Abstract
Carmine radish is famous for containing a natural red pigment (red radish pigment). However, the expression of anthocyanin biosynthesis-related genes during the dynamic development stages of the fleshy roots in carmine radish has not been fully investigated. Here, based on HPLC quantification of anthocyanin levels from our previous study, young fleshy roots of the carmine radish “Hongxin 1” obtained at the dynamic development stages of fleshy roots (seedling stage (SS), initial expansion (IE), full expansion (FE), bolting stage (BS), initial flowering stage (IFS), full bloom stage (FBS) and podding stage (PS)) were used for RNA-Seq. Approximately 126 comodulated DEGs related to anthocyanin biosynthesis (common DEGs in the dynamic growth stages of fleshy roots in carmine radish) were identified, from which most DEGs appeared to be likely to participate in anthocyanin biosynthesis, including two transcription factors, RsMYB and RsRZFP. In addition, some related proteins, e.g., RsCHS, RsDFR, RsANS, RsF′3H, RsF3GGT1, Rs3AT1, RsGSTF12, RsUFGT78D2 and RsUDGT-75C1, were found as candidate contributors to the regulatory mechanism of anthocyanin synthesis in the fleshy roots of carmine radish. In addition, 11 putative DEGs related to anthocyanin synthesis were evaluated by qRT-PCR via the (2-ΔΔCT) method; the Pearson correlation analysis indicated excellent concordance between the RNA-Seq and qRT-PCR results. Furthermore, GO enrichment analysis showed that “anthocyanin-containing compound biosynthetic process” and “anthocyanin-containing compound metabolic process” were commonly overrepresented in the dynamic growth stages of fleshy roots after the initial expansion stage. Moreover, five significantly enriched pathways were identified among the DEGs in the dynamic growth stages of fleshy roots in carmine radish, namely, flavonoid biosynthesis, flavone and flavonol biosynthesis, diterpenoid biosynthesis, anthocyanin biosynthesis, and benzoxazinoid biosynthesis. In conclusion, these results will expand our understanding of the complex molecular mechanisms of anthocyanin biosynthesis in the fleshy roots of carmine radish and the putative candidate genes involved in this process.
Keywords: Radish (Raphanus sativus L.), Differential expression genes (DEGs), KEGG pathway enrichment, Anthocyanins, Anthocyanin biosynthesis
Introduction
Anthocyanins are water-soluble pigments that produce red to purple colors in nature and belong to the flavonoid group (Khoo et al., 2017). Most research has reported that anthocyanins, which are used as beneficial food additives worldwide, could ameliorate major public health threats (such as cardiovascular disease, inflammation, diabetes and obesity) caused by synthetic food additives (Yousuf et al., 2015; He & Giusti, 2010). Currently, several regulatory genes associated with the anthocyanin biosynthetic pathway have been extensively characterized and found to be conserved among flowering plants (Bajpai et al., 2017). A series of enzymes have been identified to be involved in anthocyanin formation, especially chalcone isomerase (CHI), chalcone synthase (CHS), dihydroflavonol 4-reductase (DFR), flavanone 3-hydroxylase (F3H), and anthocyanidin synthase (ANS/LDOX) (Aza-González et al., 2012; Dao & Linthorst, 2011). Fukusaki et al. (2004) reported that flower color would change from blue to white through the knockdown of the CHS gene by RNA interference in Torenia hybrida. Additionally, in mildly colored pears, the DFR and ANS genes were identified as important regulators of skin pigmentation (Lepiniec et al., 2006), and various anthocyanidins could be generated from dihydroflavonols using NADPH as a cofactor through the DFR and ANS genes (Zhang et al., 2011). UDP-glucose: flavonoid 3-O-glucosyltransferase (UFGT) was also found to be involved in anthocyanin biosynthesis in grape berries, and the loss of color in white grapes was shown to result from the absence of the UGFT gene (Kobayashi et al., 2001). Moreover, several glutathione S-transferases (GSTs) were found to be involved in the sequestration of anthocyanins, thereby playing vital roles in flavonoid metabolism in many plants, including Arabidopsis (Arabidopsis thaliana), apple (Malus domestica) and grape (Vitis vinifera) (Soonyoung, Seonae & Yun, 2016; Cutanda-Perez et al., 2009; Li et al., 2011). Additionally, transcription factors (TFs) related to anthocyanin biosynthetic traits act as important regulators in the normal development of an organism and in routine cellular functions (Latchman, 1993; Yusuf et al., 2012). Recently, MYB-bHLH-WDR (MBW) complexes have been demonstrated to transcriptionally regulate the genes encoding these enzymes through MYB, bHLH and WD40 repeats (MBW transcriptional complex) (Xu, Dubos & Lepiniec, 2015). In addition, regulatory genes associated with anthocyanin biosynthesis traits, such as squamosa promoter binding protein-like (SPL) (Gou et al., 2011), jasmonate zim-domain (JAZ) (Qi et al., 2011), and NAC (Zhou et al., 2015), have also been reported in Arabidopsis thaliana and blood-fleshed peach. Currently, key anthocyanin biosynthesis-related genes and some of their functions have been extensively characterized in several plants.
Recently, RNA-Seq technology has developed rapidly, and more anthocyanin biosynthesis-related genes have been documented in various fruit crops, including blueberry (Li et al., 2012), blood orange (Crifò, Petrone & Recupero, 2011) and grape (Azuma, 2017). Biosynthesis-related genes in carmine radish (Raphanus sativus L.) fleshy roots were identified based on comparative RNA-Seq technology in our previous study (Gao et al., 2020). Specifically, several major anthocyanin biosynthesis-related genes (ABRGs) involved in the regulation of anthocyanin biosynthesis were identified, such as the transcripts of RsDFR1, RsDFR2 and RsFLS (Gao et al., 2019a). However, the expression of anthocyanin biosynthesis-related genes related to anthocyanin accumulation in carmine radish (Raphanus sativus L.) fleshy roots have not been fully investigated.
Methods
Plant material and experimental design
Based on the quantification of anthocyanin levels of carmine radish “Hongxin 1” using HPLC analysis in our previous study (Gao et al., 2019b) (Fig. S1), young fleshy roots obtained from the development stage of carmine radish “Hongxin 1” were used for RNA-Seq in this study. The carmine radish “Hongxin 1” was selected and kept in a greenhouse at the experimental base of the Yangtze Normal University experiment in 2019. First, seeds of “Hongxin 1” were sown in sterile soil under normal growth conditions (23 °C, 16 h light/8 h dark) for 2 weeks. Subsequently, vernalization treatment was conducted, with the 2-week-old plants transferred to and maintained in a cold room (5 ± 1 °C, 12 h light/12 h dark) for 15 days. After vernalization treatment, the plants were grown in a growth room under normal growth conditions (23 °C, 16 h light/8 h dark). Finally, fleshy roots obtained from each stage of development (seedling stage (SS), initial expansion (IE), full expansion (FE), bolting stage (BS), initial flowering stage (IFS); full-bloom stage (FBS) and podding stage (PS)) of carmine radish “Hongxin 1” were collected for RNA-Seq, with three independent biological replicates for each stage and two technical replicates. All harvested tissue was immediately frozen in liquid nitrogen and stored at −80 °C for RNA-Seq analysis.
