Summary
Extremely low birth weight (ELBW) infants often develop an altered gut microbiota composition, which is related to clinical complications, such as necrotizing enterocolitis and sepsis. Probiotic supplementation may reduce these complications, and modulation of the gut microbiome is a potential mechanism underlying the probiotic effectiveness. In a randomized, double-blind, placebo-controlled trial, we assessed the effect of Lactobacillus reuteri supplementation, from birth to post-menstrual week (PMW)36, on infant gut microbiota. We performed 16S amplicon sequencing in 558 stool samples from 132 ELBW preterm infants at 1 week, 2 weeks, 3 weeks, 4 weeks, PMW36, and 2 years. Probiotic supplementation results in increased bacterial diversity and increased L. reuteri abundance during the 1st month. At 1 week, probiotic supplementation also results in a lower abundance of Enterobacteriaceae and Staphylococcaceae. No effects were found at 2 years. In conclusion, probiotics may exert benefits by modulating the gut microbiota composition during the 1st month in ELBW infants.
Keywords: microbiota, microbial diversity, preterm infant, extremely low birth weight infant, probiotic, supplementation, Lactobacillus, Staphylococcus, randomized controlled trial, necrotizing enterocolitis
Graphical abstract

Highlights
L. reuteri DSM 17938 dominates the microbiota in supplemented ELBW infants
Supplemented ELBW infants have higher gut microbiota richness and diversity
Supplemented infants have lower abundance of Enterobacteriaceae and Staphylococcaceae
No effects of probiotic supplementation on the gut microbiota at PMW36 and 2 years
Martí et al. demonstrate, in a randomized, double-blind, placebo-controlled trial, that daily supplementation of L. reuteri from birth to post-menstrual week 36 in ELBW preterm infants modulates the gut microbiota composition, resulting in increased bacterial diversity and high abundance of the supplemented strain during the 1st month of life.
Introduction
Although the care of preterm infants has developed dramatically during the last decades, about 23% of extremely low birth weight (ELBW) (birth weight <1,000 g) preterm infants, born in Sweden, die due to clinical complications,1 such as necrotizing enterocolitis (NEC)2 and sepsis,3 and the survivors have a high risk of long-term neurological disabilities.4 The pathogenesis of such clinical complications is multifactorial and has been attributed to the immature development of the immune system, altered intestinal epithelial barrier function, gut motility, and regulation of the microvascular circulation as well as formula feeding and antibiotic treatment.5,6 Current research interest has also focused on the composition and function of the developing gut microbiota, because an altered bacterial composition and lower diversity may be a major risk factor linked to these severe clinical complications.2, 3, 4,7 For example, increased relative abundance of Proteobacteria (class Gammaproteobacteria or family Enterobacteriaceae), decreased relative abundance of Firmicutes and Bacteroidetes, absence of Propionibacterium, and an overall decrease in microbial richness have been reported to precede the onset of NEC.2,8
Probiotic supplementation to very low birth weight (VLBW) infants has been shown to reduce the risk of complications, such as NEC9 and late-onset of sepsis.10 The effect, however, seems to be strain dependent, and there is still insufficient evidence for an effect in ELBW infants.9,10 The strain Lactobacillus reuteri DSM 17938 has been reported to reduce sepsis, feeding intolerance, and days on antibiotic treatment and improve growth and immune function in randomized-controlled trials in preterm infants.11, 12, 13, 14 Recently, it was also reported to reduce NEC in preterm infants in a strain-specific systematic review.15
The underlying mechanisms of the effectiveness of probiotics are hypothesized to be modulation of the gut microbiome, reinforcement of the intestinal barrier, and interaction with the innate and adaptive immune system.9,16,17 Competitive exclusion of pathogens either via secretion of antimicrobial intermediaries or via inhibition of pathogenic adhesion is another mechanism generally associated to L. reuteri.18,19 Despite that an effect on the gut microbiota has been one of the main hypotheses on how probiotics act in preterm infants, there are still very few reports on the gut microbiome composition in VLBW infants during the 1st month of life in randomized-controlled trials.
This study was part of a prospective randomized, double-blind, placebo-controlled, multi-center trial, in which 134 ELBW preterm infants were supplemented with either L. reuteri DSM 17938 or placebo daily from birth to post-menstrual week (PMW)36, evaluating feeding tolerance and growth.13 The primary aim of the present study was to investigate the effect of supplementation with L. reuteri DSM 17938 on the gut microbiota composition of ELBW preterm infants during the neonatal period and at a follow-up at 2 years of age. Secondary aims were to analyze the microbial composition in relation to NEC, sepsis, and growth rate. We hypothesized that L. reuteri supplementation would modulate the gut microbiota composition by, for example, enhancing the gut microbiota diversity and reducing opportunistic pathogens during the neonatal period.
Results
ELBW infant cohort
Overall, 132 infants were included in the study and a total of 558 stool samples were analyzed (Figure 1). The background and clinical characteristics of the infants included in the analyses at 1 week are displayed in Table 1 and at 2 weeks, 3 weeks, 4 weeks, PMW36, and 2 years in Table S1. The following clinical characteristics significantly differed between the L. reuteri and placebo groups: gender at 1 week and 3 weeks; delivery mode at 3 weeks; and treatment with antibiotics at 4 weeks, as well as chorioamnionitis, infants from multiple pregnancies, and total days on antibiotics at PMW36 (Table S1).
Figure 1.
Flow chart of the study
PMW36: post-menstrual week 36. aStudy product was discontinued by mistake after transfer to other hospital (n = 1). bStudy product was not administrated again by mistake after temporarily being withheld during nil oral (n = 3). cStudy product ran out temporarily at the study site (n = 3). dInsufficient amounts of DNA were recovered from extraction (n = 6). eLibrary preparation failed (n = 6). fSequencing failed (n = 12).
Table 1.
Background and clinical characteristics of extremely low birth weight preterm infants from which stool samples were collected at 1 week of age
| Variables | Statistical test | Placebo n = 54 | Lactobacillus reuteri n = 54 | p value |
|---|---|---|---|---|
| Gestational age, weeks, mean (SD) | Student’s t test | 25.5 (1.3) | 25.5 (1.3) | 0.89 |
| Birth weight, g, median (IQR) | Mann-Whitney U test | 763 (197.8) | 727.5 (172.2) | 0.15 |
| Birth weight Z score, median (IQR) | Mann-Whitney U test | −0.8 (1.3) | −0.9 (1.7) | 0.25 |
| Birth length, cm, median (IQR) | Mann-Whitney U test | 33.5 (3.5) | 33 (3.8) | 0.22 |
| Birth length Z score, median (IQR) | Mann-Whitney U test | −0.8 (1.6) | −1.6 (2.3) | 0.06 |
| Birth head circumference, cm, median (IQR) | Mann-Whitney U test | 23 (3) | 23 (2) | 0.12 |
| Birth head circumference Z score, mean (SD) | Student’s t test | −0.7 (0.8) | −1 (0.8) | 0.06 |
| Apgar score at 5 min, median (IQR) | Mann-Whitney U test | 7 (2) | 7 (4) | 0.68 |
| Apgar score at 10 min, median (IQR) | Mann-Whitney U test | 8 (2) | 8 (2.8) | 0.34 |
| Small for gestational age (weight <2 SD), n (%) | Pearson’s Χ2 test | 10 (19%) | 17 (31%) | 0.18 |
| Gender female/male, n (%) | Pearson’s Χ2 test | 19 (35%)/35 (65%) | 31 (57%)/23 (43%) | 0.03 |
| Infants from multiple pregnancy, n (%) | Pearson’s Χ2 test | 17 (31%) | 17 (31%) | 1.00 |
| Caesarean section, n (%) | Pearson’s Χ2 test | 30 (56%) | 40 (74%) | 0.07 |
| Maternal smoking, n (%) | Fisher’s exact test | 4 (7%) | 4 (7%) | 1.00 |
| Pre-eclampsia, n (%) | Fisher’s exact test | 4 (7%) | 5 (9%) | 1.00 |
| Chorioamnionitis, n (%) | Pearson’s Χ2 test | 8 (15%) | 14 (26%) | 0.23 |
| Preterm premature rupture of membranes, n (%) | Pearson’s Χ2 test | 13 (24%) | 19 (35%) | 0.29 |
| Maternal antibiotics, n (%) | Pearson’s Χ2 test | 27 (50%) | 33 (61%) | 0.33 |
| Antenatal corticosteroids, n (%) | Fisher’s exact test | 53 (98%) | 53 (98%) | 1.00 |
| Inclusion site—Stockholm/Linköping, n (%) | Pearson’s Χ2 test | 35 (65%)/19 (35%) | 35 (65%)/19 (35%) | 1.00 |
| Treatment with antibiotics within the actual week, n (%) | Pearson’s Χ2 test | 54 (100%) | 54 (100%) | NA |
| Total days on antibiotics, median (IQR) | Mann-Whitney U test | 7 (0) | 7 (0) | 0.93 |
| Total days with insulin, median (IQR) | Mann-Whitney U test | 0 (0) | 0 (0) | 0.52 |
| Insulin within the actual week, n (%) | Fisher’s exact test | 9 (17%) | 12 (22%) | 0.63 |
| Total days with corticosteroids, median (IQR) | Mann-Whitney U test | 0 (0) | 0 (0) | 0.41 |
| Corticosteroids within the actual week, n (%) | Fisher’s exact test | 4 (7%) | 2 (4%) | 0.68 |
Apgar score is missing from one infant in the L. reuteri group. See Table S1 for all the 6 time points (1 week to 2 years of age). IQR, inter-quartile range; NA, not applicable.
