Abstract
Background
Metastatic prostate cancer is a fatal disease despite multiple new approvals in recent years. Recent studies revealed that circular RNAs (circRNAs) can be involved in cancer metastasis. Defining the role of circRNAs in prostate cancer metastasis and discovering therapeutic targets that block cancer metastasis is of great significance for the treatment of prostate cancer.
Methods
The circSOBP levels in prostate cancer (PCa) were determined by qRT‐PCR. We evaluated the function of circSOBP using a transwell assay and nude mice lung metastasis models. Immunofluorescence assay and electron microscopic assay were applied to determine the phenotypes of prostate cancer cells’ migration. We used fluorescence in situ hybridization assay to determine the localization of RNAs. Dual luciferase and rescue assays were applied to verify the interactions between circSOBP, miR‐141‐3p, MYPT1, and phosphomyosin light chain (p‐MLC2).
Results
We observed that circSOBP level was significantly lower in PCa specimens compared with adjacent noncancerous prostate specimens, and was correlated with the grade group of PCa. Overexpression of circSOBP suppressed PCa migration and invasion in vitro and metastasis in vivo. CircSOBP depletion increased migration and invasion and induced amoeboid migration of PCa cells. Mechanistically, circSOBP bound miR‐141‐3p and regulated the MYPT1/p‐MLC2 axis. Moreover, the depletion of MYPT1 reversed the inhibitory effect of circSOBP on the migration and invasion of PCa cells. Complementary intronic Alu elements induced but were not necessary for the formation of circSOBP. The nuclear export of circSOBP was mediated by URH49.
Conclusion
Our results suggest that circSOBP suppresses amoeboid migration of PCa cells and inhibits migration and invasion through sponging miR‐141‐3p and regulating the MYPT1/p‐MLC2 axis.
Keywords: amoeboid migration, circRNA, circSOBP, metastasis, prostate cancer
CircSOBP is downregulated in prostate cancer.
CircSOBP governs amoeboid migration and metastasis of prostate cancer through the miR‐141‐3p/MYPT1/p‐MLC2 axis.
The nuclear export of circSOBP is mediated by URH49.

Abbreviations
- ANP
adjacent noncancerous prostate
- AUC
area under the curve
- cDNA
complementary DNA
- circRNA
circular RNA
- DAPI
4’,6‐Diamidino‐2‐phenylindole
- EMT
epithelial‐mesenchymal transition
- FBS
fetal bovine serum
- FISH
fluorescence in situ hybridization
- gDNA
genomic DNA
- HRP
horseradish peroxidase
- miRNAs
micro RNA
- MLC2
myosin light chain 2
- MYPT1
myosin phosphatase target subunit 1
- PBS
phosphate‐buffered saline
- PCa
prostate cancer
- PFA
paraformaldehyde
- PVDF
polyvinylidene fluoride
- qRT‐PCR
quantitative real‐time polymerase chain reaction
- RNA
ribonucleic acid
- RNase R
ribonuclease R
- ROC curve
receiver operating characteristic curve
- siRNA
small interfering RNA
- SOBP
sine oculis binding protein homolog
- UAP56
56 kDa U2AF65‐associated protein
- URH49
UAP56‐related helicase, 49 kDa
- WT
wild type
- 3’‐UTR
3’‐untranslated region
1. BACKGROUND
Prostate cancer (PCa) accounts for 21% of all cancers and 10% estimated cancer death in men in 2020, 1 is one of the most frequently diagnosed malignant tumors among men worldwide. Metastatic PCa is a fatal disease despite multiple new approvals in recent years, while its treatment is getting increasingly difficult. 2 , 3 The 5‐year survival rate of PCa patients with the localized or regional disease was more than 90%, while the 5‐year survival rate of those with the distant disease was about 30%. 3 Investigating the molecular mechanism of cancer metastasis and discovering therapeutic targets that block cancer metastasis is of great significance for the cure of PCa.
The cellular and molecular mechanisms of cancer metastasis are incompletely understood. The migration of invasive cells consists of mesenchymal migration and amoeboid migration. 4 , 5 Mesenchymal migration is regulated by the epithelial‐mesenchymal transition (EMT) program. The amoeboid migration is less clear, in which cancer cells show morphological plasticity, enabling them to squeeze through interstices in the extracellular matrix. Cells undergoing amoeboid migration are accompanied by the formation of amoebic protrusions, efficiently invading adjacent tissues and spreading to distal organs through the amoeboid migration. 6 , 7 Morphology of cell pseudopods, cell roundness, and increased phosphomyosin light chain 2 (p‐MLC2) can define the amoeboid migration of cancer cells. 7 , 8 , 9
Circular RNAs (circRNAs) are single strand, covalently closed RNAs that are generated by backsplicing of pre‐mRNAs and other transcripts. 10 Flanking intronic complementary sequences, including Alu elements, could lead to backsplicing and the formation of circRNAs. 11 , 12 , 13 , 14 The majority of circRNAs that are generated from exonic sequences (exonic circRNAs) are located in the cytoplasm, although the backsplicing of exons occurred in the nucleus. 15 A previous study identified key factors that mediated the nuclear export of circRNAs using RNAi screening. 16 This work suggested that the depletion of 56 kDa U2AF65‐associated protein (UAP56) led to the accumulation of long (>1200 nucleotides) circRNAs in the nucleus, while the depletion of UAP56‐related helicase, 49 kDa (URH49) caused short (<400 nucleotides) circRNAs to be enriched in the nucleus. CircRNAs are initially proposed to be “noise” generated by aberrant splicing. 17 However, recent progress has uncovered the biogenesis and biological function of circRNAs in healthy tissues and disease. 14 , 18 Recent studies revealed that circRNAs can be involved in cancer metastasis. 19 , 20 CircRNAs regulate cancer metastasis through interacting with EMT transcription factors, 21 , 22 TGF‐β‐related EMT, 23 Wnt pathway, 24 , 25 cell adhesion molecules, 26 and tumor microenvironment. 18 However, circRNAs have not been investigated in regulating the amoeboid migration in cancer cells.
In the present study, we characterized a new circRNA in PCa. The novel circRNA is derived from the exon 2 and exon 3 of the sine oculis binding protein homolog (SOBP) gene, thereby being named circSOBP (genomic location: chr6: 107824860–107827631; circBase 27 ID: hsa_circ_0001633). We investigated the function of circSOBP to the amoeboid migration of PCa through in vitro experiments. We also examined whether the loss of circSOBP induced amoeboid migration by sponging miR‐141‐3p and regulating myosin phosphatase target subunit 1 (MYPT1)/p‐MLC2 axis in PCa.
2. MATERIALS AND METHODS
2.1. Patients and sample preparations
We acquired PCa and adjacent noncancerous prostate (ANP) tissue specimens from 56 patients during radical prostatectomy at Jinshan Hospital, Fudan University from February 2017 to March 2018. The diagnosis was validated by two independent pathologists. All tissues were stored at −80°C immediately after surgical removal. The Ethics Committee of Jinshan Hospital, Fudan University approved the present study. We obtained written consent from each patient.
2.2. Cell lines
The HEK293T, RWPE‐1, DU145, PC‐3, LNCap, and 22Rv1 cell lines were obtained from FuHeng Cell Center (FuHeng, Shanghai, China). DU145 and HEK293T were maintained in DMEM‐4.5 g/L glucose (Gibco, Suzhou, Jiangsu, China). PC‐3 was cultured in DMEM/F12 (HyClone, Logan, Utah, USA). LNCap and 22Rv1 were maintained in RPMI‐1640 (Gibco). All media were supplemented with 10% fetal bovine serum (FBS) (BI, Beit Haemek, Israel), 100 U/mL penicillin, and 100 μg/mL streptomycin. RWPE‐1 was cultured in Prostate Epithelial Cell Medium (ScienCell, San Diego, CA, USA). All cell lines were cultured at 37°C containing 5% v/v CO2.
2.3. RNA interference (RNAi)
An appropriate number of cells were planted 24 h before transfection so that the monolayer cell density reached 60% at the time of transfection. Scramble small interfering RNA (siRNA) and specific siRNAs (synthesized by GenePharma, Shanghai, China) were transfected using X‐tremeGENE siRNA Transfection Reagent (Roche, Mannheim, Germany). The siRNAs were transfected at 50 pmol/mL. Sequences of the siRNAs were available in Supporting information Table S1.
2.4. CircRNA expressing lentivirus production and infection
The circSOBP expressing lentiviral vector pLCDH‐cir was obtained from GeneSeed (Guangzhou, Guangdong, China). We obtained the pSPAX2 and pMD2.G plasmids from Hanbio (Shanghai, China). For lentiviral production, we transfected pLCDH‐cir, pSPAX2, pMD2.G plasmids into HEK293T cells with LipoD293 DNA transfection reagent (SignaGene Laboratories, Rockville, Maryland, USA) following the manufacturer's protocol. We collected the viral supernatant at 1, 2, and 3 days after transfection and removed the cell debris by centrifuging at 1000 g for 5 min.