Sample preparation and library construction
Library construction was conducted with NEBNext Ultra RNA Library Prep Kits for Illumina (NEB, Ipswich, MA, USA), as follows: The mRNA was isolated with oligo(dT) magnetic beads from approximately 5 µg of total RNA and subsequently converted into short fragments by fragmentation buffer. Then, short fragments were converted into first-strand cDNAs and used as templates for second-strand synthesis with random hexamers. PCR amplification was conducted using the desired purified synthesized cDNA fragments (QiaQuick PCR kit). Ultimately, 200 bp paired-end reads were generated from the prepared library with 2 replicates using Illumina HiSeqTM 2000.
Read processing and differentially expressed gene (DEG) identification
Using Trimmomatic software, clean reads were obtained by filtering out low-quality reads (base quality ≤10) and adaptor-only reads and then trimming the remaining reads. After that, the small subunit (SSU) and large subunit (LSU) rRNA sequences were downloaded from the Silva database (Christian et al., 2013) and used for high-quality read alignment using BWA software (Li & Richard, 2009), and the mapped rRNA reads were removed by a homemade Perl script. Subsequently, we used Trinity software to de novo assemble the remaining clean reads into transcripts. In addition, BLASTx searches of the Swiss-Prot protein databases and nonredundant protein (Nr) database of the National Center for Biotechnology Information (NCBI) were used to annotate all unigenes to obtain the assembled sequences. The parameters were set as follows: E-value < 1e−10, identity > 70%, query coverage ≥ 80%, and default values for other parameters. After gene annotation, the Gene Ontology (GO) database (http://www.geneontology.org/) and Kyoto Encyclopedia of Genes and Genomes (KEGG) database (http://www.kegg.jp/) were used to annotate the functions of assembled unigenes. To assess the abundances of assembled transcripts, we first mapped the clean reads obtained from different fleshy root libraries using Bowtie2 on the de novo-assembled transcriptome. Then, quantification of the de novo assembly transcript was assessed with RSEM, and only transcripts with FPKM ≥ 1 were considered significantly expressed transcripts (Dewey & Li, 2011). Finally, differentially expressed genes (DEGs) with a corrected P-value < 0.05 between each set of compared samples were identified through screening by noiseqbio (Tarazona et al., 2012). Furthermore, DEGs related to anthocyanin biosynthesis in carmine radish were analyzed and plotted using neighbor-joining clustering with a homemade R script.
GO functional annotation and KEGG pathway analysis of comodulated differentially expressed genes (DEGs) in the growth stages of carmine radish
Co-modulated DEGs (DEGs common to all the dynamic growth stages of fleshy roots in carmine radish) were identified based on Venn diagram analysis. Subsequently, we conducted GO annotation of comodulated DEGs through the AgriGO website (http://systemsbiology.cau.edu.cn/agriGOv2/) and KEGG pathway enrichment analysis using KOBAS software (Xie et al., 2011). Their respective graphs were constructed using a homemade R scripts.
Validation of candidate DEGs involved in the growth stages of carmine radish using real-time qRT-PCR
To validate the results of the RNA-Seq analysis, 11 DEGs with substantial alterations related to the growth stages of carmine radish were chosen for validation by qRT-PCR. The primers used for qRT-PCR experiments were designed by Real-time PCR (TaqMan) Primer and Probes Design Tool (https://www.genscript.com/tools/real-time-pcr-taqman-primer-design-tool), and the radish ACTIN gene was used as a reference gene (Table S1). Following the standard protocol for the ABI 7500 system, the amplification of the candidate genes was performed using qRT-PCR in triplicate as described by Jian et al. (2015). The fold change in the expression levels of target genes was calculated using the relative quantitative method (2−ΔΔCT) (Schefe et al., 2006), and variance analysis was conducted to evaluate differences in the relative expression levels of the genes between different samples, followed by multiple comparisons using Duncan’s least significant range (LSR) tests with SPSS statistical software. Different letters indicate significant differences at the p = 0.05 level. Of those, a, b, c, d, e, f, or g indicates a significant difference from a column with no superscript letter in common (P < 0.05); columns marked with the same letter are not significantly different at the 5% level by Duncan’s multiple range test.
Results
DEGs related to the dynamic growth stages of fleshy roots in carmine radish
Here, transcriptome analysis of anthocyanin biosynthesis in the carmine radish “Hongxin 1” was conducted followed by HPLC analysis, and young fleshy roots of “Hongxin 1” at different developmental stages were selected for RNA-Seq. The cDNAs obtained from fleshy roots at seven growth phases (SS, IE, FE, BS, IFS, FBS and PS) were sequenced using Illumina sequencing technology. To harvest clean reads for further assembly, we filtered out low-quality reads (base quality ≤10) and adaptor-only reads and then trimmed the sequences to obtain high-quality reads. Subsequently, we aligned the high-quality reads to the small subunit (SSU) and large subunit (LSU) rRNA sequences and removed rRNA reads by a homemade Perl script. After removing rRNA sequences, the average percentage of clean read counts was 70% of raw tags in each library (Table 1).
Table 1. Summary of raw reads for 14 samples (two replicates) of fleshy roots of seven growth phases.
