Abstract
Biomedical imaging tools permit investigation of molecular mechanisms across spatial scales, from genes to organisms. Drosophila melanogaster, a well-characterized model organism, has benefited from the use of light and electron microscopy to understand gene function at the level of cells and tissues. The application of imaging platforms that allow for an understanding of gene function at the level of the entire intact organism would further enhance our knowledge of genetic mechanisms. Here a whole animal imaging method is presented that outlines the steps needed to visualize Drosophila at any developmental stage using microcomputed tomography (μ-CT). The advantages of μ-CT include commercially available instrumentation and minimal hands-on time to produce accurate 3D information at micron-level resolution without the need for tissue dissection or clearing methods. Paired with software that accelerate image analysis and 3D rendering, detailed morphometric analysis of any tissue or organ system can be performed to better understand mechanisms of development, physiology, and anatomy for both descriptive and hypothesis testing studies. By utilizing an imaging workflow that incorporates the use of electron microscopy, light microscopy, and μ-CT, a thorough evaluation of gene function can be performed, thus furthering the usefulness of this powerful model organism.
Introduction
Imaging methods that allow for the detailed investigation of interior structures of an object without destroying its overall 3D architecture have proven to be widely beneficial to a number of different disciplines, including physics, engineering, materials science, archaeology, paleontology, geology, and biology1, 2, 3, 4, 5, 6, 7, 8, 9. Among these nondestructive imaging methods, X-ray based platforms are especially useful due to the ability of high energy X-rays to penetrate many different sample types and materials with minimal scattering compared to visible light waves. Computed Tomography (CT), Microcomputed Tomography (μ-CT), Nanocomputed Tomography (Nano-CT), and Synchrotron Microtomography have, therefore, emerged as the primary technologies for X-ray based imaging of samples ranging from meters to microns, with millimeter to sub-micron resolution capabilities10, 11, 12, 13, 14.
While these platforms differ in their design, X-ray geometry, and components in order to balance sample size and resolution, they all rely on the same basic principle for image capture: a source of X-rays that travel through the object and are captured by a detector. Differential attenuation of the X-ray beam as it passes through varying densities within the object generates image contrast. 3D data is obtained by rotating either the sample or the detector, collecting a series of 2D projection images that are then reconstructed using algorithms into tomograms containing 3D information whose resolution is isotropic in x,y,z15. For many benchtop μ-CT scanners that utilize a cone-beam X-ray geometry to project X-rays at the object being imaged, the Feldkamp algorithm is used to accurately reconstruct the object with minimal errors16.
Resolution of a given platform is determined primarily by system parameters such as the size of the X-ray beam (spot size), scanner geometry (distance from object to X-ray source), size of the pixels on the detector, and the reconstruction algorithm employed. Additional factors, such as scanner vibrations, X-ray beam fluctuations, sample movement, and material type or chemical stain used to visualize the object can also significantly influence spatial resolution under real world imaging condidtions15.
For biomedical applications, CT and μ-CT have played a key role in advancing our understanding of anatomy, physiology, development, and disease mechanisms, serving as a tool for both human patient diagnoses and as a preclinical imaging platform for model organisms17, 18. For example, the Mouse International Phenotyping Consortium, whose goal is to identify the function of every gene in the mouse genome, utilizes μ-CT as part of their phenotyping pipeline19. Their results have been critical for understanding genes involved in development and disease processes, while also serving as an atlas for mouse anatomy and development20. Other model organisms, such as zebrafish and rats, have also fully embraced the use of μ-CT for performing whole animal phenotyping of a number of gene mutants17, 21, 22, 23.
The advantage of combining whole animal imaging with model organisms is that a mechanistic understanding of gene function for a given biological process can be fully explored. This is possible because of the well-characterized genomes and many genetic tools available in model organisms that allow for precise manipulation of gene function at distinct developmental timepoints, specific tissues, individual cells, and even subcellular organelles. These include binary expression systems such as the UAS/GAL4 system (and its many derivatives), CRISPR/Cas9, and RNAi24, 25, 26. When these genetic tools are used in conjunction with a powerful imaging pipeline consisting of electron microscopy, light microscopy (fluorescent and non-fluorescent), and whole animal imaging such as μ-CT, a thorough evaluation of molecules, cells, tissues, organs, and the entire organism can be achieved, allowing for a much deeper understanding of gene function.
This protocol focuses on the use of μ-CT in the non-mammalian model organism Drosophila melanogaster, whose myriad genetic tools have helped elucidate numerous molecular mechanisms26, 27. It was adopted from previous protocols in non-model insects1, 28, 29, 30, 31, 32, and builds off of previous μ-CT studies in Drosophila to establish a standardized protocol for its use in this animal33, 34, 35, 36, 37, 38, 39, 40, 41. The steps for successful sample preparation, imaging and analysis of fly μ-CT datasets using commercially available scanners are outlined. With this protocol, all developmental stages of the fly can be visualized at high resolution for both descriptive and hypothesis-testing studies, including taxonomy, anatomy, development, physiology, and disease27. This protocol will also be useful for imaging virtually any insect and even non-living materials that require chemical staining for image contrast to enhance visualization by μ-CT.
Protocol
1. Sample selection and cuticle preparation
- Choose the appropriate developmental timepoint (embryo, larva, pupa, or adult) for imaging.
- For embryonic stages, cage females on grape juice agar plates smeared with yeast paste and collect eggs every 30–60 min. Leave these embryos to develop at 25 °C until the proper stage is reached.
- For larval stages, collect first and second instars from timed embryo collection experiments. Directly pick wandering 3rd instars from the side of the food vial under non-crowding conditions.
- For pupal timing, collect white pre-pupae (inverted spiracles) and make note of the time when the cuticle starts to brown. Animals will progress through 15 stages of pupal development at defined timepoints following cuticle browning and can be collected accordingly42.
- Collect adults as virgins following eclosion and allow to age in a food vial for a required amount of time (e.g., 5 days for complete gut development).
