Abstract
Background and objectives
Based on some previous observational studies, there is a theory that suggests a potential relationship between Helicobacter pylori (H. pylori) colonization and celiac disease (CeD); however, the type of this relationship is still controversial. Therefore, we aimed to conduct a systematic review and meta-analysis to explore all related primary studies to find any possible association between CeD and human H. pylori colonization.
Data sources
Studies were systematically searched and collected from four databases and different types of gray literature to cover all available evidence. After screening, the quality and risk of bias assessment of the selected articles were evaluated.
Synthesis methods
Meta-analysis calculated pooled odds ratio (OR) on the extracted data. Furthermore, heterogeneity, sensitivity, subgroups, and publication bias analyses were assessed.
Results
Twenty-six studies were included in this systematic review, with a total of 6001 cases and 135512 control people. The results of meta-analysis on 26 studies showed a significant and negative association between H. pylori colonization and CeD (pooled OR = 0.56; 95% CI = 0.45–0.70; P < 0.001), with no publication bias (P = 0.825). The L’Abbé plots also showed a trend of having more H. pylori colonization in the control group. Among subgroups, ORs were notably different only when the data were stratified by continents or risk of bias; however, subgroup analysis could not determine the source of heterogeneity.
Conclusions
According to the meta-analysis, this negative association might imply a mild protective role of H. pylori against celiac disease. Although this negative association is not strong, it is statistically significant and should be further considered. Further investigations in both molecular and clinic fields with proper methodology and more detailed information are needed to discover more evidence and underlying mechanisms to clear the interactive aspects of H. pylori colonization in CeD patients.
Systematic review registration number (PROSPERO)
CRD42020167730 https://www.crd.york.ac.uk/prospero/display_record.php?RecordID=167730.
Introduction
Celiac disease (CeD) is an inflammatory autoimmune disease with small intestine enteropathy disorder triggered by gluten ingestion [1]. CeD is a multifactorial disease that has risen from the interaction between gluten and immune, genetic, and environmental factors and can be diagnosed at any age [2]. Serological prevalence of CeD is around 1% in the general population; with a slight difference between Western and Eastern countries (1% vs. 1.6%) [3–5]. The incidence and prevalence of CeD have increased over the years that could be due to better diagnostic tools, screening of high-risk individuals, and general awareness or due to the real increasing rate of the disease based on the alteration of lifestyle or gut microbiota [1,6].
H. pylori is one of the most common chronic bacterial infections worldwide, it can cause significant gastroduodenal diseases [7]. The infection affects up to 90% of the population in developing countries and around 40% in developed countries. Generally, half of the global population can have H. pylori colonization [8]. There are controversies about the probable relation between H. pylori colonization and autoimmune diseases [9]. H. pylori can modulate some immune responses and play a protective role against some other diseases such as inflammatory bowel disease (IBD) [10]. Otherwise, in genetically predisposed persons, H. pylori can influence immune responses and cause microscopic duodenal inflammation and can be related to more severe damage associated with some gastrointestinal diseases [11–14]. In this regard, some studies reported no relationship between H. pylori and CeD [15–17]; whereas, others stated that H. pylori can protect against CD [14,18]. There are different hypotheses and justification about the mechanism of interaction between H. pylori and CeD.
The hygiene hypothesis is one of the explanations for the protective role of H. pylori infection against CeD and claims that lower exposure to some infectious agents in childhood leads to an increase in the risk of autoimmune diseases such as asthma, IBD, allergic rhinitis, and eczema [13,19–22]. Different mechanisms are suggested for this hypothesis. One of them is T helper (Th) 1/Th2 deviation, which suggests that the presence of infection causes the balance of Th1/Th2 to move towards an immunosuppression state. T regulatory (Treg) lymphocytes might be induced or augmented (a phenomenon called bystander suppression) due to infection [19,22,23]. Also, there are more suggested mechanisms for this hypothesis, such as antigenic competition/homeostasis, non-antigenic ligands, and gene-environment interaction [19,22,23].
Another hypothesis is the protective role of H. pylori on the modification of gluten digested by proteases secretion, and increase of the pH of the gastric lumen, which reduces the immunogenicity of gluten [13,14,19,22–24]. Some studies suggested another theory and supported the idea that a specific virulence factor, such as CagA in H. pylori, may give the bacterium a protective role in CeD. This function of CagA may be due to the association with Treg cells in the immunoregulation mechanism [17,24–26].
The fourth important theory is related to empirical antibiotic treatment in patients suspicious to gastrointestinal bacterial overgrowth, which was eventually diagnosed as CeD. This treatment may lead to the unintentional removal of H. pylori [27–29].
Otherwise, some researchers claimed that H. pylori may be involved in the development of CeD. Mooney et al. mentioned that maybe H. pylori acts as an environmental trigger for CeD, as observed in Campylobacter infection, in a study of the US military [21,30]. However, the mechanism of this act is not properly defined. Furthermore, a study at Colombia University showed increased intraepithelial lymphocytes in the duodenal mucosa in patients with H. pylori infection that could be reversed by the eradication of H. pylori [31].
Up to now and based on our search, no systematic review and meta-analysis have been conducted about this subject. Due to the inconsistency of the previous primary studies, we aimed to prepare a systematic review to scrutinize any possible association between H. pylori colonization and CeD.
Materials and methods
Eligibility criteria
The design of the present study was based on the methodology of the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guideline [32]. This review is registered in the international prospective register of systematic reviews (PROSPERO) with registration number CRD42020167730 (Available: https://www.crd.york.ac.uk/prospero/display_record.php?RecordID=167730).
Definition
Celiac disease
CeD patients were defined if they were positive for CeD-specific antibodies (comprise autoantibodies against TG2, including endomysial antibodies (EMA), and antibodies against deamidated forms of gliadin peptides (DGP)), associated with biopsy-proven histopathology or positive genetic tests: HLA-DQ2 or HLA-DQ8 haplotypes in the primary studies.
Controls
Based on the definitions in the primary studies, controls consisted of those apparently healthy people or celiac disease was ruled out through negative serology/biopsy.
Inclusion/Exclusion criteria
Inclusion criteria included: 1) Case-control, cross-sectional, suitable cohort, and brief-report studies that contain evidence about the relationship between H. pylori and CeD, 2) Human clinical study, 3) Having at least one parameter of the relative risk effect sizes such as risk ratio (RR), odds ratio (OR), and hazard ratio (HR) or can be calculated based on the information of H. pylori status in both case and control groups.
