Abstract
The COVID-19 outbreak circulating the world is far from being controlled, and possible contamination routes are still being studied. There are no confirmed cases yet, but little is known about the infection possibility via contact with sewage or contaminated water as well as with aerosols generated during the pumping and treatment of these aqueous matrices. Therefore, this article presents a literature review on the detection of SARS-CoV-2 in human excreta and its pathways through the sewer system and wastewater treatment plants until it reaches the water bodies, highlighting their occurrence and infectivity in sewage and natural water. Research lines are still indicated, which we believe are important for improving the detection, quantification, and mainly the infectivity analyzes of SARS-CoV-2 and other enveloped viruses in sewage and natural water. In fact, up till now, no case of transmission via contact with sewage or contaminated water has been reported and the few studies conducted with these aqueous matrices have not detected infectious viruses. On the other hand, studies are showing that SARS-CoV-2 can remain viable, i.e., infectious, for up to 4.3 and 6 days in sewage and water, respectively, and that other species of coronavirus may remain viable in these aqueous matrices for more than one year, depending on the sample conditions. These are strong pieces of evidence that the contamination mediated by contact with sewage or contaminated water cannot be ruled out, even because other more resistant and infectious mutations of SARS-CoV-2 may appear.
Keywords: COVID-19, SARS-CoV-2, Coronavirus, Infectivity, Sewage, Fecal-oral contamination
Graphical abstract

1. Introduction
At the time this paper was finished, on 1 February 2021, coronavirus disease 2019 (COVID-19) is already considered a pandemic, being present in 215 countries and territories across the globe, resulting in astronomical levels of infected (102 million) and dead (2.2 million) people (WHO, 2021), with the numbers continuing to rise. Nevertheless, given the large number of infected people who do not have the symptoms of the disease, these numbers are underestimated. Thus, based on hospitalization data and ribonucleic acid (RNA) from Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) detected in municipal sewage sludge, Kaplan et al. (2020) developed a mathematical model predicting that 9.3% of the population in the New Haven metropolitan area, Connecticut, USA, had already been infected on May 2020. Were this estimate to roughly correspond to the global average level of infection, more than 700 million people would already be infected around the world.
COVID-19 is characterized as a severe respiratory syndrome occurring in humans and is caused by the SARS-CoV-2 (Bojkova et al., 2020; Chen et al., 2020b; Yan et al., 2020), in which respiratory transmission is the primary route for infection (Wu et al., 2020b), being spread by person-to-person direct or indirect contact (Heymann and Shindo, 2020; Kitajima et al., 2020; Mohseni et al., 2020). However, other contamination routes are still under discussion. At this time, a possible route of infection under avid discussion is the one mediated by sewage and natural water contaminated with feces and other excreta from people infected with SARS-CoV-2 (Amirian, 2020; Daraei et al., 2020; O'Reilly et al., 2020). In fact, although the virus has already been found in sewage as well as in rivers around the world (Ahmed et al., 2020a; Haramoto et al., 2020; La Rosa et al., 2020b; Randazzo et al., 2020b; Rimoldi et al., 2020), there is a remaining question: could people, coming into contact with these contaminated aqueous matrices, be infected by SARS-CoV-2?
The literature reported the presence of coronaviruses in feces and their ability to infect human cells (Xiao et al., 2020; Zhang et al., 2020c). Therefore, there is a growing concern about a possible fecal-oral or fecal-nasal contamination (Bivins et al., 2020; Donà et al., 2020). Although this pandemic is highly impactful, there is still little knowledge about COVID-19 in some aspects, so that, to our knowledge, up to now, there is only two reports published on the infectivity of this virus present in sewage and (or) natural water (Rimoldi et al., 2020; Westhaus et al., 2021). Despite little concrete information on this issue, many researchers warn about the possibility of infection through contact with sewage or contaminated water (Adelodun et al., 2020; Amoah et al., 2020; Arslan et al., 2020; Carraturo et al., 2020; da Silva et al., 2020; Nghiem et al., 2020; Olusola-Makinde and Reuben, 2020; Thompson et al., 2020) or with the aerosols generated in the pumping and treatment systems (Gormley et al., 2020; Gude and Muire, 2021; Harries et al., 2020; Naddeo and Liu, 2020), in toilets flushing (Ding et al., 2021; Wathore et al., 2020), and also in faulty connections of floor drains with the building's main sewer pipe (Shi et al., 2020). Moreover, grounded on evidence from China, Liu (2020) conducted a study on the transmission of the COVID-19 from the urban perspective, and based on statistical findings, pointed out that sewage contributes to the spreading of the disease. Later, researchers reinforced the potential for secondary transmission via sewage, especially in low-incoming countries that have unsatisfactory heath and sanitation infrastructure, monitoring and policies (Liu et al., 2020a; Yuan et al., 2020).
Considering the work of Rimoldi et al. (2020), who investigated the presence and infectivity of SARS-CoV-2 in samples of raw and treated wastewater from three wastewater treatment plants (WWTPs) within the Milan Metropolitan Area, Italy, as well as in their receptor water bodies, SARS-CoV-2 RNA was detected in raw, but not in treated wastewater, while samples from receiving rivers showed, in some cases, SARS-CoV-2 RNA positivity. On the other hand, virus infectivity was always null for all samples, indicating limited risks for public health. Considering the preliminary nature of the presented results, the authors also advocated a precautionary approach to risk assessment related to infectivity.
In fact, we agree with the warnings and the precautionary point of view adopted by the authors. Therefore, in the present paper, we will highlight some issues we think are key points when the infectivity of SARS-CoV-2 is evaluated in complexes matrices such as sewage. It is important to highlight that, to date, these matters have not yet been clarified in other studies. Thus, considering the problems associated with the spread and contamination with SARS-CoV-2, a deeper discussion on the infectivity of coronavirus from sewage is necessary and of paramount importance; mainly because a study (Chen et al., 2020c) indicated that the SARS-CoV-2 has become more infectious and that even more infectious strains may appear.
2. Methodology
The review was prepared grounded on a literature search for documents published, until 10 November 2020, in the following databases: Pub Med, Scopus, and ScienceDirect. The keywords employed for the search were COVID-19 or COVID or SARS-CoV-2 or Coronavirus or Severe Acute Respiratory Syndrome and Water or Wastewater or Sewage or Urine or Feces or Virus Survival, being the results presented in Table S1. In this first stage of the research, the title of the articles was evaluated, and those that could contain information about the detection and quantification of SARS-CoV-2 in water, wastewater, sewage, feces, or urine samples were selected, as well as those containing information related to the survival and infectivity of SARS-CoV-2 from these matrices. 449 articles were selected, in which the abstract was screened to verify whether they really contained the information described above, which were considered relevant for the project. At this stage, 253 articles were excluded because they did not contain the required information or were review articles, resulting in 196 eligible articles that were full text screened and used as references for this review. Another 17 articles were added due to a screening in the references of the selected articles or at the reviewers' suggestion.
3. SARS-CoV-2 characterization: morphology, structure and infection
Coronaviruses are commonly found in nature and may infect a broad range of animals and humans, causing illnesses in the respiratory, gastrointestinal, and neurological systems (Ding and Liang, 2020). They belong to the Coronaviridae family, are non-segmented and positive-sense single-stranded RNA viruses with a large genome of approximately 30 thousand nucleotides within a fragile lipid envelope (Krafcikova et al., 2020; Toyoshima et al., 2020), which, in turn, due to its delicate nature, become the virions non-viable once the envelope is damaged, although their genetic fragments can still be detected (Maal-Bared et al., 2020; Nghiem et al., 2020; Polo et al., 2020). Therefore, the ability of these enveloped viruses to infect is associated with the preservation of their capsid (envelope), that is, without it, they are non-infectious (Atkinson and Petersen, 2020). Nonetheless, reports have proposed that the SARS-CoV-2 RNA might be infectious and that the transmission of these RNAs could be a possible transmission pathway of the COVID-19 (Xu, 2020).
According to Lesimple et al. (2020), the SARS-CoV-2 virion is a small spherical particle with a diameter of 100 nm, but there are studies reporting diameters of 120 nm (La Rosa et al., 2020a), 140 nm (Foladori et al., 2020), and up to 200 nm (Chen et al., 2020d). SARS-CoV-2 has four structural proteins, namely spike surface (S) and membrane (M) glycoproteins, envelope (E), and nucleocapsid (N). The S glycoproteins, having about 9 to 12 nm long, on its surface, give the virus the appearance of a solar corona when observed by electron microscopy (Bahrami and Ferns, 2020; Zhu et al., 2020). The viral structure of a coronavirus particle is represented on Fig. 1 .
Fig. 1.
Viral structure of a coronavirus particle.
Created with BioRender.com.
The S glycoproteins are composed of two subunits, S1 and S2, which are responsible for binding to human host cells that contain angiotensin-converting enzyme 2 (ACE2) receptors, and subsequently assist in viral entry (Chen et al., 2020a; Du et al., 2009; Feng et al., 2020a; Xu et al., 2020a). The S1 subunit has a receptor-binding domain that recognizes and binds to the host receptor's ACE2, and subsequent conformational changes in S2 subunit facilitate fusion between the viral membrane and the one of the host cell, releasing thus the viral RNA into the host cell (Huang et al., 2020b). After that, polyproteins are translated from the viral RNA, and replication and transcription of the viral RNA occur via protein cleavage and assembly of the replicase–transcriptase complex. Thus, viral RNA is replicated, and structural proteins are synthesized, assembled, and bundled in the host cell depleting its resources; afterward, virions are released by exocytosis (Huang et al., 2020b; Ortiz-Prado et al., 2020), a process also known as viral shedding (Nghiem et al., 2020). ACE2s are membrane glycoproteins, highly expressed in organs like lungs, arteries, kidney, heart, but in the human body their highest expression is observed in intestinal enterocytes, being these organs the most affected by SARS-CoV-2 infections (Bhowmick et al., 2019; Kumar et al., 2020b; Premkumar et al., 2020). More details on SARS-CoV-2 infection, replication cycle and assembly of virions can be found at Gralinski and Menachery (2020), Huang et al. (2020b), Ortiz-Prado et al. (2020), and Sternberg and Naujokat (2020). The mechanism of infection of the SARS-CoV-2 into human cells is represented on Fig. 2 .
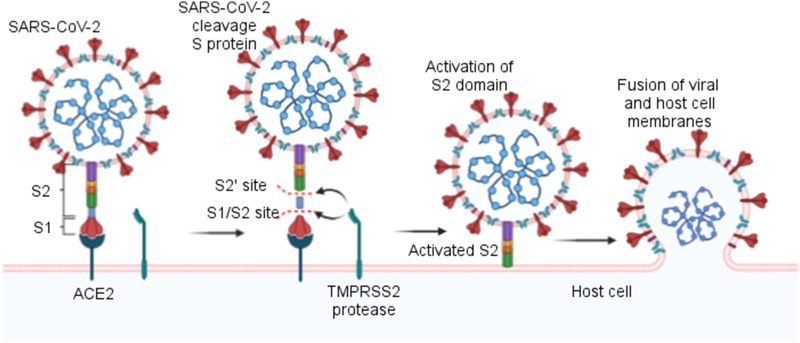
Fig. 2.
Mechanism of infection of the SARS-CoV-2 in human cells. The SARS-CoV-2 S protein binds with the ACE2 receptor on the host cell and is subsequently cleaved at the S1/S2 and S2′ sites by the TMPRSS2 protease. This activates the S2 subunit and leads to a fusion between the viral membrane and that of the host cell.
Adapted from Hartenian et al. (2020). Created with BioRender.com.
The SARS-CoV-2 has been detected in blood, respiratory secretions, sputum, tears, gastric juices, breast milk, semen, vaginal secretion, urine, and feces from symptomatic patients (Groß et al., 2020; Huang et al., 2020a; Li et al., 2020a; Lo et al., 2020; Peng et al., 2020; Wang et al., 2020b; Xia et al., 2020; Yu et al., 2020), and also in feces and other excreta from asymptomatic ones (Han et al., 2020b; Tang et al., 2020a). Furthermore, studies have reported prolonged viral shedding in feces, in which patients tested persistently positive on rectal swabs or feces, even after the nasopharyngeal test was negative (Du et al., 2020; Ling et al., 2020; Liu et al., 2020c; Xing et al., 2020; Xu et al., 2020b). Surprisingly, there are also studies reporting that the RNA test results of discharged patients returned positive during their follow-up visits (Li et al., 2020b; Tong et al., 2020). Therefore, COVID-19 is believed to be disseminated through direct contact, indirect contact via fomites (Malenovská, 2020; Pastorino et al., 2020), respiratory droplets (generated by sneezing, coughing, breathing, etc.), and possibly by aerosols (La Rosa et al., 2020a; Netz and Eaton, 2020; Smith et al., 2020; Tang et al., 2020b; Xiao et al., 2020). Recently the aerosol transmission route was confirmed by the National Health Commission of the People's Republic of China (Wang and Du, 2020), by the Center for Disease Control and Prevention (CDC) (Feng et al., 2020b), and by Lednicky et al. (2020). In addition, Fears et al. (2020) demonstrated that SARS-CoV-2 virions remained infectious for up to 16 h in respirable-sized aerosols, thus inferring that aerosols may be an important transmission route. Furthermore, it has also been reported that SARS-CoV-2 can remain infectious in the environment on a variety of surfaces for several hours or even days, such as 4 h on copper surfaces, 24 h on cardboard, two to three days on stainless steel, and three days on plastics (Nghiem et al., 2020). Yet, there are studies reporting that SARS-CoV-2 could possibly be transmitted via breastfeeding (Groß et al., 2020), vertical transmission during vaginal delivery (Carosso et al., 2020a, Carosso et al., 2020b), sexual transmission (Li et al., 2020a; Patrì et al., 2020), domestic cat transmission (Ali et al., 2020), and by contacting wastes generated from individuals affected by COVID-19 (Di Maria et al., 2020). Franklin and Bevins (2020) also discuss the possible spreading to new wild hosts such as bats, mustelids, and raccoons through the elimination of feces by infected humans and the introduction into the natural aquatic environment through the sewage treatment system.
Chen et al. (2020e) investigated SARS-CoV-2 shedding in the excreta of 42 COVID-19 patients, from which 18 (42.86%) patients remained positive for SARS-CoV-2 RNA in the feces after the pharyngeal swabs turned negative, being 6–10 days the duration of viral shedding from the feces after negative conversion in pharyngeal swabs, regardless of the disease severity. Later, other authors also reported similar fecal detection rates between patients with mild and severe COVID-19 disease (Zheng et al., 2020).
Wu et al. (2020b) reported that in 55% of the 74 infected patients studied, fecal samples remained positive for SARS-CoV-2 RNA for an average of 11.2 days after the respiratory tract samples became negative, indicating that the virus is replicating actively in the patients' gastrointestinal tract. The authors also highlight the possibility of prolonged duration of viral shedding in feces, for until five weeks after patients' respiratory samples have been negative for SARS-CoV-2 RNA, which suggests that after viral clearance in the respiratory system, fecal-oral transmission may possibly occur. Other researchers have also endorsed this approach (Hindson, 2020; Patel, 2020; Tang et al., 2020a; Xing et al., 2020; Xu et al., 2020b). In another study, Wang et al. (2020c) observed that the median duration of positive SARS-CoV-2 RNA in fecal samples was approximately nine days longer than in oropharyngeal swabs, and one patient persistently tested positive in fecal samples for 33 days after the oropharyngeal swabs tests became negative. Also, Mesoraca et al. (2020) observed patients having fecal samples positive test 25 days longer than their respiratory tract test. Recently, in a study on clinical samples from 20 patients infected with SARS-CoV-2, He et al. (2020) reported that one patient remained positive for viral RNA in the sputum and feces for about 75 and 80 days, respectively, after testing negative on nasal swab. More importantly, Lai et al. (2005) demonstrated that Severe Acute Respiratory Syndrome Coronavirus (SARS-CoV) can survive four days in diarrheal stool samples with alkaline pH, and more than seven days in respiratory samples at room temperature. Therefore, deeper studies are necessary to confirm or eliminate the hypothesis of fecal-oral transmission of SARS-CoV-2, especially for low-income countries that have low levels of sanitation, which makes sewage and surface water a possible source of spreading COVID-19 (Yeo et al., 2020).
