Abstract
The high costs of N fertilizers in the coffee production emphasizes the need to optimize fertilization practices and improve nitrogen use efficiency. Urea is widespread in nature, characterizing itself as a significant source of nitrogen for the growth and development of several organisms. Thus, the characterization of genes involved in urea transport in coffee plants is an important research topic for the sustainable production of this valuable cash crop. In the current study, we evaluated the expression of the DUR3 gene under abiotic and biotic stresses in coffee plants. Here, we show that the expression of a high-affinity urea transporter gene (CaDUR3) was up-regulated by N starvation in leaves and roots of two out of three C. arabica cultivars examined. Moreover, the CaDUR3 gene was differentially expressed in coffee plants under different abiotic and biotic stresses. In plants of cv. IAPAR59, CaDUR3 showed an increased expression in leaves after exposure to water deficit and heat stress, while it was downregulated in plants under salinity. Upon infection with H. vastatrix (coffee rust), the CaDUR3 was markedly up-regulated at the beginning of the infection process in the disease susceptible Catuaí Vermelho 99 in comparison with the resistant cultivar. These results indicate that besides urea acquisition and N-remobilization, CaDUR3 gene may be closely involved in the response to various stresses.
Keywords: Coffea ssp, Nitrogen deficiency, Gene expression, Urea transporter
Introduction
Coffee is one of the most valuable agricultural traded commodities worldwide and is grown mostly in developing countries (Tucker 2017). It is one of the most popular and frequently consumed beverage worldwide, being also an important product for trade in many non-tropical countries involved in the industrialization (Fridell 2014). The allotetraploid Coffea arabica L. (2n = 4x = 44) and the diploid C. canephora Pierre ex A. Froehner (2n = 2x = 22) are the most cultivated species in the world (International Coffee Organization, http://www.ico.org) among the 124 coffee species already described (Davis et al. 2011).
Coffee production depends on climate, soil quality and external inputs, such as agrochemicals and fertilizers. Nitrogen (N), one of the most important nutrients for crop growth and yield, are mainly available in many forms expensive commercial inorganic fertilizers (Bloom 2015). Urea [CO(NH2)2] has become the most commonly used N fertilizer in the world because of it's low production costs and high N content (http://faostat.fao.org). Urea is a small neutral molecule, which is widespread in nature, where it acts as a source of N for many organisms, including bacteria, fungi and plants (Wang et al. 2008). In coffee production, urea is the major N form supplied for the full crop cycle.
Nitrate is the preferred inorganic N source for several plant species, though ammonium has been shown to be the preferential inorganic N source for N-starved coffee roots (dos Santos et al. 2017). Plants also possess a specific system for the direct uptake of urea from the environment (Fan et al. 2017). Once absorbed, urea is hydrolyzed to form ammonium, which in turn enters into the GS/GOGAT cycle (glutamine synthetase/glutamate synthase cycle) before being assimilated in amino acids (Tegeder and Masclaux-Daubresse 2018). Urea uptake in roots can be mediated by membrane intrinsic proteins such as NIP5;1 (Yang et al. 2015), but the most important high-affinity urea transporter in roots are DUR3 homologs.
DUR3 belongs to the sodium:solute symporter (SSS) family in plants (Kojima et al. 2007). Functional complementation of the yeast dur3 mutant had been successfully used to identify and characterize the DUR3 orthologues and study the urea transport in plants (ElBerry et al. 1993; Zanin et al. 2014; Zhang et al. 2016). In Arabidopsis thaliana the AtDUR3 gene is expressed in the roots under N deficiency, and its induction increases drastically when the roots come into contact with urea (Kojima et al. 2007). Studies in algae, rice and maize confirmed DUR3 as the major transporter involved in urea acquisition and its regulation by N supply in plants (Liu et al. 2015; Beier et al. 2019). Phylogenetic analysis by De Michele et al. (2012) demonstrated a reduction of genes encoding urea transporters throughout plant evolution, eventually down to a single copy in vascular plants, which is possibly a consequence of the preference for other nitrogen sources.
Despite the agronomic importance of urea fertilizer, the detailed molecular characterization of DUR3 is restricted to model plants, such as A. thaliana, rice and maize (Kojima et al. 2007; Wang et al. 2013; Zanin et al. 2014). To the best of our knowledge, there is no study addressing urea transporters in woody species, such as in coffee, for example. Therefore, the objective of this study was to analyze transcriptional patterns of CaDUR3 in C. arabica plants under nitrogen starvation, abiotic and biotic stresses. We compared the transcriptional levels of CaDUR3 in three C. arabica cultivars under nitrogen starvation and investigated the expression patterns of the CaDUR3 gene in coffee plants under abiotic (water deficit, heat and salinity) and biotic (coffee leaf rust) stresses.
Material and methods
In silico identification of the CaDUR3 gene
We performed searches in the Brazilian Genome Project database (Vieira et al. 2006; Mondego et al. 2011) using the keyword urea transporter. A cDNA clone corresponding to a unique EST (expressed sequence tag/CA00-XX-IA2-016-H07-ECF) was identified and obtained from Cenargen/Embrapa (Brasília-DF). This sequence was compared by BLAST against Arabidopsis sequence databases (Altschul et al. 1997) and used as query sequence to find the complete CaDUR3 sequence in the NCBI database (National Center for Biotechnology Information/ https://www.ncbi.nlm.nih.gov/genome/gdv/?org=coffea-arabica&group=lamiids). After confirming the identity of the CaDUR3 gene, we used gene-specific primers (Fw CaDUR3—5′GGCCTCTTCTCCCACAATTT 3′ and Rev CaDUR3 – 5′CCTCAA ACAGCTGCTTCCAT) for amplifying the probe for Southern blot and for transcriptional analyses.
Plant materials
We used the following C. arabica cultivars throughout this study, viz., cv. IAPAR59, cv. Catuaí Vermelho 99, cv. Mundo Novo, cv. Timor hybrid CIFC 832/2 (HT). We also used C. canephora var. Conilon (Cc) and C. eugenioides (Ce) in the Southern Blot analysis. The cultivars used are specified in each procedure. The plants were grown in a greenhouse and under field conditions at the experimental facility of the Agronomic Institute of Paraná (IAPAR—latitude 23°18′ S, longitude 51°09′ W, average altitude 585 m; Londrina, Paraná State, Brazil).