The status of H. pylori should be detected by urea breath test (UBT), rapid urease test (RUT), culture, enzyme-linked immunosorbent assay (ELISA), histology, immunohistochemistry (IHC), and Polymerase chain reaction (PCR) methods. Furthermore, the CeD should be confirmed by endoscopy and biopsy ± serology tests and/or HLA DQ2/DQ8 genotyping. Articles without any of this information were excluded from further analysis.
Sources and search strategy
All relevant articles, published from 1990 to the end of the second month of 2021 were collected from the electronic database of PubMed, ProQuest, Scopus, and Web of Science. The ProQuest database was also reviewed for related dissertations, manually. Other related protocols were manually searched in PROSPERO. Besides, to expand the scope of the search and find any additional studies, we reviewed the different types of grey literatures such as meeting and conference abstracts for relevant articles (including Digestive Disease Week® (DDW), United European Gastroenterology (UEG), and American College of Gastroenterology). Moreover, we searched manually some key journals in the CeD and H. pylori fields, including "World Journal of Gastroenterology", "Journal of Pediatric Gastroenterology and Nutrition", "Gut", "Gastroenterology", and "Helicobacter". In the end, we performed a manual search in the references of the selected articles. We did not impose geographical or linguistic restrictions on our search, and non-English studies with English abstracts were also included.
The Medical subject heading (MeSH) database was used to find various terms of CeD and H. pylori. Two main keywords were “Helicobacter pylori” and “Celiac”. For better searching in the databases, we made syntaxes from a combination of free-text method, MeSH terms, the keywords, and Boolean operators (AND/OR/NOT). Moreover, calculating NNR (number need to read) helped us in evaluating the output of syntax and dedicating. The following syntax was applied in PubMed and adjusted for each search engine based on its search guidelines (Celiac OR “Celiac disease” OR Coeliac OR “Coeliac disease” OR (Disease AND Celiac) OR (Disease AND Coeliac) OR “Gluten Enteropathy” OR (Enteropathy AND Gluten) OR "Gluten Enteropathies" OR (Enteropathies AND Gluten) OR “Gluten-Sensitive Enteropathy” OR (Enteropathy AND Gluten-Sensitive) OR “Gluten-Sensitive Enteropathies” OR (Enteropathies AND Gluten-Sensitive) OR (Sprue AND Celiac) OR (Sprue AND Coeliac) OR (Sprue AND Nontropical) OR “Nontropical Sprue” OR “Celiac Sprue” OR “Coeliac Sprue” OR “Sprue” “Endemic Sprue” OR “Gluten Intolerance”) AND ("Helicobacter pylori" OR "Campylobacter pylori" OR "Campylobacter pyloridis" OR "Campylobacter pyloris" OR "Helicobacter nemestrinae" OR "Helicobacter infection" OR "HP infection" OR "HP infections" OR "Helicobacter infections" OR "Helicobacter colonization" OR "HP colonization" OR "Helicobacter colonization" OR "H Pylori" OR "enterohepatic Helicobacter" OR "EHS" OR "Helicobacter") AND 1990/01/01:2021/02/20[dp].
The search outputs were exported into the Endnote software (Version X7; Thompson Reuters Corporation, Toronto, ON, Canada) to remove any duplicate and screen articles. The first step of screening for determining the eligible primary articles was done based on the titles and abstracts. Two independent reviewers (A.A. and A.F.) evaluated the selected full-text articles and separately classified them into three relevant, irrelevant, and unsure groups, based on the eligibility criteria. Any discrepancy was supervised by a third reviewer (S.B.) and resolved via a consensus. If there was still a doubt in the selection of a study, the whole team made the final decision.
Quality and risk of bias assessment
The quality and risk of bias assessment of the selected articles were independently evaluated by two reviewers (Z.N. and N.G.), using the standard checklist of the Newcastle-Ottawa Scale (NOS) form [33]. The studies were classified into three poor, fair, and good categories based on getting a score between zero and eight in the selection, comparability, and outcome/exposure domains based on the standard guideline. Consensus and the opinion of the third reviewer (S.B) resolved any disagreements between the two reviewers.
Data extraction
Two authors (Z.N, F.A) independently performed data extraction from each paper based on a defined protocol. The data were divided into three categories and included 1) General information: the first author’s names, year of publication, journal names, country, and region; 2) The risk of bias assessment; 3) Study setting: study design, study duration, sample size and total population recruited in each study, inclusion and exclusion criteria, age group and age range, sex, the definition of CeD, source of the data, diagnostic methods, OR and 95% CI, and the ethical approval. In cases of any missing or additional necessary data, we further sent an e-mail to each author, in charge of the related study.
Data synthesis and analysis
We used the R software environment, version 4.0.2, and the “meta” package to calculate pooled OR and 95% CIs for each study. Based on the methodological heterogeneity between the studies, the random-effect model (REM) was used for the combination method. The size of the combined effect was calculated and displayed in a forest plot. The standard chi-square test (Q Cochrane test) and I2 scale evaluated the heterogeneity between studies. Combinations with an I2 scale equal or more than 70% assumed as a severe heterogeneity. In addition, a Drapery plot was drawn to show the P-value functions for the included studies, as well as pooled estimates, and to provide two-sided confidence intervals for all possible alpha levels (confidence interval function) [34,35]. For finding the cause(s) of heterogeneity, subgroup and sensitivity analyses were used. The studies were stratified based on each factor, including the median of publication year, overall age group of the participants, continent and regions of the studies, H. pylori detection method, the risk of bias assessment, as well as sampling quality. In the last subgroup, sampling, the articles were categorized into two groups (I) “Appropriate sampling” or (II) “Not appropriate sampling” groups. The studies were located in the “Not appropriate sampling” group, if the number of participants in each case and control group was less or equal than 50 people or the ratio of control to the cases was more than four times or less than 90%.
The funnel chart, followed by Begg’s test analyzed possible publication bias. Baujat and L’Abbé plots were used to show the effect of each study on the heterogeneity and overall influence of the results [36–38]. Finally, a one-out remove method was used to detect the influence of each study on the overall pooled OR.
