude larger than 3 kBT. The choice of the 3 kBT threshold is somewhat arbitrary, but the principal features of the distributions shown in Fig. S4 do not depend critically thereon. For model I (left column of Fig. S4), protein chains do not associate significantly up to εLJ = 6 kBT, whereas for εLJ ≥ 7 kBT, each protein chain binds on average to four or five other protein chains, which results in clusters like the ones shown in the top left vignette of Fig. 2. For model II (right column of Fig. S4), protein chains do not associate significantly up to εLJ = 8 kBT, whereas for εLJ ≥ 9 kBT, each protein chain binds at maximum to two other protein chains, which results in filaments like the ones shown in the top right vignette of Fig. 2.
Figure 2.
Representative snapshots extracted from simulations with 200 protein chains and εLJ = 11 kBT for model I (left column) and model II (right column), either without the DNA chain (top row) or with the DNA chain (bottom row). DNA-binding protein beads are shown in red, isomerization beads are shown in green, and other protein beads are not shown. The lines joining the centers of protein beads are shown in black. The line joining the centers of DNA beads is shown in brown (DNA beads are not shown). The blue circle is the trace of the confinement sphere. To see this figure in color, go online.
Complexes of DNA and protein chains
Let us now consider complexes formed by the DNA chain and model I and II protein chains. These complexes were obtained by first allowing the DNA chain to equilibrate inside the confinement sphere. The 200 protein chains were then introduced at random nonoverlapping positions in the confinement sphere, and the system was allowed to equilibrate again for values of εLJ ranging from 4 kBT to 12 kBT. Typical equilibration times range from 1 to 20 ms, depending on the model and the value of εLJ. For the lowest values of εLJ, equilibrated conformations display few protein-protein contacts and a limited number of DNA-protein contacts, whereas huge DNA-protein complexes are observed for larger values of εLJ. Representative snapshots of equilibrated conformations obtained with large values of εLJ are shown in the bottom row of Fig. 2. For model I (bottom left vignette of Fig. 2), the DNA chain wraps around the protein clusters, which are quite similar to those obtained without the DNA. In contrast, for model II (bottom right vignette of Fig. 2), the protein filaments and the DNA chain form thick bundles in which they align parallel to each other.
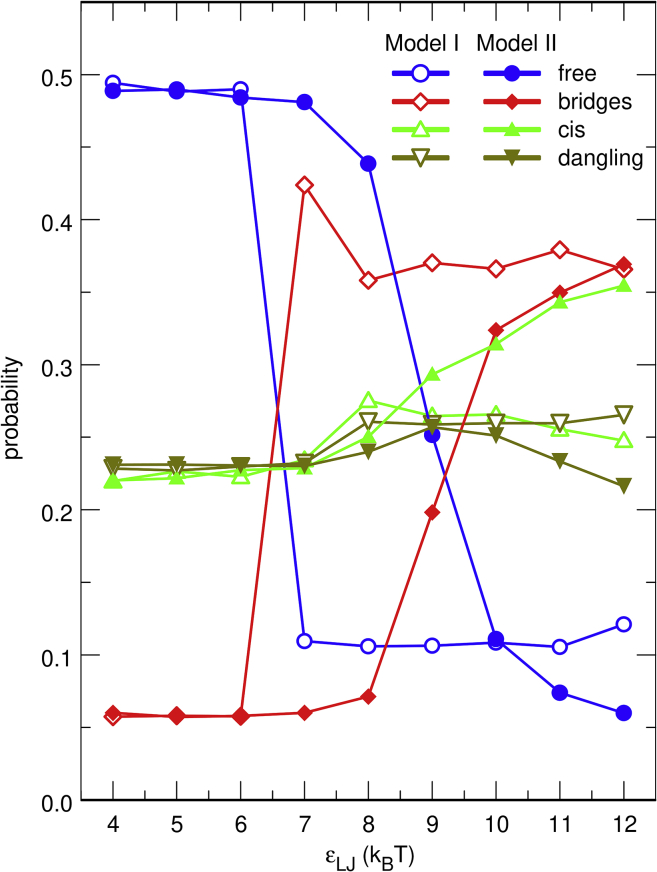
As schematized in Fig. S5, protein chains in thermodynamic equilibrium with a DNA chain can be described either as free (no contact with the DNA chain), dangling (only one extremity of the protein chain binds to the DNA chain), cis-bound (the two extremities of the protein chain bind to genomically close DNA beads), or bridging (the two extremities of the protein chain bind to genomically distant DNA beads). The evolution with increasing values of εLJ of the average fraction of the four types of protein chains is shown in Fig. 3 for model I (open symbols) and II (solid symbols). Because the two models of protein interact similarly with the DNA chain, the curves for model I and II remain superposed as long as self-association of protein chains remains negligible, that is, up to εLJ = 6 kBT. In this regime, ∼50% of the protein chains are free, ∼22% are dangling, ∼22% are cis-bound, and only ∼6% are bridging the DNA chain. However, the onset of protein self-association is accompanied in both models by a strong decrease in the number of free proteins (∼10% at εLJ = 12 kBT), which is compensated by a strong increase in the number of bridging proteins (∼35% at εLJ = 12 kBT). Evolution with increasing values of εLJ is sharper for model I than for model II.
