Abstract
Background:
Lignin is the largest natural aromatic polymer in nature and is also a unique aromatic-based biopolymer, accounting for nearly 30% of the earth’s organic carbon. Generally, lignin is regarded as waste and is mainly used as a low- value fuel that is burned to generate heat and energy to solve the problem of biomass waste; for this obstacle of lignin, highly efficient biodegradation plays a critical role in developing an environmentally friendly technique for lignin biotransformation.
Objectives:
This study intends to isolate and purify several microbial strains from nature. It also explores how their lignin degradation is able to enhance the biodegradation and recycling of biomass and the reclamation of lignin in wastewater from pulp and paper mills.
Materials and Methods:
Lignin-degrading microbial strains were isolated from soil using medium containing sodium lignosulphonate as the sole carbon source. They were then screened by aniline blue and guaiacol plate, and then the best strain was chosen and identified. The conventional one-factor method was used to optimize various parameters that affect lignin’s degradation ability.
Results:
The strain possessing the highest lignin biodegradation ability was identified and denominated as Aspergillus Flavus F-1. After optimization, the maximum degradation rate of lignin, 44.6% within 3 days, was obtained at pH 7.0, 30 ℃, 2.5 g·L-1 ammonium sulfate, 2 g·L-1 lignin and 0.5 g·L-1 glucose. The results show the LiP and Lac secreted from Aspergillus Flavus F-1 played the main role in the degradation of lignin.
Conclusion:
One microbial strain, Aspergillus Flavus F-1, was successfully isolated with a lignin-degrading ability that can cut the lignin into fragments. This provides a promising candidate for the transformation and utilization of crop waste biomass for various industrial purposes.
Keywords: Aspergillus Flavus F-1, Biomass, Degradation, Lignin
1. Background
Lignin is the largest natural aromatic polymer in nature and is also a unique aromatic-based biopolymer, accounting for nearly 30% of the earth’s organic carbon ( 1 ). Many small-molecule lignin derivatives have been widely used in chemicals, biofuels and material products, including vanillin, guaiacol, eugenol, eugenol, phenol, benzene, activated carbon as adsorbent, carbon fiber, biological products and plastics such as polyurethane and polyester ( 2 ). Nevertheless, high molecular weight and strong intramolecular and intermolecular interactions of lignin have made the degradation of lignin very difficult. The highly effective biodegradation of lignin has played a critical role in solving the problems of waste in biomass field or otherindustrial fields.
In 1983, a breakthrough was made in the field of lignin biodegradation, and the demand for fungal ligninase and its hydrogen peroxide was discovered ( 3 ). The biodegradation and biotransformation of lignin is the result of the joint action of fungi, bacteria and actinomycetes ( 4 ). Degradation of lignin by microorganisms was primarily highlighted in rot and brown-rot fungi, which were capable of producing certain extracellular ligninolytic enzymes ( 5 ). Fungal pretreatment refers to the process of decomposing lignin during the degradation of lignocellulosic biomass, thereby changing the structure of lignocellulosic biomass for subsequent processing ( 7 , 8 ). There are four main types of extracellular lignin-decomposing enzymes, namely laccase (Lac), manganese peroxidase (MnP), lignin peroxidase (LiP) and universal peroxidase, which degrade and modify lignin. Ligninolytic enzymes use oxygen as the ultimate electron acceptor and work through the oxidation and reduction processes ( 9 ). However, the direct utilization of single white-rot fungi and their enzymes was relatively difficult in industrial processes because of their slower growth rates, damageable enzymes, easier counteraction by other lower microorganisms, production costs, etc. ( 10 ). It was reported that when mold enzymes degrade complex biopolymers (such as cellulose, starch and lignin) into simpler substances (which can be absorbed by mycelium), they play a major role in the decomposition of organic substances ( 11 ), which should be also investigated for their potential in lignin degradation. It has been proved that Aspergillus flavus strain F10 can lead to noticeable color removal for detoxification in pulp and paper mills ( 12 ). This indicates that lignin degradation strains can degrade lignin effectively so it can be used in wastewater treatment.
In this article, with a focus on lignin degradation and its biodegraded products, a new isolated fungus strain F-1 with an ability to degrade lignin was characterized by its morphology and internal transcribed spacer (ITS) sequencing. Its lignin degradation capacity was also optimized in liquid culture.