For lentiviral transduction, cells should be seeded on a 24‐well plate 12 to 24 h before infection so that the monolayer cell density reached 30‐50% at infection. The culture medium was replaced with a 1:1 diluted viral supernatant and cultured overnight. For optimal infection, cells were infected with 10 μg/mL hexadimethrine bromide. After 72 h infection, cells were cultured with 3 μg/mL puromycin to kill noninfected cells.
2.5. Isolation of nuclear and cytoplasmic RNA, total RNA, and micro RNA (miRNA) extraction, and quantitative real‐time polymerase chain reaction (qRT‐PCR)
The nuclear and cytoplasmic RNA was isolated and purified using Cytoplasmic & Nuclear RNA Purification Kit (Norgen Biotek, Thorold, Ontario, Canada). Total RNA was extracted using RNA‐Quick Purification Kit (Yishan, Shanghai, China). Micro RNA (miRNAs) were extracted and purified using miRNA Isolation Kit with Spin Column (Beyotime, Shanghai, China). RNA samples were quantified and qualified using NanoDrop (Thermo Fisher Scientific, Waltham, MA, USA). For reverse transcription of total RNA, 500 ng RNA was reverse transcribed in a 10 μL system using PrimeScript RT Master Mix (Takara, Shiga, Japan). Reverse transcription of miRNAs was conducted using Mir‐X miRNA First‐Strand Synthesis Kit (Takara). Complementary DNA (cDNA) was diluted with diethylpyrocarbonate‐treated water. PCR was conducted in triplicate on the ABI 7300 Real‐Time PCR system (Thermo Fisher Scientific) using BeyoFast SYBR Green qPCR Mix (Beyotime) following the manufacturer's protocol. The cycling parameters were: 95°C for 2 min initially, 40 cycles of 95°C for 15 s followed by 60°C for 30 s. We analyzed the PCR results using the ΔΔCT method. 28 Sequences of primers are available in Supporting information Table S2.
2.6. Cell viability, colony formation, and apoptosis assay
We planted an appropriate number of cells in 96‐well plates and measured viability using Cell Counting Kit‐8 (Dojindo, Kumamoto, Japan) every 24 h. All measurements were performed at least three times.
Colony formation was determined by seeding 1000 cells in a 6‐well plate. Ten days after cultivation, cells were fixed using 4% paraformaldehyde (PFA) for 15 min and stained using the 1% crystal violet solution for another 30 min. Finally, the colonies were photographed and counted.
Apoptosis was analyzed using One‐Step TUNEL Apoptosis Assay Kit (Beyotime). Images were photographed using a fluorescence inverted microscope (Olympus, Tokyo, Japan).
2.7. Migration and invasion assays
Transwell permeable supports with 8 μm pore (Corning, New York City, USA) and 24‐well plates were used to conduct migration and invasion assays.
For invasion assay, the upper chambers were coated using diluted Matrigel (Corning) at 37°C for 4 h before seeding cells. For the migration assay, the upper chambers were not coated. An appropriate number of cells were seeded with an FBS‐free medium in the upper chamber of the Transwell permeable support. For DU145, we seeded 6 × 104 cells/well for “NC versus circSOBP” groups and 3 × 104 cells/well for “si‐NC versus si‐circSOBP” groups, respectively. For PC‐3, we seeded 8 × 104 cells/well for “NC versus circSOBP” groups and 6 × 104 cells/well for “si‐NC versus si‐circSOBP” groups, respectively. The lower chamber was filled with 400 μL complete medium supplied with 100 μL FBS. Appropriate subsequent culture time varies from cell lines (24 h for DU145 and 48 h for PC‐3). The migrated cells were fixed using 4% PFA followed by staining using 1% crystal violet solution. We counted the migrated cells from three random fields.
2.8. Generation of a nude mouse model
Briefly, 1 × 107 DU145 cells stably overexpressing circSOBP and its control cells were injected into the tail vein of nude mice (n = 10 per group). Eight weeks after injection, the nude mice were sacrificed and infused with formalin. Subsequently, the lungs of the nude mice were resected and fixed with formalin. After counting the metastatic nodules, we embedded the lungs in paraffin and made paraffin sections. Finally, the paraffin sections were stained using hematoxylin and eosin staining.
2.9. Dual‐luciferase assay
The sequences of the wild‐type circSOBP and MYPT1 3’‐untranslated region (UTR), which were predicted to target miR‐141‐3p or their mutants, were synthesized and cloned into pmirGLO vector (Promega, Madison, Wisconsin, USA) by Genewiz (Shanghai, China) between the NheI and XhoI restriction sites. PC‐3 cells were transfected using X‐tremeGENE siRNA Transfection Reagent (Roche). The pmirGLO plasmid and miRNA mimic were transfected at 1 μg/mL and 50 pmol/mL, respectively. Luciferase activity was tested 48 h post‐transfection using Dual‐Lumi Luciferase Reporter Gene Assay Kit (Beyotime).
2.10. Western blot
We extracted total protein using SDS lysis buffer (Beyotime) supplemented with protease and phosphatase inhibitor cocktail (Beyotime). Total proteins were quantified using the Enhanced BCA Protein Assay Kit (Beyotime). Protein (30 μg per lane) was loaded into 12% SDS‐PAGE gel and run at 120 V for 2 h. Proteins were transferred to polyvinylidene fluoride (PVDF) membranes (Millipore, Billerica, Massachusetts, USA) at 250 mA for 1 h. PVDF membranes containing proteins were blocked by QuickBlock Blocking Buffer for Western Blot (Beyotime) at room temperature for 20 min and cut horizontally to examine multiple proteins of different sizes. Then, membranes were incubated with diluted primary antibodies (Supporting information Table S3) at 4°C overnight, followed by washing for three times. Membranes were subsequently soaked with horseradish peroxidase (HRP)‐conjugated secondary antibodies at room temperature for 1 h. Finally, blots were visualized using Immobilon Western Chemiluminescent HRP Substrate (Millipore).
2.11. Immunofluorescence staining
In order to observe the amoeboid features of cells, a 96‐well plate was coated with 1:20 diluted Matrigel Matrix (Corning) and incubated at 37°C overnight. Subsequently, cells were plated and cultured overnight. Cells were fixed for 15 min using 4% PFA followed by permeabilization using phosphate‐buffered saline (PBS) supplemented with 0.1% Triton X‐100 for 15 min. Subsequently, cells were blocked using PBS supplemented with 5% bovine serum albumin and 0.1% Tween20 for 1 h at room temperature. After blocking, cells were incubated with a primary antibody (Supporting information Table S3) at 4°C overnight. Then the cells were incubated with Alexa Fluor 555‐labeled secondary antibody at room temperature for 1 h. F‐actin was stained using iFluor 488‐conjugated phalloidin (Yeasen, Shanghai, China). 4’,6‐Diamidino‐2‐phenylindole (DAPI) was used to stain the nuclei. All pictures were photographed using a fluorescence inverted microscope (Olympus).
2.12. RNA fluorescence in situ hybridization (RNA‐FISH)
RNA‐ fluorescence in situ hybridization (FISH) assay was conducted to determine the subcellular location of circSOBP. RNA‐FISH kit and a mix of three 5’‐Cy3 labeled probes that targeted the backsplice site of circSOBP were obtained from GenePharma (Shanghai, China). We carried out the hybridization according to the manufacturer's protocol. The concentration of the probe mix was 8 μmol/L during hybridization. All pictures were obtained using a fluorescence inverted microscope (Olympus). Sequences of circSOBP probes are listed in Supporting information Table S4.
2.13. Cell roundness index calculation
The cell roundness index between 0.0 and 1.0 was calculated by dividing the shortest diameter by the longest diameter of each cell. 9 , 29 The roundness index of a perfectly round cell is 1.0.
2.14. Statistical analyses
We conducted all experiments at least three times. We performed statistical analyses using SPSS Statistics (IBM, Chicago, IL, USA) or Prism 8 (GraphPad Software, San Diego, CA, USA). Student's t‐test, one‐way ANOVA, chi‐squared test, Fisher's exact test, Mann–Whitney U test, and Wilcoxon matched‐pairs signed‐rank test were used for comparisons, as appropriate. The levels of significance were set at: *P < .05, **P < .01, and ***P < .001.
