Abstract
Myeloid cell leukemia-1 (Mcl-1) is a validated and attractive target for cancer therapy. Over-expression of Mcl-1 in many cancers allows cancer cells to evade apoptosis and contributes to their resistance to current chemotherapeutics. In this study, more than thirty coumarin derivatives with different substituents were designed and synthesized, and their Mcl-1 inhibitory activities evaluated using a fluorescence polarization-based binding assay. The results showed that the catechol group was a key constituent for Mcl-1 inhibitory activity of the coumarins, and methylation of the catechol group led to decreased inhibitory activity. The introduction of a hydrophobic electron-withdrawing group at the C-4 position of 6,7-dihydroxycoumarin, enhanced Mcl-1 inhibitory capacity, and a hydrophilic group in this position was unbeneficial to the inhibitory potency. In addition, the introduction of a nitrogen-containing group to the C-5 or C-8 position, which allowed an intramolecular hydrogen bond, was also unfavorable for Mcl-1 inhibition. Among all coumarins tested, 4-trifluoromethyl-6,7-dihydroxycoumarin (Cpd 4) displayed the most potent inhibitory activity towards Mcl-1 (Ki = 0.21 ± 0.02 μM, IC50 = 1.21 ± 0.56 μM, respectively), for which the beneficial effect on taxol resistance was also validated in A549 cells. A strong interaction between Cpd 4 and Mcl-1 in docking simulations further supported the observed potent Mcl-1 inhibition ability of Cpd 4. 3D-QSAR analysis of all tested coumarin derivatives further provides new insights into the relationships linking the inhibitory effects on Mcl-1 and the steric-electrostatic properties of coumarins. These findings could be of great value for medicinal chemists for the design and development of more potent Mcl-1 inhibitors for biomedical applications.
Keywords: Myeloid cell leukemia-1 (Mcl-1), coumarins, structure-activity relationship (SAR)
Graphical Abstract

1. Introduction
Mcl-1 (myeloid cell leukemia 1) is a member of the Bcl-2 family of proteins and plays a central role in apoptosis regulation via interacting with other Bcl-2 family members.1 Mcl-1 was established as a major regulator of carcinogenesis and its dysregulation prevents cancer cells from undergoing programmed cell death. Mcl-1 overexpression was observed in a variety of human cancers, including lung, breast, prostate, pancreatic, ovarian, and cervical cancers, and also in melanoma and leukemia.2–6 Moreover, Mcl-1 overexpression has emerged as a resistance mechanism against a number of anticancer therapies including paclitaxel and vincristine, which are widely prescribed microtubule-targeted chemotherapeutic agents and also gemcitabine,7,8 a first-line drug for pancreatic cancer. Mcl-1 overexpression also confers resistance to ABT-263, which is an inhibitor of other Bcl-2 family members Bcl-2/Bcl-xL, and is currently in clinical trials.9–11 Inhibited cell growth and induced apoptosis of cancer cells in vitro, and markedly decreased tumorigenicity in vivo were observed after specific inhibition of Mcl-1 or by Mcl-1 silencing.12 These data suggest that direct inhibition of Mcl-1 could be an effective therapeutic option for a wide variety of cancers. In recent years, several groups reported on small molecules and stapled peptides that bind to Mcl-1 in an attempt to target Mcl-1 for cancer treatment,13–17 although few of these identified Mcl-1 inhibitors showed significant downregulating effects, and none have entered into the clinic.
Coumarins are bioactive compounds of both natural and synthetic origin. There has been a growing interest in their discovery and development into drugs due to their versatile synthetic scaffold and immeasurable pharmaceutical and biological potential with minimum side effects. A number of natural coumarins, as well as coumarin derivatives that have been modified by medicinal chemists over the years, have been used as antioxidants, antivirals, monoamine oxidases, cholinesterases and aromatase inhibitors to treat various diseases and metabolic pathway disorders.18–23 Coumarins also exhibit antitumor activities at different stages of cancer formation through various mechanisms, for example blocking cell cycle, inducing cell apoptosis, modulating estrogen receptor (ER), or inhibiting the DNA-associated enzymes, such as topoisomerase.24–29 Recently, natural coumarins were demonstrated to have anticancer activities through a mechanism involving Mcl-1 inhibition,30 but the corresponding systematic study on their structure-Mcl-1 inhibitory activity relationship is lacking.
In the present study, a series of natural or synthetic coumarins with varied substitutions on their scaffolds were used to evaluate the inhibitory activity towards Mcl-1 as assessed by the FAP assay. The structure-Mcl-1 inhibitory activity relationship of these coumarins are summarized and discussed in detail. To get a deeper understanding of the interaction between these analogs and Mcl-1, the compound with the most potent inhibitory activity was docked to the binding site of Mcl-1. A 3D-QSAR model was further developed to explore the structural requirement of coumarins for potent Mcl-1 inhibition. This research could provide useful guidance to the design and optimization of potent coumarin-type Mcl-1 inhibitors for anticancer therapy.
2. Results and discussion
2.1. Chemistry
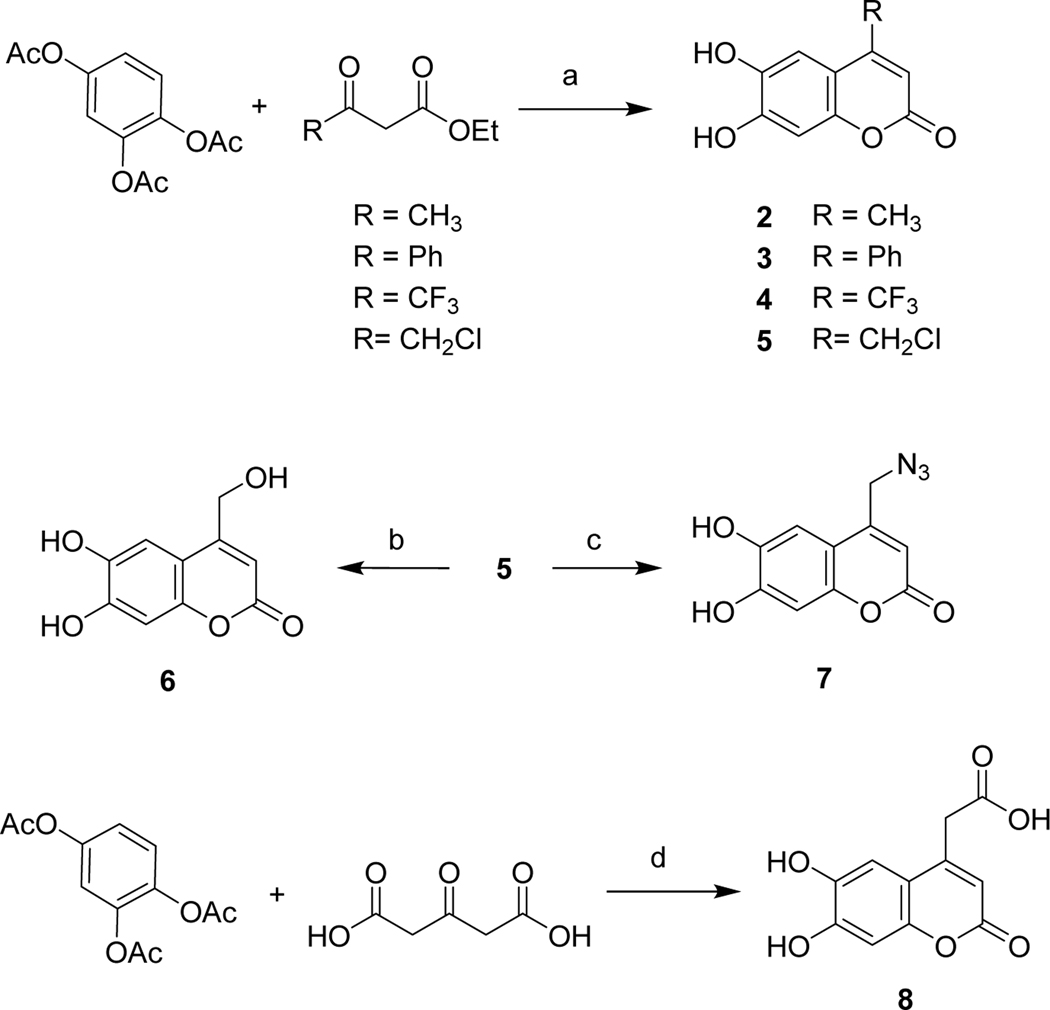
To explore the effects of different property groups and their position on hydroxycoumarin on Mcl-1 inhibition activities, a series of 4-substituted 6,7-dihydroxycoumarin derivatives were designed and prepared from 1,2,4-phenenyl triacetate via acid catalytic pechmann condensation with good yields (Scheme 1). After that, compounds 6 and 7 were obtained from chloride 5 by reflux in a mixture of DMF and water (3:10, v:v) and treated with sodium azide, respectively. 4-Carboxylic 9 was synthesized by treating 1,2,3-phenenyl triacetate with 1,3-acetonedicarboxylic acid in the present of perchloric acid.31 The substituents were selected mainly based on the electronic and lipophilic considerations defined by Craig′s plot.32 In the present study, trifluoromethyl and phenyl were selected for their electron-withdrawing and hydrophobic property; carboxyl was selected to represent electron-withdrawing and hydrophilic substituents, while groups like hydroxymethyl, azidomethyl, and chloromethyl were representative of electron-donating and hydrophilic substitution, methyl represented electron-donating and hydrophobic substituents.
Scheme 1.
Synthesis of 4-substituted 6,7-dihydroxycoumarin derivatives. Reagents and conditions: (a) HClO4, rt, 6 ~ 8 h; (b) DMF/H2O (1:3), reflux, 20 h; (c) DMF, NaN3, rt, 10 h; (d) HClO4, 50°C, 4 h.
Introduction of a nitrogen containing group as hydrogen bond acceptor on the 6,7-dihydroxycoumarins by Mannich reaction (Scheme 2) was also carried out. After formaldehyde and the corresponding secondary amine were stirred in ethanol for 0.5 h, the reaction was added to a solution of 6,7-dihydroxycoumarin/4-substituted 6,7-dihydroxycoumarin in ethanol or methanol, and stirring for 8 ~ 12 h at 25 ~ 60°C, which obtained compounds 9 ~ 16 after being purified through flash column chromatography on a silica gel. It is worth mentioning that esculetin condenses with formaldehyde and amines with the low yield and poor regioselectivity. For example, 5-substituted compounds 10 and 12 were formed simultaneously, when 8-substituted compounds 9 and 11 were prepared, respectively. While the C4-H was substituted with methyl, 4-methyl-6,7-dihydroxycoumarin condensed with formaldehyde and amines to compound 14 ~ 16 with the good yield and the excellent regioselectivity. Furthermore, monomethylation 17 and 18 were obtained from 6,7-dihydroxycoumarin via selectively methylated at 7-OH and 6-OH in a good yield with CH3I in the presence of Na2CO3 at 25 °C and NaH at 0 °C, respectively (Scheme 3).33 The nitro-esculetin 20 was obtained from dimethylation 19 through nitration reaction.