Figure 3.
Plot, as a function of εLJ, of the average fraction of free (circles), bridging (diamonds), cis-bound (triangles), and dangling (upside-down triangles) protein chains for models I (open symbols) and II (solid symbols). Each set of four open or solid symbols with the same value of εLJ was obtained from a single simulation with the DNA chain and 200 protein chains by averaging the relevant quantity over time intervals of at least 2.5 ms after equilibration. To see this figure in color, go online.
The two models differ in that the fraction of cis-bound protein chains increases up to ∼35% at εLJ = 12 kBT for model II, whereas it remains nearly constant at ∼25% for model I. However, this discrepancy merely reflects different organizations of DNA-protein clusters because the plot, as a function of εLJ, of the average fraction of protein chains that belong to clusters that bridge genomically distant DNA beads indicates that the onset of protein self-association is accompanied in both models by an increase in the fraction of such protein chains from ∼6 to 100% (see Fig. S6).
Finally, it may be worth noting that the maximal number of bridging proteins observed for model I and εLJ = 7 kBT (∼42%) is due to the fact that for this value of εLJ, which is the smallest one that leads to protein self-association for model I, the protein chains still display some ability to escape and rearrange after binding to an existing assembly. As a result, for εLJ = 7 kBT, protein chains form a single regular (nearly two-dimensional) sheet, which apparently maximizes the number of bridges, whereas they assemble in more irregular three-dimensional clusters for larger values of εLJ. This can be checked in Fig. S7, which shows typical conformations obtained for model I and εLJ = 6 kBT, 7 kBT, and 8 kBT. For model I and εLJ ≥ 8 kBT, the number of bridges remains constant within computational uncertainties.
The plots of the probability distribution for a protein chain to bind to s other protein chains, q(s), are shown in Fig. S8 for equilibrated DNA-protein systems. These plots show that the presence of the DNA chain does not significantly alter the self-association of protein chains. Indeed, model I protein chains bind on average to four or five other protein chains starting from εLJ = 7 kBT (left column of Fig. S8), whereas model II protein chains bind at maximum to two other protein chains starting from εLJ = 9 kBT (right column of Fig. S8), as is also the case without the DNA (Fig. S4). The plots of p(s), the probability distribution for a DNA-binding protein bead to bind to s DNA beads, are also shown in Fig. S8. As for q(s), it was considered that a protein bead and a DNA bead are bound if their interaction is attractive and of magnitude larger than 3 kBT. A first conclusion concerning the impact of the self-association of protein chains on DNA-protein complexes can be drawn from the comparison of the plots of p(s) and q(s) in Fig. S8. This figure reveals that the binding of protein chains to the DNA chain is boosted by protein self-association. Indeed, for both models, only ∼30% of the DNA-binding protein beads bind to a DNA bead (p(0) ≈ 0.7) for values of εLJ corresponding to weak protein self-association, that is, up to εLJ = 6 kBT for model I and εLJ = 8 kBT for model II. In contrast, ∼70% of the DNA-binding protein beads bind to at least one DNA bead (p(0) ≈ 0.3) for larger values of εLJ, for which strong protein self-association is observed.
Architectural properties of model I and II protein chains differ widely
Let us now examine in more detail the extent to which the self-association of model I and II protein chains impact their architectural properties. To this end, we studied the evolution of two quantities that describe the geometrical properties of the DNA, namely the mean radius of the coil, , and the persistence length of the DNA chain, ξ.