4. SARS-CoV-2 in sewage and natural water
Coronaviruses can be introduced into sewage by various sources, such as hand washing, sputum, vomiting (Adelodun et al., 2020; Bhowmick et al., 2019; Han et al., 2020), and mainly via viral shedding in feces of infected individuals (Wang et al., 2020b; Wei et al., 2020). In addition, urine samples from infected people were reported to contain SARS-CoV in 2003 (Lau et al., 2005; Xu et al., 2005), Middle East Respiratory Syndrome Coronavirus (MERS-CoV) in 2012 (Drosten et al., 2013), and SARS-CoV-2 in 2020 (Nomoto et al., 2020). More recently infectious viral particles were also isolated from feces (Xiao et al., 2020), sputum, oro- and nasopharyngeal swabs (Kujawski et al., 2020; Wölfel et al., 2020), and urine of COVID-19 patients (Sun et al., 2020). Thus, the viruses may enter the water systems through numerous pathways, mostly from sewage discharged from hospitals and isolation and quarantine centers (Adelodun et al., 2020; Nabi et al., 2020), but also from houses and other buildings inhabited or frequented by infected people, whether symptomatic or not (Gandhi et al., 2020; Zhang et al., 2020a).
In fact, a wide viral load variation in clinical samples from people infected with SARS-CoV-2 has been reported in the literature. For instance, around 105–108 copies/L in samples of urine (Frithiof et al., 2020; Han et al., 2020a; Jeong et al., 2020), 106–1011 copies/L in feces (Feng et al., 2021; Kim et al., 2020; Lescure et al., 2020; Xiao et al., 2020), and 104–1014 copies/L in saliva and sputum (Jeong et al., 2020; Pan et al., 2020; Yoon et al., 2020) were reported, which means that a single individual may shed billions of SARS-CoV-2 genomic copies in wastewater at once (Trottier et al., 2020).
Nevertheless, a reduction in viral load is expected when feces and other human excreta reach the sewer system, not only due to the dilution effect, but also related to the inactivation of the viruses by environmental conditions (temperature, pH, solids content), as well as by the presence of antagonists germs (Escherichia coli, Enterococcus spp., Bacillus spp., Clostridium spp., etc.), and chemicals (detergents, disinfectants) in the sewage (Carraturo et al., 2020; Foladori et al., 2020; La Rosa et al., 2020a; Larsen and Wigginton, 2020; Race et al., 2020). Incidentally, due to the recommendations for maintaining good hygiene and minimizing the number of infections, chemicals are being used in quantities far above normal. This increase achieved such a point that there are already studies (Espejo et al., 2020; Horn et al., 2020; Usman et al., 2020; Zambrano-Monserrate et al., 2020; Zhang et al., 2020b) reporting the possible impacts of these chemicals on the environment.
Thanks to the hard work developed by researchers around the world, nowadays there are already studies not only detecting but also quantifying by reverse transcription-quantitative polymerase chain reaction (RT-qPCR) the SARS-CoV-2 RNA in sewage from different countries, in which were found up to 104 copies/L in Montana (Nemudryi et al., 2020), from 104 to 105 copies/L in Detroit (Brijen et al., 2020), up to 106 copies/L in Massachusetts (Schmidt, 2020; Wu et al., 2020a), and up to 107 copies/L in Virginia (Gonzalez et al., 2020) in the USA, 60–350 copies/L in Ahmedabad — India (Kumar et al., 2020a), up to 3.4 × 104 copies/L in the United Arab Emirates (Hasan et al., 2020), 106 copies/L in Murcia — Spain (Randazzo et al., 2020b), and during the epidemic peak, up to 107 copies/L were found in the sewage from Paris — France (Wurtzer et al., 2020). Besides, Gallardo-Escárate et al. (2020) found up to 104 copies/L in sewage samples from a penitentiary and health care facilities in Chillan, a city in Southern Chile, while Gonçalves et al. (2020) found up to 107 copies/L in a hospital wastewater in Ljubljana, Slovenia. In fact, reported viral concentrations vary by several orders of magnitude, which may possibly be related to the level of infections and other specific sewage conditions (Pecson et al., 2020), such as the area served and the resulting balance between industrial, stormwater, domestic and healthcare facilities inputs.
Interestingly, there are also studies reporting the presence of SARS-CoV-2 RNA in sewage samples prior to documentation of the COVID-19 cases at the site. In that case, by re-analyzing stored samples of raw sewage that were collected between October 2019 and February 2020 in five WWTPs in northern Italy, La Rosa et al. (2021) found viral loads of 1.2 × 103, 4.1 × 103, and 2.9 × 104 copies/L in Turin, Milan and Bologna, respectively, on 10–18 December 2019. Importantly, the first autochthonous Italian case of COVID-19 was documented just on 21 February 2020. On 11 February 2020, Martin et al. (2020) detected SARS-CoV-2 RNA in sewage samples collected at a WWTP in South East England, i.e., three days before the first case was reported in that area. Randazzo et al. (2020a) detected SARS-CoV-2 RNA in WWTPs from the metropolitan Region of Valencia, Spain, in late February, when confirmed COVID-19 cases in that region were only incipient. In the same way, on 17 February 2020, Lodder and de Roda Husman (2020) detected SARS-CoV-2 RNA in sewage at Schiphol airport in Tilburg, the Netherlands, only four days after the first case of COVID-19 was confirmed in the country. Also in the Netherlands, on 5 March 2020, Medema et al. (2020) detected SARS-CoV-2 RNA in sewage from Amersfoort six days prior the announcement of the first case in the city. Samples of raw wastewater collected from April to June 2020, in 33 WWTPs of the Czech Republic, in regions with a higher prevalence of COVID-19, confirmed the presence of SARS-CoV-2 RNA in >27.3% of the WWTPs samples (Mlejnkova et al., 2020). In summary, currently, reports from different parts of the planet like Australia (Ahmed et al., 2020a), Brazil (Prado et al., 2020), Canada (D'Aoust et al., 2021), Germany (Westhaus et al., 2021), the United Arab Emirates (Albastaki et al., 2020), Sweden (Saguti et al., 2021), and the United States (Peccia et al., 2020; Sherchan et al., 2020) have shown the feasibility of detecting viral RNA in sewage and WWTP sludge, and the possibility of using these data for environmental surveillance, i.e., to monitor the prevalence of infections among the population via wastewater-based epidemiology (WBE).
Papers reported up till now indicate that coronaviruses remained active in sewage for a few hours–days (Barcelo, 2020; Hart and Halden, 2020; Heller et al., 2020; Michael-Kordatou et al., 2020; Randazzo et al., 2020a), but, depending on the environmental conditions, may persist for up to 14 days, as reported by Wang et al., 2005 after cell culture testes. Thus, whereas the wastewater residence time in sewer systems is typically <24 h (Ye et al., 2016), a substantial viral RNA load can reach the WWTPs. In this sense, Foladori et al. (2020) reported a viral RNA load in the sewage entering a WWTP with a range from 20 to 106 copies/L, which depends on the level of the epidemic. Recently, Zaneti et al. (2021) employed a quantitative microbial risk assessment (QMRA) approach to investigate the potential health risks of SARS-CoV-2 in sewage to WWTPs workers for three COVID-19 pandemic scenarios (moderate, aggressive, and extreme). The study estimated a viral RNA load in the sewage entering the WWTPs ranging from 4.14 × 104 to 5.23 × 106 copies/L, resulting in estimated risks for the aggressive and extreme scenarios (2.63 × 10−3 and 1.3 × 10−2, respectively) above the derived tolerable infection risk for SARS-CoV-2 of 5.5 × 10−4 per person per year. Conversely, also using a QMRA framework to assess the disease risks for WWTP workers, and assuming that 0.03%, 0.3%, and 3% of the wastewater-generating population is infected with SARS-CoV-2, Dada and Gyawali (2020) determined that the average disease risks for low-grade, moderate, and aggressive outbreak scenarios, respectively, are 0.036, 0.32, and 3.21 cases of disease per 1000 exposed WWTP workers.
WWTPs generally adopt conventional treatment processes, usually at a secondary or tertiary level, which, depending on the process and operational conditions, may be sufficient for the elimination of viruses. Typically, a 1.5 to 2 log10 reduction can be expected for viruses in most activated sludge processes, which depends on the solids retention time; while, in the disinfection process, inactivation of 2, 3, and 4 log10 can be achieved with dosages of 1–4, 8–16, and 20–40 mg·min/L of free chlorine or UV irradiation of 30–40, 50–70, and 70–90 mJ/cm2, respectively (Asano et al., 2007).
On 8 and 27 May 2020, Kumar et al. (2021) evaluated the SARS-CoV-2 RNA load along the WWTP of Old Pirana, Ahmedabad, India, which comprises primary, up-flow anaerobic sludge blanket (UASB), aeration pond, and polishing pond treatments. The authors reported a viral RNA load reduction higher than 1.3-log10 by the UASB treatment and no detection of viral RNA after aeration pond nor after the polishing pond. Balboa et al. (2020) investigated the presence of SARS-CoV-2 RNA in various points in the Ourense WWTP in Spain, detecting (7.5–15) × 103 copies/L in raw sewage and observing the absence of viral RNA after primary sedimentation and activated sludge treatment. Also in Spain, but in the Region of Murcia, in a similar study, Randazzo et al. (2020b) got positive results in 11% of samples after activated sludge and 100% negative to SARS-CoV-2 RNA after tertiary treatment (disinfection with sodium hypochlorite and (or) UV). On the other hand, some studies found SARS-CoV-2 RNA in primary, secondary, and even in tertiary wastewater samples. Zhang et al. (2020a) detected strikingly high levels of up to 18.7 × 103 copies/L of SARS-CoV-2 RNA in septic tanks after disinfection with 800 g/m3 sodium hypochlorite at Wuchang Cabin Hospital, in Wuhan — China. Haramoto et al. (2020) detected SARS-CoV-2 RNA with a concentration close to 2.4 × 103 copies/L in 20% of secondary-treated wastewater samples collected from a WWTP in Yamanashi, Japan. Saguti et al. (2021) detected up to 8 × 105 copies/L in raw sewage and 2–75 copies/L in treated sewage from the Rya WWTP, which is discharged into the Göta River. The WWTP, which receives sewage from Gothenburg and surrounding municipalities — Sweden, achieved a mean SARS-CoV-2 RNA reduction of 4-log10. Baldovin et al. (2020) assessed the presence of viral RNA in raw and treated sewage in the two larger WWTPs from Padua, Italy. The authors detected SARS-CoV-2 RNA in 4/9 raw sewage samples and in all two tertiary treated samples analyzed, being that both WWTPs have activated sludge as secondary treatment, and peracetic acid and UV lamps as tertiary treatment. Westhaus et al. (2021) evaluated the efficiency in removing SARS-CoV-2 RNA from three WWTPs, in the Federal State of North Rhine-Westphalia, Germany, and found no significant difference between viral load in samples of raw, secondary (activated sludge) and tertiary sewage (ozonation and filtration). Consequently, as in some cases the SARS-CoV-2 RNA has been detected after disinfection steps, it is also expected to find the viral RNA in receiving water bodies.
When changing the aqueous matrix, from sewage to natural waters, studies become even more scarce in terms of detection and (or) quantification of SARS-CoV-2, with only three investigations available so far. Although Haramoto et al. (2020) have detected SARS-CoV-2 RNA in secondary-treated wastewater, they did not detect in river water samples in Japan. On the other hand, Rimoldi et al. (2020) found viral RNA, on 14 April 2020, in all investigated water bodies (Lambro River, Vettabbia Canal, and Lambro Meridionale River) in the two provinces of Milan and Monza and Brianza, Italy. In a more recent study, Guerrero-Latorre et al. (2020) detected SARS-CoV-2 RNA in rivers from urban streams from Quito, Ecuador, where sewage is discharged directly into natural waters. On 5 June 2020, during the peak case of COVID-19, the authors have found extraordinary high SARS-CoV-2 RNA loads, ranging from 2.07 × 105 to 3.19 × 106 copies/L.
Unfortunately, in low-incoming countries, many areas still lack adequate sanitary infrastructure and wastewater treatment facilities, as well as still practice open defecation, resulting in fecal contamination of the environment and drinking water sources (Adelodun et al., 2019; Bain et al., 2014; Odih et al., 2020; Street et al., 2020). Therefore, the presence of SARS-CoV-2 in natural water may have different origins, such as inefficiency of WWTP (Rimoldi et al., 2020); release of a fraction of raw sewage into surface water, due to the lack of separation of urban runoff water from sewage, leading to combined sewage overflows (Farkas et al., 2020b; Rimoldi et al., 2020); or discharge of raw sewage directly into water bodies, which is a common practice in many parts of the world (Al Huraimel et al., 2020), as in the case of Quito — Ecuador (Guerrero-Latorre et al., 2020), Nicaragua (Vammen and Guillen, 2020), Brazil (de Freitas et al., 2020; Gomes et al., 2020; Stepping, 2016), and in the vast majority of low-income countries (Bastaraud et al., 2020).
The presence of SARS-CoV-2 in sewage may also be an additional concern in communities or regions served by combined sewer systems, as urban flooding, a common threat in rainy periods, that may cause sewage to overflow and, therefore, present new risks of spreading SARS-CoV-2 in the affected areas (Han and He, 2021). Furthermore, sewage-mediated contamination is also a concern in refugee camps (Kassem and Jaafar, 2020; Rafa et al., 2020) and in rural areas from low-incoming countries (Liu et al., 2020b; Lodder and de Roda Husman, 2020; Meng et al., 2020; Miller et al., 2020), where there is a need to share sanitary facilities, such as latrines, and due to the difficulty of maintaining a good hygiene and to access safe water. Yet, due to the disposal of sewage without proper treatment in nature, there are studies indicating recreational water as a potential transmission route for SARS-CoV-2 to humans (Cahill and Morris, 2020; Montagna et al., 2020), and also a potential transmission of SARS-CoV-2 to aquatic mammals (Mathavarajah et al., 2020; Nabi and Khan, 2020). In fact, appropriate attention should be given to the treatment of sewage, principally when the recovery and (or) reuse of such water is desired (D'Alessandro et al., 2020). The use of sewage either un-, partially or treated, for irrigation – a common practice in developing countries – is also a matter of concern (Siddiqui et al., 2020), since the virus could still be present in the treated sewage if it has not been appropriately treated. So, in view of the current situation regarding the COVID-19 pandemic, a deeper discussion on the infectivity of the SARS-CoV-2 from sewage and contaminated water, as well as on possible fecal-oral and fecal-nasal transmissions, is urgent and mandatory. Fig. 3 illustrates the main routes for SARS-CoV-2 to reach the watercourses.
Fig. 3.
The main routes for SARS-CoV-2 to reach the watercourses.
Created with BioRender.com.