Scheme 2.
Synthesis of 6,7-dihydroxycoumarins bearing Mannich bases.
Scheme 3.
Synthesis of methylated 6,7-dihydroxycoumarin derivatives. Reagents and conditions: (a) CH3I, Na2CO3, DMF, 25 °C, 4 h. (b) CH3I, NaH, DMF, 0 °C, 1 h; (c) CH3I, NaH, DMF, rt, 5 h; (d) DCM, H2SO4, HNO3, 0°C, 3h.
To further explore the effect of the existence of phenolic groups on the coumarin skeleton for Mcl-1 inhibition activities, 4-substituted 7,8-dihydroxycoumarins and their methylation were prepared by acid catalytic pechmann condensation mentioned above (Scheme 4).34 Furthermore, a few 7-hydroxycoumarin derivatives including 7-hydroxyl-4-methyl coumarin 29, 7-hydroxyl-4-methyl-8-nitro coumarin 30, 6-chloro-7-hydroxy-4-methyl coumarin 31 and 6-ethyl-7-hydroxy-4-methy coumarin 32 were also prepared as depicted in Scheme 5. In addition, some hydroxycoumarin derivatives including daphnetin 21, 7-methoxy daphnetin 25, fraxetin 33, coumarin 34, 7-hydroxycoumarin 35 and 6-hydroxycoumarin 36, which were purchased from Chengdu Pufei De Biotech Co. Ltd. (Sichuan, China), and were employed in our study in order to explore the potential structure-Mcl-1 inhibition activity relationships of hydroxyl coumarin derivatives.
Scheme 4.
Synthesis of 4-substituted 7,8-dihydroxycoumarin and methylated derivatives. Reagents and conditions: (a) HClO4, rt, 6 ~ 8 h; (b) DMF/H2O (1:3), reflux, 20 h.
Scheme 5.
Synthesis of 7-hydroxycoumarin derivatives. Reagents and conditions: (a) H2SO4, HNO3, 0°C, 3h; (b) HClO4, rt, 6 h;
2.2. Mcl-1 inhibitory activity assays
Coumarins, a class of widespread natural compounds containing fused benzene and pyrone ring systems, are an excellent reservoir of biologically active compounds.35 To investigate the potency of Mcl-1 inhibition, the binding affinities of 36 natural and synthesized coumarins to Mcl-1 protein were determined using Fluorescence polarization-based binding assay, in which the ability of inhibitors to disrupt the interaction between Mcl-1 and two different BH3 peptides, fluorescently labeled Bid and biotin-labeled Bim, was tested.
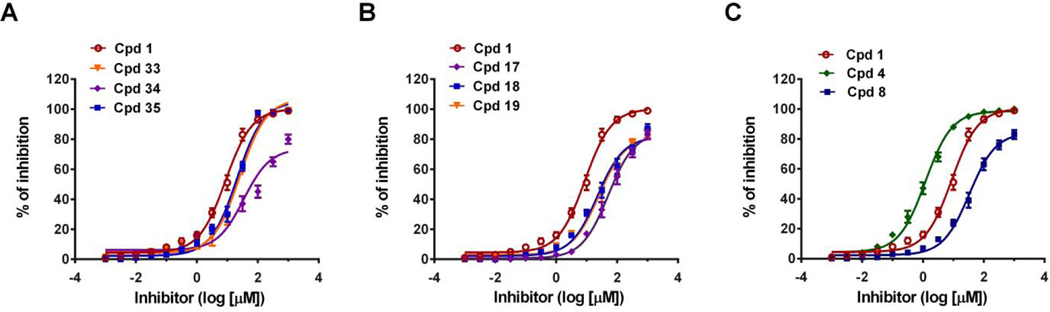
The effect of the position and number of hydroxyl groups on the coumarin scaffold toward inhibitory activity of Mcl-1 were investigated firstly. As shown in Table 1, coumarin 33 exhibited Mcl-1inhibitory activity with Ki value of 4.06 ± 0.05 μM. A 6-fold decrease in Mcl-1 inhibitory activity was observed when hydroxyl was introduced to the C-6 position of coumarin (34), while the C-7 hydroxyl substituent of coumarin (35) displayed little effect on its inhibition potency of Mcl-1. When catechol group was introduced on coumarin to form 6,7-dihydroxycoumarin 1 and 7,8-dihydroxycoumarin 21, the Mcl-1inhibitory activities were improved significantly with Ki values of 1.49 ± 0.04 μM and 1.75 ± 0.032 μM, respectively. Furthermore, when the catechol groups of compounds 1 and 21 were blocked by methylation, the inhibitory activities towards Mcl-1 were significantly reduced, with Ki values of 15.9 ± 0.03 μM, 13.4 ± 0.08 μM and 9.54±0.08 μM for the methylated products 17, 18 and 19 of compound 1, respectively. The competitive binding curves of these compounds to Mcl-1 were outlined in Fig.1A and 1B. These results indicated that catechol moieties are key groups for the Mcl-1inhibitory activities of these coumarin derivatives.
Table 1.
The IC50 and Ki values of coumarins against Mcl-1.
 | ||||||||
|---|---|---|---|---|---|---|---|---|
| Cpd | R1 | R2 | R3 | R4 | R5 | R6 | Ki (μM) | IC50 (μM) |
| 1 | H | H | H | OH | OH | H | 1.49 ± 0.04 | 8.77 ± 1.16 |
| 2 | H | CH3 | H | OH | OH | H | 1.19 ± 0.04 | 6.28 ± 1.08 |
| 3 | H | C6H5 | H | OH | OH | H | 0.65 ± 0.03 | 3.27 ± 1.77 |
| 4 | H | CF3 | H | OH | OH | H | 0.21 ± 0.02 | 1.21 ± 0.56 |
| 5 | H | CH2Cl | H | OH | OH | H | 1.04 ± 0.04 | 5.82 ± 1.18 |
| 6 | H | CH2OH | H | OH | OH | H | 0.76 ± 0.03 | 4.01 ± 0.95 |
| 7 | H | CH2N3 | H | OH | OH | H | 9.17 ± 0.06 | 34.62 ± 1.06 |
| 8 | H | CH2COOH | H | OH | OH | H | 9.36 ± 0.05 | 33.73 ± 0.86 |
| 9 | H | H | H | OH | OH | CH2N(CH3)2 | 11.55 ± 0.09 | 53.6 ± 1.68 |
| 10 | H | H | CH2N(CH3)2 | OH | OH | H | 4.79 ± 0.04 | 19.92 ± 0.94 |
| 11 | H | H | H | OH | OH | CH2N(CH2)4CHOH | 4.81 ± 0.04 | 21.79 ± 1.06 |
| 12 | H | H | CH2N(CH2)4CHOH | OH | OH | H | 2.77 ± 0.05 | 14.69 ± 3.16 |
| 13 | H | H | H | OH | OH | CH2N(CH2)4 | 5.42 ± 0.04 | 27.97 ± 1.02 |
| 14 | H | CH3 | H | OH | OH | CH2N(CH3)2 | 1.46 ± 0.07 | 7.39 ± 1.65 |
| 15 | H | CH3 | H | OH | OH | CH2N(CH2)4CHOH | 21.26 ± 0.05 | 59.22 ± 0.62 |
| 16 | H | CH3 | H | OH | OH | CH2N(CH2)4 | 2.54 ± 0.06 | 11.34 ± 1.51 |
| 17 | H | H | H | OH | OCH3 | H | 15.87 ± 0.03 | 51.48 ± 0.52 |
| 18 | H | H | H | OCH3 | OH | H | 13.45 ± 0.08 | 22.59 ± 1.49 |
| 19 | H | H | H | OCH3 | OCH3 | H | 9.54 ± 0.08 | 25.14 ± 1.53 |
| 20 | NO2 | H | H | OCH3 | OCH3 | NO2 | 2.19 ± 0.05 | 12.21 ± 1.29 |
| 21 | H | H | H | H | OH | OH | 1.75 ± 0.03 | 11.77 ± 5.12 |
| 22 | H | CH3 | H | H | OH | OH | 2.11 ± 0.03 | 11.21 ± 1.57 |
| 23 | H | CH2Cl | H | H | OH | OH | 3.19 ± 0.08 | 18.61 ± 4.33 |
| 24 | H | CH2OH | H | H | OH | OH | 5.67 ± 0.11 | 32.23 ± 7.45 |
| 25 | H | H | H | H | OCH3 | OH | 7.02 ± 0.06 | 16.93 ± 4.70 |
| 26 | H | CH3 | H | H | OCH3 | OH | 2.39 ± 0.10 | 16.93 ± 4.70 |
| 27 | H | CH2Cl | H | H | OCH3 | OH | 6.59 ± 0.10 | 30.57 ± 6.83 |
| 28 | H | CH2OH | H | H | OCH3 | OH | 7.15 ± 0.11 | 32.89 ± 7.45 |
| 29 | H | CH3 | H | H | OH | H | 22.15 ± 0.28 | 103.70 ± 0.11 |
| 30 | H | CH3 | H | H | OH | NO2 | 10.75 ± 0.07 | 26.29 ± 4.14 |
| 31 | H | CH3 | H | Cl | OH | H | 5.26 ± 0.08 | 28.57 ± 5.47 |
| 32 | H | CH3 | H | CH2CH3 | OH | H | 5.47± 0.09 | 30.88 ± 6.12 |
| 33 | H | H | H | H | H | H | 4.06 ± 0.05 | 24.38 ± 4.94 |
| 34 | H | H | H | OH | H | H | 23.86 ± 0.18 | 83.69 ± 5.64 |
| 35 | H | H | H | H | OH | H | 3.81 ± 0.07 | 21.04 ± 4.41 |
| 36 | H | H | H | OCH3 | OH | OH | 5.51 ± 0.11 | 32.1 ± 6.90 |
| (−)-gossypol | Positive control | 0.19 ± 0.03 | 1.52±0.85 | |||||
Fig. 1.