3. RESULTS
3.1. Characterization of circSOBP in PCa
To characterize PCa‐related circRNAs, our circRNA microarray across five pairs of ANP tissues and PCa tissues showed that a series of circRNAs were differentially expressed. 30 CircSOBP was chosen for the present study, the reasons were: (i) The circRNA microarray showed that circSOBP (genomic location: chr6: 107824860–107827631; circBase ID: hsa_circ_0001633) 27 was downregulated in PCa tissues. (ii) CircSOBP was of relatively high abundance, ranked 4th in PCa and 6th in ANP tissues among the 24 downregulated circRNAs (Supporting information Figure S1A); (iii) the biological function of circSOBP remained unknown.
CircSOBP is derived from exon 2 and exon 3 of the SOBP gene, thereby being named circSOBP (Figure 1A). The full length of circSOBP is shown in Supporting information Figure S1B. We designed divergent primers that amplified the backsplicing site of circSOBP and convergent primers that amplified the linear part of circSOBP as well as the SOBP gene (Figure 1A). The PCR results showed that the backsplicing site of circSOBP was only detected in cDNA but not in genomic DNA (gDNA), while the linear part of circSOBP could be detected in both cDNA and gDNA (Figure 1B,C), indicating that the sequence containing the backsplice site does not exist in the genome. In addition, circSOBP was resistant to ribonuclease R (RNase R), a 3’‐5’ exonuclease that could only digest linear transcripts but not circRNAs, indicating that circSOBP was circular (Figure 1D,E). Sanger sequencing confirmed the backsplicing site (Figure 1F). FISH (Figure 1G) indicated that circSOBP was mainly distributed in the cytoplasm. Isolation of subcellular RNAs followed by qRT‐PCR showed that circSOBP was mainly distributed in the cytoplasm (Figure 1H,I).
FIGURE 1.
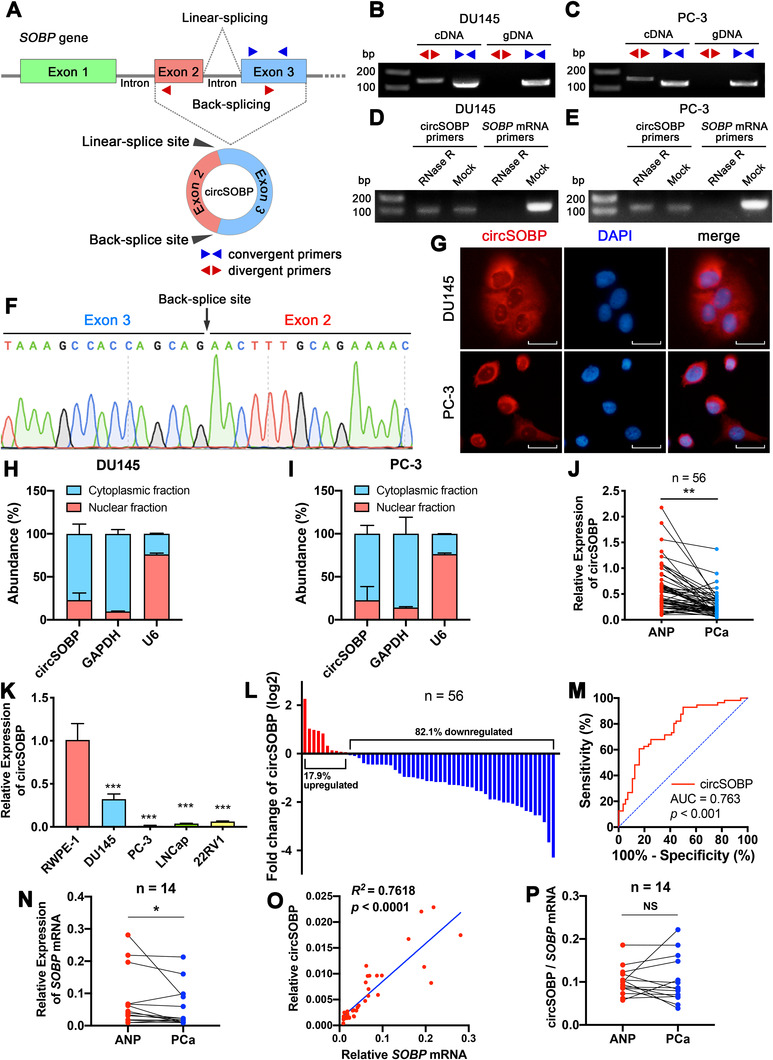
The characteristics of the circular RNA circSOBP. (A) Schematic illustration of the origin of circSOBP. Blue triangles indicate the convergent primers that amplify the linear sequence of circSOBP. Red triangles indicate the divergent primers that amplify the backsplice site of circSOBP. (B) and (C) Divergent and convergent primers were used to detect circSOBP by PCR in cDNA and gDNA of DU145 and PC‐3 cell lines. (D) and (E) CircSOBP and SOBP mRNA were detected using PCR in RNase R treated total RNAs of DU145 and PC‐3 cell lines. (F) Sanger sequencing of the backsplicing site of circSOBP. (G) The FISH assay was used to detect the subcellular localization of circSOBP (red), combined with nuclei staining using DAPI (blue). Scale bar, 30 μm. (H) and (I) Abundance of circSOBP in the nuclear and cytoplasmic fractions of DU145 and PC‐3 cells, analyzed using qRT‐PCR. GAPDH mRNA, and U6 acted as a positive control. The data are presented as the mean ± SD. (J) Expression of circSOBP in ANP and PCa tissues of 56 patients was analyzed by qRT‐PCR. **P < .01, Wilcoxon matched‐pairs signed‐rank test, n = 56. (K) Expression of circSOBP in prostate epithelial and PCa cell lines was analyzed by qRT‐PCR. The data are presented as the mean ± SD. ***P < .001 versus RWPE‐1 cells, one‐way ANOVA and Dunnett's multiple comparisons test, n = 3. (L) The percentage of upregulated or downregulated expression of circSOBP in 56 patients. (M) The ROC curve in distinguishing PCa and ANP tissues by the expression of circSOBP. (N) Expression of SOBP mRNA in ANP and PCa tissues of 14 patients was analyzed by qRT‐PCR. * P < .05, Wilcoxon matched‐pairs signed‐rank test, n = 14. (O) Scatter plot of the correlation between the expression of circSOBP and SOBP mRNA. Linear regression model, R 2 = 0.7618, P < .0001. (P) Ratio of the expression of circSOBP and SOBP mRNA in ANP and PCa tissues of 14 patients. Paired t‐test, n = 14. NS, not significant
3.2. Expression of circSOBP in PCa
Specific divergent primers were used to detect circSOBP expression in PCa cells and tissue specimens. We found circSOBP level was lower in PCa tissues (Figure 1J) and PCa cell lines (Figure 1K) than that in ANP tissues and prostate epithelial cell line RWPE‐1, respectively. Among 56 paired tissues, circSOBP expression was low in 46 (82.1%) PCa tissues compared to the paired ANP tissues (Figure 1L). The receiver operating characteristic (ROC) curve indicated that the area under the curve (AUC) of circSOBP was 0.763 (cut‐off value = 0.249, sensitivity = 0.839, 1–specificity = 0.393) (Figure 1M), indicating that circSOBP was a promising biomarker for distinguishing PCa tissues.
Detailed PCa patients’ characteristics are shown in Table 1. PCa patients were divided into PCa < ANP and PCa > ANP groups based on the relative expression of circSOBP of their PCa and ANP tissues. As shown in Table 1, circSOBP levels were significantly associated with PCa patients’ Gleason score (P = 0.0292) and grade group (P = .0118).
TABLE 1.
Correlation between circSOBP expression and clinicopathological characteristics in PCa
| Expression of circSOBP | |||
|---|---|---|---|
| Parameters | PCa < ANP (n = 46) | PCa > ANP (n = 10) | P‐value |
| Age | .3150 | ||
| ≤67 | 19 | 6 | |
| >67 | 27 | 4 | |
| PSA (ng/mL) | .7287 | ||
| ≤14.085 | 22 | 6 | |
| >14.085 | 24 | 4 | |
| Gleason score | .0292 | ||
| <7 | 0 | 2 | |
| ≥7 | 46 | 8 | |
| Grade group | .0118 | ||
| 1 | 0 | 2 | |
| 2 | 12 | 2 | |
| 3 | 10 | 4 | |
| 4 | 10 | 0 | |
| 5 | 14 | 2 | |
| pT stage | .8706 | ||
| 2 | 20 | 4 | |
| 3a | 18 | 3 | |
| 3b | 8 | 3 | |
| N stage | .6691 | ||
| 0 | 39 | 9 | |
| 1 | 7 | 1 | |
| Stage group | .9121 | ||
| I + II | 13 | 3 | |
| III + IV | 33 | 7 | |
| Nerve invasion | .7069 | ||
| Yes | 26 | 5 | |
| No | 20 | 5 | |
ANP, adjacent noncancerous prostate; PCa, prostate cancer; PSA, prostate‐specific antigen.