2. Objective
This study intended to isolate and purify several microbial strains from nature. It also explores how their lignin degradation is able to enhance the biodegradation and recycling of biomass and the reclamation of lignin in wastewater from pulp and paper mills.
3. Materials and Methods
3.1. Materials
CL (Calcium lignosulfonate, ≥96%) was purchased from Aladdin Reagent Co. Ltd, China (prepared from alkaline pulping wastewater, brown powder, carbon content 38.0-42.0%, sulfur content 4.0-6.0%, pH 8-10, water content 2.0-8.0%, filtered through 80 mesh); ABTS and BSTFA were chromatographically pure (Sigma-Aldrich), and all other chemical reagents were of analytical or higher grade.
Isolation and storage medium: potato dextrose agar (PDA) medium. Screening medium: Crawford’s solid medium containing 2.0 g·L-1 lignin (calcium lignosulfonate or Klason lignin) ( 13 ). Selection medium: guaiacol medium that was PDA containing 0.01% (w/w) guaiacol, Aniline blue medium containing 0.1 g·L-1 Aniline blue. Biodegradation medium (g·L-1): lignin 2.0, glucose 0.5, ammonium sulfate 1.5, dipotassium phosphate 1.5, potassium dihydrogen phosphate 1.5, magnesium sulfate 0.2, sodium chloride 0.2, manganese sulfate 0.02, ferric sulfate 0.05, VB1 0.002.
3.2. Enrichment and Isolation of A Lignin-Degrading Microorganism
Soil samples were collected from rice fields in the suburbs of Hefei, Anhui Province, China. The soil- dissolving solution was fully diluted, then inoculated on the screening medium and cultured at 28 °C for 5 days. First, the strains that grow fast on the plate were selected, and then they were selected and cultured on Guaiacol medium and aniline blue medium to purify the colonies. Finally, the strain was preserved, grown on a PDA slope at 4 °C, and subcultured regularly.
3.3. Identification of the Lignin-Degrading Strain
The internal transcribed spacer (ITS) sequence was assayed by Sangon Biotech Co., Ltd. (Shanghai, China). To perform gene sequence analysis, using ITS1 and ITS4 as primers, the ITS1-5.8 S-ITS2 rRNA region of the strain was amplified by polymerase chain reaction (PCR). The homologous sequences were obtained from Genbank database. Closely related sequences were aligned and manually corrected, and finally, the N-J program in MEGA 5.05 was used to construct a dendrogram ( 14 ). The stability of relationships was assessed by a bootstrap analysis of 1000 trials.
3.4. Measurement of Lignin Degrading Ratio (LDR)
Centrifuge the samples at 15984 × g for 10 minutes was used to remove mycelium, and then the lignin content was detected at a wavelength of 280 nm ( 15 ). The lignin- degrading ratio (LDR) was calculated by lignin content before fermentation (LC1) and after fermentation (LC2) as follows:
3.5. Measurement of Enzyme Activity
To determine enzyme activity, the fermentation broth was centrifuged at 3000 × g and filtrated through a 0.45 μm membrane. The 1 mL reaction solution contained 0.2 mL veratrol solution, 0.4 mL tartaric acid buffer (250 mmoL·L-1, pH 3.0), 0.4 mL culture medium or diluent, 20 μL 20 mmoL·L-1 H2O2 solution, and was reacted at 30 °C for 2 min. The absorbance value was determined at 310 nm. In the blank group, distilled water was used instead of H2O2 solution, and the other reactants remained unchanged ( 16 ).
The reaction system at room temperature contained 3.4 mL of 50 mmoL·L-1 pH 4.5 sodium lactate buffer, 0.1 mL of 1.6 mmol·L-1 MnSO4 solution, 0.4 mL of enzyme solution and 0.1 mL of 1.6 mmoL·L-1 of H2O2 to start the reaction. The absorbance was measured at 240 nm in the first 4 minutes of the reaction ( 17 ).
The laccase reaction system contained 2.3 mL of 0.2mol·L-1 acetic acid-sodium acetate buffer, 0.5 mL of crude enzyme diluent with pH = 4.5 and 0.2 mL of 1 mmol·L-1 ABTS as the substrate. The reaction was monitored by measuring the change in A420 for 5 min.
One unit of enzymes activity was defined as the amount catalyzing 1 μmol of substrates oxidized per minute of reaction ( 18 ).