The mean radius of the DNA coil, , is defined according to
| (1) |
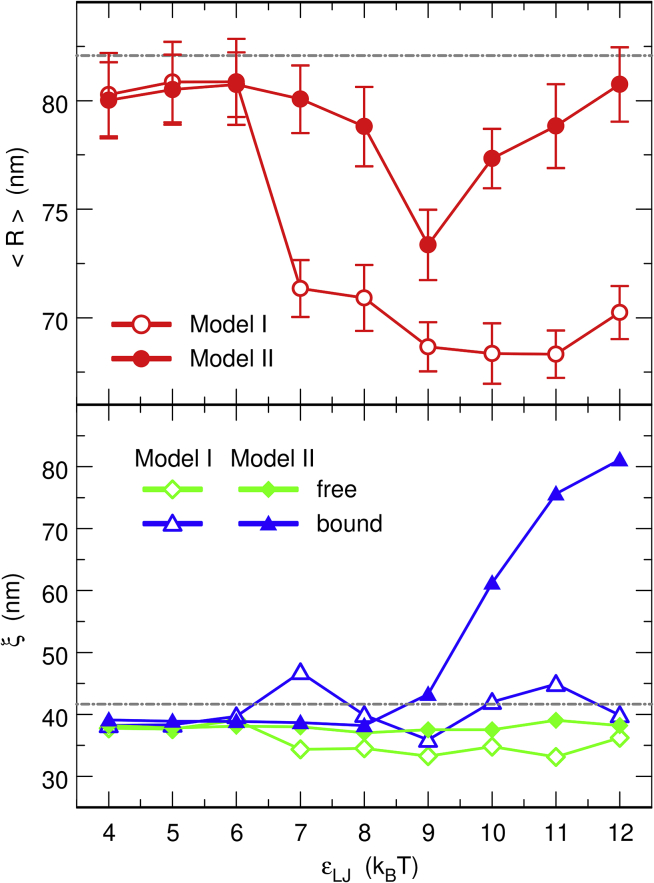
where rk denotes the position of the center of DNA bead k and rCM the position of the center of mass of the DNA coil. In the absence of protein chains, the average value = 82.1 nm results from the balance of the compressive forces exerted by the confinement sphere and the expansive forces arising from the bending rigidity of the DNA chain and the electrostatic repulsion between DNA beads. As long as protein chains self-associate only weakly, the addition of 200 of them inside the confinement sphere has little effect on the mean radius of the DNA coil, as can be checked in the top plot of Fig. 4, which shows the evolution of with increasing values of εLJ. Indeed, remains close to 80 nm for small values of εLJ. This is a direct consequence of the fact that only ∼6% of the proteins bridge the DNA chain, which is insufficient to significantly compact the DNA coil. In contrast, decreases rapidly below 70 nm for model I and values of εLJ larger than 7 kBT. This indicates that the ∼35% of protein chains that bridge the DNA chain (bottom left vignette of Fig. 2) are quite efficient in compacting the DNA coil. However, this is not the case for model II and εLJ > 9 kBT, although ∼35% of the protein chains also bridge the DNA chain. The reason is that most of these bridges localize in thick bundles similar to the one shown in the bottom right vignette of Fig. 2, in which they essentially work to maintain DNA segments parallel to each other. Such distributions of bridges are apparently not as efficient for compacting the DNA coil as the distributions of bridges for model I proteins. An exception occurs for model II and εLJ = 9 kBT, for which significant compaction of the DNA coil ( ≈ 73 nm) is observed in the top plot of Fig. 4. The reason is that this value of εLJ is the smallest one that leads to protein self-association for model II, and DNA-protein complexes are different from the thick bundles observed for larger values of εLJ. They resemble more the conformations obtained with model I, as can be checked in Fig. S9, which shows typical conformations obtained for model II and εLJ = 8 kBT, 9 kBT, and 10 kBT.
Figure 4.
Plot, as a function of εLJ, of the mean radius of the DNA coil (top) and the persistence length of the DNA chain (bottom) for models I (open symbols) and II (solid symbols). In the bottom plot, diamonds represent the values of ξfree and triangles the values of ξbound. The horizontal dot-dashed lines indicate the values of the parameters in the absence of protein chains, that is, = 82.1 nm and ξ = 41.7 nm. Each set of open and solid symbols was obtained from a single simulation with the DNA chain and 200 protein chains by averaging the relevant quantity over time intervals of at least 2.5 ms after equilibration. The error bars in the top plot represent the standard deviation of the fluctuations of . To see this figure in color, go online.
The conclusion is therefore that neither model I nor model II protein chains are efficient in compacting the DNA coil when in the monomer form. In contrast, model I protein chains significantly compact the DNA coil as soon as they self-associate (a similar result was already obtained with a different model (29)). This is not the case for model II protein chains (except for εLJ = 9 kBT), despite the fact that all protein chains interact similarly with the DNA chain.
Let us now consider the persistence length of the DNA chain, ξ, which is related to the directional correlation function C(Δ) = , where t(x) denotes the unit vector tangent to the DNA chain at curvilinear position x, according to
| (2) |
Practically, the directional correlation function can be estimated from
| (3) |
where N is a large number of DNA conformations spanning a large time interval, and tc,k is the unit vector tangent to the DNA chain at the center of bead k in conformation c. The persistence length ξ is obtained from an exponential fit of the evolution of C(Ll0) as a function of L over a certain interval of values of L. By using this procedure for 0 ≤ L ≤ 20, we obtained ξ = 41.7 nm for the DNA chain enclosed in the confinement sphere without protein chains. This value is somewhat smaller than the value estimated from the bending rigidity of the DNA chain (ξ = 50 nm). This is due to the fact that the confinement sphere imposes non-negligible additional curvature to the DNA chain because its diameter (2R0 = 240 nm) is only ∼5 times larger than the persistence length of unconstrained DNA. For equilibrated DNA-protein complexes, it is interesting to discriminate between the persistence length of DNA segments that are not bound to any protein chain (ξfree) and the persistence length of DNA segments bound to at least one protein chain (ξbound). This is easily achieved by testing at each step of the averaging procedure whether any bead of the DNA segment between beads k and k + L binds to a protein chain or not and using this segment adequately to compute either ξfree or ξbound.
