Abstract
Bats are incredibly diverse, both morphologically and taxonomically. Bats are the only mammalian group to have achieved powered flight, an adaptation that is hypothesized to have allowed them to colonize various and diverse ecological niches. However, the lack of fossils capturing the transition from terrestrial mammal to volant chiropteran has obscured much of our understanding of bat evolution. Over the last 20 years, the emergence of evo-devo in non-model species has started to fill this gap by uncovering some developmental mechanisms at the origin of bat diversification. In this review, we highlight key aspects of studies that have used bats as a model for morphological adaptations, diversification during adaptive radiations, and morphological novelty. To do so, we review current and ongoing studies on bat evolution. We first investigate morphological specialization by reviewing current knowledge about wing and face evolution. Then, we explore the mechanisms behind adaptive diversification in various ecological contexts using vision and dentition. Finally, we highlight the emerging work into morphological novelties using bat wing membranes.
Keywords: Chiroptera, wing, craniofacial, sensory adaptations, teeth
Introduction: Diversity and phylogeny of bats
Following the acquisition of powered flight, bats (Order Chiroptera) have diversified into ~1411 species that comprise ~20% of all mammals (Gunnell and Simmons, 2005; Burgin et al., 2018). While chiropteran diversity is highest in the tropics (Shi et al., 2018), bats are ubiquitous across the globe, with species inhabiting every continent except for Antarctica. Within this broad geographic distribution, bats occupy a wide array of ecological niches, with diets spanning insectivory, frugivory, nectarivory, piscivory, sanguinivory, and carnivory. This diverse range of feeding types makes bats important members of many ecosystems, where they act as pollinators, predators, and key regulators of insect populations (Hill and Smith, 1984). Bats’ diversity can also be observed in their unique morphologies and novel features; bats display greatly elongated digits, uniquely pliable bone structure, and high variance in wing membrane shape and craniofacial structure, among other unique characteristics (Hill and Smith, 1984; Dumont, 2007; Cooper and Sears, 2013). Bats also display a wide range of life spans (Wilkinson and South 2002) with at least four bat lineages exhibiting “extreme longevity” (Wilkinson and Adams, 2019) and unique immune systems (Jebb et al., 2020). The wide array of specialized niches, morphologies and life spans that characterize bats makes them an excellent study system to investigate links between development and functional morphology, ecomorphology, adaptive radiations, and macroevolution as well as senescence, aging and the immune system.
Traditionally, bats have been classified as Microchiroptera (microbats) and Megachiroptera (megabats), but molecular evidence has revealed that microbats are not a reciprocally monophyletic clade (Teeling et al., 2005). Bats have therefore been reclassified as Yinpterochiroptera and Yangochiroptera, resolving the paraphyly of Microchiroptera (Teeling et al., 2005) (Figure 1a). Yinpterochiroptera consist of the lineage previously classified as megabats (Pteropodidae) and the microbat group Rhinolophidae, while Yangochiroptera consist of the remaining three microbat lineages Emballonuroidea, Noctilionoidea, and Vespertilonoidea (Teeling et al., 2005).
Figure 1 -. Phylogeny of bats. (a) Cladogram* showing intraordinal phylogenetic relationships in Chiroptera (adapted from Teeling et al., 2018). Based on molecular data, Chiroptera is now classified into Yinpterochiroptera (orange) and Yangochiroptera (magenta), while the traditional classification into microbats and megabats is shown in grey. (b) Cladogram* showing the interordinal relationship between Chiroptera and other mammalian taxa (adapted from Jebb et al., 2020). Three of the best supported tree topologies for the relationship between the Laurasiatherian lineages Carnivora, Perrissodactyla, Cetartiodactyla, Pholidota and Chiroptera (indicated by shaded area) are shown. *Branch lengths do not indicate distance.
In relation to other mammals, molecular phylogenetic studies place bats within Laurasiatheria (Tsagkogeorga et al., 2013) as sister to Cetartiodactyla, Pholidota, Carnivora and Perissodactyla, although the interordinal relationships between these sister clades remains unresolved (Jebb et al., 2020) (Figure 1b). The ancestral relationship to Eulipotyphla (moles and shrews) as the basal clade within Laurasiatheria (Tsagkogeorga et al., 2013; Jebb et al., 2020), combined with the (limited) fossil data suggests that the ancestor to bats might have been a small, quadrupedal, insectivorous mammal with pawed limbs (Gunnell and Simmons, 2005). However, the patterns and processes of the evolutionary transition of this hypothetical ancestor into a bat remain speculative, in large part because this transition is largely uncaptured by the fossil record. Indeed, bat fossil records have the lowest skeletal completeness metrics among all tetrapods analyzed to date, with most fossils being isolated bat teeth (Brown et al., 2019). The oldest known bat fossils date from the early Eocene, approximately 52.5 million years ago; these fossils already bear remarkable morphological similarities with extant bats and when alive the animals were likely capable of powered flight (Teeling et al., 2000; Gunnell and Simmons, 2005; Simmons et al., 2008). Due to this dearth of transitional paleontological information, many researchers have begun utilizing studies of bat development and phylogenetic relationships to better understand the origin of bats and how they came to occupy such a diverse array of niches. Examining the developmental processes that generate the tremendous amount of variation in this clade has the potential to provide insights into how macroevolutionary transitions occur and what processes facilitate the origins of variation. In this article, we explore some of the insights into the processes of morphological specialization of homologous structures, morphological diversification during adaptive radiations, and the origins of morphological novelty that research in bats have provided.
Morphological specialization of homologous structures
Much of the embryological research to date using bats as model organisms has investigated the role of developmental changes in the morphological specialization of homologous structures, i.e., structures derived from a common ancestral structure. Here we discuss past and ongoing research on the developmental basis of the morphological specialization of the bat wing skeleton for flight, and of the diversification of bat cranial structures for many diets and behaviors.
Bat wings: Extended outgrowth and forelimb lengthening
Given that bats are the only mammals capable of powered flight, it is perhaps not surprising that one of their most striking and recognizable features is their wings. The earliest bats preserved in the fossil record possess the defining characteristics of the bat wing: an overall enlarged forelimb, flight membranes, and elongated skeletal elements (Gunnell and Simmons, 2005; Simmons et al., 2008). In fact, morphometric analysis of the forelimb elements of fossil and extant bats suggests that the relative length of bat forelimbs and digits has not significantly changed in 50 million years (Sears et al., 2006). With clues to the origin of chiropteran wing structures currently lacking in the fossil record, some studies have turned to developmental mechanisms to try to understand this transition, often using comparisons of bat and mouse limb, the latter being a model for limb development (Cretekos et al., 2008; Adams, 2008).
At the time of the initial cartilaginous condensation of the digits, the future skeletal elements in bat and mice forelimb are similar in size (Sears et al., 2006; Hockman et al., 2009). However, rates of chondrocyte proliferation and differentiation soon increase notably in bat long bones, and the long bones increase in relative length (Sears et al., 2006; Farnum et al., 2008). The result is that the final relative size of the bat forelimb, including the digits, is larger than that of ancestral, terrestrial mammals. Studies of the allometry of bat bones support the hypothesis that differences in the growth rates of bat long bones help drive differences in long bone proportions among bat species (López‐Aguirre et al., 2019).
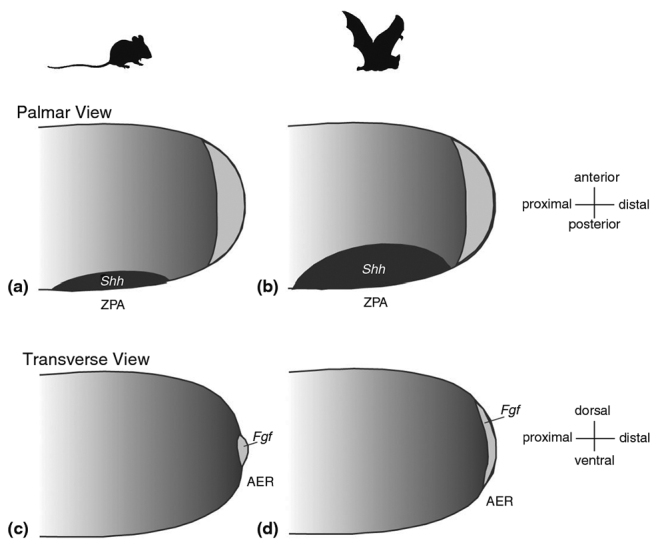
The molecular basis of the general enlargement of the bat forelimb, including long bone elongation in bat wings, has been investigated using candidate gene and genomic approaches, both individually and as part of the whole limb. Candidate gene studies of the limb demonstrate that the two main signaling centers of the limb, the apical ectodermal ridge (AER) and the zone of polarizing activity (ZPA), are up to three times larger in the developing limb buds of bats than in those of mice (Cretekos et al., 2007; Cooper et al., 2012). The expression domain of the anterior-posterior patterning gene Shh, the molecular marker of the ZPA, is initially expanded in the limb buds of bats compared to those of mice (Cooper et al., 2012). By the limb paddle stage (when the developing limb resembles a paddle), Shh expression is turned off in both bats and mice. However, Shh expression is later reinitiated in bats, but not in mice, due to a novel domain of Fgf8 expression. This leads to a feedback loop (Figure 2) whereby Fgf8 increases Shh, which increases Bmp2, which increases Grem, which in turn continues to prompt Fgf8 (Hockman et al., 2008). Both the prolonged expression and the larger expression domains of these genes are believed to ultimately contribute to limb enlargement and digit lengthening (Cooper et al., 2012). This feedback loop also contributes to the formation of a portion of the wing membrane, which will be discussed later. Candidate gene studies have also identified a posterior expansion of the expression of Hoxd13, a gene with known roles in limb development, in the bat wing relative to the mouse forelimb (Chen et al., 2005; Ray and Capecchi, 2008).