3.6. Effect of Different Factors on Lignin Degradation by Aspergillus flavus F-1
Effects of different nitrogen sources: Six inorganic nitrogen sources were selected: NH4Ac, (NH4)2SO4, NH4NO3, NH4H2PO4, NH4Cl and urea. Four organic nitrogen sources were selected: including casein, peptone, corn pulp and yeast extract. Sodium lignosulfonate was used as substrate with a concentration of 2 g·L-1 and an initial pH of 7.0, and 2% seed solution was inoculated for 5 days at 150 r·min-1 (28 °C) to determine the lignin degradation rate.
Effects of different nitrogen concentration: According to the high degradation rate of culture conditions fixed nitrogen sources, other factors remained unchanged, the effect of different nitrogen concentrations (0.5, 1.5, 2.5, 5, and 7.5 g·L-1 ) on lignin degradation by strain were studied.
Effect of different substrate concentration: Sodium lignosulfonate was used as substrate, with the concentrations set at 1, 2, 5, 10 and 20 g·L-1, and 2% seed solution was inoculated and cultured for 5 days at 150 r·min-1 (28 °C) to determine the degradation rate of lignin.
Effect of different culture temperature: Different culture temperatures were set at 20, 30, 40 and 50 °C, and the lignin degradation rate was determined by inoculating 2% seed liquid at 150 r·min-1 for 5 days.
Effects of different initial pH: 2% dilute hydrochloric acid and dilute sodium hydroxide solutions were prepared. The prepared culture was adjusted to pH of 3.0, 4.0, 5.0, 6.0, 7.0, 8.0, 9.0, 10.0 and 11.0, and the lignin degradation rate was determined by inoculating 2% seed liquid at 150 r·min-1 for 5 days.
4. Results
4.1. Isolation and Characterization of the Strain Degrading Lignin
One strain of filamentous fungi with the highest lignin- degrading enzyme activity was screened qualitatively from the collected samples. The strain F-1 had an effective ability to degrade lignin and stabilized activity. Morphologic characteristics of Aspergillus flavus strain F-1 are shown in Fig. 1. The ITS sequence of this strain and its homologous sequences were analyzed using MEGA 5.05, and a genetic relationship was also established (Fig. 2).
Figure 1.
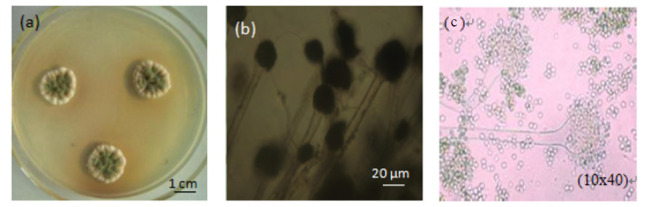

Morphologic characteristics of Aspergillus flavus strain F-1. (a) Photograph of strain F-1 colonies after 5d culture; (b) Microphotograph of strain F-1 showing the cell morphology (160×).Bar: 20 μm; (c) 10×40 times micrograph
Figure 2.
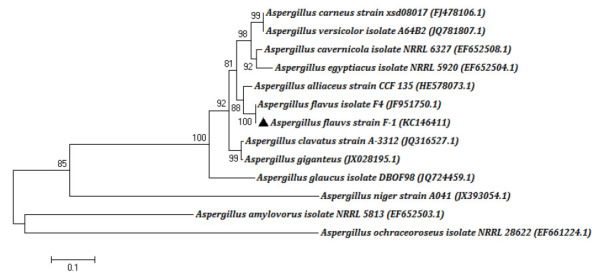
Neighbour-joining phylogenetic tree based on the ITS sequence
4.2. Biodegradation of Lignin by Aspergillus flavus Strain F-1
To determine the growth of the strain, the growth curve of Aspergillus flavus F-1 strain was measured by the cell dry weight method. As shown in Fig. 3A, Aspergillus flavus strain F-1 grew fast in the first three days and then grew slowly; after 5 days, the strain began to decline gradually. Figure 3B shows that the degradation rate of sodium lignosulfonate by Aspergillus flavus F-1 increased sharply in the first three days and then slowly increased. There were activities of lignin peroxidase (LiP) and laccase as shown in Figure 3C in biodegradation culturing solution, but MnP was not detected.
Figure 3.