- Transfer 5–50 animals to a 1.5 mL microcentrifuge tube containing 1 mL of 0.5% Triton X-100 in 1x Phosphate Buffered Saline (0.5% PBST). While tapping the tube periodically on the benchtop, incubate for 5 min at room temperature (RT) to assist in the removal of the hydrophobic coating and allow animals to become fully submerged.
-
For embryonic, larval, and pupal stages, arrest further development by placing the tube in a heat block set to 100 °C for 20 s followed by cooling at RT for 5 min.NOTE: Animals can also be flash frozen in liquid nitrogen37.
-
2. Fixation and staining
-
Remove 0.5% PBST and add 1 mL of Bouin’s Solution. Tap tube to ensure animals are fully submerged.
CAUTION: Bouin’s Solution contains formaldehyde. It may cause acute toxicity to skin and eyes if spilled and can be fatal if swallowed. Wear gloves, safety glasses and a lab coat when handling.
NOTE: Additional fixation techniques can also be employed, such as the use of ethanol. The merits of other fixatives are described in detail in ref.1, 30.- For embryo and adult samples, incubate on the benchtop for 16–20 h at RT.
- For larval and pupal samples, fix animals for 2 h at RT. Discard Bouin’s solution and wash with 1x PBS for 5 min thrice. Transfer to a multi-well dissecting dish containing 1x PBS and use a small minutien pin attached to a holder to poke a hole in the anterior and posterior cuticle being careful to not disrupt any underlying soft tissue.
- Transfer larval and pupal samples back to a microfuge tube and add 1 mL of fresh Bouin’s solution. Incubate on benchtop for 24 h at RT.
Remove Bouin’s Solution and wash sample for 30 min with 1 mL of μ-CT Wash Buffer (0.1 M Na2HPO4, 1.8% Sucrose) or 1x PBS thrice.
-
Add 1 mL of the appropriate staining solution and incubate on benchtop for 2–7 days.
NOTE: Stain choice will depend on many factors, but is generally a balance between penetration, incubation time, and resolution. In general, phosphotungstic acid (PTA) provides superior contrast and resolution of soft tissue but requires mechanical disruption of the cuticle and longer incubation times. The merits of different stain types are described in detail in ref.1.- For iodine staining, use 1 mL of 0.1 N I2KI (Lugol Solution). Incubate for 2 days. No further disruption of the adult cuticle is necessary.
- For phosphotungstic acid (PTA) staining, use 1 mL of a 0.5% (w/v) solution diluted in water. Disrupt the adult cuticle, either by removing the mouthparts or poking holes in the thorax or abdominal cuticle with a small minutien pin attached to a holder. Incubate for 5–7 days, or longer if tissue staining appears non-homogenous.
Wash with 1 mL of ultrapure water or 1x PBS for 30 min. Repeat wash step. Store animals at RT in ultrapure water or 1x PBS for up to one month.
If animals are to be scanned while hydrated, proceed directly to section 4. If longer preservation of the sample is needed, proceed to section 3 of the protocol.
3. Critical point drying (Optional)
Perform an ethanol (EtOH) dehydration series on the sample: 10%, 25%, 50%, 75%, 100%, 100%. Add 1 mL of the EtOH solution and incubate on the benchtop for 1 h for each concentration in the order stated.
After the final 100% EtOH soak, replace with fresh 100% EtOH and let sample incubate on the benchtop overnight.
-
Perform critical point drying of samples following the manufacturer’s instructions.
NOTE: Electron microscopy facilities generally have a critical point drying machine (see Table of Materials) that will perform the drying of the sample following EtOH dehydration.
Materials.
| Name | Company | Catalog Number | Comments |
|---|---|---|---|
| 100% Ethanol | For critical point drying | ||
| Bouin’s Solution | Sigma-Aldrich | HT10132 | For animal fixation |
| Critical Point Dryer | Dries samples using the critical point method; multiple options available (Balzers CPD 020 or Leica EMCPD300) | ||
| Dragonfly Software | Object Research Systems | For visualization and segmentation of micro-CT datasets; https://www.theobjects.com/dragonfly/index.html | |
| Heat Block | For microfuge tubes | ||
| Image Analysis Workstation | Should contain sufficient RAM and quality graphics card for 3D rendering | ||
| Iodine Solution (I2KI) | Fisher Scientific | SI86-1 | For staining |
| Microcomputed Tomography Scanner | Bruker | Skyscan 1172 | Cone-beam X-Ray geometry; detector is a Hamamatsu 10 MP camera with 11.54 μm pixel size. |
| Microcomputed Tomography Scanner Software | Bruker | For controling the scanner itself (e.g., performing flat field corrections, X-ray tube power, camera expsoure times, acquisition, etc.) | |
| Minutien Pins | Fine Science Tools | 26002-15 | For poking hole in cuticle |
| NRecon Image Reconstruction Software | Bruker | Used to reconstruct cross-section images from 2D projection images taken with cone-beam X-Ray geometry | |
| P10 pipet tips | Genesee Scientific | 24-120 | Sample mounting |
| Phosphate Buffered Saline | Resarch Products International | P32060-4000.0 | Dilute to 1X with water before use |
| Phosphotungstic Acid Hydrate | Sigma-Aldrich | 79690-25g | For staining |
| Pin Holder | Fine Science Tools | 26018-17 | For Minutien Pins |
| Triton X-100 | Research Products International | 111036 | To remove waxy coating from adult flies (as 0.5% PBST) |
| X-Ray Microscope | Zeiss | Xradia 520 Versa | Cone-beam X-Ray geometry featuring Fresnel zone plate objective lenses for Resoluton at a Distance (RaaD™) |
4. Sample mounting
For critically point dried samples, hot glue samples to an insect pin or other holder designed to fit within the chuck of the rotating stage or place it in a plastic or glass capillary tube (~1.0–1.25 mm inner diameter).
-
For hydrated samples, fill a P10 pipette tip with water and secure the narrow end by either heat sealing or paraffin film to prevent leakage.
CAUTION: Be sure to wrap any connections tightly so that water does not leak into the scanner and cause damage.- Transfer a single specimen to the pipette tip using forceps. Using a long, slender object, such as a dulled 20 G needle or another pipette tip, carefully push the specimen down the tip until it just contacts the wall of the pipet tip to hold it in place.