Figure 2 -. Differences in signaling centers between bat and mouse limb development. Larger expression domains in bat AER and ZPA, combined with a feedback loop caused by later re-initiation of Fgf8 and Shh, contributes to lengthening in bat forelimbs (reproduced from Cooper and Sears, 2013).
The bat wing has also become a frequent target of transcriptomic and genomic analyses. Comparisons of the transcriptomes of the developing wings of bats and forelimbs of mice and/or of the fore- and hind limbs of bats have revealed several genes with unique expression levels in the bat wing relative to the bat hind and mouse fore- limbs. These genes include the early limb bud marker Meis2, which is upregulated in the developing bat wing (Mason et al., 2015), as well as all of the 5’ HoxD genes (Hoxd9-13) including Hoxd13 (which was identified through candidate gene studies described above), Hoxa13, Tbx3, Evx2, and Fam5c (Wang et al., 2014; Maier et al., 2017). Studies restricted to developing digit tissues and their follow-up analyses have also identified several genes from the Bmp pathway, Tbx and Hox families, and others that are differentially expressed in bat wing digits. These include the genes Tbx3, Tbx15, Bmp3, RGMB, Smad1, Smad4, Nog, Hoxd8, Hoxd9, Hoxa1, Satb1, Twist1, Tmeff2, Mab21l2, and Enpp2 (Wang et al., 2010; Dai et al., 2014).
The genetic drivers underlying the patterns of bat wing gene expression described above are currently under study. Because the protein-coding regions of most patterning genes with important roles in limb development appear to be largely conserved across bats and other mammals (Chen et al., 2005; Sears et al., 2006; Cretekos et al., 2007; Ray and Capecchi, 2008), it is likely that many if not most of the morphological specializations of the bat wing have evolved through changes in gene regulation (Petit et al., 2017).
Ten years ago, two separate studies began the search for the regulatory elements behind the unique phenotypes of the bat wing. In the first study, the authors made a genetically-modified mouse in which they replaced the limb-specific transcriptional enhancer of the Prx1 locus with the orthologous sequence from the bat Carollia perspicillata (Cretekos et al., 2008). The resulting mice displayed forelimb long bones that were modestly but significantly longer than controls. These results are consistent with the phenotypes of mice lacking Prx1 function; Prx1 null mice display defects in the limb skeleton, including shortening of multiple long bone elements of the forelimb (Martin et al., 1995). The second study investigated the structure and function of a known HoxD limb enhancer, the Global Control Region (GCR) in bats, following up on the finding that HoxD13 expression differs in bat and mouse forelimbs (Ray and Capecchi, 2008). Researchers identified several sequence differences in the bat and mouse GCR, and found the bat GCR capable of driving distinct expression domains in transgenic mice relative to that of other mammals. Taken together, these results support the hypothesis that changes in gene regulation contribute to the morphological specialization of the bat forelimb for powered flight.
More recently, researchers have begun to apply a systematic approach to their study of the role of regulatory evolution in the development of the bat wing. By comparing the genomes of diverse mammals and aligning observed differences to previously assembled datasets of putative mouse limb enhancers, researchers have identified 166 bat accelerated regions (BARs) that have evolved at a faster rate in bats than other mammals (Booker et al., 2016). Five out of five of these putative bat enhancers displayed limb enhancer activity when tested in transgenic mice. These enhancers are located near five genes with known roles in limb development that are differentially expressed in bat wings: Twist2, Spry1, Shh, Spg20, and the HoxD cluster. Key among these enhancers may be BAR116, which appears to play a role in regulating the HoxD complex (Booker et al., 2016). The role of BAR116 in the divergent development of fore- and hind limbs was further tested by Yakushiji-Kaminatsui et al. (2018). This study examined the CS93 sequence in chick that is believed to be orthologous to BAR116. CD93 in chick displayed similar enhancer activity to BAR116 in bat, with more expression in the fore- than hind limbs. Furthermore, the CS93/BAR116 sequence was more conserved between chicks and bats than with mice, potentially owing to the increased disparity between fore- and hind limbs in these taxa.
ChIP-Seq assays have also been performed directly on the limbs of bats. Data from these assays has been used to identify almost 3,000 potential regulatory elements that display elevated evolutionary rates in bats and are located near genes with known roles in limb development (Eckalbar et al., 2016). While many of the enhancers driving wing development remain widely unknown, this research has great potential to identify genetic drivers of morphological specialization in bats and, by so doing, start filling in the genotype - to - phenotype map, a major goal of biology.
Bat cranial structures: Variation with diet
Bats display an extraordinary amount of morphological specialization and diversity in their crania. For example, the size and shape of the ears, nose ornaments, eyes, faces, and teeth dramatically differ among species. We present a brief overview of research into the developmental basis of cranial diversity in bats, an exciting and growing area of research. For a more thorough review of how development potentially creates diversity in mammals more broadly, please see the recent review by (Usui and Tokita, 2018).
Bats display a broad array of facial lengths that are tightly linked to the functional requirements of their diets. For example, bats that feed on nectar tend to have very long faces that facilitate their penetration into flowers, while bats that feed on hard fruit have very short faces that allow them to bite with higher force (Dumont et al., 2009; Nogueira et al., 2009). Face shape is determined during development by coordinated growth and ossification of neural crest derived ectomesenchyme in the first pharyngeal arch (which gives rise to the mandible and maxilla) and frontonasal process (Santagati and Rijli, 2003). Facial shape can be developmentally altered by changes in the migration and number of neural crest cells during early embryonic development, and/or in maturation and growth of neural crest derived skeletal elements later in development or during postnatal growth. These skeletal elements undergo ossification either via a cartilaginous template like in the appendicular skeleton (known as endochondral ossification), or directly from the ectomesenchyme, (known as membranous ossification) (Hall, 2005). Both of these ossification processes are controlled by the transcription factor Runx2 (Komori, 2018). The activity of Runx2 relies on a highly conserved Runt DNA binding domain, shared by other members of the runt-like gene family, and a unique region of repeated glutamines (Q) followed by a region of repeated alanines (A). These repeated amino acids are reflected in repeated sequences within the genome. Such simple sequence repeats are more prone to mutation than many other parts of the genome and so are thought to act in fine-tuning gene expression (Kashi and King, 2006). The QA ratio in Runx2 has been demonstrated to correlate with face length, first in domestic dogs (Fondon and Garner, 2004), and subsequently across a number of mammalian groups (Sears et al., 2007; Pointer et al., 2012; Ferraz et al., 2018). This includes phyllostomid bats, where the Runx2 QA ratio correlates with width and length of the membranous palatine bone, which in turn influences face shape and length. In bats, a low QA ratio is found in nectarivores with long and narrow faces, while a high QA ratio is found in frugivores with short and wide faces (Ferraz et al., 2018). QA repeats in the Runx2 locus may be acting to modulate the activity of the gene, since Runx2 acts early in bone development to promote differentiation of undifferentiated mesenchyme into osteoblasts, and later inhibits terminal differentiation of osteoblasts into osteocytes (reviewed in Komori, 2018). Therefore, changes in Runx2 activity will affect skeletal growth, as supported by cell based expression assays demonstrating that Runx2 activity, as determined by the expression of its target Col10a, increased in a linear fashion with QA ratio (Sears et al., 2007). Thus it is possible that this relatively changeable domain has facilitated phyllostomid morphological evolution by altering palate dimension and thus face shape, thereby allowing for the exploitation of new diets that are dependent on face shape.
The skull is a morphologically complex structure. Geometric morphometrics enable detailed and quantitative comparison of skull shape. Recent morphometric analysis of bat skull developmental and evolutionary trajectories across species suggests that variation in phyllostomid skull morphology is driven by peramorphosis, such that development of the skull is similar across species until a basal morphology is reached and then those species with a specialized form continue developing to achieve new morphologies (Camacho et al., 2019). Interestingly, bats with different morphologies seem to have subtly differing strategies for achieving peramorphosis. Generalist species and frugivores achieve their short skull form by accelerating the rate of their development, whereas nectarivores and sanguinivores diverge through a longer period of fetal and post-natal development. The hypothesis that prolonged or accelerated skull development drives morphological change in the bat crania bookends well with the hypothesized increase in Runx2 QA tandem repeats, since extended or increased osteoblast activity facilitated by Runx2 activity would be consistent with peramorphosis of skull bones.
