Summary
Optical clearing has emerged as a powerful tool for volume imaging. Although volume imaging with immunostaining have been successful in many protocols, yet obtaining homogeneously stained thick samples remains challenging. Here, we propose a method for label-free imaging of brain slices by enhancing the regional heterogeneity of the optical properties using the tissue clearing principle. We used FxClear, a method for delipidation of brain tissue, to retain a larger proportion of lipids at the white matter (WM). Furthermore, the embedding media affected the contrasts for the lipid-rich or extracellular matrix-rich areas, depending on their chemical properties. Thus, we tailored clearing conditions for the enhancement of the refractive indices (RIs) differences between gray and WM, or several pathological features. RI differences can be imaged using conventional light microscopy or optical coherence tomography. We propose that our protocol is simple, reliable, and flexible for label-free imaging, easily implementable to routine histology laboratory.
Subject areas: Physiology, Techniques in Neuroscience, Biology Experimental Methods
Graphical abstract

Highlights
-
•
Label-free imaging via enhancing RI differences in optically cleared brain
-
•
Recognition of lipid- or ECM-rich regions using tailored protocols
-
•
Easy and affordable protocols using conventional optics
Physiology; Techniques in Neuroscience; Biology Experimental Methods
Introduction
Acquiring high-resolution images from thick samples in light microscopy has been considered to have great limitations owing to the light scattering due to the high lipid contents and the opaque, dense extracellular matrix (ECM) in thick tissues (Appel et al., 2013; Bolmont et al., 2012). Thus, the advent of tissue clearing technique for the optical examination of thick tissues has revolutionized the 3D histological and pathological tissue examinations (Liebmann et al., 2016; Murata et al., 2017; Nojima et al., 2017; Olson et al., 2016; Ueda et al., 2020). There are many protocols for tissue clearing, and most of them rely on the physical strategy to reduce light scattering, which is primarily caused by the components and arrangement of materials with different refractive indices (RIs) in tissues (Chung et al., 2013; Ertürk et al., 2012; Hama et al., 2011; Ke et al., 2013; Kuwajima et al., 2013; Lee et al., 2016a; Pan et al., 2016; Renier et al., 2014; Susaki et al., 2014). To reduce photon scattering, most tissue clearing techniques remove lipids, which are the main substances causing light scattering in tissue. After lipid removal, reducing the inhomogeneity of the tissue components by immersion in RI matching medium (Chung et al., 2013; Ertürk et al., 2012; Hama et al., 2011; Ke et al., 2013; Kuwajima et al., 2013; Lee et al., 2016a; Pan et al., 2016; Renier et al., 2014; Susaki et al., 2014).
The performance of these clearing techniques varies depending on the constitution of the biological tissue. For example, tissue clearing efficiency is different across the gray matter (GM) and white matter (WM) regions of the brain because the WM region contains the lipid-rich myelin sheaths. Tissues have unique signatures for their macromolecule content and arrangement; therefore, their optical properties are distinct depending on the types and conditions of the tissues (Lee et al., 2016b; Tainaka et al., 2016; Yang et al., 2014). The contributions of these factors to the tissue-clearing efficacy are different, and the contribution of RI matching on tissue clearing efficacy is the largest in most tissues, suggesting that the properties of tissue ECM may be the primary factors influencing tissue clarity (Kim et al., 2018). Therefore, changes in the architecture and biological composition of tissues under pathological conditions, such as tissue fibrosis, can be recognized by the differential responses of the affected tissues to the tissue-clearing processes. We tested this possibility using the Active Clarity Technique (ACT) with the use of a CUBIC-based mounting solution and found that it can detect tissue fibrosis with disorganized ECM arrangements (Kim et al., 2018).
While the reduction in the RI difference is desirable after tissue clearing of thick specimens to obtain fluorescence signals (e.g., using fluorescence proteins or dyes), RI differences have been widely used for the detection of tissue or cell structural changes even without labeling (Habaza et al., 2017; Sung et al., 2009; Yi and Backman, 2012). In particular, optical coherence tomography (OCT) is one of the imaging techniques suitable for combined with optically cleared tissue (Baek et al., 2019; Sudheendran et al., 2010). OCT recognizes the intrinsic scattering properties of biological tissues and has been widely used for label-free detection of normal and pathological tissue structures (Aumann et al., 2019; Baek et al., 2019; Bizheva et al., 2007; Brezinski et al., 1996; Hanna et al., 2005). Control of intrinsic tissue scattering through the tissue clearing procedure enables high-resolution OCT imaging of neuronal bundles and blood vessel in thick brain tissue without contrast agents, and enhanced OCT signals can be obtained using tissue clearing agents (Larina et al., 2008; Ren et al., 2017; Zhong et al., 2010). However, the OCT system is not easily accessible to many laboratories, and label-free imaging of thick tissues using conventional light microscopes has been less addressed.
In this study, we aimed to further improve/examine the conditions for better recognition of the biophysical properties of the target tissues owing to the differential lipid content, ECM content, and cell compositions using conventional microscopic systems. In our optimized conditions, nerve bundles in the normal brain, brain trauma-induced (TBI) tissue fibrosis, and misfolded proteins accumulate to form amyloid plaques in the Alzheimer's disease (AD) mouse brain were sensitively recognized without any labeling. Furthermore, we discovered that the use of different RI adjusting media provided differential responses in biological specimens, which will be an additional consideration for the development of diagnostic strategies based on label-free optical imaging using the differences in RI.
