Background – Canine atopic dermatitis (cAD) is a common inflammatory and pruritic skin disease, with various treatment options. The use of topical products containing natural ingredients has proven increasingly popular. Objective – To evaluate the effects of a spray solution containing heat‐killed Lactobacillus rhamnosus and L. reuteri, on the clinical signs and cutaneous microbiota of atopic dogs. Conclusions and clinical importance – There was a significant and rapid decrease in the clinical signs associated with cAD after use of the spray. Future larger, randomized, controlled studies are needed to confirm these results and to assess the effects on the cutaneous immunity and microflora of atopic dogs.
![]()
Abstract
Background
Canine atopic dermatitis (cAD) is a common inflammatory and pruritic skin disease, with various treatment options. The use of topical products containing natural ingredients has proven increasingly popular.
Objective
To evaluate the effects of a spray solution containing heat‐killed Lactobacillus rhamnosus and L. reuteri, on the clinical signs and cutaneous microbiota of atopic dogs.
Animals
Ten privately owned, mildly affected, nonseasonally atopic dogs.
Methods and materials
The spray was applied to the ventrum every 24 h for 28 days. Clinical scores, skin barrier function and owner assessment were evaluated on day (D)0, D14, D28 and D42. The cutaneous microbiota was analysed on D0 and D28.
Results
A reduction in the total clinical score was seen at each time point (D14, P = 0.03; D28, P = 0.04; D42, P = 0.001). A reduction in the regional clinical scores was seen after D28 (P = 0.01) and D42 (P = 0.003). A significant reduction in the pruritus score was seen on D42 (P = 0.01). A lower hydration value was seen on D28 (P = 0.02) and D42 (P = 0.02) on the pinnae. A good‐to‐excellent response and an easy‐to‐use administration was reported by owners. There were no significant changes in the cutaneous microbiota after 28 days.
Conclusions and clinical importance
There was a significant and rapid decrease in the clinical signs associated with cAD after use of the spray. Future larger, randomized, controlled studies are needed to confirm these results and to assess the effects on the cutaneous immunity and microflora of atopic dogs.
Résumé
Contexte
La dermatite atopique canine (DAC) est une dermatose inflammatoire et prurigineuse fréquente avec de nombreuses options thérapeutiques. L’utilisation de topiques contenant des ingrédients naturels rencontre un succès grandissant.
Objectif
Evaluer les effets d’une solution en spray contenant Lactobacillus rhamnosus et L. reuteri tué par la chaleur, sur les signes cliniques et le microbiote cutané des chiens atopiques.
Sujets
Dix chiens de propriétaires, atteints de dermatite atopique non saisonnière modérée.
Matériels et méthodes
Le spray a été appliqué sur l’abdomen toutes les 24h pendant 28 jours. Les scores cliniques, la fonction barrière cutanée et l’estimation des propriétaires ont été évalué à jour (J) 0, J14, J28 et J42. Le microbiote cutané a été analysé à J0 et J28.
Résultats
Une diminution du score clinique total a été observé à chaque point de contrôle (J14, P = 0.03; J28, P = 0.04; J42, P = 0.001). Une diminution des scores cliniques régionaux a été vu après J28 (P = 0.01) et J42 (P = 0.003). Une diminution significative du score de prurit a été vu à J42 (P = 0.01). Une valeur d’hydratation plus faible a été vue à J28 (P = 0.02) et J42 (P = 0.02) sur le pavillon auriculaire. Une réponse bonne à excellente et une administration facile a été notée par les propriétaires. Il n’y avait pas de changement significatif dans le microbiote cutané après 28 jours.
Conclusions et importance clinique
Il y avait une diminution rapide et significative des signes cliniques associés à la DAC après utilisation du spray. D’autres études plus larges, randomisées, contrôlées sont nécessaires pour confirmer ces résultats et pour déterminer les effets sur l’immunité cutanée et la microflore des chiens atopiques.
Resumen
Introducción
la dermatitis atópica canina (cAD) es una enfermedad cutánea inflamatoria y pruriginosa común, con varias opciones de tratamiento. El uso de productos tópicos que contienen ingredientes naturales ha demostrado ser cada vez más popular.
Objetivo
evaluar los efectos de una solución en aerosol que contiene Lactobacillus rhamnosus y L. reuteri destruidos por calor, en los signos clínicos y la microbiota cutánea de perros atópicos.
Animales
diez perros de propietarios particulares, levemente afectados, atópicos no‐estacionales.
