Abstract
The COVID-19 outbreak has been causing a global health crisis since December 2019. Due to this virus declared by the World Health Organization as a pandemic, the health authorities of the countries are constantly trying to reduce the spread rate of the virus by emphasizing the rules of masks, social distance, and hygiene. COVID-19 is highly contagious and spreads rapidly globally and early detection is of paramount importance. Any technological tool that can provide rapid detection of COVID-19 infection with high accuracy can be very useful to medical professionals. The disease findings on COVID-19 images, such as computed tomography (CT) and X-rays, are similar to other lung infections, making it difficult for medical professionals to distinguish COVID-19. Therefore, computer-aided diagnostic solutions are being developed to facilitate the identification of positive COVID-19 cases. The method currently used as a gold standard in detecting the virus is the Reverse Transcription Polymerase Chain Reaction (RT-PCR) test. Due to the high false-negative rate of this test and the delays in the test results, alternative solutions are sought. This study was conducted to investigate the contribution of machine learning and image processing to the rapid and accurate detection of COVID-19 from two of the most widely used different medical imaging modes, chest X-ray and CT images. The main purpose of this study is to support early diagnosis and treatment to end the coronavirus epidemic as soon as possible. One of the primary aims of the study is to provide support to medical professionals who are most worn out and working under intense stress during COVID-19 through smart learning methods and image classification models. The proposed approach was applied to three different public COVID-19 data sets and consists of five basic steps: data set acquisition, pre-processing, feature extraction, dimension reduction, and classification stages. Each stage has its sub-operations. The proposed model performs in considerable levels of COVID-19 detection for dataset-1 (CT), dataset-2 (X-ray) and dataset-3 (CT) with the accuracy of 89.41%, 99.02%, 98.11%, respectively. On the other hand, in the X-ray data set, an accuracy of 85.96% was obtained for COVID-19 (+), COVID-19 (-), and those with Pneumonia but not COVID-19 classes. As a result of the study, it has been shown that COVID-19 can be detected with a high success rate in about less than one minute with image processing and classical learning methods. In the light of the findings, it is possible to say that the proposed system will help radiologists in their decisions, will be useful in the early diagnosis of the virus, and can distinguish pneumonia caused by the COVID-19 virus from the pneumonia of other diseases.
Keywords: COVID-19, CAD, Machine learning, X-ray, CT
1. Introduction
Coronavirus has spread very rapidly around the world since December 2019. The World Health Organization (WHO) stated on January 30, 2020, that COVID-19 caused a pandemic. According to WHO (until 17 December 2020), 72,196,732 COVID-19 cases were detected worldwide. 1.630.521 of these cases resulted in death. The vast majority of cases occurred in the Americas (30,925,241 Cases) as of December 17, 2020. America is followed by Europe (22.603.335) and southeast Asia (11.468.106) [1]. The most prominent symptoms of COVID-19 are fever and cough. Also, shortness of breath, muscle or body aches, headache, loss of taste or smell, sore throat, and diarrhea are other common symptoms [2]. As you can see, this virus can manifest itself with many symptoms. And the symptoms it shows are among the common effects that can occur in daily life due to other diseases. This situation makes it difficult to distinguish coronavirus from other diseases. A medical professional takes the following steps to diagnose COVID 19;
-
1.
It is questioned whether the patient is in contact, whether he complies with the mask, distance, and hygiene rules.
-
2.
The patient’s symptoms are analyzed. (Fever, cough, shortness of breath, diarrhea, etc.)
-
3.
RT-PCR test is applied.
-
4.
Radiological imaging is applied. In the early stages, chest radiography is used. In cases where chest radiography is insufficient, CT and ultrasonography are applied.
-
5.
Blood analysis is done and findings are evaluated.
As a result of all these procedures, whether the patient is COVID-19 is determined by the medical specialist. It is a real-time reverse transcriptase–polymerase chain reaction (RT-PCR) test taken from the SARS-CoV-2 throat swab that causes COVID-19 and is believed to be highly specific. However, this test has low sensitivity, especially in the early stages of the disease. The sensitivity rate of this test is around 60%–70% according to the studies [3], [4], [5]. This means that only 60 of the 100 positive COVID-19 patients who had a PCR test can produce true-positive results. For this reason, the disease needs to be supported by different methods such as blood analysis and medical imaging. At this stage, using radiological imaging methods together with computer-aided systems will greatly benefit medical professionals. Computed tomography (CT) and X-ray imaging techniques play a vital role in the early diagnosis and treatment of this disease [6], [7], [8], [9]. Even if false-negative results are obtained due to the low RT-PCR sensitivity of 60%–70%, it can be detected by the radiological images of the patients [10], [11]. In some studies, it has been stated that CT is a sensitive method for detecting COVID-19 pneumonia and can be considered as an auxiliary screening tool for RT-PCR [12]. It should be made clear here that radiological imaging is not a COVID-19 test. It would be more correct to consider radiological imaging as an auxiliary element to the testing process. In other words, PCR test results with low accuracy are supported by radiological imaging. Blood analysis similarly supports the PCR test results with the findings it gives.
CT findings are observed for a long time after the onset of symptoms, and patients usually have normal CT in the first 0–2 days [6]. In a study on lung CT, it was stated that the most important lung disease symptoms of patients with COVID-19 pneumonia were observed ten days after the onset of the disease [13].
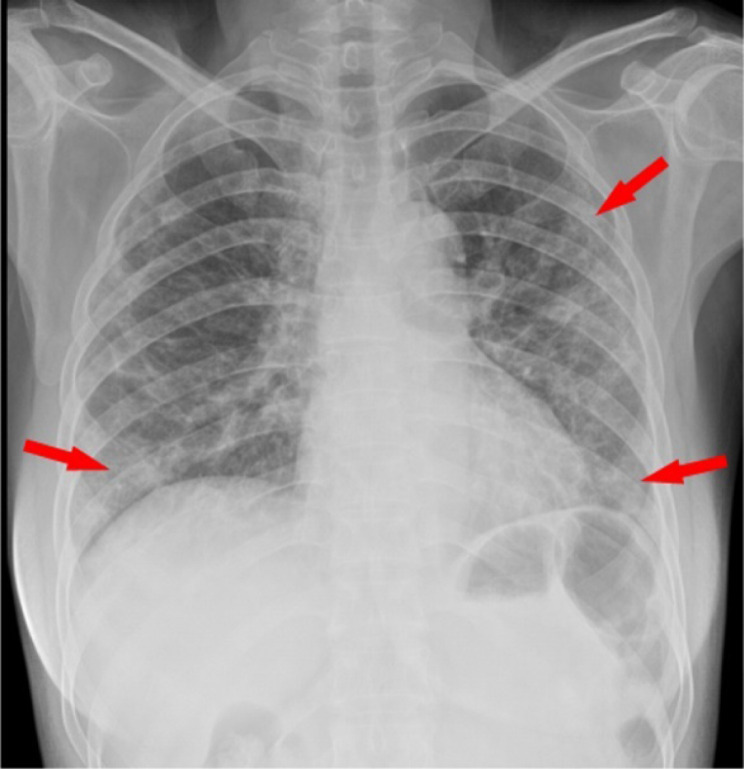
One of the most important findings of the disease is pneumonia in the lungs. For these reasons, chest radiography, computed tomography, or ultrasonography are requested from patients. The first preferred and used method is chest radiography. However, the sensitivity of chest radiography is low (30%–60%) [14]. Fig. 1 shows the image of a COVID-19 (+) patient included in the X-ray data set we used in our study. One of the most obvious signs of COVID-19 is the ground-glass opacities (GGO). In Fig. 1, the image of GGO is seen.
Fig. 1.
GGO on the Chest X-ray Image (dataset 2).
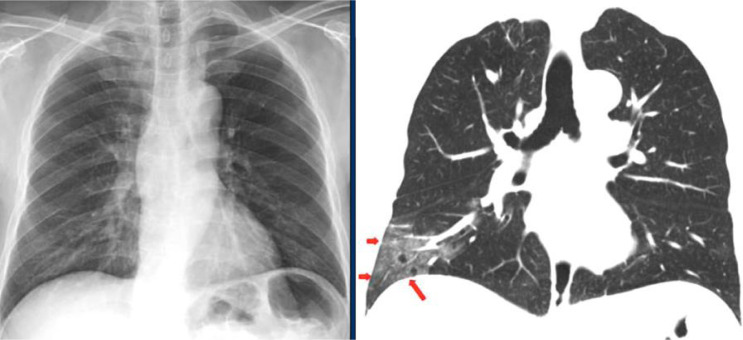
Chest radiography can be misleading in the early stages of the COVID-19 [15]. Fig. 2 shows a comparison of a chest radiograph and a CT image. GGO in the right lower lobe, indicated by red arrows on CT, is not visible on the chest radiograph taken one hour before the CT study [15].
Fig. 2.
Comparison of X-ray and CT images in the early stage of COVID-19 [15].
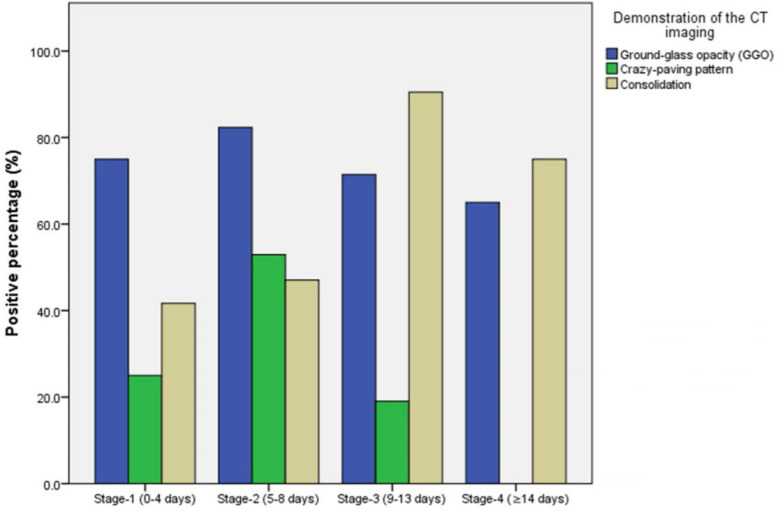
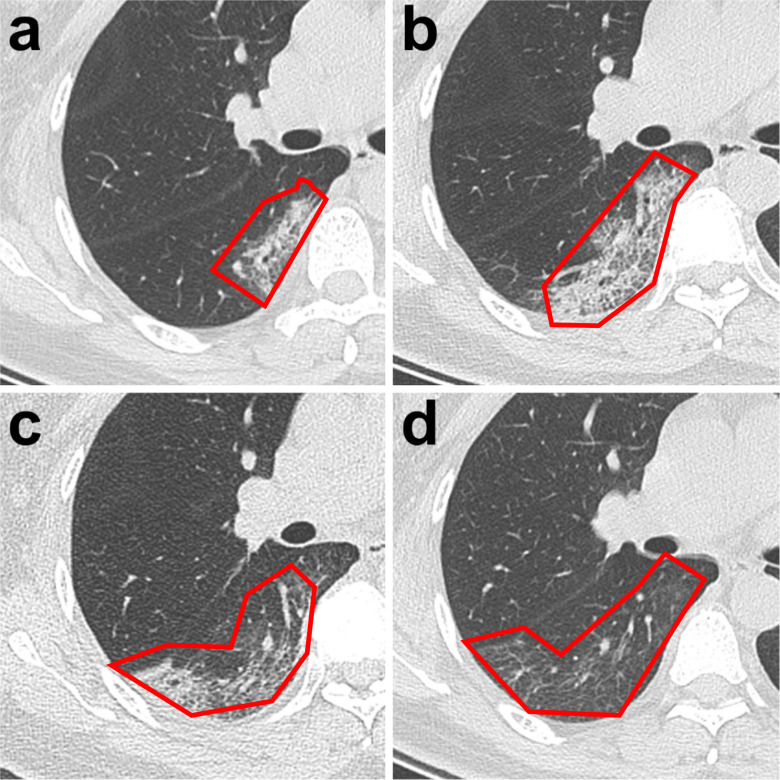
Another CT finding is irregular consolidation [16]. This finding is more common in CT scans taken after the onset of symptoms. One of the most common findings in COVID-19 cases on CT images is the crazy-paving pattern [13]. Fig. 3 shows the percentages of the findings of GGO, consolidation, and crazy-paving pattern on CT images between 0–14 days. It can be seen from Fig. 3 that GGO is at the highest rate in the early stages of the disease and consolidation is the most prominent finding between 9–14 days of the disease. Fig. 4 shows the CT images of a 47-year-old female patient diagnosed with COVID-19 on different days between 0–14 days. It can be seen from Fig. 4 that GGO is intense in the early stages of the disease, the crazy-paving pattern and consolidation increase in the following days, and the symptoms gradually disappear.