Results
Search results
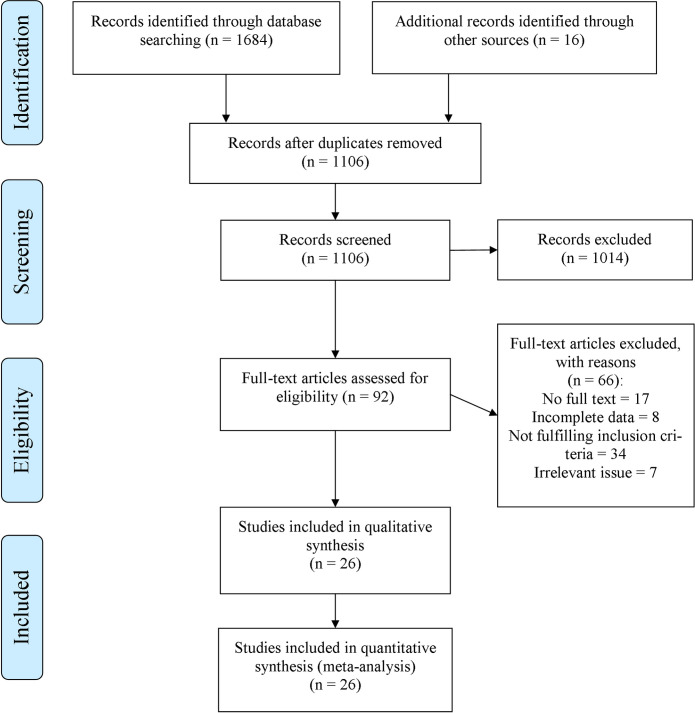
In total, the general searching step obtained 1732 papers. After removing duplicates, 1106 studies were screened, based on their titles and abstracts. Among them, 1014 articles were excluded due to not meeting the inclusion criteria. Therefore, the full-texts of 92 remained articles were completely evaluated. Finally, 26 studies were included in the systematic review and also meta-analysis (kappa (κ) = 0.84) [11,13–18,21,24–27,39–52], and the others were excluded due to a reason mentioned in Fig 1 (Fig 1). In the end, there was no suitable cohort study in the included studies.
Fig 1. Flow diagram of literature search, screening, and selection of studies for review and meta-analysis.
Characteristics of the included articles
The designs of all the included studies were case-control, with a total of 6001 cases and 135512 control people. Among the studies Lebwohl et al. had the highest study population with 130308 participants (included 2689 cases and 127619 controls) [14]. Most studies were from Turkey (N = 6) [15–18,51,52] and Italy (N = 5) [11,27,44,45,48]. Some studies had been performed only on children [13,15–17,21,44,47,51,52] or adults [18,25,27,39,43,45,48], the others evaluated both age groups [11,14,42,49,50]. Table 1 shows the details of the included articles. The NOS assessment showed that only 19.2% of the studies had good quality.
Table 1. Main characteristics of the included studies in the meta-analysis on the Helicobacter pylori infection and celiac disease (CeD).
| First author | Year | Journal title | Country | Definition of CeD | Risk of bias assessment | Case-number | Control—number | Laboratory tests of H. pylori |
|---|---|---|---|---|---|---|---|---|
| Karttunen | 1990 | Journal of Clinical Pathology | Finland | Registry | Fc | 27 | 27 | IHCf |
| Crabtree | 1992 | Journal of Clinical Pathology | England | Endoscopy + Serology + Positive response to GFDb | Pd | 99 | 250 | ELISAg |
| Feeley | 1998 | Journal of Clinical Pathology | Ireland | Registry | F | 70 | 70 | IHC |
| Diamantti | 1998 | Intestinal Disorders (poster) | Argentina | Endoscopy | P | 108 | 113 | IHC |
| Diamantti | 1999 | The American Journal of Gastroenterology | Argentina | Endoscopy + Serology + Positive response to GFD | P | 104 | 75 | IHC |
| Luzza | 1999 | Journal of Pediatric Gastroenterology and Nutrition | Italy | Endoscopy+ Positive response to GFD | Ge | 81 | 81 | IHC + RUTh + ELISA |
| Konturek | 2000 | The American Journal of Gastroenterology | Germany | Endoscopy + Serology | F | 91 | 40 | ELISA |
| Ciacci | 2000 | European Journal of Gastroenterology and Hepatology | Italy | Endoscopy | P | 187 | 76 | IHC + ELISA |
| Dettori | 2001 | Digestive and Liver Disease (Poster) | Italy | Endoscopy + Serology | G | 152 | 306 | UBTi + IHC |
| Aydogdu | 2008 | Scandinavian Journal of Gastroenterology | Turkey | Endoscopy + Serology | P | 96 | 235 | IHC |
| Rostami-Nejad | 2009 | Revista espanola de enfermedades digestivas | Iran | Endoscopy + Serology | P | 28 | 422 | IHC |
| Rostami-Nejad | 2011 | Archives of Iranian Medicine | Iran | Endoscopy + Serology | P | 24 | 226 | IHC |
| Lebwohl | 2013 | American Journal of Epidemiology | American | Endoscopy | F | 2689 | 127619 | IHC |
| Simondi | 2015 | Clinics and Research in Hepatology and Gastroenterology | Italy | Endoscopy + Serology + HLA DQ2/DQ8 genotyping | F | 73 | 404 | UBT + IHC |
| Jozefczuk | 2015 | European Review for Medical and Pharmacological Science | Poland | Endoscopy + Serology | F | 74 | 296 | UBT |
| Lasa | 2015 | Arquivos de Gastroenterologia | Argentina | Endoscopy + Serology | G | 72 | 240 | IHC |
| Jozefczuk | 2016 | European Review for Medical and Pharmacological Science | Poland | Endoscopy + Serology | F | 76 | 84 | UBT + ELISA |
| Uyanikoglu | 2016 | Euroasian Journal of Hepato-Gastroenterology | Turkey | Endoscopy | F | 31 | 592 | IHC |
| Narang | 2017 | Journal of Gastroenterology and Hepatology | India | Endoscopy + Serology + Positive response to GFD | G | 324 | 322 | IHC + RUT |
| Broide | 2017 | AGA abstracts | Germany | Endoscopy + Serology | F | 50 | 50 | UBT + IHC |
| Lucero | 2017 | Frontiers in Cellular and Infection Microbiology | Chile | Endoscopy + Serology + HLA DQ2/DQ8 genotyping | G | 66 | 50 | IHC + RUT |
| Agin | 2019 | Archives of Medical Science | Turkey | Endoscopy + Serology | P | 256 | 1012 | IHC |
| Cam | 2018 | EHMSG a | Turkey | Endoscopy | P | 141 | 150 | UBT + IHC |
| Dore | 2018 | Helicobacter | Italy | Endoscopy + HLA DQ2/DQ8 genotyping | F | 270 | 127 | UBT + IHC |
| Bayrak | 2020 | Helicobacter | Turkey | Endoscopy + Serology | F | 482 | 2060 | UBT + IHC |
| Bayrak | 2021 | Digestive Diseases | Turkey | Endoscopy + Serology | F | 278 | 505 | UBT + IHC |
aXXXIst International Workshop on Helicobacter & Microbiota in Inflammation & Cancer;
bgluten-free diet;
cFair;
dPoor;
eGood;
fImmunohistochemistry;
gEnzyme-linked immunosorbent assay;
hRapid Urease Test;
iUrease Breath Test.
