ViroMatch is an automated pipeline that takes metagenomic sequencing reads as input and performs iterative nucleotide and translated nucleotide mapping to identify viral sequences. We provide a Docker image for ViroMatch, so that users will not have to install dependencies.
ABSTRACT
ViroMatch is an automated pipeline that takes metagenomic sequencing reads as input and performs iterative nucleotide and translated nucleotide mapping to identify viral sequences. We provide a Docker image for ViroMatch, so that users will not have to install dependencies.
ANNOUNCEMENT
Next-generation sequencing (NGS) is a powerful tool that allows the comprehensive characterization of viral communities and the discovery of novel viruses (1–5). While software pipelines exist for viral detection (6–8), many rely on prohibitive memory and CPU requirements; others rely on stringent k-mer hashing that lacks sensitivity. We developed ViroMatch to analyze data sets of millions of short metagenomic sequence reads to identify viruses by sensitive sequence alignment using pragmatic system resources. We have modified and refined our analysis pipeline workflow over time, primarily applied to the analysis of vertebrate viruses (9–16). The latest version is made available to the public for the first time here.
The ViroMatch workflow is shown in Fig. 1. All reads are host filtered prior to viral assessment. Metagenomic sequences are screened for putative viral reads by nucleotide mapping (BWA-MEM [17]) and translated mapping (Diamond [18]) against a database of virus-specific reference genome sequences collected from NCBI GenBank (19). The use of both nucleotide and translated amino acid sequence alignments enables the detection of sequences that are conserved or divergent compared to reference genome sequences. This first screen is fast, but the hits include false positives; therefore, the putative viral hits are subsequently mapped to the more comprehensive NCBI nucleotide (nt) and NCBI nonredundant (nr) amino acid databases (https://www.ncbi.nlm.nih.gov/). Only sequences with an unambiguous mapping to a viral reference are counted as viral hits. Ambiguous hits—e.g., those mapping with similar scores to viruses, human, bacteria—are not counted. Ambiguous hits also include those that map to repetitive regions not suitable for determining virus positivity.
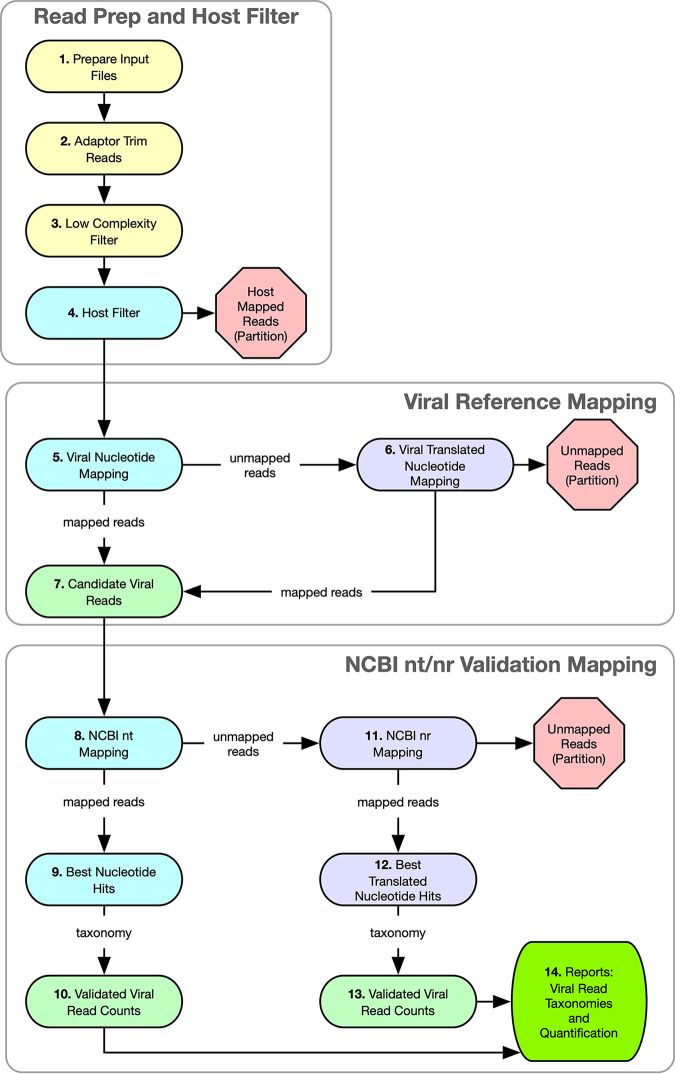
FIG 1.