Fig. 3.
Changes in the proportions of patients with GGO, crazy-paving pattern, and consolidation [13].
Fig. 4.
CT findings of COVID-19 in a 47-year-old female patient (a) A small region of GGO with partial consolidation (day 3) (b) Enlarged region of GGO with crazy-paving pattern with partial consolidation (day 7,) (c) A new area of consolidation with a small GGO (day 11) (d) the decreased GGO [13].
Looking at Fig. 4, it is possible to see that the COVID-19 virus has spread rapidly to most of the lungs within a few days. The only solution to this situation is early diagnosis and treatment. This study aims to create a computer-aided method that can support the early diagnosis and treatment of the COVID-19 virus. The use of computer-aided systems in the detection and follow-up of this disease will accelerate the diagnosis and treatment. In the last 10 years, machine learning methods based on computer-based algorithms have been used frequently in automatic diagnosis processes in the medical field. Computer-aided automatic diagnosis systems help clinicians in their decisions and shorten the diagnosis time. In particular, machine learning methods are used in many areas such as breast cancer detection [17], [18], brain tumor detection [19], heart disease [20]. Since the spread of the COVID-19 virus is very fast, the diagnosis process should be done very quickly. However, the number of experts on the subject is also limited. The rate of spread of the disease has caused the health system to collapse in certain countries [21], [22]. Therefore, it can be contributed to the solution of these problems with ML and GI methods. It can be diagnosed in the shortest time with high accuracy rates. And this can be done in less than a minute.
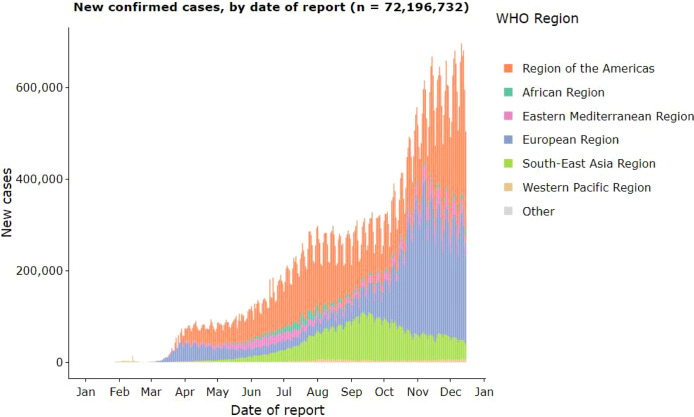
The rapid rise of the COVID-19 outbreak, the limited number of radiologists, the difficulty of providing specialist clinicians to each hospital, the insufficient number of available RT-PCR test kits, test costs, and the waiting time of test results show how important it is to use AI approaches. Fig. 5 shows how regions around the world are affected by COVID-19. When the figure is examined, it is seen that there is a continuous increase in the number of 12-month cases and the disease is most intense in America. From the graph, it is possible to see that the number of cases, which was 400,000 in October in the American region, reached 600,000 in November. This situation shows how high the spread rate of the virus is and reveals the importance of our work.
Fig. 5.
Case change (as of December 17, 2020) [1] .
Table 1 shows the countries with the highest number of cases and deaths since December 2019. With more than 16 million cases and 298,594 deaths, the United States is seen as the place most affected by COVID-19. Another remarkable point in the table is that there are approximately 9.9 million cases in India. In Brazil, this number is around 6.9 million. Although India has 3 million more cases than Brazil, the death rate is lower. This stands out as an issue that needs to be addressed. From here, it can be deduced that India follows a more effective way in diagnosis and treatment or Brazil is not able to manage the process well.
Table 1.
Highest case and death rates (as of 17 December 2020, 2020) [1].
| Country/Area/Territory | Total cases | Total deaths | |
|---|---|---|---|
| 1 | United States of America | 16,245,376 | 298,594 |
| 2 | India | 9,932,547 | 144,096 |
| 3 | Brazil | 6,927,145 | 181,835 |
| 4 | Russian Federation | 2,734,454 | 48,564 |
| 5 | France | 2,350,207 | 58,700 |
| 6 | United Kingdom of Great Britain and Northern Ireland | 1,888,120 | 64,908 |
| 7 | Italy | 1,870,576 | 65,857 |
| 8 | Spain | 1,762,212 | 48,401 |
| 9 | Argentina | 1,503,222 | 41,041 |
| 10 | Colombia | 1,434,516 | 39,195 |
As can be seen from both Fig. 5 and Table 1, the spread rate of the virus is very high. And the death rate is around 2% of the total number of cases. This is also a considerable rate. For this reason, it is deemed necessary to implement all processes that allow early diagnosis and treatment. The main motivation of this study is to produce an effective solution with high accuracy for different imaging techniques. It is possible to summarize the contributions of the study to the literature as follows;
-
1.
A new approach has been proposed that can enable early diagnosis and treatment of COVID-19 with machine learning and image processing methods.
-
2.
Successful results have been obtained with the same approach in both CT and X-ray imaging modes.
-
3.
One of the novelty aspects of the study is that it applies to different data sets. The results obtained for three data sets with different characteristics show that the study is generalizable. In other words, the proposed approach is not data set dependent. The proposed approach can also be applied to a different data set.
-
4.
To create a decision support system that can support medical professionals in their studies and help them in their decisions.
-
5.
The most dangerous aspect of the COVID-19 virus is that it can be transmitted from person to person very quickly and after it is transmitted, it can show its effect on our body very quickly. (See Fig. 4). Therefore, a study that will support early diagnosis with the help of CT and X-ray images will contribute significantly to the solution of the problem. With this study, it has been shown that classical learning methods can produce as many successful results as deep learning methods.
-
6.
While most of the studies in the literature use very small data sets, 3 different data sets with different sizes and qualities are used in this study. Also, not only binary classification, but also multi-class classification was carried out.
The rest of this paper is organized as follows. Section 2 introduces the related works while Section 3 is about materials and methods used for the proposed system. Section 4 presents the experimental results of the study and Section 5 presents discussions. Finally, the conclusion and the suggestions for future work are presented in Section 6.
2. Related works
With the COVID-19 virus affecting the world since December 2019, academic studies have started. In the past year, many studies have been carried out especially on the detection of COVID-19 with computer-aided systems. Most of the studies have been carried out using deep learning approaches that have become popular in the last few years. The first of these studies is Hemdan et al. [23] is the study where they created a deep learning model called COVIDX-Net using X-ray Images to diagnose COVID-19. In the study, they analyzed using different deep learning models such as VGG19, DenseNet201, ResNetV2, InceptionV3, InceptionResNetV2, Xception and MobileNetV2 comparatively. The results obtained showed that VGG19 and DenseNet201 models achieved the best performance with 90% success.
In the study of Barstugan et al. where classical learning methods are preferred, a new approach has been proposed for the classification of COVID-19 [24]. CT images were used in the study. They have extracted features with the help of patches of different sizes. They obtained an accuracy rate of 98.77% by classifying the obtained features with an SVM classifier. Wang and Wong designed a special deep learning-based framework called COVID-Net [25]. They applied the 1 1 convolutional deep learning method to the data sets consisting of chest X-ray images and achieved the success of 83.5%. In another study using convolutional neural networks, Maghdid et al. proposed a system for automatic diagnosis of COVID-19 pneumonia. They achieved an accuracy rate of 94.00% by making changes to existing architectures in their work [26]. Ghoshal and Tucker conducted another study using the convolutional neural network of deep learning to detect COVID-19 from X-ray images [27]. They resized the X-ray images they used in their work to 512 512. The approach they recommended achieved 92.86% accuracy.
Hall et al. [28] proposed a model with a 10-fold cross-validation strategy on X-ray images using the VGG16 deep learning model. In their studies, they resized the data set numbered [29] and used it (224 224). The method they recommended achieved an accuracy rate of 96.1%. Farooq and Hafeez [30] used a pre-trained ResNet-50 architecture for the diagnosis of COVID-19 pneumonia. To increase the success of generalization, they used the data set after preprocessing it with vertical flipping, random rotation, and different data enlargement methods. In this study, the success of COVID-19 classification was 96.23%. In the study conducted by Abbas et al. [31], the situation of being positive or negative from the COVID-19 X-ray images was evaluated. The deep transfer learning method was used in the study. The principal component analysis (PCA) method was used to reduce the high number of features in the study. The success of the system was measured with the validation, sensitivity, and specificity metrics, and success rates of 95.12, 97.91, and 91.87%, respectively, were obtained. Singh et al. In their study, [32] classified patients infected with COVID-19 as infected or not, using CT images based on multi-objective differential evolution (MODE) CNN. The sensitivity value of the system is at 95% levels. Also, the sensitivity value obtained with the system was compared with the CNN, ANFIS, and ANN methods and it was seen that even good results were obtained. Kassani et al. used both X-ray and CT images in their study. In the study, modeling has been carried out with many deep learning methods. According to the results, a 99% success rate was achieved as a result of obtaining the features of DenseNet121 architecture and training with the Bagging Tree classifier.
In addition to the above studies, the details of which are given, Table 2 shows the data of more studies. When the table is examined, it is possible to see the methods used in the studies, the display formats, the size and type of the data sets. Of course, studies on COVID-19 are not limited to this. Apart from the studies included in this study, there are also many successful studies.