Métodos y materiales
El aerosol se aplicó en la zona ventral cada 24 h durante 28 días. Las puntuaciones clínicas, la función de barrera cutánea y la valoración del propietario se evaluaron los días (D) 0, D14, D28 y D42. La microbiota cutánea se analizó en D0 y D28.
Resultados
se observó una reducción en la valoración clínica total en cada momento (D14, P = 0,03; D28, P = 0,04; D42, P = 0,001). Se observó una reducción en las valoraciones clínicas regionales después de D28 (P = 0,01) y D42 (P = 0,003). Se observó una reducción significativa en la valoración de prurito en D42 (P = 0,01). Se observó un valor de hidratación más bajo en D28 (P = 0.02) y D42 (P = 0.02) en el pabellón auricular. Los propietarios informaron de una respuesta buena a excelente y una administración fácil de usar. No hubo cambios significativos en la microbiota cutánea después de 28 días.
Conclusiones e importancia clínica
Hubo una disminución rápida y significativa de los signos clínicos asociados con la cAD después del uso del aerosol. Se necesitan estudios controlados, al azar y con mayor número de animales en el futuro para confirmar estos resultados y evaluar los efectos sobre la inmunidad cutánea y la microflora de los perros atópicos.
Zusammenfassung
Hintergrund
Die canine atopische Dermatitis (cAD) ist eine häufige entzündliche und juckende Hauterkrankung, für die es verschiedene Behandlungsmöglichkeiten gibt. Die Verwendung von topischen Produkten, die natürliche Wirkstoffe beinhalten, haben sich als zunehmend beliebt gezeigt.
Ziel
Eine Evaluierung der Wirksamkeit einer Spraylösung, welche durch Hitze‐abgetötete Lactobacillus rhamnosus und l. reuteri beinhaltete, auf die klinischen Zeichen und die kutanen Mikrobiota der atopischen Hunde.
Tiere
Zehn private nicht‐saisonal atopische Hunde , die nur mild betroffen waren.
Methoden und Materialien
Der Spray wurde 28 Tage lang alle 24 h auf dem Ventrum aufgetragen. Klinische Werte, die Funktion der Hautbarriere und eine Beurteilung der BesitzerInnen wurde am Tag (D)0, D14, D28 und D42 evaluiert. Die kutanen Mikrobiota wurden am D0 und D28 analysiert.
Ergebnisse
Eine Reduzierung des klinischen Gesamtwerts wurde zu jedem Zeitpunkt festgestellt (D14, P = 0,03; D28, P = 0,04; D42, P = 0,001). Eine Reduzierung der lokalen klinischen Werte konnte nach D28 (P = 0,01) und D42 (P = 0,003) festgestellt werden. Eine signifikante Verminderung der Juckreizwerte wurde am D42 (P = 0,01) festgestellt. Ein niedrigerer Wert wurde am D28 (P = 0,02) an den Ohren festgestellt. Die BesitzerInnen berichteten von einer guten bis ausgezeichneten Reaktion und einer leichte Methode der Anwendung. In den kutanen Mikrobiota bestanden nach 28 Tagen keine signifikanten Unterschiede.
Schlussfolgerungen und klinische Bedeutung
Nach Verwendung des Sprays gab es eine signifikante und rasche Abnahme der klinischen Zeichen der cAD. Es sind zukünftige, randomisierte, kontrollierte Studien nötig, um diese Ergebnisse zu bestätigen und die Auswirkungen auf die kutane Immunität und die Mikroflora atopischer Hunde zu bestätigen.