5. Infectivity of SARS-CoV-2 from sewage and natural water
As earlier introduced, the occurrence of SARS-CoV-2 RNA in sewage and natural water is a reality covering the planet. The detection and quantification of SARS-CoV-2 in environmental samples such as water and wastewater is mostly performed by using PCR based methods (Barcelo, 2020). Nevertheless, these methods are supported on nucleic acid amplification and the presence of fragments of the viral genome can be sufficient for the detection and even quantification of SARS-CoV-2 RNA (Corpuz et al., 2020; Xu, 2020). However, the key issue is no longer about the detection or quantification of SARS-CoV-2 RNA, but about the infectivity of these viruses when present in water matrices like natural water and sewage. Therefore, the remaining question is, are they infectious or not?
Studies using cell culture and spiking the viruses in water and (or) sewage have been conducted under controlled laboratory conditions with different coronaviruses (SARS-CoV, SARS-CoV-2, and surrogates), in order to determine how long they can survive or remain infectious in different aqueous matrices. According to Gundy et al. (2009), the Feline Infectious Peritonitis Virus (FIPV) and the Human Coronavirus 229 (HCoV 229), assayed in Crandell Reese feline kidney cell line (ATCC-94) and fetal human lung fibroblast — MRC-5 cell line (ATCC-171), respectively, achieved times to 2-log10 reduction (99% reduction (T99) of infectious viral particles) in tap water of 6.8 days at 23 °C and >87 days at 4 °C, while in sewage the T99 was 1.6–2.4 days at 23 °C. The authors concluded that the coronaviruses inactivation was strongly dependent on temperature, organic matter content, and the presence of antagonistic bacteria. Casanova et al. (2009) observed a T99 in reagent-grade water at 25 °C for two other coronaviruses, Mouse Hepatitis Virus (MHV) and Transmissible Gastroenteritis Virus (TGEV), assayed in delayed brain tumor and swine testicular culture cells, of 17 and 22 days, respectively, whereas in pasteurized settled sewage the values were even smaller, 7 and 9 days. On the other hand, at 4 °C, the reduction in infectivity was <1-log10 after four weeks, regardless of the aqueous matrix and for both studied coronaviruses. Wang et al. (2005) noted that SARS-CoV, assayed in Vero E6 cells, may survive in the sewage for 2 days at 20 °C and up to 14 days at 4 °C. In addition, viral RNA may be detected for >6 days after virus inactivation. Other authors also endorsed that the viral RNA can persist longer than infectious viral particles in different aqueous matrices (Bivins et al., 2020; Wu et al., 2020a). Ahmed et al. (2020b) observed 90% RNA reduction of MHV at 4–37 °C after 7.4–56.6 days in raw sewage, 5.6–43.1 days in autoclaved sewage, and 10.9–43.9 days in tap water, while the times for 90% RNA reduction of SARS-CoV-2 was 8.0–27.8 days in raw sewage, 5.7–43.2 days in autoclaved sewage, and 9.4–58.6 days in tap water. More recently, by using SARS-CoV-2 nCoV-WA1–2020 (MN985325.1) isolated from a clinical patient diagnosed with COVID-19 and assaying on high-passage Vero E6 cells, Bivins et al. (2020) demonstrated that the SARS-CoV-2 can survive longer in tap water than in sewage, showing times for 1-log10 reduction (90% reduction (T90) of infectious virus particles) of 1.7 and 1.5 day, at room temperature, respectively. The authors also observed that SARS-CoV-2 remained infectious for three days in the low-titer (103 TCID50/mL) tests and for the entire seven days duration of the high-titer (105 TCID50/mL) tests. In another study, Lee et al. (2020) demonstrated that the stability of SARS-CoV-2 varies according to the water matrix, so that, starting with a concentration of 104 PFU/mL in Vero cells (ATCC® CCL-81™), SARS-CoV-2 was not infectious in seawater and freshwater tests, but remained infectious for two days in tap water. In Vero cells tests, the authors determined 103 PFU/mL as the minimum infective dose of SARS-CoV-2. Finally, Yang et al. (2020a) assessed the possibility of spreading COVID-19 through environmental media, using a modified QMRA index model and data from three typical rivers (Fu, Han, and Yangtze) and watershed cities in China's Hubei province, concluding the risk index is gradually decreasing from Wuhan city to the neighboring cities, with values in the order of 10−8, 10−10, and 10−12 in the Fu, Han, and Yangtze river basins, respectively. The authors claimed for further investigation on the subject since the survival of the virus could be affected by many human and environmental factors and the researches on the risk of transmission of SARS-CoV-2 in the aquatic environment are limited so far. In Table 1 is summarized the information on the coronaviruses investigated, displaying the time to reduce the viable virions, their survival time, and RNA detection time on different aqueous matrices and under different conditions.
Table 1.
Information on the coronaviruses studied: time to reduce viable virions, their survival time, and RNA detection time on different aqueous matrices.
| Reference (alphabetical order) | Coronavirus | Aqueous matrix | Sample characteristics | Test condition | 1-Log reduction of viable virions (T90) | 2-Log reduction of viable virions (T99) | 3-Log reduction of viable virions (T99.9) | Virion survival time | RNA detection time |
|---|---|---|---|---|---|---|---|---|---|
| (Bivins et al., 2020) | SARS-CoV-2 | Tap water | Not reported | High titer (105 TCID50/mL) at 20 °C | 2.0 d | 3.9 d | – | – | – |
| Wastewater | pH 7.98, COD 153 mg/L, ammonia 13.3 mg/L, and TSS 190 mg/L | Low titer (103 TCID50/mL) at 20 °C | 2.1 d | 4.3 d | – | – | – | ||
| High titer (105 TCID50/mL) at 20, 50, and 70 °C | 1.6 d, 15 min, and 2.2 min | 3.2 d, 30 min, and 4.5 min | – | – | – | ||||
| (Casanova et al., 2009) | TGEV | Reagent-grade water | pH 6.0, turbidity 0.1 NTU | 4.5–5.8 MPN/mL at 4 and 25 °C | 110 d and 11 d | 220 d and 22 d | 330 d and 33 d | – | – |
| Pasteurized settled sewage | pH 7.6, turbidity 17.6 NTU | 24 and 4 d | 49 and 9 d | 73 d and 14 d | – | – | |||
| MHV | Reagent-grade water | pH 6.0, turbidity 0.1 NTU | 6.5–6.9 MPN/mL at 4 and 25 °C | >365 d and 9 d | >365 d and 17 d | >365 d and 26 d | – | – | |
| Pasteurized settled sewage | pH 7.6, turbidity 17.6 NTU | 35 d and 3 d | 70 d and 7 d | 105 d and 14 d | – | – | |||
| (Gundy et al., 2009) | FIPV | Tap water filtered | Tap water filtered through 0.2-μm pore size filter | 105 TCID50/mL at 4 and 23 °C | – | 87 d and 6.76 d | 130 d and 10.1 d | – | – |
| Unfiltered tap water | pH 7.8, TSS 297 mg/L, TOC 0.1 mg/L | 105 TCID50/mL at 23 °C | – | 8.32 d | 12.5 d | – | – | ||
| Secondary sewage | BOD and TSS of 5.5–22 mg/L | – | 1.62 d | 2.42 d | – | – | |||
| Primary sewage filtered | Primary sewage filtered through 0.2-μm pore size filter | – | 1.60 d | 2.40 d | – | – | |||
| Primary sewage unfiltered | BOD and TSS of 110–220 mg/L | – | 1.71 d | 2.56 d | – | – | |||
| HCoV 229E | Tap water filtered | Tap water filtered through 0.2-μm pore size filter | 105 TCID50/mL at 4 and 23 °C | – | 392 d and 6.76 d | 588 d and 10.1 d | – | – | |
| Unfiltered tap water | pH 7.8, TSS 297 mg/L, TOC 0.1 mg/L | 105 TCID50/mL at 23 °C | – | 8.09 d | 12.1 d | – | – | ||
| Secondary sewage | BOD and TSS of 5.5–22 mg/L | – | 1.85 d | 2.77 d | – | – | |||
| Primary sewage filtered | primary sewage filtered through 0.2-μm pore size filter | – | 1.57 d | 2.35 d | – | – | |||
| Primary sewage unfiltered | BOD and TSS of 110–220 mg/L | – | 2.36 d | 3.54 d | – | – | |||
| (Lee et al., 2020) | SARS-CoV-2 | Tap water | pH 7.27, turbidity 0.03 FTU, residual chlorine 1.0 mg/L | 104 and 105 PFU/mL at 23 °C | – | – | – | 3 d and 6 d | – |
| Fresh water | pH 7.38, turbidity 4.15 FTU; Inje-gun, Korea, | 104 and 105 PFU/mL at 23 °C | – | – | – | 2 d | – | ||
| Seawater | pH 8, turbidity 1.45 FTU, salinity 3.2%, Sokcho, Korea | 104 and 105 PFU/mL at 23 °C | – | – | – | 1 d | – | ||
| (Wang et al., 2005) | SARS-CoV | Sewage | Residual chlorine 1.0 mg/L | 105 TCID50/mL at 4 and 20 °C | – | – | – | >14 d and 2 d | 8 d (at 20 °C) |
| (Ye et al., 2016) | MHV | Pasteurized sewage | pH 7.6, TSS 235 mg/L, and background bacteriophage concentrations tested with E. coli ATCC 15597 800–1000 PFU/mL | 3 × 104 PFU/mL at 10 and 25 °C | 149 h and 19 h | – | – | – | – |
| untreated sewage | 36 h and 13 h | – | – | – | – |
Abbreviations: BOD: biological oxygen demand, COD: chemical oxygen demand; FIPV: feline infectious peritonitis virus; HCoV: human coronavirus; MHV: murine hepatitis virus; TGEV: transmissible gastroenteritis virus; TOC: total organic carbon, TSS: total suspended solids.
In fact, most human viruses are very difficult to maintain in vitro and their culture requires specialized equipment and trained personnel; hence, infectivity analyzes are rarely carried out on sewage or natural water samples (Farkas et al., 2020a; Maal-Bared et al., 2020). Unfortunately, to our knowledge, there are only two studies addressing the infectivity of SARS-CoV-2 from raw and treated sewage and (or) natural water (Rimoldi et al., 2020; Westhaus et al., 2021) and one study assessing the infectivity of SARS-CoV in sewage (Wang et al., 2005). As already mentioned, Rimoldi et al. (2020) assessed the presence and infectivity of SARS-CoV-2 in raw, secondary, and tertiary sewage from three WWTPs within the Milan Metropolitan Area, Italy, in addition to their receiving water bodies. The results were negative for infectivity, although SARS-CoV-2 RNA was detected in samples of raw sewage and of receiving water bodies. In that occasion, the authors precautionary have claimed for further studies to confirm the non-infectious character of sewage and natural water samples testing positive for viral RNA. In another study, Westhaus et al. (2021) investigated the presence of SARS-CoV-2 in raw and treated sewage and the infectivity in raw sewage from nine municipal WWTP in different cities of the Federal State of North Rhine-Westphalia, Germany. 24 h flow-dependent composite samples were collected, pretreated by centrifugation at 4700 ×g for 30 min, concentrated about 100 times by centrifugal ultrafiltration for 15 min at 3500 ×g using Amicon® Ultra-15 filters, and then assayed. Although the authors detected viral RNA in all samples, which ranged from 3 × 103 to 2 × 104 copies/L, the results for infectivity were null when assessed by viral outgrowth test with a potential SARS-CoV-2 host cell line in vitro. The authors indicate that sewage does not appear to be an important route of transmission, but they emphasize the importance of carrying out further studies, especially regarding the methodological validation for enveloped viruses, given the small sample volume used in these analyzes and the huge viral load released every day by the studied WWTPs into the receiving water bodies, which ranges from 6 × 1010 to 6 × 1012 SARS-CoV-2 RNA copies per day per WWTP. Moreover, Feng et al. (2021) also stated that the presence of viral RNA does not confirm the existence of infectious viral particles, but indicates that possibly some could be present. Wang et al. (2005) also did not observe infectious viruses — SARS-CoV was studied in that time, when assessing samples of raw sewage from the 309th Hospital of the Chinese People's Liberation Army. On that occasion, even after negative results for infectivity, the authors suggest the sewage system as a possible transmission route, since they have demonstrated the virus can survive for two days in the sewage at 20 °C and up to 14 days at 4 °C. So, the matter is, in which physical form were the viruses from samples having no infectivity? Were they intact, with capsid, or there were only RNA strands and other viral fragments in the analyzed samples?
Aiming to prevent the deterioration of the virus, Rimoldi et al. (2020) carefully carried out the infectivity assays without concentrating the samples, since it has already been reported on the literature (Ye et al., 2016) that the addition of chemical compounds or mechanical stress, that usually occur in the concentration stage, could harm the viability of virions. Nevertheless, the authors (Rimoldi et al., 2020) did not quantify the viral load found in any sample and carried out instantaneous sampling at 1.00 p.m.. These two aspects are extremely relevant in virus analysis, especially concerning the infectivity analysis, even because the authors might have collected the grab samples at a time when the sewage flow rate and (or) the fecal load were minimal.
The selection of the collection time is essential for the case of grab sampling because, in this sampling method, it is generally desired to collect in daily peaks of the sewage flow rate, which, in the case of most cities, occurs in the morning or evening (Amoah et al., 2020; Michael-Kordatou et al., 2020). Alternatively, to be more precise, the choice of time for grab sampling should be based on a study with indicators of human fecal load in the sewage (Corpuz et al., 2020), such as fecal coliforms (Wang et al., 2020a), antibiotics (Sims and Kasprzyk-Hordern, 2020), etc., for example, and thus could be correlated with the peak of SARS-CoV-2 viral shedding. On the other hand, 24-h composite sampling could also be done but, in this case, it would result in average daily values, making it impossible to discriminate any peak value recorded in the sampling period (Bishop et al., 2020), in addition to possibly be able to impair the persistence of the virus infectivity.
Another important factor to consider when performing infectivity analysis, especially for enveloped viruses, is the pre-treatment of the samples. Rimoldi et al. (2020) pre-filtered the samples on glass fiber filters (0.7 μm nominal pore size, 47 mm diameter) until filter clogging and then further filtrated 50 mL on nitrocellulose Millipore filters (0.22 μm nominal pore size, 47 mm diameter), focusing on the detection and isolation of SARS-CoV-2 just on the 0.22 μm filtered fraction.
It is of common knowledge that to carry out virus analysis the samples must be pre-treated, especially those from environmental nature, and that filtration in a 0.22 μm pore-size filter is a routine practice. Conversely, it is also well known that the practices were established for analysis of non-enveloped viruses and are being adapted for enveloped viruses (Carducci et al., 2020; La Rosa et al., 2020b; Lu et al., 2020; Venugopal et al., 2020; Yang et al., 2020b), as is occurring with SARS-CoV-2. In addition, studies indicate that filtration through 0.22 μm pore-size filters can reduce the viral load and, thus, impair the infectivity analysis of enveloped viruses, especially those from the Coronaviridae family.
In the study conducted by Ye et al. (2016) 26% of an enveloped virus belonging to the Coronaviridae family, murine hepatitis virus (MHV) (100 nm particle size), were adsorbed by the wastewater solids (medium-strength municipal wastewater having 235 mg/L of total suspended solids (TSS)) at equilibrium in approximately 15 min. Subsequently, this fraction of viruses adsorbed to the suspended solids were retained in the 0.22 μm pore-size filter. Then, the filtrate fractions of the samples were concentrated 100 times by ultrafiltration, and infective viruses in the concentrates were measured with plaque assays, resulting in a mean recovery of 25.1% for infectious MHV. In summary, only after the authors have concentrated the samples 100 times the recovery of 25% of the infectious viruses was accomplished. In an earlier study, Wang et al. (2005) investigated a virus concentration method based on the adsorption on electropositive filter media particle columns, elution with broth at pH 7.2, and precipitation by 10% polyethylene glycol. They analyzed 100 mL of sewage spiked with SARS-CoV and, after the concentration stage, a mean recovery efficiency of 1.02% was achieved.