The importance of Runx2 in face length and width offers a candidate for the regulation of bat craniofacial morphology through gene regulation. Another possible candidate for this is the paired homeobox transcription factor Pax9. This transcription factor has a well-established role in craniofacial development, including the development of teeth (Peters et al., 1998), mandibular bone (Anthwal et al., 2015), the lip (Nakatomi et al., 2010), and palate (Peters et al., 1998; Zhou et al., 2013). Phillips and colleagues (2013) found a conserved region of the 3’ untranslated region (UTR) of the Pax9 gene containing Musashi-binding elements (MBE) in phyllostomid and vespertilionid bats. Musashi proteins are known to regulate a range of cellular processes, including stem cell fate decisions and cellular differentiation, by the inhibitory binding mRNA via MBEs. As such, MBEs may play a role in the control of Pax9 gene expression level. The number of MBEs vary across phyllostomid bats with variable facial morphologies, but not across vespertilionids. This suggests that, at least in phyllostomids, variation in Pax9 may contribute to the generation of facial diversity. While many other genes are known to be important in facial morphogenesis in mammals, their role in bat facial diversification remains largely unexplored.
Another remarkable facial variation present in some bat species are facial clefts, either midline clefts or paramedian clefts, where there are two clefts on either side of the face and a remnant midline tissue. Orr and colleagues (2016) undertook an analysis of facial clefts in bats and concluded that such clefts resemble some cleft lip and palate disorders seen in human patients in that the cleft is not accompanied by other phenotypes elsewhere in the body. This is in contrast to mouse transgenic models of facial clefts such as knockouts of the Tgf-beta or Fgf signaling pathways, where many other structures of the organism are impacted. However, bats' clefts also differ in some ways from those seen in human clinical cases. Bat clefts are only found in the hard tissues of the face, whereas human clefts are found in both soft and hard tissues. Furthermore, the hard tissue clefts of bats are bridged by fibrous tissue (Orr et al., 2016) that, along with the intact soft tissues, probably ensures that suckling is possible during early postnatal life. The identity of the fibrous bridging material in clefts is not known. Possibly be the remnants of the skeletogenic mesenchyme arrested before ossification, and as such clefts in bats are not due to failure in midline fusion of the orofacial primordium during embryonic development (Orr et al., 2016). This hypothesis is supported by the absence of soft tissue clefts in bats. Modulation of ossification may therefore be driving the specification of these clefts in bats.
Diversification during adaptive radiation
The concept of adaptive radiation provides a theoretical framework for explaining the taxonomic and functional diversity of organisms in response to ecological selective pressures (Simpson, 1944). In this framework, the availability of ecological opportunities, i.e. ecological niches that become available through the colonization of new environments, the extinction of competitors, or the acquisition of key innovations, is linked to rapid diversification (Yoder et al., 2010). Often described as a classic example of an adaptive radiation, the acquisition of flight in bats is commonly thought to have opened up many new ecological opportunities and thereby enabled their diversification (Speakman and Racey, 1991; Simmons and Geisler, 1998; Fenton, 2010). However, while flight is likely to have played a crucial role in bats diversification, other adaptations have also likely contributed, including the specializations of the skull discussed above. In this section, we explore research in two additional systems that have likely played important roles in the adaptive radiations of bats: vision and dentition.
Adaptive radiations and sensory systems - an example from bat vision
Among novel phenotypic traits that are commonly thought to promote diversification (Yoder et al., 2010), sensory adaptations are considered key innovations that enable access to hitherto inaccessible ecological resources. For example, trichromatic vision in primates enables fruit discrimination against a background of leaves, and echolocation in cetaceans enables orientation and hunting in low-light environments (Jacobs, 2009; Steeman et al., 2009; Teeling et al., 2012; Geisler et al., 2014). Bats are known to possess many key sensory adaptations; perhaps the most well-known of which being their highly specialized hearing system that allows the true echolocation that helps most species navigate, forage, and/or hunt in the dark (recently reviewed in Yohe and Brand, 2018). However, research over the past 15 years has begun to highlight the importance of adaptations in other sensory structures to the evolutionary success of bats (Davies et al., 2013), including variation in the sizes of the cochlea (Davies et al., 2013), eyes (Heffner and Heffner, 1992; Eklöf, 2003), olfactory epithelium and olfactory bulb, as well as in the vomeronasal epithelium and accessory olfactory bulb (Bhatnagar and Smith, 2007; Smith et al., 2012; Yohe et al., 2017; Yohe and Brand, 2018). Of particular note, studies of the evolution of bat vision in the last two years have provided unprecedented insights into the role of sensory adaptations in bat evolution (Gutierrez et al., 2018b; Kries et al., 2018; Wu et al., 2018; Li et al., 2018; Simoes et al., 2019; Sadier et al. 2018). Because of this, we further discuss the evolution of the visual systems of bats.
Most mammals possess three visual opsins: rhodopsin (RHO) in rods, long-wave (M/L) sensitive opsin (OPN1LW) in L-cones, and short-wave (S) sensitive opsin (OPN1SW) in S-cones (Yokoyama and Radlwimmer, 1999, Yokoyama, 2000, Zhao et al., 2009). While many nocturnal animals are monochromatic (i.e., completely color‐blind as a result of only having rhodopsin and one cone opsin), others possess dichromacy and are capable of some chromatic distinctions (Peichl, 2005; Jacobs, 2009; Zhao et al., 2009; Jacobs, 2012; Veilleux et al., 2013). However, up to recently, our knowledge of bat vision has remained obscure. The first studies of the large eyes of flying foxes were based on microscope observations and suggested that their retinas contained only rods and no cones (Fritsch, 1911; Kolmer, 1910; Neuweiler, 1962). For a long time, 20many other species, in particular Yangochiropterans one with small eyes, were considered nearly blind. This vision started to change almost a century later when the eyes of flying foxes were reexamined, revealing that these bats also possess cones, albeit at a low cone/rod ratio, suggesting that flying foxes potentially have color vision (Jacobs, 1993) and another study tested the response of bats to different wavelengths of light (Joshi and Chandrashekaran, 1985).
More recently, the growing availability of bat genomic data have resulted in a renewed interest in bat vision. The first study that examined the molecular aspects of opsin genes in bats investigated the S and M/L opsin genes of two species of Yinpterochiroptera and one species of Yangochiroptera sequenced at that time (Wang et al., 2004). All of these species were found to possess both M/L and S opsins (Zhao et al., 2009) suggesting that these species of bats, at least, are dichromatic. Then, a number of studies began to investigate opsin sequences and electrophysiological vision capabilities in bats. Using immunohistochemistry (IHC), sequencing, and electrophysiological experiments, Müller et al. (2009) found that both M/L and S opsins are expressed and functional in two plant-visiting phyllostomid species, and that differences in cone densities in these species are potentially linked to differences in visual acuity. A separate study in four genera of Yinpterochiroptera that also used IHCs revealed that, while all species likely possess both dim and day light vision, these genera likely vary in their ability to see in color: Pteropus possesses both S and M/L opsins, and thus dichromatic vision, whereas Rousettus, Eidolon, and Epomorphorus only express M/L opsin and, consequently, are monochromatic (Müller et al., 2007). Interestingly, the authors of this study linked these differences in photoreceptor composition to the roosting preferences of these genera, as Pteropus is often more exposed to daylight than the others. By studying multiple species, these studies revealed that bat vision is more complex than previously thought, and established tight links between the visual capabilities of bats and their ecological niche specializations, a link that has been documented in other mammals (Douglas and Jeffery, 2014).
Building on these discoveries, research in the last decade has investigated the relationship between ecological pressures and vision capabilities in multiple bat species. Results of these studies, most of which have been based on gene sequences alone and fairly limited taxonomic sampling, suggest that the S opsin gene is likely functional across Yangochiroptera but lost in some Yinpterochiropteran lineages (Winter et al., 2003; Feller et al., 2009; Zhao et al., 2009; Müller et al., 2009; Butz et al., 2015; Gorresen et al., 2015). Because inferences from amino acid sequence analyses and action spectra suggest that bat S opsins are sensitive to UV light, researchers proposed that the loss of S-opsins could have profound impacts on bat visual acuity. Accordingly, researchers hypothesized the retention of S-opsins is possibly related to the demands of visual processing in mesopic, or low-light, conditions (Zhao et al., 2009), and/or plant visiting (Müller et al., 2007; Feller et al., 2009; Müller et al., 2009; Butz et al., 2015). However, the limited sampling of these studies hindered their ability to rigorously test links between vision capabilities and ecological parameters, in particular within the Yangochiroptera.