Microbial community composition and structure differs between the placebo and probiotic groups during the 1st month of life
The L. reuteri group had significantly higher bacterial richness, diversity, and evenness than the placebo group during the 1st month of life, except for richness at 4 weeks (Figure 2). The bacterial community composition (β-diversity) significantly differed between the two groups during the 1st month of life (analysis of similarities [ANOSIM], p = 0.001; Figures 3A–3D). Lactobacillus, Staphylococcus, and Klebsiella were the main genera contributing to the differences in bacterial composition. Lactobacillus was clearly associated with the L. reuteri group during the 1st month of life, although Staphylococcus and Klebsiella were associated with the placebo group at 1 week, 3 weeks, and 4 weeks and 2 weeks and 3 weeks, respectively. At 1 week, the inclusion site (Stockholm or Linköping) also affected the bacterial community composition (ANOSIM; p = 0.001). The difference between the two study groups was still significant after stratifying the data according to the inclusion site (ANOSIM; p = 0.001). At PMW36 and 2 years, there were no significant differences in bacterial community structure (α-diversity; Figure 2) and composition (β-diversity; Figures 3E and 3F) between the two study groups.
Figure 2.
Gut microbiota α-diversity of ELBW preterm infants supplemented with L. reuteri or placebo
Boxplots (median with 25% and 75% percentiles and 1.5× the interquartile range; diamond shape depicts the mean) showing the α-diversity (Shannon index), richness (observed ASVs), and evenness (Pielou’s evenness index) from 1 week to 2 years of life in ELBW preterm infants supplemented with L. reuteri (Lr) or placebo (Pl). PMW36, post-menstrual week 36; w, week; y, year. ∗∗∗p < 0.001, ∗∗p < 0.01, and ∗p < 0.05 with Mann-Whitney U test and p value adjustment for multiple comparisons with the method from Benjamini and Hochberg.
Figure 3.
Clustering of the gut microbiota composition (β-diversity) of ELBW preterm infants supplemented with L. reuteri or placebo
Non-metric multidimensional scaling (NMDS) of bacterial community composition from 1 week to 2 years of life across ELBW preterm infants supplemented with L. reuteri or placebo.
(A–F) The ASVs that significantly contributed to the variance explained (envfit(); p < 0.01 and R2 > 0.3) were classified at genus level, and only one genus for all ASVs pointing toward the same direction was displayed (Table S6). At 1 week, (A) the ellipses (confidence level 0.95) show Linköping and Stockholm because inclusion site also affected the bacterial community composition and Lactobacillus had different effects depending on the site.
(G–I) The abundance of L. reuteri DSM 17938 (qPCR data) significantly (envfit(); p < 0.01 and R2 > 0.3) correlated to bacterial community composition in the placebo group. Weight, length, and head circumference were adjusted for gestational age using the standard deviation score (Z score). ∗∗∗p < 0.001 with ANOSIM and p value adjustment for multiple comparisons with the method from Benjamini and Hochberg.
Among the potential confounding factors mentioned above, treatment with antibiotics at 4 weeks, but not the other factors, was significantly associated with lower richness and diversity (Table S2) as well as different bacterial community composition (β-diversity). After stratifying the data, diversity and evenness only differed significantly between the L. reuteri and placebo group in infants that received antibiotics at 4 weeks (Table S2).
Lower relative abundance of Staphylococcaceae and Enterobacteriaceae in the probiotic group during the 1st week of life
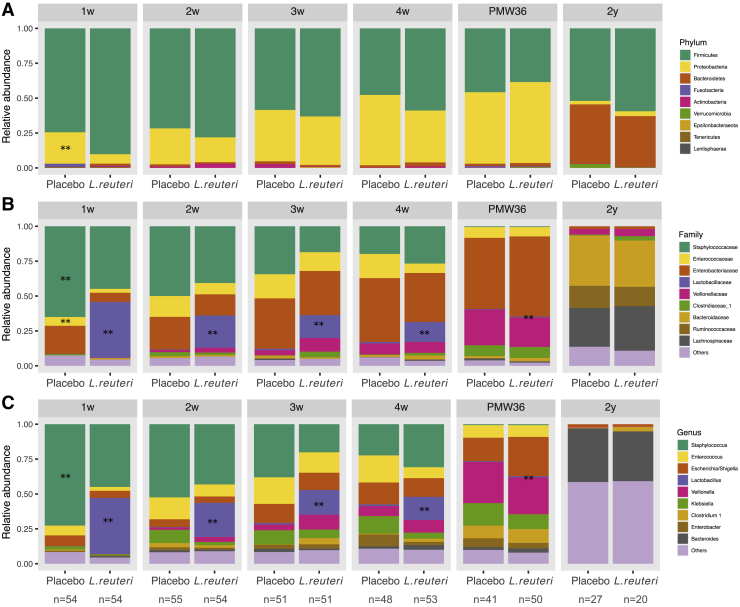
We identified a total of 491, 566, 753, 798, 778, and 1,326 bacterial amplicon sequencing variants (ASVs) at 1 week, 2 weeks, 3 weeks, 4 weeks, PMW36, and 2 years, respectively. Firmicutes and Proteobacteria were the most abundant bacterial phyla during the neonatal period (1 week to PMW36), although Bacteroidetes increased and Proteobacteria decreased in relative abundance at 2 years (Figure 4A). Staphylococcus was the dominant genus in both study groups, and Lactobacillus was the second most abundant genus in the L. reuteri group during the 1st month of life (Figure 4C), with both genera having a tendency toward decreasing in relative abundance over time. Enterococcus and Escherichia/Shigella also had a high relative abundance during the entire neonatal period, although Veillonella became dominant at PMW36. Bacteroides were present at very low levels during the neonatal period but dominated the bacterial community at 2 years in both study groups. Bifidobacteria were detected at low relative abundance (<1%) and thus belonged to the group “others” (Figure 4C).
Figure 4.
Taxonomic composition of the gut bacteria in ELBW preterm infants supplemented with L. reuteri or placebo
Relative abundance of the dominant taxa is displayed at phylum (A), family (B), and genus (C) level. At family and genus levels, the taxa with a relative abundance of <1% across all samples and time points are grouped in “others.” ∗∗ indicates taxa that significantly differed in relative abundance between the L. reuteri and placebo groups (LEfSE; p = 0.01; Table S3).
Inference of differential relative abundance between the two study groups was analyzed with linear discriminant analysis effect size (LEfSE). At 1 week, the placebo group had significantly higher relative abundance of Proteobacteria at phylum level, Staphylococcaceae and Enterobacteriaceae at family level, and Staphylococcus at genus level, although the L. reuteri group had significantly higher abundance of Lactobacillus during the 1st month of life and at PMW36 (Figure 4; Table S3). Among potential pathogens, only Staphylococcus significantly differed between the two groups. The prevalence and relative abundance of the most common pathogens found in the dataset are summarized in Table S4.