When Table 2 is examined, it is seen that most of the studies were carried out with deep learning methods. Similarly, it is seen that the studies in the table work on a single data set. From this point of view, it would not be wrong to say that our study has operated on three different data sets and added a novelty to existing studies. Besides, the successful results obtained in both CT and X-ray imaging modes, despite their different structures, can be considered as the new aspects of our study.
Table 2.
Related works on COVID-19 in the literature.
| Author(s) | Method | Imaging type | Size of data |
|---|---|---|---|
| Ozturk et al. [33] | DarkCovidNet | Chest X-ray | 125 COVID-19 (+), 500 No Findings 500 Pneumonia |
| Hemdan et al. [23] | COVIDX-Net | Chest X-ray | 25 COVID-19 (+), 25 COVID-19 (−) |
| Barstugan et al. [24] | DWT + SVM | CT | 53 COVID-19 (+), 97 COVID-19 (−) |
| Wang et al. [25] | COVID-Net | Chest X-ray | 358 COVID-19 (+), 8,066 no pneumonia, 5,538 non-COVID19 pneumonia. |
| Maghdid et al. [26] | Deep Transfer Learning | X-ray and CT | 85 X-ray and 203 CT COVID-19 (+), 85 X-ray and 153 CT COVID-19 (−) |
| Ghoshal et al. [27] | Dropweights based Bayesian Convolutional Neural Networks (BCNN) | X-ray | Normal: 1583, Bacterial Pneumonia: 2786, non-COVID-19 Viral Pneumonia: 1504, COVID-19: 68 |
| Abbas et al. [31] | Deep Transfer Learning, PCA | X-ray | 116 X-ray Covid-19 (+), 80 X-ray Covid-19 (−) |
| Farooq and Hafeez [30] | ResNet-50 | X-ray | 660 patients with nonCOVID-19 viral pneumonia cases, 68 COVID-19 radiographs 1203 patients with negative pneumonia |
| Singh et al. [32] | Multi-Objective Differential Evolution Based CNN | CT | |
| Soares et al. [34] | eXplainable Deep Learning classification approach | CT | 1252 COVID-19 (+), 1230 COVID-19 (−) |
| Yang et al. [35] | Multi-Task Learning and Contrastive Self-Supervised Learning | CT | 349 COVID-19 (+), 397 COVID-19 (−) |
| Wei et al. [36] | 3D ResNet-18 | CT | 305 COVID-19 (+), 872 Community Acquired Pneumonia (CAP), 1498 Non-pneumonia |
| Hu et al. [37] | Weakly Supervised Multi-scale Learning Framework | CT | 150 Covid-19 (+), 300 community-acquired pneumonia (CAP) and non-pneumonia (NP) |
| Wu et al. [38] | Deep Learning Based | CT | 400 COVID-19 (+) Cases, 350 COVID-19 (−) Cases |
| Sun et al. [39] | Adaptive Feature Selection guided Deep Forest | CT | 1495 Covid-19 (+), 1027 community-acquired pneumonia (CAP) |
| Jaiswal et al. [40] | DenseNet201 based deep transfer learning | CT | 1262 COVID-19 (+) Cases, 1230 COVID-19 (−) Cases |
| Abraham et al. [41] | Multi-CNN and Bayesnet Classifier | X-ray | 453 COVID-19 (+), 497 COVID-19 (−) |
| Altan and Karasu [42] | 2D curvelet transform, chaotic salp swarm algorithm, and deep learning technique | X-ray | 263 COVID-19 (+), 1609 COVID-19 (−), 1614 viral pneumonia |
| Nour et al. [43] | Deep Features and Bayesian Optimization | X-ray | 219 COVID-19 (+), 1341 COVID-19 (−), 1345 viral pneumonia |
| Kassani et al. [44] | MobileNet, DenseNet, Xception, ResNet, InceptionV3, InceptionResNetV2, VGGNet, NASNet | X-ray and CT | 117 X-ray and 20 CT images COVID-19 (+), 117 X-ray and 20 CT images COVID-19 (−) |
| Ardakani et al. [45] | K-Nearest Neighbor, Naïve Bayes, Support Vector Machine (SVM), and Ensemble | CT | 306 COVID-19 (+), 306 COVID-19 (−) |
| Zhou et al. [46] | Ensemble Deep Learning Model | CT | 500 COVID-19 (+), 500 COVID-19 (−) |
| Gupta et al. [47] | Integrated Stacking InstaCovNet-19 model | X-ray | 361 COVID-19 (+), 365 Normal (−), 362 Pneumonia |
| Aslan et al. [48] | CNN-based transfer learning–BiLSTM | X-ray | 219 COVID-19 (+), 1341 COVID-19 (−), 1345 viral pneumonia |
3. Materials and methods
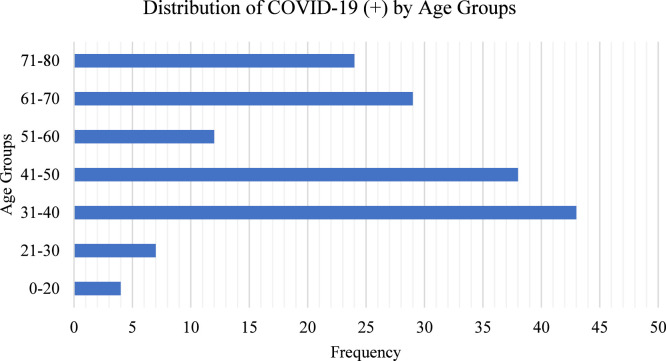
Three different data sets were used in our study. Our first dataset used in our study consists of CT images [35]. The data set includes 349 Covid-19 (+) images of 216 patients, 397 Covid-19 (-) images. While 169 people have age information, 137 people have gender information. Positive numbers according to age groups are seen in Fig. 6. While 37% of those with COVID-19 (+) are female, 63% are male.
Fig. 6.
Age distributions of Data Set-1.
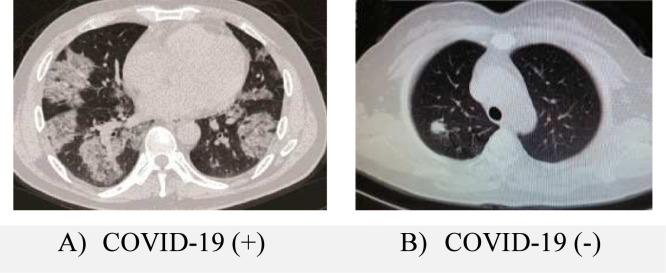
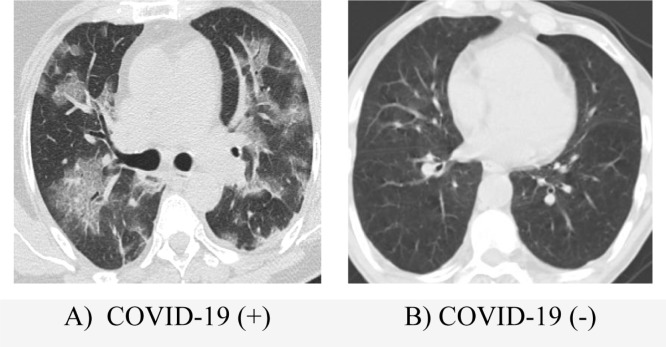
An example of positive and negative COVID-19 images in the data set is shown in Fig. 7.
Fig. 7.
Sample images from data set-1.
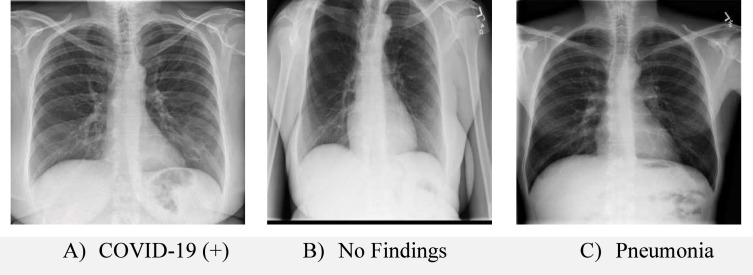
Another data set consists of X-ray images. The data set includes 125 images that are positive for COVID-19. 43 of these images belong to females and 82 of them belong to males. Besides, there are 500 clean lung X-ray images without any findings and 500 images with pneumonia but not COVID. This data set was created by combining images from different data sets [29], [49] in study number [33]. As stated in study number [33], there is no detailed information about the data set. In the data set, 26 of the positive ones have age information and their average is 55. Sample images of this data set are shown in Fig. 8.
Fig. 8.
Sample images from data set-2.
Our third and final data set is a data set consisting of CT images, which has considerably more samples than our other two data sets [34]. This dataset created in study number [34] consists of a public COVID-19 CT scan dataset that includes 1252 CT scans with COVID-19 (+) and 1230 CT scans with COVID-19 (-). The data set was collected from patients in Sao Paulo, Brazil. The images in the data set belong to a total of 60 people, 32 of whom are male and 28 of them are female [34]. While 30 of these people are COVID-19 (+), the remaining 30 are negative. Apart from this, there is no other information about the data set in the study. Sample COVID-19 positive and negative images of the data set are shown in Fig. 9.
Fig. 9.
Sample images from data set-2.
Details of the 3 different data sets we used in our study are shown in Table 3. It is possible to access information on gender distributions in the used data sets, imaging types, number of positive and negative patients, and the countries where the data sets are collected from Table 3.
Table 3.
Information about the data sets used in the study.
| Image type | # of COVID-19 (+) | # of COVID-19 (−) | Gender distribution | Location | |
|---|---|---|---|---|---|
| Dataset-1 | CT | 349 | 397 | %63 Male | China |
| %37 Female | |||||
| Dataset-2 | X-ray | 125 | 500 Clear | %66 Male | Mixed |
| 500 Not COVID-19 but Pneumonia | %34 Female | ||||
| Dataset-3 | CT | 1252 | 1230 | %53 Male | Sao Paulo, Brazil |
| %47 Female | |||||
3.1. Pre-processing
There are three different data sets that we used in our study. The images in each data set were obtained from different sources. For example, while 127 images with COVID-19 (+) in dataset-2 were obtained from the study of Cohen JP [29], 500 healthy images and 500 images with Pneumonia but not COVID-19 were obtained from the dataset created by Wang et al. [49]. For this reason, data sets consisting of images of different sizes obtained from different sources are obtained. Besides, some of the images in the data sets are in jpeg format, while some are in png format. The first process of our preprocessing stage is the resizing process. The dimensions of each of the 3 different data sets we used in our study are different from each other. The sizes of the images in each data set also differ from each other. For this reason, each data set was evaluated within itself and resized. The process here is to normalize the images contained in the data according to their height and width. In the normalization process, first of all, the available images are categorized according to their width and height. Then the frequency of each width and height value is obtained. The final values have been obtained by obtaining the weighted average of the height and width values according to these frequencies. The reason for using the frequency of each image here is to try to get a value closer to those dimensions in the resizing process by increasing the weight if there are more than certain image sizes. The Histogram of the Oriented Gradients (HOG) and Local Binary Patterns (LBP) methods make feature extraction by shifting the image by cell size. For this reason, values that can be divided by cell size were selected in the resizing process of the images. The height and width values obtained after the resizing of the data sets are shown in Table 4 below.