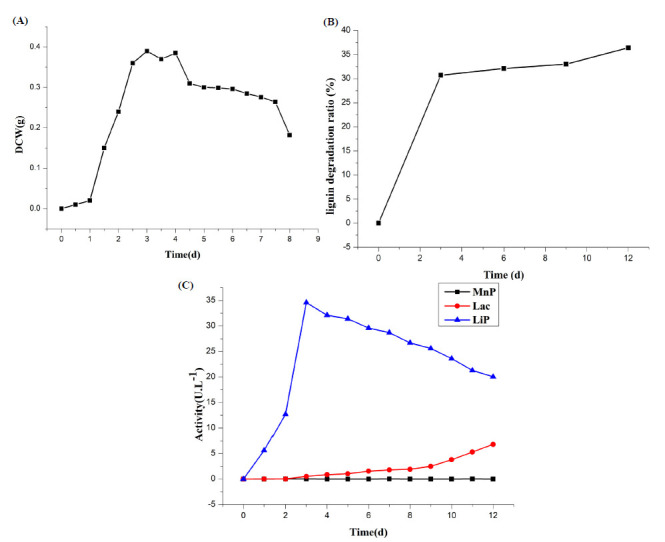
Cultural characteristics of Aspergillus flavus strain F-1. (A) Growth curve of strain F-1, described by DCW (dry cell weight); (B)Lignin degradation curve of strain F-1 in 12 d liquid cultu; (C) Enzyme production curve of strain F-1
4.3. Optimization of Various Parameters
4.3.1. Nitrogen Source
Various organic and inorganic nitrogen sources were screened to obtain the nitrogen source with the best lignin-degradation ability. The effect of different nitrogen sources on lignin degradation by Aspergillus flavus F-1 is shown in Table 1: the degradation effect of Aspergillus flavus F-1 reached 36.93% when (NH4)2SO4 was used as nitrogen source.
Table 1.
Optimization of Nitrogen Source.
| Nitrogen source species | lignin degradation ratio(%) |
|---|---|
| NH4Ac | 26.13±0.47 |
| (NH4)2SO4 | 36.93±0.56 |
| NH4NO3 | 25.43±0.41 |
| NH4H2PO4 | 29.62±0.49 |
| NH4Cl | 27.53±0.47 |
| urea | 19.51±0.41 |
| casein | 12.5±0.37 |
| peptone | 22.22±0.40 |
| corn pulp | 8.98±0.31 |
| yeast extract | 16.93±0.38 |
4.3.2 Nitrogen Sources Concentration
Nitrogen sources’ concentration was also an important factor affecting lignin degradation rate. The effects of different nitrogen sources’ concentrations on lignin degradation by Aspergillus flavus F-1 are shown in Figure 4A. The concentration of a nitrogen source among those investigated was 2.5 g·L-1 of (NH4)2SO4 .
Figure 4.
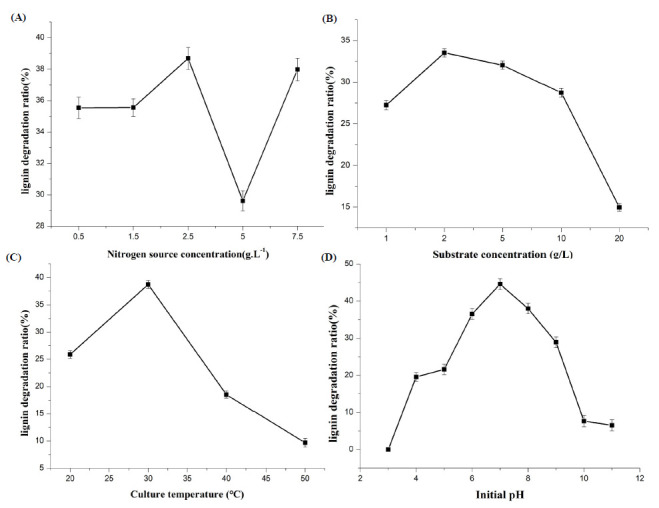
Effect of different conditions on lignin degradation by Aspergillus flavus F-1.(A) Nitrogen source concentration influence of lignin degradation on Aspergillus flavus F-1; (B) Substrate concentration influence of lignin degradation on Aspergillus flavus F-1; (C) Culture temperature influence of lignin degradation on Aspergillus flavus F-1; (D) Initial pH influence of lignin degradation on Aspergillus flavus F-1
4.3.3.Substrate Concentration
As shown in Figure 4B, it was shown that the concentration of the substrate affects the degradation of lignin depending on the strain. The appropriate substrate concentration was selected as 2 g·L-1 of lignin.
