SUMMARY
T and B cells are the two known lineages of adaptive immune cells. Here, we describe a previously unknown lymphocyte that is a dual expresser (DE) of TCR and BCR and key lineage markers of both B and T cells. In type 1 diabetes (T1D), DEs are predominated by one clonotype that encodes a potent CD4 T cell autoantigen in its antigen binding site. Molecular dynamics simulations revealed that this peptide has an optimal binding register for diabetogenic HLA-DQ8. In concordance, a synthetic version of the peptide forms stable DQ8 complexes and potently stimulates autoreactive CD4 T cells from T1D patients, but not healthy controls. Moreover, mAbs bearing this clonotype are autoreactive against CD4 T cells and inhibit insulin tetramer binding to CD4 T cells. Thus, compartmentalization of adaptive immune cells into T and B cells is not absolute, and violators of this paradigm are likely key drivers of autoimmune diseases.
In Brief
Type I diabetes patients have unique TCR- and BCR-positive lymphocytes, in which a public BCR encodes a potent autoantigen that stimulates autologous CD4 T cells and may contribute to autoimmunity.
Graphical Abstract

INTRODUCTION
B and T cells are the two main lymphocytes of the adaptive immune system that work in concert to maintain host defense or cause autoimmunity in susceptible individuals. Expression of the B cell receptor (BCR) defines B cells, and the T cell receptor (TCR) defines T cells. Both antigen receptors have similar structures and highly diverse repertoires (Wardemann et al., 2003). The BCR (surface immunoglobulin [Ig]) is a heterodimer composed of heavy (IGH) and light (IGL) chains, whereas the αβ TCR heterodimer is composed of TCRα and TCRβ chains. Each receptor has a hypervariable region containing V (variable), D (diversity in case of IGH and TCRβ), and J (joining) gene segments randomly selected from large pools of unarranged segments and recombined to generate a complementarity-determining region (CDR3) that denotes the specificity of each clonotype and comprises its antigen binding site. The diversity is enhanced by N1 and N2 nucleotide additions/deletions at the V-D and D-J junctions, respectively. Theoretically, up to 1018 unique BCRs or TCRs can be generated during the development of B cells in bone marrow and T cells in thymus (Sewell, 2012). The diversity is essential for protecting a host against virtually any pathogen, but it also leads to generation of autoreactive B and T cells that cause autoimmune diseases in genetically susceptible individuals. Currently, there is no cure for autoimmune diseases. A major reason is the limited knowledge about identities of autoreactive lymphocytes and autoantigens that cause their activation. In this study, we describe an unexpected population of lymphocytes that are dual expressers (DEs) of TCR and BCR and lineage markers of both B and T cells. These violators of the paradigm that the adaptive immune cells are either T or B cells appear to be involved in mediating autoimmunity.
One of the extensively examined and yet poorly understood aspects of autoimmunity are autoantigens that drive activation of autoreactive T cells. This question is heavily investigated in the context of type 1 diabetes (T1D), which is considered a prototypical autoimmune disease, and insulin is viewed as the primary autoantigen that drives activation of diabetogenic T cells. Yet presentation of naturally processed epitopes of the insulin B:9–23 peptides (SHLVEALYLVCGERG) by major histocompatibility class II (MHC II) molecules to CD4 T cells remains paradoxical. This is because development of T1D is tightly linked to polymorphism at β57 position of the β chain of the HLA (human leukocyte antigen)-DQ (DQ8 and DQ2) MHC II molecules that replaces aspartic acid (D) by alanine (β57D−) in 90% of T1D patients. These DQ alleles favor antigenic peptides with negatively charged resides at positions 9 (P9) and 1 (P1) (Chang and Unanue, 2009). Nonetheless, naturally processed insulin B:9–23 peptide binds poorly to DQ8 at two registers that place nonacidic residues (V or A) at P1 and (G or R) at P9. Posttranslational modification has been suggested as a mechanism that enhances insulin bindings to DQ8 molecules and its antigenicity. In support of this hypothesis, substitution of arginine (R) at P9 by glutamic acid (R22E) generates a potent mimotope that is 100-fold more potent than native insulin B:9–23 (Wang et al., 2018). Moreover, combining R22E with A14E substitution at the P1 position generates a superagonist that is about 1,000-fold more potent than native insulin (Wang et al., 2018). Insulin peptides with such modifications, however, are yet to be identified in vivo.
In this study, we identified a potent neoantigen with optimal anchor residues for DQ8. The neoantigen is encoded in the CDR3 idiotype (referred to as x-Id) of the heavy-chain clonotype of DEs in T1D subjects. Synthesized x-Id peptide forms stable DQ8 complexes and potently stimulates autoreactive CD4 T cells from T1D, but not healthy controls. Moreover, x-clonotype-bearing monoclonal antibodies (mAbs) stimulated CD4 T cells, and competitive tetramer binding assay suggests the x-mAb cross-activates insulin-specific CD4 T cells. Collectively, our findings show that compartmentalization of T and B cells is not absolute, and violators of this paradigm could be drivers of autoimmunity.
RESULTS
A Rare Subset of Lymphocytes Coexpresses T and B Cell Lineage Markers and Expands in T1D
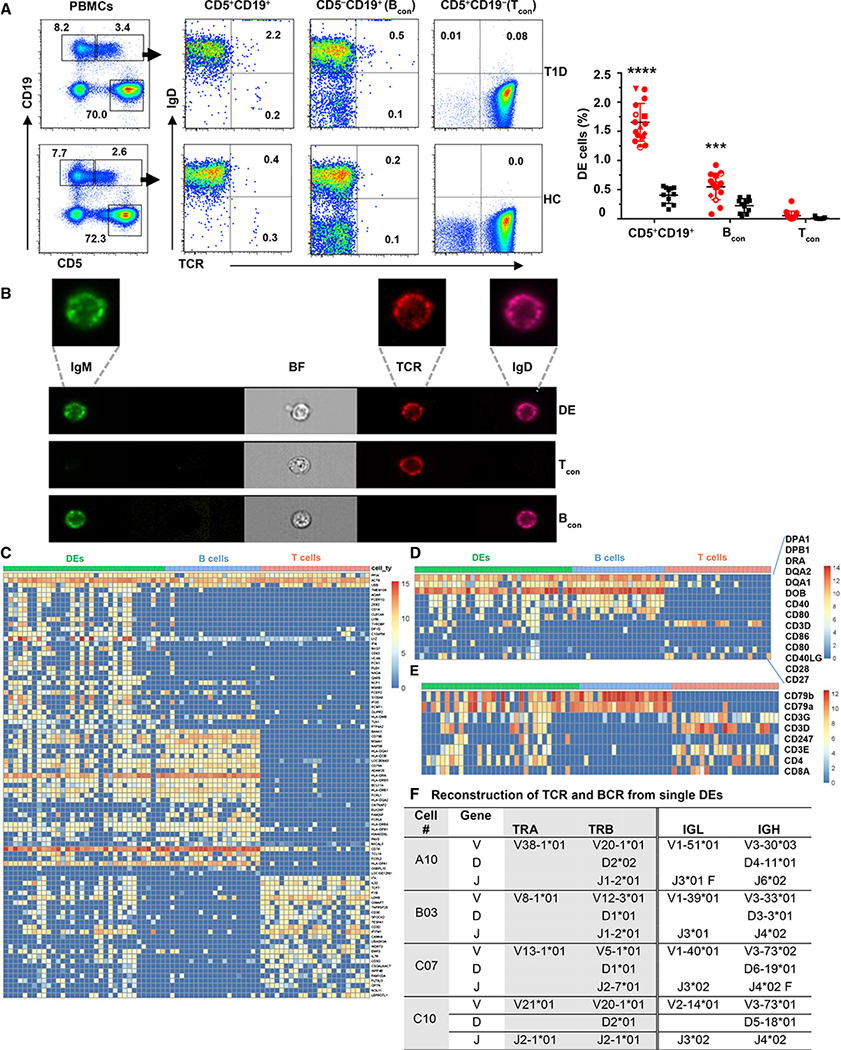
We identified a rare subpopulation of lymphocytes that coexpresses the BCR and TCR and resides mainly in the CD5+ CD19+ compartment in peripheral blood of individuals examined (Figures 1A and S1 for gating strategy). The majority of these DEs expressed IgD and IgM and were identified phenotypically as CD5+ CD19+ TCRβ+ IgD+ cells (Figure 1A). A minor subset of CD5+ CD19+ TCRβ+ were IgD− that expressed IgG, IgA, or IgM, suggesting they were class-switched DEs (see Figure 2D below). Few DEs were present among CD5− CD19+ Bcon cells and were not analyzed further. Instead, we focused on IgD+ and IgD− DEs found in the CD5+ CD19+ compartment. DEs were significantly more frequent in T1D subjects than in healthy controls (HCs) (Figure 1A; see subject characteristics in Table S1). We visualized coexpression of the IgD, IgM, and TCR in DEs at the single-cell level using flow cytometric imaging, AMNIS (Figure 1B). Thus, although rare, DEs could be pathophysiologically important due to their expansion in T1D.
Figure 1. A Rare Subset of Lymphocytes Coexpresses TCR and BCR and Expands in T1D.
(A) Representative dot plots show coexpression of IgD and TCR among gated CD5+ CD19+ cells in T1D (top panel) and HC (bottom panel) subjects. Numbers indicate percentages in quadrants. Graph shows cumulative data (mean ± SEM). Each dot represents one donor. T1D (red, n = 16), HC (black, n = 11); ***p < 0.001, ****p < 0.0001 by two-way ANOVA with Sidak’s multiple comparisons test. See also Figure S1.
(B) Representative AMNIS images show coexpression of IgD, TCR, and IgM by gated single-IgD+ DEs versus their differential expression in Bcon and Tcon cells in three T1D subjects (n = 32 DE cells). BF, bright field. See also Figure S2.
(C) Heatmap of genes differentially expressed by DEs, Bcon, or Tcon cells. Top row shows cell types. Subsequent three rows show expression of ACTB, PPIA, and UBB housekeeping genes followed by the top 30 genes preferentially expressed in each cell type. The color scale indicates the gene expression in log2 (RSEM + 1). Note that DEs differentially express large numbers of genes that are absent or low in Bcon and Tcon cells. DEs share expression of markers of B and T cells with respective cell type. Data from 34 DEs (green), 20 Bcon cells (blue), and 23 Tcon cells (orange).
(D) Heatmap shows DEs’ shared expression of indicated lineage markers with respective cell type Tcon or Bcon cells.
(E) Heatmap shows DEs shared expression of Igα (CD79a) and Igβ (CD79b) with Bcon cells and CD3 subunits with Tcon cells. CD247 is CD3zeta.
(F) Reconstruction of BCR and TCR in four DEs. No dual expression of BCR and TCR noted among Tcon and Bcon cells. See Table S2.
See also Figures S1 and S2 and Tables S1 and S2.
Figure 2. TCR-Activated DEs Maintain Their Dual Phenotype and Upregulate MHC and Costimulatory Molecules.
(A) TCR activation leads to the upregulation of CD69 by IgD+ and IgD− cells. Left dot plots show gating of CD5+ CD19+ cells and Bcon and Tcon cells in anti-CD3/CD28 (top panel) and unstimulated control (bottom panel) cultures. Middle dot plots show expression of TCR and IgD by gated subsets. Overlays and graph show CD69 expression by gated IgD+ (red) and IgD− (navy blue) and Tcon cells (green) and Bcon cells (blue) in activated and control cultures. Each dot represents one donor (n = 5); ****p < 0.0001 by two-way ANOVA with Tukey’s multiple comparisons test.
(B) TCR activation leads to the proliferation of IgD+ and IgD− DE subsets and Tcon cells as determined by CFSE dilution. Open histograms denote unstimulated cultures.
(C) Upregulation of HLA-DR and DQ by TCR-activated DEs. Note that Bcon cells were present in control, but not activated cultures. Graphs show MFI (mean ± SEM) for HLA-DR (left) and HLA-DQ (right). Each dot represents one donor (n = 4); **p < 0.01 by two-way ANOVA with Sidak’s multiple comparisons test.
See Figure S3 for upregulation of costimulatory molecules.
D) DEs maintain Ig isotype phenotypes after 7 days of anti-CD3/CD28 stimulation.
See also Figures S2, S3, and S4.
To investigate the dual phenotype of DEs at single-cell resolution, we examined their transcriptomes using single-cell RNA sequencing (scRNA-seq). We sorted individual DEs and Bcon and Tcon cells from peripheral blood mononuclear cells (PBMCs) of T1D#1 and analyzed their transcriptomes using the plate-based SMART-seq2 protocol (Tirosh and Suvà, 2018; Tirosh et al., 2016). A total of 77 cells (34 DEs, 20 Bcon, and 23 Tcon) passed quality control based on the expression of at least two out of three housekeeping genes. Results show that DEs have uniquely expressed genes along with genes encoding lineage markers of T and B cells. Figure 1C shows the top 30 genes differentially expressed by DEs compared to Tcon or Bcon cells. In addition, it shows the top 30 differentially expressed genes by Bcon and Tcon cells and their expression by DEs. The results confirm the crossover phenotype at single-cell resolution.
We highlighted shared expression of selected lineage markers of B and T cells by DEs (Figures 1D and 1E). These were also visualized at the protein level using AMNIS (Figures 1B and S2) and fluorescence-activated cell sorting (FACS) (Figures 2C, S2, and S3). We also analyzed DEs for the invariant components of BCR (CD79a and CD79b) and TCR (CD3γ, CD3ε, CD3δ, and CD247) that are responsible for transducing activation signals. In line with our FACS, AMNIS, and functional data, DEs shared expression of CD79a and CD79b with Bcon cells and CD3γ, CDε, CD3δ, and CD247 (CD3ζ) with Tcon cells (Figure 1E).
Finally, we used BCR assembly from single cells (BASIC) bioinformatics software to reconstruct recombined BCR and TCR genes expressed in DEs using Bcon and Tcon cells as controls. We detected contigs of both BCR and TCR in DEs and, as expected, contigs of BCR genes in Bcon and contigs of TCR genes in Tcon cells (Table S2). We used the IMGT/V-QUEST software and examined reconstructed sequences for V(D)J usage. Many single DEs successfully reconstructed one (22 cells) or both (18 cells) BCR chains (Table S2A). In addition, several (8 cells) had fully assembled TCRβ variable (Table S2B). Importantly, four individual DEs coexpressed fully assembled BCRs (heavy and light chains) together with fully assembled TCRβ chain with TCRVα (Figure 1F). As specificity controls, there were no assembled TCR chains in Bcon cells or BCR chains in Tcon cells. Collectively, these results provide proof of principle of the existence of a hybrid lymphocyte that combines features of T and B cells.