Results
FxClear-based tissue delipidation for the higher contrast imaging of WM
Previously, we have reported the FxClear protocol, which utilizes detergent-based delipidation of paraformaldehyde-fixed samples (Choi et al., 2019). Delipidation by the FxClear protocol does not allow for the complete removal of lipids; therefore, the resultant samples were less transparent (Figure 1A). However, they appeared to show increased contrast between the GM and lipid-rich WM compared to that of the ACT samples that exhibited uniform clearing with more extensive delipidation (Figure 1B). We confirmed that the less transparent regions in FxClear-processed brain slices were well-matched with the nerve bundles in the WM and striatum, which are labeled by TuJ1 and MBP antibodies (Figure 1C). As a primary test, we simply monitored how the contrast between the GM and WM changes during RI tuning. We selected fructose and iodixanol as RI matching compounds, which are highly soluble in water and display high RI ranging from 1.43 to 1.47. The reduction of RI of the matching solution to the suboptimal level progressively enhanced the contrast between the transparencies of the GM and WM in both ACT- and FxClear-processed samples followed by RI matching immersion (Figure 1D). Interestingly, when we used fructose as an RI matching medium, the transparency of the GM and WM was significantly affected by the RI value in FxClear-processed samples, and the transparency ratio of GM to WM (GM/WM) was larger with the FxClear-processed brain slices than with the ACT-processed samples (Figures 1D and S1). These results indicate that the transparency of the tissue is dependent on multiple factors including the characteristics of tissues (GM vs. WM), tissue processing protocol (ACT vs. FxClear), and RI value.
Figure 1.
FxClear-based tissue processing for label-free brain imaging
(A and B) Comparison of mouse brain slices (1 mm) processed with ACT or FxClear protocol. Digital image of optically cleared tissue with RI matching (A) and non-matching (B).
(C) Correlative trans-illumination imaging (gray or pseudocolored in green) of FxClear-processed samples with immunostaining of nerves (Tuj1, red) and myelin sheets (MBP, green). Scale bar = 1 mm. Inset, Scale bar = 50 μm.
(D) Images after the immersion of ACT or FxClear-processed brain slices in fructose solution with different RI (1.39–1.47) for 1 hr. Quantification of %Transparency in gray matter (GM) and white matter (WM). Data are represented as mean ± SD. Unpaired two-tailed t-tests on data between the GM and WM, n = 6, ns, not significant, ∗p < 0.05, ∗∗p = 0.0033, ∗∗∗p = 0.0007, ∗∗∗∗p < 0.0001. Scale bar = 100 μm.
Effects of RI matching media on the imaging of WM
Next, we tested the effects of different RI matching reagents (fructose, sucrose, histodenz, and iodixanol) on the contrast of the transparencies between the GM and WM (Figure 2). Although we adjusted all the media with the same RI (1.43), there were significant differences in transparency depending on the degree of delipidation and the kinds of media, resulting in a different transparency ratio with significantly higher ratio enhancement by RI matching with iodixanol (Figures 2A–2C and S2A). The use of iodixanol, which showed the highest GM:WM ratio, provided a strong contrast that was sufficient for visualization of the nerve bundles in 1 mm serial slices of the mouse brain with trans-illumination light microscopy or stereo microscopy (Figures S2B and S2C). Iodixanol also exhibited RI-dependent transparency of the WM and GM, similar to that by fructose, but we found that the ratio of GM/WM in different RI values remained constant (ratio = 1.5 with ACT, 2.8 with FxClear-processed tissues; Figure S1). The hippocampal regions of the human brain slices were also visualized with FxClear and iodixanol incubation, showing high contrast of the nerve fiber areas (Figures 2Dand 2E). Notably, the laminar organization of the parahippocampal gyrus can easily be determined by this method (Figure 2Eb). Collectively, these results indicate that different RI adjusting media exhibit different features and can be chosen for reduction or enhancement of tissue RI differences. We chose two reagents, fructose, and iodixanol, for further comparison.
Figure 2.
RI media-dependent differences of the image contrast
(A) Images after immersion of FxClear-processed mouse brain slices in RI ( = 1.43) adjustment media (fructose, sucrose, histodenz, and iodixanol) for 1 hr. Note that the dark structures in the images are tungsten sticks to immobilize the brain slice during the imaging process. Scale bar = 100 μm.
(B) Quantification of %Transparency in gray matter (GM) and white matter (WM). Data are represented as mean ± SD. Unpaired two-tailed t-tests on data between the GM and WM, n = 6, ∗∗p < 0.01, ∗∗∗∗p < 0.0001.
(C) The ratio of %Transparency of the GM and WM. Data are represented as mean ± SD. One-way ANOVA with Dunnett's post-hoc test (sucrose as control) on data, n = 6–7 individual RI solution, ∗p = 0.0263, ∗∗∗p = 0.0006.
(D) Human brain slices (1 mm) containing hippocampal formation immersed in iodixanol after FxClear process.
(E) Stereo microscopic images of cleared human hippocampal tissues. Abbreviations are DG, dentate gyrus; CA1, Cornu Ammonis 1; mf, mossy fiber. Scale bar = 1mm.
Comparison of de-lipidation methods for label-free imaging with RI differences
To compare whether other de-lipidation methods used in other clearing techniques are applicable to label-free imaging, we selected two clearing methods, CUBIC and iDISCO (Renier et al., 2014; Susaki et al., 2014). We modified the de-lipidation protocols from these methods to adopt them in our procedure. In the case of CUBIC, the RI of reagent 1 is approximately 1.435 because it contains 25% N,N,N′,N′-tetrakis(2-hydroxypropyl) ethylenediamine which exhibits both RI adjusting and delipidation effects. Thus, we simply used reagent 1 and incubated the brain slices for 24 hr. In the case of iDISCO, dichloromethane (DCM) treatment is critical for the delipidation, and we dehydrated the brain slices with methanol ascending series, incubated the slices with DCM briefly (15-min) and rehydrated them with methanol descending series, and immersed them in iodixanol. Figure 3A shows the time spending in each treatment step of the clearing protocols. The samples were incubated in the RI matching solutions after delipidation, and the transmittance images were photographed using conventional light microscopes (Figure 3B). CUBIC- and iDISCO-processed brain slices show high tissue transparency (Figure 3C) and good contrast between GM and WM, while CUBIC-processed slices exhibited significantly inferior contrast comparing to the FxClear- or iDISCO-processed brain slices (Figure 3D).