4.3.4.Culture Temperature
Temperature affects the growth of the strain, so the effect of different culture temperatures on lignin degradation by Aspergillus flavus F-1 was explored. As shown in Figure 4C, the appropriate temperature for lignin degradation was chosen as 30°C.
4.3.5. Initial pH
The effect of initial medium pH on the lignin degradation rate of Aspergillus flavus F-1 was studied within the range of pH 3-11. As shown in Figure 4D, the best degradation medium initial pH value was 7.0.
5. Discussion
The F-1 strain grows well in the PDA medium, and the color of the colony on the surface ranges from white, light pink and gray to blue-green. Microscopic observations of appearance (Fig. 1B) shown that the mycelium is transparent, branched and separated, with smooth walls and a width of 3-12 μm. As seen in Figure 1C, the inverted microscope (10×40) showed that the mycelium had a septum, was branched and had no false root; the spore pedicle was erect; the terminal sporangium had a rough surface of spherical conidial spore formed on the terminal sporangium; and the sporangium was flask or nearly globose.
The sequence has been registered in the NCBI GenBank database, and the accession number is KC146411. The BLAST analysis shows that the partial ITS rDNA of F-1 had 100% identity with those of Aspergillus flavus isolate F4 (JF951750.1). Based on the phylogenetic analysis, this strain was identified as Aspergillus flavus and named Aspergillus flavus strain F-1.
Compared with Figure 3A and Figure 3B, the degradation rate reached 30.7% in three days and then increased slowly. The maximum biodegradation of lignin (CL) was up to 36.4% at 12 days of incubation. The degradation of lignin increased with the growth of mycelium during the early and middle growth cycles; meanwhile, in the late growth cycles, the degradation degree still increased when Aspergillus flavus stopped growing. This phenomenon might have caused a series of free radical chain reactions to triggered the oxidation of the substrate and the degradation of lignin by ligninolytic enzymes of strain F-1 while the nutrition was exhausted.
Previous reports showed that white-rot fungi were the most effective microorganisms in nature to degrade lignin, and the degradation rate was 30%~40% when the longest culture period is 30 days (19). In contrast, the lignin degradation rate of Aspergillus flavus F-1 reached 36.4% in only 3 days, which greatly reduced the culture cycle. Moreover, the growth rate of Aspergillus flavus F-1 was fast, and it was easy to culture. As a result, Aspergillus flavus F-1 has the potential to be a highly effective lignin-degrading strain.
It has been shown that fungi that produce laccase and Lip or only laccase can efficiently degrade lignin. Lignin is degraded by side-chain oxidation, demethylation/ demethoxylation and aromatic ring breaking under the action of this degradation enzyme system.
As shown in Figure 3C, MnP activity was not detected in the strain F-1, and the activity of Lip increased quickly and reached its peak at 3 days, while the activity of Lac increased extremely slowly in the early and middle growth cycles and only occurred actively in the late growth cycle. It was clear that Lip played a major role in lignin degradation during the first three days. Three days later, although the activity of Lip decreased, the activity of Lac increased slowly and steadily. Therefore, the cooperation of two enzymes could have caused lignin degradation after three days, which made the rate increase, as shown in Figure 3 (b). It can be concluded that Lip and Lac were crucial factors for lignin degradation of Aspergillus flavus F-1. Experiments show that inorganic nitrogen sources have higher degradation efficiency for lignin. Because inorganic nitrogen sources are available nitrogen sources, they are more conducive to the utilization of strains. It was also possible that the strain mainly used the organic nitrogen source as a carbon source at the early stage of growth and only began to decompose and utilize the sole carbon source, sodium lignosulfonate, after nitrogen source consumption, which resulted in the existence of competitive inhibition of substrate.
It can be seen from Figure 4B that there was no significant difference in lignin degradation of Aspergillus flavus F-1 under different nitrogen source concentrations, but the overall trend showed that the degradation effect of Aspergillus flavus F-1 at lower nitrogen source concentrations was slightly better than that at higher nitrogen source concentrations.