TCR-Activated DEs Maintain Their Dual Phenotype and Upregulate MHC and B Cell Costimulatory Molecules
Next, we examined functionality of TCRs expressed on DEs and phenotypic and functional consequences of their crosslinking. Consistent with scRNA-seq results, DEs expressed the CD3ε signaling subunit (Figure S2E), indicating a functional TCR/CD3 complex. To test this notion, we activated PBMCs with anti-CD3/CD28 for 7 days and analyzed different subsets for CD69 upregulation. Because of the rarity of DEs in HCs, we did our experiments using PBMCs from T1D subjects unless stated otherwise. Analysis of activated cultures show that CD19+ CD5− Bcon cells regressed (consistent with being bystanders) and could not be properly analyzed for CD69 expression. Consequently, most remaining CD19+ cells were present within the CD5intermediate gate (87.3% ± 11.7%; n = 5), expressed TCR, and included IgD+ and IgD− DEs (Figure 2A). TCR stimulation led to significant upregulation of CD69 by IgD+ and IgD− DEs as compared to DEs in control cultures (Figure 2A). Tcon cells that made the bulk of cultures also upregulated CD69, but significantly less than DEs. Thus, TCRs on DEs are not only functional molecules, but highly responsive to TCR crosslinking. However, since our analysis of DEs was performed in bulk cultures of PBMCs (due to difficulty of growing purified DEs), we could not determine whether all responses of DEs were due to direct anti-CD3 and anti-CD28 stimulation or partially due to indirect effects caused by activated conventional T cells.
In a second set of experiments, we stimulated PBMCs with anti-CD3/CD28 and visualized proliferation using the carboxy-fluorescein succinimidyl ester (CFSE) dilution assay (Figure 2B). Consistent with the above results, Bcon cells regressed in activated cultures, and most remaining CD19+ cells expressed an intermediate level of CD5. Both IgD+ and IgD− DEs, similar to Tcon cells, divided robustly in response to anti-CD3/CD28 stimulation as indicated by CFSE dilution (Figure 2B). Consistent with scRNA-seq results, FACS analysis confirmed that DEs expressed HLA-DR (DR) and HLA-DQ (DQ) molecules and indicated pan-B and pan-T cell markers. Anti-CD3/CD28 stimulation led to significant upregulation of DR and DQ molecules (Figure 2C) and pan-B cell costimulatory markers, whereas pan-T cell markers did not significantly change (Figure S3A). Most IgD+ DEs cells maintained coexpression of IgM, and none switched to IgA or IgG, whereas IgD− DEs remained mixtures of IgG+, IgA+, and IgM+ cells, and none expressed IgE (Figure 2D). Differential expression of Ig isotypes by DEs shows that DEs do not suffer from generalized dysregulated gene expression. In addition, activated DEs expressed CD45RA, and none switched to CD45RO (Figure S3B). Likewise, consistent with transcriptome analysis, DEs differentially expressed CD4 and CD8 coreceptors, while some were CD4 and CD8 double-negative (Figures S2C and S2D). It is also noteworthy that DEs, particularly IgD− cells, produced IL-10 and IFN-γ when stimulated via PMA (phorbol 12-myristate 13-acetate) and ionomcyin or with anti-CD3 and anti-CD28 (see Figure S4). These results indicate that TCRs and BCRs remain stably coexpressed on DEs after TCR-mediated expansion.
We also examined functionality of BCRs expressed on DEs. FACS analysis confirmed expression of CD79a (Igα) and CD79b (Igβ) by DEs, indicating a functional receptor (Luisiri et al., 1996; Reth, 1992; Yu and Chang, 1992). To directly test this possibility, we determined whether crosslinking of IgM induces phosphorylation of CD79a in DEs. This experiment was possible because almost all IgD+ DEs and a fraction of IgD− DEs were IgM+ (Figure 2D). We stimulated PBMCs with F(ab)2 goat anti-human IgM and measured CD79a phosphorylation at indicated time points. Anti-IgM stimulation led to extended phosphorylation of CD79a in IgD+ DE cells (Figure S2F) and to a lesser extent in IgD− DEs (expected given that many IgD− DEs expressed IgG or IgA; see Figure 2D). On the other hand, Bcon cells transiently but significantly phosphorylated CD79a. As a specificity control, Tcon cells did not show any signal in response to anti-IgM stimulation. These results show that BCR complexes expressed by DEs are functional molecules.
Analysis of TCR and BCR Repertoires of DE Cells
Next, we sorted DEs and analyzed their clonotypic TCRβ (TRBV) and BCR (IGHV) repertoires and compared to repertoires of conventional T and B cells using genomic DNA and high-throughput ImmunoSEQ (Adaptive Biotech). This analysis was important to determine clonality of TCRs and BCRs expressed in DEs and to rule out any role for unforeseen artifacts at the protein level, such as antigen receptor trogocytosis (LeMaoult et al., 2007) in imparting DEs with their hybrid phenotype.
Restricted TCRVβ Usage by DE Cells
We first analyzed TRBV repertoires of eight DE samples (four IgD+ and four IgD− subsets) from three T1D subjects (T1D#1, #4, and #5). The subjects were unrelated and sequenced at different points in time. The sorting strategy and purity of sorted subsets are shown (Figures S5A and S5B). DEs exhibited restricted TCRVβ usage (Figures S5C and S5E and Tables S3A–S3C). In general, the IgD+ clonotypes used between 18 and 31 Vβs, and the IgD− clonotypes used 5 and 29 Vβs, whereas Tcon cells used almost all 55 Vβ genes. The paucity of DE cells in HCs did not allow sorting and deep sequencing except from one donor (HC#1) who expressed the DQ2 risk allele but was negative for islet autoantibodies, IAAs (Table S1C). We analyzed both IgD+ and IgD− DE cells from HC#1. The IgD+ subset in HC#1 used only 7 Vβs, and the IgD− subset used 5 Vβs, as compared to usage of 55 Vβ genes used by Tcon cells (Figure S5F, Venn diagram, and Table S3D). The skewed TCR repertoires of DEs provide another line of evidence of their distinctiveness. The results also argue against the possibility that TCRs detected on the surface of DEs were derived from Tcon cells that had been non-specifically conjugated with Bcon cells, because in such a case, the Vβ usage should reflect the diverse repertoire of Tcon cells. In addition, nonspecific conjugates between B and T cells are usually short lived and last only a few minutes (Okada et al., 2005). Thus, DEs, unlike Tcon cells in the same subjects, have a skewed TCR repertoire with limited Vβ usage.
Analyses of IGHV Repertoires of DEs Identify a Public Dominant Clonotype in T1D Subjects
Next, we analyzed the IGHV repertoires of IgD+ and IgD− DEs and Bcon cells from three T1D (#1, #2, #3) subjects using the sorting strategy described above (Figures S5A and S5B). We obtained results from four out of six DE samples: IgD+ cells from T1D#1, the IgD− cells from T1D#2, and both the IgD+ and IgD− cells from T1D#3. We obtained IGHV sequences of Bcon cells from each subject. There was predominant usage of the IGHV04-b gene recently named IGHV4–38-2 (Watson et al., 2013) by DEs in the three T1D subjects. It was used by 95% of IgD+ cells in T1D#1, 22% (top clonotype) of IgD− cells in T1D#2, and 88% of IgD+ and IgD− cells in T1D#3 (Figures 3A–3C). In contrast, the VH04-b gene was used by less than 1% of Bcon cells and ranked by usage number 27/76, 31/77, and 28/82 in the three subjects, respectively (Table S4). Moreover, there was no significant overlap in VH usage by DEs and Bcon cells. In fact, the top 10 VH genes used by Bcon cells (particularly T1D#1 and T1D#3) either were entirely absent or constituted minor components of DE repertoires (Figures S5G–S5J). A complete list of the VH genes used by DE cells and Bcon cells in the three subjects is shown (Table S4). Furthermore, the VH genes used by DEs, unlike their counterparts on Bcon cells, were mainly of germline configuration (Figure 3D). The distinct BCR properties of DEs rules out cross-contamination of DEs by Bcon cells. In addition, the results indicate commonality between DEs, at least in a subset of T1D patients represented by those analyzed in this study.
Figure 3. IGHV Repertoires of DEs Are Predominated by One Clonotype in T1D Subjects.
(A–C) Venn diagrams show VH gene usage by IgD+ (red) and IgD− (yellow) DEs and Bcon cells (blue) in (A) T1D#1, (B) #2, and (C) #3 patients. Graphs show percentages of the top 10 VH genes (or all 7 VH genes, in the case of T1D#2) used by IgD+ or IgD− DEs as compared to Bcon cells in each patient. Arrows point to the IGHV-04-b+ gene segment, which was predominantly used by IgD+ and IgD− DEs in the three patients.
(D) Graph shows absolute number of mutations per VH gene in DEs and Bcon cells in the three T1D subjects. Each dot represents an individual VH gene.
(E) Schematic shows the VH(N1)D(N2)JH structure with the nucleotide and amino acid sequences of the CDR3 of the x-clonotype.
(F) Venn diagram shows that the x-clonotype is one of two (red) clonotypes shared among Bcon cells of the three T1D subjects.
(G) Venn diagram shows diverse VH gene usage by IgD+ (red) and IgD− (yellow) DEs comparable to that of Bcon (blue) in HC#1. Graph shows percentages of the top 10 VH genes used by IgD+ DEs as compared to IgD− DEs and Bcon cells.
(H) Comparison of CDR3 sequences of IGHV04-b+ clonotypes in the three T1D subjects and HC#1. Asterisk indicates gap in sequence. Note the highly conserved usage of VH04-b and JH04–01*02 by DEs in all subjects.
(I) Number of mutations per VH gene in DE cells and Bcon cells. Each dot represents one VH gene.
(J) Schematic shows primers used for detection of x-clonotype in peripheral blood of genotyped T1D and HCs.
Table shows detection of x-clonotype in PBMC cDNAs of T1D and HC subjects using sequence-specific primers. Note that x-clonotype is detectable in DQ7+ (β57D+ isoform of DQ8), but not DQ8+ and DQ2+ HCs. A second probe with astringent reverse primer design produced similar results (Table S7).
Intrigued by the predominant usage of the IGHV04-b by DEs, we analyzed them for clonality. Remarkably, IGHV04-b+ DEs were comprised of a single clonotype that used the same VH, DH, and JH segments and N1 and N2 junctions in the three subjects, resulting in a CDR3 with identical nucleotide and amino acid sequences (Figure 3E). The CDR3 (CARQEDTAMVYYFDYW) is encoded by rearranged IGHV04-b, IGHD05–18, and IGHJ04–01*02 (Figure 3E and Table S5)—hereafter referred to as the x-clonotype. The dominance of the x-clonotype in the unrelated patients is unlikely to be coincidental given the extreme diversity of BCR repertoire (Trück et al., 2015; Venturi et al., 2008)—case in point, the clonal diversity of Bcon cells in the three subjects (see Tables S5E–S5G). To confirm the identity of the x-clonotype, we sorted single DEs from T1D#1 and analyzed for IGHV expression using multiplex PCR (Smith et al., 2009). We detected the exact x-clonotype nucleotide sequence in 7 out of 7 DE cells (Figures 6 and 7 and data not shown), confirming the high-throughput sequencing results that identified the x-clonotype in 95% of DEs in T1D#1.
Figure 6. Verification of Dual Expression of BCR and TCR by DEs Using an EBV-Immortalized Clone.
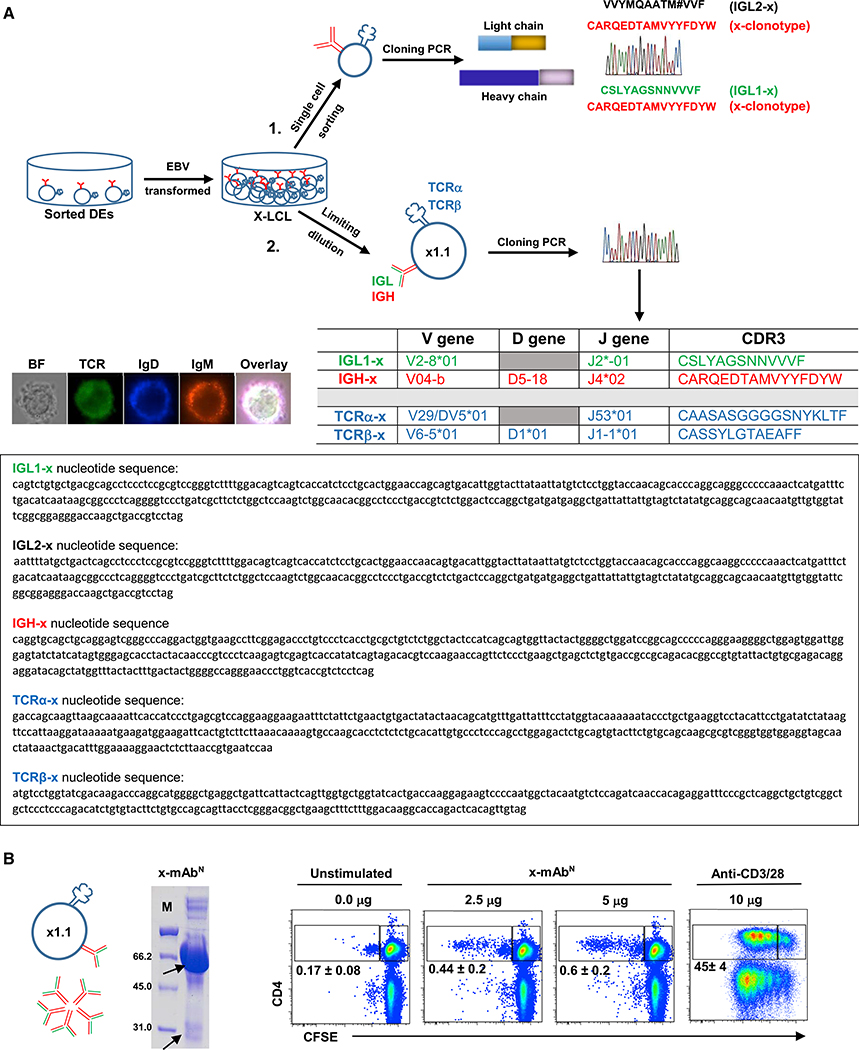
(A) Schematic depicts generation of lymphoblastoid cell line (x-LCL) and analysis of its cells for encoded antibody using two approaches: (1) cloning from sorted single cells (n = 7 cells) that yielded two antibodies that shared expression of the x-clonotype paired with one of two light chains (IGL1-x, IGL2-x) and (2) usage of limiting dilution to generate the x1.1 clone and use of PCR cloning to amplify transcripts of BCR and TCRαβ followed by usage of IMGT/V-QUEST to identify VDJ usage and CDR3 sequences. Nucleotide sequences of cloned receptors are shown. Representative images show coexpression of TCR (green), IgD (blue), and IgM (red) by a single EBV-transformed IgD+ x1.1 cell. BF, bright field. Images were taken by EVOS M7000 microscope using objective lens at 40×.