Inference of differential relative abundance between the two inclusion sites revealed that, at 1 week, the genera Escherichia/Shigella were significantly more abundant in Stockholm and, at 3 weeks, Enterococcus was significantly more abundant in Linköping. At 1 week, Escherichia/Shigella was detected in three infants (prevalence 8%) with a mean relative abundance of 1% in Linköping and in 29 infants (prevalence 41%) with a mean relative abundance of 7% in Stockholm. At 3 weeks, the prevalence of Enterococcus in Linköping was 95% (36 infants), although in Stockholm, it was 74% (52 infants), with a mean relative abundance of 27% and 9%, respectively.
High abundance of L. reuteri DSM 17938 in the probiotic group
To specifically investigate the prevalence and abundance of the supplemented L. reuteri DSM 17938 in stool, we used strain-specific quantitative PCR (qPCR). During the neonatal period (1 week to PMW36), the prevalence of L. reuteri DSM 17938 was significantly higher in the L. reuteri compared to the placebo group (Figure 5A; Fisher’s exact test; adjusted p < 0.001). At 2 years, the strain was only detected in one infant from the placebo group. The abundance of L. reuteri DSM 17938 was also significantly higher in the L. reuteri group compared to the placebo group during the neonatal period (Figure 5B), and the abundance within the L. reuteri group was lower at PMW36 compared to 1 week, 2 weeks, 3 weeks, and 4 weeks (Figure 5B). Furthermore, L. reuteri DSM 17938 abundance was significantly associated with the gut bacterial community at 2 weeks, 3 weeks, and 4 weeks in the infants receiving the probiotic (Figures 3G–3I).
Figure 5.
Prevalence and abundance of L. reuteri DSM 17938
Prevalence (A) and abundance (B) of supplemented L. reuteri DSM 17938 in L. reuteri and placebo groups at different time points.
(A) Percentage of infants with a stool sample positive for the supplemented strain.
(B) Boxplots (median with 25% and 75% percentiles and 1.5× the interquartile range) show the abundance as the number of L. reuteri DSM 17938 bacteria per 1 g wet feces. Colored dots indicate the L. reuteri DSM 17938 abundance in individual stool samples positive for the supplemented L. reuteri strain; the number of L. reuteri DSM 17938 bacteria per 1 g wet feces for infants with a L. reuteri negative stool sample was set to 1 for graphical display; (n) indicates the number of infants with a stool sample positive for supplemented L. reuteri DSM 17938; (N) indicates the total number of infants with a stool sample in the L. reuteri or placebo group at the indicated time point.
Prevalence and abundance between groups were compared using Fisher’s exact tests and Mann-Whitney U tests, respectively, and adjusted for multiple comparisons with the method from Benjamini and Hochberg. Significant differences in L. reuteri DSM 17938 abundance in the L. reuteri group across neonatal time points (1 week to PMW36) were tested for with a Kruskal-Wallis test with Dunn post hoc test and p value adjustment for multiple comparisons with the method from Benjamini and Hochberg. ∗∗∗p < 0.001; ∗∗p < 0.01.
Microbial biodiversity in relation to NEC, sepsis, and growth
Sub-analyses comparing the microbiota of NEC and sepsis cases with those of healthy controls were performed, because probiotics have been shown to reduce NEC and sepsis, and an altered bacterial composition and lower diversity have been reported to precede the onset of these diseases in preterm infants. In total, 15 out of 132 infants had NEC (eight in the placebo group and seven in the L. reuteri group), and 48 had culture-proven sepsis (23 in the placebo group and 25 in the L. reuteri group). Nine of the infants had both. Stool samples within 1 week prior to the onset of NEC were obtained from seven infants (four in the placebo group and three in the L. reuteri group) and prior to the onset of sepsis from 28 infants (14 in the placebo group and 14 in the L. reuteri group). The α-diversity and differential abundance of taxa in the NEC and sepsis cases were compared with two matched controls per index case. The selection criteria for the controls were, in a preferred order: supplementation; location; gender; gestational age; delivery mode; and treatment with antibiotics. There were no significant differences in α-diversity indices (Table S2) or relative abundance (Figure S1), with one exception: the genus Clostridium was significantly more relatively abundant in the sepsis group (LEfSE; p < 0.01).
In the original trial, the probiotic group had better head growth from birth to 4 weeks than the placebo group.13 In the present study, higher bacterial diversity at 1 week and higher richness at 2 weeks were associated with significantly better head growth rate at 4 weeks (Figures 6A and 6B) but explained only 6% and 3% of the variance, respectively. The bacterial composition at 1 week and 3 weeks was related to head growth at 4 weeks and PMW36 (R2 range = 7%–14%), as well as to weight gain at 2 weeks, 4 weeks, and PMW36 (R2 range = 6%–19%; Figures 6C and 6D). There was no correlation between the growth rate and the relative abundance of L. reuteri during the 1st month of life (data not shown). Additionally, we applied the sparse microbial causal mediation model (SparseMCMM) in order to test whether the microbial community mediated the effect of probiotic supplementation on the growth parameters, but we found no significant mediation effect (Table S5).
Figure 6.
Correlation between growth parameters and ELBW preterm infant gut microbiota composition
(A and B) Head growth until 4 weeks of life correlated to microbial diversity at 1 week (simple linear regression; p = 0.007; adjusted R2 = 0.06; A) and microbial richness at 2 weeks (simple linear regression; p = 0.035; adjusted R2 = 0.03; B).
(C and D) Head (head) and weight (weight) growth rate significantly (envfit(); p < 0.05 and R2 0.1–0.2) correlated to the microbial community composition at 1 week (C) and 3 weeks (D) of life.
Discussion
This randomized-controlled study shows that infants supplemented daily with L. reuteri DSM 17938 had increased bacterial diversity and different bacterial community composition as compared to the infants in the placebo group. These differences did not remain after the 1st month of life, i.e., at PMW36 and at the follow-up at 2 years. The prevalence of the supplemented strain was high during the neonatal period, and its abundance correlated with the infant gut bacterial composition at 2 weeks to 4 weeks. Moreover, Lactobacillus spp. dominated the gut microbiota in the probiotic group during the 1st month of life and also correlated with the bacterial composition of the probiotic group, thus likely explaining most of the observed differences. This high relative abundance of Lactobacillus, and more specifically the supplemented L. reuteri DSM 17938, during the 1st month of life in the probiotic group was probably due to a low gut bacterial load in ELBW infants during the 1st weeks of life.20,21 Moreover, the high relative abundance of Lactobacillus at 1 week might also explain why there were significant differences in the relative abundance of other bacterial taxa. The decrease in the abundance of L. reuteri DSM 17938 at PMW36 despite the ongoing probiotic supplementation was most likely explained by the maturation of the colonic microbiota and an increase in anaerobic bacteria,20 resulting in increased competition for the niche and nutrients.22 Facultative anaerobes like Streptococci and Lactobacilli are the dominating taxa in the upper part of the human intestine, although obligate anaerobes are dominating the microbiota in the colon.23 Therefore, it is possible that L. reuteri continued to be a dominating strain in the small intestine, acting on the epithelium, also in the end of the neonatal period, despite the low levels in the stool samples.
The study is consistent with previous reports of an aberrant gut microbiota in VLBW infants during the neonatal period,21 including high relative abundance of Proteobacteria, low relative abundance of Bacteroidetes at phylum level, high relative abundance of Staphylococcus, E. coli, Klebsiella, and Enterococcus at genus level, and almost no Bifidobacteria, despite that the infants were fed exclusively with breast milk until they reached a weight of at least 2,000 g. The low abundance of Bifidobacteria in the preterm infants may be due to its susceptibility to the antibiotics frequently administered to these children,24 although it also has been proposed that gut immaturity and low gestational age in itself may explain the low prevalence of Bifidobacteria in preterm infants.20 Increased relative abundance of Proteobacteria and decreased relative abundance of Firmicutes and Bacteroidetes and a lack of Propionibacterium have been reported to precede the onset of NEC.2,8 Interestingly, despite only being significant at 1 week, the relative abundance of Proteobacteria and its family Enterobacteriaceae were lower in the probiotic group. Moreover, the difference in bacterial composition (β-diversity) between the two study groups was partially explained by Klebsiella, a genus in the Enterobacteriaceae family, in the placebo group during the 1st month of life.