| Sample | Total_Read_Count | rRNA_Read_Count | Clean_Read_Count | Clean_Rate (%) |
|---|---|---|---|---|
| FBS_root_1 | 33,990,858 | 7,871,020 | 26,119,838 | 76.84 |
| FBS_root_2 | 33,990,858 | 8,386,243 | 25,604,615 | 75.33 |
| BS_root_1 | 33,401,149 | 7,702,113 | 25,699,036 | 76.94 |
| BS_root_2 | 33,401,149 | 7,587,241 | 25,813,908 | 77.28 |
| IE_root_1 | 28,574,219 | 4,463,319 | 24,110,900 | 84.38 |
| IE_root_2 | 28,574,219 | 4,663,468 | 23,910,751 | 83.68 |
| SS_root_1 | 20,191,382 | 504,415 | 19,686,967 | 97.50 |
| SS_root_2 | 20,191,382 | 560,164 | 19,631,218 | 97.23 |
| IFS_root_1 | 24,547,530 | 5,534,772 | 19,012,758 | 77.45 |
| IFS_root_2 | 24,547,530 | 5,976,321 | 18,571,209 | 75.65 |
| FE_root_1 | 14,316,434 | 3,200,500 | 11,115,934 | 77.94 |
| FE_root_2 | 14,316,434 | 3,092,331 | 11,224,103 | 78.40 |
| PS_root_1 | 20,673,951 | 6,219,688 | 14,454,263 | 70.08 |
| PS_root_2 | 20,673,951 | 5,789,807 | 14,884,144 | 72.01 |
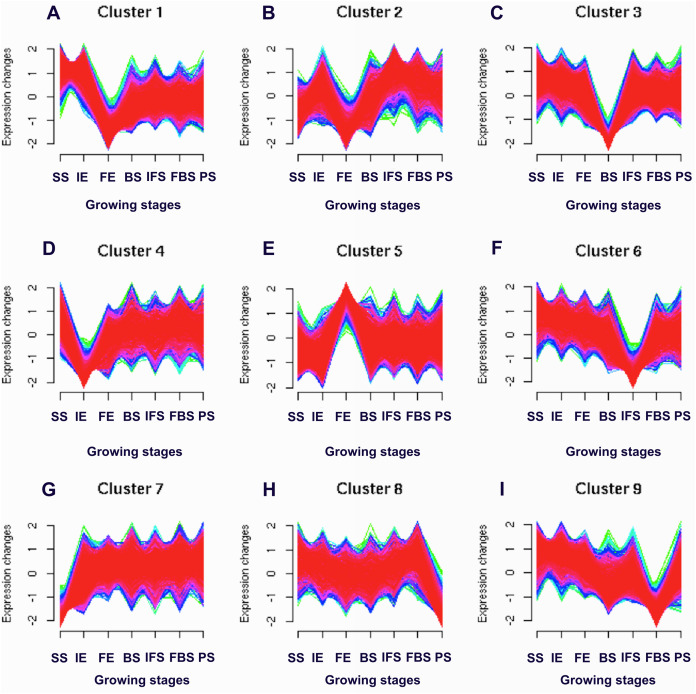
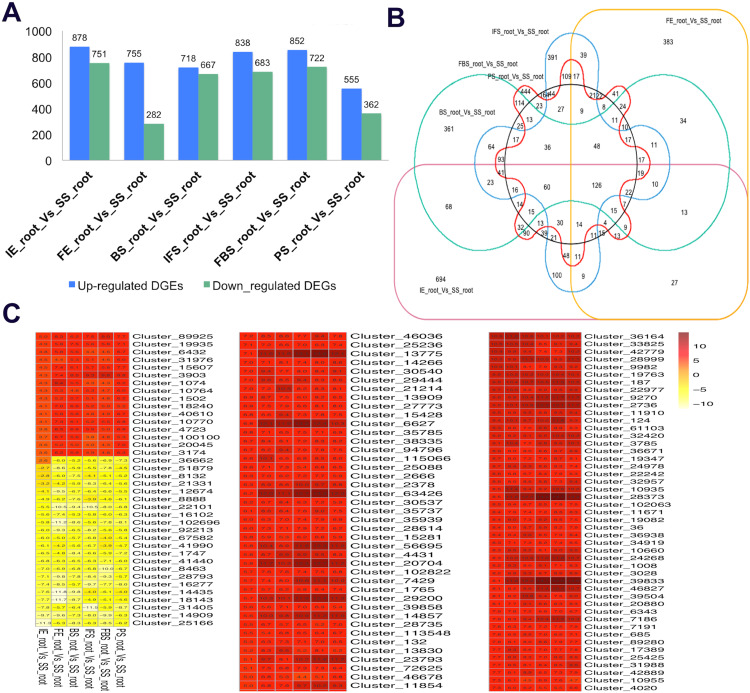
To identify the candidate genes related to the dynamic growth stages of fleshy roots in “Hongxin 1” radish, we normalized the expression levels of all the globally expressed genes for further analysis, and the results showed that highly distinct gene expression profiles exist in the dynamic growth stages of fleshy roots (Fig. 1; Table S2). Cluster analysis according to the gene expression dynamics was performed by the k-means method, and differentially expressed genes were divided into 9 clusters associated with distinct expression profiles. More interestingly, we found that the putative candidate genes belonging to Cluster 8 were consistent with the dynamics of the anthocyanin profiles of fleshy roots during the development of carmine radish, while the putative candidate genes categorized into Cluster 9 had the opposite pattern. Furthermore, DEGs related to anthocyanin accumulation in carmine radish (Raphanus sativus L.) fleshy roots were identified between other developmental periods (“IE_root”, “FE_root”, “BS_root”, “IFS_root”, “FBS_root” and “PS_root”) and the “SS_root” period. Compared with “SS_root”, 1,629, 1,037, 1,385, 1,521, 1,574 and 917 DEGs were generated in each different periods of development, including both upregulated DEGs (878, 755, 718, 838, 852 and 555 transcripts) and downregulated DEGs (751, 282, 667, 683, 722 and 362 transcripts) (Fig. 2A). Of those, 126 comodulated DEGs were identified based on a Venn diagram (Fig. 2B), and we used a heatmap to display the expression pattern changes of comodulated DEGs (common DEGs involved in the dynamic growth stages of fleshy roots in carmine radish) with different colors (Fig. 2C). Moreover, we found that some comodulated DEGs showed similar expression trends in the dynamic growth stages of fleshy roots, which were again consistent with anthocyanin profiles of the fleshy roots, such as a series of functional enzymes that acted as important regulators in anthocyanin biosynthesis, including dihydroflavonol 4-reductase (RsDFR: Cluster_13775), flavonoid 3′-monooxygenase (RsF3′H: Cluster_4431), leucoanthocyanidin dioxygenase (RsLDOX: Cluster_3903) and chalcone synthase (RsCHS: Cluster_39833), as well as some regulatory enzymes, such as anthocyanidin 3-O-glucoside 2′″-O-xylosyltransferase (Rs3GGT1: Cluster_9270), coumaroyl-CoA:anthocyanidin 3-O-glucoside-6″-O-coumaroyltransferase 1-like (Rs3AT1: Cluster_46827), UDP-glycosyltransferase 75C1-like (RsUGT75C1: Cluster_2736) and UDP-glycosyltransferase 78D2-like (RsUGT78D2: Cluster_11854). In addition, transport proteins and transcription factors, namely, glutathione S-transferase F12 (RsGSTF12: Cluster_24268), MYB transcription factor (RsMYB2: Cluster_28373), and zinc finger, RING-type protein (RsRZEP: Cluster_7186), were identified (Fig. 2C; Table S3).
Figure 1. Soft clusters of normalized expression levels for all global expressed genes involved in the dynamics growing stages of fleshy roots in carmine radish.
A total of nine clusters (A-I) with the different level of expression changes were identified. Horizontal axis represents growing stages of fleshy roots (“SS_root”, “IE_root”, “FE_root”, “BS_root”, “IFS_root”, “FBS_root” and “PS_root”). The vertical axis represents expression changes.
Figure 2. Transcriptional changes of DEGs involved in the dynamics growing stages of fleshy roots in carmine radish.