(B) Naturally produced x-mAbN stimulates CD4 T cells from T1D. Coomassie-blue-stained gel shows production of x-mAbN by the x1.1 clone. Arrows point to heavy and light chains of x-mAbN (of IgM isotype). Representative plots show dilution of CFSE by CD4 T cells activated by soluble x-mAbN. Numbers indicate percentages (mean ± SEM) of CFSElow CD4 T cells (n = 3).
Figure 7. Recombinant x-mAbR Cross-activates Insulin-Reactive CD4 T Cells.
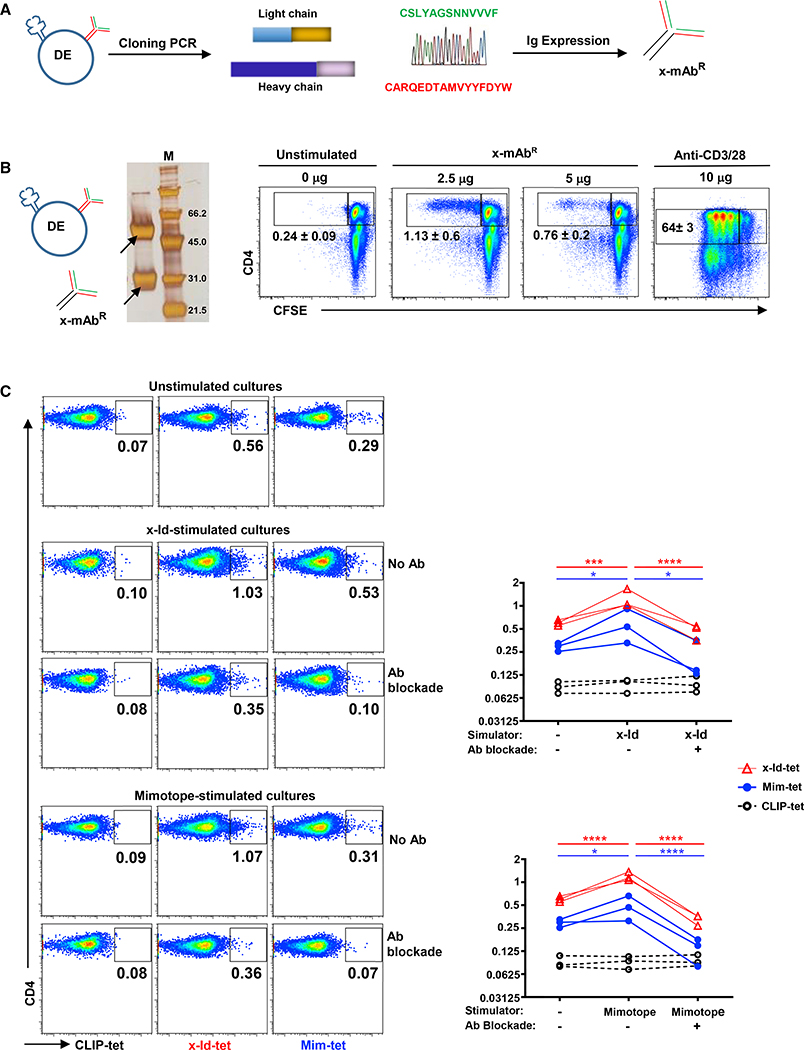
(A) Schematic depicts amplification, cloning, and CDR3 sequences of the light and heavy chain of x-mAbR from a single DE cell and expression using IgG-AbVec and Igλ-AbVec expression vectors.
(B) Silver-stained reduced gel shows heavy and light chains (arrows) of the x-mAbR. Representative plots show dilution of CFSE by activated CD4 T cells stimulated with immobilized x-mAbR. Numbers indicate percentages (mean ± SEM) of CFSElow CD4 T cells (n = 5).
(C) Binding inhibition indicates overlapping of x-Id and mimotope-reactive CD4 T cells. x-mAbR inhibits binding of mim-tet and x-Id-tet to CD4 T cells that had been activated with x-Id or mimotope. PBMCs were cultures for 7 days in the presence of absence of x-Id or mimotope peptide. Top dot plots show that CD4 T cells expanded by the x-Id-peptide are detectable not only by x-Id-tet, but also by mim-tet. Reciprocally, CD4 T cells expanded by the mimotope peptide are detectable by both the x-Id-tet and mim-tet. CLIP-Tet was used to measure background staining, and x-Id-tet+ or mim-tet+ in unstimulated cultures identify precursor frequencies. Bottom dot plots show that preincubating with cells with x-mAbR inhibits tetramer staining. Left graph shows frequency of tetramer+ CD4 T cells in different cultures of x-Id peptide-stimulated cultures. Right graph shows data from mimoptope-stimulated cultures. Each line represents one donor. Blockade with x-mAbR inhibited tetramer binding (n = 3); *p < 0.01, ***p < 0.001, ****p < 0.0001 by two-way ANOVA with Tukey’s multiple comparisons test.
We also detected the x-clonotype among Bcon cells in the three T1D subjects, but it was one of several small clonotypes that used the IGHV04-b gene (8 in T1D#1, 39 in T1D#2, and 17 in T1D#3) (Tables S5E–S5G). Furthermore, the identical amino acid sequence of the x-clonotype was generated by several Bcon clonotypes that used multiple VH genes in T1D#1 (VH04-b, VH03–11, VH01–69, VH01–46, Vh05–51, and VH01–18), T1D#2 (VH04-b and VH-04–39), and T1D#3 (VH04-b, VH03–53, VH01–02, and VH1–69) patients (see Tables S5E–S5G). Generation of an identical CDR3 amino acid sequence by different VDJ rearrangements (convergent recombination) is a characteristic of public TCRs shared between at least two individuals (Venturi et al., 2008). In this regard, the x-clonotype was only one of two IGHV clonotypes (Figure 3F) (the other less dominant one has a CAGGHNYGIKSYW CDR3 sequence) shared by Bcon cells in the three T1D subjects. Thus, the x-clonotype predominated repertoires of DEs cells and was one of only two clonotypes shared between Bcon cells of three T1D patients.
The x-Clonotype Is Absent from Repertoires of DEs of a Healthy Subject and Public Database
To shed further light into DEs and prevalence of the x-clonotype, we were able to obtain and analyze repertoires of IgD+ and IgD− DEs and compare them to those of Bcon cells in HC#1. We found that the repertoires of DEs in HC#1 were as diverse as those of Bcon cells (Figure 3G and Table S6). Usage of IGHV04-b gene was rare (≤0.015%) in IgD+ and IgD− cells and Bcon cells of HC#1 (Table S6A). More importantly, the x-clonotype was absent from repertoires of IgD+, IgD−, and Bcon cells of HC#1 (Tables S6B–S6D). Nonetheless, IGHV04-b+ IgD+ cells in HC#1 as in T1D subjects were comprised of one clonotype that used the IGHJ04–01*2 gene, but not the DH05–18 gene, and their CDR3 sequence (CARQRFWSGPLFDYW) partly matched (bold-faced) that of x-clonotype. IGHV04-b+ IgD− DE cells, however, were comprised of five clonotypes that used the IGHJ04–01*2, but not DH05–18 (Figure 3H). Furthermore, DE clonotypes in HC#1, as in T1D patients, were of germline configuration, albeit with few somatic mutations (Figure 3I). Thus, repertoires of DEs in HC#1, unlike in the three T1D subjects, were diverse and did not include the x-clonotype.
Survey of public database showed that the x-clonotype was absent from the high-resolution immunoSEQ database (37 million unique BCR sequences) of naive and memory B cells from healthy subjects (DeWitt et al., 2016) and that of insulin-binding Bcon cells (IBCs) from T1D and control subjects (Seay et al., 2016)—potential reasons discussed below. Survey of NCBI protein database, however, discovered a highly overlapping sequence (RQENFDTAMVYYF) derived from the variant surface glycoprotein (VSG; 1125.4290) of Trypanosoma brucei. Boldfaced letters indicate overlapping residues. VSGs are potent antigenic stimulators of B cells and T-independent IgM response (Mansfield, 1994). We conclude that the x-clonotype is rarely used as indicated by its absence from the available database of IGHV sequences of B cells, including IBCs.
x-Clonotype Is Detectable in Peripheral Blood Using Sequence-Specific PCR Probes
We sought to detect the x-clonotype in peripheral blood using two sets of sequence-specific primers. In both cases, we used a forward primer specific for the VH04-b gene. In the first set (probe 1), it was paired with a reverse primer complementary to the entire CDR3 sequence (see diagram in Figure 3J). Since our primary goal was to confirm the presence of the idiotype by an independent mechanism, we confined our analysis to a limited number of T1D and HC subjects (Tables S1B and S1C). We detected the x-clonotype in 4/8 T1D and 3/8 HCs. Detection of the x-clonotype in HCs prompted us to determine the HLA and IAA profiles of participants. As expected, all T1D subjects carried at least one risk allele (DQ2 or DQ8, hereafter referred to collectively as β57D−/+) with one participant also expressing DQ7 (the disease-neutral isoform, DQB:3*01, of DQ8 that expresses D at β57). All T1D subjects were positive for at least one IAA, and all HCs were negative for IAAs. Interestingly, the three x-clonotype+ HCs expressed DQ7, a DQ8 isoform that expresses D at β57. We did not detect the x-clonotype in the three HCs carrying at least one risk allele, i.e., β57D−/+, a result consistent with the absence of the x-clonotype from the high-throughput IGHV sequences of β57D−/+ HC#1 described above (Figure 3G). In probe 2, to impart more stringent specificity, we used a reverse primer complementary to the N2-J and downstream JH sequence (see diagram, Table S7A). Again, we identified the x-clonotype in 4/9 T1D subjects and 4/14 HCs (Table S7B and S7C). These subjects had not been genotyped or checked for IAAs. However, based on the results of the first probe, we speculate that x-clonotype+ HCs would include IAA/DQ7+ and at-risk IAA+/β57D−/+ individuals. Investigation in the future of well-characterized large cohorts would determine whether expression of the x-clonotype serves as a biomarker for T1D.
Molecular Dynamics Simulations Identify the x-Clonotype as an Optimal Peptidome for DQ8
As mentioned above, combining R22E at P9 and A14E at P1 substitutions generates an insulin superagonist with high affinity for DQ8 as shown in a recently published crystal structure (Wang et al., 2018). Alignment analysis predicted that the x-clonotype could include a DQ8 binding epitope with acidic residues (E or D) at the P1 and P9 positions, similar to that of the superagonist. To test this prediction and further characterize CDR3 peptide-HLA loading, we conducted computational biophysical modeling of epitope-HLA binding, which has been successfully used in several previous studies (Chowell et al., 2018; Hölzemer et al., 2015; Joglekar et al., 2018; Xia et al., 2014). The binding complexes of HLA with the CDR3 peptide (CARQEDTAMVYYFDYW, core epitope underlined) and superagonist (SHLVEELYLVAGEEG), as well as their associated binding energies, are shown in (Figure 4). We first ran molecular dynamics (MD) simulations of three HLA-epitope complexes (CDR3, superagonist, healthy control CARQRFWSGPLFDYW) to assess stability of the bound epitopes (Figure S6 and STAR Methods). We found that all three epitopes remained bound to the HLA without register shifts from their initial anchoring sites; however, the HC epitope destabilized the HLA-β subunit (backbone root mean squared deviation [RMSD] > 4 Å; Figure S6), indicating unfavorable binding. Therefore, subsequent analyses were limited to the CDR3 peptide and superagonist. The final bound structures are shown in Figure 4A for the CDR3 peptide (side and top views, respectively) and Figure 4B for the superagonist, with both displaying strong binding to DQ8. Free energy perturbation (FEP) calculations reveal that the CDR3 peptide binds even stronger than the superagonist (Figures 4C and 4D). In general, the FEP method can compute the binding affinity difference of alchemically mutating from one epitope to another (Chowell et al., 2018; Hölzemer et al., 2015; Joglekar et al., 2018; Xia et al., 2014) (STAR Methods). However, given that the epitope sequences diverge greatly in the current case with only one conserved residue (Val10), we computed the binding affinity change (ΔG) for the mutation of each epitope to a neutral polyglycine (peptide backbone) intermediate, which serves as a reference point. The relative binding affinity between different epitopes can be easily calculated from the relative differences of the values of ΔG for each epitope. Our results show that the CDR3 peptide has more favorable binding affinity than the superagonist by −2.3 ± 2.8 kcal/mol (Figure 4C). Decomposition of the binding affinity reveals a −4 kcal/mol van der Waals interaction preference for CDR3 binding over the superagonist (Figure 4D). This is supported by a simple interaction energy comparison as shown in Figure 4E, where the CDR3 peptide displays a stronger van der Waals interaction than the superagonist. Overall, these binding affinity results agree well with the above in vitro experimental binding assays where the CDR3 peptide was found to be a more potent autoantigen.
Figure 4. HLA-CDR3 Peptide Binding.
(A and B) HLA molecule loaded with (A) CDR3 (x-Id) peptide (CARQEDTAMVYYFDYW) and (B) superagonist (SHLVEELYLVAGEEG) from Wang et al., 2018. HLA-α is shown in cyan cartoon, HLA-β is shown in silver cartoon, and epitope residues are colored by type: white, hydrophobic; green, polar; blue, basic; red, acidic.