Binding affinities of (A) Cpd 1, 33, 34, 35, (B) Cpd 1, 17, 18, 19, and (C) Cpd 1, 4, 8 to Mcl-l by FP assay.
For 6,7-dihydroxycoumarin derivatives, the order of Mcl-1 inhibitory activity among compounds with varying substituents in the 4-position was as follows: trifluoromethyl 4 (Ki = 0.21 ± 0.02 μM) > phenyl 3 (Ki = 0.65±0.03 μM) > hydroxymethyl 6 (Ki = 0.76 ±0.03 μM) > chloromethyl 5 (Ki = 1.04 ± 0.04 μM) > methyl 2 (Ki = 1.19 ± 0.04 μM) > hydrogen 1 (Ki = 1.49 ±0.04 μM) > azidomethyl 7 (Ki = 9.17 ±0.06 μM) ≈ carboxyl 8 (Ki = 9.36 ±0.05 μM). This trend indicated that hydrophobic and electron–withdrawing groups in 4-position of 6,7-dihydroxycoumarin 1 were beneficial for Mcl-1 inhibitory activity, while hydrophilic substitution in 4-position was unfavorable for the potency of Mcl-1inhibition. As shown in Fig. 1C, when 4-hydrogen of parent 1 was replaced by trifluoromethyl group or carboxyl group, the IC50 of parent 1 (8.77 ± 1.16 μM) was decreased to 1.21 ± 0.56 μM or increased to 33.7 ± 0.86 μM, respectively. In order to increase the metabolic stability of coumarin analogues, nitrogen-containing groups including dimethylamino, pyrrolidinyl and piperidinol as the hydrogen bond acceptor were introduced at the 5-position or 8-position of 6,7-dihydroxycoumarin derivatives.31 All the nitrogen-containing derivatives tested exhibited lower inhibitory activities than parent 1 except Cpd 14, among which more than 7-fold decrease of inhibitory activity was observed for 8-piperidinol-6,7-dihydroxycoumarin 15.
Similar trend in potency was also observed for 7,8-dihydroxycoumarin derivatives, as blockage of the catechol group by methylation was unbeneficial for Mcl-1inhibitory activity regardless of the types of substitution on 4-postion (21 (1.75 ± 0.03 μM) > 25 (7.02 ± 0.06 μM), 22 (2.11 ± 0.03 μM) > 26 (2.39 ± 0.10 μM), 23 (3.19 ± 0.08 μM) > 27 (6.59 ± 0.10 μM), 24 (5.67 ± 0.11 μM) > 28 (7.15 ±0.11 μM)). It’s worth noting that the substitutes at 4-position showed different effects on Mcl-1 inhibitory activity of 7,8-dihydroxycoumarins compared to that of 6,7-dihydroxycoumarins, with the following order of inhibitory potency: methyl 22, 26 > chloromethyl 23, 27 > hydroxymethyl 24, 28. Furthermore, methyl group introduced at C-4 position of 7-hydroxycoumarin 35 led to a 5-fold decrease in inhibitory activity of Mcl-1, while this decrease in inhibitory activity could be alleviated by introduction of a chlorine, ethyl or nitro group to the C-6 or C-8 position of compound 35. In addition, a known inhibitor gossypol was tested under identical conditions as a positive control.36 The result indicated that the most potent Mcl-1 inhibitor obtained, Cpd 4, possessed comparable inhibitory activity against Mcl-1 to that of the positive control (Table 1). The structure-activity relationships (SAR) of these coumarin derivatives as Mcl-1 inhibitors are summarized in Fig. 2, which could be helpful for medical chemists to design and develop more potent Mcl-1 inhibitors for biomedical applications.
Fig. 2.
SAR summary of coumarin derivatives.
2.3. Cytotoxicity assay
Most solid tumor malignancies show high expression levels of Mcl-1and Mcl-1 levels are correlated with resistance to anti-cancer therapeutics. A549 cells, which also express relatively high levels of Mcl-1, demonstrated resistance to taxol, a widely prescribed microtubule-targeted agent.37 To validate whether the coumarin type Mcl-1 inhibitors has the ability to enhance the apoptotic response induced by other anti-cancer drugs known to prime cells for apoptosis, here we investigated the effect of taxol and Cpd 4, the most potent Mcl-1 inhibitor, alone or in combination on the viability of A549 cell lines. As shown in Fig. 3, cell killing activity of taxol toward A549 cell lines was significantly enhanced by combination with Cpd 4, with a more than 10 fold decreased IC50 value (0.0025 μM) observed for the combination compared with that for taxol alone (0.034 μM). These results demonstrated that Cpd 4 is potentially useful as a reversal agent in lung cancer therapy.
Fig. 3.
The IC50 concentrations of taxol or/and Cpd 4 detected in A549 cell lines and the parental cell lines.
2.4. Molecular Docking Analysis
To gain a deeper understanding of the interaction mode between Mcl-1 and the coumarins, the most potent Mcl-1 inhibitor-Cpd 4 with a Ki value of 0.204 μM, was selected and subjected to docking at the hydrophobic binding pocket of Mcl-1. 79 low-energy conformational poses were generated from docking runs and the best conformation was selected based on the docking score after investigation of ensemble of ligand poses. Good alignment was observed between the crystal ligand Mcl-1 and the low-energy conformational pose of Cpd 4. The docked ligand Cpd 4 was surrounded by Met231, Leu235, Leu246, Val249, Met250, Val253, Phe254, Thr266, Leu267, Phe270, Gly271, Val274, Leu290, and Ile294 residues at the Mcl-1 binding pocket which cover a larger area of interaction (Fig. 4A). A similar binding region of Mcl-1 was also reported previously when N-substituted 1-hydroxy-4-sulfamoyl-2-naphthoate compounds were docked into the crystal structure of Mcl-1.38
Fig. 4.
Docking simulation of the crystal structure of Mcl-1 and the stereo diagram of the ligand Cpd 4 aligned in its active site. (A) The possible binding pose of Cpd 4 in Mcl-1 active site. (B) The interactions (dotted lines with distances in Å) between the highly active compound- Cpd 4-and binding site residues of Mcl-1.
Further graphical inspection of bonded and non-bonded contacts shows that the π–π interactions stabilizes the predicted low-energy conformational pose of Cpd 4 inside the binding pocket of Mcl-1. As shown in Fig. 4B, the π– π interactions take place between A ring and B ring of Cpd 4 that insert deep into the binding pocket with the side-chain benzene ring of Phe270, enhancing the complex’s stability by helping anchor Cpd 4 in place. The distances between residues involved in the π–π interactions are also depicted in Fig. 4B. Despite the strong π– π interactions, the predicted low-energy conformational pose of Cpd 4 was further stabilized by a series of interactions between A and B rings of Cpd 4 and the alkyl groups of Met231 and Met250. Furthermore, the rest of the surrounding hydrophobic residues provide additional strength for complex stability via non-bonded interactions. All these interactions may contribute to the relatively high binding affinity of Cpd 4 to Mcl-1 protein observed in FP assay. These results of docking simulation are also in good agreement with the SAR analysis, given that both the hydroxyl in B ring and the hydrophobic groups in A ring could be beneficial for Mcl-1 inhibition as these substituents could interact favorably with specific residues of Mcl-1 protein. In addition, to explore why the introduction of 6-hydroxyl or 7-hydroxyl alone showed little impact on the Mcl-1inhibitory activity of coumarin while the introduction of 6,7-dihydroxyls led to a significant improvement of inhibitory activity of coumarin towards Mcl-1, Cpd 33, Cpd 34, Cpd 35 and Cpd 1 were docked to the binding pocket of Mcl-1. As shown in Fig. S1 and Table S1, the more favored conformation formed between Cpd 1 and Mcl-1indicated by Libdock score and binding energy, as well as the relatively shorter interaction distances between residues involved in the bonded interactions for Cpd 1 compared to those for the other three coumarins, may contribute to the significantly improved Mcl-1 inhibitory activity for Cpd 1.
2.5. Establishment of 3D QSAR model
Quantitative Structure-Activity Relationship (QSAR) studies have been extensively applied to explore the correlations between biological activities and molecular descriptors for different classes of compounds.39,40 In this investigation, 3D QSAR models were built using Drug Discovery Studio to acquire a systematic SAR profile on the coumarin derivates as Mcl-1 inhibitors. Among 36 coumarin-type Mcl-1 inhibitors, the training and test sets were chosen to develop the 3D QSAR model (Fig. 5). Coefficient determination (r2), external validation coefficient (q2) and root mean squared error (RMSE) of 0.977, 0.968 and 0.392 were achieved for training set, and r2 of 0.672, q2 of 0.633 and RMSE of 0.501 were obtained for test set, indicating that the built 3D QSAR model was acceptable.
Fig. 5.
Plots of experimental vs. predicted Mcl-1 inhibitory activities of training set and test set. (A) Plot for training set. (B) Plot for test set.
Also, the compounds aligned with the iso-surfaces of the 3D QSAR model coefficients on van der Waals grids and electrostatic potential grids are shown in Fig. 6. In the electrostatic map, red contours around regions where high electron density (negative charge) is favorable for Mcl-1 inhibition, and blue contours represent areas where low electron density (partial positive charge) is expected to increase activity (Fig. 6A). Similarly, the green-yellow steric contours depicted in Fig. 6B illustrate areas where steric bulk is predicted to increase (green) or decrease (yellow) activity. The resulting mappings suggested that both 3-D steric and electronic interactions could strongly affect the inhibitory effects of coumarins on Mcl-1. Notably, these findings agreed well with the experimental data, such as the substitutional groups on the aromatic B ring or heterocyclic A ring ask for high negative charged groups, and the small substitutional groups on the two rings are suitable. These findings provide new insights into the fine correlations between the inhibitory effects against Mcl-1 and the steric-electrostatic properties of coumarins, which could be helpful for the rational design of novel coumarins as potent Mcl-1 inhibitors.
Fig. 6.
(A) 3D-QSAR model coefficients on electrostatic potential grids. Blue represents positive coefficients; red represents negative coefficients. (B) 3D-QSAR model coefficients on van der Waals grids. Green represents positive coefficients; yellow represents negative coefficients.