(A) Statistic of differentially expression genes (including up-regulated and down-regulated in each comparison groups) in the dynamics growing stages of fleshy roots (“IE_root”, “FE_root”, “BS_root”, “IFS_root”, “FBS_root” and “PS_root”), compared with “SS_root” group. (B) Venny graph of co-modulated DEGs (Common DEGs in the dynamic growing stages of fleshyroot in carmine radish). (C) Clustering and heat map of common differentially expressed (Co-modulated genes) based on the expression profiles in the dynamics growing stages of fleshy roots (“IE_root”, “FE_root”, “BS_root”, “IFS_root”, “FBS_root” and “PS_root”), compared with “SS_root” group.
Functional annotation of DEGs related to the dynamic growth stages of fleshy roots in carmine radish
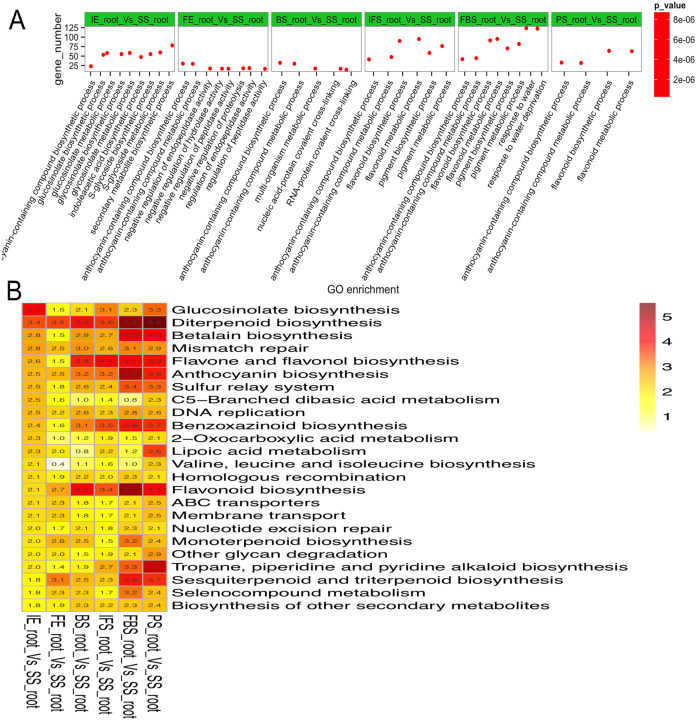
To explore the regulatory mechanisms of DEGs related to the dynamic growth stages of fleshy roots in carmine radish, GO annotation and KEGG pathway enrichment of those putative DEGs were conducted. The results indicated that “anthocyanin-containing compound biosynthetic process (GO:0009718)” and “anthocyanin-containing compound metabolic process (GO:0046283)” were commonly overrepresented in the dynamic growth stages of fleshy roots after the initial expansion (i.e., 40 days after planting), “flavonoid biosynthetic process (GO:0009813)” and “flavonoid metabolic process (GO:0009812)” were overrepresented in IFS, FBS and PS, but “pigment biosynthetic process (GO:0046148)” and “pigment metabolic process (GO:0042440)” were overrepresented in IFS and FBS. Moreover, we found that the GO terms “glucosinolate biosynthetic process (GO:0019758)” and “glucosinolate metabolic process (GO:0019757)” were overrepresented in IE (Fig. 3A; Table S4). KEGG pathway enrichment analysis identified five significantly enriched pathways related to anthocyanin synthesis traits in carmine radish, including flavonoid biosynthesis, flavone and flavonol biosynthesis, diterpenoid biosynthesis, anthocyanin biosynthesis, and benzoxazinoid biosynthesis (Fig. 3B; Table S5).
Figure 3. Functional enrichment analysis of differentially expressed genes (DEGs) related to the dynamics growing stages of fleshy roots in carmine radish.
(A) Enriched GO terms of DEGs related to the dynamics growing stages of fleshy roots in radish. GO terms are plotted on the ordinate, and the enrichment factor (rich factor) is plotted on the abscissa. The color of points represents the q-value, and the size of points represents the number of DEGs mapped to the reference pathway. Legends for the color scale of q-values and size-scaling of the number of DEGs are shown to the right of the plot. (B) Pathway enrichment analysis among differentially expressed genes related to anthocyanin synthesis in radish. Enriched KEGG pathway terms divided by the dynamics growing stages (“IE_root”, “FE_root”, “BS_root”, “IFS_root”, “FBS_root” and “PS_root”), compared with “SS_root”. Red color indicates statically overrepresented.
Validation of DEGs related to anthocyanin synthesis traits through qRT-PCR
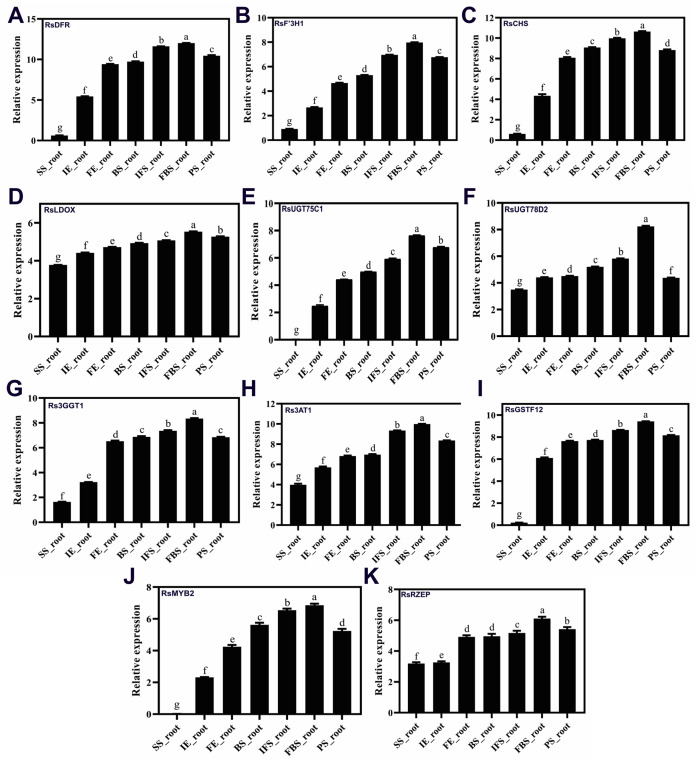
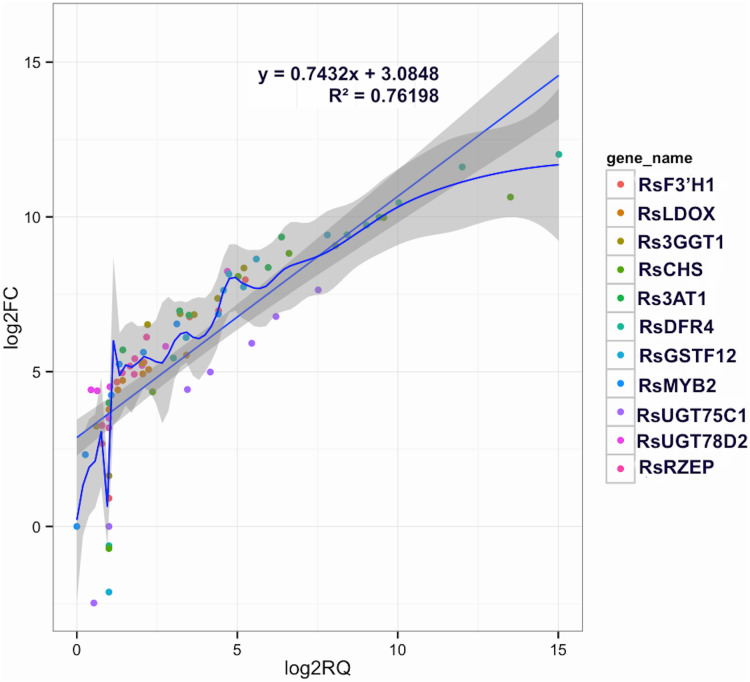