Association between H. pylori and CeD
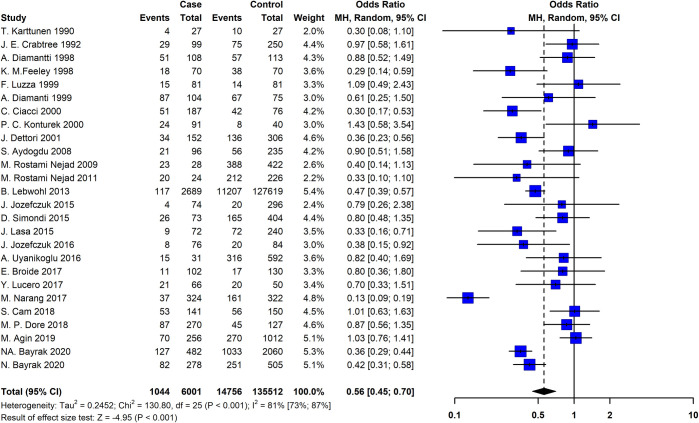
By analyzing a combination of 26 studies that addressed the association between H. pylori colonization and CeD, a negative association was found (pooled OR = 0.56; 95% CI = 0.45 − 0.70; P ≤ 0.001). The REM and Chi-squared method showed a significant statistical heterogeneity between the studies I2 = 81%, P ≤ 0.001, and X2 = 130, P ≤ 0.001, respectively (Fig 2).
Fig 2. The Forest plot of the meta-analysis on the association of the Helicobacter pylori and celiac disease.
The random-effects model was used to calculate pooled ORs with 95% CI. CI, Confidence interval; OR, Odds ratio.
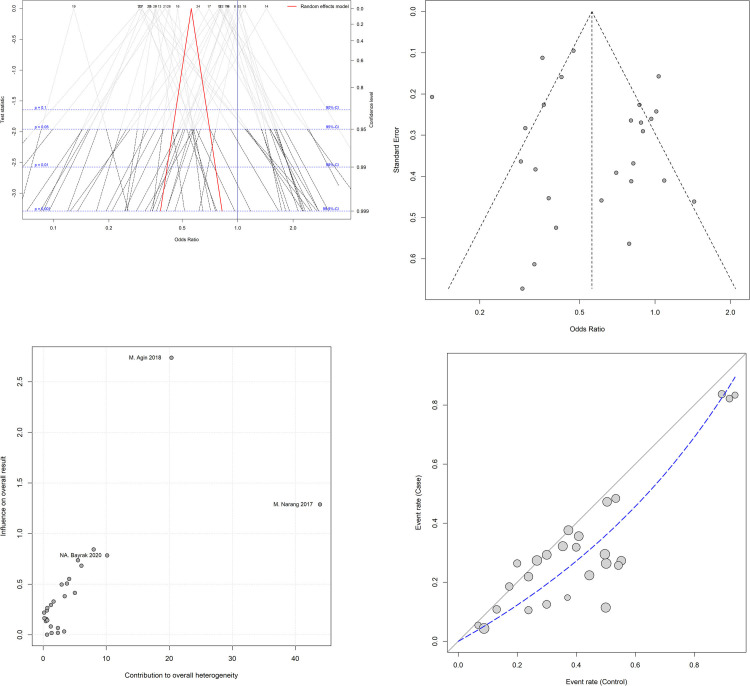
Fig 3A shows a Drapery plot, presenting that the estimated effect is less than 1 in the majority of studies, similarly as shown in the Funnel plot. As the horizontal dashed lines show the CIs for common alpha levels (0.1, 0.05, 0.01, 0.001), at any level of prediction interval (90, 95, 99, 99.9%), the main triangle line (the red-bolded line in the online version) did not intersect the null effect line (Incidence Rate Ratio (IRR) = 1). According to the results of the Funnel plot and Begg’s tests, there was no significant publication bias (P = 0.825, Fig 3B). The Baujat plot (Fig 3C) showed that the study of Narang et al. contributes more than the other studies on the overall heterogeneity and influences the pooled effect size. The L’Abbé plots (Fig 3D) also showed a trend of having more H. pylori colonization in the control group.
Fig 3.
(A) The drapery plot shows p-value functions for the included studies as well as pooled estimates and the two-sided confidence intervals for all possible alpha levels (confidence interval function). Each number represents a study. The red-bolded line (in the online version; bold line in the printed version) indicates the range of pooled odds ratio (OR) in each alpha level. (B) The Funnel chart shows no significant publication bias, as all studies are in a symmetric scheme. (C) The Baujat plot shows the effect of each study on the heterogeneity and overall influence of the results. As can be seen in the graph, the study of Narang et al. contributes more than the other studies to the overall heterogeneity. Each circle represents a study. (D) The L’Abbé plot shows more positive results for H. pylori colonization in the control group. The diagonal (x = y) oblique line represents the odds ratio (OR) equal to one. The dashed line indicates the OR of the studies. The size of each circle indicates the assigned random weight of each study. As the picture shows, most studies show an OR of less than one.
Subgroup and sensitivity analyses
The heterogeneity between studies was evaluated by multiple subgroup analyses in the six subgroups. Based on the continents of the studies, there were few studies in Asia, which made the results unreliable for the statistical analysis [13,49,50]. Europe has a higher OR (OR = 0.63, 95% CI = 0.49 − 0.82) than America (OR = 0.55, 95% CI = 0.40 − 0.76) and Asia (OR = 0.23, 95% CI = 0.10 − 0.52). However, further evaluation is needed due to the low rate of studies, especially in Asia. Other subgroups could not determine the source of heterogeneity (Table 2).
