Abstract
Background
In COVID-19 patients, information regarding superinfection, antimicrobial assessment, and the value of metagenomic sequencing (MS) could help develop antimicrobial stewardship.
Method
This retrospective study analyzed 323 laboratory-confirmed COVID-19 patients for co-infection rate and antimicrobial usage in the Shanghai Public Health Clinical Center (SPHCC) from January 23rd to March 14th 2020. The microbiota composition was also investigated in patients with critically severe COVID-19.
Results
The total population co-infection rate was 17/323 (5.3%) and 0/229 (0), 4/78 (5.1%), and 13/16 (81.3%) for the mild, severe, and critically severe subgroups, respectively. Proven fungal infection was significantly associated with a higher mortality rate (p = 0.029). In critically severe patients, the rate of antimicrobials and carbapenem usage were 16/16 (100%) and 13/16 (81.3%), respectively, in which the preemptive and empiric antimicrobial days accounted for 51.6% and 30.1%, respectively. Targeted therapy only accounted for 18.3%. MS was implemented to detect non-COVID-19 virus co-existence and the semi-quantitative surveillance of bacteremia, with clear clinical benefit seen in cases with MS-based precision antimicrobial management. Airway microbiome analysis suggested that the microbiota compositions in critically severe COVID-19 patients were likely due to intubation and mechanical ventilation.
Conclusions
In the SPHCC cohort, we observed a non-negligible rate of super-infection, especially for the critically ill COVID-19 patients. Fungal co-infection requires intensive attention due to the high risk of mortality, and the clinical benefit of MS in guiding antimicrobial management warrants further investigation.
Keywords: COVID-19, Superinfection, Antimicrobials, Escalation, Metagenomic sequencing
Introduction
According to the Surviving Sepsis Campaign guidelines, information regarding bacterial co-infection or secondary infection in COVID-19 patients is currently limited. The recommendation for empirical antimicrobials use remains unestablished.1 , 2 A recent meta-analysis of 22 studies reported that overall bacterial co-infection rates in COVID-19 were 7% and 14% in the intensive care unit (ICU). On the contrary, the prevalence rate of antibiotics prescription to COVID-19 patients was 70%–100% with different severity.3 Therefore, further studies focusing on super-infection and antibiotic usage are needed to improve COVID-19 antimicrobial stewardship.
Novel techniques for pathogen detection in COVID-19 cases could be critical for versatile co-infection surveillance and appropriate antimicrobial therapy. Microbial metagenomic next-generation sequencing (MS) emerges as a fast-evolving technology, which allows the identification of pathogens and microbiome information simultaneously within 24 h. Based on previous research4 , 5 and our experience, MS has a superior detection sensitivity than conventional microbial testing methods, thus being more advantageous in the present-day microbial surveillance and antimicrobial appraisals.
In this Shanghai cohort of 323 COVID-19 patients, we aimed to evaluate the pathogens of co-infection, antimicrobial usage, and airway microbiome characteristics with MS testing.
Materials and methods
Study population
This observational study was carried out at the Shanghai Public Health Clinical Center (SPHCC), the referral hospital for all laboratory-confirmed COVID-19 patients in Shanghai. Medical history of consecutive COVID-19 cases between January 23rd and April 28th, 2020, were collected and stratified as mild, severe, and critically severe according to the Diagnosis and Treatment of Novel Coronavirus Pneumonia Guidelines.6 The conventional method for pathogen identification was applied in all patients with suspected co-infection. Starting from February 22nd, plasma and airway samples of 10 critically severe cases were tested with MS according to clinical practitioners' judgment, with MS-based microbial surveillance conducted in approximately 5-day intervals. Co-infection incidence rates, as well as the pathogen distribution, were compared in cases with different severity. Additionally, microbe-based antimicrobial evaluation and MS-based airway microbiome assessment were studied in critically severe patients. The study protocol was approved and supervised by the institutional ethics board of SPHCC.