Southern blotting
The extraction of genomic DNA from leaves of C. arabica cv. Hybrid of Timor CIFC 832/2, C. canephora, C. eugenioides and C. arabica cv. IAPAR59 was performed using the protocol of Doyle and Doyle (1987). Aliquots (20 ug) of genomic DNA were digested overnight at 37 °C with the enzyme DraI, fractionated on 0.7% agarose gel electrophoresis and transferred onto a positively charged Hybond N+ membrane (Amersham Biosciences) according to the standard protocols. The probe was obtained by amplification of the clone CA00-XX-IA2-016-H07-EC.F (primers Fw CaDUR3 5′-GGCCTCTTCTCCCACAATTT-3′ and Rev CaDUR3 5′-CCTCAAACAGCTGCT TCCAT-3′) and random primed labeled with α-dCTP32 for 24 h at 37 °C. The image was captured using fluorescent image analyzer FLA 3000—series (Fuji Photo Film CO, Lts. Tokyo, Japan).
Drought, saline and heat stresses
Detailed procedures of the stress experiments were described in previous studies from our group (dos Santos et al. 2011, 2015). For these experiments we used 20 months-old C. arabica cv IAPAR59 plants grown in 15 L pots equally filled with substrate (3 soil: 1 sand: 1 organic compound). Mature leaves in the drought stress experiment were collected according to the water potential readings: no stress (NS; ± −1.36 MPa), moderate stress (MS; ± −2.35 MPa) and severe stress (SS; ± −4.3 MPa).
In the saline stress experiment, the plants were irrigated on the first day of the treatment with 50 mM NaCl and with 100 mM NaCl on the second day to avoid osmotic shock. From the third day until the end of the experiment (25 days), the plants were irrigated daily with 150 mM NaCl. Mature leaves were sampled after 0, 6, 12 and 25 days after the beginning of the salt treatment. Both, the drought and saline stresses were performed in a greenhouse regulated to 25 ± 2 °C on natural day/night cycle.
Plants used for the heat stress experiment were cultivated in a growth chamber (12/12 h light/dark cycle, 25/23 °C day/night temperature, 45% humidity, and photosynthetic photon flux density of ~ 250 µmol m−2·s−1, respectively. After initial acclimation (7 days), the temperature was elevated to 37 °C for five days with daily irrigation maintenance. The leaf samples were collected 3 and 5 days after the beginning of the stress. The three biological replicates were represented by pools of lateral roots and leaves at the same developmental stage. Subsequently, samples from all the treatments were immediately frozen in liquid nitrogen and stored at –80 °C until analysis.
N-starvation
The basic procedures of the N-starvation experiment using C. arabica cv. IAPAR59 plants were according to de Carvalho et al. (2013) and dos Santos et al. (2019). Initially, seeds were germinated in greenhouse using boxes containing autoclaved sand and were irrigated twice weekly with Clark nutritive solution (Clark et al. 1975). After 4 months, we selected uniform plants by size uniformity with 3 leaf pairs. Plants were then transferred to a nutritive solution system with constant aeration. In brief, plants were submitted to a 4-week period of acclimation. After this period, plants were transferred to a modified Clark solution without N. Lateral roots and leaf samples were harvested after 1 day and 10 days in the N-free solution. As experimental control for each time point, were used plants under normal N levels. Experiments were conducted twice with a minimum of three biological replicates per experiment. Biological replicates were represented by pools of lateral roots and leaves of at least nine plants at the same developmental stage. After harvesting, all samples were frozen immediately in liquid nitrogen and stored at –80 °C until further analysis.
Urea quantification in N-starved plants
For the urea determination, leaves and roots were macerated in liquid nitrogen, suspended in milliQ water autoclaved, homogenized and then centrifuged at 10.000 rpm at 4 °C for 10 min. The supernatant was filtered through a glass wool, transferred to a new tube and kept on ice until the time of analysis. The reaction mixture was composed of 750 µL phosphate buffer (KH2PO4) 0.1 M pH 7.5, 200 µL of plant extract and 50 µL of 50 mM urease solution. The reaction was incubated in a water bath at 37 °C for 40 min and quantified according to Weatherburn (1967) using urea as standard, and measuring absorbance at 625 nm.
Hemileia vastatrix infection
The experiment was conducted in a greenhouse at the Coffee Center “Alcides Carvalho”, (Instituto Agronômico de Campinas—IAC, Campinas, Brazil) as described in Baba et al. (2020). In this experiment, six months-old plants with uniform leaf size and development were used. A suspension of Hemileia vastatrix race II uredospores (final concentration of 1 mg/mL) was inoculated on the abaxial surface of completely expanded leaves of C. arabica cv. IAPAR59 and cv. Catuaí Vermelho 99 which are resistant and susceptible to coffee rust, respectively. Leaves were also mock inoculated with water as controls. Subsequently, the plants were transferred to a humid chamber, covered by a plastic and maintained in the dark for 24 h, at 22 °C with high relative humidity. Samples were collected at 0, 12, 24 and 48 h post-inoculation (HPI), and each treatment was composed of three biological replicates represented by pools of leaves at the same developmental stage. All leaves collected were frozen immediately in liquid nitrogen and stored at –80 °C until further analysis.
RNA extraction and cDNA synthesis
Before initiating the N-starvation, abiotic an biotic stresses experiments, we sampled fully expanded leaves, flower buds, flowers, plagiotropic shoots, secondary roots, endosperm and pericarp from mature fruits of the C. arabica cv. IAPAR59 plants cultivated under field conditions in order to have an overall assessment of the CaDUR3 transcriptional profiles.