In 2018, multiple studies on bat vision featuring larger taxonomic sampling of multiple independent lineages representing diverse ecologies (e.g. blood feeding, plant-visiting species) were published. These studies revealed evidence of OPN1SW pseudogenization events in both Yangochiroptera and Yinpterochiroptera (Kim et al., 2008; Kries et al., 2018; Wu et al., 2018; Li et al., 2018; Simoes et al., 2019), suggesting that UV vision loss is more widespread in bats than previously thought. In most of these studies the functionality of OPN1SW was based only on analyses of DNA and/or RNA sequence, and not on the S opsin protein. However, one study combined investigation of the conservation and expression of opsin gene sequences with the localization of M/L and S opsin proteins in 56 species of Yangochiroptera (Sadier et al., 2018). Results of this study suggest that, while the DNA sequences of S opsin genes most commonly appear functional, the gene is often not translated and/or transcribed (Figure 3). This occurs in multiple independent lineages, suggesting that bat lineages take several routes to a loss of S opsin function.
Figure 3 -. Color vision diversity in Phyllostomids (adapted from Sadier et al., 2018). Cones are shown on flat-mounted retinas in four different bat species representative of the cone diversity in Phyllostomids. To visualize L opsin (green) and S opsin (magenta), flat-mounted retinas were probed with two antibodies against L and S opsins. L opsin expressing cones were found in all bats, while S opsins were only seen in fruit visiting Carollia and Artibeus species, indicating cone type and diversity vary between species occupying different dietary niches. Diet is indicated with a pictogram.

To link the loss of S opsin function with ecological adaptation, studies tested for trade-offs between vision state and other sensory adaptations (Gutierrez et al., 2018b; Kries et al., 2018; Wu et al., 2018; Li et al., 2018; Simoes et al., 2019), and for correlations between vision state and various ecological parameters (Wu et al., 2018; Sadier et al., 2018; Gutierrez et al., 2018b; Simoes et al., 2019). These findings revealed that UV vision is lost in many bat species, and not just in high duty cycle (HDC) echolocators (Gutierrez et al., 2018b; Simoes et al., 2019), and does not support the existence of a trade-off between the evolution of HDC echolocation and loss of functional S opsin. However, other studies have found some support for an association of frugivory (Sadier et al., 2018) and roosting behavior (Müller et al., 2007) with S opsin retention in bats, and yet others for a tradeoff between UV vision and infrared sensing in the common ancestor of vampire bats (Kries et al., 2018; Li et al., 2018). Bat rhodopsin has also been found to show evidence of adaptation to dim light conditions (Gutierrez et al., 2018a). Very recently (Davies et al. 2020) found that the nectar‐feeding lineages of noctilionoids and the Stenodermatinae subfamily of fig‐eating bats exhibit molecular adaptations resulting from fine-tuning of pre-existing visual adaptations, suggesting that noctilionoids which use visual cues for identifying food and roosts and orientation were preadapted to colonized these new ecological niches. These exciting findings reveal that vision has taken a major role during bat adaptation and diversification and opens a new field of research to investigate.
Adaptive radiations and morphological diversification: Insights from bat teeth
Among vertebrates in general and mammals in particular, teeth are one of the most variable organs of the body, being highly diverse in terms of number, shape, and size. One of the main factors suspected of driving this diversity is dietary ecology, which is thought to have shaped dental morphology with some dental traits evolving numerous times in association with diet (Lucas, 1979, Hunter and Jernvall, 1995; Lucas and Peters, 2007). Indeed, intensive study of mammals from unrelated groups has uncovered many convergences, with highly similar dentitions characterizing distinct groups with similar diets (Evans et al., 2007).
Bats exhibit diverse dentitions associated with their many dietary types (Freeman, 1988; Freeman, 1998) (Figure 4). While the ancestral bat was likely insectivorous (Gunnell and Simmons, 2005), subsequent bat lineages have evolved many different diets: insectivory, carnivory, sanguivory, omnivory, nectarivory, and frugivory. Foundational work on bat teeth diversity has been primarily performed by Patricia Freeman. By studying 108 species from 78 genera from both Yinpterochiroptera and Yangochiroptera, Freeman (1998, and reviewed in Swartz et al., 2003) characterized the diversity of bat teeth. She showed that the relative tooth area is highly variable between Yinpterochiroptera and Yangochiroptera, as well as within Yinpterochiroptera (particularly in carnivorous species), and linked some differences in form to diet. Insectivorous bats possess a triangular jaw with regularly spaced teeth, nectarivorous bats an elongated jaw with long and narrow teeth, and frugivorous bats a short jaw with unequal teeth. Within frugivores, stenodermatid bats have an extreme phenotype with an extremely short face and rounded jaw (Freeman, 1998). Among them, Centurio senex displays a striking example of the extreme phenotype of stenodermatid bats, with its extremely short face and small canines.
Figure 4 -. Jaw and dental diversity of Noctilionoid bats. Members of the group occupy every possible dietary niche found in bats. In line with this, jaw size and shape as well as tooth shape, size and proportion are highly variable in Noctilionoid bats. In fact, the jaw and tooth types shown represent most of the breadth of the diversity found in both bat superfamilies. Diet is indicated with a pictogram. Scale: 10 mm.

Much of the subsequent research on bat dentition and diet has focused on Neotropical bats because of their incredible variation. Bats of the Noctilionidea, which includes the family Phyllostomidae (~200 species of New World leaf-nosed bats and allies within the suborder Yangochiroptera), have received particular attention because they are the only bat group in which all possible bat diets (insectivory, carnivory, piscivory, sanguivory, omnivory, nectarivory and frugivory) have been documented. This group diversified approximately 40 million years ago (Rojas et al., 2013; Rossoni et al., 2017) and today exhibits morphological adaptations linked to their broad dietary specializations (Monteiro and Nogueira, 2011; Dumont et al., 2012; Davies et al., 2013; Hayden et al., 2014; Yohe et al., 2015). For example, bats from this group display a diversity of biting behaviors and bite forces that facilitate their broad range of diets (Freeman, 1998; Dumont, 1999; Santana et al., 2011; Dumont et al., 2012). Biting behavior varies most substantially among frugivorous species of phyllostomids (Dumont, 1999). Specialized frugivores use one side of their mouth to bite their food, unspecialized frugivores use both sides, and omnivores have been shown to modulate their biting behavior in response to food hardness. Other studies have also linked the molar complexity of this group to their feeding performance (Evans et al., 2007; Santana et al., 2011). Taken together, these studies illuminate the diversity of diet, teeth morphology and biting behavior in bats making them an excellent group to study the diversification of an organ in relationship with its dietary ecological niche.
Because of this, bat teeth represent a great model to study the developmental basis of morphological transitions from one diet, and tooth type, to another, a shift that occurs recurrently and rapidly in mammals. Future studies should build upon the extensive foundational work that has been done in this promising study system and investigate the developmental mechanisms underlying the evolutionary diversification of bat teeth.
Origins of morphological novelty - Insights from bat wing membranes
In elongating their forearms and digits, bats created a scaffold for the dynamic, collapsible, and compliant tissue that comprises the membranes of the bat wing. The functional combination of membrane tension and elasticity has allowed for unparalleled flight capability in bats, which in turn has provided the basis for their remarkable diversification (Simmons, 2005; Teeling et al., 2005).
The bat wing consists of three membranes (patagia): dactylopatagium, between the digits; plagiopatagium, between the fifth digit to the ankle; and the propatagium, a smaller membrane on the anterior portion of the stylopod and zeugopod (Figure 5a). Additionally, many bats also possess a membrane between the hind limbs, the uropatagium (Swartz et al., 1996). Size and shape of these patagia can vary greatly between species (Figure 5b) and are often tightly correlated with both flight style and diet (Norberg, 2012), ranging from high speed in insectivores to high maneuverability in nectarivores and frugivores (Wetterer et al., 2000; Gunnell and Simmons, 2012). Insectivores have long, narrow plagiopatagia that are adapted for increased speed during aerial hawking; whereas frugivores tend to have short, broad membranes that allow for increased maneuverability in cluttered vegetation environments. In general, the combined forelimb membranes generate lift, while the uropatagia provides stability and can have a functional role in catching insects during midair hunting.
Figure 5 -. The anatomy and development of the bat wing (a) Outline of extended forelimb and wing membranes in chiropterans demonstrating the skeleton and membranes (patagia) of the wing (from Swartz et al., 1996). (b) Examples of variation in size and shape of patagia (from Norberg and Rayner, 1987). (c) Early limb development in bats compared to mice showing early similarities in limb development and divergence in growth and maturation. (from Cretekos et al., 2008).