To confirm the RNA-Seq results, eleven putative DEGs (Cluster_13775, Cluster_4431, Cluster_3903, Cluster_39833, Cluster_9270, Cluster_46827, Cluster_2736, Cluster_11854, Cluster_24268, Cluster_28373 and Cluster_7186) were selected and subjected to qRT-PCR analysis (Fig. 4). The changes in expression of these 11 putative DEGs related to anthocyanin synthesis were measured by log2-fold change, and the Pearson correlation analysis indicated excellent concordance between RNA-Seq and qRT-PCR (R2 = 0.76198) (Table S6; Fig. 5). Here, transcripts of dihydroflavonol 4-reductase (RsDFR: Cluster_13775), flavonoid 3′-monooxygenase (RsF3′H: Cluster_4431), chalcone synthase (RsCHS: Cluster_39833) and leucoanthocyanidin dioxygenase (RsLDOX: Cluster_3903) were identified and validated using qRT-PCR. The transcripts of RsDFR, RsF3′H and RsCHS, but not RsLDOX, were significantly upregulated in the different growth stages of fleshy roots among other different development periods from seedling stage to full-bloom stage but decreased in the podding stage, showing consistency with the dynamics of anthocyanin profiles of fleshy root in development of carmine radish (Figs. 4A, 4B, 4C and 4D); similar patterns were observed for UDP-glycosyltransferase 75C1-like (RsUGT75C1: Cluster_2736) and UDP-glycosyltransferase 78D2-like (RsUGT78D2: Cluster_11854), which have been reported to be involved in the methylation of anthocyanidins, resulting in stable compounds. We found that the expression of RsUGT75C1 was significantly different from that of RsUGT78D2 during the fleshy root growth development stages compared with the seedling stage (Figs. 4E and 4F). In addition, anthocyanidin 3-O-glucoside 2′″-O-xylosyltransferase (Rs3GGT1: Cluster_9270) and coumaroyl-CoA:anthocyanidin 3-O-glucoside-6″-O-coumaroyltransferase 1-like (Rs3AT2: Cluster_9270) might act as key regulatory enzymes for the formation of anthocyanins in carmine radish. We found that Rs3GGT1 was changed significantly compared with Rs3AT2 (Figs. 4G and 4H). Moreover, transport proteins and transcription factors such as glutathione S-transferase F12 (RsGSTF12: Cluster_24268), MYB transcription factor (RsMYB2: Cluster_28373), and zinc finger, RING-type protein (RsRZEP: Cluster_7186) were also identified. The transcripts of RsGSTF12 and RsRZEP were not significantly differentially expressed in the other growth stages of fleshy roots compared with the seedling stage (Figs. 4I and 4K), but we found that the transcript of RsMYB2 (Fig. 4J) was significantly upregulated in the fleshy root growth stage and decreased in podding stage compared with other development periods from seedling stage to full-bloom stage. This pattern was consistent with the dynamics of anthocyanidin profiles across the fleshy root development stages in the carmine radish.
Figure 4. (A–K) Transcriptional analysis of anthocyanin synthesis-related genes (ASRGs) identified in fleshy roots obtained from the dynamics development stage of fleshy roots (seedling stage (SS), initial expansion (IE), full-expansion (FE), bolting stage (BS), initial.
flowering stage (IFS); full-bloom stage (FBS) and podding stage (PS)) in carmine radish “Hongxin 1” using qRT-PCR. Relative gene expression levels were normalized against actin transcript levels, and log2 scale for fold change of gene expression in the development stage of fleshy roots comprising of “SS”, “IE”, “FE”, “BS”, “IFS”, “FBS” and “PS” was shown. The standard error calculated from three biological replicates and significant (P < 0.05) difference identified by uncorrected Fisher’s LSD test in multiple comparisons after two-way ANOVA are indicated by error bars and stars, respectively.
Figure 5. Correlations of transcript levels of candidate DEGs related to anthocyanin synthesis traits in carmine radish between RNA-seq and qPCR data.
Validation of candidate Co-modulated DEGs involved in the dynamics growing stages of fleshy roots in carmine radish using qRT-PCR and then correlation between RNA-seq and qPCR data were conducted. Each RNA-seq expression data was plotted against that from quantitative real-time PCR and fit into a linear regression. Both x- and y-axes were shown in log2 scale and each color represented a different gene.