- Cover the opening of the pipette tip with paraffin film to prevent evaporation of water during long scans.
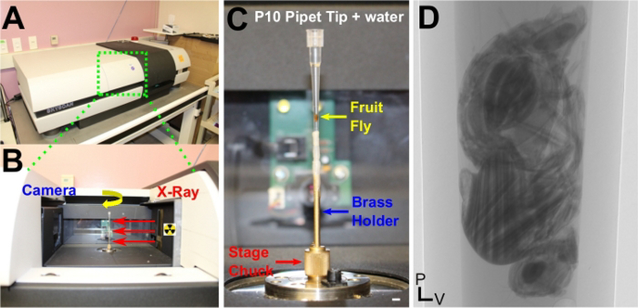
Figure 1: Overview of scanner design and sample mounting for μ-CT.
(A) A commercial benchtop μ-CT scanner. (B) View inside the scanner. The X-ray source (right) emits X-Rays that pass through the sample located on a rotating stage (yellow arrow). Attenuation of these X-rays generate image contrast as they pass through the sample and onto the detector, which consists of a scintillation screen that converts X-rays to photons and a standard CCD camera (left). (C) Mounting an adult fruit fly for hydrated imaging in water. The connection points between the pipette tip and the brass holder are wrapped in paraffin film to prevent leakage and potential damage to the scanner. The stage chuck is also highlighted. Note that the pipette tip was positioned slightly off-axis, which led to a longer scan time and reduced resolution in the final reconstruction. (D) A single 2D projection image of an adult female fly; hundreds to thousands of these projections are acquired during a scan along the rotation axis and are used for reconstruction to generate tomograms containing isotropic resolution and accurate 3D information. Scale Bars (C) = 2 mm. P, Posterior; V, Ventral. This figure has been modified from Schoborg et al.40..
5. Scanning
-
Perform any necessary calibrations of the machine prior to imaging for the day.
NOTE: These will vary by manufacturer and it is recommended to consult with an application specialist to determine the proper steps. Please see the Table of Materials for specific information on the setup and software used in this protocol. Generally, calibrations such as a stage axis alignment to remove any ‘wobble’ associated with the chuck being off-axis relative to the rotating stage and performing flat-field corrections to ensure uniform background pixel intensities on the camera provide optimal imaging conditions for the best resolution and datasets that are comparable across multiple scans.
-
Open the scanner door to gain access to the rotating stage chuck by clicking on the ‘Open Door’ icon in the upper left corner of the software. Attach the sample by tightening the collar around the base of the sample holder.
NOTE: Use gentle pressure when attaching the sample to the rotating chuck to maintain scanner alignment.
-
Set scanning parameters in the software controlling the scanner for optimal resolution and contrast.
NOTE: These will need to be determined empirically as the X-ray source, camera/detector, and geometry of each scanner will vary by manufacturer. Additional information for selecting the best parameters can also be found here43, as well as from the manufacturer’s application specialist.- Open the X-Ray source power control (Options | X-ray Source). Use the slider bars to set X-ray voltage to 30–40 kV and current to 100–110 μA to produce an X-ray beam with 3–4W of power and a small spot size for enhanced resolution.
- Use the Acquisition Modes dialog (Options | Acquisition Modes) to set camera exposure time to 500–800 ms.
- Use the slider bar located next to the magnifying glass icon at the bottom of the software to set the desired image pixel size (~700 nm to 4 μm), depending on camera settings and position. This determines the number of projection images that are acquired during the scan, with more projections leading to enhanced resolution but longer scan times (see Representative Results).
- Click and drag the slider bar located next to the rotating arrow at the bottom of the software to move the sample along its 360° rotation path. Ensure that the sample stays within the field of view.
- Click on the ‘Start Acquisition’ icon (blue circle arrow icon) at the top of the software. A dialog box appears that allows additional scanning parameters to be set, and the file and output folder where the scan will be saved to be named.
- Set the random movement to 10 and average 4–6 frames. The rotation step is automatically calculated depending on the camera settings used.
Begin scan by clicking ‘OK’ on the Acquisition dialog box. A second progress bar dialog appears that shows the scan time. The scanner will now acquire a series of projection images (Figure 1D) of the specimen along the rotation path and does not need to be monitored.
6. Reconstruction
-
To generate the tomograms, perform image reconstruction using the projection images.
NOTE: While almost all reconstruction of images from cone beam geometry scanners rely on the Feldkamp algorithm16, individual parameters will vary depending on the software implementation and should be determined empirically. The following settings, specific for a commercially available software (see Table of Materials) can be used as a guide. Parameters such as misalignment compensation, ring artifact reduction, and beam hardening correction (0% for most fly samples) are performed in an iterative fashion to choose the best values for the final reconstruction. For advanced users who would like more control over the reconstruction process, see the MATLAB interface here44.
Perform an initial image alignment by utilizing a shift-correction algorithm based on reference scans45 (Actions | X/Y alignment with a reference scan).
- Fine tune each reconstruction parameter (Start | Fine tuning). This activates a ‘Parameter fine-tuning’ dialog box.
-
Fine tune the alignment by clicking ‘Next’ to ‘PostAlignment’. Set the number of iterations to five and the parameter step to 1.0. Click the Start button to generate a series of previews. Select the image that is properly aligned.NOTE: A properly aligned image will not be blurry or display ‘shearing’ artifacts where an otherwise continuous structure (such as the cuticle) appears split.
- Fine tune the ring artifact reduction by clicking next to it. Set the number of iterations to five and the parameter step to 1.0. Click the Start button to generate a series of previews. Select the image that contains the fewest number of rings.
-
Once the above parameters have been optimized to give the best image, ensure that the selected values are properly represented in the Settings tab (Settings).
Adjust any final brightness and contrast values using the histogram, file parameters such as bit depth and type, and utilize a Region of Interest (ROI) encompassing only the structures of interest (e.g., whole fly or head only) to reduce computational time (Output).
Begin the reconstruction (Start | Start). If multiple reconstructions are required, add the current image to the batch manager (Start | Add to batch) and repeat steps 6.2–6.5 for the remaining images.