Table 4.
Information about the data sets width and height.
| Width | Height | |
|---|---|---|
| Dataset-1 | 400 | 300 |
| Dataset-2 | 525 | 525 |
| Dataset-3 | 375 | 255 |
The next preprocessing step is to perform the gray level conversion of all images. Such a process has been carried out by the necessity of the images in similar formats to run the computer-aided application. Also, this process did not reduce success and significantly reduced transaction cost and time cost. In the pre-treatment phase, the image sharpening process was applied to increase the clarity of the image and to increase success.
3.2. Feature extraction and dimension reduction
HOG, GLCM, SIFT, and LBP methods were used in the feature extraction phase in our study. HOG and LBP methods provided the most successful results among these four different methods. For this reason, the results of the HOG and LBP methods will be given in the next part of the study.
Histogram of Oriented Gradients (HOG) use the gradient values and orientation angles of pixels in the feature extraction process. Local histograms are obtained with gradient values and orientations, and the image is represented in this way [50]. To determine the HOG features; first, the edges are determined (1). The gradient magnitude and gradient orientation angle is then calculated as seen in 2.
| (1) |
| (2) |
and :Edges are determined by applying horizontal and vertical Sobel filters, G: gradient magnitude, : gradient orientation angle
While extracting the HOG features, the best cell-size was determined to be 15. This means that the window is divided into 15 × 15 cells in x and y directions and descriptors are calculated for each cell. In this work, the UoCTTI variant was used for HOG. This means that HOG computes the four-dimensional texture energy feature as well as directed and undirected gradients but reduces the result to 31 dimensions [51]. As an example, for 400 300 images using cell size 15, the size of the HOG features is 26 20 31 (16120 size feature vector for an image).
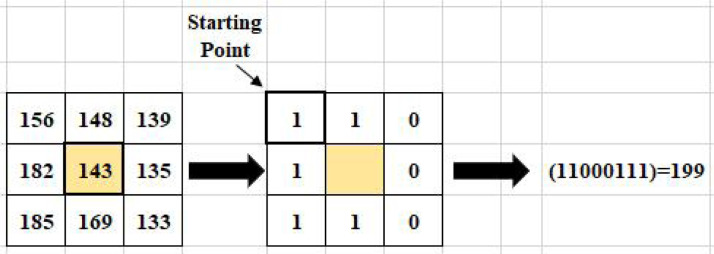
Another feature extraction method used in the study is the Local Binary Pattern (LBP), a nonparametric method. It was originally proposed as a tissue pattern analysis technique [52]. The basic functioning of this method is to establish bilateral relationships between the center and neighboring pixels. This method matches and labels neighboring pixels based on central pixel values in a 3 3 frame. The neighboring pixel takes the value 1 if it is greater than or equal to the central pixel, otherwise, it takes the value 0. Thus, an 8-bit code is generated for each pixel in the LBP neighborhood. Then, an identifier with the LBP code is created on each image as shown in Fig. 10. These operations are calculated using Eq. (3);
| (3) |
central pixel, neighbors of central pixel, the distance of central pixel to the, number of neighbors traded.
Fig. 10.
An example of generating an LBP identifier code.
In this study, PCA [53] is applied for feature selection and the 500 eigenvectors with the highest eigenvalues are selected. PCA is a method of expressing the variance structure of p number of variables as less number and linear components of these variables [54]. In other words, PCA is a statistical technique used to reduce the size of data by selecting the most important features that provide maximum information about the data set. The PCA method provides advantages in terms of removing the correlation between features, contributing to performance, and reducing overfitting. We can say that the necessity of standardizing the data, the loss of information if the correct number of principal components is not selected, and the formation of less interpretable principal components as disadvantages. Uses the eigenvalues and eigenvectors of the covariance matrix to find the linear components of the p variable in the data matrix. After feature extraction and selection steps, the classification process was started.
3.3. Classification phase
At this stage of the study, k-NN, SVM, Bag of Tree, and Kernel Extreme Learning Machine (K-ELM) methods were used to train the selected features and perform the classification process. These methods are among the classical learning methods and can produce very successful results in both two-class and multi-class learning. With the increasing popularity of deep learning methods in recent years, the rate of using classical learning methods has decreased. However, this does not mean that classical learning methods produce unsuccessful results. In our study, it has been shown that classical learning methods can produce successful results as well as deep learning methods. A study with low complexity and rapid results with classical learning methods has been revealed.
The first method we used in our study is the k-NN method, which is used in many different areas and produces simple but highly successful results. The k-NN method is a nonparametric classification method [55] and is frequently used in signal and image processing applications [56], [57], [58]. k-NN is a method based on the classification of objects according to their nearest examples in the feature space [55]. The k-NN algorithm is easy to use and apply method. The training process for this algorithm consists solely of storing the feature vectors and tags of the training images. In the classification process, the untagged query point is simply assigned to the tag of its nearest neighbor. Typically, the object is classified by the majority of votes based on the tags of its nearest neighbors. If , the object is classified as the class of the object nearest to it. Different metrics are used to determine the distances of the neighbors. The most common metrics are Manhattan, Euclidean, and Minkowski metrics. Let and Y be data points and n is the sample size. To find the distance between these data points, Minkowski distance is calculated as follows;
| (4) |
In this formula, we can manipulate the distance metric by changing the -value. For this reason, the Minkowski distance is also called the norm. In this formula, if the -value is 1, the distance metric is Manhattan, and if it is 2, it is Euclidean. The most important point where the k-NN method is disadvantageous is the number of starting numbers. Besides, we can list the slowness of the algorithm, high computational cost, and high memory requirement as other disadvantages. We can say that its application is simple, it can be applied in both classification and regression problems, and it allows not only binary classification but also multiple classifications as advantageous methods.
Another classification method used in the study is the SVM method. SVM method is one of the most commonly used methods in both image processing and medical image processing [59]. Although the foundations of the SVM method date back to the 60s, it has reached its current state in 1995 [60]. We can define the SVM method as a vector-based classification method that finds a hyper-plane between two classes to ensure that the data in each class is at the maximum distance from each other [61]. It is an effective method in cases where the number of dimensions is higher than the number of samples and in high dimensional spaces. Its features such as the ability to use different kernel functions and the efficient use of memory are other aspects where it is advantageous. We can say that it is disadvantageous in situations such as low performance in large data sets and noisy data and high time complexity. Various studies have also been done to overcome these disadvantages of SVM [62], [63].
The third classification method we use in our study is the Bag of Tree method, which we can define as an improved decision tree method. The Bag of Tree method uses a group of decision trees for classification or regression operations. A more effective method has been created by using weak decision trees together to create a community, i.e. bagging. Each tree in the community is grown on a copy drawn independently of the input data. Instances not included in this copy are classified as out of the bag. The basis of the Bag of Tree method is a random forest algorithm [64]. The bag of tree method is a method based on the Ensemble Learning technique. Because of this aspect, it reduces the overfitting problem and variance in decision trees. This increases the accuracy. Also, we can say that it is advantageous to work well with both categorical and continuous variables and to be used for both classification and regression problems. It is a method used frequently in image processing and signal processing due to its advantages [65], [66]. We can list the disadvantages of very high transaction complexity and long training time requirement.
The extreme learning machine (ELM) is a new learning algorithm for a single hidden layer feed-forward neural network (SLFN) [67], [68]. In this method, the input weights and biases of ELM are selected randomly and output weights are determined analytically. In 2006, Huang et al. compared the performance of SVM, ELM, and back propagation-based SLFN methods in terms of training time and accuracy [69]. The weight and bias values in the input layer are determined randomly, independent of the data. Output weights are calculated analytically as shown in (5).
| (5) |
where is the output weights vector between the output layer and hidden layers, is the output vector of the hidden layer, is nonlinear piecewise continuous function, randomly generated input values. In our study, the sigmoid activation function was used due to its widespread use in the literature.
Sigmoid Function:
| (6) |
Here, from the input layer to the hidden layer, x is an input sample, a is the weight value and b is the bias value. The {a, b} pair is randomly generated. ELM minimizes training error and the norm of output weights [67], [69]. Minimized: and where H is the hidden layer output matrix;
| (7) |
The least-squares method was used instead of standard optimization methods in the application phase of ELM [70].
| (8) |
is the tag matrix and is the inverse of the Moore–Penrose generalized hidden layer output matrix. [ (),…, h ()] is obtained with the training sample. Here, the function of the Moore–Penrose generalized latent layer output matrix inverse is to minimize the L2 norms of . A regularization coefficient is included in the optimization procedure to increase the robustness and generalization capability of the ELM. Therefore, given a kernel, the weight set is learned as:
| (9) |
The system was modeled with linear, polynomial, and radial basis function kernels, and since it was seen that the most successful result was obtained with radial basis function, RBF kernel was used in the study. The classification stage is the last stage of our work. 10-fold cross validity in the classification process. The parameter values of all methods used in the study are shown in Table 5.
Table 5.
The purpose and parameters of the methods used in the study.