Discussion
Anthocyanins are major color pigments and play diverse physiological roles in plants. Currently, more than 550 different anthocyanins have been isolated from diverse plants and categorized by the extent of hydroxylation in the flavonoid B ring (Ferreira, Slade & Marais, 2005). Studies have demonstrated that anthocyanin compounds are synthesized through the methylation, glycosylation and acylation of the basic flavonol structure. Several structural genes and enzymes related to anthocyanin and flavonoid biosynthesis have been reported in most plant species. Previous studies have reported that anthocyanins are first formed from phenylalanine by a series of phenylpropanoid metabolism enzymes, such as chalcone synthase (CHS), the key enzyme in the first step of flavonoid synthesis, followed by chalcone isomerase (CHI), which leads to the formation of flavanones by closing the C-ring (Grotewold, 2006). After that, proanthocyanidins important for the formation of anthocyanidins were produced through anthocyanidin synthase (ANS) and dihydroflavonol 4-reductase (DFR), and subsequently, anthocyanins were produced through the action of UDP-flavonoid glycosyltransferases (UFGTs) (Xie, 2003). In our previous study, de novo transcriptome sequencing of radish (Raphanus sativus L.) fleshy roots showed that the majority of anthocyanin biosynthesis-related genes (ABRGs) are involved in the regulation of anthocyanin biosynthesis. Of those, RsPAL2, RsCHS-B2, RsDFR1, RsDFR2, RsFLS, RsMT3 and RsUFGT73B2-like were identified as significantly associated with anthocyanin biosynthesis. RsDFR1 and RsDFR2 were most highly enriched in HX-3, and RsFLS, RsDFR1 and RsDFR2 were enriched in WG-3, but RsFLS was downregulated in HX-3 and WG-3. We proposed that RsDFR1, RsDFR2 and RsFLS might act as key regulators in the anthocyanin biosynthesis pathway (Gao et al., 2019a). In addition, several differentially expressed genes related to anthocyanin biosynthesis in carmine radish (Raphanus sativus L.) fleshy roots were identified using comparative RNA-Seq, including RsCHS, RsCHI, RsANS, RsMT2-4, RsUF3GT, glutathione S-transferase F12, RsUFGT78D2-like and RsUDGT-75C1-like, and these genes likely significantly contribute to the regulation of anthocyanin biosynthesis in radish cultivars (Gao et al., 2020). The dihydroflavonol 4-reductase (RsDFR: Cluster_13775), flavonoid 3′-monooxygenase (RsF3’H: Cluster_4431), leucoanthocyanidin dioxygenase (RsLDOX: Cluster_3903) and chalcone synthase (RsCHS: Cluster_39833) genes were identified and validated using qRT-PCR. These genes were significantly upregulated during the dynamic growth stages of fleshy roots relative to the other development periods from the seedling stage to the full-bloom stage but decreased in the podding stage, consistent with the dynamics of the anthocyanidin profile. In addition, the expression of UFGT genes has also been identified as an important regulatory mechanism involved in the anthocyanin biosynthetic pathway (Cutanda-Perez et al., 2009; Griesser et al., 2008) as UGFTs attach sugar moieties to the anthocyanin aglycone to stabilize anthocyanidin. In this study, UDP-glycosyltransferase 75C1-like (RsUGT75C1: Cluster_2736) and UDP-glycosyltransferase 78D2-like (RsUGT78D2: Cluster_11854) were significantly upregulated in different fleshy root types with the development of dynamic growth stages of fleshy roots from the seedling stage to the full-bloom stage but decreased in the podding stage. Further modifications involving glycosylation, acylation or methylation can be made; for example, anthocyanin is formed through a reaction catalyzed by a cyanidin 3-O-glycosyltransferase (Ni et al., 2018). Here, anthocyanidin 3-O-glucoside 2′″-O-xylosyltransferase (Rs3GGT1: Cluster_9270), coumaroyl-CoA:anthocyanidin 3-O-glucoside-6″-O-coumaroyltransferase 1-like (Rs3AT2: cluster_9270) were expressed as key regulatory enzymes for the formation of anthocyanins in carmine radish. In addition, transport proteins and transcription factors comprising glutathione S-transferase F12 (RsGSTF12: Cluster_24268), MYB transcription factor (RsMYB2: Cluster_28373), and zinc finger, RING-type protein (RsRZEP: Cluster_7186) were identified. Based on genetic and biochemical evidence, several GSTs were found to be involved in anthocyanin transport (Zhao & Dixon, 2010). Bz2 (encoded by the GST gene) was first found to act as an important regulator in the vacuolar transfer of anthocyanins in Zea mays (Marrs, Alfenito & Lloyd, 1995). In addition, anthocyanin accumulation and pigment mislocalization were found to be induced by mutations in GST-encoding genes in Arabidopsis (Kitamura, Shikazono & Tanaka, 2010). Here, we found that RsGSTF12 was significantly upregulated in different stages of fleshy root development. Moreover, MYB acts as a central regulator for determining the variation in anthocyanin production (Espley et al., 2009). A previous study demonstrated that the transcription of both early (CHS, CHI, F3′H and FLS) and late (DFR, ANS and ANR) flavonoid biosynthesis genes in the anthocyanin pathway was directly activated by R2R3-type MYB proteins and the MYB-bHLH-WD40 complex (Xu, Dubos & Lepiniec, 2015). Here, RsMYB2 was found to be significantly over-expressed and remarkably positively correlated with red pigment content in the dynamic development of fleshy roots in carmine radish. We therefore infer that RsMYB2 transcription factors might specifically activate flavonoid biosynthesis genes (RsCHS, RsF3′H, RsDFR) to regulate anthocyanin biosynthesis in carmine radish. However, the molecular mechanism of RsCHS, RsF3′H, and RsDFR regulation by RsMYB2 needs further study.
Conclusion
RNA-Seq technology was applied to identify the key anthocyanin biosynthesis-related genes involved in the regulation of anthocyanin biosynthesis during the dynamic growth stages of fleshy roots in carmine radish. Of those, two transcription factors, RsMYB2 and RsRZFP, as well as some related functional genes, e.g., RsCHS, RsDFR, RsLDOX, RsF′3H, RsF3GGT1, Rs3AT1, RsGSTF12, RsUFGT78D2 and RsUDGT-75C1, may contribute to the regulatory mechanism of anthocyanin synthesis. In addition, Pearson correlation analysis indicated excellent concordance between RNA-Seq and qRT-PCR experiments. Moreover, qRT-PCR showed that RsCHS, RsDFR, RsF′3H, RsUDGT-75C1, RsF3GGT1, Rs3AT1, RsMYB2 and RsRZFP were significantly upregulated and showed a remarkable positive correlation with the red pigment content in fleshy roots at different stages. We proposed that RsCHS, RsDFR, RsF′3H, RsUDGT-75C1, RsF3GGT1, Rs3AT1, and RsMYB2 might act as key regulators in the anthocyanin biosynthesis pathway.
Supplemental Information
qRT-PCR data for 11 selected DEGs related to anthocyanin biosythesis.
List of Abbreviations
- ASRGs
Anthocyanin synthesis-related genes
- PAL
Phenylalanine ammonia-lyase
- C4H
Cinnamate 4-hydroxylase
- 4CL
4-coumarate: CoA ligase
- CHS
Chalcone synthase
- CHI
Chalcone isomerase
- F3H
Flavanone 3-hydroxylase
- F35H
Flavonoid 3,5 –hydroxylase
- DFR
Dihydroflavonols 4-reductase
- ANS
Anthocyanin synthase
- UFGT
UDP-glucose Flavonoid 3-glucosyltransferase
- FNS
Flavone synthase
- FLS
Flavonol synthase
- MT
Metallothionein-like protein
- SS
seedling stage
- IE
initial expansion
- FE
full expansion
- BS
bolting stage
- IFS
initial flowering stage
- FBS
full-bloom stage
- PS
podding stage
- DEGs
differentially expressed genes
- KEGG
Kyoto Encyclopedia of Genes and Genomes
- FPKM
fragments per kilobase of transcript per million mapped reads
- GO
Gene Ontology
- NCBI
National Center for Biotechnology Information
- BP
biological process
- MF
molecular function
- CC
cellular component
Funding Statement
This research was supported by the Normal Program for the Science and Technology Research Program of Sichuan Municipal Education Commission (2020YJ0415), the Natural Science Foundation of CSTB (cstc2019jcyj-msxmX0652), and the Science and Technology Plan Projects of Fuling District (FLKJ, 2018BBB3009). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Contributor Information
Jian Gao, Email: gaojian_genomics@163.com.
Hua Peng, Email: penghua623@foxmail.com.
Additional Information and Declarations
Competing Interests
The authors declare that they have no competing interests.
Author Contributions
Xia Song conceived and designed the experiments, performed the experiments, prepared figures and/or tables, and approved the final draft.