The evolution of ξfree and ξbound with increasing values of εLJ is shown in the bottom plot of Fig. 4. Not surprisingly, ξfree remains close to 40 nm for all values of εLJ. Moreover, addition of 200 protein chains inside the confinement sphere has little effect on ξbound as long as protein chains do not self-associate significantly. This indicates that the ∼22% of protein chains that bind to the DNA chain in cis do not significantly increase its rigidity. In contrast, ξbound increases rapidly up to ∼80 nm for model II and values of εLJ larger than 10 kBT. This confirms that the thick bundles composed of DNA segments maintained parallel to each other and bridged by protein segments are quite rigid, as could be anticipated from their almost rectilinear shape (bottom right vignette of Fig. 2). No increase in ξbound is observed for model I, however, even for large values of εLJ, as can be checked in Fig. 4. Although the network of protein chains formed for εLJ ≥ 7 kBT is quite efficient in compacting the DNA coil, it is flexible enough for the numerous cross-links not to significantly alter the persistence length of DNA segments bound to protein chains.
The conclusion is consequently that neither model I nor model II protein chains are efficient in altering the persistence length of the DNA coil when in the monomer form. In contrast, when model II protein chains self-associate, the persistence length of DNA segments localized in the thick bundles formed by DNA-protein complexes is twice as large as that of free DNA segments. This is, however, not the case for model I protein chains, despite the fact that all protein chains interact similarly with the DNA chain.
Discussion
In this work, we studied the properties of two models describing nonspecific interactions between circular DNA and nucleoid proteins. The DNA-protein interaction potential is the same for the two models and was kept constant in all simulations. In contrast, when the strength of protein-protein interactions is large enough, model I proteins self-associate in the form of clusters, whereas model II proteins form filaments. The strength of protein-protein interactions was varied systematically in the simulations to check the impact of protein self-association on the geometrical and mechanical properties of DNA-protein complexes. The two models display characteristic features:
-
•
for the two models, binding of the proteins to the DNA increases strongly when proteins self-associate, although the strength of DNA-protein interactions is kept constant;
-
•
when in the monomer form, neither model I nor model II proteins are efficient in compacting the DNA coil or increasing the rigidity of the DNA;
-
•
clusters of model I proteins significantly compact the DNA coil, but this is not the case for filaments of model II proteins, although all proteins interact similarly with the DNA; and
-
•
filaments of model II proteins significantly increase the rigidity of the DNA, but this is not the case for clusters of model I proteins, although all proteins interact similarly with the DNA.
These models consequently suggest that the self-association of nucleoid proteins may have a rich and profound impact on their architectural properties. This claim and the models proposed here are supported by a set of experimental results.
First, many of the nucleoid proteins can self-associate and are present in cells in polymeric forms. For example, H-NS proteins form dimers at low concentrations but assemble into larger multimers at higher concentrations (25,30,31). Other members of the H-NS family, like StpA, can also self-associate (31,32). As for the models proposed here, cooperative binding of H-NS to DNA is related to protein-protein interactions (33). The resulting filaments of H-NS proteins bound to the DNA substrate are clearly seen in crystallographic experiments (34). It is believed that such protein filaments block DNA accessibility and are the structural basis for gene silencing (35,36), which is one of the main roles of H-NS in the cells. It has, however, been shown that the simple coverage of the DNA substrate by H-NS proteins at high concentrations is not sufficient and that the capacity of proteins to self-associate is crucial for the regulation of gene expression; derivatives of H-NS that are unable to oligomerize fail in silencing genes (37, 38, 39, 40).
Moreover, two nucleoid proteins present in the stationary phase, Dps and CbpA, can also self-associate, and experiments have shown that their aggregation and the compaction of the DNA are parallel phenomena (41,42). Unlike H-NS (26), Dps molecules do not align in filaments in co-crystals, but rather are packed in pseudohexagonal layers (43). The layers slide along the DNA direction and enable the formation of grooves for DNA accommodation (43). Similarly, partition proteins ParB in vivo first bind to the specific parS site and then spread; that is, they simultaneously self-assemble stochastically and bind to the DNA away from the parS site, thus bridging the DNA (44, 45, 46). Spreading ability is required, as ParB mutants that lack this ability are also defective in partition (47).
According to this short digest of experimental results, it appears that model I adequately captures the main features of Dps, CbpA, and ParB, which must assemble in clusters to bind to and compact the DNA molecule. The case of H-NS is more complex. Owing to the crystallographic structure in (34), which displays H-NS filaments aligned parallel to the DNA molecule, as well as the experimental observation that the persistence length of DNA/H-NS complexes may be as large as 130 nm at low divalent cation concentrations (48), it is tempting to conclude that model II provides a correct description of DNA/H-NS interactions in this salt regime. However, the fact that H-NS proteins form filaments when bound to the DNA is not a proof that they also do so when the substrate is lacking (20). The switch from the DNA-stiffening mode to the DNA-bridging mode of H-NS at higher divalent cation concentrations (19) may also be tentatively interpreted as an indication that the self-association properties of H-NS switch from the model II type to the model I type. In this respect, we note with interest that it has recently been shown that environmental variations have a direct effect on the self-association properties of H-NS (49). More work, however, is clearly needed to ascertain whether the switch from the DNA-stiffening mode to the DNA-bridging mode of H-NS is due to a decrease in the strength of DNA-protein interactions, as proposed in (20); a variation in the self-association mode of proteins, as suggested by this work; or variations in the geometry of the H-NS molecule (50).