Total RNA extraction from all the samples was performed using the standard CTAB method using the same procedures of dos Santos et al. (2011). RNAs extracted were checked for integrity by agarose gel electrophoresis and then treated with DNase (RNase-free, Invitrogen) to remove genomic DNA contamination. The purity and concentration of the extracted RNAs were determined using a NanoDrop® ND-1000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). The cDNAs synthesis were carried out using a SuperScript III Reverse Transcriptase kit (Invitrogen, Carlsbad, CA, USA) in a final volume of 20 μL using 5 μg of total RNA, according to the manufacturer's instructions.
Expression analysis
CaDUR3-specific primers were designed using Primer Express v3.0.1. Specific primers (Tm = 60 °C) were designed from the EST (CA00-XX-IA2-016-H07-ECF) to ensure the amplification of a single ~ 85 bp PCR product (Fw CaDUR3—5′GGCCTCTTCTCCCACAATTT 3′ and Rev CaDUR3—5′CCTCAAACAGCTGCTTCCAT 3′). The specificity of the primers was confirmed by PCR and by melting curves to verify the presence of nonspecific products. The mean amplification efficiency of the primer pair (98%) was assessed by the program LinRegPCR (Ruijter et al. 2009). RT-qPCR was analyzed in a 7500 Fast Real-Time PCR System (Applied Biosystems, Foster City, CA, USA). The PCR mixture (10 µL total volume) included 1 µL cDNA, 5 µL SYBR Green Master Mix (Life Technologies, USA), 0.5 µL sequence-specific forward primer (10 µM), 0.5 µL reverse primer (10 µM) and 3 µL water. The RT-qPCR program conditions an initial step of 95 °C for 10 min, followed by 40 cycles of 95 °C for 35 s and 60 °C for 60 s. The relative expression levels of the CaDUR3 gene were calculated based on (1 + E)−ΔΔct method (Livak and Schmittgen 2001) and normalized using the gene CaEF1α, as recommended in de Carvalho et al. (2013). All RT-qPCR experiments were performed in triplicate in three independent experiments.
Statistical analysis
The experimental design used in all experiments (abiotic stresses, H. vastatrix infection and N starvation) was the completely randomized design with three biological replications. Data from RT-qPCR and urea content were submitted to analysis of variance (ANOVA) to determine differences between treatments at p < 0.05. Means were separated according to Duncan’s test (p < 0.05). The data were normalized using the Box-Cox power transformation (Box and Cox 1964). All statistical analyses were performed using the easyanova (Arnhold 2013) and MASS (Venables and Ripley 2002) packages of R statistical software (R Core Team, 2019).
Results
CaDUR3 identification
Using urea transporter as the keyword, the in silico survey within the Brazilian Coffee Genome Database (http://www.lge.ibi.unicamp.br/cafe/; Vieira et al. 2006) resulted in only one EST (CA00-XX-IA2-016-H07-EC.F). This sequence corresponds to the accession number GW440609 deposited in NCBI database. Primarily, the analysis of the corresponding cDNA clone by the BLAST tools (http://www.ncbi.nlm.nih.gov/blast) showed that it matches the sequence of the DUR3 gene of the sodium:solute symporter (SSS) family (Kojima et al. 2007). Using this sequence as query, we identified the complete sequence of the DUR3 gene in the C. arabica genome deposited in the NCBI database (LOC113707287—NCBI Accession number), which is located on chromosome 8 and codes for a mature protein of 678 amino acids (XP_027085342). By comparing this sequence with the known sequences of A. thaliana genome (TAIR: https://www.arabidopsis.org), we verified that the CaDUR3 presented high similarity (84%) with AT5G45380.1 (AtDUR3; Kojima et al. 2007).
The cDNA probe detected a single band on Southern blots of genomic DNA of C. eugenioides digested and two strong hybridization signals in C. canephora, C. arabica cv. IAPAR59 and in the Hybrid Timor accession with the restriction enzyme DraI (Fig. 1). C. arabica is derived from an allopolyploidization (CaEa genome) event between the parental diploids C. canephora (C genome) and C. eugenioides (E genome) (Lashermes et al. 1999, 2000), therefore the extra band observed in the C. arabica, C. canephora and in the interspecific hybrid is due to differences in the enzyme restriction profiles of the genomic DNA between the two parental species. Indeed, the analysis of the DUR3 gene sequences of C. arabica and C. canephora on the available databases showed a single internal restriction site for the enzyme DraI, which resulted in the expected Southern Blot profiles. Therefore, according to the restriction pattern observed, the DUR3 is likely present as a single copy gene within in all species examined here.
Fig. 1.

Southern Blot analysis. The genomic DNA of C. arabica cv. Timor hybrid CIFC 832/2 (HT), C. canephora (Cc), C. eugenioides (Ce) and C. arabica cv. IAPAR59 were digested with the restriction enzyme DraI. The probe used was the PCR product obtained from the amplification of the cDNA clone CA00-XX-IA2-016-H07-EC.F that corresponds to a partial sequence of the CaDUR3 gene
CaDUR3 expression in C. arabica organs
Initially, we verified the transcript levels of the CaDUR3 gene in seven organs of C. arabica plants (cv. IAPAR59) cultivated under field conditions. The presence of CaDUR3 transcripts was observed in all tissues, with particularly high-level expression in roots (Fig. 2).
Fig. 2.
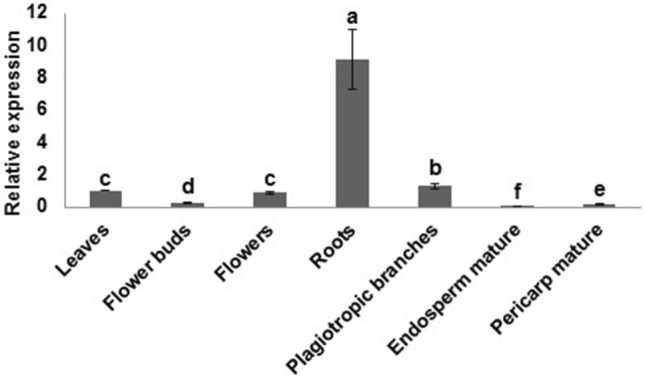
Relative expression of CaDUR3 in different tissues of the C. arabica cv. IAPAR59. The internal calibrator used in this experiment is it's leaf. Transcript abundances were normalized using the expression of the CaEF1α. Values are presented as mean ± standard deviation (n = 3)
Transcription patterns of CaDUR3 under N-starvation
In order to quantify the variation in transcript abundance of CaDUR3, we performed RT-qPCR analysis on plants of three C. arabica cultivars subjected to N-starvation for 1 and 10 days (Fig. 3). There was a considerable variation in the expression of CaDUR3 in leaves of the three cultivars (IAPAR59, Mundo Novo and Catuaí Vermelho 99). Nonetheless, our results showed that CaDUR3 transcript levels increased after long-term N-starvation (10 days) when compared to the N-sufficient condition (day 0) in all cultivars, with the levels of CaDUR3 expression being highest in leaves of the cv. IAPAR59 (approximately eight times more that the calibrator) (Fig. 3). We also showed that CaDUR3 transcripts substantially increased in roots of this cultivar after long exposure to N-starvation (Fig. 4).
