Abstract
Thymallus arcticus grubei is a specific grayling species that naturally occurs in the Xinjiang Irtysh River basin, China. The complete mitochondrial genome of The Xinjiang arctic grayling T. arcticus grubei was first determined (Accession number LC168675), with a total length of 16,658 nucleotides, of which 15,587 nucleotides are coding DNA, and 1071 nucleotides are non-coding DNA. It has the common feature with those of other grayling with respect to genome structure and gene arrangement. It contained 13 protein-coding genes, 2 ribosomal RNA genes, 22 transfer tRNA genes, and 1 control region (D-loop region). The complete mitochondrial genome of Xinjiang arctic grayling T. arcticus grubei provides an important data set for further study on its classification.
Keywords: Xinjiang arctic grayling, Thymallus arcticus grubei, mitochondrial genome
Thymallus arcticus grubei, which is classified as Thymallus, Salmonidae, Salmoniformes, Protacanthopterygii, Euteleostei is a specific grayling species that naturally occurs in Xinjiang Irtysh River basin, China (Liu et al. 2016). Commonly, T. arcticus grubei is classified as one of the subspecies of arctic grayling. However, there is still a lack of molecular data of mitochondrial DNA sequence to prove this opinion. So, it is important to obtain the whole mitogenome of T. arcticus grubei for the further study. The specimen of T. arcticus grubei, named as Thyag-01 based on the phylogenetic analysis, was collected from Irtysh River, Xinjiang, China (48°12′ N, 86°40′ E) in 2014 and stored in Xinjiang Fisheries Research Institute, Urumqi, China. Genomic DNA was extracted from Xinjiang arctic grayling muscle according to Liu et al. (2012, 2014). The mitochondrial genome was amplified using 25 pars of primers, designed based on mitogenome sequence of the arctic grayling, T. arcticus arcticus (Accession number KJ866481) and the Mongolian grayling, T. brevirostris (Accession number NC_027412). Protein-coding genes were analyzed by ORF Finder (http://www.ncbi.nlm.nih.gov/gorf/gorf.html) using the invertebrate mitochondrial code. The tRNA genes were identified by ARWEN (Laslett & Canback 2008) and tRNA-scan SE (Lowe & Eddy 1997).
The complete T. arcticus grubei mitochondrial genome (Accession number LC168675) was 16,658 nucleotides long, of which 15,587 nucleotides are coding DNA, and 1071 nucleotides are non-coding DNA. The mitochondrial genome of T. arcticus grubei was found to contain 13 protein-coding genes, 2 ribosomal RNA genes, 22 transfer RNA genes, and 1 control region. Its structure and composition is also comparable to other vertebrates (Yasuike et al. 2010; Liang et al. 2012; Liu et al. 2013; Ma et al. 2015; Xu et al. 2015; Shan & Liu 2015). A phylogenetic tree was constructed based on the comparison of the complete mitochondrial genome sequences with other Thymallus species using Neighbour-Joining method (Figure 1). On average, the AT content of the T. arcticus grubei mtDNA was 55.18%. To investigate the nucleotide bias, skew for a given strand was calculated as (A − T)/(A + T) or (G − C)/(G + C) (Perna & Kocher 1995). The AT and GC skews for the T. arcticus grubei mitochondrial genome were 0.015 and −0.225, respectively; this finding indicated that the strand that encoded genes contained more A and C than T and G, and this skew was evidence of codon usage bias. Of the 13 protein-coding genes identified in the T. arcticus grubei mitochondrial genome, all but cox1 used conventional start codons, ATG. The ORFs of six protein-coding genes, ND1, cox1, Atp8, Atp6, nad4L, and nad5, ended with a TAA codon, and only one gene, nad6, ended with TAG, respectively. In addition, six genes, nad2, cox2, cox3, nad3, nad4, and cytb, ended with incomplete stop codon T or AT. The 12S rRNA gene was 946-bp long and was flanked by the tRNA-phe and tRNA-val, while the 16S rRNA was 1679 bp long and was flanked by tRNA-val and tRNA-leu. The intergenic region between 12S rRNA and 16S rRNA was found to be 72 bp and to contain only one tRNA (tRNA-val). The lengths of 22 tRNA coding genes range in size from 67 to 75 nucleotides. The control region was located between tRNA-pro and tRNA-phe with a length of 1003 bp.