Metagenomic next-generation sequencing and analysis
Plasma and airway samples were both tested by MS. The experiment process was illustrated in our former research.5 Briefly, RNA and DNA were extracted using a TIANamp Micro DNA Kit (DP316, Tiangen Biotech). RNA reverse transcription and DNA libraries construction through an end-repair method were processed. Then, the procedure for DNB was used before analysis using the BGI Sequencer platform (BGI Genomics, Shenzhen, China). High-quality sequencing data were generated by removing low-quality, low-complexity, and short (length <35 bp) reads, followed by computational subtraction of human host sequences mapped to the human reference genome (hg19) using Burrows-Wheeler alignment. The filtered data were classified by simultaneous alignment to 4 microbial genome databases consisting of viruses, bacteria, fungi, and parasites. As for criteria for a positive MS result, coverage rate and abundance were used as the measurement parameter.
Definition
Co-infection
According to criteria of the International Sepsis Forum,7 low or moderate clinical suspicion of infection with microbiological evidence by either culture or MS was defined as co-infection, and cases with definite symptoms of infection and isolation of likely pathogens by culture were classified as proven infections. Some of the critically ill patients might experience multiple co-infection events. Indicators of clinical suspicion based on infectious symptoms such as purulent sputum, significant procalcitonin (PCT) elevation, recurrent fever, worsened oxygenation index, radiology signs, and so on.
Virus colonization and activation
Virus activations were diagnosed if viral load by MS increased 5 times in less than 1 week, or the virus detections were considered colonization.8
Statistical analysis
Comparative analysis was conducted by Fisher exact test or Kruskal–Wallis test for variables where appropriate. From sequencing reads obtained from MS, we calculated alpha diversity by Shannon index. Principal coordinate analysis plots were applied for visualization of Unifrac distances between groups. We selected bacterial genera that rank top 90% accumulative abundance from each sample and performed samples clustering using Euclidean distances with heatmap package in R (ggplot 2). Permutational multivariate analysis of variance (PERMANOVA) was used for comparing the contribution of various factors to the airway microbiome difference among various groups.
Results
Co-infection rate in mild, severe, and critically severe cases
Among 323 patients with COVID-19 pneumonia enrolled in our study, 17/323 (5.3%) were diagnosed with co-infection. Co-infection rates in mild, severe, and critically severe groups were 0/229 (0), 4/78 (5.1%), and 13/16 (81.3%), respectively (Fig. 1 A). In 16 critically severe cases, demographic information and clinical parameters, such as extracorporeal membrane oxygenation (ECMO), mortality, and glucocorticoid usage are also described in Table 1 .
Figure 1.
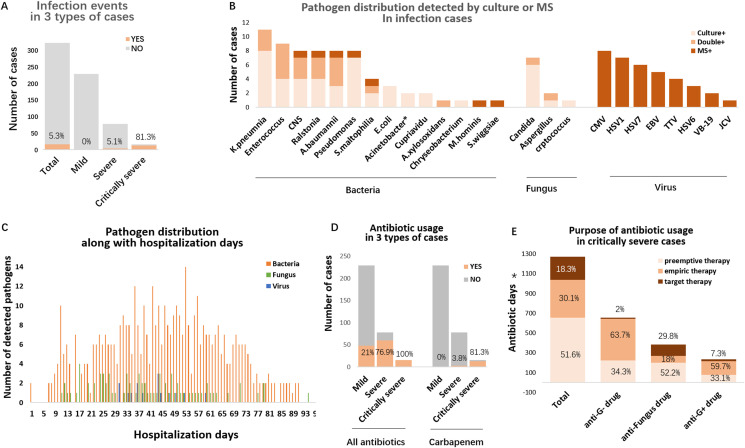
Information on co-infection and antimicrobial usage. A. Incidence rate of co-infection events in mild, severe, and critically severe cases. B. Pathogens of co-infection events detected by MS or culture. culture +: positive only by culture, MS+: positive only by MS, double+: positive by both culture and MS, CNS: coagulase-negative staphylococcus. ∗ Acinetobacter other than A. baumannii. C. Positive pathogen number along with hospitalization day. The peak time of bacterial co-infection events was between 40 and 60 hospitalized days. D. Rate of overall antibiotic and carbapenem usage in mild, severe, and critically severe cases. E. Antibiotic days for different purposes of antibiotic usage in critically severe cases. ∗Because critically severe patients were hospitalized for a prolonged time and received frequent antimicrobial modulation, antimicrobials days were used to compare various therapeutic purposes. G+, gram-positive microbes; G-, gram-negative microbes.
Table 1.