Conclusion
In this work, we used coarse-grained modeling to investigate the impact of the self-association of nucleoid proteins on their architectural properties. The simulations suggest that this impact is probably strong and that different modes of self-association may result in different architectural capabilities of the proteins. Self-association is therefore a property of the proteins that is worth considering when trying to understand how they shape the DNA coil.
To conclude, we would like to mention that models similar to those discussed in this work have recently been proposed to study the formation of the bacterial nucleoid through the demixing of DNA and nonbinding globular macromolecules (51, 52, 53, 54); the preferential localization of the nucleoid inside the cell (26); the mechanism of facilitated diffusion, by which proteins search for their targets along the DNA sequence (55, 56, 57); and the requirements for DNA-bridging proteins to act as topological barriers of the bacterial genome (23). All these models are compatible, and it is possible to combine two (or more) of them to get a more complete and realistic description of bacterial cells (22). This point is crucial because the effects of different processes taking place simultaneously in living cells are not simply additive, and the outcome may be difficult to predict when considering only the effects of each mechanism taken separately (22,29). For example, both DNA and macromolecule demixing and DNA supercoiling contribute to the compaction of the bacterial DNA, but the total compaction of the DNA coil is the sum of the two contributions only in a limited range of values of macromolecular concentration and superhelical density, whereas their interplay is much more complex outside from this range (22). In this respect, it will certainly be instructive in future work to use the models discussed in this study to investigate the interplay of nucleoid proteins and macromolecular crowders (29,52, 53, 54) or transcription factors (55, 56, 57), or DNA supercoiling and topological insulators (23).
Editor: Yamini Dalal.
Footnotes
Supporting Material can be found online at https://doi.org/10.1016/j.bpj.2020.12.006.
Supporting Citations
References (58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70) appear in the Supporting Material.
Supporting Material
References
- 1.Stonington O.G., Pettijohn D.E. The folded genome of Escherichia coli isolated in a protein-DNA-RNA complex. Proc. Natl. Acad. Sci. USA. 1971;68:6–9. doi: 10.1073/pnas.68.1.6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Worcel A., Burgi E. On the structure of the folded chromosome of Escherichia coli. J. Mol. Biol. 1972;71:127–147. doi: 10.1016/0022-2836(72)90342-7. [DOI] [PubMed] [Google Scholar]
- 3.Browning D.F., Grainger D.C., Busby S.J.W. Effects of nucleoid-associated proteins on bacterial chromosome structure and gene expression. Curr. Opin. Microbiol. 2010;13:773–780. doi: 10.1016/j.mib.2010.09.013. [DOI] [PubMed] [Google Scholar]
- 4.Dillon S.C., Cameron A.D.S., Dorman C.J. Genome-wide analysis of the H-NS and Sfh regulatory networks in Salmonella Typhimurium identifies a plasmid-encoded transcription silencing mechanism. Mol. Microbiol. 2010;76:1250–1265. doi: 10.1111/j.1365-2958.2010.07173.x. [DOI] [PubMed] [Google Scholar]
- 5.Dillon S.C., Dorman C.J. Bacterial nucleoid-associated proteins, nucleoid structure and gene expression. Nat. Rev. Microbiol. 2010;8:185–195. doi: 10.1038/nrmicro2261. [DOI] [PubMed] [Google Scholar]
- 6.Johnson R.C., Johnson L.M., Gardner J.F. Major nucleoid proteins in the structure and function of the Escherichia coli chromosome. In: Higgins N.P., editor. The Bacterial Chromosome. ASM; 2005. pp. 65–132. [Google Scholar]
- 7.Dame R.T. The role of nucleoid-associated proteins in the organization and compaction of bacterial chromatin. Mol. Microbiol. 2005;56:858–870. doi: 10.1111/j.1365-2958.2005.04598.x. [DOI] [PubMed] [Google Scholar]
- 8.Luijsterburg M.S., White M.F., Dame R.T. The major architects of chromatin: architectural proteins in bacteria, archaea and eukaryotes. Crit. Rev. Biochem. Mol. Biol. 2008;43:393–418. doi: 10.1080/10409230802528488. [DOI] [PubMed] [Google Scholar]
- 9.Azam T.A., Ishihama A. Twelve species of the nucleoid-associated protein from Escherichia coli. Sequence recognition specificity and DNA binding affinity. J. Biol. Chem. 1999;274:33105–33113. doi: 10.1074/jbc.274.46.33105. [DOI] [PubMed] [Google Scholar]
- 10.Pinson V., Takahashi M., Rouviere-Yaniv J. Differential binding of the Escherichia coli HU, homodimeric forms and heterodimeric form to linear, gapped and cruciform DNA. J. Mol. Biol. 1999;287:485–497. doi: 10.1006/jmbi.1999.2631. [DOI] [PubMed] [Google Scholar]