7. Image analysis
-
Visualize tomograms in two and three dimensions and perform further morphometric analysis with freeware or commercial software.
NOTE: Details of the software used in this protocol is available in the Table of Materials. Other software packages that are capable of evaluating μ-CT datasets include freeware options such as FIJI46, Seg3D (www.seg3D.org)47, and ITK-SNAP48, plus commercial software (e.g., AMIRA). Other applications that employ machine-learning based algorithms to semi-automate the segmentation process can help speed workflows and reduce human bias (e.g., Biomedisa49).
Import the tomogram file into the software (File | Import Image Files). The metadata associated with the file should automatically load into the window but can also be set manually to match the image properties.
To segment a structure of interest, click on the Segment tab on the left-hand side of the screen. Create a new Region of Interest (ROI) (Basic | New), give it a name and select an appropriate color.
Define the threshold range that encompasses the structure of interest (Range | Define Range) using the slider bar.
Select an appropriate 2D or 3D ROI painter tool mode (ROI Painter | Paintbrush option).
-
To paint an area defined by the threshold; press and hold the Ctrl key while holding the left mouse key. To remove an area, press and hold the Shift key while holding the left mouse key.
NOTE: To change the size of circular or square paintbrush, simply scroll the mouse wheel while holding either the Ctrl or Shift keys.
Continue to paint the structure of interest throughout its entire Z-volume.
Convert the ROI into a Mesh (Export | To a Mesh | Normal).
Highlight the mesh overlay by clicking on it in the Data Properties and Settings panel in the upper right-hand corner.
-
Smooth the mesh using an appropriate number of iterations (Smooth Mesh | Iterations).
NOTE: Use the same mesh smoothing iteration value for all images that are to be compared.
Ensure that the measurements of the mesh ROI (Surface Area, Volume and Feret Diameter) are displayed in the Information panel on the right-hand side of the screen for basic morphometric analysis. Additional analysis can be performed using the Object Analysis Module.
-
Render the mesh object and the entire tomogram image and visualize in 3D. Use the built-in Movie Maker to generate a video of the object (Right Mouse Click | Show Movie Maker) using individual frames from the viewer (Add Key).
NOTE: Several visual enhancements can also be applied to the movie using the visual effects tab in the right-hand panel to highlight certain features, etc.
Export the video for viewing using the Export Animation button in the Movie Maker.
Representative Results
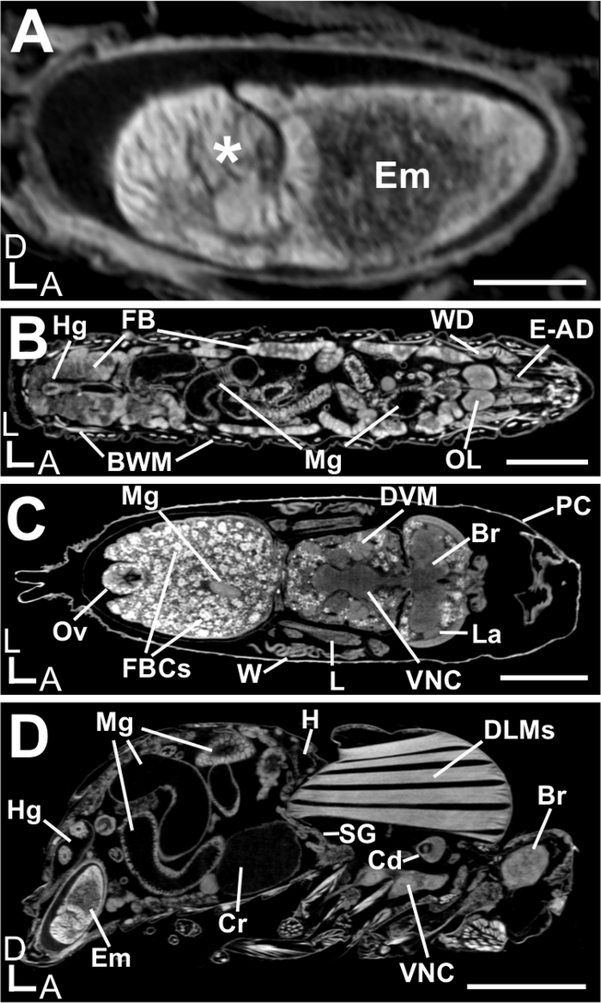
Figure 2 shows images of an embryo, 3rd instar larvae, pupae at the pharate adult stage (P7), and an adult female fly stained with iodine and imaged hydrated in water using a commercial benchtop scanner. Excellent preservation and even staining of delicate tissue are apparent, allowing all major organs to be readily identified and used for morphometric analysis and 3D visualization.
Figure 2: ll Drosophila melanogaster life cycle stages, imaged by μ-CT.
A Samples stained with iodine and imaged hydrated in water. Shown is a single 2D slice. (A) An embryo that has completed the early stages of gastrulation (asterisk). (B) A third instar larva. (C) A P7 pharate adult during metamorphosis. (D) An adult female. Various organs are highlighted: BWM, body wall muscles; Br, brain; Cd, cardia; Cr, crop; DLMs, dorsal longitudinal muscles; DVM, dorsal ventral muscles; E-AD, eye-antennal disc; Em, embryo; FB, fat bodies; FBCs, fat body cells; H, heart; Hg, hindgut; La, lamina; L, leg; Mg, midgut; OL, brain optic lobe; Ov, ovipositor; PC, pupal cuticle; SG, salivary glands; VNC, ventral nerve cord; W, wing; WD, wing disc. Scale Bars (A) = 100 μm; (B)-(D) = 500 μm. D, Dorsal; A, Anterior; L, Left. Scanning parameters: Source to Sample Distance (mm): (A, D) 36.5, (B) 48.8, (C) 40.3. Source to Camera Distance (mm): (A, D) 350, (B, C) 285. Camera Pixel Size (μm): (A-D) 11.6. Image Pixel Size (μm): (A, D) 1.2, (B) 1.9, (C) 1.7.