Of the four membranes of the bat wing, only one, the dactylopatagium, has an obvious homology in non-bat mammals, while the other three are seemingly novel traits. The dactylopatagium is also the only bat wing membrane for which the underlying developmental origins have been reasonably resolved. Mice, and most other mammals, undergo interdigital tissue regression following condensation of the digits (Figure 5c). In bats, the continued presence of Fgf8 combined with the Bmp inhibiting effects of Grem prevent the activation of the apoptotic pathway that would otherwise degrade the interdigital tissues and lead to freed digits (Weatherbee et al., 2006; Cooper and Sears, 2013). The result is the retention of interdigital tissues into the adult as the dactylopatagium (Figure 5c).
Beyond the developmental origins of the dactylopatagium, the developmental processes leading to the formation of the patagia and their subsequent diversification into a multitude of sizes and shapes remain largely unknown. One potential clue to the drivers of these phenomena was the discovery by Tokita et al. (2012) that expression of Fgf10 continues in the anterior-proximal portion of the bat forelimb mesenchyme, long after expression in other taxa (e.g., Mus, Monodelphis, and Gallus) had ceased. However, while Fgf10 signaling is active in areas where future wing membranes develop, its expression is restricted to the area immediately adjacent to the wing musculature. This suggests that some other mechanism(s) are responsible for the outgrowth and formation of the membrane itself.
As truly novel structures, the patagia (plagio-, pro-, and uro- patagia) beyond the dactylopatagia lack any known homology within mammals. These novel patagia play a significant role in chiropteran flight abilities, and so to comprehensively understand how flight has evolved in bats we need to understand the origin and diversification of these novel membranes.
Conclusions
Through a series of adaptive radiations, bats have evolved to be highly diverse. This diversity makes bats a model system for addressing biological questions on topics including biomechanics, morphological evolution, longevity, developmental evolution, sensory system adaptations, and many others. In this review, we have provided an overview of how evolution and development have molded some of the most unique morphological specializations of bats.
The elongated forelimb bones that support bat wings display higher growth rates than those of other mammals. This accelerated growth is preceded by formation of ZPA’s and AER’s in bat forelimbs that are three times larger than those of mice, coupled with changes in the associated Shh and Fgf8 feedback loop. Transcriptomic and genomic analyses have also revealed significant upregulation of HoxD genes and several other affiliated pathways in bat forelimbs relative to the forelimbs of other mammals. Evidence is starting to accumulate that changes in gene regulation strongly contribute to these differences in gene expression and the highly specialized forelimb structures of bats.
The cranial features of bats (ears, nose ornaments, and faces) can vary dramatically from species to species. Facial length is tightly correlated with diet type and activity of the Runx2 transcription factor, and regulation of Pax9 expression has been linked to bat facial diversity as well. Alterations in overall skull shape among bats have been shown to be accomplished through the acceleration and extension of development. Meanwhile, the facial clefting that some bats exhibit is likely due to modulation of ossification, leaving soft tissues unaffected.
Bat sensory systems, especially vision, have also been found to vary much more from species to species than previously assumed. Until fairly recently, most bat species were thought to have monochromatic vision, possessing rods and perhaps only L- or S- opsins. However, newer studies have shown that surprising numbers of bat species possess both L- and S-cones, and thus likely have dichromatic vision. This phenotype has been linked to ecological specializations (diet and roosting habits). Several lineages have also been found to have lost UV vision through multiple routes of S- opsin loss of function, including post-transcriptional processes.
Bats have evolved many feeding specializations, and the dentition of different groups has been tightly linked with diet type. Both tooth area and proportion are highly variable among bat species, with molar complexity and biting behavior also varying substantially from bat to bat and species to species. Because of their diversity and strong ties to diet, bat teeth have the potential to be a great model for understanding how changes in ecological niche lead to diversification of biological structures.
In addition to their numerous specializations in existing morphological structures, bats also possess truly novel morphological structures in their plagio-, pro-, and uro- patagia. While the developmental processes that have shaped the evolution of another wing membrane, the dactylopatagium, have been explored, the processes driving the formation of these novel membranes remain unknown. Given the importance of these membranes to powered flight in bats, future study of the development of these novel membranes is critical to our understanding of the evolution of powered flight and has the potential to provide insights into the processes behind the evolution of novel traits, an outstanding question in evolutionary biology.
Each of the areas discussed here have ample room for future research, and there is much more work to be done to uncover the full range of developmental mechanisms underlying the specializations and diversity of bat morphology. Time and time again, bats have proven to be a model system for study of the developmental basis of morphological specialization, evolutionary novelty, and adaptive radiation. We look forward to the many answers bats have yet to reveal.
References
- Adams RA. Morphogenesis in bat wings: linking development, evolution and ecology. Cells Tissues Organs. 2008;187:13–23. doi: 10.1159/000109960. [DOI] [PubMed] [Google Scholar]
- Anthwal N, Peters H, Tucker AS. Species-specific modifications of mandible shape reveal independent mechanisms for growth and initiation of the coronoid. Evodevo. 2015;6:35. doi: 10.1186/s13227-015-0030-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bhatnagar KP, Smith TD. Light microscopic and ultrastructural observations on the vomeronasal organ of Anoura (Chiroptera : Phyllostomidae) Anat Rec. 2007;290:1341–1354. doi: 10.1002/ar.20601. [DOI] [PubMed] [Google Scholar]
- Booker BM, Friedrich T, Mason MK, VanderMeer JE, Zhao J, Eckalbar W, Logan M, Illing N, Pollard KS, Ahituv N. Bat accelerated regions identify a bat forelimb specific enhancer in the HoxD locus. PLoS Genet. 2016;12:e1005738. doi: 10.1371/journal.pgen.1005738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown EE, Cashmore DD, Simmons NB, Butler RJ. Quantifying the completeness of the bat fossil record. Palaeontology. 2019;62:757–776. [Google Scholar]
- Burgin CJ, Colella JP, Kahn PL, Upham NS. How many species of mammals are there? J Mammal. 2018;99:1–14. [Google Scholar]
- Butz E, Peichl L, Müller B. Cone bipolar cells in the retina of the microbat Carollia perspicillata. J Comp Neurol. 2015;523:963–981. doi: 10.1002/cne.23726. [DOI] [PubMed] [Google Scholar]
- Chen CH, Cretekos CJ, Rasweiler JJ, IV, Behringer RR. Hoxd13 expression in the developing limbs of the short‐tailed fruit bat, Carollia perspicillata. Evol Dev. 2005;7:130–141. doi: 10.1111/j.1525-142X.2005.05015.x. [DOI] [PubMed] [Google Scholar]
- Camacho J, Heyde A, Bhullar BS, Haelewaters D, Simmons NB, Abzhanov A. Peramorphosis, an evolutionary developmental mechanism in neotropical bat skull diversity. Dev Dynam. 2019;248:1129–1143. doi: 10.1002/dvdy.90. [DOI] [PubMed] [Google Scholar]
- Cooper LN, Cretekos CJ, Sears KE. The evolution and development of mammalian flight. Wiley Interdiscip Rev Dev Biol. 2012;1:773–779. doi: 10.1002/wdev.50. [DOI] [PubMed] [Google Scholar]
- Cooper LN, Sears KE. How to grow a bat wing. In: Adams R, Pederson S, editors. Bat evolution, ecology, and conservation. Springer; New York: 2013. pp. 3–20. [Google Scholar]