Fig. 3.
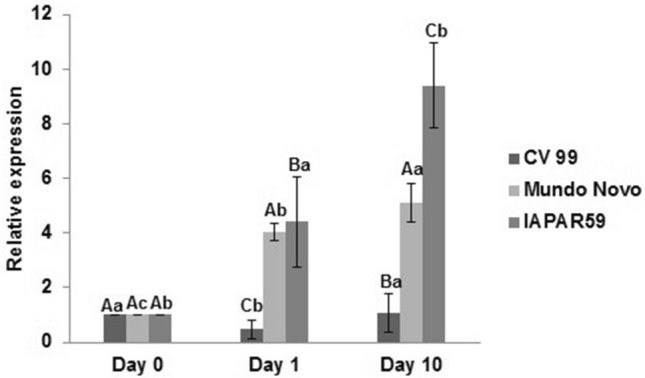
Relative expression of CaDUR3 gene in coffee leaves of plants under nitrogen starvation of the C. arabica cv. IAPAR59, C. arabica cv. Mundo Novo and C. arabica cv. Catuaí Vermelho 99. Transcript abundances were normalized against the expression of the CaEF1α gene and treatment day 0 was used as calibrator samples (relative gene expression = 1). Values are presented as mean ± standard deviation (n = 3). Averages followed by the same capital letter, compare differences between the C. arabica cultivars at the same day of N starvation treatment, did not differ significantly by the Duncan test (p < 0.05). Averages followed by the same lowercase letter, compare differences between different days of N starvation treatment within each cultivars, did not differ significantly by the Duncan test (p < 0.05)
Fig. 4.

Relative expression of CaDUR3 gene in coffee roots of the C. arabica cv. IAPAR59 under nitrogen starvation. Transcript abundances were normalized using the expression of the CaEF1α gene and day 0 as calibrator sample (relative gene expression = 1). Values are presented as mean ± standard deviation (n = 3)
Urea content in leaves and roots of C. arabica
Regarding the leaf and root urea content under N-starvation conditions, we observed that the highest concentration was in leaf tissues of the C. arabica cv. IAPAR59 (Fig. 5). The N contents in roots were very low in all treatments. The concentrations were practically the same from the beginning to the end of the experiment.
Fig. 5.

Urea content in leaves and roots of C. arabica cv. IAPAR59 plants grown under N starvation. Values are presented as mean ± standard error (n = 3). Averages followed by the same capital letter, compare differences between leaves and roots at the same day of N starvation treatment, did not differ significantly by the Duncan test (p < 0.05). Averages followed by the same lowercase letter, compare differences between the tissues (leaves and roots) at different days of N starvation treatment, did not differ significantly by the Duncan test (p < 0.05)
Transcription patterns of CaDUR3 under abiotic stresses
Abiotic stresses are among the main factors affecting food production worldwide and may affect nitrogen metabolism. In this sense, we sought to investigate the transcriptional patterns of the CaDUR3 gene in leaves of C. arabica cv. IAPAR59 plants in response to salt stress, high temperature and drought (Fig. 6). We detected 2- and threefold increase in transcript levels in plants under moderate and severe water deficits treatments compared with those under normal irrigation conditions, respectively (Fig. 6a). In heat-stressed plants, the expression of the CaDUR3 gene was upregulated when compared to the control treatment (day 0—no stress) (Fig. 6b). There were significant differences in the transcript levels, mainly at day 5 of heating at 37 °C. Under conditions of salt stress, the transcriptional profile of the CaDUR3 gene after the different periods under salinity condition (150 mM NaCl solution) was differentially modulated. The results showed in Fig. 6c, demonstrated that in the first two sampling periods (6 and 12 days) there was a drastic reduction in the number of transcripts (downregulation). However, the expression level of CaDUR3 increased moderately in leaf tissues in 25 days after NaCl treatment (Fig. 6c).
Fig. 6.
Relative expression of CaDUR3 in leaves of C. arabica cv. IAPAR59 under abiotic stresses. a Drought experiment: no stress (NS; ±-1.36 MPa), moderate stress (MS; ±-2.35 MPa), severe stress (SS; ±-4.3 MPa); b Heat experiment: no stress (Day 0), 3 and 5 days exposed to 37 °C; c Saline experiment: no stress (Day 0), 6, 12 and 25 days after onset of the saline stress (150 mM NaCl). Transcript abundances were normalized against the expression of the CaEF1α gene, and the treatments NS (drought stress) and day 0 (heat and saline stress) were used as controls
Transcription patterns of CaDUR3 after H. vastatrix infection
The infection of coffee leaves with race II of H. vastatrix modulated differential CaDUR3 gene expression patterns depending on the cultivar tested (Fig. 7). In a rust-resistant cultivar (IAPAR59), a gradual decline of transcript abundance was observed until 48 h post-inoculation. On the contrary, the susceptible cultivar (Catuaí Vermelho 99) showed a peak of CaDUR3 mRNA abundance in their leaves at the first sampling time (12 h post-inoculation), with a considerable decline in the next two sampling periods (24 and 48 h post-inoculation) (Fig. 7).
Fig. 7.