Importantly, the gut microbiota was assessed every week during the 1st month of life in the present study, as this is the critical time window when most cases of sepsis, NEC, and feeding intolerance have their onsets in VLBW infants.25 There are two previous randomized-controlled trials in VLBW infants that have reported analyses of the gut microbiota composition. In the ProPrems trial, supplementation with two Bifidobacterium strains and a Streptococcus strain was associated with lower abundance of Enterococcus, although the analysis did not discriminate when exactly Enterococcus was lower during the neonatal period.26 In the PiPS trial, the gut microbiota was only analyzed at PMW36 and, as in the present study, there were no differences in diversity or bacterial taxa between the two study groups at that age.27 Recently, an observational study in which infants born before gestational week 34 were supplemented with a combination of Bifidobacterium and Lactobacillus confirmed our result.28 Besides an increased relative abundance of the supplemented Bifidobacterium and Lactobacillus strains, they reported a reduction of genera within the Enterobacteriaceae family during the whole neonatal period and a weak reduction of Clostridium in the end of the neonatal period. Although being consistent with our results, the differences between the two study groups seemed to be larger than in our trial. However, the case mix (i.e., differences in selection criteria, background factors, and treatments) also differed from our trial by including infants born at a later gestational age, and more importantly, their control infants were recruited from other hospitals than the supplemented infants.
Our study showed that gut microbiota can differ between two inclusion sites, which emphasizes the importance of a prospective randomized-controlled design with a stratification by inclusion site as, for example, nutrition could differ somewhat between different clinics. Another potential confounder of the probiotic intervention was antibiotic treatment, which was very common during the first 4 weeks of life in this cohort. Rougé et al.24 suggested that antibiotic treatment hindered colonization of the probiotic strains Bifidobacterium longum BB536 and Lactobacillus rhamnosus GG in ELBW infants, leading to a lack of effect of the probiotic supplement on time to reach full enteral feeding. Also, Costeloe et al.29 described an association of antibiotic treatment with decreased colonization by probiotic Bifidobacterium breve BBG-001 and a lack of effect on morbidities in very preterm infants in the PiPS trial. In contrast, the abundance of L. reuteri DSM 17938 did not seem to be impeded by antibiotic treatment in this study, which may be explained by its intrinsic resistance against a variety of antibiotic agents, including aminoglycosides and vancomycin.30
Cross-contamination has been proposed to explain the lack of effect in probiotic trials, such as the PiPS study.29 Indeed, L. reuteri was detected in the placebo group in the present study but only rarely and at very low levels as compared to the probiotic group. Thus, it is unlikely that cross-contamination caused a false negative effect in this study.
The original trial did not find any effect of L. reuteri supplementation on the primary clinical outcome feeding tolerance, but probiotic-supplemented infants had a better growth rate of the head during the 1st month of life compared to the placebo group.13 The gut microbiota has been proven to affect growth and brain function in gnotobiotic animal models31,32 and has been associated with obesity in adults33,34 and growth in infants.35 The growth rate during the 1st month of life had a significant but weak correlation with the microbiota composition and structure but no correlation with L. reuteri. Taxonomic characterization by itself, however, is unable to reveal the functional potential of the microbiota in preterm infants, which is necessary to understand the mechanisms underlying the effects of the gut microbiome on growth and nutrition.36 Future metatranscriptomic and metabolic profiling of the microbiota in relation to growth in ELBW infants is therefore warranted.
In conclusion, daily supplementation of L. reuteri DSM 17938 in ELBW preterm infants modulated the gut bacterial composition, with increased bacterial diversity and a high abundance of the supplemented probiotic during the 1st month of life. Major effects on the other bacterial taxa were only observed during the 1st weeks of life, with lower relative abundance of Staphylococcaceae and Enterobacteriaceae in the probiotic than the placebo group. No differences in the gut microbiota composition remained at the follow-up at 2 years of age.
Limitations of study
A limitation of our study is that only stool samples were collected and analyzed, although many of the interactions between the gut microbiota and the intestinal mucosa and immune system are taking place in the small intestine. However, it would have been unethical to obtain samples invasively from ELBW preterm infants. The supplemented L. reuteri strain has been identified in intestinal biopsy specimens from ileum in adults, where it was seen to interact with the immune system by inducing a significantly higher amount of CD4-positive T lymphocytes in the ileal epithelium,37 although in vitro studies have shown L. reuteri to prime dendritic cells to produce increased levels of anti-inflammatory interleukin-10 (IL-10) and inhibit the proliferation of bystander effector T cells.38 Another limitation is that neither the original trial nor this sub-study were designed and powered to detect any significant effect on NEC or sepsis, and thus, findings from previous trials2,8 could not be confirmed in the present study. Moreover, we intentionally only included ELBW infants in this study, and thus, it limits the generalization of our findings to this patient group. A methodological limitation is that 16S rRNA gene sequencing produces compositional data and limits the analysis to relative abundances of bacterial taxa. To at least partially overcome this limitation, we applied qPCR to quantify the absolute abundance of the supplemented probiotic. Another limitation of the 16S rRNA gene sequencing data is that the taxonomical classification is limited in its accuracy to assign species.
STAR★Methods
Key Resources Table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Bacterial and virus strains | ||
| Lactobacillus reuteri DSM 17938 for probiotic use | BioGaia AB | https://www.biogaia.com/product-country/product-country-2163/ |
| Lactobacillus reuteri DSM 17938 for DNA extraction to generate the standards for qPCR runs | Stefan Roos, Swedish Department of Molecular Sciences, University of Agricultural Sciences | N/A |
| DNA mock control ATCC MSA-2002 | http://www.atcc.org/?geo_country=us | 20 Strain Even Mix Whole Cell Material (ATCC® MSA 2002TM) |
| Biological samples | ||
| Preterm infant stool samples (Table S10: metadata) | This paper | N/A |
| Critical commercial assays | ||
| QIAamp PowerFecal DNA kit (50 preps) | QIAGEN | Cat No./ID: 12830-50 |
| 16S Metagenomic Sequencing Library Preparation | Illumina | Part # 15044223 Rev. B |
| Nextera XT Index kit v 2 (96 indexes, 384 samples) | Illumina | FC-131-2001 |
| Agencourt AMPure XP, 450 mL | Beckman Coulter | A63882 |
| 2xKAPA HiFi HotStart ReadyMix | Roche | KK2601 |
| PhiX Control Kit v3 | Illumina | FC-110-3001 |
| MiSeq Reagent Kit v3 (600-cycle) | Illumina | MS-102-3003 |
| EZ1 DNA Tissue Kit | QIAGEN | Cat# 953034 |
| SsoFast™ EvaGreen® Supermix | Bio-Rad | Cat# 1725201 |
| Deposited data | ||
| Fastq.gz files from 16S rRNA gene sequencing | This paper | Accession number ENA: PRJEB3653 |
| Metadata | This paper | Table S10 |
| Oligonucleotides | ||
| 16S rRNA primers pair 341F/805R (See 16S Metagenomic Sequencing Library Preparation) | Klindworth et al.39 | N/A |
| L. reuteri 1694 gene forward primer 5¢-TTAAGGATGCAAACCCGAAC-3¢ | Romani et al.40 | N/A |
| L. reuteri 1694 gene reverse primer 5¢- CCTTGTCACCTGGAACCACT −3¢ | Romani et al.40 | N/A |
| Software and algorithms | ||
| CFX Manager™ Software version 3.1 | Bio-Rad | Cat# 1845000 |
| bbduk.sh bbmap/38.08 | Bushnell41 | https://sourceforge.net/projects/bbmap/ |
| FastQC/0.11.5 and MultiQC/1.7 | Ewels et al.42 | https://github.com/ewels/MultiQC |
| DADA2 Pipeline Tuytorial Dada2 version1.10.1 | Callahan et al.43 | https://benjjneb.github.io/dada2/tutorial.html |
| SILVA database version 132 | German Network for Bioinformatics Infrastructure | https://www.arb-silva.de/documentation/release-123/ |
| R Console 3.5.0 | The R project for Statistical Computing | https://cran.r-project.org/bin/macosx/ |
| DESeq2 R package version 1.28.2 | Love et al.44 | https://bioconductor.org/packages/release/bioc/html/DESeq2.html |
| diverse R package version 0.1.5 | Guevaraet al.45 | https://github.com/mguevara/diverse |
| phyloseq R package version 1.32.0 | McMurdie and Holmes46 | https://github.com/joey711/phyloseq |
| Other | ||
| Placebo (Maltodextrin in oil suspension) | BioGaia AB | N/A |
Resource availability
Lead contact
Further information and requests for resources and reagents should be directed to and will be fulfilled by the Lead Contact, Magalí Martí (magali.marti.genero@liu.se).