Table 2. Different subgroup analysis by stratifying the data.
| Subgroups | Na of articles | N of cases | N of controls | ORb | P-Value | Chi-Square | I2 | Tau2 | |
|---|---|---|---|---|---|---|---|---|---|
| Combination of all studies | 26 | 6001 | 135512 | 0.56 | <0.001 | 130.80 | 81 | 0.2452 | |
| Year | Before 2014 | 13 | 3756 | 129540 | 0.56 | 0.001 | 32.47 | 63 | 0.1350 |
| 2014 & after 2014 | 13 | 2245 | 5972 | 0.56 | <0.001 | 98.17 | 88 | 0.3804 | |
| Overall age | Children (Less than 18) | 10 | 1910 | 4875 | 0.57 | <0.001 | 92.64 | 90 | 0.4505 |
| Adult (More than 18) | 9 | 894 | 1919 | 0.60 | 0.004 | 22.59 | 65 | 0.1785 | |
| All age | 7 | 3197 | 128718 | 0.52 | 0.08 | 11.41 | 47 | 0.0734 | |
| Continent | Europe | 18 | 2586 | 6445 | 0.63 | <0.001 | 75.43 | 77 | 0.2138 |
| America | 5 | 3039 | 128097 | 0.55 | 0.15 | 6.81 | 41 | 0.0516 | |
| Asia | 3 | 376 | 970 | 0.23 | 0.06 | 5.67 | 65 | 0.3481 | |
| Sampling | Appropriate sampling | 14 | 1929 | 3794 | 0.57 | <0.001 | 97.94 | 87 | 0.4459 |
| Inappropriate sampling | 12 | 4072 | 131718 | 0.55 | 0.001 | 30.2 | 64 | 0.0983 | |
| H. Pylori detection tests | IHCc & ELISAd | 20 | 5307 | 134275 | 0.58 | <0.001 | 77.88 | 76 | 0.1598 |
| UBTe & RUTf | 6 | 694 | 1234 | 0.52 | <0.001 | 45.88 | 89 | 0.9442 | |
| Risk of bias | Poor | 9 | 1043 | 2559 | 0.72 | 0.009 | 20.23 | 60.0 | 0.1231 |
| Fair | 12 | 4263 | 131954 | 0.55 | 0.001 | 31 | 65.0 | 0.0873 | |
| Good | 5 | 695 | 999 | 0.39 | <0.001 | 31.84 | 87.0 | 0.6152 | |
aNumber;
bOdds ratio;
cImmunohistochemistry;
dEnzyme-linked immunosorbent assay;
eUrease Breath Test;
fRapid Urease Test.
Based on the one-out removed method, the removal of any of the studies could not significantly affect the pooled results. The sensitivity analysis was used to find possible outlier studies with more influence on the results. In this regard, the studies with poor design or not appropriate sampling (the most influential factors in the methodology) were omitted, and the overall effect size and level of heterogeneity were recalculated. The overall OR decreased from 0.56 (95% CI = 0.45 − 0.76) to 0.40 (95% CI: 0.26 − 0.62) but no change in the heterogeneity was observed. The results of the sensitivity analysis confirmed that the correct methodology was significantly associated with a lower amount of overall OR (Table 3).
Table 3. Subgroups sensitivity analysis.
| Models | Subgroups | Na of articles | N of cases | N of controls | ORb | P-Value | Chi-Square | I2 | Tau2 |
|---|---|---|---|---|---|---|---|---|---|
| All studies | 26 | 6001 | 135512 | 0.56 | <0.001 | 130.80 | 81 | 0.2452 | |
| Without “poor” quality studies | 17 | 4958 | 132953 | 0.49 | <0.001 | 75.27 | 79 | 0.1973 | |
| Europe | 13 | 1807 | 4722 | 0.57† | <0.001 | 38.58 | 67 | 0.1433 | |
| America | 3 | 2827 | 127909 | 0.47 | 0.40 | 1.84 | 0 | 0 | |
| Asia | 1 | 324 | 322 | 0.13 | NA | NA | NA | NA | |
| Appropriate sampling | 9 | 1229 | 2034 | 0.40 | <0.001 | 38.76 | 79 | 0.3288 | |
| Inappropriate sampling | 8 | 3729 | 130919 | 0.62 | <0.001 | 26.81 | 74 | 0.1195 | |
| Without “not appropriate sampling” studies | 14 | 1929 | 3794 | 0.54 | <0.001 | 97.94 | 87 | 0.4459 | |
| Poor | 5 | 700 | 1760 | 0.98 | 0.98 | 0.4 | 0 | 0 | |
| Fair | 5 | 600 | 1085 | 0.45 | 0.32 | 4.67 | 14 | 0.0200 | |
| Good | 4 | 629 | 949 | 0.34 | <0.001 | 26.14 | 89 | 0.6170 | |
| Europe | 11 | 1425 | 3119 | 0.66 | <0.001 | 37.91 | 74 | 0.1925 | |
| America | 2 | 180 | 353 | 0.56 | 0.04 | 4.33 | 77 | 0.3658 | |
| Asia | 1 | 324 | 322 | 0.13 | NA | NA | NA | NA | |
| Without “poor” quality and “not appropriate sampling” studies | 9 | 1229 | 2034 | 0.40 | <0.001 | 38.76 | 79 | 0.3288 | |
| Fair | 5 | 600 | 1085 | 0.45 | 0.32 | 4.67 | 14 | 0.0200 | |
| Good | 4 | 629 | 949 | 0.34 | <0.001 | 26.14 | 89 | 0.6170 | |
| Europe | 7 | 833 | 1472 | 0.48 | 0.11 | 10.3 | 42 | 0.0691 | |
| America | 1 | 72 | 240 | 0.33 | NA | NA | NA | NA | |
| Asia | 1 | 324 | 322 | 0.13 | NA | NA | NA | NA |
anumber;
bodds ratio;
†The bolded items show a statistical difference in OR of region subgroup.
Discussion
Most information about the relationship between H. pylori colonization and CeD is inferred from case-control studies and more prospective cohort and /or systematic review are needed to provide the strongest evidence about this association. In this regard, the present study is the first systematic review and meta-analysis that aimed to clarify any possible association between H. pylori colonization and CeD. The meta-analysis showed a mild negative relationship between H. pylori and CeD, and confirmed the findings of some previous studies, showing H. pylori was significantly more prevalent in apparently healthy people than celiac patients [11,13,14].
The protective effect of H. pylori on some allergic and atopic diseases and other inflammatory diseases could be considered as starting point of the debates about the association between CeD and H. pylori [10,17,18,21].
For clearing the possible protective effect of the immune-suppressive mechanisms of H. pylori on CeD, the first step is the specification of the chronological order of CeD and H. pylori colonization. As there was not any cohort study on this subject, there is no strong evidence to show a cause and effect relationship between these two and a protective effect of H. pylori on the occurrence of CeD. Therefore, conducting a cohort study should be considered as a top priority in further investigations of this subject.