- 11.Wang S., Cosstick R., Gumport R.I. The specific binding of Escherichia coli integration host factor involves both major and minor grooves of DNA. Biochemistry. 1995;34:13082–13090. doi: 10.1021/bi00040a020. [DOI] [PubMed] [Google Scholar]
- 12.Gulvady R., Gao Y., Yan J. A single molecule analysis of H-NS uncouples DNA binding affinity from DNA specificity. Nucleic Acids Res. 2018;46:10216–10224. doi: 10.1093/nar/gky826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Stella S., Cascio D., Johnson R.C. The shape of the DNA minor groove directs binding by the DNA-bending protein Fis. Genes Dev. 2010;24:814–826. doi: 10.1101/gad.1900610. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Peterson S.N., Dahlquist F.W., Reich N.O. The role of high affinity non-specific DNA binding by Lrp in transcriptional regulation and DNA organization. J. Mol. Biol. 2007;369:1307–1317. doi: 10.1016/j.jmb.2007.04.023. [DOI] [PubMed] [Google Scholar]
- 15.Ali Azam T., Iwata A., Ishihama A. Growth phase-dependent variation in protein composition of the Escherichia coli nucleoid. J. Bacteriol. 1999;181:6361–6370. doi: 10.1128/jb.181.20.6361-6370.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Azam T.A., Hiraga S., Ishihama A. Two types of localization of the DNA-binding proteins within the Escherichia coli nucleoid. Genes Cells. 2000;5:613–626. doi: 10.1046/j.1365-2443.2000.00350.x. [DOI] [PubMed] [Google Scholar]
- 17.Vora T., Hottes A.K., Tavazoie S. Protein occupancy landscape of a bacterial genome. Mol. Cell. 2009;35:247–253. doi: 10.1016/j.molcel.2009.06.035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Song D., Loparo J.J. Building bridges within the bacterial chromosome. Trends Genet. 2015;31:164–173. doi: 10.1016/j.tig.2015.01.003. [DOI] [PubMed] [Google Scholar]
- 19.Liu Y., Chen H., Yan J. A divalent switch drives H-NS/DNA-binding conformations between stiffening and bridging modes. Genes Dev. 2010;24:339–344. doi: 10.1101/gad.1883510. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Joyeux M. Role of salt valency in the switch of H-NS proteins between DNA-bridging and DNA-stiffening modes. Biophys. J. 2018;114:2317–2325. doi: 10.1016/j.bpj.2018.02.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Zhao H., Winogradoff D., Papoian G.A. The oligomerization landscape of histones. Biophys. J. 2019;116:1845–1855. doi: 10.1016/j.bpj.2019.03.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Joyeux M. Bacterial nucleoid: interplay of DNA demixing and supercoiling. Biophys. J. 2020;118:2141–2150. doi: 10.1016/j.bpj.2019.09.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Joyeux M., Junier I. Requirements for DNA-bridging proteins to act as topological barriers of the bacterial genome. Biophys. J. 2020;119:1215–1225. doi: 10.1016/j.bpj.2020.08.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Ono S., Goldberg M.D., Ladbury J.E. H-NS is a part of a thermally controlled mechanism for bacterial gene regulation. Biochem. J. 2005;391:203–213. doi: 10.1042/BJ20050453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Ceschini S., Lupidi G., Angeletti M. Multimeric self-assembly equilibria involving the histone-like protein H-NS. A thermodynamic study. J. Biol. Chem. 2000;275:729–734. doi: 10.1074/jbc.275.2.729. [DOI] [PubMed] [Google Scholar]
- 26.Joyeux M. Preferential localization of the bacterial nucleoid. Microorganisms. 2019;7:204. doi: 10.3390/microorganisms7070204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Joyeux M., Vreede J. A model of H-NS mediated compaction of bacterial DNA. Biophys. J. 2013;104:1615–1622. doi: 10.1016/j.bpj.2013.02.043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Joyeux M. Equilibration of complexes of DNA and H-NS proteins on charged surfaces: a coarse-grained model point of view. J. Chem. Phys. 2014;141:115102. doi: 10.1063/1.4895819. [DOI] [PubMed] [Google Scholar]
- 29.Dias R.S. Role of protein self-association on DNA condensation and nucleoid stability in a bacterial cell model. Polymers (Basel) 2019;11:1102. doi: 10.3390/polym11071102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Smyth C.P., Lundbäck T., Ladbury J.E. Oligomerization of the chromatin-structuring protein H-NS. Mol. Microbiol. 2000;36:962–972. doi: 10.1046/j.1365-2958.2000.01917.x. [DOI] [PubMed] [Google Scholar]
- 31.Leonard P.G., Ono S., Ladbury J.E. Investigation of the self-association and hetero-association interactions of H-NS and StpA from Enterobacteria. Mol. Microbiol. 2009;73:165–179. doi: 10.1111/j.1365-2958.2009.06754.x. [DOI] [PubMed] [Google Scholar]