Figure 3.
Comparison of delipidation methods for label-free imaging
(A) Tissue delipidation protocols of different clearing methods for 1mm-thick-brain slices. ETC, electrophoretic tissue clearing; DCM, dichloromethane; RIs, refractive indices.
(B) Trans-illumination (upper) or stereo (bottom) microscopic images of 1-mm-thick adult mouse brain slices after delipidation. Scale bar = 1 mm.
(C) Quantification of % transparency in gray matter (GM) and white matter (WM). Data are represented as mean ± SD. Unpaired two-tailed t-tests on data between the GM and WM, n = 6 individual method, ∗∗∗p < 0.001.
(D) The ratio of % transparency of the GM and WM. Data are represented as mean ± SD. One-way ANOVA with Dunnett's post-hoc test (FxClear as control) on data, n = 6 individual method, ∗p < 0.05.
Optimization of the size expansion for the simple analysis
Previously, we have reported that FxClear-based delipidation by electrophoretic tissue clearing (ETC) causes mild expansion of the tissues, and subsequent washing with phosphate-buffered saline (PBS) and incubation with RI adjustment media (CUBIC-mount) gradually returned the samples to the original size (Kim et al., 2018). Unlike CUBIC-mount, fructose and iodixanol showed an opposite trend in their effect on the tissue size (Figure S3A). Incubation of ETC samples with fructose shrunk the samples considering the original size with increasing ionic strength of the washing buffer. Iodixanol inversely expanded the tissue regardless of the ionic strength of the washing buffer (Figure S3B). Considering that tissue swelling is associated with (hyper-)hydration of the tissues, the size differences appeared to be caused by the dissimilar ability of fructose and iodixanol to promote tissue hydration. Changes in the tissue size also affect the transparency of the tissues (Kim et al., 2018); therefore, we used 0.1× PBS washing condition, which provided a minimal difference in the tissue sizes in the fructose and iodixanol groups.
Observation of FxClear-processed samples using OCT
Next, we quantified the light penetration depth and the contrast between the GM and WM using a lab-made swept-source OCT system (Figure S4A). In the OCT images, the contrast between the GM and WM was more evident in the brain slices subjected to RI matching using fructose and iodixanol than those subjected to non-RI matching (Figure 4A). For the contrast measurement, we selected four regions from the en-face image at a certain depth position that can cover the entire region of the GM and WM. The en-face images of each sample were extracted from each volume data at a depth position where the GM and WM could be clearly distinguished (Fix and FxClear: 0.26 mm from the top surface; fructose and iodixanol: 0.53 mm from the top surface). Owing to the high contrast, optical coherence microscopy (OCM) imaging showed the clearest and highly contrasted image of fiber bundles in the specimen processed with FxClear and immerged in iodixanol (Figures 4B and S4B). Next, we analyzed the penetration depth of light by imaging samples of 2 mm thickness with the depth range-enhanced OCT system. The surface of the sample in all sub-volume from the OCT volume data was flattened, and we obtained a signal profile according to the depth at the GM and WM regions by averaging the flattened sub-volume data in the lateral direction (Figure 4C). In this analysis, we found that the high attenuation rate of the fixed sample limited the imaging depth to approximately 1.5 mm, whereas the FxClear process increased the imaging depth to >2.5 mm by reducing the attenuation rate. Immersion of FxClear samples in fructose or iodixanol further reduced the attenuation rate. Although the signal strength at the surface of the specimen was substantially lower than that of the fixed or FxClear samples, the immersion of the samples in the RI matching solution maintained the maximal imaging depth at approximately 2.5 mm (Figures 4D and 4E). In particular, the immersion of FxClear samples in fructose or iodixanol further reduced the attenuation rate because of the improved signal-to-noise ratio (Figure 4F). Taken together, these data suggest that RI matching including delipidation improves imaging depth and quality in label-free imaging systems.
Figure 4.
OCT and OCM images of mouse brain slices, and the results of their analysis
(A) en-face intensity (left) and pseudo-colored (right) images obtained after averaging OCT volume data in the z-direction.
(B) OCM images showing fiber bundles (image size: 320 μm × 320 μm).
(C) Averaged signal profiles according to z-direction yellow boxed region in (A) each condition. Signal decreases from near 0 mm owing to the light scattering by the tissue and increases from near 2 mm by noise. Sudden reductions in signal strengths in the shaded area in the graphs indicate the bottom margin of the imaging system.
(D and E) Signal attenuation obtained from (C) and light penetration depth, respectively. The penetration depth is defined as the depth that became 0 dB after subtracting the background signals from the averaged signal profiles. Data are represented as mean ± SD. One-way ANOVA with Dunnett's post-hoc test (Fixation as control) on GM and WM data, n = 3 individual condition, ns, not significant, ∗∗p < 0.01, ∗∗∗p = 0.004, ∗∗∗∗p < 0.0001.
(F) The contrast between the GM and WM. Data are represented as mean ± SD. One-way ANOVA with Dunnett's post-hoc test (Fixation as control) on data, n = 3 individual condition, ns, not significant (p = 0.0766), ∗∗∗∗p < 0.0001.