Demographic data.
| Total, n = 16 |
Die, n = 7 |
Survive, n = 9 |
p value | |
|---|---|---|---|---|
| Males, No(%) | 13 (81.3) | 4 (57.1) | 9 (100) | 0.247 |
| Age, median (IQR), y | 70.5 (25–88) | 79 (25–88) | 68 (57–80) | 0.186 |
| Median (IQR), day | ||||
| Length of hospital stay | 50.5 (4–98) | 49 (4–69) | 52 (26–98) | 0.203 |
| Ventilation | 35.5 (3–87) | 49 (3–60) | 23 (8–87) | 0.916 |
| ECMO | 39 (22–47) | 37.5 (26–47) | 40 (22–47) | 0.858 |
| Glucocorticoids usage | 15 (0–63) | 16 (0–63) | 14 (5–40) | 1 |
| Duration of positive MERS-COV2 | 36 (13–53) | 38.5 (36–41) | 30 (13–53) | 0.354 |
| Infection indicators, median (IQR) | ||||
| Significant PCT elevation | 1.5 (1–3) | 1 (1–3) | 2 (1–3) | 0.954 |
| Fever | 1 (0–5) | 1.5 (0–5) | 1 (0–3) | 0.583 |
| Decreasing oxygenation index | 1 (1–4) | 1 (1–2) | 1 (1–4) | 0.641 |
| CT indication of superinfection | 0 (0–4) | 0 (0–1) | 0 (0–4) | 0.953 |
| CD4 count<500 | 2 (1–6) | 2 (1–6) | 2 (1–5) | 0.584 |
| No.(%) | ||||
| Comorbiditiesa | 11 (68.8) | 4 (57.1) | 7 (77.8) | 1 |
| Coinfection eventb | 12 (75) | 5 (71.4) | 7 (77.8) | 1 |
| Proven infection | 5 (31.3) | 4 (57.1) | 1 (11.1) | 0.049 |
| Pathogen of improven infection | ||||
| Enterobacteriaceae | 2 (40) | 1 (25) | 1 (100) | 0.849 |
| G+ microbe | 1 (20) | 1 (25) | 0 (0) | 0.242 |
| Fungus | 3 (60) | 3 (75) | 0 (0) | 0.029 |
| Anti-CD treatmentc | 0 (0–24) | 0 (0–24) | 0 (0–8) | 0.274 |
Abbreviations: ECMO, extracorporeal membrane oxygenation; NFB,nonfermentative bacteria; IQR,interquartile range; CD, Clostridium difficile;G+, gram positive microbe; G-, gram negative microbe; MS, metagenomic sequencing.
Comorbidities listed here are hypertension,cerebral infarction,hypothyroidism, cardiovascular disease, kidney disease, cardiovascular disease, cancer, pulmonary disease, schizophrenia, diabetes.
Criteria based on the International Sepsis Forum definitions.
Empirical treatment according to clinical sign.
Pathogens of co-infection by culture and MS
In all the 17 co-infection patients, pathogen detection was achieved by culture method. In 10 of 17 cases, MS was also applied for comprehensive microbial screening. The highest microbe yield was from the airway, followed by urinary tract and plasma (Fig. S1).
Bacteria
For bacterial infection, Klebsiella pneumonia was most frequently detected, followed by Enterococcus and Coagulase-negative Staphylococcus. Fastidious pathogens co-infection such as S.wiggsiae and M. hominis were also revealed by MS.
Fungi
As for the fungal infection events, we found a total of 7 Candida, 2 Aspergillus, and 1 Cryptococcus co-infection event (Fig. 1B). Six patients were diagnosed with candiduria, and all survived. The other 3 patients all died in ICU: one patient was diagnosed with invasive pulmonary Aspergillus, one patient with candidemia, and one patient with bloodstream infection of Cryptococcus and Aspergillus. The latter 3 cases were considered proven fungal co-infection events, significantly associated with the mortality rate as revealed by the univariate analysis (p = 0.029, Table 1).
Virus
MS revealed a surprisingly high prevalence of virus colonization (8/10, 80%) and activation (5/10, 50%), including cytomegalovirus (CMV), herpes simplex virus (HSV), Epstein–Barr virus (EBV), Torque teno virus (TTV), Human Parvovirus B19 (HPVB19), and JC polyomavirus (JCV) (Figure 1, Figure 2 A). Interestingly, MS showed that 5 patients had experienced multiple episodes of virus activation during the prolonged ICU stay (Fig. 2B and C).