- 32.Suzuki C., Kawazuma K., Nojiri H. Oligomerization mechanisms of an H-NS family protein, Pmr, encoded on the plasmid pCAR1 provide a molecular basis for functions of H-NS family members. PLoS One. 2014;9:e105656. doi: 10.1371/journal.pone.0105656. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Giangrossi M., Wintraecken K., de Vries R. Probing the relation between protein-protein interactions and DNA binding for a linker mutant of the bacterial nucleoid protein H-NS. Biochim. Biophys. Acta. 2014;1844:339–345. doi: 10.1016/j.bbapap.2013.11.010. [DOI] [PubMed] [Google Scholar]
- 34.Arold S.T., Leonard P.G., Ladbury J.E. H-NS forms a superhelical protein scaffold for DNA condensation. Proc. Natl. Acad. Sci. USA. 2010;107:15728–15732. doi: 10.1073/pnas.1006966107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Lim C.J., Lee S.Y., Yan J. Nucleoprotein filament formation is the structural basis for bacterial protein H-NS gene silencing. Sci. Rep. 2012;2:509. doi: 10.1038/srep00509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Lim C.J., Whang Y.R., Yan J. Gene silencing H-NS paralogue StpA forms a rigid protein filament along DNA that blocks DNA accessibility. Nucleic Acids Res. 2012;40:3316–3328. doi: 10.1093/nar/gkr1247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Spurio R., Falconi M., Gualerzi C.O. The oligomeric structure of nucleoid protein H-NS is necessary for recognition of intrinsically curved DNA and for DNA bending. EMBO J. 1997;16:1795–1805. doi: 10.1093/emboj/16.7.1795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Winardhi R.S., Fu W., Yan J. Higher order oligomerization is required for H-NS family member MvaT to form gene-silencing nucleoprotein filament. Nucleic Acids Res. 2012;40:8942–8952. doi: 10.1093/nar/gks669. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Winardhi R.S., Yan J., Kenney L.J. H-NS regulates gene expression and compacts the nucleoid: insights from single-molecule experiments. Biophys. J. 2015;109:1321–1329. doi: 10.1016/j.bpj.2015.08.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Yamanaka Y., Winardhi R.S., Yamamoto K. Dimerization site 2 of the bacterial DNA-binding protein H-NS is required for gene silencing and stiffened nucleoprotein filament formation. J. Biol. Chem. 2018;293:9496–9505. doi: 10.1074/jbc.RA117.001425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Ceci P., Cellai S., Chiancone E. DNA condensation and self-aggregation of Escherichia coli Dps are coupled phenomena related to the properties of the N-terminus. Nucleic Acids Res. 2004;32:5935–5944. doi: 10.1093/nar/gkh915. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Cosgriff S., Chintakayala K., Grainger D.C. Dimerization and DNA-dependent aggregation of the Escherichia coli nucleoid protein and chaperone CbpA. Mol. Microbiol. 2010;77:1289–1300. doi: 10.1111/j.1365-2958.2010.07292.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Dadinova L.A., Chesnokov Y.M., Shtykova E.V. Protective Dps-DNA co-crystallization in stressed cells: an in vitro structural study by small-angle X-ray scattering and cryo-electron tomography. FEBS Lett. 2019;593:1360–1371. doi: 10.1002/1873-3468.13439. [DOI] [PubMed] [Google Scholar]
- 44.Sanchez A., Cattoni D.I., Bouet J.-Y. Stochastic self-assembly of ParB proteins builds the bacterial DNA segregation apparatus. Cell Syst. 2015;1:163–173. doi: 10.1016/j.cels.2015.07.013. [DOI] [PubMed] [Google Scholar]
- 45.Broedersz C.P., Wang X., Wingreen N.S. Condensation and localization of the partitioning protein ParB on the bacterial chromosome. Proc. Natl. Acad. Sci. USA. 2014;111:8809–8814. doi: 10.1073/pnas.1402529111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Funnell B.E. ParB partition proteins: complex formation and spreading at bacterial and plasmid centromeres. Front. Mol. Biosci. 2016;3:44. doi: 10.3389/fmolb.2016.00044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Graham T.G.W., Wang X., Loparo J.J. ParB spreading requires DNA bridging. Genes Dev. 2014;28:1228–1238. doi: 10.1101/gad.242206.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Amit R., Oppenheim A.B., Stavans J. Increased bending rigidity of single DNA molecules by H-NS, a temperature and osmolarity sensor. Biophys. J. 2003;84:2467–2473. doi: 10.1016/S0006-3495(03)75051-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Shahul Hameed U.F., Liao C., Arold S.T. H-NS uses an autoinhibitory conformational switch for environment-controlled gene silencing. Nucleic Acids Res. 2019;47:2666–2680. doi: 10.1093/nar/gky1299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Qin L., Bdira F.B., Dame R.T. Structural basis for osmotic regulation of the DNA binding properties of H-NS proteins. Nucleic Acids Res. 2020;48:2156–2172. doi: 10.1093/nar/gkz1226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Joyeux M. Compaction of bacterial genomic DNA: clarifying the concepts. J. Phys. Condens. Matter. 2015;27:383001. doi: 10.1088/0953-8984/27/38/383001. [DOI] [PubMed] [Google Scholar]