(C) Change in binding affinity for mutating from polyglycine to the epitope for the CDR3 peptide and superagonist.
(D) Binding affinity decomposition into vdW and electrostatics (coulomb) for the CDR3 (x-Id) peptide and superagonist.
(E) van der Waals interaction energy between the HLA and epitope from MD simulation.
(F and G) Percentage of epitope residues buried in HLA for (F) CDR3 (x-Id) peptide (CARQEDTAMVYYFDYW) and (G) superagonist (SHLVEELYLVAGEEG) from Wang et al., 2018. The sequence in bold is the core epitope sequence discussed in the text.
(H) Average fluctuation (RMSF) for each residue in Å.
(I) Detailed structure of buried salt bridges between CDR3 peptide and HLA. Basic residues are in blue, acidic are in red, and epitope backbone are in tan.
(J) (Left) Overlay of most representative epitope conformations for the CDR3 peptide (light blue) and superagonist (red), with tyrosine residues in pockets 6 and 7 for the CDR3 peptide highlighted. (Right) Side view showing similar P1, P4, and P9 agreement but large differences elsewhere.
In (C)–(D), error bars are standard error across six replicas. In (E)–(H), error bars are standard error from dividing the last 250 ns of MD simulation into five sections.
See also Figure S6.
Further in-depth structural analyses reveal several beneficial binding characteristics of this super potent CDR3 peptide (Figures 4F–4J). The importance of the anchor residues at sites 1, 4, 6, and 9 of the CDR3 core epitope is clearly visible from contact analyses (Figures 4F and 4G). Interestingly, the tyrosine residues of CDR3 site 7 and the superagonist site 3 hold the largest normalized contact area, making extensive contacts with aromatic and hydrophobic HLA residues (see Figures S6G and S6H). Residue fluctuations, as presented in Figure 4H, often hint at which residues are rigorously bound to the HLA. Despite not being in the “core epitope,” the N-terminal CDR3 residues bound favorably to the HLA once the Arg3 and Asp6 formed a robust-yet-intricate buried salt bridge complex with β86E and α52R of the HLA (Figure 4I). In contrast, the N-terminal superagonist residue SHL (serine-histidine-leucine) not only had lower contact numbers with the HLA (Figure 4G), but also displayed larger fluctuations throughout the simulation (Figures 4H–4J). Furthermore, in addition to the tyrosine residue of CDR3 at position 7 (Y[P7]) as mentioned above (making strong π–π stacking with β47Y and hydrophobic interactions with β67V; Figure S6G), the core tyrosine residue at position 6(Y[P6]) also contributed favorably to binding by forming favorable π–π interactions with β11F, β30Y, and β61W (Figures 4J and S6F). Taken together, we conclude the strong π–π stacking and hydrophobic interactions from CDR3 peptide’s aromatic residues Y(P6) and Y(P7) contributed favorably to stronger binding with HLA, while the large fluctuation of the superagonist N terminus contributed slightly unfavorably toward its lower binding affinity. Taken together, results of MD simulations (MDSs) show that the CDR3 peptide (x-Id) appears to have optimal anchor residues for binding to DQ8.
The CDR3 Sequence of x-Clonotype Is a Potent CD4 T Cell Epitope
Expansion of the x-clonotype-expressing DEs in T1D and the unique DQ8 binding properties of the x-Id peptide suggest a connection to the disease pathogenesis. We considered and excluded the possibility that the x-clonotype encodes an IAA, because IAAs generally use VH06, have net positive charges, and have long CDR3 (Smith et al., 2015). In contrast, the x-clonotype uses VH04 and has a net negative charge (−2.01) and normal CDR3 length. Moreover, as mentioned above, the x-clonotype is absent from published sequences of IBCs. An alternative hypothesis was that the x-clonotype encodes a previously unknown DQ8-restricted CD4 T cell neoantigen. This hypothesis is directly supported by the results of MDS analysis (see Figure 4). To functionally test this idea, we examined the ability of two idiotypic peptides to form stable DQ8 complexes using a gentle SDS-PAGE assay (Kim et al., 2013; Sadegh-Nasseri and Germain, 1991). One peptide is the full CDR3 sequence, CARQEDTAMVYYFDYW (x-Id), and the second is a truncated version (TP-Id) that lacked cysteine (C) at the C-terminal and tryptophan (W) at the N terminus—ARQEDTAMVYYFDY. We used native insulin B: 9–23 and the mimotope as controls (Nakayama et al., 2015). We also tested binding of the CDR3 peptide (CARQRFWSGPLFDYW) of the IGVH04-b+ IgD+ clonotype from HC#1. Both idiotypic peptides (x-Id and TP-Id) were able to bind soluble DQ8 molecules forming SDS-stable complexes, as did the mimotope (Figure 5A). In contrast, the HC (h-Id) and native insulin peptides did not form detectable DQ8 complexes. These results show that x-idiotype (x-Id) from T1D, but not HC (h-Id), can form SDS-stable complexes with DQ8. Using CFSE proliferation assay and CD69 upregulation, we demonstrated that x-Id/DQ8 complexes, similar to mim/DQ8 complexes, were potent stimulators of CD4 T cells from DQ8+ T1D subjects. Responders included T1D#1, in which most of DEs expressed the IGHV04-b+ clonotype, hence an autoreactive response. Consistent with their poor binding to DQ8, native insulin and h-Id peptides generated no significant responses. The proliferative responses, as expected, were inhibited by anti-DQ8 mAb (Figure 5B). Importantly, x-Id/DQ8 complexes induced weak or no responses from healthy subjects, indicating that their high reactivity is associated with T1D (Figure 5B). In addition, most of CFSElow CD4 T cells upregulated CD69 as compared to their CFSEhi counterparts (Figure 5C). Similar MHC II-dependent responses results were obtained when the x-Id peptide was used to pulse PBMCs (Figure S7). These results identify, for the first time, a potent T cell autoantigen that is encoded in the idiotype of a public BCR.
Figure 5. x-Id Peptide Forms Functional HLA-DQ8 Complexes that Stimulate CD4 T Cells.
(A) Representative silver-stained SDS gel shows binding of indicated peptides to soluble DQ8 to form stable heterodimers. Arrows indicate p/DQαβ dimers and DQα and DQβ monomers, respectively. The results are from one of three independent experiments with similar results.
(B) x-Id/DQ8 complexes stimulate proliferation of CD4 T cells from DQ8+ T1D. Representative dot plots show CFSE dilution by gated CD4 T cells among PBMCs from in T1D or HC subjects that were stimulated with indicated peptide-DQ8 complexes. Numbers indicate percentages of gated CFSElow CD4 T cells. Dot plots on the right show inhibition of proliferation by anti-DQ mAb. Graph shows cumulative data from three DQ8+ T1D and three HC subjects (n = 3); *p < 0.05 by two-way ANOVA with Sidak’s multiple comparisons test.
(C) Overlays show upregulation of CD69 by gated CFSElow CD4 T cells (red line) versus CFSEhi CD4 T cells (green line) in each subject group. Numbers indicate percentages (mean ± SEM) of CFSElow CD4 T cells.
See also Figure S7.
Verification of Dual Expression of BCR and TCR by DEs Using an EBV-Immortalized Clone
Next, we successfully generated an Epstein-Barr virus (EBV)-immortalized lymphoblastoid cell line (x-LCL) from sorted DEs that were isolated from T1D#1 and transformed as described in STAR Methods (Caputo and Flytzanis, 1991; Hui-Yuen et al., 2011). Given that almost all DEs in T1D#1 expressed the x-clonotype (see Figure 3), we used x-LCL cells to clone and characterize the antibody encoding the x-clonotype. We sorted single cells from cultured x-LCL and examined each for transcripts of heavy and light chains. We confirmed expression of the x-clonotype in 7/7 single x-LCL cells using the same primers used for detection of the x-clonotype in fresh DE cells (Figure 6A). Analysis of transcripts of the light chain identified two light chains: a dominant productive light chain 1 (IGL1-x with CDR3: CSLYAGSNNVVVF) and a minor non-productive light chain 2 (IGL2-x) that was detected in two cells that could possibly expressed productive light chain(s) that were not picked by the PCR. Alternatively, IGL1-x and IGL2-x may be representing the same light chains, because the IGL2-x has the same nucleotide sequence of the IGL1-x except for missing 3 nt, which could be caused by PCR or reading errors. Thus, the antibodies expressed in DEs, at least in T1D#1, used the x-clonotype paired with at least one productive light chain.
Subsequently, we took advantage of x-LCL cells to verify coexpression of BCR and TCR in DE cells. We used limiting dilution (0.3 cell/well) and generated one clone (referred to as x1.1) that was confirmed by cloning PCR to express the IGL1-x light chain paired with the x-clonotype (Figure 6A). In addition, we have visualized coexpression of TCR, IgD, and IgM at the single-cell level using florescence imaging. Finally, using RT-PCR and nested PCR, we successfully amplified and cloned a TCRαβ composed of TRBV6–5*01/D1*01, JB1–1*01 (TCRβ-x), and TRAV29/DV5*01/J53*01 (TCRα-x) from cells of the x1.1 clone (Figure 6A). Detection of fully rearranged and expressed BCR and TCR chains from cells of the x1.1 clone confirms their dual expression in DEs.
x-Id-Peptide and Insulin Mimotope Recognize Overlapping Subpopulations of CD4 T Cells
Antibodies can activate T cells by being sources of soluble autoantigens (Khodadoust et al., 2017), and idiotypic-specific CD4 T cells have been described in multiple sclerosis (Hestvik et al., 2007) and lupus (Aas-Hanssen et al., 2014). As shown above, the x-Id peptide can serve as an autoantigen by forming functional complexes with DQ8 molecules (see Figure 5). Cultured x1.1 cells secreted copious amounts of an x-clonotype-encoding antibody, herein referred to as x-mAbN. We assessed the ability of the x-mAbN to stimulate CD4 T cells from T1D subjects using the CFSE proliferation assay (Figure 6B). Addition of soluble x-mAbN to PBMCs resulted in potent proliferation as indicated by the percentage of CFSElow CD4 T cells. We have also generated a recombinant x-mAb of IgG1 isotype (referred to x-mAbR). We cloned the variable regions of the heavy and light chains from sorted single-DE cells isolated from T1D# 1 and fused into human IgG vectors as described in the STAR Methods (Figure 7A). Similar to its natural counterpart, immobilized x-mAbR activated CD4 T cells in a CFSE proliferation assay (Figure 7B). Thus, both naturally produced and recombinant x-mAbs are stimulatory for CD4 T cells.
To examine the relationship between x-Id- and insulin-reactive CD4 T cells, we generated DQ8 tetramers loaded with the x-Id peptide (x-Id-tetramer) or the insulin mimotope (mim-tetramer). We used CLIP-DQ8 tetramer as a specificity control. We cultured PBMCs from T1D subjects in the presence or absence of x-Id peptide or insulin mimoptope for 7 days and analyzed each culture for the presence of tetramer-positive CD4 T cells (Figure 7C). We detected x-Id and mimotope specific CD4 T cells in both unstimulated and stimulated cultures, using CLIP-tetramer staining as a negative control. Furthermore, percentages of tetramer positive cells were significantly higher in stimulated cultures as compared to unstimulated cultures. More importantly, the x-Id-tetramer was able to detect CD4 T cells expanded by the insulin mimotope and the opposite was true for the mim-tetramer which detected CD4 T cells expanded by the x-Id peptide. These results suggested the mimotope and x-Id peptide stimulated the same or overlapping subpopulations of CD4 T cells. In direct support of this notion, preincubating with x-mAb significantly inhibited binding of the mim-tetramer to activated CD4 T cells (Figure 7C). Thus, it appears that x-clonotype cross-activates insulin-specific CD4 T cells.
DISCUSSION
This study describes rare lymphocytes (DEs) that clonally expand in T1D subjects and bear lineage markers of both B and T cells, epitomized by the expression of TCR and BCR. Clonally expanded DEs encode a potent autoantigen (x-autoantigen) in the variable region of the Ig heavy chain with an optimal register for the diabetogenic DQ8 molecule. The x-autoantigen peptide forms functional complexes with DQ8 molecules that robustly stimulate CD4 T cells from T1D, but not HC subjects. In addition, x-clonotype-bearing mAbs also stimulate CD4 T cells. Competitive binding inhibition analysis indicates that the x-mAb and insulin mimotope stimulate overlapping T cell subpopulations (Figure 7C). Taken together, these findings indicate that compartmentalization of T and B cells is not absolute, and violators of this paradigm could be key drivers of autoimmunity.
Features of the x-Autoantigen and Implications for T1D
The −Id peptide has an optimal binding register for DQ8 with acidic residues at P1 and P9 as revealed by MDS analysis. Significance of having acidic residues at these anchor positions is indicated by transformation of native insulin B:9–23 peptide into a potent superagonist by substituting glutamic acids for alanine at P1 and arginine at P9. In addition, BCR idiotypes are frequently processed and presented in the context of MHC II as neoantigens (Khodadoust et al., 2017). Moreover, idiotype-specific CD4 T cells have been described in multiple sclerosis (Hestvik et al., 2007) and lupus (Aas-Hanssen et al., 2014), but cell types producing these antibodies have not been identified. These observations, taken together, suggest that DEs could the source of idiotypic autoantigens in autoimmune diseases. Furthermore, our findings that x-mAbN and x-mAbR are potent stimulators of CD4 T cells in T1D subjects suggest a previously unknown role for antibodies in driving pathogenesis of T1D.
Expression of Functional TCR on DE Cells and Pathogenic Implications
Expression of the TCR on DEs could have important implications. For one, it gives DEs the ability to expand and increase their numbers upon TCR stimulation. In addition, crosslinking of TCR on DEs leads to upregulation of MHCII and costimulatory molecules, including CD40 and CD80/CD86, thereby converting DEs into potentially potent antigen-presenting cells APCs. These features will be pathogenically relevant if future analysis shows that DEs process and present x-Id peptide on surface DQ molecules and/or use membrane Ig to stimulate autoreactive CD4 T cells. TCR stimulation of DEs also leads to production of cytokines that can modulate immune responses. Future studies should investigate antigen specificities of TCRs expressed on DEs, including against common pathogens.