Beyond our subgroup analyses, there are more interactive aspects of the pathophysiology of CeD and pathogenesis of H. pylori that could help us to explore the causes of the heterogeneity. Both CeD and H. pylori cause inflammation and immune response in the GI tract and also H. pylori could escape human immune defense via different mechanisms that suppress both innate and adaptive immune response [53]. H. pylori can evade host immune response via activating signaling pathway, sustain intracellularly in macrophages and induction of apoptosis, inhibition of leukocyte migration, and producing arginase enzyme.
Recent studies reported that Vacuolating cytotoxin (VacA) of H. pylori has an immunomodulatory role by inhibiting integrin-linked kinase (ILK) [54]. VacA also intervenes with the antigen presentation of MHC class II [55] and it directly suppresses T-cells rather than antigen presenting cells (APC) [56]. It is possible that these suppressive mechanisms produced by H. pylori, affect the autoimmune response to gluten and reduce the level of gluten enteropathy in CeD. Considering the notion of a balance between immune suppressive mechanisms of H. pylori and autoimmune response in CeD, it seems more probable to observe a protective effect against autoimmune response among patients who are infected with a type of H. pylori.
In this systematic review and meta-analysis related to the association of H. pylori colonization and CeD, our search strategy prepared a comprehensive source of articles that contains any study with related evidence. Subgroup and sensitivity analyses showed that two factors (sampling method and the quality of article) could be more effective and important for case-control studies and it is better to be addressed in future studies. In addition, the analyses showed that this protective effect of H. pylori on CeD was more observed in Asian countries. After removing the studies with poor quality and non-appropriate sampling, the pooled OR was still statistically lower in a study from Asia, in comparison to both studies from Europe and America. However, heterogeneity was still more than 70% in this continent subgroup. Unfortunately, after removing the studies in sensitivity analysis, only one study from Asia remained, and the obtained results could not completely determine the source of heterogeneity. Here, we suggest conducting well-designed studies in Asia, especially in the East Asian population to clarify the role of ethnicity in the association of H. pylori colonization and CeD.
The present study had some limitations. First, just a few numbers of the selected articles had a good-quality method and many of them did not consider many confounding factors in the design of the study such as March type, the severity of celiac, H. pylori virulence, and IgA deficiency that could affect the results. Second, based on the results, the heterogeneity of outcomes was high, and the subgroup analysis could not recognize the source of it completely. Third, most of the studies were conducted in Western countries, especially in Turkey and Italy. There was no article from Africa. Moreover, only three articles reported the situation in Asia. Studying other ethnicities can be a suitable target for further investigations.
Conclusion
The meta-analysis of 26 clinical studies included 6001 CeD patients from both children and adult age-groups showed a mild protective role of H. pylori against celiac disease. The main obstacle on the way of placing the role of H. pylori in CeD under scrutiny is the shortage of accurate details in the clinical studies. To clear the relationship between H. pylori and CeD, it is suggested to implement cohort studies and also further primary studies with good methodology. Extending the scope of study about this subject to the molecular context may prepare more answers to the current controversies.
Supporting information
(DOC)
(DOCX)
Data Availability
The data underlying the results presented in the study are available from supporting information files.
Funding Statement
Golestan University of Medical Sciences, Gorgan, Iran has supported this study (grant number: GOUMS111462). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Lebwohl B, Sanders DS, Green PHR. Coeliac disease. Lancet. 2018;391: 70–81. 10.1016/S0140-6736(17)31796-8 [DOI] [PubMed] [Google Scholar]
- 2.Green PHR, Cellier C. Celiac disease. N Engl J Med. 2007;357: 1731–1743. 10.1056/NEJMra071600 [DOI] [PubMed] [Google Scholar]
- 3.Parzanese I, Qehajaj D, Patrinicola F, Aralica M, Chiriva-Internati M, Stifter S, et al. Celiac disease: From pathophysiology to treatment. World J Gastrointest Pathophysiol. 2017;8: 27. 10.4291/wjgp.v8.i2.27 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Singh P, Arora S, Singh A, Strand TA, Makharia GK. Prevalence of celiac disease in Asia: A systematic review and meta-analysis. J Gastroenterol Hepatol. 2016;31: 1095–1101. 10.1111/jgh.13270 [DOI] [PubMed] [Google Scholar]
- 5.Al-Toma A, Volta U, Auricchio R, Castillejo G, Sanders DS, Cellier C, et al. European Society for the Study of Coeliac Disease (ESsCD) guideline for coeliac disease and other gluten‐related disorders. United Eur Gastroenterol J. 2019;7: 583–613. 10.1177/2050640619844125 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Lebwohl B, Rubio-Tapia A. Epidemiology, Presentation, and Diagnosis of Celiac Disease. Gastroenterology. 2021;160: 63–75. 10.1053/j.gastro.2020.06.098 [DOI] [PubMed] [Google Scholar]
- 7.Klitorakis I, Stanciu C. Prevalence of Helicobacter pylori infection in patients with gastroesophageal reflux disease. Rev medico-chirurgicalǎ a Soc Medici şi Nat din Iaşi. 2010;114: 80–84. [PubMed] [Google Scholar]
- 8.Tonkic A, Tonkic M, Lehours P, Mégraud F. Epidemiology and Diagnosis of Helicobacter pylori Infection. Helicobacter. 2012;17: 1–8. 10.1111/j.1523-5378.2012.00975.x [DOI] [PubMed] [Google Scholar]
- 9.Smyk DS, Koutsoumpas AL, Mytilinaiou MG, Rigopoulou EI, Sakkas LI, Bogdanos DP. Helicobacter pylori and autoimmune disease: Cause or bystander. World J Gastroenterol. 2014;20: 613–629. 10.3748/wjg.v20.i3.613 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Shirzad-Aski H, Besharat S, Kienesberger S, Sohrabi A, Roshandel G, Amiriani T, et al. Association Between Helicobacter pylori Colonization and Inflammatory Bowel Disease: A Systematic Review and Meta-Analysis. J Clin Gastroenterol. 2020. 10.1097/MCG.0000000000001415 [DOI] [PubMed] [Google Scholar]
- 11.Ciacci C, Squillante A, Rendina D, Limauro S, Bencivenga C, Labanca F, et al. Helicobacter pylori infection and peptic disease in coeliac disease. Eur J Gastroenterol Hepatol. 2000;12: 1283–1287. 10.1097/00042737-200012120-00004 [DOI] [PubMed] [Google Scholar]
- 12.Krigel A, Turner KO, Makharia GK, Green PHR, Genta RM, Lebwohl B. Ethnic variations in duodenal villous atrophy consistent with celiac disease in the United States. Clin Gastroenterol Hepatol. 2016;14: 1105–1111. 10.1016/j.cgh.2016.04.032 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Narang M, Puri AS, Sachdeva S, Singh J, Kumar A, Saran RK. Celiac disease and Helicobacter pylori infection in children: Is there any Association? J Gastroenterol Hepatol. 2017;32: 1178–1182. 10.1111/jgh.13654 [DOI] [PubMed] [Google Scholar]
- 14.Lebwohl B, Blaser MJ, Ludvigsson JF, Green PHR, Rundle A, Sonnenberg A, et al. Decreased risk of celiac disease in patients with Helicobacter pylori colonization. Am J Epidemiol. 2013;178: 1721–1730. 10.1093/aje/kwt234 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Aydogdu S, Cakir M, Ali Yuksekkaya H, Tumgor G, Baran M, Arikan C, et al. Helicobacter pylori infection in children with celiac disease. Scand J Gastroenterol. 2008;43: 1088–1093. 10.1080/00365520802101846 [DOI] [PubMed] [Google Scholar]
- 16.Cam S, Ugras M. Children with Celiac disease have less H. pylori gastritis. Helicobacter. Wiley 111 River St, Hoboken 07030–5774, NJ USA; 2018.