In addition, we determined the correlation between circSOBP and SOBP mRNA levels in PCa. The expression of SOBP mRNA was lower in PCa tissue specimens than that in ANP (Figure 1N). The linear regression model suggested that the increased expression of circSOBP was associated with the increased SOBP mRNA (R 2 = 0.7618, P < .0001)(Figure 1O). However, there is no difference in the ratio of circSOBP and SOBP mRNA in ANP and PCa (Figure 1P).
3.3. CircSOBP modulation alters PCa migration and invasion in vitro and metastasis in vivo
Next, circSOBP's biological functions were determined. We constructed DU145 (Supporting information Figure S2A) and PC‐3 (Supporting information Figure S2B) cells stably overexpressing circSOBP using lentiviral transduction. CircSOBP expression was significantly overexpressed (Figure 2A), without boosting the expression of SOBP mRNA (Figure 2B). The upregulation of circSOBP did not affect cell growth (Figure 2C,D) but inhibited migration and invasion (Figure 2E,F).
FIGURE 2.
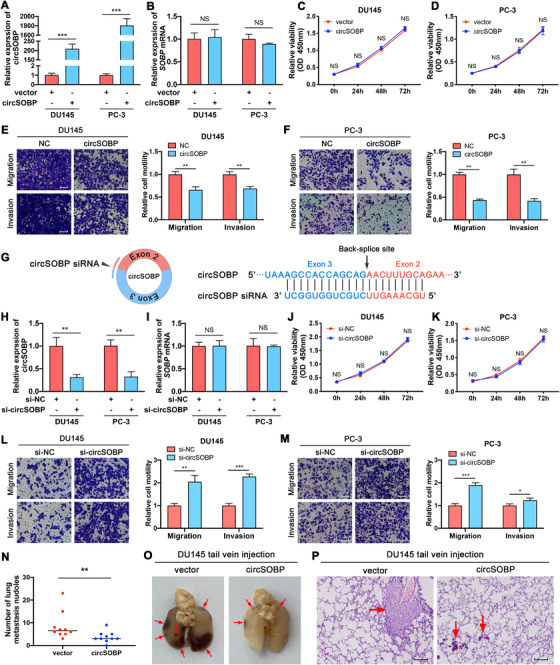
Effects of forced circSOBP expression on viability, migration, and invasion of PCa cells. (A) and (B) Expression of circSOBP and SOBP mRNA in circSOBP‐overexpressing DU145 and PC‐3 cells analyzed by qRT‐PCR. The data are presented as the mean ± SD. ***P < .001, Student's t‐test, n = 3. (C) and (D) Effect of overexpression of circSOBP on cell viability of DU145 and PC‐3 cells. The data are presented as the mean ± SD. Student's t‐test, n = 3. (E) and (F) Effect of overexpression of circSOBP on migration and invasion of DU145 and PC‐3 cells. Scale bar, 100 μm. The data are presented as the mean ± SD. **P < .01, Student's t‐test, n = 3. (G) Schematic illustration of the specific siRNA targeting the backsplicing site of circSOBP. (H) and (I) Expression of circSOBP and SOBP mRNA in circSOBP‐depleted DU145 and PC‐3 cells analyzed by qRT‐PCR. The data are presented as the mean ± SD. **P < .01, Student's t‐test, n = 3. (J) and (K) Effect of circSOBP depletion on cell viability of DU145 and PC‐3 cells. The data are presented as the mean ± SD. Student's t‐test, n = 3. (L) and (M) Effect of circSOBP depletion on migration and invasion of DU145 and PC‐3 cells. Scale bar, 100 μm. The data are presented as the mean ± SD. *P < .05, **P < .01, ***P < .001, Student's t‐test, n = 3. (N) Effect of circSOBP overexpression on the numbers of lung metastatic nodules of nude mice. Black lines in the middle of the dots depict the median. **P < .01, Mann–Whitney test, n = 10. (O) Metastatic nodules in the mouse lungs. Red arrows indicate the metastatic nodules. (P) Hematoxylin‐eosin staining of the metastatic nodules. Scale bar, 50 μm. Red arrows indicate the metastatic nodules. NC, normal control. NS, not significant
We designed siRNA targeting the backsplicing site of circSOBP (Figure 2G), which efficiently depleted circSOBP in DU145 and PC‐3 cell lines (Figure 2H), without altering the expression of SOBP mRNA (Figure 2I). CircSOBP depletion did not affect cell growth (Figure 2J,K). However, migration and invasion of DU145 (Figure 2L) and PC‐3 (Figure 2M) cell lines were promoted after the depletion of circSOBP expression.
We used the mouse lung metastasis model to investigate the biological function of circSOBP in vivo. DU145 cells stably overexpressing circSOBP formed more lung metastatic nodule than the control cells (Figure 2N). Moreover, lung metastases formed by DU145 cells stably overexpressing circSOBP were larger than those formed by control cells (Figure 2O,P).
Furthermore, we explored the effects of overexpressing or depleting circSOBP on apoptosis and colony formation of PCa cells. TUNEL staining was used to detect apoptosis. CircSOBP modulation did not induce apoptosis of DU145 (Supporting iformation Figure S2C) and PC‐3 cells (Supporting information Figure S2D). Similarly, overexpression or depletion of circSOBP did not alter colony formation of PCa cells (Supporting information Figure S2E‐G).
3.4. Depletion of circSOBP induces amoeboid migration of PCa cells
It is known that the mesenchymal migration of cancer cells regulated by EMT is a key process in regulating metastasis of cancer. Given circSOBP modulation alters migration and invasion of PCa cells, we assessed whether circSOBP could regulate EMT by detecting E‐cadherin and vimentin. However, overexpression of circSOBP did not alter the expression of E‐cadherin or vimentin (Supporting information Figure S3A‐E), indicating that circSOBP could not regulate the process of EMT. In addition to EMT, cancer cells can also metastasize by amoeboid migration. Therefore, we further determined whether circSOBP could alter the process of amoeboid migration of PCa cells.
Next, we attempted to induce amoeboid migration of PCa cells by depleting circSOBP using RNA interference. The migration of PCa cells on the Matrigel matrix includes mesenchymal migration and amoeboid migration. We observed five migration phenotypes of PCa cells using a phase‐contrast microscope or fluorescence assay: cells with blebs, stable bleb, lamellipodia, filopodia, or stable adhesion (Figure 3A). Among them, cells with blebs and stable bleb are phenotypes of amoeboid migration, while cells with lamellipodia and filopodia are phenotypes of mesenchymal migration. 7 , 9 , 31 Stable adhesion refers to cells that are stationary and morphologically unchanging. The morphology of cell pseudopods was also identified by electron microscopy (Figure 3B). The incidence of blebs and stable blebs was more predominant in circSOBP depleted DU145 (Figure 3C) and PC‐3 (Figure 3D) cells. In addition, roundness, morphology, and increased p‐MLC2 are characteristics of amoeboid migration. 8 , 9 We found that circSOBP depleted cells were rounder by calculating the roundness index (Figure 3E,F). The morphology and p‐MLC2 expression of circSOBP‐depleted DU145 and PC‐3 cells cultured on Matrigel matrix were analyzed using immunofluorescence assay (Figure 3G), indicating that p‐MLC2 was increased in circSOBP depleted cells (Figure 3H,I).
FIGURE 3.