- Cretekos CJ, Deng JM, Green ED, Program NCS, Rasweiler JJ, Behringer RR. Isolation, genomic structure and developmental expression of Fgf8 in the short-tailed fruit bat, Carollia perspicillata. Int J Dev Biol. 2007;51:333–338. doi: 10.1387/ijdb.062257cc. [DOI] [PubMed] [Google Scholar]
- Cretekos CJ, Wang Y, Green ED, Martin JF, Rasweiler JJ, Behringer RR. Regulatory divergence modifies limb length between mammals. Genes Dev. 2008;22:141–151. doi: 10.1101/gad.1620408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dai M, Wang Y, Fang L, Irwin DM, Zhu T, Zhang J, Zhang S, Wang Z. Differential expression of Meis2, Mab21l2 and Tbx3 during limb development associated with diversification of limb morphology in mammals. PLoS One. 2014;9:e106100. doi: 10.1371/journal.pone.0106100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davies KTJ, Bates PJJ, Maryanto I, Cotton JA, Rossiter SJ. The evolution of bat vestibular systems in the face of potential antagonistic selection pressures for flight and echolocation. PLoS One. 2013;8:e61998. doi: 10.1371/journal.pone.0061998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davies KT, Yohe LR, Almonte J, Sánchez MK, Rengifo EM, Dumont ER, Sears KE, Dávalos LM, Rossiter SJ. Foraging shifts and visual preadaptation in ecologically diverse bats. Mol Ecol. 2020;29:1839–1859. doi: 10.1111/mec.15445. [DOI] [PubMed] [Google Scholar]
- Douglas R, Jeffery G. The spectral transmission of ocular media suggests ultraviolet sensitivity is widespread among mammals. Proc Biol Sci. 2014;281:20132995. doi: 10.1098/rspb.2013.2995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dumont ER. The effect of food hardness on feeding behaviour in frugivorous bats (Phyllostomidae): an experimental study. J Zool. 1999;248:219–229. [Google Scholar]
- Dumont ER. Feeding mechanisms in bats: variation within the constraints of flight. Integr Comp Biol. 2007;47:137–146. doi: 10.1093/icb/icm007. [DOI] [PubMed] [Google Scholar]
- Dumont ER, Herrel A, Medellin RA, Vargas-Contreras JA, Santana SE. Built to bite: cranial design and function in the wrinkle-faced bat. J Zool. 2009;279:329–337. [Google Scholar]
- Dumont ER, Davalos LM, Goldberg A, Santana SE, Rex K, Voigt CC. Morphological innovation, diversification and invasion of a new adaptive zone. Proc Biol Sci. 2012;279:1797–1805. doi: 10.1098/rspb.2011.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eckalbar WL, Schlebusch SA, Mason MK, Gill Z, Parker AV, Booker BM, Nishizaki S, Muswamba-Nday C, Terhune E, Nevonen KA, et al. Transcriptomic and epigenomic characterization of the developing bat wing. Nat Genet. 2016;48:528–536. doi: 10.1038/ng.3537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eklöf J. Vision in echolocating bats. Göteborg University; Gothenburg: 2003. 107 PhD Thesis. [Google Scholar]
- Evans AR, Wilson GP, Fortelius M, Jernvall J. High-level similarity of dentitions in carnivorans and rodents. Nature. 2007;445:78–81. doi: 10.1038/nature05433. [DOI] [PubMed] [Google Scholar]
- Farnum CE, Tinsley M, Hermanson JW. Forelimb versus hindlimb skeletal development in the big brown bat, Eptesicus fuscus: functional divergence is reflected in chondrocytic performance in autopodial growth plates. Cells Tissues Organs. 2008;187:35–47. doi: 10.1159/000109962. [DOI] [PubMed] [Google Scholar]
- Feller KD, Lagerholm S, Clubwala R, Silver MT, Haughey D, Ryan JM, Loew ER, Deutschlander ME, Kenyon KL. Characterization of photoreceptor cell types in the little brown bat Myotis lucifugus (Vespertilionidae) Comp Biochem Physiol B. 2009;154:412–418. doi: 10.1016/j.cbpb.2009.08.006. [DOI] [PubMed] [Google Scholar]
- Fenton MB. Convergences in the diversification of bats. Curr Zool. 2010;56:454–468. [Google Scholar]
- Ferraz T, Rossoni DM, Althoff SL, Pissinatti A, Paixao-Cortes VR, Bortolini MC, Gonzalez-Jose R, Marroig G, Salzano FM, Goncalves GL, et al. Contrasting patterns of RUNX2 repeat variations are associated with palate shape in phyllostomid bats and New World primates. Sci Rep. 2018;8:7867. doi: 10.1038/s41598-018-26225-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fondon JW, Garner HR. Molecular origins of rapid and continuous morphological evolution. Proc Natl Acad Sci U S A, 2004;101:18058–18063. doi: 10.1073/pnas.0408118101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Freeman P. Form, function, and evolution in skulls and teeth of bats. In: Kunz TH, Racey PA, editors. Bat Biology and Conservation. Smithsonian Institution Press; Washington, D.C.: 1998. pp. 140–156. [Google Scholar]
- Freeman PW. Frugivorous and animalivorous bats (Microchiroptera) - dental and cranial adaptations. Biol J Linn Soc. 1988;33:249–272. [Google Scholar]
- Fritsch G. Contributions to the histology of the eye of the Pteropus. Z Wiss Zool. 1911;98:288–296. [Google Scholar]
- Geisler JH, Colbert MW, Carew JL. A new fossil species supports an early origin for toothed whale echolocation. Nature. 2014;508:383–386. doi: 10.1038/nature13086. [DOI] [PubMed] [Google Scholar]
- Gorresen PM, Cryan PM, Dalton DC, Wolf S, Bonaccorso FJ. Ultraviolet vision may be widespread in bats. Acta Chiropt. 2015;17:193–198. [Google Scholar]
- Gunnell GF, Simmons NB. Fossil evidence and the origin of bats. J Mammal Evol. 2005;12:209–246. [Google Scholar]
- Gunnell GF, Simmons NB. Evolutionary history of bats: fossils, molecules and morphology. Cambridge University Press; Cambridge,: 2012. [Google Scholar]
- Gutierrez ED, Castiglione GM, Morrow JM, Schott RK, Loureiro LO, Lim BK, Chang BSW. functional shifts in bat dim-light visual pigment are associated with differing echolocation abilities and reveal molecular adaptation to photic-limited environments. Mol Biol Evol. 2018;35:2422–2434. doi: 10.1093/molbev/msy140. [DOI] [PubMed] [Google Scholar]
- Gutierrez ED, Schott RK, Preston MW, Loureiro LO, Lim BK, Chang BSW. The role of ecological factors in shaping bat cone opsin evolution. Proc R Soc B-Biol Sci. 2018;285:20172835. doi: 10.1098/rspb.2017.2835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hall BK. Bones and cartilage: Developmental and evolutionary skeletal biology. Elsevier Academic Press; Boston,: 2005. 792 [Google Scholar]
- Hayden S, Bekaert M, Goodbla A, Murphy WJ, Davalos LM, Teeling EC. A cluster of olfactory receptor genes linked to frugivory in bats. Mol Biol Evol. 2014;31:917–927. doi: 10.1093/molbev/msu043. [DOI] [PubMed] [Google Scholar]
- Heffner RS, Heffner HE. Visual factors in sound localization in Mammals. J Comp Neurol. 1992;317:19–232. doi: 10.1002/cne.903170302. [DOI] [PubMed] [Google Scholar]
- Hill JE, Smith JD. Bats: A natural history. Cambridge University Press; Cambridge,: 1984. 243 [Google Scholar]
- Hockman D, Cretekos CJ, Mason MK, Behringer RR, Jacobs DS, Illing N. A second wave of Sonic hedgehog expression during the development of the bat limb. Proc Natl Acad Sci U S A. 2008;105:16982–16987. doi: 10.1073/pnas.0805308105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hockman D, Mason MK, Jacobs DS, Illing N. The role of early development in mammalian limb diversification: A descriptive comparison of early limb development between the natal long‐fingered bat (Miniopterus natalensis) and the mouse (Mus musculus) Dev Dyn. 2009;238:965–979. doi: 10.1002/dvdy.21896. [DOI] [PubMed] [Google Scholar]
- Hunter JP, Jernvall J. The hypocone as a key innovation in mammalian evolution. Proc Natl Acad Sci U S A. 1995;92:10718–10722. doi: 10.1073/pnas.92.23.10718. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacobs GH. The distribution and nature of colour vision among the mammals. Biol Rev. 1993;68:413–471. doi: 10.1111/j.1469-185x.1993.tb00738.x. [DOI] [PubMed] [Google Scholar]
- Jacobs GH. Evolution of colour vision in mammals. Philos Trans R Soc Lond B Biol Sci. 2009;364:2957–2967. doi: 10.1098/rstb.2009.0039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacobs GH. The evolution of vertebrate color vision. Adv Exp Med Biol. 2009;739:156–172. doi: 10.1007/978-1-4614-1704-0_10. [DOI] [PubMed] [Google Scholar]