Autofluorescence-based imaging of FxClear-based de-lipidated whole tissue using selective plane illumination microscope
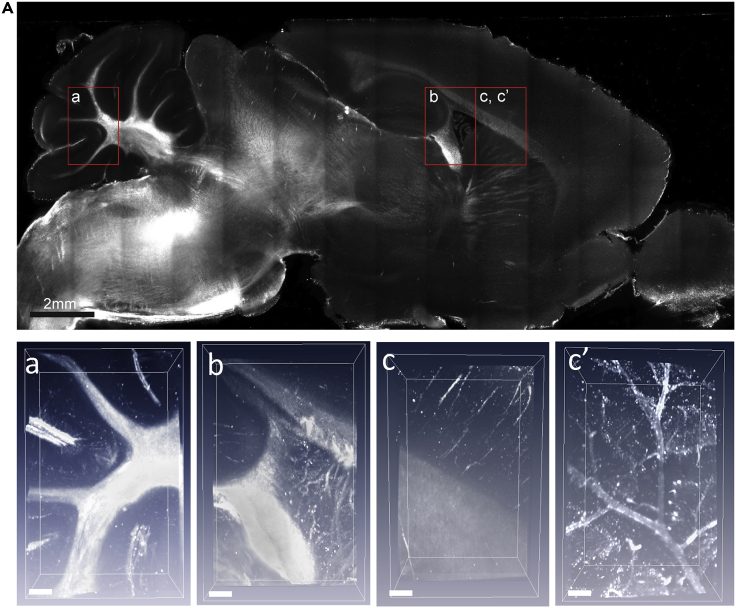
While RI difference in FxClear-processed tissue was successfully captured and imaged with the above optical devices, we wondered whether autofluorescence in the WM can be also imaged from the tissues, as previously demonstrated with other optical clearing methods (Morozov et al., 2010; Ren et al., 2017; Renier et al., 2016). Using a home-made selective plane illumination microscope (SPIM) based on the Huisken et al. (Huisken and Stainier, 2007), FxClear-processed entire hemi-sectioned mouse brain was successfully imaged (Figure 5A and Video S1). We observed the fiber projection profiles in the brain. In high-resolution imaging with SPIM, detailed structures of fibers projections in the cerebellum (Figure 5A), hippocampus (Figure 5B), and corpus callosum (Figure 5C) were visualized (Video S2). In addition, vasculature in the cerebral cortex was also noticeable (Figure 5C'), which was less profound in our previous RI-based imaging. Thus, it appeared that autofluorescence is more sensitive to imaging vascular structures in the brain. Hence, FxClear-based delipidated tissue could be applied for label-free 3D imaging based on their regional differences in RI and autofluorescence.
Figure 5.
SPIM imaging of an intact mouse brain hemisphere
(A) A sagittal view of the mouse brain hemisphere. Scale bar = 2 mm (a–c) High-magnification views of the red boxed regions in (A). (a–ć) Three-dimensional rendering of the selected subvolume in (A) Scale bar = 200 μm.
Differentiation of injury areas via different RI adjustment media
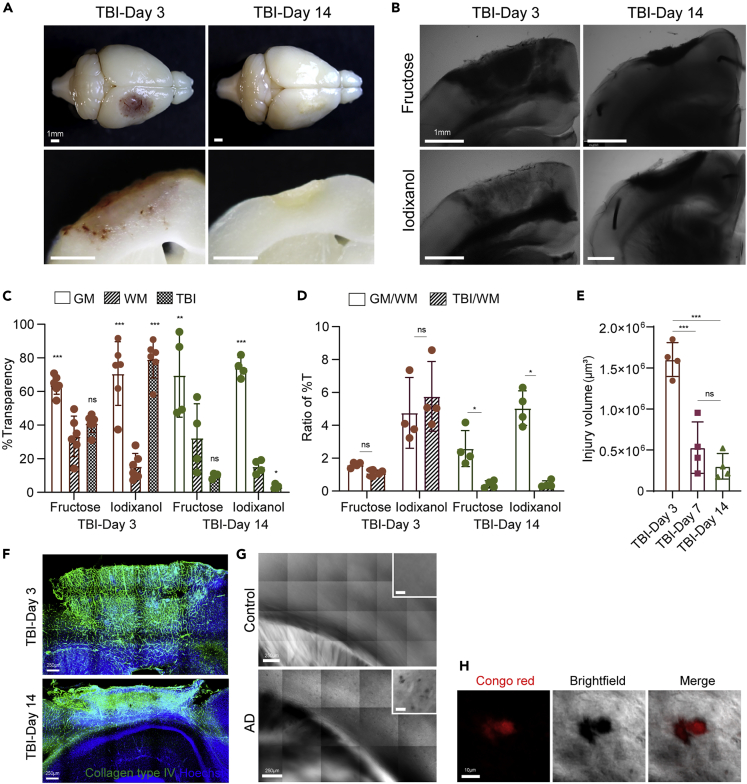
Next, we tested whether a new protocol can be used for label-free identification of various pathological signatures in biological tissues. We first damaged the cerebral cortex with a liquid nitrogen-frozen probe to induce TBI, which caused rapid bleeding and progressive recovery with permanent tissue damage (Figure 6A). Our optimized protocol sensitively recognized the damaged area as early as 3 days after TBI. In fructose medium, the TBI injury spot became dark and clearly distinguishable from the injury periphery. The injured regions in iodixanol became brighter with spotty dark areas in TBI day 3 samples, which were remnants of blood clots after the cardiac perfusion procedure, but the clots appeared to have been removed in TBI day 7 samples (Figures 6A, 6B, and S6A). By 2 weeks after injury, the damaged area became increasingly opaque, and transparency markedly reduced in both RI media (Figures 6B and 6C). Accordingly, the ratio of the TBI regions and WM (TBI/WM) progressively decreased, whereas the GM/WM ratio remained relatively stable (Figure 6D). Owing to the high contrast, the injured area was easily identifiable, and we measured the time course changes in the injury volume over 2 weeks, demonstrating the value of the method for the quantitative and qualitative evaluation (Figure 6E).To associate TBI-induced brain damage and optical alterations, we used typical markers representing the injury penumbra (accumulation of activated microglia labeled with Iba1 antibody) and injury periphery (activation of astrocytes labeled by GFAP antibody) and explored their distribution (Figure S6B). These labeling clearly distinguished the different sub-regions, but FxClear-based transparency imaging did not efficiently segregate these sub-regions. Collagen type IV staining as a marker for ECM deposition/rearrangement was well-matched with the transparency imaging, suggesting that these optical features are primarily affected by ECM condensation (Figure 6F).