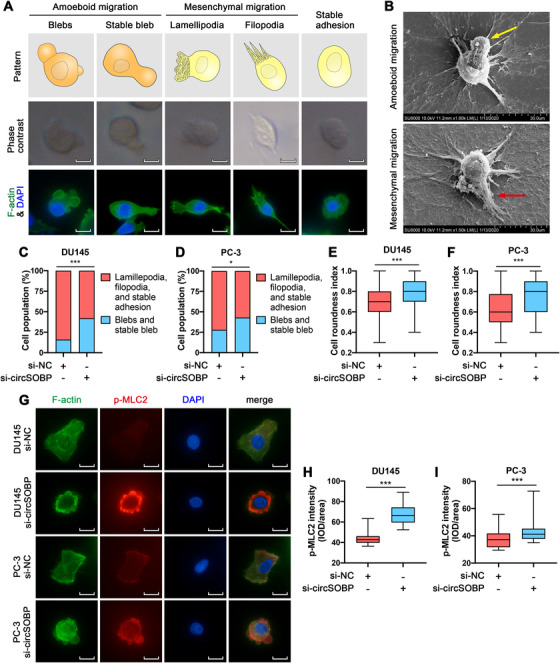
Depletion of circSOBP induces amoeboid features of PCa cells. (A) The morphology of cells undergoing mesenchymal migration or amoeboid migration on the Matrigel matrix. Scale bar, 20 μm. (B) Electron microscopic images of cells undergoing mesenchymal and amoeboid migration on the Matrigel matrix. The yellow arrow indicates a stable bleb. The red arrow indicates filopodia. Scale bar, 30 μm. (C) and (D) The percentage of circSOBP‐depleted DU145 and PC‐3 cells undergoing different phenotypes of migration on the Matrigel matrix compared with control cells. *P < .05, ***P < .001, Fisher's exact test. (E) and (F) Cell roundness of circSOBP‐depleted DU145 and PC‐3 cells. Boxplots depict median, 25th and 75th percentile, min‐max whiskers. ***P < .001, Mann–Whitney test, n = 3. (G) The morphology of circSOBP‐depleted DU145 and PC‐3 cells analyzed using immunofluorescence assay, representative images. Scale bar, 20 μm. (H) and (I) Quantification of p‐MLC2 expression from immunofluorescence assay. Boxplots depict median, 25th and 75th percentile, min‐max whiskers. ***P < .001, Mann–Whitney test, n = 3. NC, normal control
3.5. CircSOBP regulates MYPT1/p‐MLC2 axis by sponging miR‐141‐3p
We further explored the mechanism by which circSOBP regulates the amoeboid migration of PCa. Previous studies have revealed that circRNAs could sponge miRNAs and abrogate their functions. 14 , 32 We predicted six potential miRNAs sponged by circSOBP using RegRNA 2.0 database. 33 Next, we used a dual‐luciferase assay to determine the interaction between circSOBP and miRNAs, showing that circSOBP could bind miR‐141‐3p (Figure 4A). However, circSOBP overexpression did not alter the expression of the predicted miRNAs (Supporting information Figure S4). MiR‐141‐3p was upregulated in PCa (n = 52) compared with normal specimens (n = 495) (Figure 4B). RNA FISH assay indicated the colocalization of circSOBP and miR‐141‐3p in DU145 and PC‐3 cells (Figure 4C). To further verify whether circSOBP could sponge miR‐141‐3p, dual‐luciferase plasmids with wild‐type and mutant circSOBP sequences were constructed (Figure 4D). MiR‐141‐3p was able to suppress the luciferase activity in cells with wild‐type circSOBP but not in the cells with mutant circSOBP (Figure 4E).
FIGURE 4.
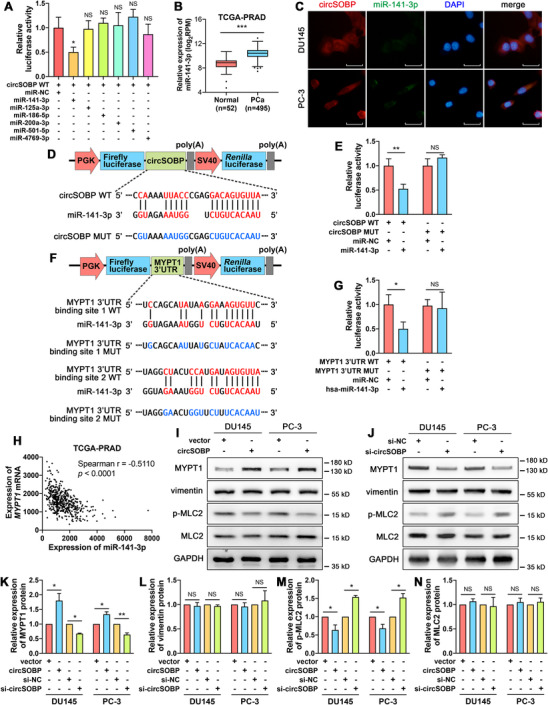
CircSOBP regulates the MYPT1/p‐MLC2 axis by sponging miR‐141‐3p. (A) Dual‐luciferase assay was used to identify the direct interaction between circSOBP and six predicted miRNAs. The data are presented as the mean ± SD. *P < .05 versus miR‐NC, one‐way ANOVA and Dunnett's multiple comparisons test, n = 3. (B) Expression of miR‐141‐3p in PCa and normal tissues in TCGA database. Boxplot depicts median, 25th and 75th percentile, min‐max whiskers. ***P < .001, Mann–Whitney test. (C) The FISH assay was used to identify the subcellular colocalization of circSOBP (red) and miR‐141‐3p (green), combined with nuclei staining using DAPI (blue). Scale bar, 30 μm. (D) Wild‐type and mutated binding site of miR‐141‐3p in circSOBP. (E) Dual‐luciferase assay was used to identify the direct interaction between circSOBP and miR‐141‐3p. The data are presented as the mean ± SD. **P < .01, Student's t‐test, n = 3. (F) Binding sites of miR‐141‐3p and their mutants in 3’UTR of MYPT1 mRNA. (G) Dual‐luciferase assay was used to identify the direct interaction between the 3’‐UTR of MYPT1 mRNA and miR‐141‐3p. The data are presented as the mean ± SD. *P < .05, Student's t‐test, n = 3. (H) Correlation between the expression of miR‐141‐3p and MYPT1 mRNA in the TCGA‐PRAD database. (I) Effect of circSOBP overexpression on the expression of related proteins of DU145 and PC‐3 cells. (J) Effect of circSOBP depletion on the expression of related proteins of DU145 and PC‐3 cells. (K)‐(N) Densimetric analysis of the blots in (I) and (J), n = 3 independent experiments, GAPDH was used as the loading control. The data are presented as the mean ± SD. *P < .05, **P < .01, Student's t‐test. MUT, mutant. NC, normal control. NS, not significant. WT, wild type
We predicted MYPT1 as the downstream target of circSOBP/miR‐141‐3p, the reasons were as follows: (i) miR‐141‐3p targeted 3’UTR of MYPT1 mRNA in accordance with the prediction of the ENCORI database (http://starbase.sysu.edu.cn), which were derived from CLIP‐seq data 34 ; (ii) MYPT1 was downregulated in PCa according to the data from the Cancer Genome Atlas (TCGA) Research Network (https://www.cancer.gov/tcga), which is consistent with predicted expression trend of our study; (iii) cells lacking MYPT1 exhibit a hyperblebbing phenotype, which lead to amoeboid behaviors 35 ; (iv) amoeboid migration requires a high level of p‐MLC2, 8 , 9 while MYPT1 mediates dephosphorylation of p‐MLC2. 36 , 37 , 38
Dual‐luciferase plasmids with wild‐type and mutant MYPT1 mRNA 3’UTR were used to verify the interaction between miR‐141‐3p and MYPT1 3’UTR (Figure 4F). We confirmed that miR‐141‐3p could bind to MYPT1 3’UTR (Figure 4G). A negative correlation was found between miR‐141‐3p and MYPT1 mRNA (Spearman r = −0.5110, P < .0001) in PCa (Figure 4H) according to the TCGA data.
Besides, we found that overexpression of circSOBP could upregulate the expression of MYPT1 and downregulate the expression of p‐MLC2, but could not alter the expression of vimentin or total MLC2 (Figure 4I, K‐N). At the same time, depletion of circSOBP could downregulate MYPT1 and upregulate p‐MLC2 without altering vimentin or total MLC2 (Figure 4J, K‐N). Overexpression or depletion of circSOBP did not alter the expression of MLC2 mRNA (Supporting information Figure S3F,G).
3.6. Depletion of MYPT1 reverses the inhibitory effect of circSOBP on migration and invasion of PCa cells
The data from the TCGA Research Network showed the low expression of MYPT1 in PCa (n = 52) compared with normal specimens (n = 498) (Figure 5A). The ROC curve indicated that the AUC of MYPT1 was 0.875 (cut‐off value = 11.020, sensitivity = 0.788, 1–specificity = 0.162) (Figure 5B), indicating that MYPT1 was a promising biomarker for distinguishing PCa tissues. MYPT1 was downregulated among the paired samples (n = 52) (Figure 5C). MYPT1 was downregulated in 45 (82.1%) PCa tissues compared with the paired normal specimens (Figure 5D).
FIGURE 5.