As a result of the studies above referred, some questions were left open. If no concentration stage was applied, would the recovery from infectious viruses be at most 0.25%? When did the viruses lose their infectivity, in the pre-filtration stage or in the concentration stage?
Answering these questions was not the objective of the researches above referred but in fact it brought about these new questions on the filtration steps for the removal of the wastewater suspended solids prior the infectivity assays and how make them viable for enveloped viruses in complex samples. On the other end, other studies isolated infectious SARS-CoV-2 from feces of a patient with severe COVID-19 by filtering the samples through 0.45 μm pore-size filters (Xiao et al., 2020).
Considering the cautiousness necessary to preserve the integrity of enveloped viruses such as SARS-CoV-2 in infectivity analyzes, the choice of the filter to be used in the sample pre-treatment seems to have an important role in this type of analysis. Besides, microfiltration is expected to remove the viral load by up to 2-log10 (Asano et al., 2007). Therefore, selecting a filter with a pore size of 0.45 μm instead of 0.22 μm seems to be more appropriate, when the main objective is the analysis of the infectivity of SARS-CoV-2 in water and sewage. However, these two filters can remove not only the sample suspended solids but also the virus fraction that is adsorbed to sample solids, as reported by Ye et al. (2016). The 0.22 μm pore-size filter, being narrower, has a greater capacity for retaining the virus fraction that is not adsorbed to solids, because, as previously reported, SARS-CoV-2 has a particle size of 100–200 nm, which is very close to the pore size of the filter, which in this case is 220 nm. Moreover, it is also important to determine TSS in the samples, because the higher the solids content, the greater the potential amount of virus adsorbed on the solids and the lower the virus concentration in the filtrate. Furthermore, suspended solids can adsorb on the membrane surface, clogging or reducing the pores of the filter (de Souza et al., 2020), further increasing virus retention and reducing its concentration in the filtrate. It is possible to carry out an analogy between the behavior of viral particles and small solutes during filtration operations with membranes (filters) with a pore size of 0.22 μm. Using a 0.22 μm pore-size membrane as a tertiary treatment of domestic sewage, Gómez et al. (2007) observed the membrane retained about 60% of bisphenol A, a small solute with molar mass of 229 g/mol and Stokes radius (Zhao et al., 2015) of approximately 1.1 nm. In addition, the shear force imposed on a viral particle during a filtration operation using a 0.22 μm pore-size membrane (filter) is greater than that using a 0.45 μm pore-size one, which increases the possibility of capsid rupture and, consequently, the possible virus inactivation.
Therefore, to ensure the absence of infectivity in sewage and other complex aqueous matrices, complementary analyzes showing the physical structure of the viruses, if they are intact, with capsid, or if there are only RNA strands and other viral fragments in the analyzed samples, is of paramount importance for public health, including the definition of new public policies in the area of sanitation, especially in low-incoming countries that have low rates of sanitation, and, consequently, high rates of infections related to waterborne diseases.
How these issues may then be improved? Currently, many world-renowned research groups have high-resolution image acquisition equipment available to elucidate this problem, such as electron microscopes. These instruments allow the acquisition of images of the viruses, enabling to check if their physical structure is intact (with capsid) or deteriorated, corroborating on the evaluation of the non-infectivity of the investigated sample. Otherwise, in some cases, a negative SARS-CoV-2 infectivity test could be interpreted as a mere false negative.
For instance, Hindson (2020) employed electron microscopy to demonstrate that SARS-CoV-2 virions were live, i.e. infectious, in stool samples from two patients testing positive for RNA viral. Santarpia et al. (2020) conducted a study during the initial isolation of 13 individuals with COVID-19 at the University of Nebraska Medical Center, collecting air and surface samples to examine the viral shedding from isolated individuals. By using transmission electron microscopy (TEM) on samples from the windowsill, the authors confirmed the presence of intact SARS-CoV-2 virions after three days of cell culture. Other studies have also effectively used TEM to detect live virions in samples collected from stool (Wang et al., 2020b; Xiao et al., 2020; Zhang et al., 2020c) and human airway epithelial cells (Zhu et al., 2020). These studies, therefore, show that, despite being more laborious and expensive, by using TEM it is possible to visualize the virions and thus guarantee whether they are infectious or not. In fact, it is well known that cell culture is the gold standard for testing viral viability, but due to the capsid weakness of enveloped viruses, electron microscopy could or even should be used as a complementary analysis to cell culture in the analyzes of infectivity of enveloped viruses such as coronaviruses. The association of these techniques would allow confirming whether the viruses used in cell culture have their structure intact or if there are only viral fragments. Fig. 4 shows Coronaviruses images acquired with TEM.
Fig. 4.
Visualization of Coronaviruses with transmission electron microscopy: A) MERS-CoV, B) SARS-CoV-2, and C) Human Coronavirus 229E.
Photos A and B credit to Dr. Cynthia Goldsmith and Dr. Azaibi Tamin, and Photo C credit to Dr. Fred Murphy. These images comes from the Centers for Disease Control and Prevention's (CDC) Public Health Image Library (PHIL), identification numbers 15908 (https://phil.cdc.gov/Details.aspx?pid=15908), 23640 (https://phil.cdc.gov/Details.aspx?pid=23640), and 15523 (https://phil.cdc.gov/Details.aspx?pid=15523), respectively.
6. Concluding remarks and recommendations
The available literature on the pathways for spread and infectivity of SARS-CoV-2 in environmental aqueous matrices (natural water and sewage) investigated to date were reviewed, indicating a wide availability of documentation on dissemination but just two papers addressing the infectivity of SARS-CoV-2 on sewage and (or) natural water samples. Despite the lack of data on the survival and infectivity of SARS-CoV-2 in water and sewage and the unconfirmed contagion via these aqueous matrices, the infectivity of these viruses must be thoroughly investigated to assess potential health risks, mainly regarding the possible transmission via fecal-oral and virus-laden aerosol borne-routes. This information would be extremely important, especially for communities with poor sanitation, where people are likely to be exposed to contaminated water or even raw sewage.
In fact, the humanity is building the knowledge about the infectivity of SARS-CoV-2, what is very important to combat, control, and mitigate the COVID-19 pandemic effects, for both purposes: to save lives and the global economy. Otherwise, the pandemic could last for a long time, causing much greater impacts than it has already caused. In the meantime, we must fight the ongoing pandemic by acting quickly but cautiously, assessing all possible routes of transmission. In the particular case of infectivity of SARS-CoV-2 in sewage and water, we believe that subsequent studies should address the following points:
-
•
Assess the behavior of different filters in the pre-filtration stage of the sample, monitoring the pressure applied to determine the shear rate. Hence, choosing the filter that allows the passage of the largest fraction of live enveloped viruses, but that is also capable of removing the suspended solids from the sample.
-
•
Investigate filtration in a different mode of operation, membrane tangential filtration like micro/ultrafiltration. The control of the operating parameters of tangential flow rate and transmembrane pressure would allow virus adsorption minimization, as well as keeping the virus intact.
-
•
Investigate the possibility of using submerged membranes at low pressures in the pre-filtration (microfiltration) and concentration (ultrafiltration) steps.
-
•
Investigate the influence of different angular velocities during centrifugation and centrifugal filtration when used as pre-treatment or concentration steps, in the inactivation of enveloped viruses, and thus determine the shear force required for capsid rupture.
-
•
Determine the proportion of active (infectious) viruses in each sample-processing step, confirming the analysis by cell culture tests.
-
•
Develop alternative techniques that allow confirming the physical integrity (viability) of enveloped viruses.
-
•
Finally, develop and standardize pre-treatment and concentration methodologies specifically for analyzing enveloped viruses in complex aqueous matrices such as sewage and natural water. Importantly, the recovery efficiency should be confirmed by cell culture tests and not just via PCR analyzes.
The following is the supplementary data related to this article.
Bibliographic search results in Pub Med, SCOPUS, and ScienceDirect databases.
CRediT authorship contribution statement
Alexandre Giacobbo: Writing – original draft, Conceptualization, Investigation, Writing – review & editing. Marco Antônio Siqueira Rodrigues: Conceptualization, Investigation, Writing – review & editing. Jane Zoppas Ferreira: Conceptualization, Investigation, Writing – review & editing. Andréa Moura Bernardes: Writing – original draft, Conceptualization, Investigation, Writing – review & editing. Maria Norberta de Pinho: Conceptualization, Investigation, Writing – review & editing.
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgments
The authors gratefully acknowledge to the Brazilian funding agencies (CAPES, CNPq, FINEP and FAPERGS); to the Portuguese Funding Agency, Fundação para a Ciência e Tecnologia (FCT), CeFEMA [grant number 325UID/CTM/04540/2013]; and to the Programa Iberoamericano de Ciencia y Tecnología para el Desarollo (CYTED) by the financial support.
Editor: Damia Barcelo
References
- Adelodun B., Odedishemi F., Segun M., Choi K.S. Dosage and settling time course optimization of Moringa oleifera in municipal wastewater treatment using response surface methodology. Desalin. Water Treat. 2019;167:45–56. doi: 10.5004/dwt.2019.24616. [DOI] [Google Scholar]
- Adelodun B., Odedishemi F., Gbemisola R., Olalekan H., Choi K. Snowballing transmission of COVID-19 (SARS-CoV-2) through wastewater: any sustainable preventive measures to curtail the scourge in low-income countries? Sci. Total Environ. 2020;742:140680. doi: 10.1016/j.scitotenv.2020.140680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ahmed W., Angel N., Edson J., Bibby K., Bivins A., Brien J.W.O., Choi P.M., Kitajima M., Simpson S.L., Li J., Tscharke B., Verhagen R., Smith W.J.M., Zaugg J., Dierens L., Hugenholtz P., Thomas K., V, Mueller J.F. First confirmed detection of SARS-CoV-2 in untreated wastewater in Australia: a proof of concept for the wastewater surveillance of COVID-19 in the community. Sci. Total Environ. 2020;728:138764. doi: 10.1016/j.scitotenv.2020.138764. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ahmed W., Bertsch P.M., Bibby K., Haramoto E., Hewitt J., Huygens F., Gyawali P., Korajkic A., Riddell S., Sherchan S.P., Simpson S.L., Sirikanchana K., Symonds E.M., Verhagen R., Vasan S.S., Kitajima M., Bivins A. Decay of SARS-CoV-2 and surrogate murine hepatitis virus RNA in untreated wastewater to inform application in wastewater-based epidemiology. Environ. Res. 2020;191:110092. doi: 10.1016/j.envres.2020.110092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Al Huraimel K., Alhosani M., Kunhabdulla S., Stietiya M.H. SARS-CoV-2 in the environment: modes of transmission, early detection and potential role of pollutions. Sci. Total Environ. 2020;744:140946. doi: 10.1016/j.scitotenv.2020.140946. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Albastaki A., Naji M., Lootah R., Almeheiri R., Almulla H., Almarri I., Alreyami A., Aden A., Alghafri R. First confirmed detection of SARS-COV-2 in untreated municipal and aircraft wastewater in Dubai, UAE: the use of wastewater based epidemiology as an early warning tool to monitor the prevalence of COVID-19. Sci. Total Environ. 2020;143350 doi: 10.1016/j.scitotenv.2020.143350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ali M., Zaid M., Saqib M.A.N., Ahmed H., Afzal M.S. SARS-CoV-2 and the hidden carriers: sewage, feline, and blood transfusion. J. Med. Virol. 2020;92:2291–2292. doi: 10.1002/jmv.25956. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Amirian E.S. Diseases potential fecal transmission of SARS-CoV-2: current evidence and implications for public health. Int. J. Infect. Dis. 2020;95:363–370. doi: 10.1016/j.ijid.2020.04.057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Amoah I.D., Kumari S., Bux F. Coronaviruses in wastewater processes: source, fate and potential risks. Environ. Int. 2020;143:105962. doi: 10.1016/j.envint.2020.105962. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arslan M., Xu B., Gamal El-Din M. Transmission of SARS-CoV-2 via fecal-oral and aerosols-borne routes: environmental dynamics and implications for wastewater management in underprivileged societies. Sci. Total Environ. 2020;743:140709. doi: 10.1016/j.scitotenv.2020.140709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Asano T., Burton F.L., Leverenz H., Tsuchihashi R., Tchobanoglous G. 1st ed. McGraw-Hill Professional Publishing; New York: 2007. Water Reuse: Issues, Technologies, and Applications. [Google Scholar]
- Atkinson B., Petersen E. SARS-CoV-2 shedding and infectivity. Lancet. 2020;395:1339–1340. doi: 10.1016/S0140-6736(20)30868-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bahrami A., Ferns G.A. Genetic and pathogenic characterization of SARS-CoV-2: a review. Future Virol. 2020 doi: 10.2217/fvl-2020-0129. [DOI] [Google Scholar]