Immediate Translational Impacts and Future Significance
Our findings lay the groundwork for examining whether the x-autoantigen could serve as a T1D risk biomarker. DQ8 is a major risk factor for T1D that is carried by about 90% of patients. However, only a fraction of DQ8+ subjects develops T1D. It’s therefore interesting that in the very limited numbers of subjects examined, the x-clonotype was detected in DQ8+ T1D patients, but not in DQ8+ HCs. The x-clonotype was also absent in HCs carrying other DQ alleles. The exceptions were subjects carrying DQ7 (β57D+ the neutral isoform of DQ8), which favors a positively charged residue at P9 and cannot optimally bind the x-clonotype, at the least in the same register that the DQ8 molecule does. Screening of well-characterized at-risk subjects will address the value of the x-clonotype as a biomarker.
Future studies should include extensive scRNA-seq analysis to determine whether DE cells represent a distinct new cell type or a subpopulation of an existing cell type. In addition, the short read length (38 bp) prevented us from accurately analyzing CDR3 sequences of DEs, and we plan to use read length of at least 50 bp in the future (Rizzetto et al., 2017). Moreover, it will be important to determine antigen specificities of the TCRs and BCRs in DEs and mechanisms allowing their dual expression. Furthermore, if the x-clonotype proved to be a key driver of T1D, its conserved germline sequence will be useful in developing antigen-specific therapeutic strategies.
STAR★METHODS
CONTACT FOR REAGENT AND RESOURCE SHARING
Further information and requests for resources and reagents should be directed to and will be fulfilled by the Lead Contact, Abdel Rahim A. Hamad (ahamad@jhmi.edu).
EXPERIMENTAL MODEL AND SUBJECT DETAILS
Human subjects
Peripheral blood samples were obtained from donors using protocols approved by the Johns Hopkins Institutional Review Board. All donors provided written informed consent and sample size estimation was not employed. All T1D subjects met the American Diabetes Association criteria for classification and were recruited at Johns Hopkins Comprehensive Diabetes Center. Donors with no T1D are classified as healthy controls (HCs) and were recruited from normal volunteers. Patients were males and females. The clinical characteristics of donors are summarized in (Table S1A). The study was conducted in accordance with the declaration of Helsinki principles. Peripheral blood mononuclear cells (PBMCs) were freshly isolated using Ficoll-paque density centrifugation (GE Healthcare) gradient. Islet autoantibodies profiles and HLA genotypes of subjects (Tables S1B and S1C), whose repertoires were analyzed by high-throughput was performed at Autoantibody/HLA Core Service Center at the Barbara Davis Center for Childhood Diabetes.
Cell lines
HEK293 (female) were obtained from Thermo Fisher Scientific and cultured in basal media (An aliquot of 250 mL each of sterile RPMI and DMEM; 3.75 mL of antibiotic/antimycotic and 5 mL each of L-glutamine (200 mM), 100 × Nutridoma and sodium pyruvate (100 mM) was used. Basal media must be made fresh every 7 d. L-Glutamine can be stored at −20° C for up to 1 year, Nutridoma can be stored at room temperature (20–25° C) for up to 1 year and sodium pyruvate can be stored for up to 6 months at 4° C. Cells were incubated at 37° C in humidified air containing 5% CO2 and transfections were performed using polyplus jet-prime transfection reagent (Polypus transfection) according to the manufacturer’s protocols.
Bacterial strains
The bacterial strains used in this study are listed in the Key Resources Table. LB agar or LB broth (Invitrogen) were used for bacterial growth at 37° C. Supplements, 50 mM (20 mg/mL) X-Gal (fisher scientific) was applied directly to the top of the agar plates for blue/white screening. Antibiotics were added to media at the following concentrations: 50 μg/mL of ampicillin. In all procedure, 37°C shaking (225 rpm) and non-shaking incubator have been applied. Glycerol stocks of each culture have been provided by transferring 300 mL of 1:1 sterile LB/glycerol and 700 mL of the confluent culture to a 2 mL tube, mix well and freeze at −80° C.
KEY RESOURCES TABLE.
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| APC anti-human CD5 (L17F12) | BioLegend | Cat#364016; RRID: AB_2565726 |
| FITC anti-human CD5 (L17F12) | BioLegend | Cat#364022; RRID: AB_2566248 |
| PE/Cy7 anti-human CD5 (UCHT2) | BioLegend | Cat#300622; RRID: AB_2275812 |
| BV421 anti-human CD19 (HIB19) | BD Biosciences | Cat#562440; RRID: AB_11153299 |
| PerCP anti-human CD19 (HIB19) | BioLegend | Cat#302228; RRID: AB_893272 |
| V450 anti-human CD19 (HIB19) | BD Biosciences | Cat#560353; RRID: AB_1645564 |
| AF-488 anti-human TCRαβ (IP26) | BioLegend | Cat#306712; RRID: AB_528967 |
| Pacific Blue anti-human TCRαβ (IP26) | BioLegend | Cat#306716; RRID: AB_1953257 |
| APC anti-human TCRαβ (IP26) | BioLegend | Cat#306718; RRID: AB_10612569 |
| PE anti-human IgD (IA6-2) | BioLegend | Cat#348204; RRID: AB_10553900 |
| PE/Cy7 anti-human IgD (IA6-2) | BioLegend | Cat#348210; RRID: AB_10680462 |
| FITC anti-human IgM (MHM-88) | BioLegend | Cat#314506; RRID: AB_493009 |
| PE-Cy7 anti-human IgG (G18-145) | BD Biosciences | Cat#561298; RRID: AB_10611712 |
| PE anti-human IgG (G18-145) | BD Biosciences | Cat#555787; RRID: AB_396122 |
| PE anti-human IgA (IS11-8E10) | Miltenyi Biotec | Cat#130-093-128; RRID: AB_1036158 |
| PerCP/Cy5.5 anti-human IgE (MHE-18) | BioLegend | Cat#325512; RRID: AB_11219590 |
| APC/Cy7 anti-human CD69 (FN50) | BioLegend | Cat#310914; RRID: AB_314849 |
| APC/Cy7 anti-human CD45 (HI30) | BioLegend | Cat#304014; RRID: AB_314402 |
| APC/efluor anti-human CD79a (HM47) 780 | eBioscience | Cat#47-0792-41; RRID: AB_2573967 |
| PerCP/Cy5.5 anti-human CD79b (3A2-2E7) | BioLegend | Cat#341408; RRID: AB_2075869 |
| FITC anti-human CD4 (OKT-4) | eBioscience | Cat#50-939-1; RRID: AB_1633390 |
| PE/Cy7 anti-human CD4 (A161A1) | BioLegend | Cat#357410; RRID: AB_2565662 |
| APC anti-human CD4 (OKT-4) | BioLegend | Cat#317416; RRID: AB_571945 |
| PerCP-Cyanine5.5 anti-human CD4 (RPA-T4) | eBioscience | Cat#13557660; RRID: AB_1518744 |
| PE anti-human CD8a (HIT8a) | eBioscience | Cat#12-0089-41; RRID: AB_10804878 |
| FITC anti-human CD40 (5C3) | eBioscience | Cat#11-0409-41; RRID: AB_1272137 |
| PE anti-human CD154 (CD40L) (24-31) | BioLegend | Cat#310806; RRID: AB_314829 |
| FITC anti-human CD80 (2D10.4) | eBioscience | Cat#50-967-7; RRID: AB_10853194 |
| PE anti-human CD86 (IT2.2) | eBioscience | Cat#12-0869-41; RRID: AB_10804536 |
| BV421 anti-human CD3 (UCHT1) | BD Biosciences | Cat#562426; RRID: AB_11152082 |
| PerCP anti-human CD3 (UCHT1) | BioLegend | Cat#300428; RRID: AB_893298 |
| Alexa Fluor 700 anti-human CD28 (CD28.2) | BioLegend | Cat#302920; RRID: AB_528786 |
| PE anti-human CD28 (CD28.2) | BioLegend | Cat#302902; RRID: AB_314304 |
| PE anti-human HLA-DQ (HLADQ1) | BioLegend | Cat#318106; RRID: AB_604129 |
| PE anti-human HLA-DR (L243) | BioLegend | Cat#307606; RRID: AB_314684 |
| Anti-human HLA-DQ (SPV-L3) | Abcam | Cat#ab23632; RRID: AB_447580 |
| Purified anti-human HLA-DR (L243) | BioLegend | Cat#307602; RRID: AB_314680 |
| APC/Cy7 anti-human CD27 (O323) | BioLegend | Cat#302816; RRID: AB_571977 |
| PE/Cy7 anti-human CD38 (HB-7) | eBioscience | Cat#356608; RRID: AB_2561904 |
| InVivoMab anti-human CD3 (OKT-3) | Bio X Cell | Cat#BE0001-2; RRID: AB_1107632 |
| InVivoMab anti-human CD28 (9.3) | Bio X Cell | Cat#BE0248; RRID: AB_2687729 |
| PE anti-human IL-10 (JES3-9D7) | eBioscience | Cat#12-7108-81; RRID: AB_466178 |
| eFluor 450 anti-human IFN-γ (4S.B3) | eBioscience | Cat#48-7319-41; RRID: AB_2043867 |
| AffiniPure F(ab’)2 fragment Goat anti-human IgM, Fcsu fragment specific | Jackson ImmunoResearch | Cat#109-006-064; RRID: AB_2337548 |
| Bacterial and Virus Strains | ||
| DH5α competent cells | Invitrogen | Cat#18265017 |
| JM109 competent Cells | Promega | Cat#L2005 |
| Epstein-Barr Virus B95-8 | American Type Cell Culture (ATCC) | Cat#ATCC VR-1492 |
| Biological Samples | ||
| Human peripheral blood samples | This paper (Table S1) | N/A |
| Chemicals, Peptides, and Recombinant Proteins | ||
| CFSE | eBioscience | Cat#65-0850-84 |
| eBioscience Streptavidin PE | eBioscience | Cat#12-4317 |
| CpG ODN 2006 (ODN 7909) | InvivoGen | Cat#tlrl-2006-1 |
| Nutridoma-SP (basal media) | Sigma-Aldrich | Cat#11011375001 |
| Protein A Agarose Beads | Thermo Fisher Scientific | Cat#15918014 |
| x-Idiotype peptide (x-Id) (CARQEDTAMVYYFDYW) | This paper | N/A |
| Truncated peptide (TP-Id) (ARQEDTAMVYYFDY) | This paper | N/A |
| HC peptide (h-Id) (CARQERFWSGPLFDYW) | This paper | N/A |
| Insulin peptide (Ins) (SHLVEALYLVCGERG) | This paper | N/A |
| Insulin Mimotope peptide (Mim) (SHLVEALYLVCGEEG) | Crawford et al., 2011 | N/A |
| Biotinylated-monomers DQB1*03:02/DQA1*03:01 | NIH Tetramer Core Facility | N/A |
| Critical Commercial Assays | ||
| Qiaquick Gel Extraction Kit | QUIGEN | Cat#28704 |
| Qiaprep Spin Miniprep Kit | QUIGEN | Cat#27106 |
| Qiaprep Plasmid Maxi Kit | QUIGEN | Cat#12162 |
| OneStep RT-PCR Kit | QUIGEN | Cat#210212 |
| Monarch Total RNA Miniprep Kit | BioLabs | Cat#T2010S |
| QIAamp DNA Blood Mini Kit | QUIGEN | Cat#51104 |
| RNeasy Mini Kit | QUIGEN | Cat#74104 |
| RevertAid First Strand cDNA Synthesis Kit | QUIGEN | Cat#K1621 |
| Pierce Protein A Columns | Pierce | Cat#20356 |
| EZQ Protein Quantitation Kit | Invitrogen | Cat#R33200 |
| Pro-DetectTM Rapid Antibody Isotyping kit, human | Thermo Fisher Scientific | Cat#A38552 |
| Deposited Data | ||
| RNA-seq from DEs, T cells and B cells | GEO Data Repository | GEO: GSE129112 |
| Immuno-Seq | Adaptive Biotechnologies | 10.21417/RA042019 |
| IGH-x | This paper; GenBank | MK764540 |
| IGL1-x | This paper; GenBank | MK764541 |
| IGL2-x | This paper; GenBank | MK764542 |
| TCRA-x | This paper; GenBank | MK764543 |
| TCRB-x | This paper; GenBank | MK764544 |
| Experimental Models: Cell Lines | ||
| HEK293A cells | Thermo Fisher Scientific | Cat#R705-07 |
| IRR-MRC-5 iRRADIATED Fibroblast | American Type Cell Culture (ATCC) | Cat#ATCC 55-X |
| Oligonucleotides | ||
| GCTGGAGTGGATTGGGAGTA (Forward) Probe1 | This paper | N/A |
| CCCAGTAGTCAAAGTAGTAAACCATA (Reverse) Probe1 | This paper | N/A |
| GCTGGAGTGGATTGGGAGTA (Forward) Probe2 | This paper | N/A |
| TCCCTGGCCCCAGTAGTCAAAGTAGTA (Reverse) Probe2 | This paper | N/A |
| RT-PCR and Cloning-PCR primers used for amplification of Heavy and Light chains of single DE cells and x-mAbR | Smith et al., 2009 | N/A |
| RT-PCR and Nested PCR primers used for amplification of TCRα and TCRβ chains of x1.1 DE clone | Eugster et al., 2013 | N/A |
| Recombinant DNA | ||
| AbVec -Igγ, expression vectors | Smith et al., 2009 | N/A |
| AbVec –Igλ, expression vectors | Smith et al., 2009 | N/A |
| pGEM-T Easy Vector Systems | Promega | Cat#A1360 |
| Software and Algorithms | ||
| FlowJo | TreeStar | https://www.flowjo.com/solutions/flowjo |
| GraphPad Prism | GraphPad Software | https://www.graphpad.com/scientific-software/prism/ |
| ImmunoSEQ Analyzer 2.0 software | Adaptive Biotechnologies | N/A |
| IDEAS6.2 | MilliporeSigma | http://www.emdmillipore.com/US/en |
| BASIC: BCR and TCR assembly from single cell RNA-seq | Toyota Technological Institute at Chicago | https://ttic.uchicago.edu/~aakhan/BASIC/ |
| IMGT/V-QUEST | IMGT, the international ImMunoGeneTics information system | http://www.imgt.org/IMGT_vquest/share/textes/ |
METHOD DETAILS
Flow cytometric analysis
Cell phenotypes were analyzed by flow cytometry (Dai et al., 2015; Martina et al., 2016). Briefly, single cell suspensions were surface-stained for 20 min on ice with predetermined optimal concentrations of indicated fluorochrome-conjugated antibodies (Key Resources Table). Acquired samples (5 × 105 to 1 × 106 live events) were properly compensated using single color stains. Data analysis, gating, and graphical presentation were done using FlowJo software (TreeStar). Doublets were excluded from analysis using FSC-Height versus FSC-Width and SSC-Height versus SSC-Width plots. Multiple specificity controls were used. These included human FcR blocking reagent (Miltenyi Biotec), Fluorescence-Minus One (FMO) for CD5, CD19, TCR, IgD, dump gating, and isotype controls. In addition, when applicable, irrelevant cell types were used as internal biological controls and in the case of in vitro stimulation, we used unstimulated cultures as negative controls.