Materials availability
This study did not generate new unique reagents.
Data and code availability
The 16S rRNA dataset generated during this study is available at the European Nucleotide Archive : PRJEB36531. The code generated during this study is available at: https://github.com/magge30/PROPEL-ELBW-16S.
Infant metadata, qPCR data and ENA accession numbers are included in Table S10.
Experimental model and subject details
Human preterm infant cohort
134 infants born between gestational age 23+0 and 27+6 with a birth weight below 1,000 g were enrolled between 2012 and 2015 at two level III neonatal intensive care units (NICUs) (Astrid Lindgren Children’s Hospital, Stockholm, and Linköping University Hospital, Linköping, Sweden). See Table S1 for the background and clinical characteristics of the infants included in the study.
Human study design
The present study was part of a prospective randomized, double-blind, placebo-controlled, multi-center trial evaluating the effect of oral supplementation with the probiotic strain L. reuteri DSM 17938 in ELBW infants. The clinical outcomes have been published before13. Randomization was stratified by study center. The infants were characterized using comprehensive clinical data in a study-specific case report form from birth until PMW36 (Table 1). Weight, length and head circumference were recorded at birth, two weeks of age (2w), four weeks of age (4w), and at PMW36. In order to adjust for gestational age, the standard deviation score (z-score) for each measurement was calculated using Niklasson’s growth chart47. Growth rate was calculated as the difference in z-score between the later measurements and birth. Necrotising enterocolitis was staged according to Bell’s criteria48, and all cases of stage II or greater were recorded. A sepsis diagnosis required positive blood and/or cerebral spinal fluid culture, clinical deterioration and laboratory inflammatory response. Due to the 100% coverage of breast milk donor banks all infants were fed exclusively with breast milk until they reached a weight of at least 2,000 g. Protein and lipid fortification was based individually on analyses of the macronutrient and energy content of the breast milk given to each infant. Oral feeding started during the first day of life and increased gradually at a rate specified in clinical guidelines. Mother’s own milk and/or donor milk was analyzed for macronutrient and energy content. Targeted fortification was based on the guidelines of the European Society of Paediatric Gastroenterology, Hepatology and Nutrition (ESPGHAN)49. Breast milk fortification with bovine protein fortifier started when the enteral feeds had reached 100 mL/kg/day. The conduct complied with the principles of the International Conference on Harmonisation guidelines for Good Clinical Practice (ICH-GCP).
A total of 558 stool samples, collected from 132 infants, were included in the study. Samples were collected weekly during the first four weeks of life (1w, 2w, 3w and 4w), at PMW36, and at a follow-up at 2y (Figure 1). The samples were stored in sterile tubes at −20°C (short-term) and subsequently at −80°C until analysis. The scientist performing the microbial analyses was blinded until statistical analyses started.
Supplementation
Daily supplementation of L. reuteri DSM 17938 (1.25 × 108 bacteria/day) or placebo started within three days of age and continued until PMW36. L. reuteri DSM 17938 was provided in oil drops consisting of sunflower oil, medium chain triglyceride oil and silicon dioxide. The placebo was maltodextrin provided in an identical oil suspension and it was not possible to differentiate the placebo from the active product by smell, taste or visual appearance. The study product was administered via the gastric tube or via mouth (if the nasogastric tube had been removed) but was withheld during periods when infants were nil orally. The drops were flushed down by at least 0.3 mL breast milk after the administration in the gastric tube. The study products were provided by BioGaia AB (Stockholm, Sweden) in identical oil suspensions. The strain L. reuteri DSM 17938 has been derived from the strain L. reuteri ATCC 55730 by removing two plasmids carrying tet(W) tetracyclin and lnu(A) lincosamide resistance genes50. Originally the mother strain L. reuteri ATCC 55730 was isolated from the breast milk of a Peruvian mother. The manufacturer checked the quality of the study product regularly, and the concentration of L. reuteri was within the stipulated limits in all batches used in the trial.
Ethical approval
Written informed consent was obtained from both parents. The study was approved by the Ethics Committee for Human Research in Linköping, Sweden (Dnr 2012/28-31, Dnr 2012/433-32).
Method details
DNA extraction of infant stool samples
Total DNA was extracted from 0.10 ± 0.03 g of stool samples using the QIAamp PowerFecal DNA kit (QIAGEN, Hilden, Germany) on the QIAcube instrument (QIAGEN) according to the manufacturer’s instructions, with slight modifications: after heating, samples were disrupted for 5 min at 50 Hz with a TissueLyzer II (QIAGEN) in order to better capture Bifidobacteria51, and after subsequent centrifugation, the procedure was automatized using the QIAamp PowerFecal DNA program for stool and biosolid on a QIAcube. DNA concentrations were measured with Qubit dsDNA HS Assay kits (Thermo Fisher Scientific, Waltham, MA) according to the manufacturer’s instructions, and DNA was stored at −20°C.
16S rRNA gene sequencing
The 16S Metagenomic Sequencing Library Preparation, Preparing 16S Ribosomal RNA Gene Amplicons for the Illumina MiSeq System (Part # 15044223 Rev. B) was used to prepare the 16S RNA gene amplicons, which uses the primer pairs 341F/805R targeting the V3-V4 hypervariable region of 16S rRNA genes39 and the Nextera XT Index Kit (Illumina, San Diego, CA). The V3-V4 hypervariable region is widely applied in gut microbiota studies and considered among the least biased40, it is also recognized as a good region for a proper detection of Bifidobacteria51. The amplicon PCR protocol was modified to 30 cycles. The final pooled normalized libraries (4 nM), including a DNA mock control (ATCC MSA-2002), were diluted and denatured to 10 pM and spiked with 20% PhiX library (10 pM). A paired-end 300 bp sequencing run (600 cycles) was performed using the MiSeq platform (Illumina), with MiSeq Reagent Kit v3 chemicals.
Demultiplexed .fastq files were quality-filtered and trimmed, adaptor-trimmed and PhiX-filtered using bbduk.sh bbmap/38.0841. Reads were quality-trimmed to Q35 on the 5′ end and to Q30 on the 3′ end using Phred algorithm, sequences with lengths between 100 to 300 bp were retained. Phred scores were examined using MultiQC42. Trimmed sequences were further processed following the DADA2 Workflow with dada2 version 1.10.1 (https://benjjneb.github.io/dada2/tutorial.html 43) to generate an ASV table and assign taxonomy using a Naive Bayes classifier trained on the V3-V4 region of reference sequences (99% similarity) from SILVA version 132 (https://www.arb-silva.de). Merged reads with a length between 100 and 500 bp were kept and pseudo pooling was used for sample inference. ASVs identified as Archaea (2 ASVs), Eukaryote (119 ASVs) and Cyanobacteria (5 ASVs), as well as ASVs that were not identified at kingdom level (102 ASVs) were filtered out, as well as ASVs detected in only one sample and with less than 30 reads. After ASV filtering, we obtained a total of 33,883,590 sequencing reads, which belonged to 4,547 ASVs (Table S8). Two outliers (a 1w-sample in the L. reuteri group with 674,453 reads and a 3w-sample in the placebo group with 216,935 reads) were rarefied to the same number of reads as the sample with the third-most number of reads (134,932 reads).
Synthetic mock community
A synthetic mock microbial community (20 Strain Even Mix Whole Cell Material (ATCC® MSA-2002)) was prepared alongside the samples and it was used to determine the prevalence filtering threshold (amplicon sequences variants (ASV) detected in only one sample and with less than 30 reads) in order to remove potential contaminants (Table S7).
DNA extraction from L. reuteri cultures
To obtain L. reuteri DNA for standard curves in qPCR assays, L. reuteri DSM 17938 bacteria (courtesy of Dr Stefan Roos) were cultured anaerobically in 10 mL De Man, Rogosa, and Sharpe (MRS) broth at 36°C for 24 h, and DNA was extracted using the EZ1 DNA Tissue kit (QIAGEN). For DNA extraction, broth cultures were centrifuged at 3,000 rpm for 10 min at 4°C. Pellets were resuspended in 500 μL buffer G2, transferred to glass bead tubes, and shaken with a TissueLyzer II for 1 min at 30 Hz. Two hundred μL lysate were subjected to automatized DNA extraction using the protocol for purification of DNA from bacterial culture samples on an EZ1 Advanced XL robot (QIAGEN). DNA concentrations were measured with Qubit dsDNA HS Assay kits (Thermo Fisher Scientific, Waltham, MA) according to the manufacturer’s instructions, and DNA was stored at −20°C.