The suitable substrate concentration was more favorable to the degradation of lignin. Because of its surfactant properties, sodium lignosulfonate adsorbed on its surface and was surrounded by complex reticular structure, which was not favorable for the absorption of a small amount of glucose to promote growth, resulting in slow growth. Figure 4B shows that the lower the substrate content, the higher the degradation rate, reaching about 30%. However, while the substrate concentration was higher, the degradation system became viscous because of the colloidal property of sodium lignosulfonate swelling in the medium, which affected the dissolution and transfer of oxygen.
The degradation rate of lignin showed a trend of first increasing and then decreasing with the increase in temperature. It can be seen from Figure 4C that the degradation rate of lignin was the highest at 30 °C. When the temperature was lower than its optimal temperature, the metabolism of microorganisms became active with the increase in temperature, and when the temperature was higher beyond its optimal temperature, the microorganisms would die, leading to the lower degradation rate of lignin.
Neutral pH greatly affects the growth of microorganisms and the membrane transport of nutrients. Studies on optimizing the pH of the medium show that the maximum lignin degradation rate is reached at pH 7.0, which may be due to the extreme pH directly affecting the pH of the microbial cell cytoplasm, which in turn affects the growth and enzyme productivity of the microbe. As shown in Figure 4D, the strain F-1 could degrade lignin in a wide range of acid–base culture conditions. When the pH of the culture system was low, the growth of the strain was seriously inhibited. It may be the case that the environment of strong alkali or strong acid destroyed cell membrane permeability and intracellular enzyme activity, thus affecting the metabolism of cells and adversely affecting the utilization of lignin by the strain. Therefore, pH 7.0 was selected as the degradation medium’s initial pH, and under the best conditions, it could cause 44.6% of the lignin to fragment within 3 days.
6. Conclusion
The strain of Aspergillus flavus F-1 from soil presented ligninolytic enzyme activities and could degrade 44.6% of the lignin into fragmentation at pH 7.0 and 30 °C in 3 days. Aspergillus flavus F-1 mainly produced lignin peroxidase and laccase. It could be Lip that played a major role in lignin degradation at first and then the cooperation of two enzymes that especially worked in lignin degradation after three days. The degradation characteristics of the strain showed that Aspergillus flavus strain F-1 could be a nitrogen-deficient strain, and at lower nitrogen source concentration it was slightly better than that at higher nitrogen source concentration. Aspergillus flavus F-1 proved a good candidate for the transformation and utilization of crop waste resources. At the same time, this work also provided a feasible way for the utilization of lignin in papermaking black liquor.
Acknowledgement
Authors express truly thanks for the support from Natural Science Research Key Project (2011AJZR0919); they also thank the Young Teachers’ Development Fund (2011HGQC1045) and College Students’ Innovative Fund (2012CXCY458, 2012CXCY464) of Hefei University of Technology. Analysis and Test Center of University of Science and Technology of China is appreciated.
References
- 1.Fatai B, Takanori F, Louise H. Isolation of novel lignin degrading enzymes and lignin degradation products from bacteria and fungi. NEW BIOTECHNOL. 2014; 20(31S): S191. doi: 10.1016/j.nbt.2014.05.938. [DOI] [Google Scholar]
- 2.Hao Z, Li S, Sun J, Song L, Fang Z. Efficient visible-light-driven depolymerization of oxidized lignin to aromatics catalyzed by an iridium complex immobilized on mesocellular silica foams. Appl Catal B-Environ. 2018; 237:366–372. doi: 10.1016/j.apcatb.2018.05.072. [DOI] [Google Scholar]