3. Conclusion
In summary, more than thirty coumarin derivatives with different substituents were designed and synthesized. The Mcl-1 inhibitory activities of these compounds were assayed and the potential structure–inhibition relationships of coumarin derivatives were summarized and discussed. The results demonstrated that the catechol group in the coumarin scaffold was a key group for Mcl-1 inhibitory activity, while introduction of a hydrophobic and electron-withdrawing group at the C-4 position of coumarins was beneficial for the Mcl-1 inhibition effect. Among all the tested compounds, 4-trifluoromethyl-6,7-dihydroxycoumarin (Cpd 4) exhibited the strongest inhibitory activity against Mcl-1, and its beneficial effect on taxol resistance was validated in A549 cell lines. The strong interaction between Cpd 4 and Mcl-1 protein in docking simulation further supported the observed potent Mcl-1 inhibitory ability of Cpd 4. Meanwhile, a 3D-QSAR model was well established for these coumarin derivatives, and the results demonstrated that the steric bulk and the electrostatic potential of substituted groups in the coumarin scaffold were key determinants for Mcl-1 inhibitory activity. Among all the tested compounds, Cpd 4 exhibited the strongest Mcl-1 inhibitory activity, which meet all the preferred structural requirements according to the 3D-QSAR model. All these findings could be helpful for medicinal chemists to design and develop potent anticancer agents by targeting Mcl-1 based on the structural modifications of coumarin derivatives.
4. Materials and methods
4.1. Reagents and materials
A Bid BH3 peptide of 21-residue (residues 79–99) with a 6-carboxy-fluorescein succinimidyl ester fluorescence tag (FAM-Bid) was synthesized by HD Biosciences (Shanghai, China). Recombinant Mcl-1 protein Escherichia coli BL21 was synthesized and purified following methods described earlier.41,42
4.2. Chemicals and Measurements
The 1H NMR and 13C NMR spectra were recorded on a Bruker ARX (Bruker, Rheinstetten, Germany) 400 MHz or Bruker AMX 400 MHz or 600 MHz spectrometer in dimethyl sulfoxide (DMSO-d6) if not noted otherwise, and the chemical shifts were expressed as ppm using trimethylsilane (TMS) as an internal reference. High-resolution mass spectral (HRMS) analyses were measured on a TripleTOF™ Mass Spectrometer (TripleTOF 5600, SCIEX, Foster City, USA). Coumarin derivatives were analyzed using an ultra-fast liquid chromatography spectrometry system (Shimadzu, Kyoto, Japan) equipped with two LC-20AD pumps, a DGU-20A3 vacuum degasser, a SIL-20ACHT auto-sampler, a CTO-20AC column oven, and an SPD-M 20A diode-array detector (DAD). All reagents used in the synthesis were obtained commercially and used without further purification. The reactions were monitored using thin layer chromatography (TLC) on glass packed precoated silica gel GF254 plates. Flash column chromatography was performed using silica gel (200–300 mesh), which was purchased from the Qingdao Ocean Chemical Co. Ltd (Qingdao, China). Compounds 1, 21, 25, 33, 34, 35 and 36 were purchased from Chengdu Pufei De Biotech Co. Ltd. (Sichuan, China).
General procedure for the synthesis of 4-substituted esculetins (2 ~ 5).
To a mixture of 1,2,4-phenenyl triacetate (5.0 mmol) and properly substituted β-ketoesters (10 mmol) was added drop-wise perchloric acid (5.0 mL) at room temperature and stirred for 6 ~ 8 h. After completion of the reaction, as indicated by TLC monitoring, the reaction mixture was poured slowly into 100 mL ice-water with stirring. The resultant suspension was filtered and the collected solid washed with water and dried, then the crude compound was recrystallized from methanol.
6,7-dihydroxy-4-methyl-2H-chromen-2-one (2).
According to the general procedure, 1,2,4-phenenyl triacetate was reacted with ethyl acetoacetate in the presence of perchloric acid, and crude compound was recrystallized from methanol to produce compound 2 as light white solid, 87% yield. 1H NMR (400 MHz, DMSO-d6) δ: 2.31 (s, 3H, CH3), 6.09 (s, 1H, COCH=C), 6.73 (s, 1H, ArH), 7.00 (s, 1H, ArH), 9.34 (s, br, 1H, ArOH), 10.19 (s, br, 1H, ArOH); 13C NMR (100 MHz, DMSO-d6) δ: 18.68, 103.13, 109.89, 110.86, 111.96, 143.24, 148.17, 150.58, 153.68, 161.08; ESI-MS: M = 192, found 191.0 [M-H]-.
6,7-dihydroxy-4-phenyl-2H-chromen-2-one (3).
According to the general procedure, 1,2,4-phenenyl triacetate was reacted with ethyl benzoylacetate in the presence of perchloric acid, and crude compound was recrystallized from methanol to generate compound 3 as white solid, 91% yield. 1H NMR (400 MHz, DMSO-d6) δ: 6.12 (s, 1H, COCH=C), 6.80 (s, 1H, ArH), 6.84 (s, 1H, ArH), 7.50–7.58 (m, 5H, ArH) 9.43 (s, br, 1H, ArOH), 10.26 (s, br, 1H, ArOH); ESI-MS: M = 254, found 253.0 [M-H]-.
6,7-dihydroxy-4-trifluoromethyl-2H-chromen-2-one (4).
According to the general procedure, 1,2,4-phenenyl triacetate was reacted with ethyl 4,4,4-trifluoroacetoacetate in the presence of perchloric acid, and crude compound was recrystallized from methanol to produce compound 4 as light yellow solid, 63% yield.1H NMR (400 MHz, DMSO-d6) δ: 6.71 (s, 1H, COCH=C), 6.86 (s, 1H, ArH), 7.03 (s, 1H, ArH), 9.78 (s, br, 1H, ArOH), 10.61 (s, br, 1H, ArOH); 13C NMR (100 MHz, DMSO-d6) δ: 104.01, 105.05, 108.89, 112.12, 120.94, 123.68, 144.03, 149.59, 152.08, 159.62; ESI-MS: M = 246, found 245.1 [M-H]-.
6,7-dihydroxy-4-chloromethyl-2H-chromen-2-one (5).
According to the general procedure, 1,2,4-phenenyl triacetate was reacted with ethyl 4-chloroacetoacetate in the presence of perchloric acid, and crude compound was recrystallized from methanol to generate compound 5 as white solid, 78% yield. 1H NMR (400 MHz, DMSO-d6) δ: 4.90 (s, 2H, CH2Cl), 6.40 (s, 1H, COCH=C), 6.78 (s, 1H, ArH), 7.12 (s, 1H, ArH), 9.48 (s, br, 1H, ArOH), 10.39 (s, br, 1H, ArOH). ESI-MS: M = 225.5, found 224.9 [M-H]- and 226.9 [M-H]-.
6,7-dihydroxy-4-hydroxymethyl-2H-chromen-2-one (6).
Chloride 5 (0.5 g, 2.2 mmol) was dissolved in a mixture of DMF (3 ml) and H2O (10 ml) with stirring and then refluxed for 20 h. After completion of the reaction as indicated by TLC, the mixture was diluted with water. The resultant suspension was filtered and the collected solid was washed with water and dried, then crude compound was recrystallized from methanol to produce hydroxymethyl 6 (380 mg, 83%) as light brown solid. 1H NMR (400 MHz, DMSO-d6) δ: 4.63 (s, 2H, CH2OH), 6.21 (s, 1H, COCH=C), 6.75 (s, 1H, ArH), 6.94 (s, 1H, ArH), 9.31 (s, br, 1H, ArOH), 10.19 (s, br, 1H, ArOH); ESI-MS: M = 208, found 208.9 [M+H]+.
4-(azidomethyl)-6,7-dihydroxy-2H-chromen-2-one (7).
Chloride 5 (0.5 g, 2.2 mmol) was dissolved in a mixture of DMF (6.0 ml), then NaN3 (0.3 g 4.5 mmol) was added and stirred overnight. After completion of the reaction as indicated by LC-MS, the mixture was diluted with water. The resultant suspension was filtered and the collected solid was washed with water and dried, then crude compound was recrystallized from methanol to generate azidomethyl 7 in 90% yield as light brown solid. 1H NMR (400 MHz, d-DMSO) δ: 4.73 (s, 2H, CH2N3), 6.26 (s, 1H, COCH=C), 6.78 (s, 1H, ArH), 6.98 (s, 1H, ArH), 9.49 (s, br, 1H, ArOH), 10.38 (s, br, 1H, ArOH). ESI-MS: M = 233, found 234.2 [M+H]+.
2-(7,8-dihydroxy-2-oxo-2H-chromen-4-yl)acetic acid (8).
According to the general procedure mentioned above, 1,2,4-phenenyl triacetate was reacted with 1,3-acetonedicarboxylic acid in the presence of perchloric acid, and crude compound was recrystallized from methanol to produce compound 8 in 70 % yield as white solid. 1H NMR (400 MHz, DMSO) δ 12.75 (s, 1H), 10.11 (s, 1H), 9.35 (s, 1H), 7.03 (d, J = 8.4 Hz, 1H), 6.81 (d, J = 8.4 Hz, 1H), 6.22 (s, 1H), 3.80 (s, 2H). 13C NMR (101MHz, DMSO) δ: 37.88, 112.43,112.58,112.71, 116.03, 132.74, 143.95, 149.93, 150.99, 160.63,171.23; ESI-MS: M = 236, found 235.1 [M-H]-.
General procedure for the synthesis of 5/8-substituted esculetin Mannich bases (9 ~ 16).
To a solution of the appropriate amines (0.013 mol) in 30 mL of absolute methanol was added aqueous formaldehyde (37%) (0.015 mol). After 0.5 h of gentle refluxing at 50°C, esculetin/4-substituted esculetin was added (0.01 mol), dissolved in 50 mL of absolute methanol. The duration of the reaction was 8 ~ 12 h and the reaction was monitored by TLC.
6,7-dihydroxy-8-((dimethylamino)methyl)-2H-chromen-2-one (9) and 6,7-dihydroxy-5-((dimethylamino)methyl)-2H-chromen-2-one (10).
According to the general procedure, esculetin was reacted with aqueous formaldehyde and aqueous dimethylamine. After completion of the reaction as indicated by TLC, the solvent was removed under reduced pressure. The residue was applied to flash column chromatography (silicagel) to generate compound 9 and compound 10 with the yield of 17% and 6%, respectively.
Compound 9: 1H NMR (400 MHz, DMSO-d6) δ: 2.49 (s, 6H, 2CH3), 4.03 (s, 2H, CH2), 6.03 (d, J=8.0 Hz, 1H, COCH=C), 6.86 (s, 1H, ArH), 7.79 (d, J=12.0 Hz, 1H, PhCH=C), 8.16 (s, 1H, ArOH); ESI-MS: M = 235, found 236.1 [M+H]+.