- Jebb D, Huang Z, Pippel M, Hughes GM, Lavrichenko K, Devanna P, Winkler S, Jermiin LS, Skirmuntt EC, Katzourakis A, et al. Six new reference-quality bat genomes illuminate the molecular basis and evolution of bat adaptations. Nature. 2020;583:578–584. doi: 10.1038/s41586-020-2486-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Joshi D, Chandrashekaran M. Spectral sensitivity of the photoreceptors responsible for phase shifting the circadian rhythm of activity in the bat, Hipposideros speoris. J Comp Physiol A. 1985;156:189–198. [Google Scholar]
- Kashi Y, King DG. Simple sequence repeats as advantageous mutators in evolution. Trends Genet. 2006;22:253–259. doi: 10.1016/j.tig.2006.03.005. [DOI] [PubMed] [Google Scholar]
- Kim TJ, Jeon YK, Lee JY, Lee ES, Jeon CJ. The photoreceptor populations in the retina of the greater horseshoe bat Rhinolophus ferrumequinum. Cells. 2008;26:373–379. [PubMed] [Google Scholar]
- Kolmer W. Zur Kenntnis des Auges der Macrochiropteren. Z Wiss Zool. 1910;97:91–104. [Google Scholar]
- Komori T. Runx2, an inducer of osteoblast and chondrocyte differentiation. Histochem Cell Biol. 2018;149:313–323. doi: 10.1007/s00418-018-1640-6. [DOI] [PubMed] [Google Scholar]
- Kries K, Barros MAS, Duytschaever G, Orkin JD, Janiak MC, Pessoa DMA, Melin AD. Colour vision variation in leaf-nosed bats (Phyllostomidae): Links to cave roosting and dietary specialization. Mol Ecol. 2018;27:3627–3640. doi: 10.1111/mec.14818. [DOI] [PubMed] [Google Scholar]
- Li LF, Chi H, Liu HN, Xia Y, Irwin DM, Zhang S, Liu Y. Retention and losses of ultraviolet-sensitive visual pigments in bats. Sci Rep. 2018;8:11933. doi: 10.1038/s41598-018-29646-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- López‐Aguirre C, Hand SJ, Koyabu D, Son NT, Wilson LA. Prenatal allometric trajectories and the developmental basis of postcranial phenotypic diversity in bats (Chiroptera) J Exp Zool B Mol Dev Evol. 2019;332:36–49. doi: 10.1002/jez.b.22846. [DOI] [PubMed] [Google Scholar]
- Lucas P. The dental-dietary adaptations of mammals. N Jb Palaont Mh. 1979;8:486–512. [Google Scholar]
- Lucas PW, Peters CR. Function of postcanine tooth crown shape in mammals. In: Teaford MF, Smith MM, Ferguson MWJ, editors. Development, Function and Evolution of Teeth. Cambridge University Press; Cambridge,: 2007. pp. 282–289. [Google Scholar]
- Maier JA, Rivas-Astroza M, Deng J, Dowling A, Oboikovitz P, Cao X, Behringer RR, Cretekos CJ, Rasweiler JJ, Zhong S, et al. Transcriptomic insights into the genetic basis of mammalian limb diversity. BMC Evol Biol. 2017;17:86. doi: 10.1186/s12862-017-0902-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin JF, Bradley A, Olson EN. The paired-like homeo box gene MHox is required for early events of skeletogenesis in multiple lineages. Genes Dev. 1995;9:1237–1249. doi: 10.1101/gad.9.10.1237. [DOI] [PubMed] [Google Scholar]
- Mason MK, Hockman D, Curry L, Cunningham TJ, Duester G, Logan M, Jacobs DS, Illing N. Retinoic acid-independent expression of Meis2 during autopod patterning in the developing bat and mouse limb. Evodevo. 2015;6:6. doi: 10.1186/s13227-015-0001-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Monteiro LR, Nogueira MR. Evolutionary patterns and processes in the radiation of phyllostomid bats. BMC Evol Biol. 2011;11:137. doi: 10.1186/1471-2148-11-137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Müller B, Goodman SM, Peichl L. Cone photoreceptor diversity in the retinas of fruit bats (Megachiroptera) Brain Behav Evol. 2007;70:90–104. doi: 10.1159/000102971. [DOI] [PubMed] [Google Scholar]
- Müller B, Glösmann M, Peichl L, Knop GC, Hagemann C, Ammermüller J. Bat eyes have ultraviolet-sensitive cone photoreceptors. PloS One. 2009;4:e6390. doi: 10.1371/journal.pone.0006390. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakatomi M, Wang XP, Key D, Lund JJ, Turbe-Doan A, Kist R, Aw A, Chen Y, Maas RL, Peters H. Genetic interactions between Pax9 and Msx1 regulate lip development and several stages of tooth morphogenesis. Dev Biol. 2010;340:438–449. doi: 10.1016/j.ydbio.2010.01.031. [DOI] [PubMed] [Google Scholar]
- Neuweiler G. Bau und Leistung des Flughundauges (Pteropus giganteus gig. Brünn.) Z Vergl Physiol. 1962;46:13–56. [Google Scholar]
- Nogueira MR, Peracchi AL, Monteiro LR. Morphological correlates of bite force and diet in the skull and mandible of phyllostomid bats. Funct Ecol. 2009;23:715–723. [Google Scholar]
- Norberg UM. Vertebrate flight: mechanics, physiology, morphology, ecology and evolution. Springer-Verlag; Berlin Heidelberg,: 2012. 308 [Google Scholar]
- Norberg UM, Rayner JMV. Ecological morphology and flight in bats (Mammalia; Chiroptera): Wing adaptations, flight performance, foraging Strategy and echolocation. PhilosTrans R Soc Lond B Biol Sci. 1987;316:335–427. [Google Scholar]
- Orr DJA, Teeling EC, Puechmaille SJ, Finarelli JA. Patterns of orofacial clefting in the facial morphology of bats: a possible naturally occurring model of cleft palate. J Anat. 2016;229:657–672. doi: 10.1111/joa.12510. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peichl L. Diversity of mammalian photoreceptor properties: Adaptations to habitat and lifestyle? Anat Rec. 2005;287a:1001–1012. doi: 10.1002/ar.a.20262. [DOI] [PubMed] [Google Scholar]
- Peters H, Neubuser A, Kratochwil K, Balling R. Pax9-deficient mice lack pharyngeal pouch derivatives and teeth and exhibit craniofacial and limb abnormalities. Genes Dev. 1998;12:2735–2747. doi: 10.1101/gad.12.17.2735. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Petit F, Sears KE, Ahituv N. Limb development: a paradigm of gene regulation. Nat Rev Genet. 2017;18:245–258. doi: 10.1038/nrg.2016.167. [DOI] [PubMed] [Google Scholar]
- Phillips CD, Butler B, Fondon JW, Mantilla-Meluk H, Baker RJ. Contrasting evolutionary dynamics of the developmental regulator PAX9, among bats, with evidence for a novel post-transcriptional regulatory mechanism. PLoS One. 2013;8:e57649. doi: 10.1371/journal.pone.0057649. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pointer MA, Kamilar JM, Warmuth V, Cheater SGB, Delsuc F, Mundy N, Asher RJ, Bradley BJ. RUNX2 tandem repeats and the evolution of facial length in placental mammals. BMC Evol Biol. 2012;12:103. doi: 10.1186/1471-2148-12-103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ray R, Capecchi M. An examination of the Chiropteran HoxD locus from an evolutionary perspective. Evol Dev. 2008;10:657–670. doi: 10.1111/j.1525-142X.2008.00279.x. [DOI] [PubMed] [Google Scholar]
- Rojas D, Mancina CA, Flores-Martinez JJ, Navarro L. Phylogenetic signal, feeding behaviour and brain volume in Neotropical bats. J Evol Biol. 2013;26:1925–1933. doi: 10.1111/jeb.12190. [DOI] [PubMed] [Google Scholar]
- Rossoni DM, Assis APA, Giannini NP, Marroig G. Intense natural selection preceded the invasion of new adaptive zones during the radiation of New World leaf-nosed bats. Sci Rep. 2017;7:11076. doi: 10.1038/s41598-017-08989-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sadier A, Davies KTJ, Yohe LR, Yun K, Donat P, Hedrick BP, Dumont ER, Davalos LM, Rossiter SJ, Sears KE. Multifactorial processes underlie parallel opsin loss in neotropical bats. Elife. 2018;7:e37412. doi: 10.7554/eLife.37412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Santagati F, Rijli FM. Cranial neural crest and the building of the vertebrate head. Nat Rev Neurosci. 2003;4:806–818. doi: 10.1038/nrn1221. [DOI] [PubMed] [Google Scholar]
- Santana SE, Geipel I, Dumont ER, Kalka MB, Kalko EKV. All you can eat: high performance capacity and plasticity in the common big-eared bat, Micronycteris microtis (Chiroptera: Phyllostomidae) PLoS One. 2011;6:e28584. doi: 10.1371/journal.pone.0028584. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sears KE, Behringer RR, Rasweiler JJ, Niswander LA. Development of bat flight: morphologic and molecular evolution of bat wing digits. Proc Natl Acad Sci U S A. 2006;103:6581–6586. doi: 10.1073/pnas.0509716103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sears KE, Goswami A, Flynn JJ, Niswander LA. The correlated evolution of Runx2 tandem repeats, transcriptional activity, and facial length in Carnivora. Evol Dev. 2007;9:555–565. doi: 10.1111/j.1525-142X.2007.00196.x. [DOI] [PubMed] [Google Scholar]
- Shi JJ, Westeen EP, Katlein NT, Dumont ER, Rabosky DL. Ecomorphological and phylogenetic controls on sympatry across extant bats. J Biogeogr. 2018;45:1560–1570. [Google Scholar]