概要
背景
犬特应性皮炎(cAD)是一种常见的炎性和瘙痒性皮肤病, 有多种治疗选择。使用含有天然成分的外用产品已被证明越来越受欢迎。
目的
评价含有热灭活鼠李糖乳杆菌和罗伊氏乳杆菌的喷雾溶液对特应性患犬临床体征和皮肤微生物群的影响。
动物
10只轻度发病、非季节性特应性私家犬。
方法和材料
每24h将喷剂应用于腹部, 持续28天。在第0天(D)、D14、D28和D42进行临床和皮肤屏障功能评分和主人评分。在D0和D28分析皮肤微生物群。
结果
在每个时间点均观察到临床总评分降低(D14,P = 0.03;D28,P = 0.04;D42,P = 0.001)。D28(P = 0.01)和D42(P = 0.003)后观察到外部临床评分降低。在D42观察到瘙痒评分显著降低(P = 0.01)。在D28(P = 0.02)和D42(P = 0.02)观察到耳廓上的水合值较低。犬主人给出了良好至极佳的药效和给药方便的反馈。28天后皮肤菌群无明显变化。
结论和临床重要性
使用喷剂后, 与cAD相关的临床体征显著快速减少。未来需要更大规模、随机、对照研究来证实这些结果, 并评估对特应性犬皮肤免疫和微生物群落的影响。
要約
背景
犬アトピー性皮膚炎(cAD) は、一般的な炎症性および掻痒性皮膚疾患であり、さまざまな治療オプションがある。天然成分含有外用製品の使用はますます人気があることが証明されている。
目的
本研究の目的は、アトピー犬の臨床徴候と皮膚微生物叢に対する、熱殺菌したLactobacillus rhamnosuおよびL. reuteri含有スプレー液の効果を評価することであった。
被験動物
個人所有の、軽症、非季節性アトピー犬10頭。
材料と方法
スプレーを24時間ごとに28日間腹側に塗布した。臨床スコア、皮膚バリア機能、および所有者の評価を、 (D)0、D14、D28、およびD42日に評価した。皮膚微生物叢はD0およびD28に解析した。
結果
各時点で総臨床スコアの低下が見られた (D14、P = 0.03; D28、P = 0.04; D42、P = 0.001) 。 D28(P = 0.01) およびD42(P = 0.003) 後に、局部の臨床スコア低下が見られた。掻痒スコアの有意な低下がD42で見られた (P = 0.01) 。D28(P = 0.02)およびD42(P = 0.02) で、耳介のより低い水和値が見られた。所有者から良〜優れた反応および使いやすい管理が報告された。 28日後、皮膚微生物相に有意な変化は認められなかった。
結論と臨床的重要性
スプレー使用後、cADに関連する臨床徴候が有意かつ急速に減少した。これらの結果を確認し、アトピー犬の皮膚免疫および微生物叢への影響を評価するには、将来のより大規模なランダム化比較試験が必要である。
Resumo
Contexto
A dermatite atópica canina (DAC) é uma doença cutânea inflamatória e pruriginosa comum, com várias opções de tratamento. O uso de produtos tópicos contendo ingredientes naturais tem se mostrado cada vez mais popular.
Objetivo
Avaliar os efeitos de uma solução em spray contendo Lactobacillus rhamnosus e L. reuteri inativados pelo calor, nos os sinais clínicos e na microbiota cutânea de cães atópicos.
Animais
Dez cães atópicos não sazonais de propriedade privada, levemente afetados.
Métodos e materiais
O spray foi aplicado no abdômen a cada 24 horas por 28 dias. Escores clínicos, função de barreira cutânea e avaliação do proprietário foram avaliadas nos dias (D) 0, D14, D28 e D42. A microbiota cutânea foi analisada em D0 e D28.
Resultados
Observou‐se uma redução no escore clínico total em todos os momentos (D14, P = 0,03; D28, P = 0,04; D42, P = 0,001). Observou‐se uma redução nos escores clínicos regionais após D28 (P = 0,01) e D42 (P = 0,003). Uma redução significativa no escore de prurido foi observada no D42 (P = 0,01). Observou‐se um valor de hidratação inferior no D28 (P = 0,02) e no D42 (P = 0,02) no pavilhão auricular. Os proprietários relataram resposta boa a excelente e fácil administração. Não houve alterações significativas na microbiota cutânea após 28 dias.
Conclusões e importância clínica
Houve diminuição significativa e rápida dos sinais clínicos associados à DAC após o uso do spray. Futuros estudos maiores, randomizados e controlados são necessários para confirmar esses resultados e avaliar os efeitos sobre a imunidade cutânea e microflora de cães atópicos.
Introduction
Atopic dermatitis (AD) is one of the most common chronic inflammatory skin diseases in dogs. Genetic and immune system alterations along with defects of the skin barrier and modifications of the cutaneous microflora have been reported in atopic dogs. 1 , 2 , 3 , 4 Multiple treatment options are available; 5 , 6 there is a growing interest in the use of alternative products based on so‐called natural ingredients. 7
In human AD there are reports of the potential positive effects of topical applications of heat‐killed beneficial bacteria. 8 , 9 , 10 In particular, the use of topical products containing heat‐killed bacteria (Vitreoscilla filiformis, Lactobacillus johnsonii and L. reuteri) has been reported to show a significant decrease in clinical signs and pruritus in both atopic people and mice. 8 , 9 , 10 , 11 There was an overall reduction in the Staphylococcus aureus population reported in one study. 10 Some studies report that the use of rapidly killed beneficial bacteria (e.g. lactobacilli) can ameliorate the clinical signs of human AD and positively stimulate the local immunity, to the point of influencing the local microbiome. 12 , 13 , 14
The objective of this preliminary, noncontrolled, prospective clinical trial was to assess the effects of a spray version of a veterinary product containing heat‐killed lactobacilli (L. rhamnosus and L. reuteri) marketed as an adjuvant therapy for allergic dogs (LinkSkin spray, DRN Inc.; Cremona, Italy) on the clinical signs of canine (c)AD (primary outcome), the skin barrier and the cutaneous microbiota (secondary outcomes).