Compound 10: 1H NMR (400 MHz, d-DMSO) δ: 2.25 (s, 6H, 2CH3), 3.81 (s, 2H, CH2), 6.17 (d, J=12 Hz, 1H, COCH=C), 6.70 (s, 1H, ArH), 8.10 (d, J=12 Hz, 1H, PhCH=C), 8.15 (s, 1H, ArOH), ESI-MS: M = 235, found 236.1 [M+H]+.
6,7-dihydroxy-8-((4-hydroxypiperidin-1-yl)methyl)-2H-chromen-2-one (11) and 5-((4-hydroxypiperidin-1-yl)methyl)-6,7-dihydroxy-2H-chromen-2-one (12).
According to the general procedure, esculetin was reacted with aqueous formaldehyde and 4-hydroxy piperidine. After completion of the reaction as indicated by TLC, the solvent was removed under reduced pressure. The residue was applied to flash column chromatography (silicagel) to produce compound 11 and compound 12 with the yield of 12% and 3%, respectively.
Compound 11: 1H NMR (400 MHz, DMSO-d6) δ: 1.48 (m, 2H, CH2), 1.82 (m, 2H, CH2), 2.49 (m, 2H, NCH2), 2.91 (m, 2H, NCH2), 3.62 (m, 1H, CHOH), 4.00 (s, 2H, CH2), 6.08 (d, J=8 Hz, 1H, COCH=C), 6.89 (s, 1H, ArH),7.81 (d, J=8 Hz, 1H, PhCH=C); 13C NMR (100 MHz, DMSO-d6) δ: 33.50, 49.72, 52.56, 106.77, 108.87, 109.71, 110.38, 142.79, 144.87, 146.79, 153.60, 160.62; ESI-MS: M = 291, found 292.1 [M+H]+; HRMS (ESI): C15H17NO5 calcd for [M + H]+: 292.1185, found: 292.1181.
Compound 12: 1H NMR (400 MHz, DMSO-d6) δ: 1.68 (m, 2H, CH2), 1.92 (m, 2H, CH2), 3.09 (m, 2H, NCH2), 3.29 (m, 2H, NCH2), 3.76 (m, 1H, CHOH), 4.44 (s, 2H, CH2), 6.27 (d, J=8 Hz, 1H, COCH=C), 6.97 (s, 1H, ArH), 8.33 (d, J=8 Hz, 1H, PhCH=C); 13C NMR (100 MHz, DMSO-d6) δ: 30.38, 48.53, 49.61, 103.75, 110.68, 111.56, 112.92, 141.95, 143.54, 148.89, 150.28, 160.18; ESI-MS: M = 291, found 292.1 [M+H]+; HRMS (ESI): C15H17NO5 calcd for [M + H]+: 292.1185, found: 292.1183.
6,7-dihydroxy-8-(pyrrolidin-1-ylmethyl)-2H-chromen-2-one (13).
According to the general procedure, esculetin was reacted with aqueous formaldehyde and pyrrolidine. After completion of the reaction as indicated by TLC, the solvent was removed under reduced pressure. The residue was applied to flash column chromatography (silicagel) to afford compound 13 with the yield of 18 % as light brown solid. 1H NMR (400 MHz, DMSO-d6) δ: 1.85 (m, 4H, CH2 CH2), 2.88 (t, J = 4.8 Hz 4H, NCH2 CH2), 4.17 (s, 2H, CH2), 5.97 (d, J=8.0 Hz, 1H, COCH=C), 6.82 (s, 1H, ArH), 7.76 (d, J=8.0 Hz, 1H, PhCH=C); 13C NMR (100 MHz, DMSO-d6) δ: 23.05 (2C), 49.88, 52.84(2C), 105.98, 107.16, 107.75, 109.05, 143.49, 144.90, 147.58, 156.57, 160.86; ESI-MS: M = 261, found 262.1 [M+H]+; HRMS (ESI): C14H15NO4 calcd for [M + H]+: 262.1079, found: 262.1072.
8-((dimethylamino)methyl)-4-methyl-2H-chromen-2-one (14).
According to the general procedure, 4-methyl esculetin was reacted with aqueous formaldehyde and aqueous dimethylamine. After completion of the reaction, as indicated by TLC, the mixture was diluted with water. The resultant suspension was filtered and the collected solid washed with water and dried, then crude compound was recrystallized from methanol to generate compound 14 as yellow solid with 78% yield. 1H NMR (400 MHz, DMSO-d6) δ: 2.30 (s, 3H, CH3), 2.42 (s, 6H, 2CH3), 4.00 (s, 2H, CH2), 5.99 (s, 1H, COCH=C), 6.92 (s, 1H, ArH);13C NMR (100 MHz, DMSO-d6) δ: 18.87, 43.83, 54.51, 106.93, 107.48, 108.82, 109.27, 143.39, 146.82, 154.21, 155.03, 160.98; ESI-MS: M = 249, found 250.0 [M+H]+; HRMS (ESI): C13H15NO4 calcd for [M + H]+: 250.1079, found: 250.1074.
6,7-dihydroxy-8-((4-hydroxypiperidin-1-yl)methyl)-4-methyl-2H-chromen-2-one (15).
According to the general procedure, 4-methyl esculetin was reacted with aqueous formaldehyde and 4-hydroxy piperidine. After completion of the reaction as indicated by TLC, the mixture was diluted with water. The resultant suspension was filtered and the collected solid washed with water and dried, then crude compound was recrystallized from methanol to produce compound 15 as light yellow solid with 80% yield. 1H NMR (400 MHz, DMSO-d6) δ: 1.47 (m, 2H, CH2), 1.81 (m, 2H, CH2), 2.31 (s, 1H, CH3), 2.45 (m, 2H, NCH2), 2.87 (m, 2H, NCH2), 3.61 (m, 1H, CHOH), 3.99 (s, 2H, CH2), 6.04 (s, 1H, COCH=C), 6.94 (s, 1H, ArH); 13C NMR (100 MHz, DMSO-d6) δ: 18.30, 33.61, 49.77, 52.88, 107.02, 107.58, 109.18, 109.77, 142.49, 145.82, 152.80, 153.61, 160.32; ESI-MS: M = 305, found 306.1 [M+H]+; HRMS (ESI): C16H19NO5 calcd for [M + H]+: 306.1341, found: 306.1337.
6,7-dihydroxy-8-(pyrrolidin-1-ylmethyl)-4-methyl-2H-chromen-2-one (16).
According to the general procedure, 4-methyl esculetin was reacted with aqueous formaldehyde and pyrrolidine. After completion of the reaction as indicated by TLC, the mixture was diluted with water. The resultant suspension was filtered and the collected solid washed with water and dried, then crude compound was recrystallized from methanol to afford compound 16 as yellow solid with 88% yield.1H NMR (400 MHz, DMSO-d6) δ: 1.84 (m, 4H, 2CH2), 2.29 (s, 3H, CH3), 2.83 (m, 4H, 2NCH2), 4.15 (s, 2H, CH2), 5.96 (s, 1H, COCH=C), 6.89 (s, 1H, ArH); 13C NMR (100 MHz, DMSO-d6) δ: 18.89, 23.60, 50.17, 53.33, 106.97, 108.32, 108.70, 143.65, 146.90, 154.23, 155.87, 161.07; ESI-MS: M = 275, found 276.1 [M+H]+; HRMS (ESI): C15H17NO4 calcd for [M + H]+: 276.1236, found: 276.1230.
6-hydroxy-7-methoxy coumarin (17).
To the solution of 6,7-dihydroxycoumarins (500 mg, 2.81 mmol) in DMF, Na2CO3 (3 eq.) was added at 15°C and stirred for 0.5 h. iodomethane (525 μL) was added dropwise to the reaction mixture at 15°C, maintaining the temperature at 20°C for 4 h. The reaction mixture was poured into water-ice and acidified with 2N hydrochloric acid. The reaction mixture was extracted with ethyl acetate (50 mL x 3). The combined organic layer was washed with water and brine, dried over anhydrous sodium sulfate, and evaporated in vacuo. The residue was purified by column chromatography using silica gel with petroleum ether- dichloromethane- acetone as mobile phase to obtain compound 17 in 75% yield as white solid. 1H NMR (400 MHz, DMSO-d6) δ 3.82 (s, 3H, OCH3), 6.23 (d, J = 9.5 Hz, 1H, COCH=C), 7.00 (s, 2H, Ar-H), 7.89 (d, J = 9.5 Hz, 1H, C=CH). M = 192, found 193 [M+H]+.
6-methoxy-7-hydroxycoumarin (18).
To the solution of 6,7-dihydroxycoumarins (500 mg, 2.81 mmol) in DMF, NaH (60% suspension in oil) (7.0 mmol) was added at 0°C under dry argon and allowed to stir for 0.5 h for the anion generation. iodomethane (210 μL) was added dropwise to the reaction mixture, maintaining the temperature at 0–5°C until the reaction was completed, as indicated by TLC (VDCM: VMeOH = 50: 1). The reaction mixture was poured into water-ice and acidified with 2N hydrochloric acid. The resultant suspension was filtered and the collected solid was washed with water and dried, then crude compound was recrystallized from methanol to afford compound 18 in about 90% yield as white solid. 1H NMR (400 MHz, DMSO-d6) δ 3.82 (s, 3H, OCH3), 6.22 (d, J = 9.5 Hz, 1H, COCH=C), 6.78 (s, 1H, Ar-H), 7.22 (s, 1H, Ar-H), 7.91 (d, J = 9.5 Hz, 1H, C=CH), 10.29 (d, br, 1H, Ar-OH). M = 192, found 191.1 [M-H]-, found 193 [M+H]+.
6,7-dimethoxylcoumarin (19).
To the solution of 6,7-dihydroxycoumarin (500 mg, 2.81 mmol) in DMF, NaH (60% suspension in oil) (7.0 mmol) was added at 0°C under dry argon and allowed to stir for 0.5 h for the anion generation. iodomethane (300 μL) was added dropwise to the reaction mixture and stirred for 5 h at room temperature. The reaction mixture was poured into water-ice and acidified with 2N hydrochloric acid. The resultant suspension was filtered and the collected solid was washed with water and dried, then crude compound was recrystallized from methanol to produce compound 19 in about 95% yield as white solid. 1H NMR (400 MHz, d-DMSO) δ: 3.80 (s, 3H, CH3O), 3.86 (s, 3H, CH3O), 6.29 (d, J = 8.0 Hz, 1H, COCH=C), 7.06 (s, 1H, ArH), 7.25 (s, 1H, ArH), 7.95 (d, J = 8.0 Hz, 1H, PhCH=C).13C NMR (100 MHz, d-DMSO) δ: 56.34, 56.63, 100.50, 109.38, 111.64, 113.11, 144.79, 146.31, 149.88, 152.99, 161.01. ESI-MS: M = 206, found 207.1 [M+H]+.