ViroMatch pipeline workflow. (1 to 3) Input sequences are prepared for processing. Sequencing adaptors are excised, low-quality base pairs are trimmed from read ends, and the resultant too-short reads are removed. Low-complexity and repetitive base pairs are soft masked. (4) The reads are mapped to the host reference genome. Reads that map to the host are partitioned for later referral. Nonhost reads are promoted for mapping to the virus-only database. (5 to 7) The reads are first mapped to a virus-only reference genome database using nucleotide alignment. Reads that did not map are subsequently mapped to a translated nucleotide version of the same virus-only database. Reads that did not map at all are partitioned for later referral. The mapped reads (both nucleotide and translated nucleotide) are collected for validation mapping. (8 to 13) Candidate viral reads are validated by mapping to comprehensive NCBI nt and nr references in an iterative approach similar to steps 5 to 7. The best hit for each read is chosen using a predefined algorithm that determines viral positivity and taxonomic classification. (14) Viral read counts and taxonomic classifications are compiled into reports.
Upon completion, ViroMatch provides reports detailing viral taxonomic classification and quantification of mapped reads. Report read counts have been mapped to the virus-only reference database, have undergone validation of candidate viral reads against local NCBI nt and nr reference databases, have been taxonomically classified, and have been filtered by best-hit logic. Only reads that have passed all of these steps are considered viral identities.
The system requirements related to disk space and memory vary depending on the size of the samples being processed; however, we recommend a minimum of 16 Gb of memory. We have run ViroMatch on thousands of samples, ranging from 1 to 200 million reads processed per sample. As an example, ViroMatch processed ∼11 million reads using a single core and 16 Gb of RAM with a total runtime of 8:00:06. ViroMatch uses Snakemake (20) to transparently organize and run all of its steps.
Data availability.
ViroMatch is available through an executable Docker (21) image (https://hub.docker.com/r/twylie/viromatch), which contains all necessary code and third-party dependencies. ViroMatch’s required reference genome databases are also available for download (https://twylie.github.io/viromatch/download_and_install/databases/) using Globus (22, 23) data transfer software. ViroMatch pipeline source code is also available through GitHub (https://github.com/twylie/viromatch) and is provided under the MIT license. For download and installation instructions and complete usage documentation, please visit the ViroMatch website at https://twylie.github.io/viromatch/.
ACKNOWLEDGMENT
The following grants have provided funding in part for the development of ViroMatch: the National Institutes of Health Clinical and Translational Science (NIH CTSA UL1TR002345) and the National Institutes of Health research grant number R01HD095986. The funders had no role in the study design, data collection and interpretation, or the decision to submit the work for publication.
REFERENCES
- 1.Zárate S, Taboada B, Yocupicio-Monroy M, Arias CF. 2017. Human virome. Arch Med Res 48:701–716. doi: 10.1016/j.arcmed.2018.01.005. [DOI] [PubMed] [Google Scholar]
- 2.Garmaeva S, Sinha T, Kurilshikov A, Fu J, Wijmenga C, Zhernakova A. 2019. Studying the gut virome in the metagenomic era: challenges and perspectives. BMC Biol 17:84. doi: 10.1186/s12915-019-0704-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ogilvie LA, Jones BV. 2015. The human gut virome: a multifaceted majority. Front Microbiol 6:918. doi: 10.3389/fmicb.2015.00918. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Lecuit M, Eloit M. 2013. The human virome: new tools and concepts. Trends Microbiol 21:510–515. doi: 10.1016/j.tim.2013.07.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Wylie KM, Weinstock GM, Storch GA. 2013. Virome genomics: a tool for defining the human virome. Curr Opin Microbiol 16:479–484. doi: 10.1016/j.mib.2013.04.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Wood DE, Lu J, Langmead B. 2019. Improved metagenomic analysis with Kraken 2. Genome Biol 20:257. doi: 10.1186/s13059-019-1891-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Naccache SN, Federman S, Veeraraghavan N, Zaharia M, Lee D, Samayoa E, Bouquet J, Greninger AL, Luk K-C, Enge B, Wadford DA, Messenger SL, Genrich GL, Pellegrino K, Grard G, Leroy E, Schneider BS, Fair JN, Martínez MA, Isa P, Crump JA, DeRisi JL, Sittler T, Hackett J, Jr, Miller S, Chiu CY. 2014. A cloud-compatible bioinformatics pipeline for ultrarapid pathogen identification from next-generation sequencing of clinical samples. Genome Res 24:1180–1192. doi: 10.1101/gr.171934.