Imaging flow cytometry (AMNIS)
Freshly isolated PBMCs were stained with FITC-conjugated anti-TCRαβ, PE-conjugated anti-IgD, APC-conjugated anti-CD5, and BV421-conjugated anti-CD19 and analyzed at X60 magnification on an Image Stream flow cytometer (Amnis corporation) with low flow rate/high sensitivity using INSPIRE software. For each sample, 10,000 events were acquired. Single color controls were used for creation of a compensation matrix, to set the optimal laser power for each fluorochrome and to avoid saturation of the camera. The compensation matrix was applied to all files to correct for spectral cross-talk. Positive cutoff values were calculated on the basis of the bright detail similarity (BDS) background of TCRαβ and an irrelevant signal (for example, side scatter). Image analysis was performed with the IDEAS 6.2 software package using bright field images to set cell boundary and gating on internalized events. Compensated data files were analyzed using a gating strategy that involved selecting focused cells on the basis of gradient RMS and an aspect ratio that was consistent with single events and devoid of debris or multi-cellular events (doublets). T cell and B cell singlets were successfully identified using this strategy and the selection of good quality, focused singlets within the viewing window allowed refining of final gating. After the gating of Tcon and Bcon cells, individual IgD+ DE cells were identified based on their surface profile (CD19+CD5+TCR+IgD+) and analyzed for the indicated markers. Bright field imagery was collected with an LED-based bright field illuminator. Each plot was manually adjusted so that the machine noise generated at the beginning of acquisition was set to zero.
Single cell RNA-seq data generation and processing
FACS sorted single cells (see Figures S5A and S5B for sorting strategy) were processed with the Smart-seq2 protocol (Picelli et al., 2014) with the following modifications. RNA purification was performed prior to reverse transcription using RNAClean XP beads (Beckman Coulter). cDNA was amplified with 21 PCR cycles followed by DNA cleanup with AMPure XP beads. Libraries were prepared using the Nextera XT Library Prep kit (Illumina) using custom barcode adapters. Uniquely barcoded Libraries were sequenced together on a NextSeq 500 sequencer (Illumina).
Bioinformatic analysis of scRNA-seq Data
QC checks were performed on the scRNA-seq data with R bioconductor package scater following the methods described by Lun et al. (Lun et al., 2016). The QC metrics included library size, number of features expressed, proportions of ERCC spike-in controls, and three empty wells that were included in the experimental design as negative controls. In addition to the three empty wells, 18 out of 93 biological (B, T and DE) cells had either log-library sizes and/or log-transformed number of expressed transcripts blow the respective medians by more than 3 median absolute deviations (MADs) and were filtered out as low quality outlier samples. Another DE-cell D07 had a library size below the maximum of the three empty wells and was viewed as a low quality sample. Among the 19 low-quality biological cell samples, 12/45 are DE cells, 6/24 Bcon cells and 1/24 Tcon cells. All the 19 cells had library sizes lower than or comparable to the empty wells. 64 out of the 74 good quality samples have a sequencing depth of 1–3 million reads and are deemed to reach saturation while the other 10 samples have a depth between 0.7–1 million reads, good for the detection of large majority of genes (Michel et al., 2012; Wu et al., 2014; Ziegenhain et al., 2017). The sequencing assay kit also included 12 ERCC spike in controls. The 19 low quality cells had a pattern of spike-in ERCC proportions similar to the good quality ones above and did not show any increase. Assuming the majority of cells are of high quality, it suggests there is little loss of endogenous RNA in all the cells. Taken together, the analyses above suggest good overall quality of the scRNA-seq experiment.
Following a biology-guided strategy, we limited downstream analysis of the scRNA-Seq data to cells in which at least two of three housekeeping genes (PPIA, ACTB, and UBB) were detected as expressed, defined as having log2 (RSEM value + 1) > 0. This resulted in 77 high quality cells. Genes preferentially expressed in either Bcon cells, Tcon cells, or DE cells were identified using the Template Matching method, which tests for an association between each profile and an artificial profile that represents an ideal, cluster- or condition-specific, profile using the Pearson’s product moment correlation coefficient (Pavlidis and Noble, 2001). Multiple testing corrections were performed using Holm’s method (Holm, 1979). To identify BCR and TCR transcripts expressed in the single cells, we used BASIC (Canzar et al., 2017). We annotated each cell according to whether BASIC was able to assemble BCR or TCR transcripts.
Polyclonal TCR stimulation
Freshly isolated PBMCs were placed onto the wells of 24-well tissue culture plates (106 cells in 1 mL complete culture medium) in the presence or absence of anti-CD3/CD28 beads (106 bead/well) and incubated at 37° C and 5% CO2. Alternatively, plates coated with anti-CD3 (10 μg/mL) and anti-CD28 (10 μg/mL) were used (Yoneshiro et al., 2017). After 7 days in culture, viable cells were harvested, counted using trypan blue, and analyzed for the expression of indicated molecules using a BD LSRII flow cytometer. Absolute cell numbers were determined by multiplying the frequency of the indicated subset by the viable cell count.
CFSE proliferation assay
Freshly isolated PBMCs were washed twice with warm (37° C) 1× PBS to remove serum that affect staining and the cells were resuspended in warm (37° C) 1× PBS at a density of 1.5–2.0 × 106 cells/mL. Cells were labeled with 1 μM CFSE (eBioscience) for 1–2 min at 37° C with continuous vortexing. The labeling reaction was quenched by adding chilled complete culture media. CFSE-labeled cells were washed in 1× PBS, resuspended in complete media, and plated into 24-well tissue culture plates (1.5–2.0 × 106 cells/well in 1 mL complete culture medium). To evaluate functionality of HLA-DQ8 molecules, we immobilized DQ8 molecules loaded with indicated peptides (x-Id, TP-Id, mimotope, native insulin and CDR3 peptide from IgD+ DE from HC#1 (referred to as h-Id) into wells of 24-well plates (10 μM) and examined their ability to stimulate CFSE-labeled CD4 T cells from among PBMCs. In parallel experiments, we activated cultures in the presence (20 uM) of mouse anti-HLA-DQ (SPV-L3; Abcam) and anti-HLA-DR (L243;Abcam) to assess MHC restriction. In similar manner, CFSE-labeled cells were also stimulated in the presence or absence of the above-indicated peptides (10 μM) as soluble antigen. Alternatively, in a separate experiment, to evaluate the mAb-specific proliferative response, purified mAbR and mAbN (described later in the method) concentration of 2.5 and 5ug, immobilized into the wells of 24-well plates, and used to stimulate CFSE-labeled PBMCs. CFSE labeled cells without stimulation and with CD3–28 stimulation were taken as specific negative and positive controls respectively. After 7 days of incubation, cultures were stained as indicated in Figure legends and proliferation assessed by determining frequency of CFSElow CD4 T cells.
Intracellular Cytokine analysis
Single cell suspensions were stimulated for 4 h at 37°C in 5% CO2 with phorbol 12-myristate 13-acetate (PMA) (50 ng/mL) and ionomycin (500 ng/mL) in the presence of Golgi-Plug (Saxena et al., 2017; Xiao et al., 2011). Intracellular cytokine analysis was performed using the manufacturers’ instructions. Briefly, surface-stained samples were fixed, permeabilized and incubated with mAbs against indicated intracellular cytokines for 30 min on ice, washed, acquired and analyzed using the above described strategy.
Anti-IgM stimulation
Freshly isolated PBMCs (1X106) in RPMI-1640 medium were allowed to rest at 37° C in CO2 incubator for 30 min before stimulation. BCR signaling was induced by crosslinking the BCR (Wang et al., 2014) with 10 μg/mL goat F(ab′) 2 anti-IgM (Jackson ImmunoResearch) at 37° C in CO2 incubator for indicated time points. Time course analysis was achieved by adding anti-IgM to each sample in reverse time points and fixing all samples in unison. To determine basal levels of phosphorylation, parallel cultures of unstimulated PBMCs were fixed at time zero. To detect phosphorylated CD79a (pIgα), cells were fixed (1.5% paraformaldehyde, 5 min, room temperature), permeabilized (90% methanol, 10 min, 4° C), and stained with rabbit antibodies specific for pCD79A (Igα, Tyr82) followed by PE–conjugated goat anti-rabbit IgG (Jackson ImmunoResearch Laboratories).
Cell sorting and DNA extraction for high-throughput sequencing of IGHV and TRB
For repertoire analysis, IgD+ and IgD− DE cells, Bcon, and Tcon cells were sorted from freshly isolated PBMCs using a FACSAria II (BD Biosciences, Bedford, MA). The sorting strategy and purity of isolated cell populations are shown in (Figures S5A and S5B). Two sorts were performed from each donor and (except HC#2) were performed at different time points with one sort used for IGHV and the second for TRB analysis. Donor characteristics, including islet autoantibodies and HLA genotypes, are shown in (Table S1). Autologous Bcon cells were used as controls for IGVH analysis and Tcon cells for TRB analysis. Briefly, freshly isolated PBMCs were stained for CD19, CD5, IgD, and TCRαβ for 30 min on ice, washed thoroughly, and suspended in a pre-sort buffer (BD Biosciences). Propidium iodide (PI) was added immediately prior to sorting to exclude non-viable cells. Sorted cells were collected in RPMI medium supplemented with 50% FBS on ice. IgD+ DE cells were identified as CD19+CD5+IgD+TCRβ+ (800–1000 cells per sort) and IgD− DE cells as CD19+CD5+IgD−TCRβ+ cells (100–200 cells per sort). Bcon cells were identified as CD19+CD5−TCRβ− and Tcon cells as CD19−CD5+TCRβ+ cells. Total DNA was directly extracted from sorted cells using QIAmp DNA blood mini Kit (QIAGEN) according to the manufacturer’s instructions. DNA from sorted IgD+ and IgD− DE cells, Bcon cells and Tcon cells were used for BCR or TCRBV sequencing as described in text.
High-throughput immune SEQ and data analysis
Analyses of IGHV and TRBV clonotypes were performed on genomic DNA from each sorted cell type using the immunoSEQ platform at survey level resolution (Adaptive Biotechnologies). The immunoSEQ assay combines multiplex PCR with high-throughput sequencing and sophisticated bioinformatics pipeline for CDR3 region analysis (Carlson et al., 2013; DeWitt et al., 2016; Robins et al., 2009). Samples were amplified from 40 ng to 100 ng genomic DNA per sample. Attempts to sequence IgD− from T1D#1 and IgD+ cells from T1D#2 were unsuccessful. TCRβ and IGH sequences are available at Adaptive Biotechnologies (DOI: https://doi.org/10.21417/RA042019).
Raw ImmunoSeq data from individual samples were processed with ImmunoSeq Analyzer 2.0 software (Adaptive Biotechnologies). Measurement metrics of processed data were exported in the tsv file format and analyzed using the R platform. Clones of uncertain vGene identity or out-of-frame were excluded from downstream analysis. For vGene in each cell type in individual samples, counts of distinct cell clones were obtained by summing the metric of the “estimated number of cell genomes present in the sample,” upon which the corresponding percentages were calculated. The percentage quantification provides a uniform basis for the vGene (VH and Vβ) usages to be fairly and consistently compared across the different cell types and samples, minimizing any effects that could result from sequencing different and very tiny numbers of DE subset cells. Percentages were visualized with bar plots to make straight comparisons of vGene usages between different cell types. The presence or absence of vGenes in the different cell subsets was determined on the basis of the vGene usages. Unique and shared vGenes among different cell subsets were identified and displayed in Venn diagrams using the functions of R Limma package. The vGene mutations are identified based on alignment with the IMGT database, upon which the differences from germlines are marked, counted and recorded in the column of “vAlignSubstitution-Count” of the Raw ImmunoSeq data tsv spreadsheet. The vGene mutation values were further summed per gene and displayed with a combination of boxplot and scatterplot using R.
String searches with the amino acid sequences of the selected clones were performed to determine their presence in individual subjects. A string search of the amino acid sequence of the invariant clonotype, “CARQEDTAMVYYFDYW” was also performed with an R script against a public ImmunoSEQ database of 37 million unique BCR sequences of naive and memory Bcon cells, which was downloaded from https://clients.adaptivebiotech.com/immuneaccess and that of IBC examined from nPOD Adaptive Immune Repertoire (https://public.tableau.com/profile/npod.adaptive.immune.repertoire#!/).
PCR probes for detection of x-clonotype in peripheral blood
To determine whether the x-Id clonotype can be detected in peripheral blood, we designed and used two PCR probes for analysis of PBMCs. In the first probe we used a VH04-b-specific sense primer (5′- GCTGGAGTGGATTGGGAGTA-3′) paired with antisense primer (5′- CCCAGTAGTCAAAGTAGTAAACCATA-3′) complementary to the entire CDR3 region (see diagram, Figure 3J). In the second probe, the VH04-b-specific primer was paired with a reverse primer (5′-TCCCTGGCCCCAGTAGTCAAAGTAGTA-3′) that span JH04 and ended at the N2 region (see diagram, Figure S7). Briefly, RNA was extracted from fresh PBMCs using the RNeasy mini kit (Quigen), and analyzed by NanoDrop (ND-1000 spectrophotometer) to assess purity and measure concentration. Reverse transcription (RT) PCR was performed on approximately 1 μg of purified RNA to prepare cDNA using the RevertAid First Strand cDNA Synthesis Kit (Thermofisher) according to the kit protocol: RNA was incubated with 5X reaction mix, random hexamer primer, and RevertAid M-MuLV RT (200 U/μL) enzyme mix in a final volume of 20 μL at 25°C for 10 min, followed by 42°C for 60 min and inactivation at 70°C for 5 min. Positive (GAPDH-specific primers) and negative (reaction mixture without RT enzymes) control reactions were used to verify specificity of cDNA synthesis. PCR reaction (2 μL cDNA in a total volume of 25 μL prepared using 2X QIAGEN HotStarTaq master mix) was performed under the following conditions: initial denaturation at 95°C for 3 min, 95°C for 30 s, 54°C for 30 s, 40 cycles at 72°C for 1 min followed by a final extension step at 72°C for 10 min using a thermocycler (BioRad T100). PCR product was visualized as a band size of 200 bp on 1.2% agarose gel and the band was excised, purified using PCR purification kit (Quigen) and Sanger sequenced at the Johns Hopkins Medical Institute GRCF sequencing core. Sequences were analyzed using the Immunogenetics IMGT/V-QUEST software: http://imgt.org/IMGT_vquest/share/textes/.