6,7-dimethoxy-3,8-dinitro-2H-chromen-2-one (20).
To a solution of 6,7-dimethoxylcoumarin 19 (0.5 g, 2.4 mmol) in DCM (10.0 ml) was added drop-wise a mixture of concentrated sulfuric acid and fuming nitric acid (4 ml) at 0°C, After completion of the reaction as indicated by TLC, the reaction mixture was poured slowly into a mixture of ice-water (50 ml) with stirring. The resultant suspension was filtered and the collected solid was washed with water and dried, then the crude compound was recrystallized from methanol to afford 20 as yellow solid. 1H NMR (400 MHz, d-DMSO) δ: 3.91 (s, 3H, CH3O), 4.01 (s, 3H, CH3O), 7.62 (s, 1H, ArH), 8.77 (s, 1H, PhCH=C). 13C NMR (100 MHz, d-DMSO) δ: 58.60, 62.99, 102.69, 104.28, 134.56, 137.36, 139.70, 142.02, 151.78, 152.65, 159.79. HRMS: M=296, found 327.0461 [M+MeOH-H]−.
7,8-dihydroxy-4-methyl-2H-chromen-2-one (22).
To a mixture of 1,2,3-phenenyl triacetate (5.0 mmol) and ethyl acetoacetate (7.5 mmol) was added drop-wise perchloric acid (6.0 ml) at room temperature and stirred for 6 h. After completion of the reaction as indicated by TLC, the reaction mixture was poured slowly into a mixture of ice-water (50 ml) with stirring. The resultant suspension was filtered and the collected solid was washed with water and dried, then the crude compound was recrystallized from methanol to produce compound 22 as a light white solid with a yield of 78%. 1H NMR (600 MHz, DMSO) δ 10.05 (s, 1H), 9.28 (s, 1H), 7.09 (d, J = 8.6 Hz, 1H), 6.82 (d, J = 8.6 Hz, 1H), 6.13 (d, J = 1.1 Hz, 1H), 2.36 (s, 3H); 13C NMR (151 MHz, DMSO) δ 160.67, 154.40, 149.87, 143.79, 132.62, 115.95, 113.23, 112.58, 110.65, 18.72; HRMS (ESI) for C10H8O4, Calcd 192.0423, found 192.0355 [M-H]-.
7,8-dihydroxy-4-chloromethyl-2H-chromen-2-one (23).
To a mixture of 1,2,3-phenenyl triacetate (5.0 mmol) and ethyl 4-chloroacetoacetate (7.5 mmol) was added drop-wise perchloric acid (6.0 ml) at room temperature and stirred for 6 h. After completion of the reaction as indicated by TLC, the reaction mixture was poured slowly into a mixture of ice-water (50 ml) with stirring. The resultant suspension was filtered and the collected solid was washed with water and dried, then the crude compound was recrystallized from methanol to produce compound 23 as a white solid with a yield of 69%. 1H NMR (600 MHz, DMSO) δ 10.19 (s, 1H), 9.40 (s, 1H), 7.18 (d, J = 8.7 Hz, 1H), 6.85 (d, J = 8.7 Hz, 1H), 6.42 (s, 1H), 4.94 (s, 2H); 13C NMR (151 MHz, DMSO) δ 160.55, 151.86, 150.23, 144.15, 132.92, 115.95, 112.79, 111.42, 110.57, 41.95; HRMS (ESI) for C10H7ClO4, Calcd 226.0033, found 224.9967 [M-H]-.
7,8-dihydroxy-4-hydroxymethyl-2H-chromen-2-one (24).
Chloride C23 (0.5 g, 2.2 mmol) was dissolved in a mixture of DMF (3 mL) and H2O (10 mL) with stirring and refluxed for 20 h. After TLC indicated that the reaction was complete, the mixture was diluted with water. The resultant suspension was filtered, and the collected solid was washed with water and dried. The crude compound was recrystallized from methanol to yield hydroxymethyl 24 as a light brown solid with a yield of 80%. 1H NMR (600 MHz, DMSO) δ 10.03 (s, 1H), 9.30 (s, 1H), 7.01 (d, J = 8.7 Hz, 1H), 6.78 (d, J = 8.6 Hz, 1H), 6.24 (t, J = 1.4 Hz, 1H), 5.55 (t, J = 5.5 Hz, 1H), 4.69 (d, J = 4.2 Hz, 2H); 13C NMR (151 MHz, DMSO) δ 161.03, 157.70, 149.75, 143.82, 132.75, 114.75, 112.65, 110.73, 106.85, 59.56; HRMS (ESI) for C10H8O5, Calcd 208.0372, found 207.0304 [M-H]-.
8-hydroxy-7-methoxy-4-methylcoumarin (26).
To a mixture of 3-methoxybenzene-1,2-diol (0.5 g, 3.5 mmol) and ethyl acetoacetate (0.95 ml, 7.0 mmol) was added drop-wise perchloric acid (3.0 ml) at room temperature and stirred for 6 h. After completion of the reaction as indicated by TLC, the reaction mixture was poured slowly into a mixture of ice-water (50 ml) with stirring. The resultant suspension was filtered and the collected solid was washed with water and dried, then the crude compound was recrystallized from methanol to produce 26 as light white solid. 1H NMR, δ: 2.38 (s, 3H, CH3), 3.89 (s, 3H, OCH3), 6.20 (s, 1H, COCH=C), 7.03 (d, J = 8 Hz, 1H, Ar-H), 7.21 (d, J = 8 Hz, 1H, Ar-H), 9.38 (s, 1H, OH). 13C NMR, δ: 160.4, 154.3, 154.1, 148.1, 134.7, 120.9, 113.5, 113.4, 110.8, 61.1, 18.7. M = 206, found 205.1 [M-H]-, found 206.9 [M+H]+. IR (KBr), ṽ/cm–1: 3552, 3447, 3362, 2968, 2840, 1663, 1613, 1574, 1511, 1457, 1444, 1385.
4-(chloromethyl)-8-hydroxy-7-methoxy-2H-chromen-2-one (27).
To a mixture of 3-methoxybenzene-1,2-diol (0.5 g, 3.5 mmol) and ethyl 4-chloroacetoacetate (1.0 ml, 7.0 mmol) was added drop-wise perchloric acid (5.0 ml) at room temperature and stirred for 6 h. After completion of the reaction as indicated by TLC, the reaction mixture was poured slowly into a mixture of ice-water (50 ml) with stirring. The resultant suspension was filtered and the collected solid was washed with water and dried, then the crude compound was recrystallized from methanol to generate 27 as light white solid. 1H NMR (400 MHz, d-DMSO) δ: 3.90 (s, 3H, OCH3), 4.97 (s, 2H, CH2Cl), 6.48 (s, 1H, COCH=C), 7.08 (d, J = 8 Hz, 1H, ArH), 7.30 (d, J = 8 Hz, 1H, ArH), 9.51 (s, br, 1H, ArOH); 13C NMR (100 MHz, d-DMSO) δ: 41.90, 56.79, 108.87, 111.88, 112.67, 115.63, 134.13, 143.20, 151.34, 151.60, 160.36. ESI-MS: m/z 241 and 243 [M+H]−
8-hydroxy-7-methoxyl-4-hydroxymethyl-2H-chromen-2-one (28).
Chloride 27 (0.4 g, 1.7 mmol) was dissolved in a mixture of DMF (3 mL) and H2O (10 mL) with stirring and refluxed for 20 h. After TLC indicated that the reaction was complete, the mixture was diluted with water. The resultant suspension was filtered, and the collected solid was washed with water and dried. The crude compound was recrystallized from methanol to yield hydroxymethyl 28 as a light brown solid with a yield of 83%.1H NMR (400 MHz, d-DMSO) δ: 3.87 (s, 3H, OCH3), 4.70 (d, J = 4 Hz, 2H, CH2), 5.59 (s, 1H, OH),6.30 (s, 1H, COCH=C), 6.96 (d, J = 8 Hz, 1H, ArH), 7.09 (d, J = 8 Hz, 1H, ArH), 9.39 (s, br, 1H, ArOH); 13C NMR (100 MHz, d-DMSO) δ: 56.68, 59.56, 108.07, 108.67, 112.10, 114.39, 133.98, 142.88, 150.93, 157.35, 160.87. ESI-MS: m/z 223 [M+H]+
4-methyl-7-hydroxy-8-nitrocoumarin (30).
To a stirred solution of 4-methyl-7-hydroxy coumarin (1.5 g, 8.5 mmol) in concentrated sulphuric acid (30 mL), a solution of concentrated nitric acid (6.0 mL) in concentrated sulphuric acid (6.0 mL) was added at such a rate as to keep the temperature below 5 oC. After warming to 20 oC, the reaction mixture was poured into a stirred mixture of ice cold water. The yellow solid that separated was filtered and thoroughly washed with water. The product was recrystalized first from glacial acetic acid and then methanol to obtain compound 30 in 78% yield. 1H NMR (400 MHz, DMSO) δ 12.18 (s, 1H), 7.81 (d, J = 9.0 Hz, 1H), 7.05 (d, J = 8.9 Hz, 1H), 6.32 (d, J = 1.1 Hz, 1H), 2.41 (d, J = 1.1 Hz, 3H). 13C NMR (101 MHz, DMSO) δ 158.57, 154.05, 152.86, 146.09, 128.62, 113.52, 112.46, 111.69, 18.74.
6-chloro-7-hydroxy-4-methyl-2H-chromen-2-one (31).
To a mixture of 4-chlorobenzene-1,3-diol (5.0 mmol) and ethyl acetoacetate (7.5 mmol) was added drop-wise perchloric acid (6.0 ml) at room temperature and stirred for 6 h. After completion of the reaction as indicated by TLC, the reaction mixture was poured slowly into a mixture of ice-water (50 ml) with stirring. The resultant suspension was filtered and the collected solid was washed with water and dried, then the crude compound was recrystallized from methanol to produce compound 31 with a yield of 65%.1H NMR (400 MHz, DMSO) δ 11.37 (s, 1H), 7.76 (s, 1H), 6.90 (s, 1H), 6.21 (d, J = 1.1 Hz, 1H), 2.38 (d, J = 1.1 Hz, 3H). 13C NMR (101 MHz, DMSO) δ 160.30, 156.71, 153.46, 153.37, 126.59, 117.30, 113.30, 111.78, 103.77, 18.56.