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Zhao G, Wu G, Lim ES, Droit L, Krishnamurthy S, Barouch DH, Virgin HW, Wang D. 2017. VirusSeeker, a computational pipeline for virus discovery and virome composition analysis. Virology 503:21–30. doi: 10.1016/j.virol.2017.01.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Wylie KM, Wylie TN, Cahill AG, Macones GA, Tuuli MG, Stout MJ. 2018. The vaginal eukaryotic DNA virome and preterm birth. Am J Obstet Gynecol 219:189.e1–189.e12. doi: 10.1016/j.ajog.2018.04.048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Wylie KM, Wylie TN, Buller R, Herter B, Cannella MT, Storch GA. 2018. Detection of viruses in clinical samples by use of metagenomic sequencing and targeted sequence capture. J Clin Microbiol 56:e01123-18. doi: 10.1128/JCM.01123-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Chu S, Wylie TN, Wylie KM, Johnson GC, Skidmore ZL, Fleer M, Griffith OL, Bryan JN. 2020. A virome sequencing approach to feline oral squamous cell carcinoma to evaluate viral causative factors. Vet Microbiol 240:108491. doi: 10.1016/j.vetmic.2019.108491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Wylie TN, Wylie KM, Herter BN, Storch GA. 2015. Enhanced virome sequencing using targeted sequence capture. Genome Res 25:1910–1920. doi: 10.1101/gr.191049.115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Eskew AM, Stout MJ, Bedrick BS, Riley JK, Omurtag KR, Jimenez PT, Odem RR, Ratts VS, Keller SL, Jungheim ES, Wylie KM. 2020. Association of the eukaryotic vaginal virome with prophylactic antibiotic exposure and reproductive outcomes in a subfertile population undergoing in vitro fertilisation: a prospective exploratory study. BJOG 127:208–216. doi: 10.1111/1471-0528.15951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Wylie KM, Mihindukulasuriya KA, Zhou Y, Sodergren E, Storch GA, Weinstock GM. 2014. Metagenomic analysis of double-stranded DNA viruses in healthy adults. BMC Biol 12:71. doi: 10.1186/s12915-014-0071-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Wylie KM, Mihindukulasuriya KA, Sodergren E, Weinstock GM, Storch GA. 2012. Sequence analysis of the human virome in febrile and afebrile children. PLoS One 7:e27735. doi: 10.1371/journal.pone.0027735. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Dharnidharka VR, Ruzinova MB, Chen C-C, Parameswaran P, O'Gorman H, Goss CW, Gu H, Storch GA, Wylie K. 2019. Metagenomic analysis of DNA viruses from posttransplant lymphoproliferative disorders. Cancer Med 8:1013–1023. doi: 10.1002/cam4.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Li H. 2013. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv 13033997 [q-bio.GN]. https://arxiv.org/abs/1303.3997.
- 18.Buchfink B, Xie C, Huson DH. 2015. Fast and sensitive protein alignment using DIAMOND. Nat Methods 12:59–60. doi: 10.1038/nmeth.3176. [DOI] [PubMed] [Google Scholar]
- 19.Benson DA, Cavanaugh M, Clark K, Karsch-Mizrachi I, Lipman DJ, Ostell J, Sayers EW. 2013. GenBank. Nucleic Acids Res 41:D36–D42. doi: 10.1093/nar/gks1195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Köster J, Rahmann S. 2012. Snakemake—a scalable bioinformatics workflow engine. Bioinformatics 28:2520–2522. doi: 10.1093/bioinformatics/bts480. [DOI] [PubMed] [Google Scholar]
- 21.Merkel D. 2014. Docker: lightweight Linux containers for consistent development and deployment. Linux J 2014:2. [Google Scholar]
- 22.Foster I. 2011. Globus Online: accelerating and democratizing science through cloud-based services. IEEE Internet Comput 15:70–73. doi: 10.1109/MIC.2011.64. [DOI] [Google Scholar]
- 23.Allen B, Bresnahan J, Childers L, Foster I, Kandaswamy G, Kettimuthu R, Kordas J, Link M, Martin S, Pickett K, Tuecke S. 2012. Software as a service for data scientists. Commun ACM 55:81–88. doi: 10.1145/2076450.2076468. [DOI] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
ViroMatch is available through an executable Docker (21) image (https://hub.docker.com/r/twylie/viromatch), which contains all necessary code and third-party dependencies. ViroMatch’s required reference genome databases are also available for download (https://twylie.github.io/viromatch/download_and_install/databases/) using Globus (22, 23) data transfer software. ViroMatch pipeline source code is also available through GitHub (https://github.com/twylie/viromatch) and is provided under the MIT license. For download and installation instructions and complete usage documentation, please visit the ViroMatch website at https://twylie.github.io/viromatch/.