- Simmons NB. Order Chiroptera. In: Wilson DE, Reeder DM, editors. Mammal Species of the World: A Taxonomic and Geographic Reference. Johns Hopkins University Press; Baltimore: 2005. pp. 312–529. [Google Scholar]
- Simmons NB, Geisler JH. Phylogenetic relationships of Icaronycteris, Archaeonycteris, Hassianycteris, and Palaeochiropteryx to extant bat lineages, with comments on the evolution of echolocation and foraging strategies in Microchiroptera. B Am Mus Nat Hist. 1998;235:1–182. [Google Scholar]
- Simmons NB, Seymour KL, Habersetzer J, Gunnell GF. Primitive Early Eocene bat from Wyoming and the evolution of flight and echolocation. Nature. 2008;451:818–821. doi: 10.1038/nature06549. [DOI] [PubMed] [Google Scholar]
- Simoes BF, Foley NM, Hughes GM, Zhao H, Zhang S, Rositer SJ, Teeling EC. As blind as a bat? Opsin phylogenetics illuminates the evolution of color vision in bats. Mol Biol Evol. 2019;36:54–68. doi: 10.1093/molbev/msy192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simpson GG. Tempo and mode in evolution. Columbia University Press; New York: 1944. 217 [Google Scholar]
- Smith TD, Eiting TP, Bhatnagar KP. A quantitative study of olfactory, non-olfactory, and vomeronasal epithelia in the nasal fossa of the bat Megaderma lyra. J Mammal Evol. 2012;19:27–41. [Google Scholar]
- Speakman JR, Racey PA. No cost of echolocation for bats in flight. Nature. 1991;350:421–423. doi: 10.1038/350421a0. [DOI] [PubMed] [Google Scholar]
- Steeman ME, Hebsgaard MB, Fordyce RE, Ho SYW, Rabosky DL, Nielsen R, Rahbek C, Glenner H, Sorensen MV, Willerslev E. Radiation of extant cetaceans driven by restructuring of the oceans. Syst Biol. 2009;58:573–585. doi: 10.1093/sysbio/syp060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Swartz S, Groves M, Kim H, Walsh W. Mechanical properties of bat wing membrane skin. J Zool. 1996;239:357–378. [Google Scholar]
- Swartz SM, Freeman PW, Stockwell EF. Ecomorphology of bats: Comparative and experimental approaches relating structural design to ecology. In: Kunz TH, Fenton MB, editors. Bat Ecology. University of Chicago Press; Chicago: 2003. pp. 257–300. [Google Scholar]
- Teeling EC, Scally M, Kao DJ, Romagnoli ML, Springer MS, Stanhope MJ. Molecular evidence regarding the origin of echolocation and flight in bats. Nature. 2000;403:188–192. doi: 10.1038/35003188. [DOI] [PubMed] [Google Scholar]
- Teeling EC, Springer MS, Madsen O, Bates P, O'brien SJ, Murphy WJ. A molecular phylogeny for bats illuminates biogeography and the fossil record. Science. 2005;307:580–584. doi: 10.1126/science.1105113. [DOI] [PubMed] [Google Scholar]
- Teeling EC, Dool S, Springer MS. Phylogenies, fossils and functional genes: the evolution of echolocation in bats. In: Gunnell GF, Simmons NB, editors. Evolutionary History of Bats: Fossils, Molecules and Morphology. 2012. pp. 1–22. [Google Scholar]
- Teeling EC, Vernes SC, Dávalos LM, Ray DA, Gilbert MTP, Myers E, Bat1K Consortium Bat biology, genomes, and the Bat1K Project: To generate chromosome-level genomes for all living bat species. Annu Rev Anim Biosci. 2018;6:23–46. doi: 10.1146/annurev-animal-022516-022811. [DOI] [PubMed] [Google Scholar]
- Tokita M, Abe T, Suzuki K. The developmental basis of bat wing muscle. Nat Commun. 2012;3:1–9. doi: 10.1038/ncomms2298. [DOI] [PubMed] [Google Scholar]
- Tsagkogeorga G, Parker J, Stupka E, Cotton JA, Rossiter SJ. Phylogenomic analyses elucidate the evolutionary relationships of bats. Curr Biol. 2013;23:2262–2267. doi: 10.1016/j.cub.2013.09.014. [DOI] [PubMed] [Google Scholar]
- Usui K, Tokita M. Creating diversity in mammalian facial morphology: a review of potential developmental mechanisms. Evodevo. 2018;9:15. doi: 10.1186/s13227-018-0103-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Veilleux CC, Louis EE, Bolnick DA. nocturnal light environments influence color vision and signatures of selection on the OPN1SW Opsin gene in nocturnal lemurs. Mol Biol Evol. 2013;30:1420–1437. doi: 10.1093/molbev/mst058. [DOI] [PubMed] [Google Scholar]
- Wang D, Oakley T, Mower J, Shimmin LC, Yim S, Honeycutt RL, Tsao H, Li W-H. Molecular evolution of bat color vision genes. Mol Biol Evol. 2004;21:295–302. doi: 10.1093/molbev/msh015. [DOI] [PubMed] [Google Scholar]
- Wang Z, Dong D, Ru B, Young RL, Han N, Guo T, Zhang S. Digital gene expression tag profiling of bat digits provides robust candidates contributing to wing formation. BMC Genomics. 2010;11:619. doi: 10.1186/1471-2164-11-619. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Z, Dai M, Wang Y, Cooper KL, Zhu T, Dong D, Zhang J, Zhang S. Unique expression patterns of multiple key genes associated with the evolution of mammalian flight. Proc Biol Sci. 2014;281:20133133. doi: 10.1098/rspb.2013.3133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weatherbee SD, Behringer RR, Rasweiler JJ, Niswander LA. Interdigital webbing retention in bat wings illustrates genetic changes underlying amniote limb diversification. Proc Natl Acad Sci U S A. 2006;103:15103–15107. doi: 10.1073/pnas.0604934103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wetterer AL, Rockman MV, Simmons NB. Phylogeny of phyllostomid bats (Mammalia: Chiroptera): data from diverse morphological systems, sex chromosomes, and restriction sites. Bull Am Mus Nat Hist. 2000;248:1–200. [Google Scholar]
- Wilkinson GS, Adams DM. Recurrent evolution of extreme longevity in bats. Biol Lett. 2019;15:20180860. doi: 10.1098/rsbl.2018.0860. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilkinson GS, South JM. Life history, ecology and longevity in bats. Aging Cell. 2002;1:124–131. doi: 10.1046/j.1474-9728.2002.00020.x. [DOI] [PubMed] [Google Scholar]
- Winter Y, Lopez J, von Helversen O. Ultraviolet vision in a bat. Nature. 2003;425:612–614. doi: 10.1038/nature01971. [DOI] [PubMed] [Google Scholar]
- Wu JW, Jiao HW, Simmons NB, Lu Q, Zhao HB. Testing the sensory trade-off hypothesis in New World bats. Proc R Soc B. 2018;285:20181523. doi: 10.1098/rspb.2018.1523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yakushiji-Kaminatsui N, Lopez-Delisle L, Bolt CC, Andrey G, Beccari L, Duboule D. Similarities and differences in the regulation of HoxD genes during chick and mouse limb development. PLoS Biol. 2018;16:e3000004. doi: 10.1371/journal.pbio.3000004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yoder JB, Clancey E, Des Roches S, Eastmen JM, Gentry L, Godsoe W, Hagey TJ, Jochimsen D, Oswalk BP, Robertson J, et al. Ecological opportunity and the origin of adaptive radiations. J Evol Biol. 2010;23:1581–1596. doi: 10.1111/j.1420-9101.2010.02029.x. [DOI] [PubMed] [Google Scholar]
- Yohe LR, Abubakar R, Giordano C, Dumont E, Sears KE, Rossiter SJ, Davalos LM. Trpc2 pseudogenization dynamics in bats reveal ancestral vomeronasal signaling, then pervasive loss. Evolution. 2017;71:923–935. doi: 10.1111/evo.13187. [DOI] [PubMed] [Google Scholar]
- Yohe LR, Brand P. Evolutionary ecology of chemosensation and its role in sensory drive. Curr Zool. 2018;64:525–533. doi: 10.1093/cz/zoy048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yohe LR, Velazco PM, Rojas D, Gerstner BE, Simmons NB, Davalos LM. Bayesian hierarchical models suggest oldest known plant-visiting bat was omnivorous. Biol Lett. 2015;11:20150501. doi: 10.1098/rsbl.2015.0501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yokoyama S. Molecular evolution of vertebrate visual pigments. Prog Retin Eye Res. 2000;19:385–419. doi: 10.1016/s1350-9462(00)00002-1. [DOI] [PubMed] [Google Scholar]
- Yokoyama S, Radlwimmer FB. The molecular genetics of red and green color vision in mammals. Genetics. 1999;153:919–932. doi: 10.1093/genetics/153.2.919. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao HB, Rossiter SJ, Teeling EC, Li CJ, Cotton JA, Zhang SY. The evolution of color vision in nocturnal mammals. Proc Natl Acad Sci U S A. 2009;106:8980–8985. doi: 10.1073/pnas.0813201106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou J, Gao Y, Lan Y, Jia SH, Jiang RL. Pax9 regulates a molecular network involving Bmp4, Fgf10, Shh signaling and the Osr2 transcription factor to control palate morphogenesis. Development. 2013;140:4709–4718. doi: 10.1242/dev.099028. [DOI] [PMC free article] [PubMed] [Google Scholar]