Methods and materials
Animal welfare documentation
All dogs were recruited at the authors’ institution with previous approval from the University of Florida and a signed owner consent form.
Canine population
Naturally affected, privately owned dogs with mild‐to‐moderate [Canine Atopic Dermatitis Extent and Severity Index, 4th iteration CADESI‐04 with scores of 10 to 59] 15 nonseasonal AD were enrolled in the study. Diagnosis of cAD was based on published guidelines. 16 All dogs received monthly flea prevention.
Exclusion criteria
Dogs with clinical evidence or a history of malignant neoplasia, metabolic/endocrine, parasite or other allergic skin conditions (food/contact) were excluded. Dogs with superficial infections, diagnosed based on published guidelines, 17 were excluded. Owners withdrew drugs for their dog using the following periods: oral and topical antibiotics and antifungal agents (four weeks), corticosteroids (eight weeks for depo‐injectable, four weeks for oral, three weeks for topical), antihistamines (two weeks), ciclosporin (four weeks), lokivetmab (eight weeks) and oclacitinib (one week). Dogs receiving allergen‐specific immunotherapy for less than one year before enrolment were excluded from the study.
Intervention
The treatment consisted of a water‐based spray solution containing a mixture of lactobacilli (L. reuteri and L. rhamnosus) together with the culture media containing metabolites produced by the bacteria, tamarind extract and polyphenols. This solution had undergone Tyndallization, a process used to sterilize food from heat‐resistant endospores and bacteria, based on the use of heat to reach boiling point (~100°C) for 15 min for three consecutive days. This methodology, although designed to allow endospores to germinate and be killed, allows minimal degradation of the bacteria walls with minimal alteration of the immunological properties of the bacteria. 12 , 13 , 14
Other topical and/or systemic treatments including antimicrobial, humectant or antipruritic, were not allowed during the study. Owners sprayed the glabrous areas (axillae, ventral thorax, inguinal area and medial thighs) every 24 h for 28 days, irrespective of the distribution of the dog’s skin disease. Licking and subsequent alteration of the cutaneous microbiota was prevented by distracting the dog and, when unsupervised, placing a T‐shirt on the dog, throughout the 28 days of the study. The use of the T‐shirt was preferred to the use of an E‐collar for ethical reasons and tolerability. Each dog was clinically evaluated by the same investigator on Day (D)0, D14, D28 and D42.
Clinical assessment
The primary outcome of this study was to assess the clinical efficacy of the spray for cAD. On D0, D14, D28 and D42, the severity of clinical signs was assessed using the CADESI‐04 score system recording 20 body sites that are commonly affected in AD (total CADESI‐04). These areas were scored on a scale between 0 and 3 (0,none;1, mild; (2, moderate; 3, severe) with a maximum score of 180. 15 A separate CADESI‐04 score (regional) was calculated adding the score of only the treated areas (axillae, ventral thorax, inguinal area and medial thighs). At each visit, the pruritus was scored by the owners using a pruritus Visual Analog Scale (pVAS). 18 , 19 The percentage of dogs with a CADESI‐04 N (<10) and with a pVAS10‐N (<2) was calculated. 20 An owner global assessment of treatment efficacy (OGATE) score was recorded using a scale between 0 and 4 (0, no response; 1, poor; 2, fair; 3, good; 4, excellent). The percentage of dogs with an OGATE‐G2E (>2) was calculated in a similar way to the CADESI‐04 N. 20 An “ease to administer” score by the owners was recorded, based on asking the owners if the product was not easy, easy, or very easy to use. 21 Parameters considered by the owners included ease‐of‐spray, skin residues, stickiness and time of absorption.
Skin barrier function evaluation
As secondary outcome, the skin barrier function was assessed in all dogs on D0, D14, D28 and D42; assessing skin hydration via corneometry and pH. These parameters were analysed on the inguinal, axillary, pedal and aural surfaces using a corneometer and expressed as microsiemens values [µS] (Corneometer CM825, Courage + Khazaka electronic GmbH; Cologne, Germany) and a pH meter (Skin‐pH‐meter PH 905, Courage + Khazaka electronic GmbH) as described previously. 22 Before each evaluation, if present, the T‐shirt was removed and dogs were allowed to acclimatise to the testing room for 30 min. Temperature and humidity settings were maintained at 25 ± 5°C and 50 ± 10% for the entire duration of the study and as recommended by the manufacturer to optimize the performance of the instruments.