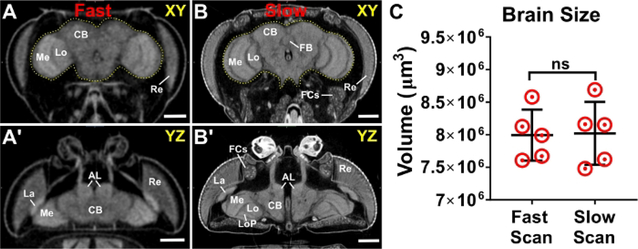
Typically, scans that acquire fewer projection images of the specimen provide lower resolution than scans that acquire more projection images, with the tradeoff being time spent scanning. Although scan times will vary by instrument and other scanning parameters, scans that acquire a few hundred projections (~3 μm image pixel size) takes roughly 30 min per specimen, whereas scans consisting of thousands of projection images (700 nm-1.25 μm image pixel size) can take 8–16 h. A comparison of the same adult fly headcase taken in both ‘slow’ (thousands of projections) and ‘fast’ (hundreds of projections) scan mode is shown in Figure 3. Importantly, morphometric analyses do not differ between ‘slow’ and ‘fast’ scans (Figure 3C)40. Our imaging pipeline, therefore, utilizes fast scans to generate a sufficiently large sample size for morphometric analysis, and slow scans to visualize any morphological or anatomical defects at higher resolution. Using the software (Step 7), any tissue or organ system of interest can be segmented and used for morphometric analysis and visualized in 3D using the movie maker (Movie 1).
Figure 3: Scanning parameters and image resolution do not alter morphometric analyses.
An adult head scanned using both (A, A’) ‘fast’ scanner settings (hundreds of projections) and (B, B’) ‘slow’ scanner settings (thousands of projections). The brain is outlined in yellow. (C) Brain volume measurements from slow and fast scans. Highlighted structures: AL; antennal lobe; CB, central brain; FB, fan shaped body; FCs, fat cells; La, lamina; Lo, lobula; LoP, lobula plate; Me, medulla; Re, retina. n = 5, Welch’s t-test. ns = not significant. Scale bars = 100 μm. Scanning parameters: Source to Sample Distance (mm): (A) 44.4, (B) 36.5. Source to Camera Distance (mm): (A) 348 (B) 350. Camera Pixel Size (μm): (A-B) 11.6. Image Pixel Size (μm): (A) 2.95, (B) 1.2. This figure has been modified from Schoborg et al.40..
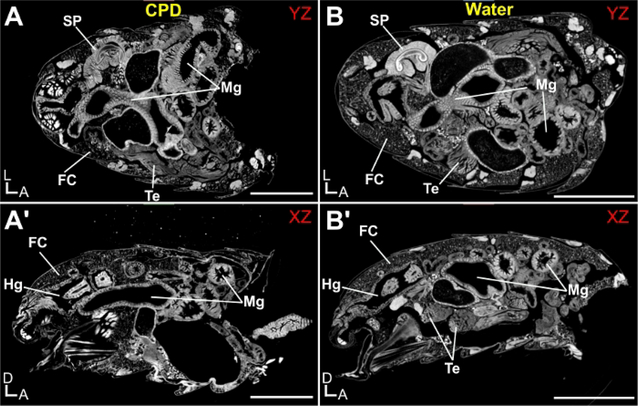
Figure 4 shows an example of the fly abdomen stained with PTA and imaged hydrated (water) or following critical point drying (CPD) on an X-ray microscope (Table of Materials). The detail afforded by a combination of the PTA and the capabilities of this platform is readily evident in these images, such that individual epithelia cells of the midgut and sperm bundles within the testes are easily resolved. While the CPD image shows marginally increased resolution compared to the hydrated sample, better preservation of the ultrastructure of delicate tissues (such as the fat cells near the cuticle) is achieved with hydrated samples (Movie 2).
Figure 4: Drosophila melanogaster abdomen imaged by X-ray Microscopy.
Abdomens were stained with 0.5% PTA and imaged hydrated (water) or following critical point drying (CPD). (A) Critical Point Dried abdomen, shown from the YZ perspective and (A’) XZ perspective. (B) Hydrated abdomen, shown from the YZ perspective and (B’) XZ perspective. Various organs are highlighted: FC, fat cells; Hg, hindgut; Mg, Midgut; SP, Sperm Pump; Te, Testes. Scale Bars (A) = 250 μm. D, Dorsal; A, Anterior; L, Left. Scanning parameters: Source to Sample Distance (mm): (A) 6.7, (B) 7. Source to Camera Distance (mm): (A) 28 (B) 29.5. Objective: (A-B) 4X. Image Pixel Size (μm): (A-B) 0.65.
Movie 1: A third instar larva, rendered in 3D using the Movie Maker in Dragonfly. Highlighted organs include the brain (yellow), eye-antennal imaginal discs (red), fat body (blue) and the body wall muscles (green).
Movie 2: Comparison of samples imaged in water or follow critical point drying. Abdomens stained with 0.5% PTA are shown. Both abdomens were scanned with identical image pixel size settings (0.65 μm). A series of 2D slices are shown in a ‘Z-stack’ format (YZ) starting at the dorsal surface and ending at the ventral surface of the abdomen. Organs highlighted: FC, fat cells; Mg, Midgut; SP, Sperm Pump; Te, Testes. Scanning parameters: Source to Sample Distance (mm): (A) 6.7, (B) 7. Source to Camera Distance (mm): (A) 28 (B) 29.5. Objective: (A-B) 4X. Image Pixel Size (μm): (A-B) 0.65.
Discussion
Visualizing intact Drosophila melanogaster at all developmental stages has remained a challenge, primarily due to the incompatibility of light microscopy with the thick, pigmented cuticle found in this animal. While other whole animal imaging methods, such as Magnetic Resonance Imaging (MRI), Optical Coherence Tomography (OCT), and ultramicroscopy coupled with tissue clearing have been used with success in flies50, 51, 52, 53, 54, μ-CT presents a number of advantages that make it ideal for whole animal imaging of this organism13, 15, 30. X-rays easily penetrate the pigmented cuticle and their small wavelength allows for sub-micron imaging. Labeling requires minimal investment in widely available chemicals and no specialized bench skills13. μ-CT scanners are also commercially available, and costs are comparable to light microscopy platforms, while also being more attractive to wider range of disciplines (Geology, Paleontology, Engineering, etc.) that can also benefit from its availability at an institution. Synchrotron X-Ray sources can also be used for high resolution μ-CT imaging of fixed and living insects31, 55, 56, but are less accessible than commercial benchtop scanners.