Figure 6.
Label-free imaging of various pathological tissue
(A) Images of the whole (top) and sliced (bottom) mouse brain 3 and 14 days after cryogenic TBI. Scale bar = 1 mm.
(B) Brightfield images of brain slices (3- or 14-days after injury) immersed in fructose or iodixanol media. Scale bar = 500 μm.
(C) Quantification of %Transparency in GM, WM, and TBI region. Data are represented as mean ± SD, One-way ANOVA with Dunnett's post-hoc test (WM as control) on data, n = 4–7 individual condition, ∗p = 0.035, ∗∗p = 0.0036, ∗∗∗p < 0.001, ns, not significant.
(D) Quantification of %Transparency ratios of GM:WM and TBI/WM in each group. Data are represented as mean ± SD. Unpaired two-tailed t-tests on data between individual pairs, n = 4–7, ns, not significant, ∗p < 0.05.
(E) Quantification of injury region volume change after the injury. Data are represented as mean ± SD. One-way ANOVA with Bonferroni's multiple comparison test on data, n = 4 individual group, ns, not significant, ∗∗∗p < 0.001.
(F) Immunostaining of brain slices obtained 3 or 14 days after TBI: Collagen type IV (green) and Hoechst (blue). Scale bar = 250 μm.
(G) Comparison of label-free images between the normal and 5xFAD AD model mouse brain slice. Insets show large magnification views. Scale bar = 250 μm. Scale bar in insets = 25 μm.
(H) High magnified images of Aβ in an AD brain slice with Congo red staining (red). Scale bar = 10 μm.
Next, we tested whether our protocol can capture the pathological features such as misfolded protein aggregates, such as amyloid-beta (Aβ) plaques in AD mouse brain. Figure 6G shows stitched wide-field transillumination images of representative cortex coronal plane from control or 5xFAD AD model mouse brain which were FxClear-processed and immersed in fructose media, demonstrating that Congo red-labeled Aβ-plaques were selectively found only in the AD mouse brain (Video S3). A large magnification image clearly showed that the Congo red-labeled area can also be distinguished with dark spots in the transillumination imaging (Figure 6H).
Finally, we examined whether this protocol discriminates against the implanted human glioblastoma (GBM), which has a distinct cellular identity from nearby the mouse brain tissue. However, the implanted GBM cells were not associated with ectopic vasculature or disruption of ECM arrangements and were not recognized by the transparency difference in both fructose and iodixanol solutions (Figure S6C).
Discussion
In this study, we optimized the tissue-clearing protocol for the assessment of brain structures without specific labeling. By using simple transillumination or sophisticated imaging systems such as OCT and OCM, we were able to discriminate nerve fiber bundles that have high lipid content and brain traumatic regions that exhibit blood clots, misfolded protein accumulation such as Aβ plaques, and deposition of ECM proteins, but not implanted GBM cells. High-contrast, label-free imaging was primarily achieved with optimization of delipidation and RI adjustment conditions. To visualize the RI difference-based tissue structures, we used three different optical methods: simple trans-illumination imaging, stereo microscopic imaging, and OCT. Because trans-illumination or stereo microscopic imaging systems are basic equipment for most histological laboratories, our protocol provides a simple and affordable approach to achieve label-free information of brain structures. Loss of depth information would restrict the application of this method to some degree, but this limitation can be overcome partially using stereo microscopy, and the use of OCT provided the best quantifiable results. Quantification of the images obtained from OCT revealed that FxClear with RI adjustment in iodixanol provided the best-fit contrast and sufficient imaging of 3 mm depth without labeling. In addition, a recently introduced imaging system with solely based on the autofluorescence allowed axonal bundles, blood vessels, and neuronal fiber in the mouse brain and structural proteins such as collagen, elastin in mouse lung without labeling (Morozov et al., 2010; Ren et al., 2017; Renier et al., 2016). Similarly, we found that FxClear-based delipidated mouse brain hemisphere can also be utilized for 3D imaging with these autofluorescence signals (Figure 5). Thus, we propose that our current protocol is flexible to obtain various label-free images by different optical devices.
Delipidation in FxClear protocol resulted in better contrast between the GM and WM than other protocols offering better delipidation such as ACT (Lee et al., 2016a). Delipidation in FxClear protocol is executed by electrophoresis with a buffer containing 2% sodium dodecyl sulfate (SDS) micelles, and more than 30% of lipids remain in the tissue after the completion of electrophoresis (Choi et al., 2019). SDS cannot remove all types of lipids equally in the biological specimen, resulting in lipid species remaining even after near completion of the reactions. Although charged lipids such as phospholipids are easily removable by SDS, the removal of neutral lipids and fat bodies is less effective (Jensen and Berg, 2017). Also, the dense hydrophobic assembly of lipids reduces the surface–volume ratio, resulting in poor interaction with SDS micelles. Therefore, both the quality and quantity of lipids in the tissue may affect the efficiency of SDS-based delipidation, and fiber-dense regions are less efficiently resolved by SDS-based clearing. In direct comparisons with CUBIC- and iDISCO-based delipidation procedures, the FxClear-based procedure provided rapid, simple protocol with high RI contrast. Although CUBIC-based protocol provided slightly inferior RI contrasts, it provides simplest single step solution. However, because RI adjustment and de-lipidation is not separable in this approach, RI contrast is inevitably changing depending on the incubation time. The iDISCO-based procedure requires toxic materials and complex steps although this protocol also provided high RI contrast comparable to the FxClear-based procedure. Therefore, the de-lipidation methods can be selected based on the tissues and the purpose of the experiments.