Figure 2.
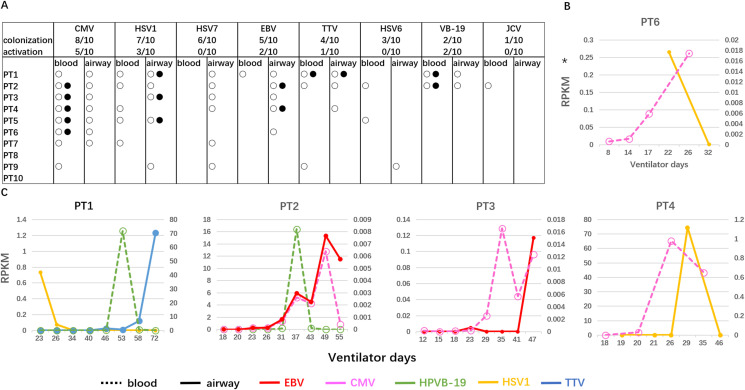
Virus prevalence in 10 critically severe cases. A. Virus prevalence in 10 critically severe patients by MS technique. ○Virus colonization,●virus activation, PT: patient number. B, C. Moreover, MS showed that 5 patients had experienced multiple episodes of virus activation during prolonged ventilation. ∗RPKM: reads number/(gene length/1000∗total numbers/1000000). RPKM can be statistically compared among different virus types. Abbreviation. CMV: cytomegalovirus, HSV: herpes simplex virus, EBV: Epstein–Barr virus, TTV: Torque teno virus, HPVB19: Human Parvovirus B19, and JCV: JC polyomavirus.
Pathogen distribution along with hospitalization days
We found that bacterial co-infection events' peak time was between 40 and 60 hospitalized days, while fungal and virus infection events occurred in a sporadic manner (Fig. 1C).
Assessment of antimicrobial usage
The rate of all antimicrobials and carbapenem usage were 48/229 (21.0%) and 0% in mild patients, 60/78 (76.9%) and 3/78 (3.8%) in severe patients, 16/16 (100%) and 13/16 (81.3%) in critically severe cases (Fig. 1D).
According to the treatment purpose, antimicrobial usage in critically ill cases was classified into preemptive, empiric, and target therapy. Preemptive and empiric antimicrobial days accounted for 51.6% and 30.1%, while targeted therapy days only accounted for 18.3% of the antimicrobial usage (Fig. 1E).
The clinical benefit of MS testing for guiding antimicrobial therapy
Due to the high sensitivity and semi-quantitative nature of MS, we observed a potential clinical benefit of the MS-guided strategy for bloodstream infection surveillance and antimicrobial decision in the abovementioned 10 critically severe patents. For instance, in a patient (PT3) with elevated procalcitonin level on ventilator day 12th, the airway detection of E. faecium, S. maltophilia, and K. pneumonia by both culture and MS failed to clarify whether these bacteria were colonization or pathogenic. Despite the negative result from multiple blood cultures, MS of plasma suggested a pronounced elevation of E. faecium relative abundance, followed by a drop-down after vancomycin target therapy. A relief of the patient's symptoms and procalcitonin level (red zone in Fig. 3 A) was also noticed. Conversely, in another patient (PT2) with elevated procalcitonin and recurrent airway A. baumanii detections, we observed a low plasma level of A. baumanii by MS, which remained minimally changed at least two weeks. The procalcitonin then gradually alleviated following a simple wait-and-see strategy, preventing broad-spectrum antibiotics' unnecessary use (red zone in Fig. 3B).
Figure 3.