Relative expression of CaDUR3 gene in coffee leaves infected with Hemileia vastatrix race II after 0, 12, 24 and 48 hours post-inoculation in C. arabica cv. Catuaí Vermelho 99 (susceptible) and C. arabica cv. IAPAR59 (resistant) genotypes. Transcript abundances were normalized against the expression of the CaEF1α gene and T0 was used as calibrator samples (relative gene expression = 1). Values are presented as mean ± standard deviation (n = 3). Averages followed by the same capital letter, compare differences between the C. arabica cultivars at the same hours post-inoculation, did not differ significantly by the Duncan test (p < 0.05). Averages followed by the same lowercase letter, compare differences between hours post-inoculation within each cultivars, did not differ significantly by the Duncan test (p < 0.05)
Discussion
Urea is mainly used as a source of N fertilizer in agricultural crops with high N demand (Karamos et al. 2014), such as in coffee. In plants, the first evidence of active transport of urea in planta was presented by Liu et al. (2003a). DUR3 belongs to a family of transporters for several solutes found in eukaryotes and is presented as a single copy in the plant systems evaluated so far (De Michele et al. 2012). The C. arabica is an amphidiploid (CaEa genome) species originated from the natural hybridization (estimated 553,000 to 665,000 years ago) (Yu et al. 2011; Scalabrin et al. 2020; Bawin et al. 2020) between two diploid ancestral species (C. canephora—C genome and C. eugenioides—E genome), and in its genome there is the contribution of two sub-genomes (Lashermes et al. 1999, 2000; Vidal et al. 2010). As illustrated by the Southern blot, the results in the Coffea species tested (Fig. 1) are concordant with data in other plant genomes, such as in Arabidopsis (Liu et al. 2003a), rice (OsDUR3) and maize (ZmDUR3) (Wang et al. 2012; Zanin et al. 2014).
We were able to detect the presence a low level of CaDUR3 transcripts in most analyzed tissues of the C. arabica cv. IAPAR59, indicating that the urea transporter is differentially expressed in different tissues of this species under normal field growth conditions (Fig. 2). However, the highest level of mRNA expression was observed in the roots, similarly to previous studies in other species (Zanin et al. 2014). The higher expression of this gene in root tissues most probably reflects its involvement in the mechanisms of N acquisition by the roots.
Although the roots absorb the N available in the soil solution (Witte 2011), some studies suggest that urea can be translocated from the roots to other parts of plants (Gerendás et al. 1998; Mérigout et al. 2008). In this study, changes in the transcript level of CaDUR3 in leaves of three C. arabica cultivars under nitrogen starvation were detected (Fig. 3). We suggesting that this transporter has a great potential to be considered in breeding for optimization of urea use between different genotypes and foliar fertilization. Conceivably, genotypes maintaining higher CaDUR3 gene expression may have better performance under low N availability conditions.
By observing the expression data in different coffee tissues, particularly in leaves (Fig. 2 and 3), we can reinforce the role of the CaDUR3 gene in the process of remobilization of N in coffee tissues under stress conditions. As this element is easily redistributed in plants via phloem in the form of amino acids when the N supply is not sufficient, the N of the old leaves is remobilized to other leaves and tissues (Matiz et al. 2019). Indeed, other studies have reported that DUR3 triggered a mechanism of endogenous urea remobilization during leaf senescence and germination of seeds (Bohner et al. 2015; Liu et al. 2015). Additionally, Beier et al. (2019) suggested in their study that DUR3 participates in the N redistribution from leaves to panicles in rice.
Our study revealed a differentiated relationship of CaDUR3 gene expression with urea content present in leaves of cv. IAPAR59 (Figs. 3, 5). Possibly, the urea was rapidly assimilated and therefore there was no difference expressive in content. Additionally, we can suggest that the coffee tree also showed a "facilitated diffusion" in the acquisition of urea under stress conditions. Our hypothesis is that the CaDUR3 in coffee plants may be the limiting transporter for absorption of urea through the roots to the leaves. It is important to emphasize that the limiting factors for use efficiency of urea may be associated with its translocation and assimilation capacities. In the plant cell, the assimilation of urea is possibly slower than for ammonium or nitrate, as it depends on the action of the enzyme urease (Wang et al. 2008). Moreover, when perturbation occurs in N metabolism, urea levels may deliberately increase and be translocated to other tissues (Masclaux-Daubresse et al. 2010).
The expression of the CaDUR3 gene increased almost fourfold in roots of cv. IAPAR59 after a prolonged N-deficiency period (Fig. 4), showing that DUR3 is not only up-regulated by urea, but also by extended periods of nutrient depletion.
Drought, salinity and extreme temperatures are among the main environmental restrictions for coffee crop cultivation. Under such conditions, changes occur in the plant's ability to absorb, transport and assimilate the ions needed for its growth, including nitrogen.
Here, we were able to show that the CaDUR3 is differentially modulated in coffee plants under the drought, heat and saline stresses (Fig. 3a–c). It is well known that abiotic stresses lead to major changes in nitrogen metabolism and are accompanied by significant changes in the expression of various genes involved in the plant's ability to absorb, transport and assimilate this nutrient. Previous research showed that tonoplast intrinsic proteins (TIPs), and aquaporins that facilitate the movement of low-molecular weight molecules such as urea, boron and ammonia, among others (Chaumont and Tyerman 2017), play a role in the response to abiotic stresses.
Few studies to date have attempted to evaluate genes related to mineral nutrient uptake and defense response to pathogens, especially in perennial crops. However, it is known that nutritional imbalances associated with N can influence negatively to increase of disease in plants (Roca et al. 2018; Thalineau et al. 2018). Also, studies have demonstrated that the source of N can also influence the severity of a disease (Pageau et al. 2006; Gupta et al. 2013). Here, we showed that the expression of CaDUR3 was higher in H. vastratix infected leaves of the susceptible C. arabica cultivar (CV 99) when compared with the tolerant genotype (IAPAR59) in almost all sampling points during infection (Fig. 7). This may indicate that the higher level of expression of CaDUR3 gene may reflect the impaired physiological and metabolic status of the susceptible genotype during infection (Baba et al. 2020).