- Bain R., Cronk R., Hossain R., Bonjour S., Onda K., Wright J., Yang H., Slaymaker T., Hunter P., Prüss-Ustün A., Bartram J. Global assessment of exposure to faecal contamination through drinking water based on a systematic review. Trop. Med. Int. Heal. 2014;19:917–927. doi: 10.1111/tmi.12334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Balboa S., Mauricio-Iglesias M., Rodríguez S., Martínez-Lamas L., Vasallo F.J., Regueiro B., Lema J.M. The fate of SARS-CoV-2 in wastewater treatment plants points out the sludge line as a suitable spot for incidence monitoring. medRxiv. 2020 doi: 10.1101/2020.05.25.20112706. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baldovin T., Amoruso I., Fonzo M., Buja A., Baldo V., Cocchio S., Bertoncello C. SARS-CoV-2 RNA detection and persistence in wastewater samples: an experimental network for COVID-19 environmental surveillance in Padua, Veneto Region (NE Italy) Sci. Total Environ. 2020;143329 doi: 10.1016/j.scitotenv.2020.143329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barcelo D. An environmental and health perspective for COVID-19 outbreak: meteorology and air quality influence, sewage epidemiology indicator, hospitals disinfection, drug therapies and recommendations. J. Environ. Chem. Eng. 2020;8:104006. doi: 10.1016/j.jece.2020.104006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bastaraud A., Cecchi P., Handschumacher P., Altmann M., Jambou R. Urbanization and waterborne pathogen emergence in low-income countries: where and how to conduct surveys? Int. J. Environ. Res. Public Health. 2020;17:480. doi: 10.3390/ijerph17020480. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bhowmick G.D., Dhar D., Dhar D., Nath D., Ghangrekar M.M., Banerjee R., Das S., Chatterjee J. Coronavirus disease 2019 (COVID-19) outbreak: some serious consequences with urban and rural water cycle. NPJ Clean Water. 2019;3:1–8. doi: 10.1038/s41545-020-0079-1. [DOI] [Google Scholar]
- Bishop N., Jones-Lepp T., Margetts M., Alvarez D., Keil D.E. Wastewater-based epidemiology pilot study to examine drug use in the Western United States. Sci. Total Environ. 2020:140697. doi: 10.1016/j.scitotenv.2020.140697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bivins A., Greaves J., Fischer R., Yinda K.C., Ahmed W., Kitajima M., Munster V.J., Bibby K. Persistence of SARS-CoV-2 in water and wastewater. Environ. Sci. Technol. Lett. 2020 doi: 10.1021/acs.estlett.0c00730. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bojkova D., Klann K., Koch B., Widera M., Krause D., Ciesek S., Cinatl J., Münch C. Proteomics of SARS-CoV-2-infected host cells reveals therapy targets. Nature. 2020;583:469–472. doi: 10.1038/s41586-020-2332-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brijen M., Xavier F., John N., Anna M., Irene X. SARS-CoV-2 in Detroit wastewater. J. Environ. Eng. 2020;146:6020004-1–6020004-6. doi: 10.1061/(ASCE)EE.1943-7870.0001830. [DOI] [Google Scholar]
- Cahill N., Morris D. Recreational waters – a potential transmission route for SARS-CoV-2 to humans? Sci. Total Environ. 2020;740:140122. doi: 10.1016/j.scitotenv.2020.140122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carducci A., Federigi I., Liu D., Thompson J.R., Verani M. Making waves: coronavirus detection, presence and persistence in the water environment: state of the art and knowledge needs for public health. Water Res. 2020;179:115907. doi: 10.1016/j.watres.2020.115907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carosso A., Cosma S., Benedetto C. Vaginal delivery in COVID-19 pregnant women: anorectum as a potential alternative route of SARS-CoV-2 transmission. Am. J. Obstet. Gynecol. 2020;223:612. doi: 10.1016/j.ajog.2020.06.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carosso A., Cosma S., Borella F., Marozio L., Coscia A., Ghisetti V., Di Perri G., Benedetto C. Pre-labor anorectal swab for SARS-CoV-2 in COVID-19 pregnant patients: is it time to think about it? Eur. J. Obstet. Gynecol. Reprod. Biol. 2020;249:98–99. doi: 10.1016/j.ejogrb.2020.04.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carraturo F., Del Giudice C., Morelli M., Cerullo V., Libralato G., Galdiero E., Guida M. Persistence of SARS-CoV-2 in the environment and COVID-19 transmission risk from environmental matrices and surfaces. Environ. Pollut. 2020;265:115010. doi: 10.1016/j.envpol.2020.115010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Casanova L., Rutala W.A., Weber D.J., Sobsey M.D. Survival of surrogate coronaviruses in water. Water Res. 2009;43:1893–1898. doi: 10.1016/j.watres.2009.02.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen B., Tian E., He B., Tian L., Han R., Wang S., Xiang Q., Zhang S. Overview of lethal human coronaviruses. Signal Transduct. Target. Ther. 2020;5:1–16. doi: 10.1038/s41392-020-0190-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Jiahui, Wang R., Wang M., Wei G.-W. Mutations strengthened SARS-CoV-2 infectivity. J. Mol. Biol. 2020;432:5212–5226. doi: 10.1016/j.jmb.2020.07.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Jun, Lu H., Melino G., Boccia S., Piacentini M., Ricciardi W., Wang Y., Shi Y., Zhu T. COVID-19 infection: the China and Italy perspectives. Cell Death Dis. 2020;11:438. doi: 10.1038/s41419-020-2603-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen N., Zhou M., Dong X., Qu J., Gong F., Han Y., Qiu Y., Wang J., Liu Y., Wei Y., Xia J., Yu T., Zhang X., Zhang L. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet. 2020;395:507–513. doi: 10.1016/S0140-6736(20)30211-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Y., Chen L., Deng Q., Zhang G., Wu K., Ni L., Yang Y., Liu B., Wang W., Wei C., Yang J., Ye G., Cheng Z. The presence of SARS-CoV-2 RNA in the feces of COVID-19 patients. J. Med. Virol. 2020;833–840 doi: 10.1002/jmv.25825. [DOI] [PubMed] [Google Scholar]
- Corpuz M.V.A., Buonerba A., Vigliotta G., Zarra T., Ballesteros F., Campiglia P., Belgiorno V., Korshin G., Naddeo V. Viruses in wastewater: occurrence, abundance and detection methods. Sci. Total Environ. 2020;140910 doi: 10.1016/j.scitotenv.2020.140910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dada A.C., Gyawali P. Quantitative microbial risk assessment (QMRA) of occupational exposure to SARS-CoV-2 in wastewater treatment plants. Sci. Total Environ. 2020;142989 doi: 10.1016/j.scitotenv.2020.142989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- D’Alessandro D., Gola M., Appolloni L., Dettori M., Fara G.M., Rebecchi A., Settimo G., Capolongo S. COVID-19 and living space challenge. Well-being and public health recommendations for a healthy, safe, and sustainable housing. Acta Bio Med. Atenei Parm. 2020;91:61–75. doi: 10.23750/abm.v91i9-S.10115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- D’Aoust P.M., Mercier E., Montpetit D., Jia J.-J., Alexandrov I., Neault N., Baig A.T., Mayne J., Zhang X., Alain T., Langlois M.-A., Servos M.R., MacKenzie M., Figeys D., MacKenzie A.E., Graber T.E., Delatolla R. Quantitative analysis of SARS-CoV-2 RNA from wastewater solids in communities with low COVID-19 incidence and prevalence. Water Res. 2021;188:116560. doi: 10.1016/j.watres.2020.116560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Daraei H., Toolabian K., Kazempour M., Javanbakht M. The role of the environment and its pollution in the prevalence of COVID-19. J. Inf. Secur. 2020;81:e168–e169. doi: 10.1016/j.jinf.2020.06.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- da Silva R.R., dos Santos M.B., dos Santos A.D., Tavares D.D.S., dos Santos P.L. Coronavirus disease and basic sanitation: too early to be worried? J. Braz. Soc. Trop. Med. 2020;53:1–5. doi: 10.1590/0037-8682-0345-2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Freitas D.A.F., Kuwajima J.I., dos Santos G.R. Water resources, public policies and the COVID-19 pandemic. Ambient. Água Interdiscip. J. Appl. Sci. 2020;15:e2540. doi: 10.4136/1980-993X. [DOI] [Google Scholar]
- de Souza D.I., Giacobbo A., Fernandes E.D.S., Rodrigues M.A.S., de Pinho M.N., Bernardes A.M. Experimental design as a tool for optimizing and predicting the nanofiltration performance by treating antibiotic-containing wastewater. Membranes (Basel) 2020;10:1–15. doi: 10.3390/membranes10070156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Di Maria F., Beccaloni E., Bonadonna L., Cini C., Confalonieri E., La Rosa G., Milana M.R., Testai E., Scaini F. Minimization of spreading of SARS-CoV-2 via household waste produced by subjects affected by COVID-19 or in quarantine. Sci. Total Environ. 2020;743:140803. doi: 10.1016/j.scitotenv.2020.140803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ding S., Liang T.J. Is SARS-CoV-2 also an enteric pathogen with potential fecal-oral transmission? A COVID-19 virological and clinical review. Gastroenterology. 2020;159:53–61. doi: 10.1053/j.gastro.2020.04.052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ding Z., Qian H., Xu B., Huang Y., Miao T., Yen H.-L., Xiao S., Cui L., Wu X., Shao W., Song Y., Sha L., Zhou L., Xu Y., Zhu B., Li Y. Toilets dominate environmental detection of severe acute respiratory syndrome coronavirus 2 in a hospital. Sci. Total Environ. 2021;753:141710. doi: 10.1016/j.scitotenv.2020.141710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Donà D., Minotti C., Costenaro P., Da Dalt L., Giaquinto C. Fecal-oral transmission of SARS-CoV-2 in children. Pediatr. Infect. Dis. J. 2020;39:e133–e134. doi: 10.1097/INF.0000000000002704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drosten C., Seilmaier M., Corman V.M., Hartmann W., Scheible G., Sack S., Guggemos W., Kallies R., Muth D., Junglen S., Müller M.A., Haas W., Guberina H., Röhnisch T., Schmid-Wendtner M., Aldabbagh S., Dittmer U., Gold H., Graf P., Bonin F., Rambaut A., Wendtner C.-M. Clinical features and virological analysis of a case of Middle East respiratory syndrome coronavirus infection. Lancet Infect. Dis. 2013;13:745–751. doi: 10.1016/S1473-3099(13)70154-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Du L., He Y., Zhou Y., Liu S., Zheng B.J., Jiang S. The spike protein of SARS-CoV — a target for vaccine and therapeutic development. Nat. Rev. Microbiol. 2009;7:226–236. doi: 10.1038/nrmicro2090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Du W., Yu J., Liu X., Chen H., Lin L., Li Q. Persistence of SARS-CoV-2 virus RNA in feces: a case series of children. J. Infect. Publ. Health. 2020;13:926–931. doi: 10.1016/j.jiph.2020.05.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Espejo W., Celis J.E., Chiang G. Environment and Covid-19: pollutants, impacts, dissemination, management and recommendations for facing future epidemic threats. Sci. Total Environ. 2020;141314 doi: 10.1016/j.scitotenv.2020.141314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Farkas K., Hillary L.S., Malham S.K., McDonald J.E., Jones D.L. Wastewater and public health: the potential of wastewater surveillance for monitoring COVID-19. Curr. Opin. Environ. Sci. Heal. 2020;17:14–20. doi: 10.1016/j.coesh.2020.06.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Farkas K., Walker D.I., Adriaenssens E.M., McDonald J.E., Hillary L.S., Malham S.K., Jones D.L. Viral indicators for tracking domestic wastewater contamination in the aquatic environment. Water Res. 2020;181:115926. doi: 10.1016/j.watres.2020.115926. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fears A.C., Klimstra W.B., Duprex P., Hartman A., Weaver S.C., Plante K.S., Mirchandani D., Plante J.A., Aguilar P., V, Fernández D., Nalca A., Totura A., Dyer D., Kearney B., Lackemeyer M., Bohannon J.K., Johnson R., Garry R.F., Reed D.S., Roy C.J. Persistence of severe acute respiratory syndrome coronavirus 2 in aerosol suspensions. Emerg. Infect. Dis. 2020;26:2168–2171. doi: 10.3201/eid2609.201806. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feng B., Xu K., Gu S., Zheng S., Zou Q., Xu Y., Yu L., Lou F., Yu F., Jin T., Li Y., Sheng J., Yen H.-L., Zhong Z., Wei J., Chen Y. Multi-route transmission potential of SARS-CoV-2 in healthcare facilities. J. Hazard. Mater. 2021;402:123771. doi: 10.1016/j.jhazmat.2020.123771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feng W., Newbigging A.M., Le C., Pang B., Peng H., Cao Y., Wu J., Abbas G., Song J., Wang D., Cui M., Tyrrell D.L., Zhang X., Zhang H., Le X.C. Molecular diagnosis of COVID-19: challenges and research needs. Anal. Chem. 2020;92:10196–10209. doi: 10.1021/acs.analchem.0c02060. [DOI] [PubMed] [Google Scholar]
- Feng Y., Marchal T., Sperry T., Yi H. Influence of wind and relative humidity on the social distancing effectiveness to prevent COVID-19 airborne transmission: a numerical study. J. Aerosol Sci. 2020;147:105585. doi: 10.1016/j.jaerosci.2020.105585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Foladori P., Cutrupi F., Segata N., Manara S., Pinto F., Malpei F., Bruni L., La Rosa G. SARS-CoV-2 from faeces to wastewater treatment: what do we know? A review. Sci. Total Environ. 2020;743:140444. doi: 10.1016/j.scitotenv.2020.140444. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Franklin A.B., Bevins S.N. Spillover of SARS-CoV-2 into novel wild hosts in North America: a conceptual model for perpetuation of the pathogen. Sci. Total Environ. 2020;733:139358. doi: 10.1016/j.scitotenv.2020.139358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Frithiof R., Bergqvist A., Järhult J.D., Lipcsey M., Hultström M. Presence of SARS-CoV-2 in urine is rare and not associated with acute kidney injury in critically ill COVID-19 patients. Crit. Care. 2020;24:587. doi: 10.1186/s13054-020-03302-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gallardo-Escárate C., Valenzuela-Muñoz V., Núñez-Acuña G., Valenzuela-Miranda D., Benaventel B.P., Sáez-Vera C., Urrutia H., Novoa B., Figueras A., Roberts S., Assmann P., Bravo M. The wastewater microbiome: a novel insight for COVID-19 surveillance. Sci. Total Environ. 2020;142867 doi: 10.1016/j.scitotenv.2020.142867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gandhi M., Yokoe D.S., Havlir D., V Asymptomatic transmission, the Achilles’ heel of current strategies to control Covid-19. N. Engl. J. Med. 2020;382:2158–2160. doi: 10.1056/NEJMe2009758. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gomes F.M.D.S., Santo M.C.C.D.E., Gryschek R.C.B., Bertolozzi M.R., França F.O.D.S. Access to drinking water and sewage treatment in Brazil: a challenge for the control of waterborne infectious diseases. Rev. Inst. Med. Trop. Sao Paulo. 2020 doi: 10.1590/S1678-9946202062071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gómez M., Garralón G., Plaza F., Vílchez R., Hontoria E., Gómez M.A. Rejection of endocrine disrupting compounds (bisphenol A, bisphenol F and triethyleneglycol dimethacrylate) by membrane technologies. Desalination. 2007;212:79–91. doi: 10.1016/j.desal.2006.11.004. [DOI] [Google Scholar]