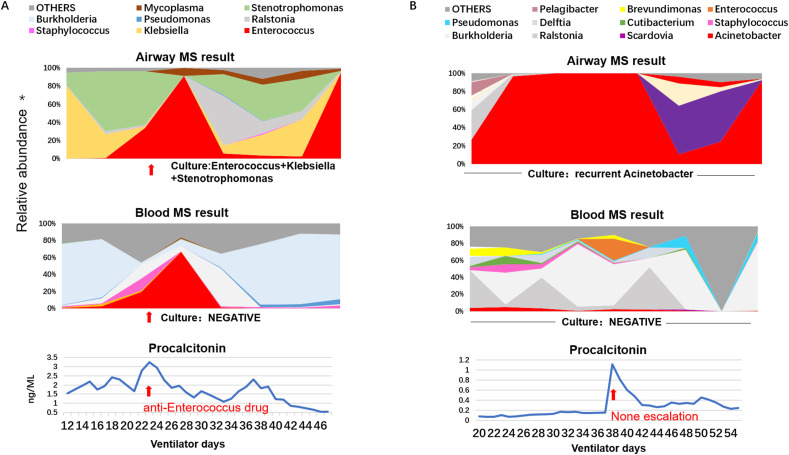
Clinical benefit of MS in typical patients. A. PT3. The red arrow indicates the time point at which procalcitonin is elevated. On this point, airway pathogens identified by MS and culture were Enterococcus, Stenotrophomonas, and Klebsiella (red, green, and yellow zone in MS). Blood pathogens by culture were negative, while MS surveilled increment of Enterococcus abundance (red zone), which may be the culprit of procalcitonin elevation. Target therapy of vancomycin decreased procalcitonin level. B. PT2. Conversely, A. baumanii was continuously detected in the airway by culture and MS (red zone). However, MS surveilled a stably low level of blood abundance (red zone), which restrained from using unnecessary broad-spectrum antibiotics despite procalcitonin fluctuation. ∗The bacteria loads were described as relative abundance in MS. Some pathogens, such as Burkholderia, Ralstonia and Delftia, were considered contamination rather than true pathogens.
MS also confirmed multiple virus activation in 5 patients (Fig. 2B and C). The presence of fever and/or elevated inflammatory biomarkers in these patients might otherwise lead to empirical antibiotic use, commonly seen in the ICUs.
Comparative analysis of airway microbiome by MS
MS analysis of 50 COVID-19 airway samples demonstrated minimal differences in the microbiome between patients with different prognoses and patients with a different number of ventilator days (Fig. S2). Further comparison of bronchoalveolar fluid (BALF) and endotracheal aspiration (ETA) showed no difference in the diversity profile and microbiome composition of airway genera between the two sample types (p = 0.77), indicating that sampling of BALF is not superior to ETA as a non-invasive procedure (Fig. S3 and S4).
Furthermore, we sought to explore the similarity in airway microbiomes of critically severe COVID-19 (all with intubation) patients with those of other patients, including those intubated for non-COVID-19 diseases (n = 20) or non-incubation viral pneumonia (n = 31), and non-incubation non-infectious diseases (n = 23). Hierarchical clustering analysis demonstrated that the microbial composition of the critically severe COVID-19 was closely clustered and was dissimilar to the microbiomes of other types of viral pneumonia (non-incubation) or non-infectious diseases (non-incubation). In contrast, the latter two were interleaved without obvious segregation (Fig. 4 A). Therefore, we hypothesized that the airway microbiome of critically severe COVID-19 patients' airway microbiome was dominated by intubation, rather than the COVID-19 per se. As expected, the alpha diversity of critically severe-COVID-19 patients was significantly lower than that of non-intubated patients (other viral pneumonia and non-infectious disease groups combined) (p < 0.001, p = 0.0044); however, it was similar to that of the intubated non-COVID-19 group (p = 0.75) (Fig. 4B). Similarly, PCoA analysis indicated that there was a significant difference between the three groups (p < 0.001), with the greatest difference observed between the non-intubated patients versus the other two groups with intubation (Fig. 4C). Using the PERMANNOVA test, we found that the R-square between COVID-19 patients and intubated non-COVID-19 patients was 0.095 (p = 0.001), significantly lower than that between COVID-19 patients and non-intubated patients (R-square = 0.173, p = 0.001). Furthermore, the comparison of the relative abundance composition at the genera level revealed a higher relative abundance of Acinetobacter (p < 0.001), Klebsiella (p = 0.001), Pelomonas (p = 0.005), Ralstonia (p < 0.0001), and Sphingomonas (p = 0.005) in COVID-19 patients. Conversely, the relative abundance of Actinomyces (p = 0.002), Haemophilus (p = 0.002), Neisseria (p < 0.0001), Prevotella (p < 0.001), Streptococcus (p < 0.001) and Veillonella (p < 0.001) was less in the COVID-19 samples.