Molecular dynamics simulations
The new peptide system was built from a crystal structure of an insulin B chain epitope bound to HLA-DQ8 (PDB: 1JK8) (Sharp, 2012). The insulin epitope sequence was mutated to the new peptide epitope sequence using the Mutator plugin from VMD, ensuring the new peptide epitope was in the desired register. The CDR3 epitope from HC#1 was also built from the insulin-bound epitope structure (PDB: 1JK8), following the same protocol as the new peptide system. The super-agonist system was built from the crystal structure of an insulin mimotope bound to HLA-DQ8 (PDB: 5UJT) (Wang et al., 2018). For this system, in addition to mutating the epitope to match the super-agonist, both HLA chains α and β were mutated to match the sequence of the HLA in the insulin crystal structure (PDB: 1JK8). More specifically, besides distal residues, residue 72C of HLA-α was mutated to Isoleucine to match the 1JK8 HLA sequence. Each system was solvated in a TIP3P water box and then charged, neutralized, and ionized with 100mM concentration using Na+ and Cl−.
Following system creation, each system underwent at least 20,000 steps of conjugate-gradient minimization to hold protein atoms fixed, followed by at least 10,000 steps of minimization allowing all the atoms to move. The systems were subsequently equilibrated for 1ns at 310K using a 2fs timestep. Production MD simulations were run for 500ns using a 2fs timestep. A Langevin thermostat maintained the temperature at 310K. The CHARMM36 force field (Best et al., 2012) was used for protein parameters. The Particle Mesh Ewald (PME) method was used to compute long-range electrostatics with the electrostatics and van der Waals cutoff of 12Å. All simulations were run using NAMD2.11. For the MD simulations, only the last 250ns were used for analysis, dividing the trajectory into 5 parts.
The contact area was computed using solvent accessible surface area (SASA) calculation in Gromacs tools with a water radius of 1.4Å. Van der Waals interaction energy was computed using NAMDEnergy. The electrostatics energy was biased due to the absence of solvent screening and was left out of the interaction energy. The RMSD and RMSF were computed using Gromacs tools. Averages and error bars for contact area, interaction energy, and RMSF were computed by taking the last half (250ns) of the MD simulations and dividing them into 5 sections with 50ns each and taking the average of each section as a measurement in the sample. Error bars shown are standard error.
Free energy perturbation
Binding affinity was calculated via the free energy perturbation (FEP) method. The final structures of the production MD simulations were selected for FEP computation. We computed free energy perturbation calculations for the bound (HLA + epitope) and free states (epitope only) with 6 replicas for each calculation. Due to the extensive sequence differences between epitopes, we mutated the epitopes to a neutral, intermediate sequence of polyglycine the length of the epitope. The dual topology was implemented using the Mutator plugin from VMD. Each system was slowly mutated from the epitope to polyglycine using λ increments of maximum 0.04 with smaller increments toward the ends, totaling at least 34 FEP windows for each system. Each FEP window was run for 1ns, leading to well over 800ns simulation (6 replicas × 34 windows × 2 states (complex + free) × 2 epitopes). Electrostatics was switched on starting at λ = 0.1. Convergence at each window was assessed by comparing values across replicas. NAMD2.11 with the CHARMM36 protein force field and TIP3P water model were used for FEP calculations, matching the MD work. From observing the FEP trajectories, the polyglycines do not shift registers but maintain the starting register of the epitope. Free energy error bars are standard errors.
Analysis of x-Id peptide binding to DQ8 using gentle SDS-PAGE assay
Gentle SDS-PAGE was used to assess formation of stable complexes between peptides and HLA-DQ8 molecules as previously described (Kim et al., 2013; Sadegh-Nasseri and Germain, 1991). Briefly, 0.5 μM of HLA-DQ8 monomers (provided by NIH tetramer core facility) were treated with thrombin to cleave and remove CLIP peptide. Empty monomers were incubated in the absence or presence of 100 μM of indicated peptides (x-Id, TP-Id, mimotope (R22E), native insulin B:9–23, and h-Id) at 37°C for 72 h in citrate phosphate buffer, pH 5.5 with 1 mM PMSF and 0.025% NaN3. Reactions were neutralized, mixed with equal volumes of SDS-PAGE sample buffer containing 0.1% SDS (final concentration) and placed for 15 min at room temperature and run on 10% PAGE gels and silver-stained using a standard protocol. To assess stability some samples were boiled for 3 min, which resulted in degradation of complexes (data not shown).
Generation of EBV-immortalized DE x1.1 clone
To generate immortalized DE cells, we sorted IgD+ DE cells from freshly isolated PBMCs using a FACSAria II using described strategy (Figure 6A). Sorted cells were seeded at 10, 25, 50 or 100 cells per well of 96-well microplates that had been coated 24 h earlier with irradiated fibroblasts (ATCC® 55-X). Cultures were pulsed with 2.5 μg/mL CpG ODN 2006 (ODN7909) and EBV supernatant stock (ATCC® VR-1492) from B95–8 cells (Caputo and Flytzanis, 1991; Hui-Yuen et al., 2011). Cultures were maintained by replacing half of culture medium with fresh medium every 5 to 7 days. Immortalized cells were visible after 8 days in cultures seeded with 50 or 100 DE cells. We selected on lymphoblastoid cell line (hereafter referred to as x-LCL) for subsequent analysis. In one set of experiments, we sorted single cells from x-LCL and examined for expression of Ig heavy and light chains. In a second set, we used x-LCL to generate x1.1 clone by limiting dilution (0.3 cell/well) as described (Hamad et al., 1994). Cells of the x1.1 clone were used for analysis of BCR and TCR and spontaneous antibody production.
Analysis of x1.1 clone for coexpression of BCR and TCRαβ
We used two approaches to ensure single cell clonality of immmortalized DE cell populations, outgrown monoclonal DE cells were sorted on 96 well microplates using a FACSAria II (BD Biosciences, Bedford, MA) as described above containing RNA catch buffer (Smith et al., 2009).
Cloning and expression of BCR from x1.1 clone and fresh single DE cells
We used the same protocol that was developed by Smith et al. (Smith et al., 2009) for analysis of BCR expression from single cells of the x1.1 clone and freshly sorted single DE cells. Briefly, individual cells were sorted into wells of 96-well PCR plate loaded with catch buffer containing RNase inhibitor to perform RT-PCR using OneStep RT-PCR Kit (QIAGEN). Two primers were utilized to amplify all VH4 gene family members and 8 primers for amplification of genes encoding lamda chain. Cloning PCR of the heavy chain was performed using primers that incorporate the cloning restriction sites and place VDJ heavy chain and constant region genes in frame within the cloning vector (AbVec-hIgG1). Cloning PCR products were purified using Monarch PCR & DNA Cleanup Kit (New England, BioLabs) and visualization as a band of approximately 400 bp in 1.5% agarose gel. Insert and vector were digested with AgeI and SalI and purified as described above. A three-fold molar excess of insert to vector were used to transform DH5α cells. Positive colonies were picked, cultured and plasmid extracted by QIAprep Spin Miniprep Kit (QIAGEN) followed by sequencing using the AbVec primer. The lambda chain was cloned using the same procedure except that insert and vector were digested with AgeI and XhoI and cloned into AbVec- Igλ. Complete sequences of the variable regions were used to identify VDJ usage and CDR3 by IMGT/V-QUEST software.
Expression and analysis of recombinant x-mAbR from single DE cells
Following purification and digestion, amplified cDNAs of the antibody variable genes from single cells were cloned into expression vectors containing human IgG (AbVec-hIgG1) and Igλ (AbVec- Igλ) constant regions (Smith et al., 2009). Briefly, AbVec-hIgG1 containing the heavy- and AbVec- Igλ containing light-chain Ig genes were co-transfected into the HEK293A cell line using polyplus jet-prime transfection (Polypus transfection) and manufacturer’s instruction. Transfected 293A cells were allowed to secrete antibodies in serum-free basal media for 4 to 5 days and mAbR was purified using immobilized protein A columns (Pierce). Antibody expression and purity was verified by SDS-PAGE, and purified antibody concentrations were determined using the EZQ Protein Quantitation Kit (Invitrogen).
Cloning of α and β chains of TCR from the x1.1 clone
The genes for TCRα and TCRβ chains were cloned using a modified version of the method described by Eugster et al. (Eugster et al., 2013). Briefly, total RNA was isolated from cells of the x1.1 clone using RNA extraction kit (Biolabs). cDNA was prepared and mixed with degenerate primers for the α and β chains using OneStep RT-PCR Kit (QIAGEN) and used to amplify the α and β chains by nested-PCR using specific primers for the α chain and β chains, separately. Amplified products were visualized on 1.5% agarose gel and cloned into pGEM-T Easy vector (Promega). DNA was extracted using plasmid extraction kit (QIAGEN) and Sanger-sequenced using the M1 primer (Eugster et al., 2013). Complete sequences of the variable regions were used to identify VDJ usage and CDR3 by the IMGT/V-QUEST software.
Characterization of the x-mAbN produced by x1.1 clone
Cells of x1.1 clones were expanded in complete medium for 3–4 days, washed with PBS and cultured in basal media (Key Resource Table) for five days. Secreted mAbN was detected in supernatants by using SDS-PAGE. Isotype of secreted mAb was determined as IgM using Pro-Detect Rapid Antibody Isotyping Assay human Kit (Thermo Fisher). Antibody concentration was determined using the EZQ Protein Quantitation Kit (Invitrogen).
Tetramer preparation and staining
We made three DQ8 tetramers using biotinylated HLA-DQ monomer provided by (NIH Tetramer Core Facility) using method described by Crawford et al. (Crawford et al., 2011). Briefly, one tetramer is made of HLA-DQ with x-Id peptide (x-Id tet), one complexed with insulin mimotope (mim-tet) and the third one complexed with CLIP (CLIP-tet). Clip was removed using thrombin and empty monomers were loaded with 0.2 mg/mL of either x-Id or Insulin-mimotope peptide. Loading was conducted at 37°C for 72 h in the presence of 2.5 mg/mL (0.25%) n-octyl-β-D-glucopyranoside and 1 mM Pefabloc SC (Sigma-Aldrich). Peptide-loaded HLA-DQ monomers were tetramerized with PE-conjugated streptavidin (eBioscience) at a molar ratio of 1:4, respectively. HLA-DQ/CLIP monomer was tetramerized as control negative in staining. The successful formation of tetramer complexes were verified by gentle SDS-PAGE. Tetramer staining was performed incubating PBMCs with 2 μg /mL HLA class II tetramer for 1 hr at room temperature in FACS buffer. Antibody specific for surface CD4, TCR and CD19 have been used and samples were acquired. The data was analyzed as described above (Dai et al., 2015).
QUANTIFICATION AND STATISTICAL ANALYSIS
Description of experimental replicates and sample sizes are describe in Figures legends. Statistical significance of the results was performed using Prism 6 (GraphPad Software). Analysis was performed using independent samples t test or a paired sample Student’s t test as appropriate. The results were expressed as the mean ± SEM unless stated otherwise. p ≤ 0.05 was considered as statistically significant.
DATA AND SOFTWARE AVAILABILITY
The gene sequencing data of x-mAb (Heavy and Light chains) reported in this study are deposited in the Gene Bank under accession codes IGH-x: MK764540; IGL1-x: MK764541; IGL2-x: MK764542; TCRA-x: MK764543; TCRB-x: MK764544. The RNA-sequencing data presented in this paper are available in the GEO: GSE129112. Immuno-Seq data are available on https://clients.adaptivebiotech.com/pub/ahmed-2019-cell under DOI accession number: https://doi.org/10.21417/RA042019: see (Key Resources Table) for data availability.
Supplementary Material
Highlights.
Dual expressers (DEs) are new lymphocytes that coexpress functional BCR and TCR
A single BCR clonotype (x-clonotype) predominates DEs in T1D subjects
x-clonotype encodes a potent autoantigen with an optimal register for HLA-DQ8
x-mAb secreted by DEs is a potent stimulator of insulin-specific CD4 T cells
ACKNOWLEDGMENTS
We thank participants enrolled in this study and their families for their blood donation. Special thanks to Sarah Longenecker of Johns Hopkins for recruiting and consenting subjects and the NIH Tetramer Core Facility for providing DQ8 monomers. We thank Dr. Katie Boland of Adaptive Biotechnologies (Seattle, WA) for depositing the immuno-seq data to immuneACCESS Data—Adaptive Biotechnologies. We thank Norman Barker of Pathology Photography, Digital Imaging, & Computer Graphics at Johns Hopkins Medicine for production of gel images and Jinglong Ding and Michelle Beaucher from ThermoFisher Scientific for imaging DE clones using the EVOS M7000 Imagine System. Special thanks to Dr. Joseph Margolick, Director of Flor Cytometry Core at the Bloomberg School of Public Health. This work was supported by NIH, United States, grant R0 AI099027 (A.R.A.H.) and the Norman Raab Foundation, United States. A.G. is supported by NIH, United States T32HD044355.
Footnotes
DECLARATION OF INTERESTS
The authors declare no conflict of interests.
SUPPLEMENTAL INFORMATION
Supplemental Information can be found online at https://doi.org/10.1016/j.cell.2019.05.007.