This protocol provides an efficient way to obtain μ-CT images of fly adults, pupa, larva and cellularized embryos. Note that for many of the steps outlined above, alternative methods can also be applied to prepare samples for imaging. Other studies have provided a detailed comparison of different fixation, labeling, and drying steps for use in insects and those interested in adopting this technique are encouraged to evaluate the merits of each approach1, 4, 13, 29, 30, 57. While this protocol is relatively straightforward, a few helpful suggestions are presented.
First, care should be taken when disrupting the cuticle of intact specimens such that underlying soft tissues are not significantly disrupted. It is important to let larval and early pupal stages undergo fixation for 2 hours in Bouin’s solution before poking. This will stiffen the tissue and limit the amount of hemolymph that will ooze out of the cuticle holes, which can alter organ architecture. Individual body segments (head, thorax and abdomen) of the adult can be separated if the structure(s) of interest are located there. It is recommended to use a scalpel to cleanly slice through these segments rather than pulling them apart with forceps, which could disrupt the 3D architecture of the gut or central nervous system, for example. As for timing, adults generally need only 16 hrs. for complete fixation, whereas larval and pupal stages need 24 h. Also, if iodine or PTA staining appears uneven, the sample can be placed back in solution to incubate longer until even staining is achieved. Finally, hydrated samples should not be placed at 4 °C, as this seems to induce the formation of air bubbles within the body cavity after warming to room temperature.
Second, sample mounting will vary by instrument, stage type and whether the sample needs to remain hydrated or has been critical point dried. If hydrated, ensure the sample does not leak and possibly destroy the scanner. When mounting the sample inside a pipette tip, be sure to push gently with a dulled object until the specimens encounters slight resistance and can’t move. Pushing too hard can lead to cuticle deformation and underlying structural defects. Also, be sure that the sample is aligned in the holder as close to the axis of rotation as possible. Any wobble will increase scan times due to the larger field of view and reduce the resolution of the final tomogram following reconstruction.
Third, scanner settings for acquiring projection images will also vary by instrument. To maximize the resolution capabilities of the scanner, the X-ray beam spot size should be as small as possible (5–10 μm). This can be achieved by balancing X-Ray voltage and current settings such that the total power is 3–4 W. With these settings and the appropriate exposure time on the camera, proper X-ray beam attenuation by the sample and optimal image contrast can be achieved. The use of aluminum or copper filters between the object and the X-ray source can be used to fine-tune the optimal X-ray energy settings for the best image contrast or attenuate the beam sufficiently for higher powered sources to be used. As for image resolution, this will depend on many different variables, including stain type, number of projection images, image pixel size, camera position, sample movement, scanner vibrations and reconstruction parameters. A bar pattern phantom (QRM GmbH) containing known size markers can help evaluate spatial resolution for a given scanner and camera setting.
It is also worth evaluating the merits of imaging critical point dried or hydrated samples. Sombke et al. performed a comparative assessment of the two methods and found critical point drying to be superior for μ-CT applications involving arthropods30. However, benefits of hydrated samples are that animals are subjected to less chemical and mechanical exposure that could lead to both quantitative and morphological artifacts. This also tends to preserve delicate tissues better than CPD. However, hydrated samples have a much shorter shelf life and should be imaged no later than one month after fixation since tissue degradation and reduced image quality becomes obvious at that point. Also, the resolution of hydrated samples will be slightly less than a critical point dried sample, because X-rays must also penetrate through both a plastic pipette tip and the surrounding liquid (water or buffer). Critical Point Dried samples can be preserved for much longer periods of time, especially when kept on Drierite. They also can be placed directly in the X-ray beam path by simply gluing the wings or legs to an insect pin and placing it in the stage chuck, simplifying the mounting process. However, the extensive ethanol dehydration of these samples can lead to tissue shrinkage and loss of delicate tissue architecture, which is why it is important to perform a range of increasing EtOH concentrations to minimize these effects. Nonetheless, it should be noted that all forms of chemical treatment, including paraformaldehyde fixation and even iodine staining can cause tissue shrinkage58, 59. While neither method will provide measurements of ‘actual organ size’ in a living fly, morphometric measurements are still valid when comparing mutant and wildtype animals so long as the fixation, staining, and drying steps are carried out identically for both sets of samples—preferably in parallel.
In conclusion, μ-CT provides a useful whole animal imaging tool for Drosophila33, 34, 35, 36, 37, 38, 39, 40, 41. Many other studies have showcased the power of this technology for understanding various aspects of insect taxonomy, ecology, physiology, development, and anatomy that can help inform future studies in flies1, 28, 30, 31, 32, 55, 56, 57. Combined with the genetic and light microscopy tools already widely used in this organism, μ-CT can position itself within an experimental pipeline that allows for a deeper understanding between genotype and phenotype.
Acknowledgments
None of this would have been possible without the support of Nasser Rusan. I would like to thank H. Doug Morris, Danielle Donahue, and Brenda Klaunberg of the NIH Mouse Imaging Facility and Ben Ache of Micro Photonics for training and helpful discussion. I also thank Mansoureh Norouzi Rad of Zeiss for scanning abdomen samples on the Xradia 520 Versa. Lauren Smith, Samantha Smith, and Rachel Ng also helped with scanning. Mike Marsh of Object Research Systems provided Dragonfly technical support. I am also grateful for support from the National Heart, Lung, and Blood Institute (1K22HL137902-01) and Start Up Funds from the University of Wyoming. I also thank the anonymous reviewers for their helpful suggestions and comments.
Footnotes
Disclosures
The author declares no competing or financial interests.