Overall, in this study we demonstrated the multiplicity of factors that modify the mRNA abundance of the CaDUR3 gene. Although DUR3 play a fundamental role in the high affinity urea acquisition by the roots, the CaDUR3 was shown to be transcriptionally present in other vegetative and reproductive tissues of C. arabica. Here, we also showed that this gene was upregulated by exposure of C. arabica plants to different abiotic stresses and depending on the genotype resistance, to H. vastratix infection. From an agricultural point of view, this gene may be considered a strong candidate for coffee breeding programs aiming at the efficient use of nitrogen with the objective of increasing the quality and production of coffee fruits.
Conclusions
To our knowledge, this is the first study on the transcriptional changes of the DUR3 gene in C. arabica plants. Our expressional analysis of CaDUR3 has provided new understanding for the molecular functions of this gene in C. arabica plants, especially under biotic and abiotic stresses. Considering the importance of urea as a source of nitrogen in coffee production, and that its absorption may be dependent on the combined performance of high transport (DUR3) and low affinity systems, further analysis using heterologous gene expression would be essential for determining the contribution of each transporter system in plants subject to abiotic and biotic stresses under field conditions.
Acknowledgments
This research was supported by of the Brazilian Coffee Research Consortium and National Council of Technological and Scientific Development (CNPq). Special acknowledge to Dr. Anderson Rotter Meda for participating in the beginning of these studies.
Compliance with ethical standards
Conflict of interest
The author declare that they have no conflict of interest.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25(17):3389–3402. doi: 10.1093/nar/25.17.3389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arnhold E. Package in the R environment for analysis of variance and complementary analyses. Braz J Vet Res Anim Sci. 2013;50(6):488–492. [Google Scholar]
- Baba VY, Braghini MT, dos Santos TB, de Carvalho K, Soares JDM, Ivamoto-Suzuki ST, Maluf MP, Padilha L, Paccola-Meirelles LD, Pereira LF, Domingues DS. Transcriptional patterns of Coffea arabica L. nitrate reductase, glutamine and asparagine synthetase genes are modulated under nitrogen suppression and coffee leaf rust. PeerJ. 2020;8:e8320. doi: 10.7717/peerj.8320. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bawin Y, Ruttink T, Staelens A, Haegeman A, Stoffelen P, Mwanga JCIM, Roldán-Ruiz I, Honnay O, Janssens SB (2020) Phylogenomic analysis clarifies the evolutionary origin of Coffea arabica L. bioRxiv. 10.1111/jse.12694
- Beier MP, Fujita T, Sasaki K, Kanno K, Ohashi M, Tamura W, Konishi N, Saito M, Imagawa F, Ishiyama K, Miyao A, Yamaya T, Kojima S. The urea transporter DUR3 contributes to rice production under nitrogen-deficient and field conditions. Physiol Plant. 2019;167(1):75–89. doi: 10.1111/ppl.12872. [DOI] [PubMed] [Google Scholar]
- Bohner A, Kojima S, Hajirezaei M, Melzer M, von Wirén N. Urea retranslocation from senescing Arabidopsis leaves is promoted by DUR3-mediated urea retrieval from leaf apoplast. Plant J. 2015;81:377–387. doi: 10.1111/tpj.12740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Box GEP, Cox DR. An analysis of transformations (with discussion) R Stat Soc Ser B Methodol. 1964;26:211–252. [Google Scholar]
- Bloom AJ. The increasing importance of distinguishing among plant nitrogen sources. Curr Opin Plant Biol. 2015;25:10–16. doi: 10.1016/j.pbi.2015.03.002. [DOI] [PubMed] [Google Scholar]
- Chaumont F, Tyerman SD. Plant aquaporins from transport to signaling. Cham: Springer; 2017. [Google Scholar]
- Clark RB. Characterization of phosphatase of intact maize roots. J Agr Food Chem. 1975;23:458–460. doi: 10.1021/jf60199a002. [DOI] [PubMed] [Google Scholar]
- Davis AP, Tosh J, Ruch N, Fay MF. Growing coffee: Psilanthus (Rubiaceae) subsumed on the basis of molecular and morphological data; implications for the size, morphology, distribution and evolutionary history of Coffea. Bot J Linn Soc. 2011;167:357–377. [Google Scholar]
- de Carvalho K, Bespalhok Filho JC, dos Santos TB, de Souza SGH, Vieira LGE, Pereira LFP, Domingues DS. Nitrogen starvation, salt and heat stress in coffee (Coffea arabica L.): identification and validation of new genes for qPCR normalization. Mol Biotechnol. 2013;53:315–325. doi: 10.1007/s12033-012-9529-4. [DOI] [PubMed] [Google Scholar]
- De Michele R, Loqué D, Lalonde S, Frommer WB. Ammonium and urea transporter inventory of the Selaginella and Pyscomitrella genomes Front. Plant Sci. 2012;3:62. doi: 10.3389/fpls.2012.00062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- dos Santos TB, Budzinski IG, Marur CJ, Petkowicz CL, Pereira LF, Vieira LG. Expression of three galactinol synthase isoforms in Coffea arabica L. and accumulation of raffinose and stachyose in response to abiotic stresses. Plant Physiol Biochem. 2011;49:441–448. doi: 10.1016/j.plaphy.2011.01.023. [DOI] [PubMed] [Google Scholar]