| Process | Aim of use | Parameters |
|---|---|---|
| Image resizing | Preprocessing | Dataset-1 400 300 Dataset-2 525 525 Dataset-3 375 255 |
| Image Sharpening | Preprocessing | Radius 1 Strength of the sharpening effect 0.8 Threshold 0 |
| Gray level transformation | Preprocessing | Default values |
| HOG | Feature extraction | Variant HOG UoCTI Cell Size: 15 |
| LBP | Feature extraction | Window Size: 3 3 Cell Size: 15 |
| PCA | Feature selection | Algorithm: Singular value decomposition (SVD) |
| k-NN | Classification |
1 Neighbor Searcher Method exhaustive Distance minkowski Standardize 1 |
| Bag of Tree | Classification | # of Trees 100 |
| SVM | Classification | Default values |
| K-ELM | Classification | Kernel RBF # of Hidden Neurons 4096 C 1e−1 |
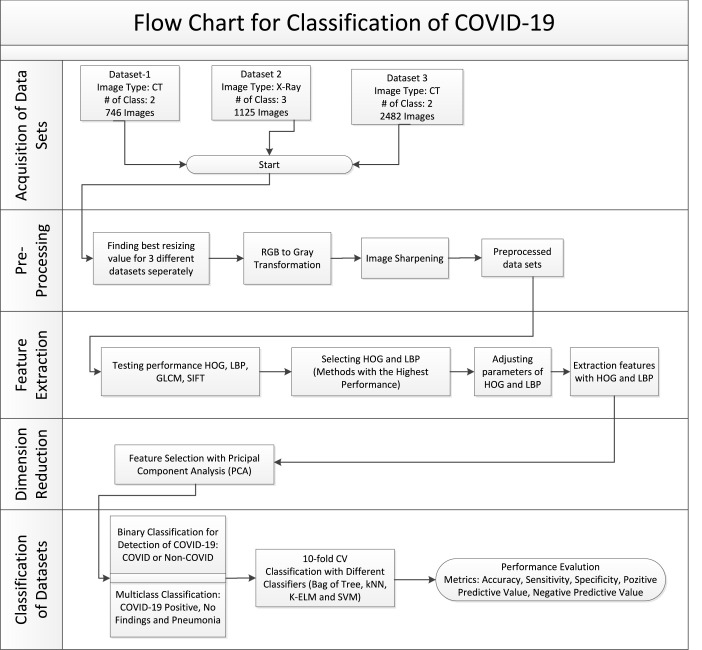
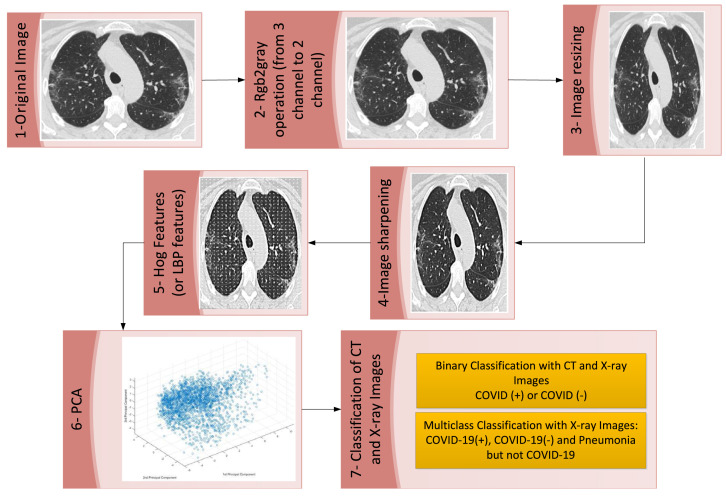
The flow diagram of our work is shown in Fig. 11. When the figure is examined, it is possible to see a structure that consists of 5 basic steps and each step has sub-steps. As can be seen from the flow chart, first of all, there is the acquisition of images for three different data sets. Then the pre-processing stage is started. In the preprocessing stage, resizing of images, image transformations, and image sharpening processes are performed. The next step after pre-processing is the feature extraction phase. There are many methods used for feature extraction in the literature. In this study, we used GLCM, HOG, LBP, and SIFT methods. According to the results obtained from the feature extraction methods, the HOG and LBP methods with the highest success were chosen. A large amount of data was obtained after the feature extraction step. For this reason, a dimension reduction process has been carried out. PCA method was used for dimension reduction. After this stage, the classification stage, which is the last stage of our study, started. Two different classification processes were carried out in the classification stage. The first of these is the binary classification process that distinguishes COVID (+) and COVID (-). Another classification process is the multiclass classification process in which the classes COVID (+), NO Findings, and Pneumonia but not COVID-19 are distinguished from each other. In the classification stage, the 10-fold cross-validation method was used to model the data. Five different metrics, namely Accuracy, Sensitivity, Specificity, Positive Predicted Value, and Negative Predicted Value, were used to evaluate the findings obtained as a result of the classification process.
Fig. 11.
Flow chart of the proposed model.
4. Results
In the experimental studies carried out, a computer with an i5 processor, GT 730 4 GB graphic card, and 16 GB RAM was used. MATLAB platform was used to realize all the stages in the flow diagram shown in Fig. 11. Accuracy, Sensitivity, Specificity, Positive Predictive Value (PPV), and Negative Predictive Value (PPV) metrics were used to evaluate the obtained classification results. The necessary formulas for the calculation of these metrics are shown in (10), (11), (12), (13), and (14).
| (10) |
| (11) |
| (12) |
| (13) |
| (14) |
TP, TN, FP, and FN mean true positive, true negative, false positive, and false negative, respectively.
It is possible to consider positive predictive value (PPV) and negative predictive value (NPV) as the clinical significance of a test. The main difference between PPV and NPV from sensitivity and specificity is that they use a prevalence [71]. Sensitivity is the percentage of true positives. Specificity is the percentage of true negatives. Accuracy is the rate at which all those with and without the disease can be correctly detected.
It was previously stated that 3 different data sets were used in our study. The classification process was carried out by dividing these data sets into training and test sets by the 10-fold cross-validation method. The binary classification process has been carried out on all data sets. Also, a multi-class classification process has been carried out in data set number 2, which consists of X-ray images.
Table 6 shows the success percentages obtained for data set number 1, which has the least number of images and uses CT images from three different data sets we used in our study. In the table, the highest rates obtained for each metric are marked in bold. It can be seen from the table that the most successful results are obtained with LBP feature extraction and modeled with a k-NN classifier. When the table is examined, it is seen that the ability to distinguish the patients (Sensitivity) is 86.53%, and the specificity is 91.94%. The correct detection rate (Accuracy) of all patients and non-patients is 89.41%.
Table 6.
Binary classification results for data set-1.
| Accuracy (%) | Sensitivity (%) | Specificity (%) | PPV (%) | NPV (%) | ||
|---|---|---|---|---|---|---|
| HOG | Bag of Tree | 80.97 | 77.36 | 84.13 | 81.08 | 80.87 |
| K-ELM | 80.16 | 77.94 | 82.12 | 79.30 | 80.89 | |
| k-NN | 84.45 | 83.95 | 84.89 | 83.00 | 85.75 | |
| SVM | 83.24 | 80.80 | 85.39 | 82.94 | 83.50 | |
| LBP | Bag of Tree | 78.02 | 65.90 | 88.67 | 83.64 | 74.73 |
| K-ELM | 80.83 | 76.50 | 84.63 | 81.40 | 80.38 | |
| k-NN | 89.41 | 86.53 | 91.94 | 90.42 | 88.59 | |
| SVM | 86.73 | 83.95 | 89.17 | 87.20 | 86.34 | |
Data set number 2 was the data set consisting of X-ray images. This dataset includes three classes: those with COVID-19 (+), those with No Findings, and those with Pneumonia but not COVID-19. The success rates obtained for this three-class data set can be seen in Table 7. Although the number of classes is three, it is an important issue that the accuracy rate is around 85.96%. Another remarkable point is that the Sensitivity (94.40%) and Specificity (100%) values are considerably higher than the binary classification data set 1 in X-ray images with three classes. From the table, it can be said that the SVM method produces more successful results than other methods in the multi-class classification process of X-ray images when looking at both healthy and sick patients in general.
Table 7.
Multi-class classification results for data set-2.
| Accuracy (%) | Sensitivity (%) | Specificity (%) | PPV (%) | NPV (%) | ||
|---|---|---|---|---|---|---|
| HOG | Bag of Tree | 80.71 | 64.80 | 99.80 | 97.59 | 95.78 |
| K-ELM | 81.60 | 82.40 | 99.40 | 94.50 | 97.83 | |
| k-NN | 76.09 | 86.40 | 99.00 | 91.53 | 98.31 | |
| SVM | 84.89 | 87.20 | 99.50 | 95.61 | 98.42 | |
| LBP | Bag of Tree | 82.84 | 71.20 | 100.00 | 100.00 | 96.53 |
| K-ELM | 80.18 | 88.00 | 99.40 | 94.83 | 98.51 | |
| k-NN | 77.51 | 94.40 | 98.80 | 90.77 | 99.30 | |
| SVM | 85.96 | 88.80 | 99.80 | 98.23 | 98.62 | |
The binary classification was also made for the X-ray images in dataset 2 in the study. Binary classification is primarily made between the COVID-19 (+) and No Findings classes. However, another binary classification has been made between the Pneumonia but non-COVID-19 class and the COVID-19 (+) class. Finally, the class of no findings and the class of non-COVID-19 Pneumonia are classified together as a class and COVID-19 (+) ones as a class. The results obtained are shown in Table 8, Table 9, Table 10, respectively. When Table 8 is examined, it is seen that the highest accuracy rate is obtained when modeled with the HOG feature extraction method and K-ELM classification method. On the other hand, a high accuracy rate of 98.88% was obtained in the process of classifying those with COVID-19 (+) and those without any symptoms in X-ray images. Besides, the highest success rates obtained for Sensitivity, Specificity, PPV, and NPV values are 96%, 100%, 100%, and 99%, respectively.
Table 8.
Binary classification (COVID vs. No Findings) results for data set 2.
| Accuracy (%) | Sensitivity (%) | Specificity (%) | PPV (%) | NPV (%) | ||
|---|---|---|---|---|---|---|
| HOG | Bag of Tree | 96.48 | 83.20 | 99.80 | 99.05 | 95.96 |
| K-ELM | 98.88 | 94.40 | 100.00 | 100.00 | 98.62 | |
| k-NN | 98.08 | 91.20 | 99.80 | 99.13 | 97.84 | |
| SVM | 98.72 | 94.40 | 99.80 | 99.16 | 98.62 | |
| LBP | Bag of Tree | 96.32 | 81.60 | 100.00 | 100.00 | 95.60 |
| K-ELM | 98.24 | 92.00 | 99.80 | 99.14 | 98.04 | |
| k-NN | 98.72 | 96.00 | 99.40 | 97.56 | 99.00 | |
| SVM | 98.72 | 96.00 | 99.40 | 97.56 | 99.00 | |
Table 9.
Binary classification (COVID vs. Pneumonia) results for data set 2.
| Accuracy (%) | Sensitivity (%) | Specificity (%) | PPV (%) | NPV (%) | ||
|---|---|---|---|---|---|---|
| HOG | Bag of Tree | 94.72 | 75.20 | 99.60 | 97.92 | 94.14 |
| K-ELM | 96.96 | 87.20 | 99.40 | 97.32 | 96.88 | |
| k-NN | 95.68 | 86.40 | 98.00 | 91.53 | 96.65 | |
| SVM | 97.76 | 92.80 | 99.00 | 95.87 | 98.21 | |
| LBP | Bag of Tree | 94.40 | 72.00 | 100.00 | 100.00 | 93.46 |
| K-ELM | 98.24 | 92.80 | 99.60 | 98.31 | 98.22 | |
| k-NN | 96.16 | 91.20 | 97.40 | 89.76 | 97.79 | |
| SVM | 98.56 | 93.60 | 99.80 | 99.15 | 98.42 | |
Table 10.
Binary classification (COVID vs. Pneumonia +No Findings) results for data set 2.
| Accuracy (%) | Sensitivity (%) | Specificity (%) | PPV (%) | NPV (%) | ||
|---|---|---|---|---|---|---|
| HOG | Bag of Tree | 95.64 | 62.40 | 99.80 | 97.50 | 95.50 |
| K-ELM | 97.69 | 84.00 | 99.40 | 94.59 | 98.03 | |
| k-NN | 97.60 | 85.60 | 99.10 | 92.24 | 98.22 | |
| SVM | 98.49 | 92.00 | 99.30 | 94.26 | 99.00 | |
| LBP | Bag of Tree | 95.20 | 56.80 | 100.00 | 100.00 | 94.88 |
| K-ELM | 98.93 | 90.40 | 100.00 | 100.00 | 98.81 | |
| k-NN | 98.13 | 93.60 | 98.70 | 90.00 | 99.20 | |
| SVM | 99.02 | 92.80 | 99.80 | 98.31 | 99.11 | |
Table 9 shows the results of the comparison of those with COVID-19 and non-COVID-19 pneumonia. 98.56% success rate was obtained in modeling with LBP and SVM. When we compare Table 8, Table 9, it determined those who did not show any findings with a rate of 96%, while non-COVID-19 pneumonia was detected with a rate of 93.60%. The highest accuracy rates (98.88%, 98.56%) obtained for Table 8, Table 9 are close to each other. In addition to these, it can be seen from Table 9 that although there are foggy images in both classes, the ability to distinguish patients with pneumonia as COVID-19 and non-COVID is quite high. This is an indication that the proposed model is very successful.