Jian Gao conceived and designed the experiments, analyzed the data, prepared figures and/or tables, authored or reviewed drafts of the paper, and approved the final draft.
Hua Peng analyzed the data, authored or reviewed drafts of the paper, and approved the final draft.
Data Availability
The following information was supplied regarding data availability:
All the raw read sequences are available at the NCBI Sequence Read Archive: PRJNA565866.
References
- Aza-González et al. (2012).Aza-González C, Herrera-Isidrón L, Núñez-Palenius HG, Vega OMDL, Ochoa-Alejo N. Anthocyanin accumulation and expression analysis of biosynthesis-related genes during chili pepper fruit development. Biologia Plantarum. 2012;57(1):49–55. doi: 10.1007/s10535-012-0265-1. [DOI] [Google Scholar]
- Azuma (2017).Azuma A. Genetic and environmental impacts on the biosynthesis of anthocyanins in grapes. Horticulture Journal. 2017;87(3):1–17. [Google Scholar]
- Bajpai et al. (2017).Bajpai A, Khan K, Muthukumar M, Rajan S, Singh NK. Molecular analysis of anthocyanin biosynthesis pathway genes and their differential expression in mango peel. Genome. 2017;61(3):157–166. doi: 10.1139/gen-2017-0205. [DOI] [PubMed] [Google Scholar]
- Christian et al. (2013).Christian Q, Elmar P, Pelin Y, Jan G, Timmy S, Pablo Y, Jörg P, Oliver GF. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Research. 2013;41(D1):590–596. doi: 10.1093/nar/gks1219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crifò, Petrone & Recupero (2011).Crifò TPI, Petrone G, Recupero GR. Lo Piero AR.: expression analysis in response to low temperature stress in blood oranges: implication of the flavonoid biosynthetic pathway. Gene. 2011;476(1–2):1–2. doi: 10.1016/j.gene.2011.02.005. [DOI] [PubMed] [Google Scholar]
- Cutanda-Perez et al. (2009).Cutanda-Perez MC, Ageorges A, Gomez C, Vialet S, Terrier N, Romieu C, Torregrosa L. Ectopic expression of VlmybA1 in grapevine activates a narrow set of genes involved in anthocyanin synthesis and transport. Plant Molecular Biology. 2009;69(6):633–648. doi: 10.1007/s11103-008-9446-x. [DOI] [PubMed] [Google Scholar]
- Dao & Linthorst (2011).Dao TTH, Linthorst HJM. Verpoorte R: chalcone synthase and its functions in plant resistance. Phytochemistry Reviews. 2011;10(3):397–412. doi: 10.1007/s11101-011-9211-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dewey & Li (2011).Dewey CN, Li B. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics. 2011;12(1):323. doi: 10.1186/1471-2105-12-323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Espley et al. (2009).Espley RV, Brendolise C, Chagne D, Kutty-Amma S, Green S, Volz R, Putterill J, Schouten HJ, Gardiner SE, Hellens RP. Multiple repeats of a promoter segment causes transcription factor autoregulation in red apples. Plant Cell. 2009;21(1):168–183. doi: 10.1105/tpc.108.059329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ferreira, Slade & Marais (2005).Ferreira D, Slade D, Marais JPJ. Flavans and proanthocyanidins. In: Harborne JB, editor. The Flavonoids. Boston: Springer; 2005. [Google Scholar]
- Fukusaki et al. (2004).Fukusaki EI, Kawasaki K, Kajiyama SI, An CI, Suzuki K, Tanaka Y, Kobayashi A. Flower color modulations of Torenia hybrida by downregulation of chalcone synthase genes with RNA interference. Journal of Biotechnology. 2004;111(3):229–240. doi: 10.1016/j.jbiotec.2004.02.019. [DOI] [PubMed] [Google Scholar]
- Gao et al. (2019a).Gao J, Li WB, Liu HF, Chen FB. De novo transcriptome sequencing of radish (Raphanus sativus L.) fleshy roots: analysis of major genes involved in the anthocyanin synthesis pathway. BMC Molecular and Cell Biology. 2019a;2019(24):45. doi: 10.1186/s12860-019-0228-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao et al. (2020).Gao J, Li WB, Liu HF, Chen FB. Identification of differential expression genes related to anthocyanin biosynthesis in carmine radish (Raphanus sativus L.) fleshy roots using comparative RNA-Seq method. PLOS ONE. 2020;15(4):e0231729. doi: 10.1371/journal.pone.0231729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao et al. (2019b).Gao J, Peng H, Chen F, Luo M, Li W. Genome-wide analysis of transcription factors related to anthocyanin biosynthesis in carmine radish (Raphanus sativus L.) fleshy roots. PeerJ. 2019b;7(1):e8041. doi: 10.7717/peerj.8041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gou et al. (2011).Gou JY, Felippes FF, Liu CJ, Weigel D, Wang JW. Negative regulation of anthocyanin biosynthesis in arabidopsis by a miR156-targeted SPL transcription factor. Plant Cell. 2011;23(4):1512–1522. doi: 10.1105/tpc.111.084525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Griesser et al. (2008).Griesser M, Hoffmann T, Bellido ML, Rosati C, Fink B, Kurtzer R, Aharoni A, Munoz-Blanco J, Schwab W. Redirection of flavonoid biosynthesis through the down-regulation of an anthocyanidin glucosyltransferase in ripening strawberry fruit. Plant Physiology. 2008;146(4):1528–1539. doi: 10.1104/pp.107.114280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grotewold (2006).Grotewold E. The genetics and biochemistry of floral pigments. Annual Review of Plant Biology. 2006;57(1):761–780. doi: 10.1146/annurev.arplant.57.032905.105248. [DOI] [PubMed] [Google Scholar]
- He & Giusti (2010).He J, Giusti MM. Anthocyanins: natural colorants with health-promoting properties. Annual Review of Food Science & Technology. 2010;1(1):163–187. doi: 10.1146/annurev.food.080708.100754. [DOI] [PubMed] [Google Scholar]
- Jian et al. (2015).Jian G, Mao L, Zhang C, Hua P, Lin H, Shen Y, Zhao M, Pan G, Zhang Z. A putative pathogen-resistant regulatory pathway between MicroRNAs and candidate target genes in maize. Journal of Plant Biology. 2015;58(4):211–219. doi: 10.1007/s12374-014-0572-5. [DOI] [Google Scholar]