References
- 1.Metscher BD MicroCT for comparative morphology: simple staining methods allow high-contrast 3D imaging of diverse non-mineralized animal tissues. BMC Physiology. 9, 11 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Rahman IA, Smith SY Virtual paleontology: computer-aided analysis of fossil form and function. Journal of Paleontology. 88 (04), 633–635 (2014). [Google Scholar]
- 3.Carlson WD Three-dimensional imaging of earth and planetary materials. Earth and Planetary Science Letters. 249 (3–4), 133–147 (2006). [Google Scholar]
- 4.Gutiérrez Y, Ott D, Töpperwien M, Salditt T, Scherber C X-ray computed tomography and its potential in ecological research: A review of studies and optimization of specimen preparation. Ecology and Evolution. 8 (15), 7717–7732 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Abel RL et al. Digital preservation and dissemination of ancient lithic technology with modern micro-CT. Computers & Graphics. 35 (4), 878–884 (2011). [Google Scholar]
- 6.Kastengren A, Powell CF Synchrotron X-ray techniques for fluid dynamics. Experiments in Fluids. 55 (3), 1686 (2014). [Google Scholar]
- 7.Boerckel JD, Mason DE, McDermott AM, Alsberg E Microcomputed tomography: approaches and applications in bioengineering. Stem Cell Research & Therapy. 5 (6), 144 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Du Plessis A, Yadroitsev I, Yadroitsava I, Le Roux SG X-Ray Microcomputed Tomography in Additive Manufacturing: A Review of the Current Technology and Applications. 3D Printing and Additive Manufacturing. 5 (3), 227–247 (2018). [Google Scholar]
- 9.Cnudde V, Boone MN High-resolution X-ray computed tomography in geosciences: A review of the current technology and applications. Earth-Science Reviews. 123, 1–17 (2013). [Google Scholar]
- 10.Zhu Y, Zhang J, Li A, Zhang Y, Fan C Synchrotron-based X-ray microscopy for sub-100nm resolution cell imaging. Current Opinion in Chemical Biology. 39, 11–16 (2017). [DOI] [PubMed] [Google Scholar]
- 11.Kampschulte M et al. Nano-Computed Tomography: Technique and Applications. RoFo: Fortschritte Auf Dem Gebiete der Rontgenstrahlen und der Nuklearmedizin. 188 (2) (2016). [DOI] [PubMed] [Google Scholar]
- 12.Seeram E Computed tomography: A technical review. Radiologic Technology. 89 (3), 279CT–302CT (2018). [PubMed] [Google Scholar]
- 13.Rawson SD, Maksimcuka J, Withers PJ, Cartmell SH X-ray computed tomography in life sciences. BMC Biology. 18 (1), 21 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Morton EJ, Webb S, Bateman JE, Clarke LJ, Shelton CG Three-dimensional x-ray microtomography for medical and biological applications. Physics in Medicine and Biology. 35 (7), 805–820 (1990). [DOI] [PubMed] [Google Scholar]
- 15.Lin ASP, Stock SR, Guldberg RE Microcomputed Tomography Springer Handbook of Microscopy. Springer; 2–2 (2019). [Google Scholar]
- 16.Feldkamp LA, Davis LC, Kress JW Practical cone-beam algorithm. Journal of the Optical Society of America A. 1 (6), 612 (1984). [Google Scholar]
- 17.Clark DP, Badea CT Micro-CT of rodents: state-of-the-art and future perspectives. Physica Medica: PM: An International Journal Devoted to the Applications of Physics to Medicine and Biology: Official Journal of the Italian Association of Biomedical Physics (AIFB). 30 (6), 619–634 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Schambach SJ, Bag S, Schilling L, Groden C, Brockmann MA Application of micro-CT in small animal imaging. Methods. 50 (1), 2–13 (2010). [DOI] [PubMed] [Google Scholar]
- 19.International Mouse Knockout Consortium, Collins FS, Rossant J, Wurst WA mouse for all reasons. Cell. 128 (1), 9–13 (2007). [DOI] [PubMed] [Google Scholar]
- 20.Wong MD, Dorr AE, Walls JR, Lerch JP, Henkelman RM A novel 3D mouse embryo atlas based on micro-CT. Development. 139 (17), 3248–3256 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Dyer EL et al. Quantifying Mesoscale Neuroanatomy Using X-Ray Microtomography. eNeuro 4 (5) (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Weinhardt V et al. Quantitative morphometric analysis of adult teleost fish by X-ray computed tomography. Scientific Reports. 8 (1), 16531 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Ding Y et al. Computational 3D histological phenotyping of whole zebrafish by X-ray histotomography. eLife. 8, e44898 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Ahmadzadeh E et al. A collection of genetic mouse lines and related tools for inducible and reversible intersectional mis-expression. Development. 147 (10) dev186650 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Koster R, Sassen WA A molecular toolbox for genetic manipulation of zebrafish. Advances in Genomics and Genetics. 2015, 151–163 (2015). [Google Scholar]
- 26.Hales KG, Korey CA, Larracuente AM, Roberts DM Genetics on the fly: A primer on the Drosophila model system. Genetics. 201 (3), 815–842 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Ugur B, Chen K, Bellen HJ Drosophila tools and assays for the study of human diseases. Disease Models & Mechanisms. 9 (3), 235–244 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Smith DB et al. Exploring miniature insect brains using micro-CT scanning techniques. Scientific Reports. 6, 21768 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Swart P, Wicklein M, Sykes D, Ahmed F, Krapp HG A quantitative comparison of micro-CT preparations in Dipteran flies. Scientific Reports. 6, 39380 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Sombke A, Lipke E, Michalik P, Uhl G, Harzsch S Potential and limitations of X-Ray micro-computed tomography in arthropod neuroanatomy: a methodological and comparative survey. The Journal of Comparative Neurology. 523 (8), 1281–1295 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Betz O et al. Imaging applications of synchrotron X-ray phase-contrast microtomography in biological morphology and biomaterials science. I. General aspects of the technique and its advantages in the analysis of millimetre-sized arthropod structure. Journal of Microscopy. 227 (Pt 1), 51–71 (2007). [DOI] [PubMed] [Google Scholar]