6-ethyl-7-hydroxy-4-methyl-2H-chromen-2-one (32).
To a mixture of 4-ethylbenzene-1,3-diol (5.0 mmol) and ethyl acetoacetate (7.5 mmol) was added drop-wise perchloric acid (6.0 ml) at room temperature and stirred for 6 h. After completion of the reaction as indicated by TLC, the reaction mixture was poured slowly into a mixture of ice-water (50 ml) with stirring. The resultant suspension was filtered and the collected solid was washed with water and dried, then the crude compound was recrystallized from methanol to produce compound 32 with a yield of 85%. 1H NMR (400 MHz, DMSO) δ 10.51 (s, 1H), 7.45 (s, 1H), 6.73 (s, 1H), 6.11 (d, J = 1.1 Hz, 1H), 2.60 (q, J = 7.5 Hz, 2H), 2.38 (d, J = 1.1 Hz, 3H), 1.16 (t, J = 7.5 Hz, 3H). 13C NMR (101 MHz, DMSO) δ 160.90, 159.39, 154.11, 153.41, 128.18, 125.60, 112.17, 110.57, 102.03, 22.98, 18.63, 14.60.
4.3. Fluorescence polarization-based binding (FP) assay
For the competitive binding assay for Mcl-1 protein, FAM-Bid peptide (10 nM) and Mcl-1 protein (50 nM) were preincubated in the assay buffer (25 mM Tris, pH 8.0; 150 mM NaCl). Each inhibitor was dissolved in DMSO to obtain a stock solution (10 mM). The stock solution was then diluted successively to get different concentrations (1000 μM, 400 μM, 100 μM, 100 μM, 40 μM, 10 μM, 4 μM, 1 μM, 0.4 μM, 0.1 μM 0.04 μM and 0.01 μM). The serial dilutions of each compound were added to the incubation mixture. After 30 min incubation, the polarization values were measured using a black 96-well plate in Spectra Max M5 Detection System. A Kd value of 13 nM for FAM-Bid binding to Mcl-1 protein was determined by saturation experiments. The Ki value for each inhibitor was calculated using the equation developed for FP-based assays by Kuntz et al. using a computer program for Ki value calculation,43 which is available at: http://sw16.im.med.umich.edu/software/calc_ki/.
4.4. Cell culture
Human lung adenocarcinoma cells (A549 cell line) was cultured in RPMI1640 medium supplemented with 10% fetal bovine serum (FBS), 60 mg/mL streptomycin and 100 mg/mL penicillin in a humidified atmosphere of a 5% CO2 at 37°C.
4.5. Cytotoxicity assay by SRB method
A549 cells were seeded in 96-well plates (6.5 × 103 cells/well) and incubated for 24 hours at 37°C in a humid atmosphere with 5% CO2 for adhesion. After this period, the medium was withdrawn and the wells were treated with different concentrations (0.0001, 0.001, 0.01, 0.1, 1, 10 and 100 μM) of Cpd 4 and taxol, either alone or in combination in 10% FBS. After 48 h, 50 μl of 50% pre-cold trichloroacetic acid was added to the wells gently. After incubation at 4°C for one h, the plates were flicked and washed five times with cold water and were then air dried. The air-dried plates were stained with 100 μl SRB and kept for 30 min at 37°C. The unbound dye was removed by four times wash with 1% acetic acid. After being air-dried, 100 μl of 10 mM Tris base (pH 10.5) was then added to the wells to solubilize the dye and the plates were shaken vigorously for 5 min. The absorbance reading was performed in a microplate reader (Powerwave HT; Biotek) at 570 nm. The percentage growth inhibition was calculated using following formula:
All experiments were performed in triplicate. The results were expressed as the means of the IC50 (drug concentration that reduced cell viability to 50% of the control).
4.6. Molecular docking
The crystal structure of Mcl-1 was obtained from the Protein Data Bank (PDB ID: 4WMR).44 The molecular docking process was performed using Discovery Studio (BIOVIA Discovery Studio 2016, Dassault Systèmes, San Diego, USA). The CHARMM 40.1 force field was used to represent the protein and ligand structures. Docking simulations were performed by a standard LibDock protocol, where protein features are referred to as hotspots. After a final energy-minimization step (allowing the ligand poses to be flexible), the top scoring ligand poses are saved. The rigid poses were placed into the active site of Mcl-1, and the hotspots are matched as triplets. The protein-ligand complexes with the highest LibDock score were taken from the docking results and depicted in full text.
4.7. 3D-QSAR Model Building
Three 3D-QSAR models were built using the corresponding package of Drug Discovery Studio. Primarily, compounds were aligned by consensus on both steric and electrostatic fields, with relative weight of 50–50%. The aligned molecules were placed in a 3D grid space, with grid spacing of 1.5 Å. The extent of the grid was set to the bounding box of all the ligands plus 6.0 Å of extension. The CHARMm force field was used. The electrostatic potential and the van derWaals potential were treated as separate terms. A +1e point charge was used as the electrostatic potential probe. Distance-dependent dielectric constant was used to mimic the solvation effect. For the van der Waals potential, a carbon atom with a 1.73 Å radius was used as a probe. The energy grid potentials were filtered to remove highly correlated descriptors (maximum descriptor correlation was set as 0.9). The energy grid potentials were filtered to remove highly correlated descriptors. Partial least squares (PLS) models were then built using these remaining descriptors, and Log (Ki for Mcl-1) was used as activity properties.
Supplementary Material
Highlights.
The inhibitory effects of a series of structurally diverse coumarins on Mcl-1 were assayed.
The structure-activity relationship of couamrins as Mcl-1 inhibitors were summarized.
Docking simulations were conducted to explore the interactions between coumarins and Mcl-1.
3D-QSAR analysis provided new insights into the relationships between the Mcl-1 inhibitory effects and the steric-electrostatic properties of coumarins.
Acknowledgement
This work was supported by National Key Research and Development Program of China (2017YFC1700200, 2017YFC1702000), National Natural Science Foundation of China (81603187), Natural Science Foundation of Shanghai (18ZR1436500), Fundamental Research Funds for the Central Universities (DUT18LK39), and National Cancer Institute Intramural Research Program (ZIABC005708).
Footnotes
Declaration of interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
The authors report no conflicts of interests.
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Wei G, Margolin AA, Haery L, Brown E, Cucolo L, Julian B, Shehata S, Kung AL, Beroukhim R, Golub TR, Cancer Cell 21 (2012) 547–562. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Song L, Coppola D, Livingston S, Cress D, Haura EB, Mcl-1 regulates survival and sensitivity to diverse apoptotic stimuli in human non-small cell lung cancer cells, Cancer Biol. Ther 4 (2005) 267–276. [DOI] [PubMed] [Google Scholar]
- 3.Ding Q, He X, Xia W, Hsu JM, Chen CT, Li LY, Lee DF, Yang JY, Xie X, Liu JC, Hung MC, Myeloid cell leukemia-1 inversely correlates with glycogen synthase kinase-3beta activity and associates with poor prognosis in human breast cancer, Cancer Res. 67 (2007) 4564–4571. [DOI] [PubMed] [Google Scholar]
- 4.Derenne S, Monia B, Dean NM, Taylor JK, Rapp MJ, Harousseau JL, Bataille R, Amiot M, Antisense strategy shows that Mcl-1 rather than Bcl-2 or Bcl-x(L) is an essential survival protein of human myeloma cells, Blood 100 (2002) 194–199. [DOI] [PubMed] [Google Scholar]
- 5.Andersen MH, Becker JC, Thor Straten P, The antiapoptotic member of theBcl-2 family Mcl-1 is a CTL target in cancer patients, Leukemia 19 (2005) 484–485. [DOI] [PubMed] [Google Scholar]
- 6.Kang MH, Wan Z, Kang YH, Sposto R, Reynolds CP, Mechanism of synergy of N-(4-hydroxyphenyl)retinamide and ABT737 in acute lymphoblastic leukemia cell lines: Mcl-1 inactivation, J. Natl. Cancer Inst (2008) 580–595. [DOI] [PubMed] [Google Scholar]
- 7.Wertz IE, Kusam S, Lam C, Okamoto T, Sandoval W, Anderson DJ, Helgason E, Ernst JA, Eby M, Liu J, Belmont LD, Kaminker JS, O’Rourke KM, Pujara K, Kohli PB, Johnson AR, Chiu ML, Lill JR, Jackson PK, Fairbrother WJ, Seshagiri S, Ludlam MJC, Leong KG, Dueber EC, Maecker H, Huang DCS, Dixit VM, Sensitivity to antitubulin chemotherapeutics is regulated by MCL1 and FBW7, Nature 471 (2011) 110–114. [DOI] [PubMed] [Google Scholar]
- 8.Wei SH, Dong K, Lin F, Wang X, Li B, Shen JJ, Zhang Q, Wang R, Zhang HZ, Inducing apoptosis and enhancing chemosensitivity to gemcitabine via RNA interference targeting Mcl-1 gene in pancreatic carcinoma cell, Cancer Chemother. Pharmacol 62 (2008) 1055–1064. [DOI] [PubMed] [Google Scholar]
- 9.van Delft MF, Wei AH, Mason KD, Vandenberg CJ, Chen L, Czabotar PE, Willis SN, Scott CL, Day CL, Cory S, Adams JM, Roberts AW, Huang DCS, The BH3 mimetic ABT737 targets selective Bcl-2 proteins and efficiently induces apoptosis via Bak/Bax if Mcl-1 is neutralized, Cancer Cell 10 (2006) 389–399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Varin E, Denoyelle C, Brotin E, Meryet-Figuiere M, Giffard F, Abeilard E, Goux D, Gauduchon P, Icard P, Poulain L, Downregulation of Bcl-xL and Mcl-1 is sufficient to induce cell death in mesothelioma cells highly refractory to conventional chemotherapy, Carcinogenesis 31 (2010) 984–993. [DOI] [PubMed] [Google Scholar]