We applied the FxClear technique as a strategy for designing the delipidation step optimized to enhance the structural contrast of the tissue. To achieve label-free imaging with thick tissue via conventional microscopes, we considered a chemical process, especially, RI matching. However, the RI values of the mounting medium showed no correlation with the tissue transparency (Figure S1). The clearing performance does not solely rely on the RI value of the medium but does reflect some chemical properties of the constituent. Among the RI matching solution, sugar-based RI medium derives moderate tissue transparency compared with RI media consisting of contrast reagents such as iodixanol. Iodixanol shows high hydrophilicity and good aqueous solubility due to the dimers of non-ionic tri-iodinated aromatic compounds (Priebe et al., 1999). Hydrophilic aromatic amides contributed to reducing the RI inhomogeneity due to the efficient solvation of the protein backbone amides (Tainaka et al., 2018). On the other hand, the hydrophilicity of the iodixanol also makes poor penetrability to the densely hydrophobic domains such as WH. Therefore, iodixanol seems to have a higher clearing contrast efficiency than fructose. In addition, iodixanol enhances the hyper-hydration of tissues and mildly expands the tissue size. The hyper-hydration effect of iodixanol is achieved by disruption of the charged ECM fiber structures and promotes media diffusivity. Conversely, the degree of hyper-hydration is affected by the hydrophilicity of the tissue. Accordingly, the tissue-clearing effect of the media would be different depending on the lipid content in the region. In comparison, fructose did not show overt tissue expansion but exhibited salt-dependent shrinkage in a given tissue.
Traumatic brain scar or Aβ deposits were efficiently detected by our protocol, especially using fructose solution. In the early phase of TBI, the injury core contained blood clots, the debris of dying cells and fibers, and acutely infiltrated immune cells, whereas ECM deposition was mild, while the blood clots were found to be cleared on day 7 of TBI. Fructose or iodixanol has no activity to remove heme-based pigments; therefore, colored blood in the tissue is recognized by trans-illumination imaging. Fructose, but not iodixanol, efficiently detected mild ECM remodeling in the early phase (3 days) TBI samples, and low light transmission was found in the TBI core region where blood clots were not detected. In our previous study with different protocols based on ACT clearing, we failed to achieve TBI core visualization at 3 days (Kim et al., 2018), indicating that our current protocol with fructose is more sensitive to the detection of brain trauma. Furthermore, our protocol provides sensitive enough contrast for recognizing micro deposits of Aβ in AD model mouse brain, suggesting the potential to use of label-free imaging for the diagnosis purposes.
Limitation of the study
Although our proposed method can be used to visualize lipid concentration and ECM deposition, which greatly affect the light scattering of the tissue, cellular heterogeneity was not recognized because we could not show the changes in cell composition in the TBI model or GBM implants. This may be because of the similar optical properties of the cells, regardless of their cellular identity, and improvement of signal detection with high-end optical devices may be required. In addition, we performed our OCT and SPIM imaging with home-made systems. Because imaging quality is greatly dependent on the optimization of imaging equipment, it was difficult to directly compare our procedures with several previously published procedures (Ren et al., 2017; Renier et al., 2016).
Resource availability
Lead contact
Further information and requests for resources and reagents should be directed to and will be fulfilled by the Lead Contact, Woong Sun (woongsun@korea.ac.kr).
Materials availability
This study did not generate new unique reagents.
Data and code availability
All data is available by contacting the lead author. The study did not generate unique code.
Methods
All methods can be found in the accompanying transparent methods supplemental file.
Acknowledgments
This research was supported by the Brain Research Program through the National Research Foundation (NRF), South Korea, which is funded by the Korean Ministry of Science, the ICT & Future Planning (NRF-2015M3C7A1028790, NRF-2017M3A9B3061308, NRF-2018R1D1A1A02086190 and NRF-2019M3E5D2A01063939). We also thank Dr Lee, Ju-Hyun for her assistance to produce 3D images.
Author contributions
W.S. designed the study and wrote the manuscript. B.L. and J.C. performed most of the tissue clearing and tissue immunolabeling. B.L. and E.L. performed confocal microscopy analysis. E.L. contributed data analysis compiled the manuscript. J.H.K. performed computational analysis. H.J.K., Y.G.K., and B.M.K. performed OCT imaging analysis and data interpretation. H.J.K. and S.G.K. conducted GBM model experimental system. K.K., Y.K., and K.C. produced SPIM imaging system and contributed to the imaging and analysis. All authors provided their feedback on the final manuscript.
Declaration of interests
K.C. is a shareholder and employee, K.K. and Y. K. are employees of Aligned Genetics, Inc.
Published: April 23, 2021
Footnotes
Supplemental information can be found online at https://doi.org/10.1016/j.isci.2021.102267.
Contributor Information
Eunsoo Lee, Email: eunssoo@ewha.ac.kr.
Woong Sun, Email: woongsun@korea.ac.kr.