Quantitative PCR
The single-copy gene Lactobacillus reuteri unknown extracellular protein lr1694 (GenBank accession number: DQ074924.1) is specific for the supplemented probiotic L. reuteri strain used in this study52. The lr1694 gene was amplified in 20 μL qPCR reactions consisting of 2 μL of 10-fold diluted DNA, 1X SsoFast™ EvaGreen® Supermix (Bio-Rad, Hercules, CA), 300 μM of forward primer (sequence: 5′ TTAAGGATGCAAACCCGAAC 3′) and 300 μM reverse primer (sequence: 5′ CCTTGTCACCTGGAACCACT 3′). The qPCR assays were performed in CFX96™ Real-Time PCR Detection Systems (Bio-Rad) using the following program: 2 min at 98°C, 40 cycles of 5 s at 98°C and 5 s at 63°C. After each qPCR run, a melting curve analysis was conducted by gradually increasing the temperature from 65°C to 95°C in steps of 0.5°C. Serial dilutions of L. reuteri DSM 17938 DNA were used for generating standard curves with 5 × 104 to 2.5 × 101 lr1694 gene copies/μL corresponding to quantification limits of 2.3 × 104 to 4.5 × 107 bacteria/g wet feces. Similar standard curve values were obtained in all runs (n = 24) (mean (95% confidence interval)): slope = −3.51 (−3.55 – −3.47), y-intercept = 42.3 (42.0 – 42.6), efficiency = 93% (91% – 95%), and R2 = 0.996 (0.995 – 0.997). Samples were analyzed in duplicates and they were re-run if the difference in Cq values from duplicates was larger than 0.3. Non-diluted or 20-fold diluted samples were used if Cq values fell outside of the range of the standard curve. If a non-diluted sample had a Cq value larger than 35, the sample was considered as negative for L. reuteri DSM 17938. Unspecific amplification or PCR inhibition were not observed. Data was normalized to the amount of feces used for DNA isolation and expressed as L. reuteri DSM 17938 bacteria per 1 g wet feces.
Quantification and statistical analysis
Background and clinical characteristics
Continuous variables with skewed distributions were analyzed with Mann-Whitney U tests, while Student’s t tests were employed for continuous variables with normal distributions. Pearson’s chi-square test was used for categorical outcome variables. Fisher’s exact test was used when the observed frequency for any cell was less than five. Statistical analyses were performed in R Console 3.5.0.
16S rRNA gene statistical analyses
Prior to β-diversity analyses, variance stabilizing transformation (VST) was applied for normalization across samples46, using the DESeq2 package in R Console 3.5.044. Bacterial community distributions across the L. reuteri and placebo groups were displayed by non-metric multidimensional scaling (NMDS) plots, and statistically tested using the analysis of similarities (ANOSIM), with 999 permutations53. Alpha-diversity was calculated using Shannon’s diversity index, Pielou’s evenness index, and richness assessed as number of observed ASVs, using the diverse package45, and statistically tested for differences between the groups (L. reuteri versus placebo, NEC cases versus matched controls, sepsis versus matched controls) using Mann-Whitney U tests. Rarefaction tests prior to α-diversity analysis were performed, concluding that rarefaction was not needed for the analysis (Table S9). Inference of differential abundance between the study groups was performed at ASV level as well as at all the different taxonomic levels, using the Linear discriminant analysis Effect Size (LEfSE)54. Correlation between α-diversity and growth parameters was explored using simple linear regression, and co-variation between β-diversity and growth parameters as well as qPCR data was assessed by fitting the growth clinical output data onto the ordination derived from the NMDS, with the envfit() function. The microbial causal mediation effect on the growth parameters was tested using the Sparse Microbial Causal Mediation Model (SparseMCMM)55. The p values for the β-diversity analyses were corrected for false discovery rate according to Benjamini & Hochberg.
Quantitative PCR statistical analyses
Statistically significant differences in prevalence of L. reuteri DSM 17938 in the two study groups were tested for using Fisher’s exact test. Mann-Whitney U tests were performed to test for differences in L. reuteri DSM 17938 abundance in the two supplementation groups. A Kruskal-Wallis test with Dunn post hoc test was used to compare the abundance of L. reuteri DSM 17938 in L. reuteri-supplemented infants across time points. The p values for the qPCR analyses were corrected for false discovery rate according to Benjamini & Hochberg. Statistical analyses were performed in R Console 3.5.0.
Additional resources
The study is registered at ClinicalTrials.gov (ID NCT01603368).
Acknowledgments
This research was funded by the Lions Forskningsfond mot folksjukdomar (Lions LiU-2019-03041), the Swedish Research Council (grant numbers 921.2014-7060 and 2016-01715_3), the Swedish Society for Medical Research, the Swedish Society of Medicine, The Ekhaga Foundation, The Research Council for South-East Sweden, ALF grants, Region Östergötland, and BioGaia AB, Stockholm, Sweden. The computations were partially performed on resources provided by the Swedish National Infrastructure for Computing (SNIC) through Uppsala Multidisciplinary Center for Advanced Computational Science (UPPMAX) under project SNIC 2020/5-336. We thank Dr. Stefan Roos, Dr. Fredrik Ingemansson, Dr. Josefin Lundström, Dr. Anders Palm, Dr. Björn Westrup, Dr. Laura Österdahl, principal research engineer Camilla Janefjord, PhDs Ulrika Holmlund and Yeneneh Haileselassie, and the study nurses Mrs. Christina Fuxin and Mrs. Karin Jansmark for their help.
Author contributions
Study design, T.A., G.M., and E.S.-E.; funding acquisition, M.M., E.S.-E., and T.A.; sample collection, G.M., E.W., and T.A.; sequencing, qPCR, and statistical analyses, M.M. and J.E.S.; bioinformatic analyses, M.M.; overall data interpretation, M.M., J.E.S., P.D.R., M.C.J., and T.A.; writing of the manuscript, M.M.; critical review of the manuscript, J.E.S., P.D.R., E.W., G.M., E.S.-E., M.C.J., and T.A.
Declaration of interests
T.A. has received honoraria for lectures and a grant for the present trial from BioGaia AB. M.C.J. has received honoraria for lectures from BioGaia AB. E.S.-E. has received honoraria for lectures and a research grant from BioGaia AB.
Published: February 22, 2021
Footnotes
Supplemental information can be found online at https://doi.org/10.1016/j.xcrm.2021.100206.