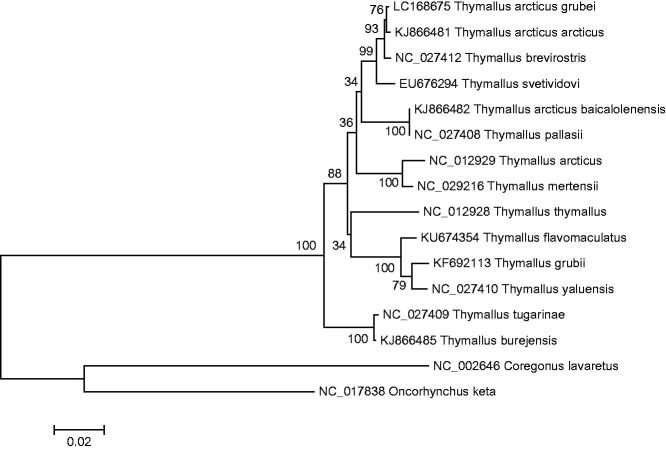
Figure 1.
A phylogenetic tree constructed based on the comparison of complete mitochondrial genome sequences of the grayling, T. arcticus grubei and other thirteen species of Thymallus genus. They are T. thymallus (European grayling), T. pallasii (East Siberian grayling), T. arcticus (Arctic grayling), T. tugarinae (Lower Amur grayling), T. burejensis (Bureya grayling), T. grubii (Amur grayling), T. arcticus arcticus (Arctic grayling), T. brevirostris (Mongolian grayling), T. yaluensis (Yalu grayling), T. mertensii (Kamchatka grayling), T. flavomaculatus (Yellow-spotted grayling), T. baicalolenensis (Baikal black grayling), and T. svetividovi (Upper Yenisei grayling). Oncorhynchus keta and Coregonus lavaretus are used as outgroup. Genbank accession numbers for all sequences are listed in the figure. The numbers at the nodes are bootstrap percent probability values based on 1000 replications.
Acknowledgments
Disclosure statement
The authors report no conflicts of interest. The authors alone are responsible for the content and writing of the article.
Funding
This work was supported by the National Natural Science Foundation of China [31560719], the Natural Science Foundation of Xinjiang Uygur Autonomous Region [2015211C261], and the High-level talent introduction project of Xinjiang Uygur Autonomous Region [201412].
References
- Laslett D, Canback B. 2008. ARWEN: a program to detect tRNA genes in metazoan mitochondrial nucleotide sequences. Bioinformatics. 24:172–175. [DOI] [PubMed] [Google Scholar]
- Liu YG, Kurokawa T, Sekino M, Tanabe T, Watanabe K. 2013. Complete mitochondrial DNA sequence of the ark shell Scapharca broughtonii: an ultra-large metazoan mitochondrial genome. Compar Biochem Physiol Part D. 8:72–81. [DOI] [PubMed] [Google Scholar]
- Lowe TM, Eddy SR. 1997. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 25:955–964. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perna NT, Kocher TD. 1995. Patterns of nucleotide composition at fourfold degenerate sites of animal mitochondrial genomes. J Mol Evol. 41:353–358. [DOI] [PubMed] [Google Scholar]
- Xu D, He CQ, Li QH, He J, Ma HM. 2015. The complete mitochondrial genome of the Daweizi pig. Mitochondrial DNA. 26:640–641. [DOI] [PubMed] [Google Scholar]
- Liu YG, Liu LX, Wang YX. 2016. A review: research progress of development of microsatellite markers and conservation genetics in Graying. Fish Sci. 35:87–92. [Google Scholar]
- Liu YG, Guo YG, Hao J, Liu LX. 2012. Genetic diversity of swimming crab (Portunus trituberculatus) populations from Shandong peninsula as assessed by microsatellite markers. Biochem Syst Ecol. 41:91–97. [Google Scholar]
- Liu YG, Liu LX, Xing SC. 2014. Development and characterization of thirteen polymorphic microsatellite loci in the Cynoglossus semilaevis. Conserv Genet Res. 6:683–684. [Google Scholar]
- Ma B, Jiang H, Sun P, Chen J. 2015. Complete mitochondrial genome of Thymallus grubii (Salmonidae: Thymallinae). Mitochondrial DNA. 26:815–816. [DOI] [PubMed] [Google Scholar]
- Yasuike M, Jantzen S, Cooper G, Leder E, Davidson W, Koop B. 2010. Grayling (Thymallinae) phylogeny within salmonids: complete mitochondrial DNA sequences of Thymallus arcticus and Thymallus thymallus. J Fish Biol. 76:395–400. [DOI] [PubMed] [Google Scholar]
- Liang HW, Hu GF, Li Z, Zou GW, Liu XL. 2012. Mitochondrial DNA sequence of yellow catfish (Pelteobagrus fulvidraco). Mitochondrial DNA. 23:170–172. [DOI] [PubMed] [Google Scholar]
- Shan WJ, Liu YG. 2015. The complete mitochondrial DNA sequence of the cape hare Lepus capensis pamirensis. Mitochondrial DNA. doi: 10.3109/19401736.2015.1101569. [DOI] [PubMed] [Google Scholar]