Supplemental information
References
- Appel A.A., Anastasio M.A., Larson J.C., Brey E.M. Imaging challenges in biomaterials and tissue engineering. Biomaterials. 2013;34:6615–6630. doi: 10.1016/j.biomaterials.2013.05.033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aumann S., Donner S., Fischer J., Müller F. Springer; 2019. Optical coherence tomography (OCT): principle and technical realization; pp. 59–85. (High Resolution Imaging in Microscopy and Ophthalmology). Chapter 3. [DOI] [PubMed] [Google Scholar]
- Baek K., Jung S., Lee J., Min E., Jung W., Cho H. Quantitative assessment of regional variation in tissue clearing efficiency using optical coherence tomography (OCT) and magnetic resonance imaging (MRI): a feasibility study. Sci. Rep. 2019;9:1–10. doi: 10.1038/s41598-019-39634-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bizheva K.K., Unterhuber A., Hermann B.M., Povazay B., Sattmann H., Fercher A.F., Drexler W., Preusser M., Budka H., Stingl A. Imaging ex vivo healthy and pathological human brain tissue with ultra-high-resolution optical coherence tomography. J. Biomed. Opt. 2007;10:011006. doi: 10.1117/1.1851513. [DOI] [PubMed] [Google Scholar]
- Bolmont T., Bouwens A., Pache C., Dimitrov M., Berclaz C., Villiger M., Wegenast-Braun B.M., Lasser T., Fraering P.C. Label-free imaging of cerebral β-amyloidosis with extended-focus optical coherence microscopy. J. Neurosci. 2012;32:14548–14556. doi: 10.1523/JNEUROSCI.0925-12.2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brezinski M.E., Tearney G.J., Bouma B.E., Izatt J.A., Hee M.R., Swanson E.A., Southern J.F., Fujimoto J.G. Optical coherence tomography for optical biopsy: properties and demonstration of vascular pathology. Circulation. 1996;93:1206–1213. doi: 10.1161/01.cir.93.6.1206. [DOI] [PubMed] [Google Scholar]
- Choi J., Lee E., Kim J.H., Sun W. FxClear, A free-hydrogel electrophoretic tissue clearing method for rapid de-lipidation of tissues with high preservation of immunoreactivity. Exp. Neurobiol. 2019;28:436–445. doi: 10.5607/en.2019.28.3.436. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chung K., Wallace J., Kim S.-Y., Kalyanasundaram S., Andalman A.S., Davidson T.J., Mirzabekov J.J., Zalocusky K.A., Mattis J., Denisin A.K. Structural and molecular interrogation of intact biological systems. Nature. 2013;497:332–337. doi: 10.1038/nature12107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ertürk A., Becker K., Jährling N., Mauch C.P., Hojer C.D., Egen J.G., Hellal F., Bradke F., Sheng M., Dodt H.-U. Three-dimensional imaging of solvent-cleared organs using 3DISCO. Nat. Protoc. 2012;7:1983. doi: 10.1038/nprot.2012.119. [DOI] [PubMed] [Google Scholar]
- Habaza M., Kirschbaum M., Guernth-Marschner C., Dardikman G., Barnea I., Korenstein R., Duschl C., Shaked N.T. Rapid 3D refractive-index imaging of live cells in suspension without labeling using dielectrophoretic cell rotation. Adv. Sci. 2017;4:1600205. doi: 10.1002/advs.201600205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hama H., Kurokawa H., Kawano H., Ando R., Shimogori T., Noda H., Fukami K., Sakaue-Sawano A., Miyawaki A. Scale: a chemical approach for fluorescence imaging and reconstruction of transparent mouse brain. Nat. Neurosci. 2011;14:1481–1488. doi: 10.1038/nn.2928. [DOI] [PubMed] [Google Scholar]
- Hanna N., Saltzman D., Mukai D., Chen Z., Sasse S., Milliken J., Guo S., Jung W., Colt H., Brenner M. Two-dimensional and 3-dimensional optical coherence tomographic imaging of the airway, lung, and pleura. J. Thorac. Cardiovasc. Surg. 2005;129:615–622. doi: 10.1016/j.jtcvs.2004.10.022. [DOI] [PubMed] [Google Scholar]
- Huisken J., Stainier D.Y. Even fluorescence excitation by multidirectional selective plane illumination microscopy (mSPIM) Opt. Lett. 2007;32:2608–2610. doi: 10.1364/ol.32.002608. [DOI] [PubMed] [Google Scholar]
- Jensen K.H.R., Berg R.W. Advances and perspectives in tissue clearing using CLARITY. J. Chem. Neuroanat. 2017;86:19–34. doi: 10.1016/j.jchemneu.2017.07.005. [DOI] [PubMed] [Google Scholar]
- Ke M.-T., Fujimoto S., Imai T. SeeDB: a simple and morphology-preserving optical clearing agent for neuronal circuit reconstruction. Nat. Neurosci. 2013;16:1154–1161. doi: 10.1038/nn.3447. [DOI] [PubMed] [Google Scholar]
- Kim J.H., Jang M.J., Choi J., Lee E., Song K.D., Cho J., Kim K.-T., Cha H.-J., Sun W. Optimizing tissue-clearing conditions based on analysis of the critical factors affecting tissue-clearing procedures. Sci. Rep. 2018;8:1–11. doi: 10.1038/s41598-018-31153-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuwajima T., Sitko A.A., Bhansali P., Jurgens C., Guido W., Mason C. ClearT: a detergent-and solvent-free clearing method for neuronal and non-neuronal tissue. Development. 2013;140:1364–1368. doi: 10.1242/dev.091844. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Larina I., Carbajal E., Tuchin V., Dickinson M., Larin K. Enhanced OCT imaging of embryonic tissue with optical clearing. Laser Phys. Lett. 2008;5:476–479. doi: 10.1117/12.808055. [DOI] [Google Scholar]