- Khoo et al. (2017).Khoo HE, Azlan A, Tang ST, Lim SM. Anthocyanidins and anthocyanins: colored pigments as food, pharmaceutical ingredients, and the potential health benefits. Food & Nutrition Research. 2017;61(1):1361779. doi: 10.1080/16546628.2017.1361779. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kitamura, Shikazono & Tanaka (2010).Kitamura S, Shikazono N, Tanaka A. TRANSPARENT TESTA 19 is involved in the accumulation of both anthocyanins and proanthocyanidins in Arabidopsis. Plant Journal. 2010;37(1):104–114. doi: 10.1046/j.1365-313X.2003.01943.x. [DOI] [PubMed] [Google Scholar]
- Kobayashi et al. (2001).Kobayashi S, Ishimaru M, Ding CK, Yakushiji H, Goto N. Comparison of UDP-glucose: flavonoid 3-O-glucosyltransferase (UFGT) gene sequences between white grapes (Vitis vinifera) and their sports with red skin. Plant Science. 2001;160(3):543–550. doi: 10.1016/S0168-9452(00)00425-8. [DOI] [PubMed] [Google Scholar]
- Latchman (1993).Latchman DS. Transcription factors: an overview. International Journal of Biochemistry & Cell Biology. 1993;74(5):417–422. [PMC free article] [PubMed] [Google Scholar]
- Lepiniec et al. (2006).Lepiniec Lïc, Debeaujon I, Routaboul J-M, Baudry A, Pourcel L, Nesi N, Caboche M. Genetics and biochemistry of seed flavonoids. Annual Review of Plant Biology. 2006;57(1):405–430. doi: 10.1146/annurev.arplant.57.032905.105252. [DOI] [PubMed] [Google Scholar]
- Li et al. (2011).Li X, Gao P, Cui D, Wu L, Parkin I, Saberianfar R, Menassa R, Pan H, Westcott N, Gruber MY. The Arabidopsis tt19-4 mutant differentially accumulates proanthocyanidin and anthocyanin through a 3′ amino acid substitution in glutathione S-transferase. Plant Cell & Environment. 2011;34(3):374–388. doi: 10.1111/j.1365-3040.2010.02249.x. [DOI] [PubMed] [Google Scholar]
- Li & Richard (2009).Li H, Richard D. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;2010(14):1754–1760. doi: 10.1093/bioinformatics/btp324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li et al. (2012).Li X, Sun H, Pei J, Dong Y, Wang F, Chen H, Sun Y, Wang N, Li H, Li Y. De novo sequencing and comparative analysis of the blueberry transcriptome to discover putative genes related to antioxidants. Gene. 2012;511(1):54–61. doi: 10.1016/j.gene.2012.09.021. [DOI] [PubMed] [Google Scholar]
- Marrs, Alfenito & Lloyd (1995).Marrs KA, Alfenito MR, Lloyd AM. Walbot V: a glutathione-s-transferase involved in vacuolar transfer encoded by the maize gene bronze-2. Nature. 1995;375(6530):397–400. doi: 10.1038/375397a0. [DOI] [PubMed] [Google Scholar]
- Ni et al. (2018).Ni X, Xue S, Iqbal S, Wang W, Gao Z. Candidate genes associated with red colour formation revealed by comparative genomic variant analysis of red- and green-skinned fruits of Japanese apricot (Prunus mume) PeerJ. 2018;6(3):e4625. doi: 10.7717/peerj.4625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qi et al. (2011).Qi T, Song S, Ren Q, Wu D, Huang H, Chen Y, Fan M, Peng W, Ren C, Xie D. The Jasmonate-ZIM-domain proteins interact with the WD-Repeat/bHLH/MYB complexes to regulate jasmonate-mediated anthocyanin accumulation and trichome initiation in Arabidopsis thaliana. Plant Cell. 2011;23(5):1795–1814. doi: 10.1105/tpc.111.083261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schefe et al. (2006).Schefe JH, Lehmann KE, Buschmann IR, Unger T, Funke-Kaiser H. Quantitative real-time RT-PCR data analysis: current concepts and the novel “gene expression’s CT difference” formula. Journal of Molecular Medicine. 2006;84(11):901–910. doi: 10.1007/s00109-006-0097-6. [DOI] [PubMed] [Google Scholar]
- Soonyoung, Seonae & Yun (2016).Soonyoung A, Seonae K, Yun HK. Glutathione S-transferase genes differently expressed by pathogen-infection in Vitis flexuosa. Plant Breeding & Biotechnology. 2016;4(1):61–70. [Google Scholar]
- Tarazona et al. (2012).Tarazona S, García F, Ferrer A, Dopazo J, Conesa A. NOIseq: a RNA-seq differential expression method robust for sequencing depth biases. University of Southampton. 2012;17(B):18. [Google Scholar]
- Xie (2003).Xie D-Y. Role of Anthocyanidin Reductase, Encoded by BANYULS in Plant Flavonoid Biosynthesis. Science. 2003;299(5605):396–399. doi: 10.1126/science.1078540. [DOI] [PubMed] [Google Scholar]
- Xie et al. (2011).Xie C, Mao X, Huang J, Ding Y, Wu J, Dong S, Kong L, Gao G, Li C-Y, Wei L. KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Research. 2011;39(suppl_2):W316–W322. doi: 10.1093/nar/gkr483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu, Dubos & Lepiniec (2015).Xu W, Dubos C, Lepiniec L. Transcriptional control of flavonoid biosynthesis by MYB-bHLH–WDR complexes. Trends in Plant Science. 2015;20(3):176–185. doi: 10.1016/j.tplants.2014.12.001. [DOI] [PubMed] [Google Scholar]
- Yousuf et al. (2015).Yousuf B, Gul K, Wani AA, Singh P. Health benefits of anthocyanins and their encapsulation for potential use in food systems: a review. Critical Reviews in Food Science and Nutrition. 2015;56(13):2223–2230. doi: 10.1080/10408398.2013.805316. [DOI] [PubMed] [Google Scholar]
- Yusuf et al. (2012).Yusuf D, Butland SL, Swanson MI, Bolotin E, Ticoll A, Cheung WA, Zhang XYC, Dickman CT, Fulton DL, Lim JS. The transcription factor encyclopedia. Genome Biology. 2012;13(3):R24. doi: 10.1186/gb-2012-13-3-r24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang et al. (2011).Zhang X, C Allan A, Yi Q, Chen L, Li K, Shu Q, Su J. Differential gene expression analysis of yunnan red pear, pyrus pyrifolia, during fruit skin coloration. Plant Molecular Biology Reporter. 2011;29(2):305–314. doi: 10.1007/s11105-010-0231-z. [DOI] [Google Scholar]
- Zhao & Dixon (2010).Zhao J, Dixon RA. The ‘ins’ and ‘outs’ of flavonoid transport. Trends in Plant Science. 2010;15(2):72–80. doi: 10.1016/j.tplants.2009.11.006. [DOI] [PubMed] [Google Scholar]
- Zhou et al. (2015).Zhou H, Lin-Wang K, Wang H, Gu C, Dare AP, Espley RV, He H, Allan AC, Han Y. Molecular genetics of blood-fleshed peach reveals activation of anthocyanin biosynthesis by NAC transcription factors. Plant Journal. 2015;82(1):105–121. doi: 10.1111/tpj.12792. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
qRT-PCR data for 11 selected DEGs related to anthocyanin biosythesis.
Data Availability Statement
The following information was supplied regarding data availability:
All the raw read sequences are available at the NCBI Sequence Read Archive: PRJNA565866.