- 17.Agin M, Batun I, Ozdemir S, Doran F, Tumgor G. Prevalence of Helicobacter pylori in Turkish children with celiac disease and its effect on clinical, histopathological, and laboratory parameters. Arch Med Sci AMS. 2019;15: 1475. 10.5114/aoms.2019.83699 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Uyanikoglu A, Dursun H, Yenice N. Is There any Association Between Celiac Disease and Helicobacter pylori? Euroasian J hepato-gastroenterology. 2016;6: 103. 10.5005/jp-journals-10018-1179 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Christen U, von Herrath MG. Infections and autoimmunity—good or bad? J Immunol. 2005;174: 7481–7486. 10.4049/jimmunol.174.12.7481 [DOI] [PubMed] [Google Scholar]
- 20.Brooks C, Pearce N, Douwes J. The hygiene hypothesis in allergy and asthma: an update. Curr Opin Allergy Clin Immunol. 2013;13: 70–77. 10.1097/ACI.0b013e32835ad0d2 [DOI] [PubMed] [Google Scholar]
- 21.Jozefczuk J, Bancerz B, Walkowiak M, Glapa A, Nowak J, Piescikowska J, et al. Prevalence of Helicobacter pylori infection in pediatric celiac disease. Age [years]. 2015;6: 3.0. [PubMed] [Google Scholar]
- 22.Okada H, Kuhn C, Feillet H, Bach J. The ‘hygiene hypothesis’ for autoimmune and allergic diseases: an update. Clin Exp Immunol. 2010;160: 1–9. 10.1111/j.1365-2249.2010.04139.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Bach J-F. The hygiene hypothesis in autoimmunity: the role of pathogens and commensals. Nat Rev Immunol. 2018;18: 105. 10.1038/nri.2017.111 [DOI] [PubMed] [Google Scholar]
- 24.Lasa J, Zubiaurre I, Dima G, Peralta D, Soifer L. Helicobacter pylori prevalence in patients with celiac disease: results from a cross-sectional study. Arq Gastroenterol. 2015;52: 139–142. 10.1590/S0004-28032015000200012 [DOI] [PubMed] [Google Scholar]
- 25.Konturek PC, Karczewska E, Dieterich W, Hahn EG, Schuppan D. Increased prevalence of Helicobacter pylori infection in patients with celiac disease. Am J Gastroenterol. 2000;95: 3682. 10.1111/j.1572-0241.2000.03421.x [DOI] [PubMed] [Google Scholar]
- 26.Lucero Y, Oyarzún A, O’Ryan M, Quera R, Espinosa N, Valenzuela R, et al. Helicobacter pylori cagA+ is associated with milder duodenal histological changes in Chilean celiac patients. Front Cell Infect Microbiol. 2017;7: 376. 10.3389/fcimb.2017.00376 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Simondi D, Ribaldone DG, Bonagura GA, Foi S, Sapone N, Garavagno M, et al. Helicobacter pylori in celiac disease and in duodenal intraepithelial lymphocytosis: active protagonist or innocent bystander? Clin Res Hepatol Gastroenterol. 2015;39: 740–745. 10.1016/j.clinre.2015.03.005 [DOI] [PubMed] [Google Scholar]
- 28.De Boissieu D, Chaussain M, Badoual J, Raymond J, Dupont C. Small-bowel bacterial overgrowth in children with chronic diarrhea, abdominal pain, or both. J Pediatr. 1996;128: 203–207. 10.1016/s0022-3476(96)70390-6 [DOI] [PubMed] [Google Scholar]
- 29.Teo M, Chung S, Chitti L, Tran C, Kritas S, Butler R, et al. Small bowel bacterial overgrowth is a common cause of chronic diarrhea. J Gastroenterol Hepatol. 2004;19: 904–909. 10.1111/j.1440-1746.2004.03376.x [DOI] [PubMed] [Google Scholar]
- 30.Mooney PD, Evans KE, Barrett SM, Arnaout A, Sanders DS. Re:" Decreased risk of celiac disease in patients with Helicobacter pylori colonization". Am J Epidemiol. 2014;179: 1275. [DOI] [PubMed] [Google Scholar]
- 31.Memeo L, Jhang J, Hibshoosh H, Green PH, Rotterdam H, Bhagat G. Duodenal intraepithelial lymphocytosis with normal villous architecture: common occurrence in H. pylori gastritis. Mod Pathol. 2005;18: 1134–1144. 10.1038/modpathol.3800404 [DOI] [PubMed] [Google Scholar]
- 32.Moher D, Liberati A, Tetzlaff J, Altman DG, Group P. Preferred reporting items for systematic reviews and meta-analyses: the PRISMA statement. PLoS med. 2009;6: e1000097. 10.1371/journal.pmed.1000097 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Wells G, Shea B, O’connell D, Peterson J, Welch V, Losos M, et al. The Newcastle-Ottawa Scale (NOS) for assessing the quality of nonrandomised studies in meta-analyses. Ottawa: Ottawa Hospital Research Institute. oxford. asp; 2011. [Google Scholar]
- 34.Birnbaum A. Confidence curves: An omnibus technique for estimation and testing statistical hypotheses. J Am Stat Assoc. 1961;56: 246–249. [Google Scholar]
- 35.Infanger D, Schmidt‐Trucksäss A. P value functions: An underused method to present research results and to promote quantitative reasoning. Stat Med. 2019;38: 4189–4197. 10.1002/sim.8293 [DOI] [PubMed] [Google Scholar]
- 36.Deeks JJ. Issues in the selection of a summary statistic for meta‐analysis of clinical trials with binary outcomes. Stat Med. 2002;21: 1575–1600. 10.1002/sim.1188 [DOI] [PubMed] [Google Scholar]
- 37.L’abbé Ka Detsky As, O’rourke K. Meta-analysis in clinical research. Ann Intern Med. 1987;107: 224–233. 10.7326/0003-4819-107-2-224 [DOI] [PubMed] [Google Scholar]