Supplemental information
References
- 1.Norman M., Hallberg B., Abrahamsson T., Björklund L.J., Domellöf M., Farooqi A., Foyn Bruun C., Gadsbøll C., Hellström-Westas L., Ingemansson F. Association between year of birth and 1-year survival among extremely preterm infants in Sweden during 2004-2007 and 2014-2016. JAMA. 2019;321:1188–1199. doi: 10.1001/jama.2019.2021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Pammi M., Cope J., Tarr P.I., Warner B.B., Morrow A.L., Mai V., Gregory K.E., Kroll J.S., McMurtry V., Ferris M.J. Intestinal dysbiosis in preterm infants preceding necrotizing enterocolitis: a systematic review and meta-analysis. Microbiome. 2017;5:31. doi: 10.1186/s40168-017-0248-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Stewart C.J., Embleton N.D., Marrs E.C.L., Smith D.P., Fofanova T., Nelson A., Skeath T., Perry J.D., Petrosino J.F., Berrington J.E., Cummings S.P. Longitudinal development of the gut microbiome and metabolome in preterm neonates with late onset sepsis and healthy controls. Microbiome. 2017;5:75. doi: 10.1186/s40168-017-0295-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Borre Y.E., O’Keeffe G.W., Clarke G., Stanton C., Dinan T.G., Cryan J.F. Microbiota and neurodevelopmental windows: implications for brain disorders. Trends Mol. Med. 2014;20:509–518. doi: 10.1016/j.molmed.2014.05.002. [DOI] [PubMed] [Google Scholar]
- 5.Jin Y.T., Duan Y., Deng X.K., Lin J. Prevention of necrotizing enterocolitis in premature infants - an updated review. World J. Clin. Pediatr. 2019;8:23–32. doi: 10.5409/wjcp.v8.i2.23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Dong Y., Speer C.P. Late-onset neonatal sepsis: recent developments. Arch. Dis. Child. Fetal Neonatal Ed. 2015;100:F257–F263. doi: 10.1136/archdischild-2014-306213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Levy M., Kolodziejczyk A.A., Thaiss C.A., Elinav E. Dysbiosis and the immune system. Nat. Rev. Immunol. 2017;17:219–232. doi: 10.1038/nri.2017.7. [DOI] [PubMed] [Google Scholar]
- 8.Lindberg T.P., Caimano M.J., Hagadorn J.I., Bennett E.M., Maas K., Brownell E.A., Matson A.P. Preterm infant gut microbial patterns related to the development of necrotizing enterocolitis. J. Matern. Fetal Neonatal Med. 2020;33:349–358. doi: 10.1080/14767058.2018.1490719. [DOI] [PubMed] [Google Scholar]
- 9.Thomas J.P., Raine T., Reddy S., Belteki G. Probiotics for the prevention of necrotising enterocolitis in very low-birth-weight infants: a meta-analysis and systematic review. Acta Paediatr. 2017;106:1729–1741. doi: 10.1111/apa.13902. [DOI] [PubMed] [Google Scholar]
- 10.Aceti A., Maggio L., Beghetti I., Gori D., Barone G., Callegari M.L., Fantini M.P., Indrio F., Meneghin F., Morelli L., Italian Society of Neonatology. Probiotics prevent late-onset sepsis in human milk-fed, very low birth weight preterm infants: systematic review and meta-analysis. Nutrients. 2017;9:E904. doi: 10.3390/nu9080904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Cui X., Shi Y., Gao S., Xue X., Fu J. Effects of Lactobacillus reuteri DSM 17938 in preterm infants: a double-blinded randomized controlled study. Ital. J. Pediatr. 2019;45:140. doi: 10.1186/s13052-019-0716-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Indrio F., Riezzo G., Tafuri S., Ficarella M., Carlucci B., Bisceglia M., Polimeno L., Francavilla R. Probiotic supplementation in preterm: feeding intolerance and hospital cost. Nutrients. 2017;9:E965. doi: 10.3390/nu9090965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Wejryd E., Marchini G., Frimmel V., Jonsson B., Abrahamsson T. Probiotics promoted head growth in extremely low birthweight infants in a double-blind placebo-controlled trial. Acta Paediatr. 2019;108:62–69. doi: 10.1111/apa.14497. [DOI] [PubMed] [Google Scholar]
- 14.Oncel M.Y., Sari F.N., Arayici S., Guzoglu N., Erdeve O., Uras N., Oguz S.S., Dilmen U. Lactobacillus reuteri for the prevention of necrotising enterocolitis in very low birthweight infants: a randomised controlled trial. Arch. Dis. Child. Fetal Neonatal Ed. 2014;99:F110–F115. doi: 10.1136/archdischild-2013-304745. [DOI] [PubMed] [Google Scholar]
- 15.van den Akker C.H.P., van Goudoever J.B., Szajewska H., Embleton N.D., Hojsak I., Reid D., Shamir R., ESPGHAN Working Group for Probiotics, Prebiotics & Committee on Nutrition Probiotics for preterm infants: a strain-specific systematic review and network meta-analysis. J. Pediatr. Gastroenterol. Nutr. 2018;67:103–122. doi: 10.1097/MPG.0000000000001897. [DOI] [PubMed] [Google Scholar]
- 16.Dermyshi E., Wang Y., Yan C., Hong W., Qiu G., Gong X., Zhang T. The “golden age” of probiotics: a systematic review and meta-analysis of randomized and observational studies in preterm infants. Neonatology. 2017;112:9–23. doi: 10.1159/000454668. [DOI] [PubMed] [Google Scholar]
- 17.Hill C., Guarner F., Reid G., Gibson G.R., Merenstein D.J., Pot B., Morelli L., Canani R.B., Flint H.J., Salminen S. Expert consensus document. The International Scientific Association for Probiotics and Prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat. Rev. Gastroenterol. Hepatol. 2014;11:506–514. doi: 10.1038/nrgastro.2014.66. [DOI] [PubMed] [Google Scholar]
- 18.Mu Q., Tavella V.J., Luo X.M. Role of Lactobacillus reuteri in human health and diseases. Front. Microbiol. 2018;9:757. doi: 10.3389/fmicb.2018.00757. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Singh T.P., Kaur G., Kapila S., Malik R.K. Antagonistic activity of Lactobacillus reuteri strains on the adhesion characteristics of selected pathogens. Front. Microbiol. 2017;8:486. doi: 10.3389/fmicb.2017.00486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.La Rosa P.S., Warner B.B., Zhou Y., Weinstock G.M., Sodergren E., Hall-Moore C.M., Stevens H.J., Bennett W.E., Jr., Shaikh N., Linneman L.A. Patterned progression of bacterial populations in the premature infant gut. Proc. Natl. Acad. Sci. USA. 2014;111:12522–12527. doi: 10.1073/pnas.1409497111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Arboleya S., Sánchez B., Milani C., Duranti S., Solís G., Fernández N., de los Reyes-Gavilán C.G., Ventura M., Margolles A., Gueimonde M. Intestinal microbiota development in preterm neonates and effect of perinatal antibiotics. J. Pediatr. 2015;166:538–544. doi: 10.1016/j.jpeds.2014.09.041. [DOI] [PubMed] [Google Scholar]
- 22.Pereira F.C., Berry D. Microbial nutrient niches in the gut. Environ. Microbiol. 2017;19:1366–1378. doi: 10.1111/1462-2920.13659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Hayashi H., Takahashi R., Nishi T., Sakamoto M., Benno Y. Molecular analysis of jejunal, ileal, caecal and recto-sigmoidal human colonic microbiota using 16S rRNA gene libraries and terminal restriction fragment length polymorphism. J. Med. Microbiol. 2005;54:1093–1101. doi: 10.1099/jmm.0.45935-0. [DOI] [PubMed] [Google Scholar]
- 24.Rougé C., Piloquet H., Butel M.J., Berger B., Rochat F., Ferraris L., Des Robert C., Legrand A., de la Cochetière M.F., N’Guyen J.M. Oral supplementation with probiotics in very-low-birth-weight preterm infants: a randomized, double-blind, placebo-controlled trial. Am. J. Clin. Nutr. 2009;89:1828–1835. doi: 10.3945/ajcn.2008.26919. [DOI] [PubMed] [Google Scholar]
- 25.Neu J., Walker W.A. Necrotizing enterocolitis. N. Engl. J. Med. 2011;364:255–264. doi: 10.1056/NEJMra1005408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Plummer E.L., Bulach D.M., Murray G.L., Jacobs S.E., Tabrizi S.N., Garland S.M., ProPrems Study Group Gut microbiota of preterm infants supplemented with probiotics: sub-study of the ProPrems trial. BMC Microbiol. 2018;18:184. doi: 10.1186/s12866-018-1326-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Millar M., Seale J., Greenland M., Hardy P., Juszczak E., Wilks M., Panton N., Costeloe K., Wade W.G. The microbiome of infants recruited to a randomised placebo-controlled probiotic trial (PiPS trial) EBioMedicine. 2017;20:255–262. doi: 10.1016/j.ebiom.2017.05.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Alcon-Giner C., Dalby M.J., Caim S., Ketskemety J., Shaw A., Sim K., Lawson M.A.E., Kiu R., Leclaire C., Chalklen L. Microbiota supplementation with Bifidobacterium and Lactobacillus modifies the preterm infant gut microbiota and metabolome: an observational study. Cell Rep. Med. 2020;1:100077. doi: 10.1016/j.xcrm.2020.100077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Costeloe K., Hardy P., Juszczak E., Wilks M., Millar M.R., Probiotics in Preterm Infants Study Collaborative Group Bifidobacterium breve BBG-001 in very preterm infants: a randomised controlled phase 3 trial. Lancet. 2016;387:649–660. doi: 10.1016/S0140-6736(15)01027-2. [DOI] [PubMed] [Google Scholar]