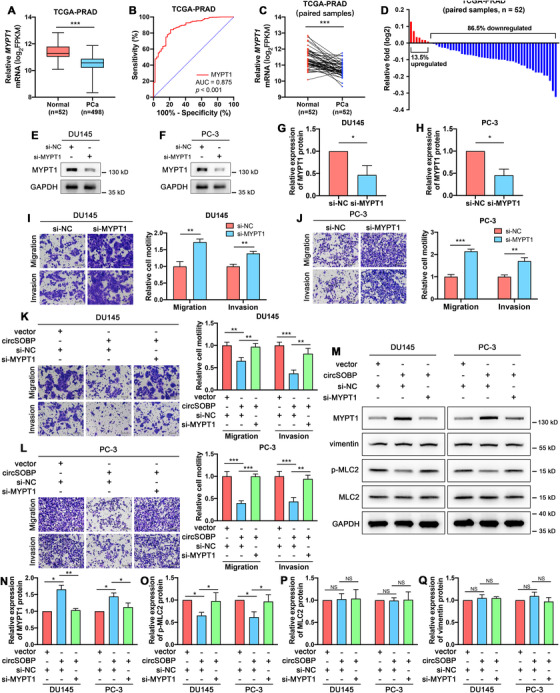
Depletion of MYPT1 reverses the inhibitory effect of circSOBP on PCa migration and invasion. (A) Expression of circSOBP in PCa and normal tissues from the TCGA database. Boxplot depicts median, 25th and 75th percentile, min‐max whiskers. ***P < .001, Mann–Whitney test. (B) The ROC curve in distinguishing PCa and ANP tissues by the expression of MYPT1. (C) Differential expression of circSOBP in 52 paired PCa and normal tissues from the TCGA database. ***P < .001, Wilcoxon matched‐pairs signed‐rank test. (D) The percentage of upregulated or downregulated expression of MYPT1 in 52 patients. (E) and (F) Efficacy of siRNA targeting MYPT1, analyzed using Western blot. (G) and (H) Densimetric analysis of the blots in (E) and (F), n = 3 independent experiments, GAPDH was used as a loading control. The data are presented as the mean ± SD. *P < .05, Student's t‐test, n = 3. (I) and (J) Effect of MYPT1 depletion on migration and invasion of DU145 and PC‐3 cell lines. The data are presented as the mean ± SD. **P < 0.01, ***P < .001, Student's t‐test, n = 3. (K) and (L) Effect of upregulated circSOBP accompanied by suppressing MYPT1 on migration and invasion of DU145 and PC‐3 cell lines. Scale bar, 100 μm. The data are presented as the mean ± SD. **P < .01, ***P < .001, one‐way ANOVA and Tukey's multiple comparisons test, n = 3. (M) Effect of upregulated circSOBP accompanied by suppressing MYPT1 on the expression of related proteins of DU145 and PC‐3 cells. (N)‐(Q) Densimetric analysis of the blots in (M), n = 3 independent experiments, GAPDH was used as the loading control. The data are presented as the mean ± SD. *P < .05, **P < .01, Student's t‐test. NC, normal control. NS, not significant
MYPT1 was successfully depleted using siRNA (Figure 5E‐H). Migration and invasion abilities of DU145 (Figure 5I) and PC‐3 (Figure 5J) cells were enhanced after the depletion of MYPT1. The depletion of MYPT1 reversed the inhibitory effect of circSOBP on migration and invasion of both DU145 (Figure 5K) and PC‐3 (Figure 5L) cell lines. Furthermore, overexpression of circSOBP boosted MYPT1 expression, while inhibited p‐MLC2. The depletion of MYPT1 could reverse the regulatory effects of circSOBP on the expression of MYPT1 (Figure 5M,N) and p‐MLC2 (Figure 5M,O). The expression of total MLC2 (Figure 5M,P) and vimentin (Figure 5M,Q) was not affected by the change of circSOBP or MYPT1.
3.7. Flanking introns mediate the circularization of circSOBP
We found that there were two Alu elements, Alu Sz and Alu Sq, located in the flanking introns of circSOBP in the SOBP gene, which might boost the formation of circSOBP. By aligning the sequences of Alu Sz and Alu Sq using the Basic Local Alignment Search Tool (https://blast.ncbi.nlm.nih.gov/Blast.cgi), it was found that they were highly reverse complementary (Figure 6A). To verify whether the circularization of circSOBP was promoted by Alu Sz and Alu Sq, we constructed five sequences in pcDNA3.1 plasmid (Figure 6B): #1, Exon 2 & 3 with wild type of flanking introns containing the Alu elements; #2, Exon 2 & 3 and flanking introns without Alu Sz; #3, Exon 2 & 3 and flanking introns without Alu Sq; #4, Exon 2 & 3 and flanking introns without both Alu elements; #5, Exon 2 & 3 without flanking introns. After the transfection of the plasmids into PC‐3 cells, the qRT‐PCR assay was used to verify the circularization of circSOBP using divergent primers and convergent primers, respectively. Interestingly, we found that the flanking introns without the Alu elements (#2‐4) could also mediate the circularization of circSOBP (Figure 6C), while the #5 plasmid could only overexpress the linear sequence of circSOBP (Figure 6C,D). Nevertheless, #2‐4 plasmids overexpressed less circSOBP than #1 plasmid, probably due to the deletion of Alu elements (Figure 6C).
FIGURE 6.
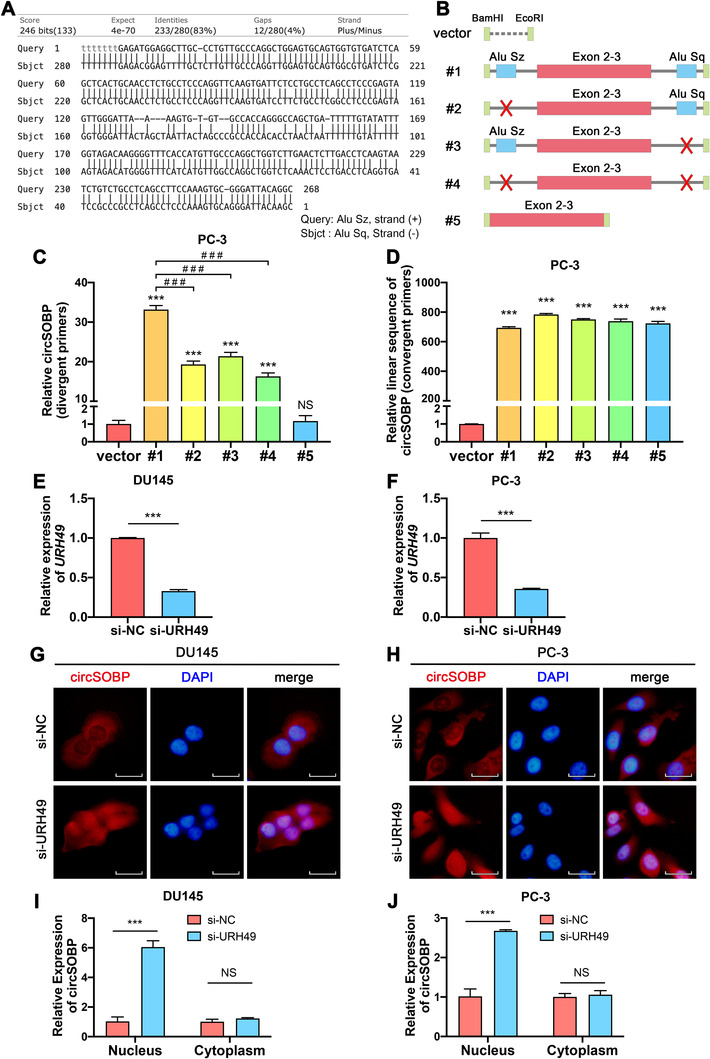
The circularization and nuclear export of circSOBP. (A) Alignment of Alu Sz and Alu Sq which were located on the flanking introns of circSOBP. (B) Schematic illustration of the plasmid constructions which were used to investigate the circularization of circSOBP. (C) Effect of the deletions of Alu sequences and flanking introns on the circularization of circSOBP analyzed using divergent primers. The data are presented as the mean ± SD. ***P < .001 comparing with vector, ###P < .001 comparing with #1, one‐way ANOVA and Dunnett's multiple comparisons test, n = 3. (D) Effect of the deletions of Alu sequences and flanking introns on the expression of the linear sequence of circSOBP analyzed using convergent primers. The data are presented as the mean ± SD. ***P < .001 comparing with vector, one‐way ANOVA, and Dunnett's multiple comparisons test, n = 3. (E) and (F) Efficacy of URH49 depletion using siRNA in DU145 and PC‐3 cell lines. The data are presented as the mean ± SD. *** P < .001, Student's t‐test, n = 3. (G) and (H) Effect of URH49 depletion on the subcellular distribution of circSOBP in DU145 and PC‐3 cells, analyzed using circRNA FISH assay. (I) and (J) Effect of URH49 depletion on the subcellular distribution of circSOBP in DU145 and PC‐3 cells, analyzed using qRT‐PCR. The data are presented as the mean ± SD. ***P < .001, Student's t‐test, n = 3. NC, normal control. NS, not significant
3.8. URH49 mediates the nuclear export of circSOBP
A previous study discovered that the depletion of URH49 caused short (<400 nucleotides) circRNAs to be enriched in the nucleus. 16 Our findings indicated that circSOBP is a short circRNA (325 nucleotides) that mainly distributes in the cytoplasm. Therefore, we further explored whether URH49 could mediate the nuclear export of circSOBP. We successfully depleted the expression of URH49 in DU145 (Figure 6E) and PC‐3 (Figure 6F) cell lines using siRNA. CircRNA FISH assay (Figure 6G,H) and isolation of the nuclear and cytoplasmic RNA followed by qRT‐PCR (Figure 6I,J) suggested that URH49 depletion led to the accumulation of circSOBP in the nucleus.