In Table 10, a class was created by combining two non-COVID classes (No Findings and non-COVID pneumonia) of X-ray images, and the results were binary classified. Here, an accuracy rate of more than 99% has been achieved.
The success results obtained as a result of applying the proposed approach to our third and last data set are shown in Table 11. Considering the results obtained on this data set, it is seen that the highest success rates are obtained with k-NN, which is a simple but effective method. The number of CT images in this dataset is 2482, which is an indication of the high generalization performance of the proposed approach.
Table 11.
Binary classification results for data set 3.
| Accuracy (%) | Sensitivity (%) | Specificity (%) | PPV (%) | NPV (%) | ||
|---|---|---|---|---|---|---|
| HOG | Bag of Tree | 86.90 | 85.38 | 88.45 | 88.27 | 85.59 |
| K-ELM | 91.70 | 91.69 | 91.70 | 91.84 | 91.55 | |
| k-NN | 97.90 | 98.40 | 97.40 | 97.47 | 98.36 | |
| SVM | 95.85 | 96.09 | 95.61 | 95.70 | 96.00 | |
| LBP | Bag of Tree | 85.21 | 82.51 | 87.96 | 87.47 | 83.15 |
| K-ELM | 90.08 | 90.02 | 90.15 | 90.30 | 89.86 | |
| k-NN | 98.11 | 98.80 | 97.40 | 97.48 | 98.76 | |
| SVM | 96.41 | 96.96 | 95.95 | 95.97 | 96.88 | |
The implementation times of the proposed approach in all data sets are shown in Table 12. When the table is examined, it is seen that the feature extraction process with the HOG method is faster than the LBP method. Among the classification methods, it is seen that the k-NN method produces much faster results than other methods. One of the most effective points of our study can be seen by looking at Table 12. The fact that COVID-19 can be detected in less than 1 min can be touted as an indicator of success alone. When Table 12 is examined, it is seen that the slowest classification method is the Bag of Tree method. The confusion matrices of the classification results given in the tables above are shown in Table 13.
Table 12.
Average time of Covid-19 diagnosis by different methods.
| Binary classification dataset-1 CT images |
Binary classification dataset-3 CT images |
Binary classification dataset-2X-ray images (COVID – No findings) |
Multiclass (3-Class) classification dataset-2X-ray images |
||||||
|---|---|---|---|---|---|---|---|---|---|
| HOG | LBP | HOG | LBP | HOG | LBP | HOG | LBP | ||
| Time (s) | Bag of Tree | 149.5 | 219.1 | 286.2 | 474.8 | 209.0 | 391.8 | 337.7 | 600.4 |
| K-ELM | 77.6 | 51.1 | 77.9 | 146.8 | 52.9 | 252.9 | 108.3 | 226.8 | |
| k-NN | 10.70 | 17.10 | 40.8 | 91.9 | 18.2 | 46.8 | 35.7 | 80.9 | |
| SVM | 32.7 | 76.2 | 151.8 | 362.3 | 27.5 | 51.5 | 155.7 | 304.5 | |
Table 13.
The confusion matrices were obtained as a result of the classification process. (Green values in the lower right corner show Accuracy, Bottom-Left blue value Sensitivity, Bottom-right blue value specificity, Right-top blue PPV, Right-bottom blue NPV values. (a)-Dataset-1, (b)-Dataset-2 (Multiclass), (c, d, e) - Dataset-2 (Binary), (f)-Dataset-3))
 |
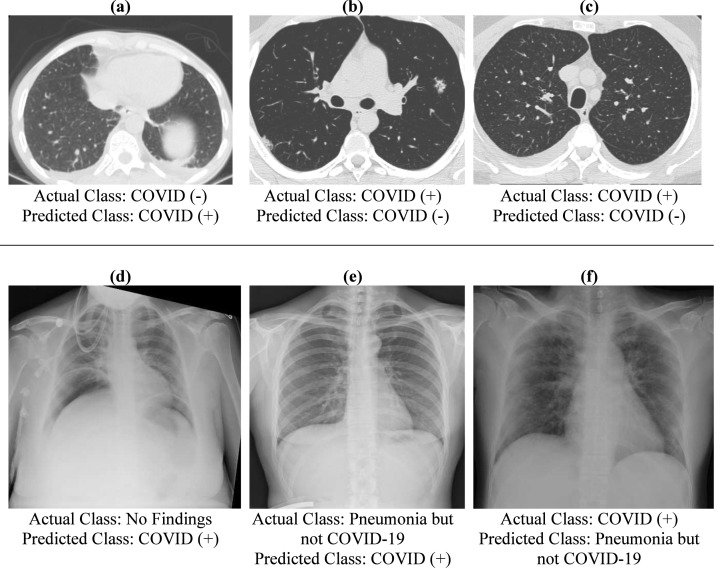
False-positive and false-negative examples seen in the confusion matrix are shown in Fig. 12. Fig. 12a shows a CT image that is not actually COVID-19 but has been incorrectly diagnosed with COVID-19. The cause of the error is thought to be because the appearance seen in the left lobe resembles the GGO seen in COVID-19. In Fig. 12b, a COVID-19 (+) CT image is classified as COVID-19 (-). Here, as the CT image is taken at the early stage of the disease, there is no obvious finding on the lungs. There is a small GGO at the bottom of the right lobe, but the system missed it. Similarly, Fig. 12c is thought to be an image taken in the early stages of the disease. It appears to be a very similar image when compared to the COVID-19 (-) images. It has been mixed for this reason. Fig. 12 d, e and f show examples of faulty classifications on X-ray images. Both images e and f have pneumonia, but one is caused by COVID and the other is caused by another disease. The system misdiagnosed these two images. In Fig. 12 d, an image with no findings was diagnosed with COVID. Here, too, it is seen that the reason is related to imaging. Looking at the image, it is seen that there are images similar to the GGO.
Fig. 12.
Examples of images detected incorrectly as a result of the study.
Fig. 13 was created to make the operations we performed in our study more understandable. In Fig. 13, the process steps and the results of the steps are displayed on a sample CT image from the data sets we use. The details will be understood more clearly when zooming on the images.
Fig. 13.
Display of the process steps of the study on the sample image.
5. Discussions
The main purpose of this study is to establish a system that can support the diagnosis and treatment process by detecting the COVID-19 virus as soon as possible. The study and the results obtained show that there is sufficient evidence to achieve this goal. Table 14 shows the performances of this study and other studies that use one of the three datasets in the literature. It can be seen clearly from Table 14 that almost all of the results obtained in our study are more successful than other studies in the literature. The only classification result with low success; while the accuracy value of the multi-class study performed for data set 2 was 85.96% in our study, it was 87.02% in the study number [32]. However, as can be seen from the Table, our study is ahead in all other metrics.
Table 14.
Comparison of the performances of this study and other studies in the literature.
| Author(s) | Dataset | Accuracy (%) | Sensitivity (%) | Specificity (%) | PPV (%) | NPV (%) |
|---|---|---|---|---|---|---|
| Ozturk et al. [33] | Dataset-2 (Multiclass) | 87.02 | 85.35 | 92.18 | 89.96 | NA |
| Ozturk et al. [33] | Dataset-2 (Binary) | 98.08 | 95.13 | 95.3 | 98.03 | NA |
| Soares et al. [34] | Dataset-3 | 97.38 | 95.53 | NA | 99.16 | NA |
| Yang et al. [35] | Dataset-1 | 89.0 | NA | NA | NA | NA |
| Jaiswal et al. [40] | Dataset-3 | 96.25 | 96.29 | 96.21 | 96.29 | NA |
| Our study | Dataset-1 | 89.41 | 86.53 | 91.94 | 90.42 | 88.59 |
| Our study | Dataset-2 (Multiclass) | 85.96 | 94.40 | 100.00 | 100.00 | 99.30 |
| Our study | Dataset-2 (Binary) | 98.88 | 96.00 | 100.00 | 100.00 | 98.42 |
| Our study | Dataset-3 | 98.11 | 98.80 | 97.40 | 97.48 | 98.76 |
Another important aspect of the study contributing to the literature is the duration of diagnosis. Table 15 shows that the COVID-19 test was detected in approximately 4 h by the RT-PCR test. Another striking issue in the same table is the diagnosis time of CT images is 21.5 min [72]. The 21.5-minute evaluation period mentioned here includes all processes such as taking the image, transmitting it to the radiology specialist by the technician, and the radiology specialist communicating its evaluation to the relevant doctor. Under normal circumstances, a CT image can be evaluated and diagnosed within seconds by an experienced radiologist. Our study aims to provide support to radiology specialists by further reducing evaluation times that are prolonged due to these processes. As we have stated in many parts of our study, one of our main goals is to provide mechanisms that can assist medical professionals in the diagnosis. COVID-19 has caused incredible levels of intensity in the medical industry all over the world. The health sector of many countries has come to a collapse level. Tents were built in the hospital gardens to cope with the intensity. It is quite natural that there may be issues that radiologists overlook in such a stressful and tiring work tempo. It is thought that our study will contribute in this sense as well.
Table 15.
Average time of Covid-19 diagnosis by different methods.
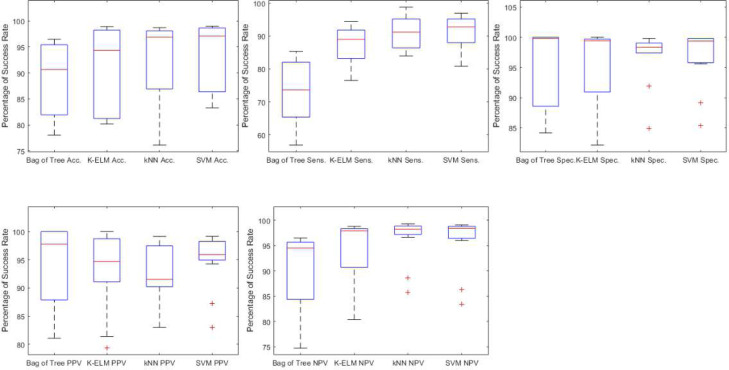
Table 6, Table 7, Table 8, Table 9, Table 10, Table 11 show the results of five different metrics obtained as a result of the classification of features obtained by four different classifiers and two different feature extractor methods. From all these tables, it seems a little difficult to analyze the success of classifiers in general. For this, the Box Plot chart shown in Fig. 14 was created. This figure has been obtained by using all the success rates in the tables mentioned. When Fig. 14 is examined, it is seen that the best results are obtained in k-NN and SVM methods according to Accuracy, Sensitivity, and NPV metrics. On the other hand, it is seen that the Bag of Tree method is good only in Specificity and PPV results, and in other metrics, it is the worst classifier. For K-ELM, it is possible to say that it is good in all metrics. It would not be wrong to say that it is the most appropriate method in overall evaluation even if it does not have the highest success rate in any of the five different metrics.