Figure 4.
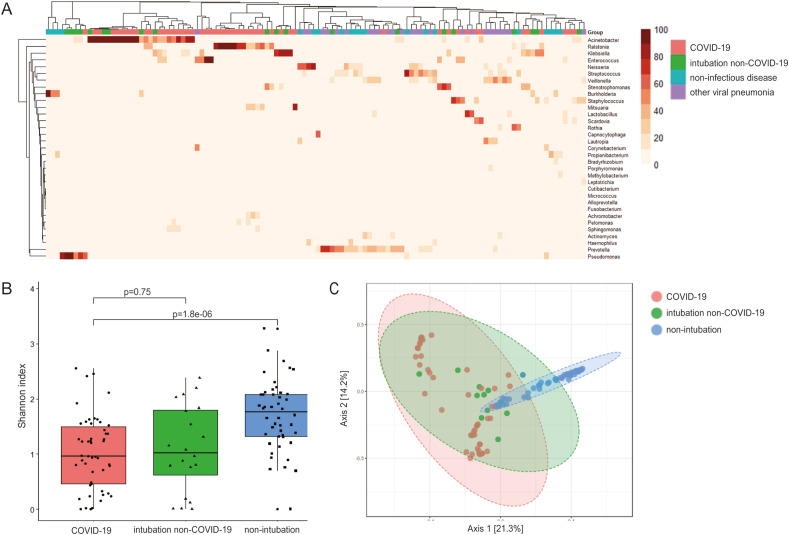
Comparative analysis of airway microbiome. A. The heatmap visualization of relative abundance at the genera level. Unsupervised hierarchical clustering reflected high relative abundance (red) of genera (rows) for each patient (columns). Column colors above the heatmap indicate whether a patient belonged to the COVID-19 (red), intubation-non-COVID-19 (green) or non-COVID-19 cohort (other viral pneumonia with blue and non-infectious disease with purple). Hierarchical clustering analysis demonstrated that the microbial composition of the critically severe COVID-19 was closely clustered, dissimilar to the other viral pneumonia or non-infectious disease. In contrast, the latter two were interleaved without obvious segregation. B. comparison of airway microbiome α-diversity. Shannon diversity index was used in subjects. Alpha diversity of critically severe-COVID-19 was significantly lower than the non-intubation cases (p < 0.001, p = 0.0044), but similar to the intubation non-COVID-19 group (p = 0.75). C. Principal coordinate analysis based on unweighted UniFrac distance. Separation among three groups was distinct (p < 0.001), while 2 intubation groups were much closer to each other than to the non-COVID-19 group.
Discussion
To the best of our knowledge, this is the first report focusing on super-infection in COVID-19 patients with the application of MS and evaluating the underlying pathogens and antimicrobial strategy comprehensively and systematically. MS is an unbiased tool for detecting versatile microbes, providing more comprehensive information for co-infection and antimicrobial evaluation.
We found that overall super-infections occurred in 5.3% of the general patient cohort and were commonly seen in critically ill patients with an infection rate of 81.3%. Additionally, our study included a prolonged follow-up duration; for the first time, it proposed 40 to 60 hospitalization days as the peak time of common bacteria detection, unlike the sporadic incidence of viral or fungal co-infection. Thus, antimicrobials were mainly reserved for critically ill patients on mechanical ventilation (100% for overall antibiotics and 81.3% for carbapenem), with the rate significantly lower in mild-severe patients (35.2%). Furthermore, in critically severe cases, target therapy with definite microbial evidence accounted for a small fraction of antibiotic days. Our results, combined with other reports about co-infection of COVID-19,9, 10, 11, 12, 13 reinforced the challenge of appropriate antibiotics use for critically ill cases and the need for a more precise antimicrobial strategy for these patients.
Notably, we found an incidence rate of proven fungal infection in our cohort (3/16) was relatively lower than that in previous studies (26%–33% in ICU patients),14 , 15 which may be potentially attributable to the frequent pre-emptive antifungal therapy in our cohort. Besides, we observed a significant association of fungal coinfection with unfavorable survival prognosis in the critically ill COVID-19. However, the potential causal relationship between fungal co-infection and mortality needs to be interpreted with caution since fungus detection might indicate extremely immunocompromised status with a poor prognosis in such patients. Therefore, whether active antifungal therapy could improve the prognosis warrants further investigation.