- 3.Isroi, Millati R, Syamsiah S, Niklasson C, Cahyanto MN, Lundquist K. Biological pretreatment of lignocelluloses with white-rot fungi and its applications: a review. Bioresources. 2011; 6(4):5224–5259. doi: 10.3390/molecules171214995. [DOI] [Google Scholar]
- 4.Lundquist K, Kirk TK, Connors WJ. Fungal degradation of kraft lignin and lignin sulfonates prepared form synthetic 14 C-lignins. Arch Microbiol. 1977; 112(3):291–296. doi: 10.1007/bf00413095. [DOI] [Google Scholar]
- 5.Li C, Zhao X, Wang A, Huber GW, Zhang T. Catalytic transformation of lignin for the production of chemicals and fuels. Chem Rev. 2015; 115(21):11559–11624. doi: 10.1021/acs.chemrev.5b00155. [DOI] [PubMed] [Google Scholar]
- 6.Ye L, Wang W, Zhou Y, Yao S, Deng L, Zeng K. Isolation, identification and in vitro screening of Chongqing orangery yeasts for the biocontrol of Penicillium digitatum on citrus fruit. Biol Control. 2017; 110:18–24. doi: 10.1016/j.biocontrol.2017.04.002. [DOI] [Google Scholar]
- 7.Masran R, Zanirun Z, Bahrin EK, Ibrahim MF, Lai YP, Abd- Aziz S. Harnessing the potential of ligninolytic enzymes for lignocellulosic biomass pretreatment. Appl Microb Biotechnol. 2016; 100(12):1–16. doi: 10.1007/s00253-016-7545-1. [DOI] [PubMed] [Google Scholar]
- 8.Nikupaavola ML, Raaska L, Itavaara M. Detection of white-rot fungi by a non-toxic stain. Mycol Res. 1990; 94(1):27–31. doi: 10.1016/s0953-7562(09)81260-4. [DOI] [Google Scholar]
- 9.Sun Z, Fridrich B, Santi AD, Elangovan S, Barta K. Bright side of lignin depolymerization: toward new platform chemicals. Chem Rev. 2018; 118(2):614–678. doi: 10.1021/acs.chemrev.7b00588. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Ming T, Kirk TK. Lignin-degrading enzyme from the hymenomycete Phanerochaete chrysosporium burds. Appl Biochem Biotech. 1983; 221(4611):661–663. doi: 10.1126/science.221.4611.661. [DOI] [PubMed] [Google Scholar]
- 11.Anand B HJ. Decolourization and biological treatment of pulp and paper mill effluent by lignin-degrading fungus Aspergillus flavus strain F10. Int J Curr Microbiol Appl Sci. 2016; 5(5):19–32. doi: 10.20546/ijcmas.2016.505.003. [DOI] [Google Scholar]
- 12.Haq I, Kumar S. Evaluation of bioremediation potentiality of ligninolytic Serratia liquefaciens for detoxification of pulp and paper mill effluent. J Hazard Mater. 2016;305(Suppl):190–199. doi: 10.1016/j.jhazmat.2015.11.046. [DOI] [PubMed] [Google Scholar]
- 13.Tamura K, Dudley J, Nei M, Kumar S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol. 2007;24(8):1596. doi: 10.1093/molbev/msm092. [DOI] [PubMed] [Google Scholar]
- 14.Yang YS, Lu H, Yuan YL, Zhao LH. Isolation and characterization of a fungus Aspergillus sp. strain F-3 capable of degrading alkali lignin. Biodegradation. 2011;22(5):1017– 27. doi: 10.1007/s10532-011-9460-6. [DOI] [PubMed] [Google Scholar]
- 15.Yang YS, Zhou JT, Lu H, Yuan YL, Zhao LH. Isolation and characterization of Streptomyces spp. strains F-6 and F-7 capable of decomposing alkali lignin. Environ Technol. 2012; 33(23):2603–2609. doi: 10.1080/09593330.2012.672473. [DOI] [PubMed] [Google Scholar]
- 16.Zeng GM, Yu HY, Huang HL, Huang DL, Chen YN, Huang GH, et al. Laccase activities of a soil fungus Penicillium simplicissimum in relation to lignin degradation. World J Microbiol Biotechnol. 2006; 22(4):317–324. doi: 10.1007/s11274-005-9025-0. [DOI] [Google Scholar]
- 17.Zhao X, Zhang L, Liu D. Biomass recalcitrance. Part II: Fundamentals of different pre-treatments to increase the enzymatic digestibility of lignocellulose. Biofuel Bioprod Bior. 2012; 6(5):561–579. [Google Scholar]
- 18.Bumpus JA, Aust SD. Biodegradation of environmental pollutants by the white rot fungus Phanerochaete chrysosporium: Involvement of the lignin degrading system. Bioessays. 2010;6(4):166–70. doi: 10.1002/bies.950060405. [DOI] [Google Scholar]
- 19.Youn HD, Hah YC, Kang SO. Role of laccase in lignin degradation by white-rot fungi. Fems Microbiol Lett. 1995;132(3):183–188. doi: 10.1016/0378-1097(95)00315-v. [DOI] [Google Scholar]