4. DISCUSSION
The present study found a low expression of circSOBP in patients with PCa. CircRNAs are ubiquitously expressed in human tissues. More than 100 000 circRNAs have been identified, however, most of these molecules are understudied. 27 , 39 Previous research has reported a differential abundance of circRNAs in cancer tissues comparing with normal, a majority were downregulated in cancer. 39 We have performed circRNA microarray using PCa tissues and ANP tissues from five patients to find differentially expressed circRNAs in PCa. 30 After verification from PCa and ANP tissues from another 56 patients, we found circSOBP showed low expression in PCa. Besides, circSOBP expression was significantly correlated with the grade group of PCa patients, indicating that circSOBP might be related to the prognosis of patients with PCa. CircSOBP is derived from exon 2 and exon 3 of the SOBP gene, its function is unknown. We found that circSOBP governed PCa migration and invasion in vitro and metastasis in vivo. The depletion of circSOBP enhanced migration and invasion of PCa cells through regulating amoeboid migration. Amoeboid migration of cancer cells can lead to metastasis of cancer.
The invasion‐metastasis cascade 4 , 40 , 41 envisions a succession of local invasion, intravasation, transit through the lymphatic and hematogenous systems, extravasation, formation of micrometastases, and colonization. Cancer cells metastasize through cell movement, mainly through mesenchymal migration and amoeboid migration. Cells undergoing mesenchymal migration express markers of EMT such as vimentin, E‐cadherin, N‐cadherin, etc. 42 Cells that perform amoeboid migration require the interaction between p‐MLC2 and the F‐actin bundle to mediate the generation of high contractile forces, thereby, allowing cells to shuttle through the interstitial space. 43 , 44 Cells undergoing amoeboid migration could be distinguished by amoeboid pseudopods, cell roundness, and increased p‐MLC2. 7 , 8 , 9 We discovered that the depletion of circSOBP led to an increase in p‐MLC2 expression, incidence of blebs, and stable blebs, and more roundness. Nevertheless, circSOBP did not shift the expression of vimentin. These results showed that circSOBP regulated amoeboid migration of PCa cells, but did not regulate the EMT‐mediated mesenchymal migration. This is the first study that discovers the relationship between circRNA and amoeboid migration of cancer.
We found that circSOBP inhibited migration and invasion by regulating amoeboid migration and altering the MYPT1/p‐MLC2 axis. CircSOBP could sponge miR‐141‐3p, thereby regulating MYPT1/p‐MLC2 expression. Previous studies have proved that individual circRNAs could act as a miRNA sponge. 32 , 45 For example, CDR1as, one of the most well‐characterized circRNAs, have conserved miRNA binding sites. 32 CDR1as is resistant to miRNA‐mediated destabilization, but it sponges miR‐7 resulting in increased miR‐7 targets. 32 We confirmed that circSOBP was located in the cytoplasm using a FISH assay, so it has the potential to function as a miRNA sponge. We found circSOBP could sponge miR‐141‐3p, thereby upregulate MYPT1 expression, which is targeted by miR‐141‐3p. MYPT1 is a subunit of myosin phosphatase which mediates the dephosphorylation of p‐MLC2, 37 , 38 while p‐MLC2 interacts with F‐actin and produces the contractile force for cell movement. 43 , 44
Complementary sequences in the genomic flanking introns could facilitate the circularization of circRNAs. 11 , 12 , 13 In human cells, most flanking intronic complementary sequences are Alu elements. 11 , 12 Previous studies found that the deletion of Alu elements significantly reduces the circularization of circRNAs. 12 , 46 We found that there were two reverse complementary Alu elements in the flanking introns of circSOBP. However, flanking introns with the deletion of Alu elements could also mediate the circularization of circSOBP, although the production of circSOBP was reduced by comparing with the wild‐type introns. This finding indicates that complementary Alu elements are not necessary for the formation of circSOBP. A possible explanation for this might be that RNA binding protein could bind to the flanking introns and boost the formation of circRNAs. 47 , 48 , 49 The knockout of circRNAs using CRISPR‐Cas9‐mediated deletion of Alu elements 50 might be unreliable for certain circRNAs.
5. CONCLUSIONS
The present study identified a novel circRNA circSOBP that was downregulated in PCa. Overexpression of circSOBP suppressed PCa cells’ migration and invasion in vitro and metastasis in vivo. CircSOBP depletion increased migration and invasion and induced amoeboid migration of PCa cells. The inhibitory effect of circSOBP on PCa cell migration and invasion was at least in part due to the regulation of the miR‐141‐3p/MYPT1/p‐MLC2 axis. Complementary intronic Alu elements could promote but would not be necessary for the formation of circSOBP. The nuclear export of circSOBP was mediated by URH49 (Figure 7). Our study discovered the role of circSOBP and the mediation of amoeboid migration in cancer cells for the first time, which may provide a promising target for the therapeutic application in patients with PCa.
FIGURE 7.

Schematic illustration of the mechanism of circSOBP regulating amoeboid migration of PCa cells. CircSOBP functioned as a sponge for miR‐141‐3p, and in turn upregulated the expression of MYPT1, which dephosphorylate p‐MLC2. This pathway governed the amoeboid migration of PCa cells
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
The present study was approved by the Ethics Committee of Jinshan Hospital, Fudan University. Written consent was obtained from each participant.
CONFLICT OF INTEREST
The authors declare that they have no conflict of interest.
Supporting information
Supporting information
Supporting information
Supporting information
Supporting information
Supporting information
Supporting information
Supporting information
Supporting information
ACKNOWLEDGMENTS
We thank Prof. Yuanfang Zhang, Department of Urology, Huashan Hospital, Fudan University, for the critical advice and comments. This work was supported by grants from the National Science Foundation of Shanghai (No. 18ZR1405800) and the Project for Key Medical Specialty Construction in Jinshan District (6th Period, Type A) (No. JSZK2019A03) to GC.
Chao F, Song Z, Wang S, et al. Novel circular RNA circSOBP governs amoeboid migration through the regulation of the miR‐141‐3p/MYPT1/p‐MLC2 axis in prostate cancer. Clin Transl Med. 2021;11:e360. 10.1002/ctm2.360
Contributor Information
Guoxiong Xu, Email: guoxiong.xu@fudan.edu.cn.
Gang Chen, Email: chgan365@126.com.