Due to the invasive support system, immunocompromised status, and frequent glucocorticoid usage in the critically ill COVID-19 patients, the prescription and escalation of broader antimicrobial were frequent in ICUs. Due to the relatively low sensitivity of microbial culture, it is common to implement a procalcitonin-based strategy for infection assessment and antimicrobial (de)escalation. However, such a strategy is empirical and has been challenged by some authors,16 , 17 due to the potential biases led by various factors, such as simultaneous supportive care, self-limitation of the disease, and inter-individual heterogeneity. In our study, we proposed, as a proof-of-concept, that combining procalcitonin and MS detection of multiple bio-specimens in short intervals is a feasible and efficient approach for the decisions of initiation or withdrawal of antimicrobial in critically severe patients. In our cohort, we reported the successful management of multiple cases. The semi-quantitative surveillance of microbes by MS enabled “timely initiate” or “rule out” antibiotic prescription for secondary bloodstream infection. In this setting, MS is specifically advantageous to traditional microbial culture, which has limited sensitivity in detecting fastidious or low viable microorganisms in the bloodstream.18
Consistent with our former study,5 MS revealed a surprisingly high rate of non-COVID-19 viral infection events in prolonged ICU dwellers. In addition to HSV and CMV viruses, we also detected TTV and HPVB19 activation, which is seldomly covered by commercially available molecular panels, and might be underestimated in previous reports. Notably, our results also suggested that patients might experience multiple viral activation periods in the ICUs, which has never been reported before. However, more studies are needed to clarify the significance and optimal management of non-COVID-19 viral infection in critically severe COVID-19 patients.
Lung microbiota alteration has recently gained increasing popularity due to its mechanistic implication in the pathogenesis of pneumonia,19 and its association with prognostic outcomes in mechanically ventilated ICU patients.20 Our study is the first report of the airway microbiome in critically ill COVID-19 patients on the metagenomic level. We found that such patients' microbiome was characterized by low diversity and more richness in non-ferment bacteria (Acinetobacter, Pelomonas, Ralstonia, and Sphingomonas). Remarkably, we proposed that these changes might be attributed to intubation and mechanical ventilation rather than COVID-19 pneumonia per se. This highlights the importance of reducing and/or shortening mechanical ventilation duration for maintaining normal airway microbiome characteristics, thus potentially preventing co-infections in COVID-19 patients.
Conclusion
In the SPHCC cohort, we observed a non-negligible rate of super-infection, especially for the critically ill COVID-19 patients. Specifically, fungal co-infection requires intensive attention due to the high risk of mortality in critically severe cases. Additionally, MS might be a feasible and efficient adjunctive tool for precision antimicrobial management in mechanically ventilated COVID-19 patients in ICUs.
Funding
Financial support from Shanghai Science and Technology Committee for restraining COVID-19 from becoming severe and number is 20411950500 and National Nature and Science Funding for precision diagnosis by metagenomics and number is 81902107.
Declaration of competing interest
The authors have disclosed that they do not have any potential conflicts of interest.
Acknowledgment
We thank the Shanghai Municipal Center for Disease Control and Prevention and BGI (The Beijing Genomics Institute) China for their support in the MS data.
Footnotes
This study was performed in the Shanghai Public Health Clinical Center.
Supplementary data to this article can be found online at https://doi.org/10.1016/j.jmii.2021.03.015.