- 38.Baujat B, Mahé C, Pignon J, Hill C. A graphical method for exploring heterogeneity in meta‐analyses: application to a meta‐analysis of 65 trials. Stat Med. 2002;21: 2641–2652. 10.1002/sim.1221 [DOI] [PubMed] [Google Scholar]
- 39.Karttunen T, Niemelä S. Lymphocytic gastritis and coeliac disease. J Clin Pathol. 1990;43: 436. 10.1136/jcp.43.5.436 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Crabtree JE, O’Mahony S, Wyatt JI, Heatley R V, Vestey JP, Howdle PD, et al. Helicobacter pylori serology in patients with coeliac disease and dermatitis herpetiformis. J Clin Pathol. 1992;45: 597–600. 10.1136/jcp.45.7.597 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Feeley KM, Heneghan MA, Stevens FM, McCarthy CF. Lymphocytic gastritis and coeliac disease: evidence of a positive association. J Clin Pathol. 1998;51: 207–210. 10.1136/jcp.51.3.207 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Diamantti A, Maino C, Vazquez H, Pedreira S, Niveloni S, Smecuol E, et al. Celiac disease (CD), lymphocytic gastritis and Helicobacter pylori (HP) infection. A controlled study. Gastroenterology. 1998;114: A363. [Google Scholar]
- 43.Diamanti A, Maino C, Niveloni S, Pedreira S, Vazquez H, Smecuol E, et al. Characterization of gastric mucosal lesions in patients with celiac disease: a prospective controlled study. Am J Gastroenterol. 1999;94: 1313–1319. 10.1111/j.1572-0241.1999.01082.x [DOI] [PubMed] [Google Scholar]
- 44.Luzza F, Mancuso M, Imeneo M, Mesuraca L, Contaldo A, Giancotti L, et al. Helicobacter pylori infection in children with celiac disease: prevalence and clinicopathologic features. J Pediatr Gastroenterol Nutr. 1999;28: 143–146. 10.1097/00005176-199902000-00009 [DOI] [PubMed] [Google Scholar]
- 45.Dettori I, Dore MP, Marras L, Nieddu S, Manca A, Massarelli G, et al. Is there anything to the reported association between Helicobacter pylori infection and celiac disease? Gut. 2001;49: A42–A42. [Google Scholar]
- 46.Broide E, Sandbank J, Scapa E, Kimchi NA, Shapiro M, Lerner A. The immunohistochemistry profile of lymphocytic gastritis in celiac disease and Helicobacter pylori infection: interplay between infection and inflammation. Mediators Inflamm. 2007;2007. Broide E, Richter V, Shirin H. Is Helicobacter Pylori Chronic Infection Associated with the Severity of Celiac Disease in Children?. Gastroenterology. 2017. April 1;152(5):S245. 10.1155/2007/81838 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Józefczuk J, Mądry E, Nowak J, Walkowiak M, Łochocka K, Banasiewicz T, et al. Conflicting results of non-invasive methods for detection of Helicobacter pylori infection in children with celiac disease-a preliminary study. Acta Biochim Pol. 2016;63: 127–130. 10.18388/abp.2015_1088 [DOI] [PubMed] [Google Scholar]
- 48.Dore MP, Salis R, Loria MF, Villanacci V, Bassotti G, Pes GM. Helicobacter pylori infection and occurrence of celiac disease in subjects HLA‐DQ 2/DQ 8 positive: A prospective study. Helicobacter. 2018;23: e12465. 10.1111/hel.12465 [DOI] [PubMed] [Google Scholar]
- 49.Rostami-Nejad M, Villanacci V, Mashayakhi R, Molaei M, Bassotti G, Zojaji H, et al. Celiac disease and Hp infection association in Iran. Rev Esp Enfermedades Dig. 2009;101: 850–854. 10.4321/s1130-01082009001200004 [DOI] [PubMed] [Google Scholar]
- 50.Rostami-Nejad M, Rostami K, Yamaoka Y, Mashayekhi R, Molaei M, Dabiri H, et al. Clinical and histological presentation of Helicobacter pylori and gluten related gastroenteropathy. Arch Iran Med. 2011;14: 115–119. [PMC free article] [PubMed] [Google Scholar]
- 51.Bayrak NA, Tutar E, Volkan B, Sahin Akkelle B, Polat E, Kutluk G, et al. Helicobacter pylori infection in children with celiac disease: Multi‐center, cross‐sectional study. Helicobacter. 2020;25: e12691. 10.1111/hel.12691 [DOI] [PubMed] [Google Scholar]
- 52.Bayrak NA, Volkan B. Helicobacter pylori infection in children with concurrent Celiac disease and type 1 Diabetes Mellitus. Dig Dis. 2021. 10.1159/000514276 [DOI] [PubMed] [Google Scholar]
- 53.Mahdi BM. Immune response to Helicobacter pylori. Trends Helicobacter pylori Infect. 2014; 79. [Google Scholar]
- 54.Yuan J, Li P, Tao J, Shi X, Hu B, Chen H, et al. H. pylori escape host immunoreaction through inhibiting ILK expression by VacA. Cell Mol Immunol. 2009;6: 191–197. 10.1038/cmi.2009.26 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Molinari M, Salio M, Galli C, Norais N, Rappuoli R, Lanzavecchia A, et al. Selective inhibition of Ii-dependent antigen presentation by Helicobacter pylori toxin VacA. J Exp Med. 1998;187: 135–140. 10.1084/jem.187.1.135 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Gebert B, Fischer W, Weiss E, Hoffmann R, Haas R. Helicobacter pylori vacuolating cytotoxin inhibits T lymphocyte activation. Science. 2003;301(5636): 1099–1102. 10.1126/science.1086871 [DOI] [PubMed] [Google Scholar]