Fig. 14.
Comparison of the metric values of the classification methods in all data sets.
6. Conclusions
In this study, COVID-19 was detected in three different data sets with accuracy rates of 89.41%, 99.02%, and 98.11%. Two of the data sets used to consist of CT images, while one consists of X-ray images. On the other hand, an accuracy of 85.96% was achieved in the X-ray dataset for those with COVID-19 (+), COVID-19 (-), and Pneumonia but not the COVID-19 class. Thus, the study has been shown to produce successful results in both CT and X-ray imaging modes. Also, the fact that the study was applied to three data sets with different characteristics shows its generalizability. This is one of the most important features of the study. As a result of the study, it has been shown that COVID-19 can be detected with a high success rate in less than a minute with image processing and classical learning methods.
A model that can classify COVID-19 patients with the help of chest CT and X-ray images is proposed in the study. In the model implemented, the 10-fold cross-validation method was used in the separation of training and test data. The training data set was used to create the model, and the test data set was used for the verification process. In the proposed model, different image processing techniques were applied to CT and X-ray images. Afterward, by applying a large number of classifiers to the preprocessed images, it was shown that the system achieved a higher success than the studies in the literature. It has been observed that k-NN and SVM, which are among the classical learning methods, can detect COVID-19 (+) very successfully. On the other hand, in the classification of both (+) and (-) data, it was observed that the K-ELM method produced relatively better results than other methods used, with a holistic perspective. In addition to all these successful results, the implementation times of the system are also remarkably good. The model creation and implementation times of the system are under 1 min. This shows that the system is highly applicable. Looking at the experimental results, it is understood that the proposed model performs faster and better than other models. Especially in a period when deep learning models have become widespread in recent years, it has been shown that very high success rates and very short implementation times can be achieved after the data set of classical learning methods are processed correctly.
COVID-19 is a highly contagious virus. For this reason, the primary way to stop the virus is to reduce the transmission rate. This can only be achieved with early diagnosis and treatment. Early diagnosis of the diseases caused by the COVID-19 virus is the common goal of both our study and other studies in the literature. For this reason, a system with both a short duration and a high success rate are inevitable. This proposed system is exactly an example of this. It would not be wrong to say that it is the most original aspect of this study that it can achieve success rates of 99% in less than 1 min.
Clinical information was not available for the individuals included in the data sets used in the study. For this reason, only image analysis was performed and it could be analyzed whether the patients had the COVID-19 virus or not. However, conditions such as being in the advanced age group and underlying diseases are matters that can change the course of the disease. Therefore, it would not be wrong to consider the lack of clinical data on patients as a limitation of our study. We will make a comprehensive evaluation by ensuring that the clinical features and laboratory examination information of the cases we include in our next study are also included. Also, there is no information on which day of the disease the images in the existing data sets belong to. This is another limitation of the study. If there is information that the radiological images of patients are taken on which day of the disease, very different and useful evaluations can be made about the course of the disease. In our future studies, it is aimed to make more detailed and different analyzes by including the information on the day of the illness as well as the radiological images of the patients. The future goal of our study is to create a comparative model by including deep learning methods. In this way, the performance and implementation times of both deep learning methods and classical learning methods will be analyzed.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- 1.World Health Organization; Geneva: 2020. WHO COVID- 19 Explorer. Available online: https://worldhealthorg.shinyapps.io/covid/, (retrieved 17th December 2020) [Google Scholar]
- 2.Carfì A., Bernabei R., Landi F. Persistent symptoms in patients after acute COVID-19. JAMA. 2020;324:603–605. doi: 10.1001/jama.2020.12603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ai T., Yang Z., Hou H., Zhan C., Chen C., Lv W., Tao Q., Sun Z., Xia L. Correlation of chest CT and RT-PCR testing in coronavirus disease 2019 (COVID-19) in China: a report of 1014 cases. Radiology. 2020 doi: 10.1148/radiol.2020200642. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Fang Y., Zhang H., Xie J., Lin M., Ying L., Pang P., Ji W. Sensitivity of chest CT for COVID-19: comparison to RT-PCR. Radiology. 2020 doi: 10.1148/radiol.2020200432. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Lan L., Xu D., Ye G., Xia C., Wang S., Li Y., Xu H. Positive RT-PCR test results in patients recovered from COVID-19. JAMA. 2020;323:1502–1503. doi: 10.1001/jama.2020.2783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Bernheim A., Mei X., Huang M., Yang Y., Fayad Z.A., Zhang N., Diao K., Lin B., Zhu X., Li K. Chest CT findings in coronavirus disease-19 (COVID-19): relationship to duration of infection. Radiology. 2020 doi: 10.1148/radiol.2020200463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Borghesi A., Maroldi R. COVID-19 outbreak in Italy: experimental chest X-ray scoring system for quantifying and monitoring disease progression. La Radiol. Med. 2020:1. doi: 10.1007/s11547-020-01200-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Jacobi A., Chung M., Bernheim A., Eber C. Portable chest X-ray in coronavirus disease-19 (COVID-19): A pictorial review. Clin. Imaging. 2020 doi: 10.1016/j.clinimag.2020.04.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Wong H.Y.F., Lam H.Y.S., Fong A.H.-T., Leung S.T., Chin T.W.-Y., Lo C.S.Y., Lui M.M.-S., Lee J.C.Y., Chiu K.W.-H., Chung T. Frequency and distribution of chest radiographic findings in COVID-19 positive patients. Radiology. 2020 doi: 10.1148/radiol.2020201160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Kanne J.P., Little B.P., Chung J.H., Elicker B.M., Ketai L.H. 2020. Essentials for radiologists on COVID-19: an update—radiology scientific expert panel, in, radiological society of north america. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Xie X., Zhong Z., Zhao W., Zheng C., Wang F., Liu J. Chest CT for typical 2019-nCoV pneumonia: relationship to negative RT-PCR testing. Radiology. 2020 doi: 10.1148/radiol.2020200343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lee E.Y., Ng M.-Y., Khong P.-L. COVID-19 pneumonia: what has CT taught us? Lancet Infect. Dis. 2020;20:384–385. doi: 10.1016/S1473-3099(20)30134-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Pan F., Ye T., Sun P., Gui S., Liang B., Li L., Zheng D., Wang J., Hesketh R.L., Yang L. Time course of lung changes on chest CT during recovery from 2019 novel coronavirus (COVID-19) pneumonia. Radiology. 2020 doi: 10.1148/radiol.2020200370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Yoon S.H., Lee K.H., Kim J.Y., Lee Y.K., Ko H., Kim K.H., Park C.M., Kim Y.-H. Chest radiographic and CT findings of the 2019 novel coronavirus disease (COVID-19): analysis of nine patients treated in Korea. Korean J. Radiol. 2020;21:494–500. doi: 10.3348/kjr.2020.0132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Ng M.-Y., Lee E.Y., Yang J., Yang F., Li X., Wang H., M.M.-s. Lui C.M., Lo C.S.-Y., Leung B., Khong P.-L. Imaging profile of the COVID-19 infection: radiologic findings and literature review. Radiol.: Cardiothoracic Imaging. 2020;2 doi: 10.1148/ryct.2020200034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Ye Z., Zhang Y., Wang Y., Huang Z., Song B. Chest CT manifestations of new coronavirus disease 2019 (COVID-19): a pictorial review. Eur. Radiol. 2020:1–9. doi: 10.1007/s00330-020-06801-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Dheeba J., Singh N.A., Selvi S.T. Computer-aided detection of breast cancer on mammograms: A swarm intelligence optimized wavelet neural network approach. J. Biomed. Inform. 2014;49:45–52. doi: 10.1016/j.jbi.2014.01.010. [DOI] [PubMed] [Google Scholar]
- 18.Saygılı A. Classification and diagnostic prediction of breast cancers via different classifiers. Int. Sci. Vocat. Stud. J. 2018;2:48–56. [Google Scholar]
- 19.Dong H., Yang G., Liu F., Mo Y., Guo Y. Annual Conference on Medical Image Understanding and Analysis. Springer; 2017. Automatic brain tumor detection and segmentation using U-net based fully convolutional networks; pp. 506–517. [Google Scholar]
- 20.A. Saygılı, A novel approach to heart attack prediction improvement via extreme learning machines classifier integrated with data resampling strategy, Konya Mühendislik Bilimleri Dergisi, 8 853-865.