Appendix A. Supplementary data
The following is the Supplementary data to this article:
References
- 1.Alhazzani W., Moller M.H., Arabi Y.M. Surviving sepsis Campaign: guidelines on the management of critically ill adults with coronavirus disease 2019 (COVID-19) Crit Care Med. 2020;48(6):e440–e469. doi: 10.1097/CCM.0000000000004363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Alhazzani W., Moller M.H., Arabi Y.M. Surviving Sepsis Campaign: guidelines on the management of critically ill adults with Coronavirus Disease 2019 (COVID-19) Intensive Care Med. 2020;46(5):854–887. doi: 10.1007/s00134-020-06022-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Lansbury L., Lim B., Baskaran V. Co-infections in people with COVID-19: a systematic review and meta-analysis. J Infect. 2020;81(2):266–275. doi: 10.1016/j.jinf.2020.05.046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Yang L., Haidar G., Zia H. Metagenomic identification of severe pneumonia pathogens in mechanically-ventilated patients: a feasibility and clinical validity study. Respir Res. 2019;20(1):265. doi: 10.1186/s12931-019-1218-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Miao Q., Ma Y., Wang Q. Microbiological diagnostic performance of metagenomic next-generation sequencing when applied to clinical practice. Clin Infect Dis. 2018;67(suppl_2):S231–S240. doi: 10.1093/cid/ciy693. [DOI] [PubMed] [Google Scholar]
- 6.新型冠状病毒肺炎诊疗方案(试行第七版) [J]. 2020. [Google Scholar]
- 7.Calandra T., Cohen J. The international sepsis forum consensus conference on definitions of infection in the intensive care unit. Crit Care Med. 2005;33(7):1538–1548. doi: 10.1097/01.ccm.0000168253.91200.83. [DOI] [PubMed] [Google Scholar]
- 8.Zou X., Tang G., Zhao X. Simultaneous virus identification and characterization of severe unexplained pneumonia cases using a metagenomics sequencing technique. Sci China Life Sci. 2017;60(3):279–286. doi: 10.1007/s11427-016-0244-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Huang C., Wang Y., Li X. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet. 2020;395(10223):497–506. doi: 10.1016/S0140-6736(20)30183-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Wu C., Chen X., Cai Y. Risk factors associated with acute respiratory distress syndrome and death in patients with coronavirus disease 2019 pneumonia in Wuhan, China. JAMA Intern Med. 2020 Jul 1;180(7):934–943. doi: 10.1001/jamainternmed.2020.0994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Yang X., Yu Y., Xu J. Clinical course and outcomes of critically ill patients with SARS-CoV-2 pneumonia in Wuhan, China: a single-centered, retrospective, observational study. Lancet Respir Med. 2020;8(5):475–481. doi: 10.1016/S2213-2600(20)30079-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Chen N., Zhou M., Dong X. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet. 2020;395(10223):507–513. doi: 10.1016/S0140-6736(20)30211-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Wang Z., Yang B., Li Q. Clinical features of 69 cases with coronavirus disease 2019 in Wuhan, China. Clin Infect Dis. 2020:a272. doi: 10.1093/cid/ciaa272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Rodriguez-Goncer I., Thomas S., Foden P. Invasive pulmonary aspergillosis is associated with adverse clinical outcomes in critically ill patients receiving veno-venous extracorporeal membrane oxygenation. Eur J Clin Microbiol Infect Dis. 2018;37(7):1251–1257. doi: 10.1007/s10096-018-3241-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Koehler P., Cornely O.A., Böttiger B.W. COVID-19 associated pulmonary aspergillosis. Mycoses. 2020;63(6):528–534. doi: 10.1111/myc.13096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Papazian L., Klompas M., Luyt C. Ventilator-associated pneumonia in adults: a narrative review. Intensive Care Med. 2020 May;46(5):888–906. doi: 10.1007/s00134-020-05980-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Albrich W.C., Harbarth S. Pros and cons of using biomarkers versus clinical decisions in start and stop decisions for antibiotics in the critical care setting. Intensive Care Med. 2015;41(10):1739–1751. doi: 10.1007/s00134-015-3978-8. [DOI] [PubMed] [Google Scholar]
- 18.Brenner T., Decker S.O., Grumaz S. Next-generation sequencing diagnostics of bacteremia in sepsis (Next GeneSiS-Trial): study protocol of a prospective, observational, noninterventional, multicenter, clinical trial. Medicine (Baltim) 2018;97(6):e9868. doi: 10.1097/MD.0000000000009868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Zakharkina T., Martin-Loeches I., Matamoros S. The dynamics of the pulmonary microbiome during mechanical ventilation in the intensive care unit and the association with occurrence of pneumonia. Thorax. 2017;72(9):803–810. doi: 10.1136/thoraxjnl-2016-209158. [DOI] [PubMed] [Google Scholar]
- 20.Dickson R.P., Schultz M.J., van der Poll T. Lung microbiota predict clinical outcomes in critically ill patients. Am J Respir Crit Care Med. 2020;201(5):555–563. doi: 10.1164/rccm.201907-1487OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.