- Gonçalves J., Koritnik T., Mioč V., Trkov M., Bolješič M., Berginc N., Prosenc K., Kotar T., Paragi M. Detection of SARS-CoV-2 RNA in hospital wastewater from a low COVID-19 disease prevalence area. Sci. Total Environ. 2020;143226 doi: 10.1016/j.scitotenv.2020.143226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gonzalez R., Curtis K., Bivins A., Bibby K., Weir M.H., Yetka K., Thompson H., Keeling D., Mitchell J., Gonzalez D. COVID-19 surveillance in Southeastern Virginia using wastewater-based epidemiology. Water Res. 2020;186:116296. doi: 10.1016/j.watres.2020.116296. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gormley M., Aspray T.J., Kelly D.A. COVID-19: mitigating transmission via wastewater plumbing systems. Lancet Glob. Heal. 2020;8 doi: 10.1016/S2214-109X(20)30112-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gralinski L.E., Menachery V.D. Return of the coronavirus: 2019-nCoV. Viruses. 2020;12:135. doi: 10.3390/v12020135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Groß R., Conzelmann C., Müller J.A., Stenger S., Steinhart K., Kirchhoff F., Münch J. Detection of SARS-CoV-2 in human breastmilk. Lancet. 2020;395:1757–1758. doi: 10.1016/S0140-6736(20)31181-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gude V.G., Muire P.J. Preparing for outbreaks — implications for resilient water utility operations and services. Sustain. Cities Soc. 2021;64:102558. doi: 10.1016/j.scs.2020.102558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guerrero-Latorre L., Ballesteros I., Villacrés-Granda I., Granda M.G., Freire-Paspuel B., Ríos-Touma B. SARS-CoV-2 in river water: implications in low sanitation countries. Sci. Total Environ. 2020;743:140832. doi: 10.1016/j.scitotenv.2020.140832. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gundy P.M., Gerba C.P., Pepper I.L. Survival of coronaviruses in water and wastewater. Food Environ. Virol. 2009;1:10–14. doi: 10.1007/s12560-008-9001-6. [DOI] [Google Scholar]
- Han H., Luo Q., Mo F., Long L., Zheng W. SARS-CoV-2 RNA more readily detected in induced sputum than in throat swabs of convalescent COVID-19 patients. Lancet Infect. Dis. 2020;20:655–656. doi: 10.1016/S1473-3099(20)30174-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han J., He S. Urban flooding events pose risks of virus spread during the novel coronavirus (COVID-19) pandemic. Sci. Total Environ. 2021;755:142491. doi: 10.1016/j.scitotenv.2020.142491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han M.S., Seong M.-W., Heo E.Y., Park J.H., Kim N., Shin S., Cho S.I., Park S.S., Choi E.H. Sequential analysis of viral load in a neonate and her mother infected with severe acute respiratory syndrome coronavirus 2. Clin. Infect. Dis. 2020 doi: 10.1093/cid/ciaa447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han M.S., Seong M.-W., Kim N., Shin S., Cho S.I., Park H., Kim T.S., Park S.S., Choi E.H. Viral RNA load in mildly symptomatic and asymptomatic children with COVID-19, Seoul, South Korea. Emerg. Infect. Dis. J. 2020;26:2497–2499. doi: 10.3201/eid2610.202449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haramoto E., Malla B., Thakali O., Kitajima M. First environmental surveillance for the presence of SARS-CoV-2 RNA in wastewater and river water in Japan. Sci. Total Environ. 2020;737:140405. doi: 10.1016/j.scitotenv.2020.140405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harries A.D., dar Berger S., Satyanarayana S., Thekkur P., Kumar A.M., V Testing wastewater to detect severe acute respiratory syndrome coronavirus 2 in communities. Trans. R. Soc. Trop. Med. Hyg. 2020;114:782–783. doi: 10.1093/trstmh/traa066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hart O.E., Halden R.U. Computational analysis of SARS-CoV-2/COVID-19 surveillance by wastewater-based epidemiology locally and globally: feasibility, economy, opportunities and challenges. Sci. Total Environ. 2020;730:138875. doi: 10.1016/j.scitotenv.2020.138875. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hartenian E., Nandakumar D., Lari A., Ly M., Tucker J.M., Glaunsinger B.A. The molecular virology of coronaviruses. J. Biol. Chem. 2020;295:12910–12934. doi: 10.1074/jbc.REV120.013930. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hasan S.W., Ibrahim Y., Daou M., Kannout H., Jan N., Lopes A., Alsafar H., Yousef A.F. Detection and quantification of SARS-CoV-2 RNA in wastewater and treated effluents: surveillance of COVID-19 epidemic in the United Arab Emirates. Sci. Total Environ. 2020;142929 doi: 10.1016/j.scitotenv.2020.142929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- He Y., Luo J., Yang J., Song J., Wei L., Ma W. Value of viral nucleic acid in sputum and feces and specific IgM/IgG in serum for the diagnosis of Coronavirus Disease 2019. Front. Cell. Infect. Microbiol. 2020;10 doi: 10.3389/fcimb.2020.00445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heller L., Mota C.R., Greco D.B. COVID-19 faecal-oral transmission: are we asking the right questions? Sci. Total Environ. 2020;729:138919. doi: 10.1016/j.scitotenv.2020.138919. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heymann D.L., Shindo N. COVID-19: what is next for public health? Lancet. 2020;395:542–545. doi: 10.1016/S0140-6736(20)30374-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hindson J. COVID-19: faecal–oral transmission? Nat. Rev. Gastroenterol. Hepatol. 2020;17:259. doi: 10.1038/s41575-020-0295-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horn S., Vogt B., Pieters R., Bouwman H., Bezuidenhout C. Impact of potential COVID-19 treatment on South African water sources already threatened by pharmaceutical pollution. Environ. Toxicol. Chem. 2020;39:1305–1306. doi: 10.1002/etc.4734. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang Yongbo, Chen S., Yang Z., Guan W., Liu D., Lin Z., Zhang Y., Xu Z., Liu X., Li Y. SARS-CoV-2 viral load in clinical samples from critically ill patients. Am. J. Respir. Crit. Care Med. 2020;201:1435–1438. doi: 10.1164/rccm.202003-0572LE. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang Yuan, Yang C., Xu X., Xu W., Liu S. Structural and functional properties of SARS-CoV-2 spike protein: potential antivirus drug development for COVID-19. Acta Pharmacol. Sin. 2020;41:1141–1149. doi: 10.1038/s41401-020-0485-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jeong H.W., Kim S.-M., Kim H.-S., Kim Y.-I., Kim J.H., Cho J.Y., Kim S., Kang H., Kim S.-G., Park S.-J., Kim E.-H., Choi Y.K. Viable SARS-CoV-2 in various specimens from COVID-19 patients. Clin. Microbiol. Infect. 2020 doi: 10.1016/j.cmi.2020.07.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaplan E.H., Wang D., Wang M., Malik A.A., Zulli A., Peccia J. Aligning SARS-CoV-2 indicators via an epidemic model: application to hospital admissions and RNA detection in sewage sludge. Health Care Manag. Sci. 2020 doi: 10.1007/s10729-020-09525-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kassem I.I., Jaafar H. The potential impact of water quality on the spread and control of COVID-19 in Syrian refugee camps in Lebanon. Water Int. 2020;45:423–429. doi: 10.1080/02508060.2020.1780042. [DOI] [Google Scholar]
- Kim J.-M., Kim H.M., Lee E.J., Jo H.J., Yoon Y., Lee N.-J., Son J., Lee Y.-J., Kim M.S., Lee Y.-P., Chae S.-J., Park K.R., Cho S.-R., Park S., Kim S.J., Wang E., Woo S., Lim A., Park S.-J., Jang J., Chung Y.-S., Chin B.S., Lee J.-S., Lim D., Han M.-G., Yoo C.K. Detection and isolation of SARS-CoV-2 in serum, urine, and stool specimens of COVID-19 patients from the Republic of Korea. Osong Publ. Heal. Res Perspect. 2020;11:112–117. doi: 10.24171/j.phrp.2020.11.3.02. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kitajima M., Ahmed W., Bibby K., Carducci A., Gerba C.P., Hamilton K.A., Haramoto E., Rose J.B. SARS-CoV-2 in wastewater: state of the knowledge and research needs. Sci. Total Environ. 2020;739:139076. doi: 10.1016/j.scitotenv.2020.139076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krafcikova P., Silhan J., Nencka R., Boura E. Structural analysis of the SARS-CoV-2 methyltransferase complex involved in RNA cap creation bound to sinefungin. Nat. Commun. 2020;1–7 doi: 10.1038/s41467-020-17495-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kujawski S.A., Wong K.K., Collins J.P., Epstein L., Killerby M.E., Midgley C.M., Abedi G.R., Ahmed N.S., Almendares O., Alvarez F.N., Anderson K.N., Balter S., Barry V., Bartlett K., Beer K., Ben-Aderet M.A., Benowitz I., Biggs H.M., Binder A.M., Black S.R., Bonin B., Bozio C.H., Brown C.M., Bruce H., Bryant-Genevier J., Budd A., Buell D., Bystritsky R., Cates J., Charles E.M., Chatham-Stephens K., Chea N., Chiou H., Christiansen D., Chu V., Cody S., Cohen M., Conners E.E., Curns A.T., Dasari V., Dawson P., DeSalvo T., Diaz G., Donahue M., Donovan S., Duca L.M., Erickson K., Esona M.D., Evans S., Falk J., Feldstein L.R., Fenstersheib M., Fischer M., Fisher R., Foo C., Fricchione M.J., Friedman O., Fry A., Galang R.R., Garcia M.M., Gerber S.I., Gerrard G., Ghinai I., Gounder P., Grein J., Grigg C., Gunzenhauser J.D., Gutkin G.I., Haddix M., Hall A.J., Han G.S., Harcourt J., Harriman K., Haupt T., Haynes A.K., Holshue M., Hoover C., Hunter J.C., Jacobs M.W., Jarashow C., Joshi K., Kamali T., Kamili S., Kim L., Kim M., King J., Kirking H.L., Kita-Yarbro A., Klos R., Kobayashi M., Kocharian A., Komatsu K.K., Koppaka R., Layden J.E., Li Y., Lindquist S., Lindstrom S., Link-Gelles R., Lively J., Livingston M., Lo K., Lo J., Lu X., Lynch B., Madoff L., Malapati L., Marks G., Marlow M., Mathisen G.E., McClung N., McGovern O., McPherson T.D., Mehta M., Meier A., Mello L., Moon S., Morgan M., Moro R.N., Murray J., Murthy R., Novosad S., Oliver S.E., O’Shea J., Pacilli M., Paden C.R., Pallansch M.A., Patel M., Patel S., Pedraza I., Pillai S.K., Pindyck T., Pray I., Queen K., Quick N., Reese H., Reporter R., Rha B., Rhodes H., Robinson S., Robinson P., Rolfes M.A., Routh J.A., Rubin R., Rudman S.L., Sakthivel S.K., Scott S., Shepherd C., Shetty V., Smith E.A., Smith S., Stierman B., Stoecker W., Sunenshine R., Sy-Santos R., Tamin A., Tao Y., Terashita D., Thornburg N.J., Tong S., Traub E., Tural A., Uehara A., Uyeki T.M., Vahey G., Verani J.R., Villarino E., Wallace M., Wang L., Watson J.T., Westercamp M., Whitaker B., Wilkerson S., Woodruff R.C., Wortham J.M., Wu T., Xie A., Yousaf A., Zahn M., Zhang J., Covid-19 Investigation Team Clinical and virologic characteristics of the first 12 patients with coronavirus disease 2019 (COVID-19) in the United States. Nat. Med. 2020;26:861–868. doi: 10.1038/s41591-020-0877-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kumar M., Patel A.K., Shah A., V, Raval J., Rajpara N., Joshi M., Joshi C.G. First proof of the capability of wastewater surveillance for COVID-19 in India through detection of genetic material of SARS-CoV-2. Sci. Total Environ. 2020;746:141326. doi: 10.1016/j.scitotenv.2020.141326. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kumar M., Kuroda K., Patel A.K., Patel N., Bhattacharya P., Joshi M., Joshi C.G. Decay of SARS-CoV-2 RNA along the wastewater treatment outfitted with Upflow Anaerobic Sludge Blanket (UASB) system evaluated through two sample concentration techniques. Sci. Total Environ. 2021;754:142329. doi: 10.1016/j.scitotenv.2020.142329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kumar V., Singh S.B., Singh S. COVID-19: environment concern and impact of Indian medicinal system. J. Environ. Chem. Eng. 2020;8:104144. doi: 10.1016/j.jece.2020.104144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- La Rosa G., Bonadonna L., Lucentini L., Kenmoe S., Suffredini E. Coronavirus in water environments: occurrence, persistence and concentration methods — a scoping review. Water Res. 2020;179:115899. doi: 10.1016/j.watres.2020.115899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- La Rosa G., Iaconelli M., Mancini P., Bonanno Ferraro G., Veneri C., Bonadonna L., Lucentini L., Suffredini E. First detection of SARS-CoV-2 in untreated wastewaters in Italy. Sci. Total Environ. 2020;736:139652. doi: 10.1016/j.scitotenv.2020.139652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- La Rosa G., Mancini P., Bonanno Ferraro G., Veneri C., Iaconelli M., Bonadonna L., Lucentini L., Suffredini E. SARS-CoV-2 has been circulating in northern Italy since December 2019: evidence from environmental monitoring. Sci. Total Environ. 2021;750:141711. doi: 10.1016/j.scitotenv.2020.141711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lai M.Y.Y., Cheng P.K.C., Lim W.W.L. Survival of severe acute respiratory syndrome coronavirus. Clin. Infect. Dis. 2005;41:e67–e71. doi: 10.1086/433186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Larsen D.A., Wigginton K.R. Tracking COVID-19 with wastewater. Nat. Biotechnol. 2020;38:1151–1153. doi: 10.1038/s41587-020-0690-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lau S.K.P., Che X.-Y., Woo P.C.Y., Wong B.H.L., Cheng V.C.C., Woo G.K.S., Hung I.F.N., Poon R.W.S., Chan K.-H., Peiris J.S.M., Yuen K.-Y. SARS coronavirus detection methods. Emerg. Infect. Dis. 2005;11:1108–1111. doi: 10.3201/eid1107.041045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lednicky J.A., Lauzardo M., Fan Z.H., Jutla A., Tilly T.B., Gangwar M., Usmani M., Shankar S.N., Mohamed K., Eiguren-Fernandez A., Stephenson C.J., Alam M.M., Elbadry M.A., Loeb J.C., Subramaniam K., Waltzek T.B., Cherabuddi K., Morris J.G., Jr., Wu C.-Y. Viable SARS-CoV-2 in the air of a hospital room with COVID-19 patients. Int. J. Infect. Dis. 2020;100:476–482. doi: 10.1016/j.ijid.2020.09.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee Y.J., Kim J.H., Choi B.-S., Choi J.-H., Jeong Y.-I. Characterization of Severe Acute Respiratory Syndrome Coronavirus 2 stability in multiple water matrices. J. Korean Med. Sci. 2020;35 doi: 10.3346/jkms.2020.35.e330. [DOI] [PubMed] [Google Scholar]
- Lescure F.-X., Bouadma L., Nguyen D., Parisey M., Wicky P.-H., Behillil S., Gaymard A., Bouscambert-Duchamp M., Donati F., Le Hingrat Q., Enouf V., Houhou-Fidouh N., Valette M., Mailles A., Lucet J.-C., Mentre F., Duval X., Descamps D., Malvy D., Timsit J.-F., Lina B., Van-der-Werf S., Yazdanpanah Y. Clinical and virological data of the first cases of COVID-19 in Europe: a case series. Lancet Infect. Dis. 2020;20:697–706. doi: 10.1016/S1473-3099(20)30200-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lesimple A., Jasim S.Y., Johnson D.J., Hilal N. The role of wastewater treatment plants as tools for SARS-CoV-2 early detection and removal. J. Water Proc. Eng. 2020;38:101544. doi: 10.1016/j.jwpe.2020.101544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li D., Jin M., Bao P., Zhao W., Zhang S. Clinical characteristics and results of semen tests among men with Coronavirus Disease 2019. JAMA Netw. Open. 2020;3 doi: 10.1001/jamanetworkopen.2020.8292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Y., Hu Y., Yu Y., Zhang X., Li B., Wu J., Li J., Wu Y., Xia X., Tang H., Xu J. Positive result of Sars-Cov-2 in faeces and sputum from discharged patients with COVID-19 in Yiwu, China. J. Med. Virol. 2020;92:1938–1947. doi: 10.1002/jmv.25905. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ling Y., Xu S.-B., Lin Y.-X., Tian D., Zhu Z.-Q., Dai F.-H., Wu F., Song Z.-G., Huang W., Chen J., Hu B.-J., Wang S., Mao E.-Q., Zhu L., Zhang W.-H., Lu H.-Z. Persistence and clearance of viral RNA in 2019 novel coronavirus disease rehabilitation patients. Chin. Med. J. 2020;133:1039–1043. doi: 10.1097/CM9.0000000000000774. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu D., Thompson J.R., Carducci A., Bi X. Potential secondary transmission of SARS-CoV-2 via wastewater. Sci. Total Environ. 2020;749:142358. doi: 10.1016/j.scitotenv.2020.142358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu L. Emerging study on the transmission of the Novel Coronavirus (COVID-19) from urban perspective: evidence from China. Cities. 2020;103:102759. doi: 10.1016/j.cities.2020.102759. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu L., Hu J., Hou Y., Tao Z., Chen Z., Chen K. Pit latrines may be a potential risk in rural China and low-income countries when dealing with COVID-19. Sci. Total Environ. 2020;143283 doi: 10.1016/j.scitotenv.2020.143283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu P., Cai J., Jia R., Xia S., Wang X., Cao L., Zeng M., Xu J. Dynamic surveillance of SARS-CoV-2 shedding and neutralizing antibody in children with COVID-19. Emerg. Microb. Infect. 2020;9:1254–1258. doi: 10.1080/22221751.2020.1772677. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lo I.L., Lio C.F., Cheong H.H., Lei C.I., Cheong T.H., Zhong X., Tian Y., Sin N.N. Evaluation of SARS-CoV-2 RNA shedding in clinical specimens and clinical characteristics of 10 patients with COVID-19 in Macau. Int. J. Biol. Sci. 2020;16:1698–1707. doi: 10.7150/ijbs.45357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lodder W., de Roda Husman A.M. SARS-CoV-2 in wastewater: potential health risk, but also data source. Lancet Gastroenterol. Hepatol. 2020;5:533–534. doi: 10.1016/S2468-1253(20)30087-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu D., Huang Z., Luo J., Zhang X., Sha S. Primary concentration — the critical step in implementing the wastewater based epidemiology for the COVID-19 pandemic: a mini-review. Sci. Total Environ. 2020;141245 doi: 10.1016/j.scitotenv.2020.141245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maal-Bared R., Brisolara K., Munakata N., Bibby K., Gerba C., Sobsey M., Schaefer S., Swift J., Gary L., Sherchan S., Babatola A., Bastian R., Olabode L., Reimers R., Rubin A. Implications of SARS-CoV-2 on current and future operation and management of wastewater systems. Water Environ. Res. 2020 doi: 10.1002/wer.1446. n/a. [DOI] [PubMed] [Google Scholar]