- Lee E., Choi J., Jo Y., Kim J.Y., Jang Y.J., Lee H.M., Kim S.Y., Lee H.-J., Cho K., Jung N. ACT-PRESTO: rapid and consistent tissue clearing and labeling method for 3-dimensional (3D) imaging. Sci. Rep. 2016;6:18631. doi: 10.1038/srep18631. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee E., Kim H.J., Sun W. See-through technology for biological tissue: 3-dimensional visualization of macromolecules. Int. Neurourol. J. 2016;20:S15. doi: 10.5213/inj.1632630.315. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liebmann T., Renier N., Bettayeb K., Greengard P., Tessier-Lavigne M., Flajolet M. Three-dimensional study of Alzheimer’s disease hallmarks using the iDISCO clearing method. Cell Rep. 2016;16:1138–1152. doi: 10.1016/j.celrep.2016.06.060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morozov A.N., Il'ya V.T., Kamenskii V., Fiks I., Lazutkin A., Bezryadkov D., Ivanova A., Toptunov D., Anokhin K.V. Fibreoptic fluorescent microscopy in studying biological objects. Quan. Electron. 2010;40:842. doi: 10.1070/QE2010v040n09ABEH014344. [DOI] [Google Scholar]
- Murata T., Honda T., Egawa G., Kitoh A., Dainichi T., Otsuka A., Nakajima S., Kore-eda S., Kaku Y., Nakamizo S. Three-dimensional evaluation of subclinical extension of extramammary Paget disease: visualization of the histological border and its comparison to the clinical border. Br. J. Dermatol. 2017;177:229–237. doi: 10.1111/bjd.15282. [DOI] [PubMed] [Google Scholar]
- Nojima S., Susaki E.A., Yoshida K., Takemoto H., Tsujimura N., Iijima S., Takachi K., Nakahara Y., Tahara S., Ohshima K. CUBIC pathology: three-dimensional imaging for pathological diagnosis. Sci. Rep. 2017;7:1–14. doi: 10.1038/s41598-017-09117-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olson E., Levene M.J., Torres R. Multiphoton microscopy with clearing for three dimensional histology of kidney biopsies. Biomed. Opt. Express. 2016;7:3089–3096. doi: 10.1364/BOE.7.003089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pan C., Cai R., Quacquarelli F.P., Ghasemigharagoz A., Lourbopoulos A., Matryba P., Plesnila N., Dichgans M., Hellal F., Ertürk A. Shrinkage-mediated imaging of entire organs and organisms using uDISCO. Nat. Methods. 2016;13:859–867. doi: 10.1038/nmeth.3964. [DOI] [PubMed] [Google Scholar]
- Priebe H., Aukrust A., Bjørsvik H.R., Tønseth C., Wiggen U. Stability of the X-ray contrast agent iodixanol= 3, 3′, 5, 5′-tetrakis (2, 3-dihydroxypropylcarbamoyl)-2, 2′, 4, 4′, 6, 6′-hexaiodo-N, N′-(2-hydroxypropane-1, 3-diyl)-diacetanilide towards acid, base, oxygen, heat and light. J. Clin. Pharm. Ther. 1999;24:227–235. doi: 10.1046/j.1365-2710.1999.00225.x. [DOI] [PubMed] [Google Scholar]
- Ren J., Choi H., Chung K., Bouma B.E. Label-free volumetric optical imaging of intact murine brains. Sci. Rep. 2017;7:46306. doi: 10.1038/srep46306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Renier N., Adams E.L., Kirst C., Wu Z., Azevedo R., Kohl J., Autry A.E., Kadiri L., Venkataraju K.U., Zhou Y. Mapping of brain activity by automated volume analysis of immediate early genes. Cell. 2016;165:1789–1802. doi: 10.1016/j.cell.2016.05.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Renier N., Wu Z., Simon D.J., Yang J., Ariel P., Tessier-Lavigne M. iDISCO: a simple, rapid method to immunolabel large tissue samples for volume imaging. Cell. 2014;159:896–910. doi: 10.1016/j.cell.2014.10.010. [DOI] [PubMed] [Google Scholar]
- Sudheendran N., Mohamed M., Ghosn M.G., Tuchin V.V., Larin K.V. Assessment of tissue optical clearing as a function of glucose concentration using optical coherence tomography. J. Innov. Opt. Health Sci. 2010;3:169–176. doi: 10.1142/S1793545810001039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sung Y., Choi W., Fang-Yen C., Badizadegan K., Dasari R.R., Feld M.S. Optical diffraction tomography for high resolution live cell imaging. Opt. Express. 2009;17:266–277. doi: 10.1364/OE.17.000266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Susaki E.A., Tainaka K., Perrin D., Kishino F., Tawara T., Watanabe T.M., Yokoyama C., Onoe H., Eguchi M., Yamaguchi S. Whole-brain imaging with single-cell resolution using chemical cocktails and computational analysis. Cell. 2014;157:726–739. doi: 10.1016/j.cell.2014.03.042. [DOI] [PubMed] [Google Scholar]
- Tainaka K., Kuno A., Kubota S.I., Murakami T., Ueda H.R. Chemical principles in tissue clearing and staining protocols for whole-body cell profiling. Annu. Rev. Cell Dev. Biol. 2016;32:713–741. doi: 10.1146/annurev-cellbio-111315-125001. [DOI] [PubMed] [Google Scholar]
- Tainaka K., Murakami T.C., Susaki E.A., Shimizu C., Saito R., Takahashi K., Hayashi-Takagi A., Sekiya H., Arima Y., Nojima S. Chemical landscape for tissue clearing based on hydrophilic reagents. Cell Rep. 2018;24:2196–2210. e2199. doi: 10.1016/j.celrep.2018.07.056. [DOI] [PubMed] [Google Scholar]
- Ueda H.R., Ertürk A., Chung K., Gradinaru V., Chédotal A., Tomancak P., Keller P.J. Tissue clearing and its applications in neuroscience. Nat. Rev. Neurosci. 2020;21:61–79. doi: 10.1038/s41583-019-0250-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang B., Treweek J.B., Kulkarni R.P., Deverman B.E., Chen C.-K., Lubeck E., Shah S., Cai L., Gradinaru V. Single-cell phenotyping within transparent intact tissue through whole-body clearing. Cell. 2014;158:945–958. doi: 10.1016/j.cell.2014.07.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yi J., Backman V. Imaging a full set of optical scattering properties of biological tissue by inverse spectroscopic optical coherence tomography. Opt. Lett. 2012;37:4443–4445. doi: 10.1364/OL.37.004443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhong H., Guo Z., Wei H., Si J., Guo L., Zhao Q., Zeng C., Xiong H., He Y., Liu S. Enhancement of permeability of glycerol with ultrasound in human normal and cancer breast tissues in vitro using optical coherence tomography. Laser Phys. Lett. 2010;7:315–320. doi: 10.1002/lapl.201010006. [DOI] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All data is available by contacting the lead author. The study did not generate unique code.