- dos Santos TB, Lima JE, Felicio MS, Soares JDM, Domingues DS. Genome-wide identification, classification and transcriptional analysis of nitrate and ammonium transporters in Coffea. Genet Mol Biol. 2017;40:346–359. doi: 10.1590/1678-4685-GMB-2016-0041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- dos Santos TB, Lima RBD, Nagashima GT, Petkowicz CLDO, Carpentieri-Pípolo V, Pereira LFP, Domingues DS, Vieira LGE. Galactinol synthase transcriptional profile in two genotypes of Coffea canephora with contrasting tolerance to drought. Genet Mol Biol. 2015;38(2):182–190. doi: 10.1590/S1415-475738220140171. [DOI] [PMC free article] [PubMed] [Google Scholar]
- dos Santos TB, Soares JDM, Lima JE, Silva JC, Ivamoto ST, Baba VY, Souza SGH, Lorenzetti APR, Paschoal AR, Meda AR, Nishiyama Júnior MY, De Oliveira ÚC, Mokochinski JB, Guyot R, Junqueira-de-Azevedo ILM, Figueira AVO, Mazzafera P, Júnior OR, Vieira LGE, Pereira LFP, Domingues DS. An integrated analysis of mRNA and sRNA transcriptional profiles in Coffea arabica L. roots: insights on nitrogen starvation responses. Funct Integr Genom. 2019;19(1):151–169. doi: 10.1007/s10142-018-0634-8. [DOI] [PubMed] [Google Scholar]
- Doyle JJ, Doyle JL. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull. 1987;19:11–15. [Google Scholar]
- Elberry HM, Majumdar ML, Cunningham TS, Sumrada RA, Cooper TG. Regulation of the urea active transporter gene (DUR3) in Saccharomyces cerevisiae. J Bacteriol. 1993;175(15):4688–4698. doi: 10.1128/jb.175.15.4688-4698.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fan X, Naz M, Fan X, Xuan W, Miller AJ, Xu G. Plant nitrate transporters: from gene function to application. J Exp Bot. 2017;68:2463–2475. doi: 10.1093/jxb/erx011. [DOI] [PubMed] [Google Scholar]
- Fridell G. Coffee. New York: Wiley; 2014. [Google Scholar]
- Gerendás J, Zhu Z, Sttelmacher B. Influence of nitrogen and Ni supply on nitrogen metabolism and urease activity in rice (Oryza sativa L.) J Exp Bot. 1998;49:1545–1554. [Google Scholar]
- Gupta KJ, Brotman Y, Segu S, Zeier T, Zeier J, Persijn ST, Cristescu SM, Harren FJM, Bauwe H, Fernie AR, Kaiser WM, Mur LAJ. The form of nitrogen nutrition affects resistance against Pseudomonas syringae pv. phaseolicola in tobacco. J Exp Bot. 2013;64:553–568. doi: 10.1093/jxb/ers348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karamos RE, Hanson K, Stevenson FC. Nitrogen form, time and rate of application, and nitrification inhibitor effects on crop production. Can J Plant Sci. 2014;94:425–432. [Google Scholar]
- Kojima S, Bohner A, Gassert B, Yuan L, von Wirén N. AtDUR3 represents the major transporter for high-affinity urea transport across the plasma membrane of nitrogen-deficient Arabidopsis roots. Plant J. 2007;52:30–40. doi: 10.1111/j.1365-313X.2007.03223.x. [DOI] [PubMed] [Google Scholar]
- Lashermes P, Combes MC, Robert J, Trouslot P, D’Hont A, Charrier A. Molecular characterisation and origin of the Coffea arabica L. genome. Mol Gen Genet. 1999;261:259–266. doi: 10.1007/s004380050965. [DOI] [PubMed] [Google Scholar]
- Lashermes P, Paczek V, Trouslot P, Combes MC, Couturon E, Charrier A. Single-locus inheritance in the allotetraploid Coffea arabica L. and interspecific hybrid C. arabica x C. canephora. J Hered. 2000;91:81–85. doi: 10.1093/jhered/91.1.81. [DOI] [PubMed] [Google Scholar]
- Liu LH, Ludewig U, Frommer WB, von Wirén N. AtDUR3 encodes a new type of high-affinity urea/H + symporter in Arabidopsis. Plant Cell. 2003;15:790–800***. doi: 10.1105/tpc.007120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu GW, Sun AL, Li DQ, Athman A, Gilliham M, Liu LH. Molecular identification and functional analysis of a maize (Zea mays) DUR3 homolog that transports urea with high affinity. Planta. 2015;241:861–874. doi: 10.1007/s00425-014-2219-7. [DOI] [PubMed] [Google Scholar]
- Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2−CT method. Methods. 2001;25:402–408. doi: 10.1006/meth.2001.1262. [DOI] [PubMed] [Google Scholar]
- Masclaux-Daubresse C, Daniel-Vedele F, Dechorgnat J, Chardon F, Gaufichon L, Suzuki A. Nitrogen uptake, assimilation and remobilization in plants: challenges for sustainable and productive agriculture. Ann Bot. 2010;105:1141–1157. doi: 10.1093/aob/mcq028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matiz A, Mioto PT, Mercier H (2019) Urea in plants: metabolic aspects and ecological implications 1–31. In: Progress in botany. Springer, Berlin, Heidelberg
- Mérigout P, Lelandais M, Bitton F, Renou JP, Briand X, Meyer C, Daniel-Vedele F. Physiological and transcriptomic aspects of urea uptake and assimilation in Arabidopsis plants. Plant Physiol. 2008;147:1225–1238. doi: 10.1104/pp.108.119339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mondego JMC, Vidal RO, CarazzolleMF TEK, Parizzi LP, Costa GGL. An EST-based analysis identifies new genes and reveals distinctive gene expression features of Coffea arabica and Coffea canephora. BMC Plant Biol. 2011;11:30. doi: 10.1186/1471-2229-11-30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pageau K, Reisdorf-Cren M, Morot-Gaudry JF, Masclaux-Daubress C. The two senescence-related markers, GS1 (cytosolic glutamine synthetase) and GDH (glutamate dehydrogenase), involved in nitrogen mobilization, are differentially regulated during pathogen attack and by stress hormones and reactive oxygen species in Nicotiana tabacum L. leaves. J Exp Bot. 2006;57:547–557. doi: 10.1093/jxb/erj035. [DOI] [PubMed] [Google Scholar]