- Malenovská H. Coronavirus persistence on a plastic carrier under refrigeration conditions and its reduction using wet wiping technique, with respect to food safety. Food Environ. Virol. 2020;12:361–366. doi: 10.1007/s12560-020-09447-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin J., Klapsa D., Wilton T., Zambon M., Bentley E., Bujaki E., Fritzsche M., Mate R., Majumdar M. Tracking SARS-CoV-2 in sewage: evidence of changes in virus variant predominance during COVID-19 pandemic. Viruses. 2020;12:1144. doi: 10.3390/v12101144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mathavarajah S., Stoddart A.K., Gagnon G.A., Dellaire G. Pandemic danger to the deep: the risk of marine mammals contracting SARS-CoV-2 from wastewater. Sci. Total Environ. 2020;143346 doi: 10.1016/j.scitotenv.2020.143346. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Medema G., Heijnen L., Elsinga G., Italiaander R., Brouwer A. Presence of SARS-Coronavirus-2 RNA in sewage and correlation with reported COVID-19 prevalence in the early stage of the epidemic in the Netherlands. Environ. Sci. Technol. Lett. 2020;7:511–516. doi: 10.1021/acs.estlett.0c00357. [DOI] [PubMed] [Google Scholar]
- Meng X., Huang X., Zhou P., Li C., Wu A. Alert for SARS-CoV-2 infection caused by fecal aerosols in rural areas in China. Infect. Control Hosp. Epidemiol. 2020;41:987. doi: 10.1017/ice.2020.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mesoraca A., Margiotti K., Viola A., Cima A., Sparacino D., Giorlandino C. Evaluation of SARS-CoV-2 viral RNA in fecal samples. Virol. J. 2020;17:86. doi: 10.1186/s12985-020-01359-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Michael-Kordatou I., Karaolia P., Fatta-Kassinos D. Sewage analysis as a tool for the COVID-19 pandemic response and management: the urgent need for optimised protocols for SARS-CoV-2 detection and quantification. J. Environ. Chem. Eng. 2020;116544 doi: 10.1016/j.jns.2019.116544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller M., Loaiza J., Takyar A., Gilman R. COVID-19 in Latin America: novel transmission dynamics for a global pandemic? PLoS Negl. Trop. Dis. 2020;14 doi: 10.1371/journal.pntd.0008265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mlejnkova H., Sovova K., Vasickova P., Ocenaskova V., Jasikova L., Juranova E. Preliminary study of SARS-CoV-2 occurrence in wastewater in the Czech Republic. Int. J. Environ. Res. Public Health. 2020;17:5508. doi: 10.3390/ijerph17155508. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mohseni A.H., Taghinezhad-S S., Xu Z., Fu X. Body fluids may contribute to human-to-human transmission of severe acute respiratory syndrome coronavirus 2: evidence and practical experience. Chin. Med. 2020;15:58. doi: 10.1186/s13020-020-00337-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Montagna M.T., Diella G., De Giglio O., Triggiano F., Carlucci M., Carpagnano F.L., Caggiano G. Can beaches and bathing environments represent a risk of spreading COVID-19? Ann. Ig. 2020;32:593–596. doi: 10.7416/ai.2020.2379. [DOI] [PubMed] [Google Scholar]
- Nabi G., Khan S. Risk of COVID-19 pneumonia in aquatic mammals. Environ. Res. 2020;188:109732. doi: 10.1016/j.envres.2020.109732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nabi G., Siddique R., Khan S. Detecting viral outbreaks in future using enhanced environmental surveillance. Environ. Res. 2020;188:109731. doi: 10.1016/j.envres.2020.109731. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Naddeo V., Liu H. Editorial perspectives: 2019 novel coronavirus (SARS-CoV-2): what is its fate in urban water cycle and how can the water research community respond? Environ. Sci. Water Res. Technol. 2020;6:1213–1216. doi: 10.1039/D0EW90015J. [DOI] [Google Scholar]
- Nemudryi A., Nemudraia A., Surya K., Wiegand T., Buyukyoruk M., Wilkinson R., Wiedenheft B. Temporal detection and phylogenetic assessment of SARS-CoV-2 in municipal wastewater. Cell Rep. Med. 2020;1:100098. doi: 10.1016/j.xcrm.2020.100098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Netz R.R., Eaton W.A. Physics of virus transmission by speaking droplets. Proc. Natl. Acad. Sci. 2020;117:25209–25211. doi: 10.1073/pnas.2011889117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nghiem L.D., Morgan B., Donner E., Short M.D. The COVID-19 pandemic: considerations for the waste and wastewater services sector. Case Stud. Chem. Environ. Eng. 2020;1:100006. doi: 10.1016/j.cscee.2020.100006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nomoto H., Ishikane M., Katagiri D., Kinoshita N., Nagashima M., Sadamasu K., Yoshimura K., Ohmagari N. Cautious handling of urine from moderate to severe COVID-19 patients. Am. J. Infect. Control. 2020;48:969–971. doi: 10.1016/j.ajic.2020.05.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Odih E.E., Afolayan A.O., Akintayo I., Okeke I.N. Could water and sanitation shortfalls exacerbate SARS-CoV-2 transmission risks? Am. J. Trop. Med. Hyg. 2020;103:554–557. doi: 10.4269/ajtmh.20-0462. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olusola-Makinde O.O., Reuben R.C. Ticking bomb: prolonged faecal shedding of novel coronavirus (2019-nCoV) and environmental implications. Environ. Pollut. 2020;267:115485. doi: 10.1016/j.envpol.2020.115485. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O’Reilly K.M., Allen D.J., Fine P., Asghar H. Comment the challenges of informative wastewater sampling for SARS-CoV-2 must be met: lessons from polio eradication. Lancet Microbe. 2020;2:1–2. doi: 10.1016/S2666-5247(20)30100-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ortiz-Prado E., Simbaña-Rivera K., Gómez-Barreno L., Rubio-Neira M., Guaman L.P., Kyriakidis N.C., Muslin C., Jaramillo A.M.G., Barba-Ostria C., Cevallos-Robalino D., Sanches-SanMiguel H., Unigarro L., Zalakeviciute R., Gadian N., López-Cortés A. Clinical, molecular, and epidemiological characterization of the SARS-CoV-2 virus and the Coronavirus Disease 2019 (COVID-19), a comprehensive literature review. Diagn. Microbiol. Infect. Dis. 2020;98:115094. doi: 10.1016/j.diagmicrobio.2020.115094. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pan Y., Zhang D., Yang P., Poon L.L.M., Wang Q. Viral load of SARS-CoV-2 in clinical samples. Lancet Infect. Dis. 2020;20:411–412. doi: 10.1016/S1473-3099(20)30113-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pastorino B., Touret F., Gilles M., de Lamballerie X., Charrel R. Prolonged infectivity of SARS-CoV-2 in fomites. Emerg. Infect. Dis. J. 2020;26:2256–2257. doi: 10.3201/eid2609.201788. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Patel J. Faecal shedding of SARS-CoV-2: considerations for hospital settings. J. Hosp. Infect. 2020;1253:19–20. doi: 10.1016/j.jhin.2020.05.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Patrì A., Gallo L., Guarino M., Fabbrocini G. Sexual transmission of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2): a new possible route of infection? J. Am. Acad. Dermatol. 2020;82:e227. doi: 10.1016/j.jaad.2020.03.098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peccia J., Zulli A., Brackney D.E., Grubaugh N.D., Kaplan E.H., Casanovas-Massana A., Ko A.I., Malik A.A., Wang D., Wang M., Warren J.L., Weinberger D.M., Arnold W., Omer S.B. Measurement of SARS-CoV-2 RNA in wastewater tracks community infection dynamics. Nat. Biotechnol. 2020;38:1164–1167. doi: 10.1038/s41587-020-0684-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pecson B., Gerrity D., Bibby K., Drewes J.E., Gerba C., Gersberg R., Gonzalez R., Haas C.N., Hamilton K.A., Nelson K.L., Olivieri A., Rock C., Rose J., Sobsey M. Editorial perspectives: will SARS-CoV-2 reset public health requirements in the water industry? Integrating lessons of the past and emerging research. Environ. Sci. Water Res. Technol. 2020;6:1761–1764. doi: 10.1039/D0EW90031A. [DOI] [Google Scholar]
- Peng L., Liu J., Xu W., Luo Q., Chen D., Lei Z., Huang Z., Li X., Deng K., Lin B., Gao Z. SARS-CoV-2 can be detected in urine, blood, anal swabs, and oropharyngeal swabs specimens. J. Med. Virol. 2020;92:1676–1680. doi: 10.1002/jmv.25936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Polo D., Quintela-Baluja M., Corbishley A., Jones D.L., Singer A.C., Graham D.W., Romalde J.L. Making waves: wastewater-based epidemiology for COVID-19 — approaches and challenges for surveillance and prediction. Water Res. 2020;186:116404. doi: 10.1016/j.watres.2020.116404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prado T., Fumian T.M., Mannarino C.F., Maranhão A.G., Siqueira M.M., Miagostovich M.P. Preliminary results of SARS-CoV-2 detection in sewerage system in Niterói municipality, Rio de Janeiro, Brazil. Mem. Inst. Oswaldo Cruz. 2020 doi: 10.1590/0074-02760200196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Premkumar L., Segovia-Chumbez B., Jadi R., Martinez D.R., Raut R., Markmann A., Cornaby C., Bartelt L., Weiss S., Park Y., Edwards C.E., Weimer E., Scherer E.M., Rouphael N., Edupuganti S., Weiskopf D., Tse L.V., Hou Y.J., Margolis D., Sette A., Collins M.H., Schmitz J., Baric R.S., de Silva A.M. The receptor binding domain of the viral spike protein is an immunodominant and highly specific target of antibodies in SARS-CoV-2 patients. Sci. Immunol. 2020;5:1–9. doi: 10.1126/sciimmunol.abc8413. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Race M., Ferraro A., Galdiero E., Guida M., Núñez-Delgado A., Pirozzi F., Siciliano A., Fabbricino M. Current emerging SARS-CoV-2 pandemic: potential direct/indirect negative impacts of virus persistence and related therapeutic drugs on the aquatic compartments. Environ. Res. 2020;188:109808. doi: 10.1016/j.envres.2020.109808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rafa N., Uddin S.M.N., Staddon C. Exploring challenges in safe water availability and accessibility in preventing COVID-19 in refugee settlements. Water Int. 2020:1–6. doi: 10.1080/02508060.2020.1803018. [DOI] [Google Scholar]
- Randazzo W., Cuevas-Ferrando E., Sanjuán R., Domingo-Calap P., Sánchez G. Metropolitan wastewater analysis for COVID-19 epidemiological surveillance. Int. J. Hyg. Environ. Health. 2020;230:113621. doi: 10.1016/j.ijheh.2020.113621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Randazzo W., Truchado P., Cuevas-Ferrando E., Simón P., Allende A., Sánchez G. SARS-CoV-2 RNA in wastewater anticipated COVID-19 occurrence in a low prevalence area. Water Res. 2020;181:1–8. doi: 10.1016/j.watres.2020.115942. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rimoldi S.G., Stefani F., Gigantiello A., Polesello S., Comandatore F., Mileto D., Maresca M., Longobardi C., Mancon A., Romeri F., Pagani C., Cappelli F., Roscioli C., Moja L., Gismondo M.R., Salerno F. Presence and infectivity of SARS-CoV-2 virus in wastewaters and rivers. Sci. Total Environ. 2020;744:140911. doi: 10.1016/j.scitotenv.2020.140911. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saguti F., Magnil E., Enache L., Churqui M.P., Johansson A., Lumley D., Davidsson F., Dotevall L., Mattsson A., Trybala E., Lagging M., Lindh M., Gisslén M., Brezicka T., Nyström K., Norder H. Surveillance of wastewater revealed peaks of SARS-CoV-2 preceding those of hospitalized patients with COVID-19. Water Res. 2021;189:116620. doi: 10.1016/j.watres.2020.116620. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Santarpia J.L., Rivera D.N., Herrera V.L., Morwitzer M.J., Creager H.M., Santarpia G.W., Crown K.K., Brett-Major D.M., Schnaubelt E.R., Broadhurst M.J., Lawler J., V, Reid S.P., Lowe J.J. Aerosol and surface contamination of SARS-CoV-2 observed in quarantine and isolation care. Sci. Rep. 2020;10:1–8. doi: 10.1038/s41598-020-69286-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmidt C. Watcher in the wastewater. Nat. Biotechnol. 2020 doi: 10.1038/s41587-020-0620-2. [DOI] [PubMed] [Google Scholar]
- Sherchan S.P., Shahin S., Ward L.M., Tandukar S., Aw T.G., Schmitz B., Ahmed W., Kitajima M. First detection of SARS-CoV-2 RNA in wastewater in North America: a study in Louisiana, USA. Sci. Total Environ. 2020;743:140621. doi: 10.1016/j.scitotenv.2020.140621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi K.-W., Huang Y.-H., Quon H., Ou-Yang Z.-L., Wang C., Jiang S.C. Quantifying the risk of indoor drainage system in multi-unit apartment building as a transmission route of SARS-CoV-2. Sci. Total Environ. 2020;143056 doi: 10.1016/j.scitotenv.2020.143056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Siddiqui R., Khamis M., Ibrahim T., Khan N.A. Irrigation system and COVID-19 recurrence: a potential risk factor in the transmission of SARS-CoV-2. ACS Chem. Neurosci. 2020;11:2903–2905. doi: 10.1021/acschemneuro.0c00570. [DOI] [PubMed] [Google Scholar]
- Sims N., Kasprzyk-Hordern B. Future perspectives of wastewater-based epidemiology: monitoring infectious disease spread and resistance to the community level. Environ. Int. 2020;139:105689. doi: 10.1016/j.envint.2020.105689. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith S.H., Somsen G.A., van Rijn C., Kooij S., van der Hoek L., Bem R.A., Bonn D. Aerosol persistence in relation to possible transmission of SARS-CoV-2. Phys. Fluids. 2020;32:107108. doi: 10.1063/5.0027844. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stepping K. Urban sewage in Brazil: drivers of and obstacles to wastewater treatment and reuse. Gov. Water-Energy-Food Nexus Ser. Discuss. Pap. 2016;50 [Google Scholar]