- 30.Egervärn M., Danielsen M., Roos S., Lindmark H., Lindgren S. Antibiotic susceptibility profiles of Lactobacillus reuteri and Lactobacillus fermentum. J. Food Prot. 2007;70:412–418. doi: 10.4315/0362-028x-70.2.412. [DOI] [PubMed] [Google Scholar]
- 31.Ridaura V.K., Faith J.J., Rey F.E., Cheng J., Duncan A.E., Kau A.L., Griffin N.W., Lombard V., Henrissat B., Bain J.R. Gut microbiota from twins discordant for obesity modulate metabolism in mice. Science. 2013;341:1241214. doi: 10.1126/science.1241214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Diaz Heijtz R., Wang S., Anuar F., Qian Y., Björkholm B., Samuelsson A., Hibberd M.L., Forssberg H., Pettersson S. Normal gut microbiota modulates brain development and behavior. Proc. Natl. Acad. Sci. USA. 2011;108:3047–3052. doi: 10.1073/pnas.1010529108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Goodrich J.K., Waters J.L., Poole A.C., Sutter J.L., Koren O., Blekhman R., Beaumont M., Van Treuren W., Knight R., Bell J.T. Human genetics shape the gut microbiome. Cell. 2014;159:789–799. doi: 10.1016/j.cell.2014.09.053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Dao M.C., Everard A., Aron-Wisnewsky J., Sokolovska N., Prifti E., Verger E.O., Kayser B.D., Levenez F., Chilloux J., Hoyles L., MICRO-Obes Consortium Akkermansia muciniphila and improved metabolic health during a dietary intervention in obesity: relationship with gut microbiome richness and ecology. Gut. 2016;65:426–436. doi: 10.1136/gutjnl-2014-308778. [DOI] [PubMed] [Google Scholar]
- 35.White R.A., Bjørnholt J.V., Baird D.D., Midtvedt T., Harris J.R., Pagano M., Hide W., Rudi K., Moen B., Iszatt N. Novel developmental analyses identify longitudinal patterns of early gut microbiota that affect infant growth. PLoS Comput. Biol. 2013;9:e1003042. doi: 10.1371/journal.pcbi.1003042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Younge N.E., Newgard C.B., Cotten C.M., Goldberg R.N., Muehlbauer M.J., Bain J.R., Stevens R.D., O’Connell T.M., Rawls J.F., Seed P.C., Ashley P.L. Disrupted maturation of the microbiota and metabolome among extremely preterm infants with postnatal growth failure. Sci. Rep. 2019;9:8167. doi: 10.1038/s41598-019-44547-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Valeur N., Engel P., Carbajal N., Connolly E., Ladefoged K. Colonization and immunomodulation by Lactobacillus reuteri ATCC 55730 in the human gastrointestinal tract. Appl. Environ. Microbiol. 2004;70:1176–1181. doi: 10.1128/AEM.70.2.1176-1181.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Smits H.H., Engering A., van der Kleij D., de Jong E.C., Schipper K., van Capel T.M.M., Zaat B.A., Yazdanbakhsh M., Wierenga E.A., van Kooyk Y., Kapsenberg M.L. Selective probiotic bacteria induce IL-10-producing regulatory T cells in vitro by modulating dendritic cell function through dendritic cell-specific intercellular adhesion molecule 3-grabbing nonintegrin. J. Allergy Clin. Immunol. 2005;115:1260–1267. doi: 10.1016/j.jaci.2005.03.036. [DOI] [PubMed] [Google Scholar]
- 39.Klindworth A., Pruesse E., Schweer T., Peplies J., Quast C., Horn M., Glöckner F.O. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013;41:e1. doi: 10.1093/nar/gks808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Aloisio I., Quagliariello A., De Fanti S., Luiselli D., De Filippo C., Albanese D., Corvaglia L.T., Faldella G., Di Gioia D. Evaluation of the effects of intrapartum antibiotic prophylaxis on newborn intestinal microbiota using a sequencing approach targeted to multi hypervariable 16S rDNA regions. Appl. Microbiol. Biotechnol. 2016;100:5537–5546. doi: 10.1007/s00253-016-7410-2. [DOI] [PubMed] [Google Scholar]
- 41.Bushnell B. 2019. BBMap short read aligner, and other bioinformatic tools.https://sourceforge.net/projects/bbmap [Google Scholar]
- 42.Ewels P., Magnusson M., Lundin S., Käller M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics. 2016;32:3047–3048. doi: 10.1093/bioinformatics/btw354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Callahan B.J., McMurdie P.J., Rosen M.J., Han A.W., Johnson A.J.A., Holmes S.P. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods. 2016;13:581–583. doi: 10.1038/nmeth.3869. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Love M.I., Huber W., Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550. doi: 10.1186/s13059-014-0550-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Guevara M.R., Hartmann D., Mendoza M. diverse: an R package to analyze diversity in complex systems. R J. 2016;8:60–78. [Google Scholar]
- 46.McMurdie P.J., Holmes S. Waste not, want not: why rarefying microbiome data is inadmissible. PLoS Comput. Biol. 2014;10:e1003531. doi: 10.1371/journal.pcbi.1003531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Niklasson A., Albertsson-Wikland K. Continuous growth reference from 24th week of gestation to 24 months by gender. BMC Pediatr. 2008;8:8. doi: 10.1186/1471-2431-8-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Bell M.J., Ternberg J.L., Feigin R.D., Keating J.P., Marshall R., Barton L., Brotherton T. Neonatal necrotizing enterocolitis. Therapeutic decisions based upon clinical staging. Ann. Surg. 1978;187:1–7. doi: 10.1097/00000658-197801000-00001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Koletzko B., Goulet O., Hunt J., Krohn K., Shamir R., Parenteral Nutrition Guidelines Working Group. European Society for Clinical Nutrition and Metabolism. European Society of Paediatric Gastroenterology, Hepatology and Nutrition (ESPGHAN) European Society of Paediatric Research (ESPR) 1. Guidelines on paediatric parenteral nutrition of the European Society of Paediatric Gastroenterology, Hepatology and Nutrition (ESPGHAN) and the European Society for Clinical Nutrition and Metabolism (ESPEN), supported by the European Society of Paediatric Research (ESPR) J. Pediatr. Gastroenterol. Nutr. 2005;41(Suppl 2):S1–S87. doi: 10.1097/01.mpg.0000181841.07090.f4. [DOI] [PubMed] [Google Scholar]
- 50.Rosander A., Connolly E., Roos S. Removal of antibiotic resistance gene-carrying plasmids from Lactobacillus reuteri ATCC 55730 and characterization of the resulting daughter strain, L. reuteri DSM 17938. Appl. Environ. Microbiol. 2008;74:6032–6040. doi: 10.1128/AEM.00991-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Walker A.W., Martin J.C., Scott P., Parkhill J., Flint H.J., Scott K.P. 16S rRNA gene-based profiling of the human infant gut microbiota is strongly influenced by sample processing and PCR primer choice. Microbiome. 2015;3:26. doi: 10.1186/s40168-015-0087-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Romani Vestman N., Hasslöf P., Keller M.K., Granström E., Roos S., Twetman S., Stecksén-Blicks C. Lactobacillus reuteri influences regrowth of mutans streptococci after full-mouth disinfection: a double-blind, randomised controlled trial. Caries Res. 2013;47:338–345. doi: 10.1159/000347233. [DOI] [PubMed] [Google Scholar]
- 53.Clarke K.R. Non-parametric multivariate analyses of changes in community structure. Aust. J. Ecol. 1993;18:117–143. [Google Scholar]
- 54.Segata N., Izard J., Waldron L., Gevers D., Miropolsky L., Garrett W.S., Huttenhower C. Metagenomic biomarker discovery and explanation. Genome Biol. 2011;12:R60. doi: 10.1186/gb-2011-12-6-r60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Wang C., Hu J., Blaser M.J., Li H. Estimating and testing the microbial causal mediation effect with high-dimensional and compositional microbiome data. Bioinformatics. 2020;36:347–355. doi: 10.1093/bioinformatics/btz565. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The 16S rRNA dataset generated during this study is available at the European Nucleotide Archive : PRJEB36531. The code generated during this study is available at: https://github.com/magge30/PROPEL-ELBW-16S.
Infant metadata, qPCR data and ENA accession numbers are included in Table S10.