- R Core Team (2019) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna. https://www.R-project.org/
- Roca LF, Romero J, Bohórquez JM, Alcántara E, Fernández-Escobar R, Trapero A. Nitrogen status affects growth, chlorophyll content and infection by Fusicladium oleagineum in olive. Crop Prot. 2018;109:80–85. [Google Scholar]
- Ruijter JM, Ramakers C, Hoogaars WMH, Karlen Y, Bakker O, van den Hoff MJB, Moorman AFM. Amplification efficiency: linking baseline and bias in the analysis of quantitative PCR data. Nucleic Acids Res. 2009;37:e45. doi: 10.1093/nar/gkp045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scalabrin S, Toniutti L, Di Gaspero G, Scaglione D, Magris G, Vidotto M, Pinosio S, Cattonaro F, Magni F, Jurman I, Cerutti M, Liverani FS, Navarini L, Del Terra L, Pellegrino G, Ruosi MR, Vitulo N, Valle G, Pallavicini A, Graziosi G, Klein PE, Bentley N, Murray S, Solano W, Al Hakimi A, Schilling T, Montagnon C, Morgante M, Bertrand B. A single polyploidization event at the origin of the tetraploid genome of Coffea arabica is responsible for the extremely low genetic variation in wild and cultivated germplasm. Sci Rep. 2020;10(1):4642. doi: 10.1038/s41598-020-61216-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tegeder M, Masclaux-Daubresse C. Source and sink mechanisms of nitrogen transport and use. New Phytol. 2018;217:35–53. doi: 10.1111/nph.14876. [DOI] [PubMed] [Google Scholar]
- Tucker C. Coffee culture. 2. New York: Routledge; 2017. p. 188. [Google Scholar]
- Thalineau E, Fournier C, Gravot A, Wendehenne D, Jeandroz S, Truong H-N. Nitrogen modulation of Medicago truncatula resistance to Aphanomyces euteiches depends on plant genotype. Mol Plant Pathol. 2018;19(3):664–676. doi: 10.1111/mpp.12550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Venables WN, Ripley BD. Modern applied statistics with S. 4. New York: Springer; 2002. [Google Scholar]
- Vidal EA, Araus V, Lu C, Parry G, Green PJ, Coruzzi GM, Gutiérrez RA. Nitrate-responsive miR393/AFB3 regulatory module controls root system architecture in Arabidopsis thaliana. P Natl Acad Sci. 2010;107:4477–4482. doi: 10.1073/pnas.0909571107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vieira LGE, Andrade AC, Colombo CA, Moraes AHDA, Metha Â, Oliveira ACD, Labate CA, Marino CL, Monteiro-Vitorello CB, Monte DC, Giglioti E, Kimura ET, Romano E, Kuramae EE, Lemos EGM, Almeira ERP, Jorge EC, Albuquerque EVS, Silva FR, Vinecky F, Sawazaki HE, Dorry HFA, Carrer H, Abreu IN, Batista JAN, Teixeira JB, Kitajima JP, Xavier KG, Lima LM, Camargo LEA, Pereira LFP, Coutinho LL, Lemos MVF, Romano MR, Machado MA, Costa MMC, Sá MFG, Goldman MHS, Ferro MIT, Tinoco MLP, Oliveira MC, Sluys MAV, Shimizu MM, Maluf MP, Eira MTS, Filho OG, Arruda P, Mazzafera P, Mariani PDSC, Oliveira RLBC, Harakava R, Balbao SF, Tsai SM, Mauro SMZ, Santos SN, Siqueira WJ, Costa GGL, Formighieri EF, Carazzolle MF, Pereira GAG. Brazilian coffee genome project: an ESTbased genomic resource. Braz J Plant Physiol. 2006;18:95–108. [Google Scholar]
- Wang WH, Köhler B, Cao FQ, Liu LH. Molecular and physiological aspects of urea transport in higher plants. Plant Sci. 2008;175:467–477. [Google Scholar]
- Wang WH, Köhler B, Cao FQ, Liu GW, Gong YY, Sheng S, Song QC, Cheng XY, Garnett T, Okamoto M, Qin R, Mueller-Roeber B, Tester M, Liu LH. Rice DUR3 mediates high-affinity urea transport and plays an effective role in improvement of urea acquisition and utilization when expressed in Arabidopsis. New Phytol. 2012;193:432–444. doi: 10.1111/j.1469-8137.2011.03929.x. [DOI] [PubMed] [Google Scholar]
- Wang WH, Liu GW, Cao FQ, Cheng XY, Liu BW, Liu LH. Inadequate root uptake may represent a major component limiting rice to use urea as sole nitrogen source for growth. Plant Soil. 2013;363(1–2):191–200. [Google Scholar]
- Weatherburn MW. Phenol-hypochlorite reaction for determination of ammonia. Anal Chem. 1967;39:971–974. [Google Scholar]
- Witte CP. Urea metabolism in plants. Plant Sci. 2011;180:431–438. doi: 10.1016/j.plantsci.2010.11.010. [DOI] [PubMed] [Google Scholar]
- Yang H, Menz J, Haussermann I, Benz M, Fujiwara T, Ludewig U. High and low affinity urea root uptake: involvement of NIP5;1. Plant Cell Physiol. 2015;56(8):1588–1597. doi: 10.1093/pcp/pcv067. [DOI] [PubMed] [Google Scholar]
- Yu Q, Guyot R, de Kochko A, Byers A, Navajas-Pérez R, Langston BJ, Dubreuil-Tranchant C, Paterson AH, Poncet V, Nagai C, Ming R. Microcolinearity and genome evolution in the vicinity of an ethylene receptor gene of cultivated diploid and allotetraploid coffee species (Coffea) Plant J. 2011;67:305–317. doi: 10.1111/j.1365-313X.2011.04590.x. [DOI] [PubMed] [Google Scholar]
- Zanin L, Tomasi N, Wirdnam C, Meier S, Komarova NY, Mimmo T, Cesco S, Rentsch D, Pinton R. Isolation and functional characterization of a high affinity urea transporter from roots of Zea mays. BMC Plant Biol. 2014;14:222. doi: 10.1186/s12870-014-0222-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang L, Yan J, Vatamaniuk OK, Du X. CsNIP2;1 is a plasma membrane transporter from Cucumis sativus that facilitates urea uptake when expressed in Saccharomyces cerevisiae and Arabidopsis thaliana. Plant Cell Physiol. 2016;57(3):616–629. doi: 10.1093/pcp/pcw018. [DOI] [PubMed] [Google Scholar]