- 52.Joyeux M. In vivo compaction dynamics of bacterial DNA: a fingerprint of DNA/RNA demixing? Curr. Opin. Colloid Interface Sci. 2016;26:17–27. [Google Scholar]
- 53.Joyeux M. Coarse-grained model of the demixing of DNA and non-binding globular macromolecules. J. Phys. Chem. B. 2017;121:6351–6358. doi: 10.1021/acs.jpcb.7b03011. [DOI] [PubMed] [Google Scholar]
- 54.Joyeux M. A segregative phase separation scenario of the formation of the bacterial nucleoid. Soft Matter. 2018;14:7368–7381. doi: 10.1039/c8sm01205a. [DOI] [PubMed] [Google Scholar]
- 55.Florescu A.M., Joyeux M. Description of nonspecific DNA-protein interaction and facilitated diffusion with a dynamical model. J. Chem. Phys. 2009;130:015103. doi: 10.1063/1.3050097. [DOI] [PubMed] [Google Scholar]
- 56.Florescu A.M., Joyeux M. Dynamical model of DNA-protein interaction: effect of protein charge distribution and mechanical properties. J. Chem. Phys. 2009;131:105102. [Google Scholar]
- 57.Florescu A.M., Joyeux M. Comparison of kinetic and dynamical models of DNA-protein interaction and facilitated diffusion. J. Phys. Chem. A. 2010;114:9662–9672. doi: 10.1021/jp101151a. [DOI] [PubMed] [Google Scholar]
- 58.Jian H., Vologodskii A., Schlick T. A combined wormlike-chain and bead model for dynamic simulations of long linear DNA. J. Comput. Phys. 1997;136:168–179. [Google Scholar]
- 59.Manning G.S. Limiting laws and counterion condensation in polyelectrolyte solutions. I. Colligative properties. J. Chem. Phys. 1969;51:924–933. [Google Scholar]
- 60.Oosawa F. Marcel Dekker; New York: 1971. Polyelectrolytes. [Google Scholar]
- 61.Dorman C.J., Hinton J.C., Free A. Domain organization and oligomerization among H-NS-like nucleoid-associated proteins in bacteria. Trends Microbiol. 1999;7:124–128. doi: 10.1016/s0966-842x(99)01455-9. [DOI] [PubMed] [Google Scholar]
- 62.Link A.J., Robison K., Church G.M. Comparing the predicted and observed properties of proteins encoded in the genome of Escherichia coli K-12. Electrophoresis. 1997;18:1259–1313. doi: 10.1002/elps.1150180807. [DOI] [PubMed] [Google Scholar]
- 63.Record M.T., Jr., Anderson C.F., Lohman T.M. Thermodynamic analysis of ion effects on the binding and conformational equilibria of proteins and nucleic acids: the roles of ion association or release, screening, and ion effects on water activity. Q. Rev. Biophys. 1978;11:103–178. doi: 10.1017/s003358350000202x. [DOI] [PubMed] [Google Scholar]
- 64.Mascotti D.P., Lohman T.M. Thermodynamic extent of counterion release upon binding oligolysines to single-stranded nucleic acids. Proc. Natl. Acad. Sci. USA. 1990;87:3142–3146. doi: 10.1073/pnas.87.8.3142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Fenley M.O., Russo C., Manning G.S. Theoretical assessment of the oligolysine model for ionic interactions in protein-DNA complexes. J. Phys. Chem. B. 2011;115:9864–9872. doi: 10.1021/jp204915y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Breslauer K.J., Remeta D.P., Marky L.A. Enthalpy-entropy compensations in drug-DNA binding studies. Proc. Natl. Acad. Sci. USA. 1987;84:8922–8926. doi: 10.1073/pnas.84.24.8922. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Wang S., Kumar A., Wilson W.D. Different thermodynamic signatures for DNA minor groove binding with changes in salt concentration and temperature. Chem. Commun. (Camb.) 2013;49:8543–8545. doi: 10.1039/c3cc44569k. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Meyer E.E., Rosenberg K.J., Israelachvili J. Recent progress in understanding hydrophobic interactions. Proc. Natl. Acad. Sci. USA. 2006;103:15739–15746. doi: 10.1073/pnas.0606422103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Lin M.S., Fawzi N.L., Head-Gordon T. Hydrophobic potential of mean force as a solvation function for protein structure prediction. Structure. 2007;15:727–740. doi: 10.1016/j.str.2007.05.004. [DOI] [PubMed] [Google Scholar]
- 70.Makowski M., Czaplewski C., Scheraga H.A. Potential of mean force of association of large hydrophobic particles: toward the nanoscale limit. J. Phys. Chem. B. 2010;114:993–1003. doi: 10.1021/jp907794h. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.