DATA AVAILABILITY STATEMENT
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
REFERENCES
- 1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70(1):7‐30. [DOI] [PubMed] [Google Scholar]
- 2. Sartor O, de Bono JS. Metastatic prostate cancer. N Engl J Med. 2018;378(7):645‐657. [DOI] [PubMed] [Google Scholar]
- 3. Steeg PS. Targeting metastasis. Nat Rev Cancer. 2016;16(4):201‐218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144(5):646‐674. [DOI] [PubMed] [Google Scholar]
- 5. Guan X. Cancer metastases: challenges and opportunities. Acta Pharm Sin B. 2015;5(5):402‐418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Laser‐Azogui A, Diamant‐Levi T, Israeli S, Roytman Y, Tsarfaty I. Met‐induced membrane blebbing leads to amoeboid cell motility and invasion. Oncogene. 2014;33(14):1788‐1798. [DOI] [PubMed] [Google Scholar]
- 7. Ruprecht V, Wieser S, Callan‐Jones A, et al. Cortical contractility triggers a stochastic switch to fast amoeboid cell motility. Cell. 2015;160(4):673‐685. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Georgouli M, Herraiz C, Crosas‐Molist E, et al. Regional activation of myosin II in cancer cells drives tumor progression via a secretory cross‐talk with the immune microenvironment. Cell. 2019;176(4):757‐774. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Gao Y, Wang Z, Hao Q, et al. Loss of ERalpha induces amoeboid‐like migration of breast cancer cells by downregulating vinculin. Nat Commun. 2017;8:14483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Zhang P, Zhang XO, Jiang T, et al. Comprehensive identification of alternative back‐splicing in human tissue transcriptomes. Nucleic Acids Res. 2020; 28:1779‐1789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Jeck WR, Sorrentino JA, Wang K, et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA. 2013;19(2):141‐157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Zhang XO, Wang HB, Zhang Y, Lu X, Chen LL, Yang L. Complementary sequence‐mediated exon circularization. Cell. 2014;159(1):134‐147. [DOI] [PubMed] [Google Scholar]
- 13. Liang D, Wilusz JE. Short intronic repeat sequences facilitate circular RNA production. Genes Dev. 2014;28(20):2233‐2247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Kristensen LS, Andersen MS, Stagsted LVW, Ebbesen KK, Hansen TB, Kjems J. The biogenesis, biology and characterization of circular RNAs. Nat Rev Genet. 2019;20(11):675‐691. [DOI] [PubMed] [Google Scholar]
- 15. Chen LL. The expanding regulatory mechanisms and cellular functions of circular RNAs. Nat Rev Mol Cell Biol. 2020;21(8):475‐490. [DOI] [PubMed] [Google Scholar]
- 16. Huang C, Liang D, Tatomer DC, Wilusz JE. A length‐dependent evolutionarily conserved pathway controls nuclear export of circular RNAs. Genes Dev. 2018;32(9‐10):639‐644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Cocquerelle C, Mascrez B, Hetuin D, Bailleul B. Mis‐splicing yields circular RNA molecules. FASEB J. 1993;7(1):155‐160. [DOI] [PubMed] [Google Scholar]
- 18. Zhang Q, Wang W, Zhou Q, et al. Roles of circRNAs in the tumour microenvironment. Mol Cancer. 2020;19(1):14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Shen B, Wang Z, Li Z, Song H, Ding X. Circular RNAs: an emerging landscape in tumor metastasis. Am J Cancer Res. 2019;9(4):630‐643. [PMC free article] [PubMed] [Google Scholar]
- 20. Shang BQ, Li ML, Quan HY, et al. Functional roles of circular RNAs during epithelial‐to‐mesenchymal transition. Mol Cancer. 2019;18(1):138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Meng J, Chen S, Han JX, et al. Twist1 regulates vimentin through Cul2 circular RNA to promote EMT in hepatocellular carcinoma. Cancer Res. 2018;78(15):4150‐4162. [DOI] [PubMed] [Google Scholar]
- 22. Chen X, Chen RX, Wei WS, et al. PRMT5 circular RNA promotes metastasis of urothelial carcinoma of the bladder through sponging miR‐30c to induce epithelial‐mesenchymal transition. Clin Cancer Res. 2018;24(24):6319‐6330. [DOI] [PubMed] [Google Scholar]
- 23. Wang L, Tong X, Zhou Z, et al. Circular RNA hsa_circ_0008305 (circPTK2) inhibits TGF‐beta‐induced epithelial‐mesenchymal transition and metastasis by controlling TIF1gamma in non‐small cell lung cancer. Mol Cancer. 2018;17(1):140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Chen Z, Duan X. hsa_circ_0000177‐miR‐638‐FZD7‐Wnt signaling cascade contributes to the malignant behaviors in glioma. DNA Cell Biol. 2018;37(9):791‐797. [DOI] [PubMed] [Google Scholar]
- 25. Xia L, Wu L, Bao J, et al. Circular RNA circ‐CBFB promotes proliferation and inhibits apoptosis in chronic lymphocytic leukemia through regulating miR‐607/FZD3/Wnt/beta‐catenin pathway. Biochem Biophysic Res Commun. 2018;503(1):385‐390. [DOI] [PubMed] [Google Scholar]
- 26. Yang Z, Qu CB, Zhang Y, et al. Dysregulation of p53‐RBM25‐mediated circAMOTL1L biogenesis contributes to prostate cancer progression through the circAMOTL1L‐miR‐193a‐5p‐Pcdha pathway. Oncogene. 2019;38(14):2516‐2532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Glazar P, Papavasileiou P, Rajewsky N. circBase: a database for circular RNAs. RNA. 2014;20(11):1666‐1670. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real‐time quantitative PCR and the 2(‐Delta Delta C(T)) method. Methods. 2001;25(4):402‐408. [DOI] [PubMed] [Google Scholar]
- 29. Sanz‐Moreno V, Gaggioli C, Yeo M, et al. ROCK and JAK1 signaling cooperate to control actomyosin contractility in tumor cells and stroma. Cancer Cell. 2011;20(2):229‐245. [DOI] [PubMed] [Google Scholar]
- 30. Song Z, Zhuo Z, Ma Z, Hou C, Chen G, Xu G. Hsa_Circ_0001206 is downregulated and inhibits cell proliferation, migration and invasion in prostate cancer. Artif Cells Nanomed Biotechnol. 2019;47(1):2449‐2464. [DOI] [PubMed] [Google Scholar]
- 31. Lammermann T, Sixt M. Mechanical modes of ‘amoeboid’ cell migration. Curr Opin Cell Biol. 2009;21(5):636‐644. [DOI] [PubMed] [Google Scholar]
- 32. Hansen TB, Jensen TI, Clausen BH, et al. Natural RNA circles function as efficient microRNA sponges. Nature. 2013;495(7441):384‐388. [DOI] [PubMed] [Google Scholar]
- 33. Chang TH, Huang HY, Hsu JB, Weng SL, Horng JT, Huang HD. An enhanced computational platform for investigating the roles of regulatory RNA and for identifying functional RNA motifs. BMC Bioinformatics. 2013;14(Suppl 2):S4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Li JH, Liu S, Zhou H, Qu LH, Yang JH. starBase v2.0: decoding miRNA‐ceRNA, miRNA‐ncRNA and protein‐RNA interaction networks from large‐scale CLIP‐Seq data. Nucleic Acids Res. 2014;42:D92‐D97. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Weiser DC, Row RH, Kimelman D. Rho‐regulated myosin phosphatase establishes the level of protrusive activity required for cell movements during zebrafish gastrulation. Development. 2009;136(14):2375‐2384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Ito M, Nakano T, Erdodi F, Hartshorne DJ. Myosin phosphatase: structure, regulation and function. Mol Cell Biochem. 2004;259(1‐2):197‐209. [DOI] [PubMed] [Google Scholar]
- 37. Wang X, Obeidat M, Li L, et al. TIMAP inhibits endothelial myosin light chain phosphatase by competing with MYPT1 for the catalytic protein phosphatase 1 subunit PP1cbeta. J Biol Chem. 2019;294(36):13280‐13291. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Kiss A, Erdodi F, Lontay B. Myosin phosphatase: unexpected functions of a long‐known enzyme. Biochim Biophys Acta Mol Cell Res. 2019;1866(1):2‐15. [DOI] [PubMed] [Google Scholar]
- 39. Vo JN, Cieslik M, Zhang Y, et al. The landscape of circular RNA in cancer. Cell. 2019;176(4):869‐881. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Talmadge JE, Fidler IJ. AACR centennial series: the biology of cancer metastasis: historical perspective. Cancer Res. 2010;70(14):5649‐5669. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Fidler IJ. The pathogenesis of cancer metastasis: the ‘seed and soil’ hypothesis revisited. Nat Rev Cancer. 2003;3(6):453‐458. [DOI] [PubMed] [Google Scholar]
- 42. Zeisberg M, Neilson EG. Biomarkers for epithelial‐mesenchymal transitions. J Clin Invest. 2009;119(6):1429‐1437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Wyckoff JB, Pinner SE, Gschmeissner S, Condeelis JS, Sahai E. ROCK‐ and myosin‐dependent matrix deformation enables protease‐independent tumor‐cell invasion in vivo. Curr Biol. 2006;16(15):1515‐1523. [DOI] [PubMed] [Google Scholar]
- 44. Kimura K, Ito M, Amano M, et al. Regulation of myosin phosphatase by Rho and Rho‐associated kinase (Rho‐kinase). Science. 1996;273(5272):245‐248. [DOI] [PubMed] [Google Scholar]
- 45. Xiao MS, Ai Y, Wilusz JE. Biogenesis and functions of circular RNAs come into focus. Trends Cell Biol. 2020;30(3):226‐240. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Yu J, Xu QG, Wang ZG, et al. Circular RNA cSMARCA5 inhibits growth and metastasis in hepatocellular carcinoma. J Hepatol. 2018;68(6):1214‐1227. [DOI] [PubMed] [Google Scholar]
- 47. Nilsen TW, Graveley BR. Expansion of the eukaryotic proteome by alternative splicing. Nature. 2010;463(7280):457‐463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Conn SJ, Pillman KA, Toubia J, et al. The RNA binding protein quaking regulates formation of circRNAs. Cell. 2015;160(6):1125‐1134. [DOI] [PubMed] [Google Scholar]
- 49. Ashwal‐Fluss R, Meyer M, Pamudurti NR, et al. circRNA biogenesis competes with pre‐mRNA splicing. Mol Cell. 2014;56(1):55‐66. [DOI] [PubMed] [Google Scholar]
- 50. Zhang Y, Xue W, Li X, et al. The biogenesis of nascent circular RNAs. Cell Rep. 2016;15(3):611‐624. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supporting information
Supporting information
Supporting information
Supporting information
Supporting information
Supporting information
Supporting information
Supporting information
Data Availability Statement
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.