- Sternberg A., Naujokat C. Structural features of coronavirus SARS-CoV-2 spike protein: targets for vaccination. Life Sci. 2020;257:118056. doi: 10.1016/j.lfs.2020.118056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Street R., Malema S., Mahlangeni N., Mathee A. Wastewater surveillance for Covid-19: an African perspective. Sci. Total Environ. 2020;743:140719. doi: 10.1016/j.scitotenv.2020.140719. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun J., Zhu A., Li H., Zheng K., Zhuang Z., Chen Z., Shi Y., Zhang Z., Chen S., Liu X., Dai J., Li X., Huang S., Huang X., Luo L., Wen L., Zhuo J., Li Yuming, Wang Y., Zhang L., Zhang Y., Li F., Feng L., Chen X., Zhong N., Yang Z., Huang J., Zhao J., Li Yi-min. Isolation of infectious SARS-CoV-2 from urine of a COVID-19 patient. Emerg. Microb. Infect. 2020;9:991–993. doi: 10.1080/22221751.2020.1760144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tang A., Tong Z., Wang H., Dai Y., Li K., Liu J., Wu W., Yuan C., Yu M., Li P., Yan J. Detection of novel coronavirus by RT-PCR in stool specimen from asymptomatic child, China. Emerg. Infect. Dis. J. 2020;26:1337. doi: 10.3201/eid2606.200301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tang S., Mao Y., Jones R.M., Tan Q., Ji J.S., Li N., Shen J., Lv Y., Pan L., Ding P., Wang X., Wang Y., MacIntyre C.R., Shi X. Aerosol transmission of SARS-CoV-2? Evidence, prevention and control. Environ. Int. 2020;144:106039. doi: 10.1016/j.envint.2020.106039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thompson J.R., Nancharaiah Y., V, Gu X., Lee W.L., Rajal V.B., Haines M.B., Girones R., Ng L.C., Alm E.J., Wuertz S. Making waves: wastewater surveillance of SARS-CoV-2 for population-based health management. Water Res. 2020;184:116181. doi: 10.1016/j.watres.2020.116181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tong Y., Bao A., Chen H., Huang J., Lv Z., Feng L., Cheng Y., Wang Y., Bai L., Rao W., Zheng H., Wu Z., Qiao B., Zhao Z., Wang H., Li Y. Necessity for detection of SARS-CoV-2 RNA in multiple types of specimens for the discharge of the patients with COVID-19. J. Transl. Med. 2020;18:411. doi: 10.1186/s12967-020-02580-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Toyoshima Y., Nemoto K., Matsumoto S., Nakamura Y., Kiyotani K. SARS-CoV-2 genomic variations associated with mortality rate of COVID-19. J. Hum. Genet. 2020 doi: 10.1038/s10038-020-0808-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trottier J., Darques R., Ait Mouheb N., Partiot E., Bakhache W., Deffieu M.S., Gaudin R. Post-lockdown detection of SARS-CoV-2 RNA in the wastewater of Montpellier, France. One Heal. 2020;10:100157. doi: 10.1016/j.onehlt.2020.100157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Usman M., Farooq M., Hanna K. Environmental side effects of the injudicious use of antimicrobials in the era of COVID-19. Sci. Total Environ. 2020;745:141053. doi: 10.1016/j.scitotenv.2020.141053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vammen K., Guillen S.M. Water resources of Nicaragua and COVID-19: between panic and apathy? Braz. J. Biol. doi. 2020 doi: 10.1590/1519-6984.237891. [DOI] [PubMed] [Google Scholar]
- Venugopal A., Ganesan H., Raja S.S.S., Govindasamy V., Arunachalam M., Narayanasamy A., Sivaprakash P., Rahman P.K.S.M., Gopalakrishnan A.V., Siama Z., Vellingiri B. Novel wastewater surveillance strategy for early detection of coronavirus disease 2019 hotspots. Curr. Opin. Environ. Sci. Heal. 2020;17:8–13. doi: 10.1016/j.coesh.2020.05.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang J., Du G. COVID-19 may transmit through aerosol. Ir. J. Med. Sci. 2020 doi: 10.1007/s11845-020-02218-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang J., Shen J., Ye D., Yan X., Zhang Y., Yang W., Li X., Wang Junqi, Zhang L., Pan L. Disinfection technology of hospital wastes and wastewater: suggestions for disinfection strategy during Coronavirus Disease 2019 (COVID-19) pandemic in China. Environ. Pollut. 2020;262:114665. doi: 10.1016/j.envpol.2020.114665. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang W., Xu Y., Gao R., Lu R., Han K., Wu G., Tan W. Detection of SARS-CoV-2 in different types of clinical specimens. J. Am. Med. Assoc. 2020;323:1843–1844. doi: 10.1001/jama.2020.3786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang X., Zheng J., Guo L., Yao H., Wang L., Xia X., Zhang W. Fecal viral shedding in COVID-19 patients: clinical significance, viral load dynamics and survival analysis. Virus Res. 2020;289:198147. doi: 10.1016/j.virusres.2020.198147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang X.W., Li J., Guo T., Zhen B., Kong Q., Yi B., Li Z., Song N., Jin M., Xiao W., Zhu X., Gu C., Yin J., Wei W., Yao W., Liu C., Li J., Ou G., Wang M., Fang T., Wang G., Qiu Y., Wu H., Chao F., Li J. Concentration and detection of SARS coronavirus in sewage from Xiao Tang Shan hospital and the 309th Hospital of the Chinese People’s Liberation Army. Water Sci. Technol. 2005;52:213–221. doi: 10.2166/wst.2005.0266. [DOI] [PubMed] [Google Scholar]
- Wathore R., Gupta A., Bherwani H., Labhasetwar N. Understanding air and water borne transmission and survival of coronavirus: insights and way forward for SARS-CoV-2. Sci. Total Environ. 2020;749:141486. doi: 10.1016/j.scitotenv.2020.141486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wei X.-S., Wang X., Niu Y.-R., Ye L.-L., Peng W.-B., Wang Z.-H., Yang W.-B., Yang B.-H., Zhang J.-C., Ma W.-L., Wang X.-R., Zhou Q. Diarrhea is associated with prolonged symptoms and viral carriage in Corona Virus Disease 2019. Clin. Gastroenterol. Hepatol. 2020;18:1753–1759.e2. doi: 10.1016/j.cgh.2020.04.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Westhaus S., Weber F.-A., Schiwy S., Linnemann V., Brinkmann M., Widera M., Greve C., Janke A., Hollert H., Wintgens T., Ciesek S. Detection of SARS-CoV-2 in raw and treated wastewater in Germany — suitability for COVID-19 surveillance and potential transmission risks. Sci. Total Environ. 2021;751:141750. doi: 10.1016/j.scitotenv.2020.141750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wölfel R., Corman V.M., Guggemos W., Seilmaier M., Zange S., Müller M.A., Niemeyer D., Jones T.C., Vollmar P., Rothe C., Hoelscher M., Bleicker T., Brünink S., Schneider J., Ehmann R., Zwirglmaier K., Drosten C., Wendtner C. Virological assessment of hospitalized patients with COVID-2019. Nature. 2020;581:465–469. doi: 10.1038/s41586-020-2196-x. [DOI] [PubMed] [Google Scholar]
- World Health Organization – WHO Weekly operational update on COVID-19. 2021. https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situation-reports/ [WWW Document]. URL. accessed 2.1.21.
- Wu F., Xiao A., Zhang J., Gu X., Lee W.L., Kauffman K., Hanage W., Matus M., Ghaeli N., Endo N., Duvallet C., Moniz K., Erickson T., Chai P., Thompson J., Alm E. SARS-CoV-2 titers in wastewater are higher than expected from clinically confirmed cases. mSystems. 2020;5:e00614–e00620. doi: 10.1128/mSystems.00614-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu Y., Guo C., Tang L., Hong Z., Zhou J., Dong X., Yin H., Xiao Q., Tang Y., Qu X., Kuang L., Fang X., Mishra N., Lu J., Shan H., Jiang G., Huang X. Prolonged presence of SARS-CoV-2 viral RNA in faecal samples. Lancet Gastroenterol. Hepatol. 2020;5:434–435. doi: 10.1016/S2468-1253(20)30083-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wurtzer S., Marechal V., Mouchel J.-M., Maday Y., Teyssou R., Richard E., Almayrac J.L., Moulin L. Evaluation of lockdown effect on SARS-CoV-2 dynamics through viral genome quantification in waste water, Greater Paris, France, 5 March to 23 April 2020. Euro Surv. 2020;25:2000776. doi: 10.2807/1560-7917.ES.2020.25.50.2000776. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xia J., Tong J., Liu M., Shen Y., Guo D. Evaluation of coronavirus in tears and conjunctival secretions of patients with SARS-CoV-2 infection. J. Med. Virol. 2020;92:589–594. doi: 10.1002/jmv.25725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xiao F., Sun J., Xu Y., Li F., Huang X., Li H., Zhao Jingxian, Huang J., Zhao Jincun. Infectious SARS-CoV-2 in feces of patient with severe COVID-19. Emerg. Infect. Dis. 2020;26:1920–1922. doi: 10.3201/eid2608.200681. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xing Y., Ni W., Wu Q., Li W., Li G., Wang W., Tong J., Song X., Wong G.W., Xing Q. Prolonged viral shedding in feces of pediatric patients with coronavirus disease 2019. J. Microbiol. Immunol. Infect. 2020;53:473–480. doi: 10.1016/j.jmii.2020.03.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu D., Zhang Z., Jin L., Chu F., Mao Y., Wang H., Liu M., Wang M., Zhang L., Gao G.F., Wang F.S. Persistent shedding of viable SARS-CoV in urine and stool of SARS patients during the convalescent phase. Eur. J. Clin. Microbiol. Infect. Dis. 2005;24:165–171. doi: 10.1007/s10096-005-1299-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu X., Chen P., Wang J., Feng J., Zhou H., Li X., Zhong W., Hao P. Evolution of the novel coronavirus from the ongoing Wuhan outbreak and modeling of its spike protein for risk of human transmission. Sci. China Life Sci. 2020;63:457–460. doi: 10.1007/s11427-020-1637-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu Y., Li X., Zhu B., Liang H., Fang C., Gong Y., Guo Q., Sun X., Zhao D., Shen J., Zhang H., Liu H., Xia H., Tang J., Zhang K., Gong S. Characteristics of pediatric SARS-CoV-2 infection and potential evidence for persistent fecal viral shedding. Nat. Med. 2020;26:502–505. doi: 10.1038/s41591-020-0817-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu Z. Can the novel coronavirus be transmitted via RNAs without protein capsids? J. Infect. Dev. Ctries. 2020;14:1001–1003. doi: 10.3855/jidc.13880. [DOI] [PubMed] [Google Scholar]
- Yan R., Zhang Y., Li Y., Xia L., Guo Y., Zhou Q. Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science (80-. ) 2020;367:1444–1448. doi: 10.1126/science.abb2762. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang B., Li W., Wang J., Tian Z., Cheng X., Zhang Y., Qiu R., Hou S., Guo H. Estimation of the potential spread risk of COVID-19: occurrence assessment along the Yangtze, Han, and Fu River basins in Hubei, China. Sci. Total Environ. 2020;746:141353. doi: 10.1016/j.scitotenv.2020.141353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang W., Cai C., Dai X. The potential exposure and transmission risk of SARS-CoV-2 through sludge treatment and disposal. Resour. Conserv. Recycl. 2020;162:105043. doi: 10.1016/j.resconrec.2020.105043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ye Y., Ellenberg R.M., Graham K.E., Wigginton K.R. Survivability, partitioning, and recovery of enveloped viruses in untreated municipal wastewater. Environ. Sci. Technol. 2016;50:5077–5085. doi: 10.1021/acs.est.6b00876. [DOI] [PubMed] [Google Scholar]
- Yeo C., Kaushal S., Yeo D. Enteric involvement of coronaviruses: is faecal–oral transmission of SARS-CoV-2 possible? Lancet Gastroenterol. Hepatol. 2020;5:335–337. doi: 10.1016/S2468-1253(20)30048-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yoon J.G., Yoon J., Song J.Y., Yoon S.Y., Lim C.S., Seong H., Noh J.Y., Cheong H.J., Kim W.J. Clinical significance of a high SARS-CoV-2 viral load in the saliva. J. Korean Med. Sci. 2020;35 doi: 10.3346/jkms.2020.35.e195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu F., Yan L., Wang N., Yang S., Wang L., Tang Y., Gao G., Wang S., Ma C., Xie R., Wang F., Tan C., Zhu L., Guo Y., Zhang F. Quantitative detection and viral load analysis of SARS-CoV-2 in infected patients. Clin. Infect. Dis. 2020;71:793–798. doi: 10.1093/cid/ciaa345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yuan J., Chen Z., Gong C., Liu H., Li B., Li K., Chen X., Xu C., Jing Q., Liu G., Qin P., Liu Y., Zhong Y., Huang L., Zhu B.-P., Yang Z. Sewage as a possible transmission vehicle during a Coronavirus Disease 2019 outbreak in a densely populated community: Guangzhou, China, April 2020. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2020 doi: 10.1093/cid/ciaa1494. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zambrano-Monserrate M.A., Ruano M.A., Sanchez-Alcalde L. Indirect effects of COVID-19 on the environment. Sci. Total Environ. 2020;728:138813. doi: 10.1016/j.scitotenv.2020.138813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zaneti R.N., Girardi V., Spilki F.R., Mena K., Westphalen A.P.C., Colares E.R.D.C., Pozzebon A.G., Etchepare R.G. QMRA of SARS-CoV-2 for workers in wastewater treatment plants. Sci. Total Environ. 2021;754:142163. doi: 10.1016/j.scitotenv.2020.142163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang D., Ling H., Huang X., Li J., Li W., Yi C., Zhang T., Jiang Y. Potential spreading risks and disinfection challenges of medical wastewater by the presence of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) viral RNA in septic tanks of Fangcang Hospital. Sci. Total Environ. 2020;741:140445. doi: 10.1016/j.scitotenv.2020.140445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang H., Tang W., Chen Y., Yin W. Disinfection threatens aquatic ecosystems. Science (80-. ) 2020;368:146–147. doi: 10.1126/science.abb8905. [DOI] [PubMed] [Google Scholar]
- Zhang Y., Chen C., Zhu S., Shu C., Wang D., Song J., Song Y., Zhen W., Feng Z., Wu G., Xu J., Xu W. Isolation of 2019-nCoV from a stool specimen of a laboratory-confirmed case of the coronavirus disease 2019 (COVID-19) China CDC Wkly. 2020;2:123–124. doi: 10.46234/ccdcw2020.033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao F.-B., Tang C.-C., Liu X.-Y., Shi F.-J., Song X.-R., Tian Y., Li Z.-S. Transportation characteristics of bisphenol A on ultrafiltration membrane with low molecule weight cut-off. Desalination. 2015;362:18–25. doi: 10.1016/j.desal.2015.01.048. [DOI] [Google Scholar]
- Zheng S., Fan J., Yu F., Feng B., Lou B., Zou Q., Xie G., Lin S., Wang R., Yang X., Chen W., Wang Q., Zhang D., Liu Y., Gong R., Ma Z., Lu S., Xiao Y., Gu Y., Zhang J., Yao H., Xu K., Lu X., Wei G., Zhou J., Fang Q., Cai H., Qiu Y., Sheng J., Chen Y., Liang T. Viral load dynamics and disease severity in patients infected with SARS-CoV-2 in Zhejiang province, China, January–March 2020: retrospective cohort study. BMJ. 2020;369 doi: 10.1136/bmj.m1443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu N., Zhang D., Wang W., Li X., Yang B., Song J., Zhao X., Huang B., Shi W., Lu R., Niu P., Zhan F., Ma X., Wang D., Xu W., Wu G., Gao G.F., Tan W. A novel coronavirus from patients with pneumonia in China, 2019. N. Engl. J. Med. 2020;382:727–733. doi: 10.1056/NEJMoa2001017. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Bibliographic search results in Pub Med, SCOPUS, and ScienceDirect databases.