- 11.Olberding KE, Wang X, Zhu Y, Pan J, Rai SN, Li C, Actinomycin D synergistically enhances the efficacy of the BH3 mimetic ABT-737 by downregulating Mcl-1 expression, Cancer Biol. Ther 10 (2010) 918–929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Zhang H, Guttikonda S, Roberts L, Uziel T, Semizarov D, Elmore SW, Leverson JD, Lam LT, Mcl-1 is critical for survival in a subgroup of non-small-cell lung cancer cell lines, Oncogene 30 (2011) 1963–1968. [DOI] [PubMed] [Google Scholar]
- 13.Rega MF, Wu B, Wei J, Zhang Z, Cellitti JF, Pellecchia M, SAR by interligand nuclear Overhauser effects (ILOEs) based discovery of acylsulfonamide compounds active against Bcl-x(L) and Mcl-1, J. Med. Chem 54 (2011) 6000–6013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Kim YB, Balasis ME, Doi K, Berndt N, Duboulay C, Hu CCA, Guida W, Wang HG, Sebti SM, Del Valle JR, Synthesis and evaluation of substituted hexahydronaphthalenes as novel inhibitors of the Mcl-1/BimBH3 interaction, Bioorg. Med. Chem. Lett 22 (2012) 5961–5965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Cohen NA, Stewart ML, Gavathiotis E, Tepper JL, Bruekner SR, Koss B, Opferman JT, Walensky LD, A competitive stapled peptide screen identifies a selective small molecule that overcomes MCL-1-dependent leukemia cell survival, Chem. Biol 19 (2012) 1175–1186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Stewart ML, Fire E, Keating AE, Walensky LD, The MCL-1 BH3 helix is an exclusive MCL-1 inhibitor and apoptosis sensitizer, Nat. Chem. Biol 6 (2010) 595–601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Muppidi A, Doi K, Edwardraja S, Drake EJ, Gulick AM, Wang HG, Lin Q, Rational design of proteolytically stable, cellpermeable peptide-based selective Mcl-1 inhibitors, J. Am. Chem. Soc 134 (2012) 14734–14737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kostova I, Bhatia S, Grigorov P, Balkansky S, Parmar VS, Prasad AK, Saso L, Coumarins as antioxidants, Curr. Med. Chem 18 (2011) 3929–3951; [DOI] [PubMed] [Google Scholar]
- 19.Atmaca M, Bilgin HM, Obay BD, Diken H, Kelle M, Kale E, The hepatoprotective effect of coumarin and coumarin derivates on carbon tetrachloride-induced hepatic injury by antioxidative activities in rats, J. Physiol. Biochem 67 (2011) 569–576. [DOI] [PubMed] [Google Scholar]
- 20.Emami S, Foroumadi A, Faramarzi MA, Samadi N, Synthesis and antibacterial activity of quinolone-based compounds containing a coumarin moiety, Arch. Pharm. Chem. Life Sci 341 (2008) 42–48. [DOI] [PubMed] [Google Scholar]
- 21.Manvar A, Bavishi A, Radadiya A, Patel J, Vora V, Dodia N, Rawal K, Shah A, Diversity oriented design of various hydrazides and their in vitro evaluation against Mycobacterium tuberculosis H37Rv strains, Bioorg. Med. Chem. Lett 21 (2011) 4728–4731. [DOI] [PubMed] [Google Scholar]
- 22.Sashidhara KV, Kumar A, Chatterjee M, Rao KB, Singh S, Verma AK, Palit G, Discovery and synthesis of novel 3-phenylcoumarin derivatives as antidepressant agents, Bioorg. Med. Chem. Lett 21 (2011) 1937–1941. [DOI] [PubMed] [Google Scholar]
- 23.Yuce B, Danis O, Ogan A, Sener G, Bulut M, Yarat A, Antioxidative and lipid lowering effects of 7, 8-dihydroxy-3-(4-methylphenyl) coumarin in hyperlipidemic rats, Arzneim. Forsch 59 (2009) 129–134. [DOI] [PubMed] [Google Scholar]
- 24.Wang CJ, Hsieh YJ, Chu CY, Lin YL, Tseng TH, Inhibition of cell cycle progression in human leukemia HL-60 cells by esculetin, Cancer Lett. 183 (2002) 163–168. [DOI] [PubMed] [Google Scholar]
- 25.Nasr T, Bondock S, Youns M, Anticancer activity of new coumarin substituted hydrazide-hydrazone derivatives, Eur. J. Med. Chem 76 (2014) 539–548. [DOI] [PubMed] [Google Scholar]
- 26.Vijay Avin BR, Thirusangu P, Lakshmi Ranganatha V, Firdouse A, Prabhakar BT, Khanum SA, Synthesis and tumor inhibitory activity of novel coumarin analogs targeting angiogenesis and apoptosis, Eur. J. Med. Chem 75 (2014) 211–221. [DOI] [PubMed] [Google Scholar]
- 27.Ruiz-Marcial C, Reyes-Chilpa Chilpa P, Estrada E, Reyes Esparza J, Farina GG, Rodríguez-Fragoso L, Antiproliferative, cytotoxic and antitumour activity of coumarin isolated from Calophyllum brasiliense, J. Pharm. Pharmacol 59 (2007) 719–725. [DOI] [PubMed] [Google Scholar]
- 28.Facompre M, Tardy C, Bal-Mayeu C, Colson P, Perez C, Manzanares I, Cuevas C, Bailly C, Lamellarin D: A Novel potent inhibitor of topoisomerase I, Cancer Res. 63 (2003) 7392–7399. [PubMed] [Google Scholar]
- 29.Nagarajan M, Morrell A, Fort BC, Meckley MR, Antony S, Kohlhagen G, Pommier Y, Cushman M, Synthesis and anticancer activity of simplified indenoisoquinoline topoisomerase I inhibitors lacking substituents on the aromatic rings, J. Med. Chem 47 (2004) 5651–5661. [DOI] [PubMed] [Google Scholar]
- 30.Oh BS, Shin EA, Jung JH, Jung DB, Kim B, Shim BS, Yazdi MC, Iranshahi M, Kim SH, Apoptotic effect of galbanic acid via activation of caspases and inhibition of Mcl-1 in H460 non-small lung carcinoma cells, Phytother Res. 29 (2015) 844–849. [DOI] [PubMed] [Google Scholar]
- 31.Wang P, Xia YL, Yu Y, Lu JX, Zou LW, Feng L, Ge GB, Yang L, Design, synthesis and biological evaluation of esculetin derivatives as anti-tumour agents, RSC Adv. 5 (2015) 53477–53483. [Google Scholar]
- 32.Van der Waterbeemd H, Quantitative Approaches to Structure Activity Relationships, in: Wermuth CG (Eds.), In the Practice of Medicinal Chemistry, Academic Press, London, 1996, pp. 367. [Google Scholar]
- 33.Chen C, Wang P, Zou L, Yang L, Fan Y, Hu W, Ge G, Synthesis and biological evaluation of hydroxylcoumarin derivatives as antioxidant agents, Chem. Res. Chinese U 33 (2017) 194–199. [Google Scholar]
- 34.Xia Y, Chen C, Liu Y, Ge G, Dou T, Wang P, Synthesis and structure-activity relationship of daphnetin derivatives as potent antioxidant agents, Molecules, 23 (2018) 2476–2488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Barot KP, Jain SV, Kremer L, Singh S, Ghate MD, Recent advances and therapeutic journey of coumarins: Current status and perspectives, Med. Chem. Res 24 (2015) 2771–2798. [Google Scholar]
- 36.Zhang ZC, Liu CW, Li XQ, Song T, Wu ZY, Liang XM, Zhao Y, Shen XY, Chen HB, Fragment-based design, synthesis, and biological evaluation of N-substituted-5-(4-isopropylthiophenol)-2-hydroxynicotinamide derivatives as novel Mcl-1 inhibitors, Eur. J. Med. Chem 60 (2013) 410–20. [DOI] [PubMed] [Google Scholar]
- 37.Shen Q, Li J, Mai JH, Zhang Z, Fisher A, Wu XY, Li ZQ, Ramirez MR, Chen SQ, Shen HF, Sensitizing non-small cell lung cancer to BCL-xL-targeted apoptosis, Cell Death Dis. 9 (2018) 986–998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Lanning ME, Yu WB, L Yap J, Chauhan J, Chen LJ, Whiting E, S Pidugu L, Atkinson T, Bailey H, Li W, M Roth B, Hynicka L, Chesko K, A Toth E, Shapiro P, D MacKerell A Jr, T Wilder P, Fletcher S, Structure-based design of N-substituted 1-hydroxy-4-sulfamoyl-2-naphthoates as selective inhibitors of the Mcl-1 oncoprotein, Eur. J. Med. Chem 113 (2016) 273–92. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Soderholm AA, Lehtovuori PT, Nyronen TH, Docking and three-dimensional quantitative structure-activity relationship (3D QSAR) analyses of nonsteroidal progesterone receptor ligands, J. Med. Chem 49 (2006) 4261–4268. [DOI] [PubMed] [Google Scholar]
- 40.Vujasinovic I, Paravic-Radicevic A, Mlinaric-Majerski K, Brajsa K, Bertosa B, Synthesis and biological validation of novel pyrazole derivatives with anticancer activity guided by 3D-QSAR analysis, Bioorg. Med. Chem 20 (2012) 2101–2110. [DOI] [PubMed] [Google Scholar]
- 41.Zhang Z, Song T, Zhang T, Wu G, Gao J, An L, Du G, A novel BH3 mimetic S1 potently induces Bax/Bak-dependent apoptosis by targeting both Bcl-2 and Mcl-1, Int. J. Cancer 128 (2011) 1724–1735. [DOI] [PubMed] [Google Scholar]
- 42.Wang G, Nikolovska-Coleska Z, Yang CY, Wang R, Tang G, Guo J, Shangary S, Qiu S, Gao W, Yang D, Meagher J, Stuckey J, Krajewski K, Jiang S, Roller PP, Abaan HO, Tomita Y, Wang S, Structure-based design of potent small-molecule inhibitors of anti-apoptotic Bcl-2 proteins, J. Med. Chem 49 (2006) 6139–6142. [DOI] [PubMed] [Google Scholar]
- 43.Kuntz ID, Chen K, Sharp KA, Kollman PA, The maximal affinity of ligands, Proc. Natl. Acad. Sci. U.S.A 96 (1999) 9997–10002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Clifton MC, Dranow DM, Leed A, Fulroth B, Fairman JW, Abendroth J, Atkins KA, Wallace E, Fan D, Xu G, Ni ZJ, Daniels D, Van Drie J, Wei G, Burgin AB, Golub TR, Hubbard BK, Serrano-Wu MH, A maltose-binding protein fusion construct yields a robust crystallography platform for MCL1, PLoS One 10 (2015) e0125010–e0125010. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.