REFERENCES
- Aas-Hanssen K, Funderud A, Thompson KM, Bogen B, and Munthe LA (2014). Idiotype-specific Th cells support oligoclonal expansion of anti-dsDNA B cells in mice with lupus. J. Immunol. 193, 2691–2698. [DOI] [PubMed] [Google Scholar]
- Best RB, Zhu X, Shim J, Lopes PE, Mittal J, Feig M, and Mackerell AD Jr. (2012). Optimization of the additive CHARMM all-atom protein force field targeting improved sampling of the backbone φ, ψ and side-chain χ(1) and χ(2) dihedral angles. J. Chem. Theory Comput. 8, 3257–3273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Canzar S, Neu KE, Tang Q, Wilson PC, and Khan AA (2017). BASIC: BCR assembly from single cells. Bioinformatics 33, 425–427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Caputo JG, and Flytzanis N (1991). Kink-antikink collisions in sine-Gordon and phi4 models: Problems in the variational approach. Phys. Rev. A 44, 6219–6225. [DOI] [PubMed] [Google Scholar]
- Carlson CS, Emerson RO, Sherwood AM, Desmarais C, Chung MW, Parsons JM, Steen MS, LaMadrid-Herrmannsfeldt MA, Williamson DW, Livingston RJ, et al. (2013). Using synthetic templates to design an unbiased multiplex PCR assay. Nat. Commun. 4, 2680. [DOI] [PubMed] [Google Scholar]
- Chang KY, and Unanue ER (2009). Prediction of HLA-DQ8beta cell peptidome using a computational program and its relationship to autoreactive T cells. Int. Immunol. 21, 705–713. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chowell D, Morris LGT, Grigg CM, Weber JK, Samstein RM, Makarov V, Kuo F, Kendall SM, Requena D, Riaz N, et al. (2018). Patient HLA class I genotype influences cancer response to checkpoint blockade immunotherapy. Science 359, 582–587. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crawford F, Stadinski B, Jin N, Michels A, Nakayama M, Pratt P, Marrack P, Eisenbarth G, and Kappler JW (2011). Specificity and detection of insulin-reactive CD4+ T cells in type 1 diabetes in the nonobese diabetic (NOD) mouse. Proc. Natl. Acad. Sci. USA 108, 16729–16734. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dai H, Rahman A, Saxena A, Jaiswal AK, Mohamood A, Ramirez L, Noel S, Rabb H, Jie C, and Hamad AR (2015). Syndecan-1 identifies and controls the frequency of IL-17-producing naїve natural killer T (NKT17) cells in mice. Eur. J. Immunol. 45, 3045–3051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DeWitt WS, Lindau P, Snyder TM, Sherwood AM, Vignali M, Carlson CS, Greenberg PD, Duerkopp N, Emerson RO, and Robins HS (2016). A Public Database of Memory and Naive B-Cell Receptor Sequences. PLoS ONE 11, e0160853. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eugster A, Lindner A, Heninger AK, Wilhelm C, Dietz S, Catani M, Ziegler AG, and Bonifacio E (2013). Measuring T cell receptor and T cell gene expression diversity in antigen-responsive human CD4+ T cells. J. Immunol. Methods 400–401, 13–22. [DOI] [PubMed] [Google Scholar]
- Hamad AR, Herman A, Marrack P, and Kappler JW (1994). Monoclonal antibodies defining functional sites on the toxin superantigen staphylococcal enterotoxin B. J. Exp. Med. 180, 615–621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hestvik AL, Vartdal F, Fredriksen AB, Thompson KM, Kvale EO, Skorstad G, Bogen B, and Holmoy T (2007). T cells from multiple sclerosis patients recognize multiple epitopes on Self-IgG. Scand. J. Immunol. 66, 393–401. [DOI] [PubMed] [Google Scholar]
- Holm S (1979). Simple Sequentially Rejective Multiple Test Procedure. Scand. J. Stat. 6, 65–70. [Google Scholar]
- Hölzemer A, Thobakgale CF, Jimenez Cruz CA, Garcia-Beltran WF, Carlson JM, van Teijlingen NH, Mann JK, Jaggernath M, Kang SG, Körner C, et al. (2015). Selection of an HLA-C*03:04-Restricted HIV-1 p24 Gag Sequence Variant Is Associated with Viral Escape from KIR2DL3+ Natural Killer Cells: Data from an Observational Cohort in South Africa. PLoS Med. 12, e1001900. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hui-Yuen J, McAllister S, Koganti S, Hill E, and Bhaduri-McIntosh S (2011). Establishment of Epstein-Barr virus growth-transformed lymphoblastoid cell lines. J. Vis. Exp. (57) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Joglekar AV, Liu Z, Weber JK, Ouyang Y, Jeppson JD, Noh WJ, Lamothe-Molina PA, Chen H, Kang SG, Bethune MT, et al. (2018). T cell receptors for the HIV KK10 epitope from patients with differential immunologic control are functionally indistinguishable. Proc. Natl. Acad. Sci. USA 115, 1877–1882. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khodadoust MS, Olsson N, Wagar LE, Haabeth OA, Chen B, Swaminathan K, Rawson K, Liu CL, Steiner D, Lund P, et al. (2017). Antigen presentation profiling reveals recognition of lymphoma immunoglobulin neoantigens. Nature 543, 723–727. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim A, Ishizuka I, Hartman I, Poluektov Y, Narayan K, and Sadegh-Nasseri S (2013). Studying MHC class II peptide loading and editing in vitro. Methods Mol. Biol. 960, 447–459. [DOI] [PubMed] [Google Scholar]
- LeMaoult J, Caumartin J, and Carosella ED (2007). Exchanges of membrane patches (trogocytosis) split theoretical and actual functions of immune cells. Hum. Immunol. 68, 240–243. [DOI] [PubMed] [Google Scholar]
- Luisiri P, Lee YJ, Eisfelder BJ, and Clark MR (1996). Cooperativity and segregation of function within the Ig-alpha/beta heterodimer of the B cell antigen receptor complex. J. Biol. Chem. 271, 5158–5163. [DOI] [PubMed] [Google Scholar]
- Lun AT, McCarthy DJ, and Marioni JC (2016). A step-by-step workflow for low-level analysis of single-cell RNA-seq data with Bioconductor. F1000Res. 5, 2122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mansfield JM (1994). T-cell responses to the trypanosome variant surface glycoprotein: a new paradigm? Parasitol. Today (Regul. Ed.) 10, 267–270. [DOI] [PubMed] [Google Scholar]
- Martina MN, Noel S, Saxena A, Bandapalle S, Majithia R, Jie C, Arend LJ, Allaf ME, Rabb H, and Hamad AR (2016). Double-Negative αβ T Cells Are Early Responders to AKI and Are Found in Human Kidney. J. Am. Soc. Nephrol. 27, 1113–1123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Michel ML, Pang DJ, Haque SF, Potocnik AJ, Pennington DJ, and Hayday AC (2012). Interleukin 7 (IL-7) selectively promotes mouse and human IL-17-producing γδ cells. Proc. Natl. Acad. Sci. USA 109, 17549–17554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakayama M, McDaniel K, Fitzgerald-Miller L, Kiekhaefer C, Snell-Bergeon JK, Davidson HW, Rewers M, Yu L, Gottlieb P, Kappler JW, and Michels A (2015). Regulatory vs. inflammatory cytokine T-cell responses to mutated insulin peptides in healthy and type 1 diabetic subjects. Proc. Natl. Acad. Sci. USA 112, 4429–4434. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Okada T, Miller MJ, Parker I, Krummel MF, Neighbors M, Hartley SB, O’Garra A, Cahalan MD, and Cyster JG (2005). Antigen-engaged B cells undergo chemotaxis toward the T zone and form motile conjugates with helper T cells. PLoS Biol. 3, e150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pavlidis P, and Noble WS (2001). Analysis of strain and regional variation in gene expression in mouse brain. Genome Biol. 2, RESEARCH0042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Picelli S, Faridani OR, Björklund AK, Winberg G, Sagasser S, and Sandberg R (2014). Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181. [DOI] [PubMed] [Google Scholar]
- Reth M (1992). Antigen receptors on B lymphocytes. Annu. Rev. Immunol. 10, 97–121. [DOI] [PubMed] [Google Scholar]
- Rizzetto S, Eltahla AA, Lin P, Bull R, Lloyd AR, Ho JWK, Venturi V, and Luciani F (2017). Impact of sequencing depth and read length on single cell RNA sequencing data of T cells. Sci. Rep. 7, 12781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robins HS, Campregher PV, Srivastava SK, Wacher A, Turtle CJ, Kahsai O, Riddell SR, Warren EH, and Carlson CS (2009). Comprehensive assessment of T-cell receptor beta-chain diversity in alphabeta T cells. Blood 114, 4099–4107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sadegh-Nasseri S, and Germain RN (1991). A role for peptide in determining MHC class II structure. Nature 353, 167–170. [DOI] [PubMed] [Google Scholar]
- Saxena A, Yagita H, Donner TW, and Hamad ARA (2017). Expansion of FasL-Expressing CD5+ B Cells in Type 1 Diabetes Patients. Front. Immunol. 8, 402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seay HR, Yusko E, Rothweiler SJ, Zhang L, Posgai AL, Campbell-Thompson M, Vignali M, Emerson RO, Kaddis JS, Ko D, et al. (2016). Tissue distribution and clonal diversity of the T and B cell repertoire in type 1 diabetes. JCI Insight 1, e88242. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sewell AK (2012). Why must T cells be cross-reactive? Nat. Rev. Immunol. 12, 669–677. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharp KA (2012). Statistical thermodynamics of binding and molecular recognition models. In Protein-Ligand Interactions, Mannhold R, Kubinyi H, and Folkers G, eds.. [Google Scholar]
- Smith K, Garman L, Wrammert J, Zheng NY, Capra JD, Ahmed R, and Wilson PC (2009). Rapid generation of fully human monoclonal antibodies specific to a vaccinating antigen. Nat. Protoc. 4, 372–384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith MJ, Packard TA, O’Neill SK, Henry Dunand CJ, Huang M, Fitzgerald-Miller L, Stowell D, Hinman RM, Wilson PC, Gottlieb PA, and Cambier JC (2015). Loss of anergic B cells in prediabetic and new-onset type 1 diabetic patients. Diabetes 64, 1703–1712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tirosh I, and Suvà ML (2018). Dissecting human gliomas by single-cell RNA sequencing. Neuro-oncol. 20, 37–43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tirosh I, Venteicher AS, Hebert C, Escalante LE, Patel AP, Yizhak K, Fisher JM, Rodman C, Mount C, Filbin MG, et al. (2016). Single-cell RNA-seq supports a developmental hierarchy in human oligodendroglioma. Nature 539, 309–313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trück J, Ramasamy MN, Galson JD, Rance R, Parkhill J, Lunter G, Pollard AJ, and Kelly DF (2015). Identification of antigen-specific B cell receptor sequences using public repertoire analysis. J. Immunol. 194, 252–261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Venturi V, Price DA, Douek DC, and Davenport MP (2008). The molecular basis for public T-cell responses? Nat. Rev. Immunol. 8, 231–238. [DOI] [PubMed] [Google Scholar]
- Wang J, Sohn H, Sun G, Milner JD, and Pierce SK (2014). The autoinhibitory C-terminal SH2 domain of phospholipase C-γ2 stabilizes B cell receptor signalosome assembly. Sci. Signal. 7, ra89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Y, Sosinowski T, Novikov A, Crawford F, Neau DB, Yang J, Kwok WW, Marrack P, Kappler JW, and Dai S (2018). C-terminal modification of the insulin B:11–23 peptide creates superagonists in mouse and human type 1 diabetes. Proc. Natl. Acad. Sci. USA 115, 162–167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wardemann H, Yurasov S, Schaefer A, Young JW, Meffre E, and Nussenzweig MC (2003). Predominant autoantibody production by early human B cell precursors. Science 301, 1374–1377. [DOI] [PubMed] [Google Scholar]
- Watson CT, Steinberg KM, Huddleston J, Warren RL, Malig M, Schein J, Willsey AJ, Joy JB, Scott JK, Graves TA, et al. (2013). Complete haplotype sequence of the human immunoglobulin heavy-chain variable, diversity, and joining genes and characterization of allelic and copy-number variation. Am. J. Hum. Genet. 92, 530–546. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu AR, Neff NF, Kalisky T, Dalerba P, Treutlein B, Rothenberg ME, Mburu FM, Mantalas GL, Sim S, Clarke MF, and Quake SR (2014). Quantitative assessment of single-cell RNA-sequencing methods. Nat. Methods 11, 41–46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xia Z, Chen H, Kang SG, Huynh T, Fang JW, Lamothe PA, Walker BD, and Zhou R (2014). The complex and specific pMHC interactions with diverse HIV-1 TCR clonotypes reveal a structural basis for alterations in CTL function. Sci. Rep. 4, 4087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xiao Z, Mohamood AS, Uddin S, Gutfreund R, Nakata C, Marshall A, Kimura H, Caturegli P, Womer KL, Huang Y, et al. (2011). Inhibition of Fas ligand in NOD mice unmasks a protective role for IL-10 against insulitis development. Am. J. Pathol. 179, 725–732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yoneshiro T, Matsushita M, Hibi M, Tone H, Takeshita M, Yasunaga K, Katsuragi Y, Kameya T, Sugie H, and Saito M (2017). Tea catechin and caffeine activate brown adipose tissue and increase cold-induced thermogenic capacity in humans. Am. J. Clin. Nutr. 105, 873–881. [DOI] [PubMed] [Google Scholar]
- Yu LM, and Chang TW (1992). Human mb-1 gene: complete cDNA sequence and its expression in B cells bearing membrane Ig of various isotypes. J. Immunol. 148, 633–637. [PubMed] [Google Scholar]
- Ziegenhain C, Vieth B, Parekh S, Reinius B, Guillaumet-Adkins A, Smets M, Leonhardt H, Heyn H, Hellmann I, and Enard W (2017). Comparative Analysis of Single-Cell RNA Sequencing Methods. Mol Cell 65, 631–643 e634. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The gene sequencing data of x-mAb (Heavy and Light chains) reported in this study are deposited in the Gene Bank under accession codes IGH-x: MK764540; IGL1-x: MK764541; IGL2-x: MK764542; TCRA-x: MK764543; TCRB-x: MK764544. The RNA-sequencing data presented in this paper are available in the GEO: GSE129112. Immuno-Seq data are available on https://clients.adaptivebiotech.com/pub/ahmed-2019-cell under DOI accession number: https://doi.org/10.21417/RA042019: see (Key Resources Table) for data availability.