- 32.Wipfler B, Pohl H, Yavorskaya MI, Beutel RG A review of methods for analysing insect structures - the role of morphology in the age of phylogenomics. Current Opinion in Insect Science. 18, 60–68 (2016). [DOI] [PubMed] [Google Scholar]
- 33.Chen WC et al. Toward a new insight of calcium oxalate stones in Drosophila by micro-computerized tomography. Urolithiasis. 46 (2), 149–155 (2017). [DOI] [PubMed] [Google Scholar]
- 34.Fabian B, Schneeberg K, Beutel RG Comparative thoracic anatomy of the wild type and wingless (wg1cn1) mutant of Drosophila melanogaster (Diptera). Arthropod Structure & Development. 45 (6), 611–636 (2016). [DOI] [PubMed] [Google Scholar]
- 35.Harrison JF et al. Developmental plasticity and stability in the tracheal networks supplying Drosophila flight muscle in response to rearing oxygen level. Journal of Insect Physiology. (2017). [DOI] [PubMed] [Google Scholar]
- 36.Klok CJ, Kaiser A, Socha JJ, Lee WK, Harrison JF Multigenerational effects of rearing atmospheric oxygen level on the tracheal dimensions and diffusing capacities of pupal and adult Drosophila melanogaster. Advances in Experimental Medicine and Biology. 903 (2016). [DOI] [PubMed] [Google Scholar]
- 37.Mattei AL, Riccio ML, Avila FW, Wolfner MF Integrated 3D view of postmating responses by the Drosophila melanogaster female reproductive tract, obtained by micro-computed tomography scanning. Proceedings of the National Academy of Sciences of the United States of America. 112 (27), 8475–8480 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Mizutani R, Saiga R, Takeuchi A, Uesugi K, Suzuki Y Three-dimensional network of Drosophila brain hemisphere. Journal of Structural Biology. 184 (2), 271–279 (2013). [DOI] [PubMed] [Google Scholar]
- 39.Mizutani R, Takeuchi A, Akamatsu G, Uesugi K, Suzuki Y Element-specific microtomographic imaging of Drosophila brain stained with high-Z probes. Journal of Synchrotron Radiation. 15 (Pt 4), 374–377 (2008). [DOI] [PubMed] [Google Scholar]
- 40.Schoborg TA, Smith SL, Smith LN, Morris HD, Rusan NM Micro-computed tomography as a platform for exploring Drosophila development. Development. 146 (23) (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Chaturvedi D, Prabhakar S, Aggarwal A, Atreya KB, VijayRaghavan K Adult Drosophila muscle morphometry through microCT reveals dynamics during ageing. Open Biology. 9 (6), 190087 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Bainbridge SP, Bownes M Staging the metamorphosis of Drosophila melanogaster. Journal of Embryology and Experimental Morphology. 66, 57–80 (1981). [PubMed] [Google Scholar]
- 43.Withers PJ X-ray nanotomography. Materials Today. 10 (12), 26–34 (2007). [Google Scholar]
- 44.Bleichrodt F et al. J. Easy implementation of advanced tomography algorithms using the ASTRA toolbox with Spot operators. Numerical Algorithms. 71 (3), 673–697 (2016). [Google Scholar]
- 45.Salmon PL, Liu X, Sasov A A post-scan method for correcting artefacts of slow geometry changes during micro-tomographic scans. Journal of X-ray Science and Technology. 17 (2), 161–174 (2009). [DOI] [PubMed] [Google Scholar]
- 46.Schindelin J et al. Fiji: an open-source platform for biological-image analysis. Nature Methods. 9 (7), 676–682 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.CIBC Seg3D: Volumetric Image Segmentation and Visualization. Scientific Computing and Imaging Institute (SCI). at <http://www.seg3d.org> (2016).
- 48.Yushkevich PA et al. User-guided 3D active contour segmentation of anatomical structures: significantly improved efficiency and reliability. Neuroimage. 31 (3), 1116–1128 (2006). [DOI] [PubMed] [Google Scholar]
- 49.Lösel P, Heuveline V Enhancing a diffusion algorithm for 4D image segmentation using local information. Medical Imaging 2016: Image Processing. 9784, 97842L (2016). [Google Scholar]
- 50.Null B, Liu CW, Hedehus M, Conolly S, Davis RW High-resolution, in vivo magnetic resonance imaging of Drosophila at 18.8 Tesla. Plos One. 3 (7), e2817 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Holmes J In vivo real-time optical coherence tomography imaging of Drosophila for cardiovascular research. Nature Methods. 6 (10) 5557 (2009). [Google Scholar]
- 52.Men J et al. Optical coherence tomography for brain imaging and developmental biology. IEEE journal of selected topics in quantum electronics: a publication of the IEEE Lasers and Electro-optics Society. 22 (4) 6083213 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Jährling N, Becker K, Schönbauer C, Schnorrer F, Dodt HU Three-dimensional reconstruction and segmentation of intact Drosophila by ultramicroscopy. Frontiers in Systems Neuroscience. 4, 1 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Pende M et al. High-resolution ultramicroscopy of the developing and adult nervous system in optically cleared Drosophila melanogaster. Nature Communications. 9 (1), 4731 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Socha JJ, Westneat MW, Harrison JF, Waters JS, Lee WK Real-time phase-contrast x-ray imaging: a new technique for the study of animal form and function. BMC Biology. 5, 6 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Westneat MW et al. Tracheal respiration in insects visualized with synchrotron x-ray imaging. Science. 299 (5606), 558–560 (2003). [DOI] [PubMed] [Google Scholar]
- 57.Metscher BD MicroCT for developmental biology: a versatile tool for high-contrast 3D imaging at histological resolutions. Developmental Dynamics. 238 (3), 632–640 (2009). [DOI] [PubMed] [Google Scholar]
- 58.Stowell RE Effect on tissue volume of various methods of fixation, dehydration, and embedding. Stain Technology. 16 (2), 67–83 (1941). [Google Scholar]
- 59.Vickerton P, Jarvis J, Jeffery N Concentration-dependent specimen shrinkage in iodine-enhanced microCT. Journal of Anatomy. 223 (2), 185–193 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]