- 21.Armocida B., Formenti B., Ussai S., Palestra F., Missoni E. The Italian health system and the COVID-19 challenge. Lancet Publ. Health. 2020;5 doi: 10.1016/S2468-2667(20)30074-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Ortega F., Orsini M. Governing COVID-19 without government in Brazil: Ignorance, neoliberal authoritarianism, and the collapse of public health leadership. Glob. Publ. Health. 2020;15:1257–1277. doi: 10.1080/17441692.2020.1795223. [DOI] [PubMed] [Google Scholar]
- 23.Hemdan E.E.-D., Shouman M.A., Karar M.E. 2020. Covidx-net: A framework of deep learning classifiers to diagnose covid-19 in x-ray images. arXiv preprint arXiv:2003.11055. [Google Scholar]
- 24.Barstugan M., Ozkaya U., Ozturk S. 2020. Coronavirus (covid-19) classification using ct images by machine learning methods. arXiv preprint arXiv:2003.09424. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.LINDA W. A tailored deep convolutional neural network design for detection of covid-19 cases from chest radiography images. J. Netw. Comput. Appl. 2020 doi: 10.3389/fmed.2020.608525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Maghdid H.S., Asaad A.T., Ghafoor K.Z., Sadiq A.S., Khan M.K. 2020. Diagnosing COVID-19 pneumonia from X-ray and CT images using deep learning and transfer learning algorithms. arXiv preprint arXiv:2004.00038. [Google Scholar]
- 27.Ghoshal B., Tucker A. 2020. Estimating uncertainty and interpretability in deep learning for coronavirus (COVID-19) detection. arXiv preprint arXiv:2003.10769. [Google Scholar]
- 28.Hall L.O., Paul R., Goldgof D.B., Goldgof G.M. 2020. Finding covid-19 from chest x-rays using deep learning on a small dataset. arXiv preprint arXiv:2004.02060. [Google Scholar]
- 29.Cohen J.P., Morrison P., Dao L., Roth K., Duong T.Q., Ghassemi M. 2020. Covid-19 Image data collection: Prospective predictions are the future. arXiv preprint arXiv:2006.11988. [Google Scholar]
- 30.Farooq M., Hafeez A. 2020. Covid-resnet: A deep learning framework for screening of covid19 from radiographs. arXiv preprint arXiv:2003.14395. [Google Scholar]
- 31.Abbas A., Abdelsamea M.M., Gaber M.M. 2020. Classification of COVID-19 in chest X-ray images using detrac deep convolutional neural network. arXiv preprint arXiv:2003.13815. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Singh D., Kumar V., Kaur M. Classification of COVID-19 patients from chest CT images using multi-objective differential evolution–based convolutional neural networks. Eur. J. Clin. Microbiol. Infect. Dis. 2020:1–11. doi: 10.1007/s10096-020-03901-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Ozturk T., Talo M., Yildirim E.A., Baloglu U.B., Yildirim O., Acharya U.R. Automated detection of COVID-19 cases using deep neural networks with X-ray images. Comput. Biol. Med. 2020 doi: 10.1016/j.compbiomed.2020.103792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Soares E., Angelov P., Biaso S., Higa Froes M., Kanda Abe D. 2020. SARS-CoV-2 CT-Scan dataset: A large dataset of real patients CT scans for SARS-CoV-2 identification. medRxiv. [Google Scholar]
- 35.Zhao J., Zhang Y., He X., Xie P. 2020. COVID-CT-Dataset: a CT scan dataset about COVID-19. arXiv preprint arXiv:2003.13865. [Google Scholar]
- 36.Li Y., Wei D., Chen J., Cao S., Zhou H., Zhu Y., Wu J., Lan L., Sun W., Qian T. Efficient and effective training of COVID-19 classification networks with self-supervised dual-track learning to rank. IEEE J. Biomed. Health Inf. 2020;24:2787–2797. doi: 10.1109/JBHI.2020.3018181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Hu S., Gao Y., Niu Z., Jiang Y., Li L., Xiao X., Wang M., Fang E.F., Menpes-Smith W., Xia J. Weakly supervised deep learning for covid-19 infection detection and classification from ct images. IEEE Access. 2020;8 [Google Scholar]
- 38.Wu Y.-H., Gao S.-H., Mei J., Xu J., Fan D.-P., Zhao C.-W., Cheng M.-M. 2020. JCS: An explainable COVID-19 diagnosis system by joint classification and segmentation. arXiv preprint arXiv:2004.07054. [DOI] [PubMed] [Google Scholar]
- 39.Sun L., Mo Z., Yan F., Xia L., Shan F., Ding Z., Song B., Gao W., Shao W., Shi F. Adaptive feature selection guided deep forest for covid-19 classification with chest ct. IEEE J. Biomed. Health Inf. 2020;24:2798–2805. doi: 10.1109/JBHI.2020.3019505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Jaiswal A., Gianchandani N., Singh D., Kumar V., Kaur M. Classification of the COVID-19 infected patients using densenet201 based deep transfer learning. J. Biomol. Struct. Dyn. 2020:1–8. doi: 10.1080/07391102.2020.1788642. [DOI] [PubMed] [Google Scholar]
- 41.Abraham B., Nair M.S. Computer-aided detection of COVID-19 from X-ray images using multi-CNN and Bayesnet classifier. Biocybern. Biomed. Eng. 2020;40:1436–1445. doi: 10.1016/j.bbe.2020.08.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Altan A., Karasu S. Recognition of COVID-19 disease from X-ray images by hybrid model consisting of 2D curvelet transform, chaotic salp swarm algorithm and deep learning technique. Chaos Solitons Fractals. 2020;140 doi: 10.1016/j.chaos.2020.110071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Nour M., Cömert Z., Polat K. A novel medical diagnosis model for COVID-19 infection detection based on deep features and Bayesian optimization. Appl. Soft Comput. 2020 doi: 10.1016/j.asoc.2020.106580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Kassani S.H., Kassasni P.H., Wesolowski M.J., Schneider K.A., Deters R. 2020. Automatic detection of coronavirus disease (COVID-19) in X-ray and CT images: A machine learning-based approach. arXiv preprint arXiv:2004.10641. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Ardakani A.A., Acharya U.R., Habibollahi S., Mohammadi A. Covidiag: a clinical CAD system to diagnose COVID-19 pneumonia based on CT findings. Eur. Radiol. 2020:1–10. doi: 10.1007/s00330-020-07087-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Zhou T., Lu H., Yang Z., Qiu S., Huo B., Dong Y. The ensemble deep learning model for novel COVID-19 on CT images. Appl. Soft Comput. 2021;98 doi: 10.1016/j.asoc.2020.106885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Gupta A., Anjum H., Gupta S., Katarya R. Instacovnet-19: A deep learning classification model for the detection of COVID-19 patients using chest X-ray. Appl. Soft Comput. 2020 doi: 10.1016/j.asoc.2020.106859. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Aslan M.F., Unlersen M.F., Sabanci K., Durdu A. CNN-Based transfer learning–BiLSTM network: A novel approach for COVID-19 infection detection. Appl. Soft Comput. 2021;98 doi: 10.1016/j.asoc.2020.106912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.X. Wang, Y. Peng, L. Lu, Z. Lu, M. Bagheri, R.M., Summers, Chestx-ray8: Hospital-scale chest x-ray database and benchmarks on weakly-supervised classification and localization of common thorax diseases, in: Proceedings of the IEEE conference on computer vision and pattern recognition, 2017, pp. 2097-2106.
- 50.Dalal N., Triggs B. 2005 IEEE Computer Society Conference on Computer Vision and Pattern Recognition (CVPR’05) IEEE; 2005. Histograms of oriented gradients for human detection; pp. 886–893. [Google Scholar]
- 51.Felzenszwalb P.F., Girshick R.B., McAllester D., Ramanan D. Object detection with discriminatively trained part-based models. IEEE Trans. Pattern Anal. Mach. Intell. 2009;32:1627–1645. doi: 10.1109/TPAMI.2009.167. [DOI] [PubMed] [Google Scholar]
- 52.Ojala T., Pietikäinen M., Mäenpää T. Computer Vision - ECCV 2000: 6th European Conference on Computer Vision Dublin, Ireland, June 26 – July 1, 2000 Proceedings, Part I. Springer Berlin Heidelberg; Berlin, Heidelberg: 2000. Gray scale and rotation invariant texture classification with local binary patterns; pp. 404–420. [Google Scholar]
- 53.Jolliffe I.T. Principal Component Analysis. Springer; 1986. Principal component analysis and factor analysis; pp. 115–128. [Google Scholar]
- 54.Wold S., Esbensen K., Geladi P. Principal component analysis. Chemometr. Intell. Lab. Syst. 1987;2:37–52. [Google Scholar]
- 55.Arya S., Mount D.M., Netanyahu N.S., Silverman R., Wu A.Y. An optimal algorithm for approximate nearest neighbor searching fixed dimensions. J. ACM. 1998;45:891–923. [Google Scholar]
- 56.Glowacz A., Glowacz Z. Recognition of images of finger skin with application of histogram, image filtration and K-NN classifier. Biocybern. Biomed. Eng. 2016;36:95–101. [Google Scholar]
- 57.Glowacz A., Glowacz Z. Diagnosis of the three-phase induction motor using thermal imaging. Infrared Phys. Technol. 2017;81:7–16. [Google Scholar]
- 58.J. Kim, B. Kim, S. Savarese, Comparing image classification methods: K-nearest-neighbor and support-vector-machines, in: Proceedings of the 6th WSEAS international conference on Computer Engineering and Applications, and Proceedings of the 2012 American conference on Applied Mathematics, 2012, pp. 48109-42122.
- 59.Kayaaltı Ö., Aksebzeci B.H., Karahan İ.Ö., Deniz K., Öztürk M., Yılmaz B., Kara S., Asyalı M.H. Liver fibrosis staging using CT image texture analysis and soft computing. Appl. Soft Comput. 2014;25:399–413. [Google Scholar]
- 60.Cortes C., Vapnik V. Support vector machine. Mach. Learn. 1995;20:273–297. [Google Scholar]
- 61.Schölkopf B., Smola A.J., Bach F. MIT press; 2002. Learning with Kernels: Support Vector Machines, Regularization, Optimization, and beyond. [Google Scholar]
- 62.Cervantes J., García Lamont F., López-Chau A., Rodríguez Mazahua L., Sergio Ruíz J. Data selection based on decision tree for SVM classification on large data sets. Appl. Soft Comput. 2015;37:787–798. [Google Scholar]
- 63.Lee D. LS-GKM: a new gkm-SVM for large-scale datasets. Bioinformatics. 2016;32:2196–2198. doi: 10.1093/bioinformatics/btw142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Breiman L. Random forests. Mach. Learn. 2001;45:5–32. [Google Scholar]
- 65.Tang T.T., Zawaski J.A., Francis K.N., Qutub A.A., Gaber M.W. Image-based classification of tumor type and growth rate using machine learning: a preclinical study. Sci. Rep. 2019;9:12529. doi: 10.1038/s41598-019-48738-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Zhang J., Lin F., Xiong P., Du H., Zhang H., Liu M., Hou Z., Liu X. Automated detection and localization of myocardial infarction with staked sparse autoencoder and treebagger. IEEE Access. 2019;7:70634–70642. [Google Scholar]
- 67.Huang G.-B., Zhu Q.-Y., Siew C.-K. Extreme learning machine: Theory and applications. Neurocomputing. 2006;70:489–501. [Google Scholar]
- 68.Huang G.-B., Zhou H., Ding X., Zhang R. Extreme learning machine for regression and multiclass classification. IEEE Trans. Syst. Man Cybern. B. 2011;42:513–529. doi: 10.1109/TSMCB.2011.2168604. [DOI] [PubMed] [Google Scholar]
- 69.Huang G.-B., Zhu Q.-Y., Siew C.-K. Neural Networks, 2004. Proceedings. 2004 IEEE International Joint Conference on. IEEE; 2004. Extreme learning machine: a new learning scheme of feedforward neural networks; pp. 985–990. [Google Scholar]
- 70.Huang G.-B., Zhou H., Ding X., Zhang R. Extreme learning machine for regression and multiclass classification. IEEE Trans. Syst. Man Cybern. B. 2012;42:513–529. doi: 10.1109/TSMCB.2011.2168604. [DOI] [PubMed] [Google Scholar]
- 71.Wong H.B., Lim G.H. Measures of diagnostic accuracy: sensitivity, specificity, PPV and NPV. Proc. Singapore Healthc. 2011;20:316–318. [Google Scholar]
- 72.Huang Z., Zhao S., Li Z., Chen W., Zhao L., Deng L., Song B. The battle against coronavirus disease 2019 (COVID-19): emergency management and infection control in a radiology department. J. Amer. College Radiol. 2020 doi: 10.1016/j.jacr.2020.03.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Won J., Lee S., Park M., Kim T.Y., Park M.G., Choi B.Y., Kim D., Chang H., Kim V.N., Lee C.J. Development of a laboratory-safe and low-cost detection protocol for SARS-CoV-2 of the coronavirus disease 2019 (COVID-19) Exp. Neurobiol. 2020;29:107. doi: 10.5607/en20009. [DOI] [PMC free article] [PubMed] [Google Scholar]