Skin microbiota collection
Another secondary outcome was the assessment of the cutaneous microbiota before (D0) and after treatment (D28). Two skin swabs, one from the groin and one from both axillae, were collected using swab applicators (Isohelix DNA Buccal Swabs, Cell Projects Ltd; Harrietsham, Kent, UK) before the acclimatisation period. Each swab applicator was rubbed on the skin 20 times (10 strokes per swab side) per anatomical site, as reported previously. 23 In addition, an extra swab agitated for 20 s in the air was submitted as a negative control, to control for environmental contaminants. All samples were stored immediately at –80°C until processed.
Library construction of microbial samples
For each extraction, 5 ng gDNA was used for 16S library construction using Quick‐16S NGS Library Prep Kit (Zymo Research Corp.; Irvine, CA, US). PCR amplification was performed using V3–V4 primers (95°C for 10 min, 20 cycles of (95°C for 30 s, 55°C for 30 s and 72°C for 3 min). The PCR products were quantified by real‐time PCR with a Bio‐Rad CFX Real‐Time PCR Detection System (Bio‐Rad Laboratories; Hercules, CA, USA) and cleaned with the Zymo enzymatic cleanup kit. Each library was barcoded with five cycles of PCR amplification and cleaned with the Select‐a‐Size MagBeads (Zymo Research Corp.). All 48 libraries were pooled together for one MiSeq 2 x 250 cycles run.
Statistical analyses
Clinical and skin barrier assessment. Data were analysed using the intention‐to‐treat analysis (ITT) with the last value carried forward. Data collection was tested for normality using the Shapiro–Wilk test (alpha level = 0.05). Then repeated measures ANOVA (skin hydration and pH) or Friedman’s test (CADESI‐04 and pVAS) followed by Dunnett’s or Dunn’s multiple comparison test were performed to evaluate the behaviour of each variable over time. A P‐value of <0.05 was considered statistically significant. All statistical analyses were performed using prism 6.09 statistical software (GraphPad Software Inc.; La Jolla, CA, USA).
Data analysis of microbial samples. Reads of 16S RNA (V3–V4 region) acquired from the Illumina MiSeq system were cleaned up with the cutadapt program (v2.8; https://cutadapt.readthedocs.io/en/stable/) 24 to trim off sequencing adaptors, low‐quality bases and potential errors introduced during sequencing. Only sequences with length ≥60 bp were included in the analyses. The cleaned paired‐end reads were merged and analysed using Quantitative Insight Into Microbial Ecology (QIIME, v1.9.1; http://qiime.org/). 25
The merged 16S rRNA sequences were grouped into operational taxonomic units (OTUs) (minimum OTU cluster size = 2; OTU similarity = 0.97) using both the open reference and closed reference OTU picking strategy and classified using the SILVA 16S/18S reference database. 26 The potential chimeric sequences and OTUs were detected and filtered at 97% cluster identity by Vsearch (v2.13.4; https://manpages.debian.org/stretch/vsearch/vsearch.1). 27 Taxonomies were summarized at multiple taxonomic levels (L2–L6) as the OTU tables with the samtools and R‐based scripts developed at the Interdisciplinary Center for Biotechnology Research, University of Florida. The sequencing coverage was evaluated by rarefaction analysis and the estimated species richness and diversity indices were calculated with the alpha‐diversity analysis. The differences of the microbial compositions between different samples were compared with the beta‐diversity analysis, including the principal coordinate analysis (PCoA) based on the Bray–Curtis distance model in R/phyloseq. In addition, the quantitative comparison of taxa at different taxonomic levels between the different groups was performed based on the raw OTU tables using the DEB application. 28
Results
Dogs
Ten dogs (five males, five females) were enrolled for this study. The average age was 5.3 ± 3.6 years. The average weight was 28.2 ± 24 kg. Breeds included great Dane (2), mixed‐breed dogs (3), pit bull terrier (2), and one each of German shepherd dog, Jack Russell terrier and French bulldog. One mixed‐breed dog was withdrawn from the study after D28 owing to relocation of the owner.
Clinical assessment
Compared to D0, a significant reduction in the total CADESI‐04 was seen at each time point (P = 0.028; P = 0.036; P = 0.001, respectively) (Figure S1). In particular, on D0, a median (range) total CADESI‐04 score of 16.5 (13–32) was recorded. This score decreased to 8.5 (3–25) on D14, 9.5 (3–18) on D28 and 6.5 (0–14) on D42. A CADESI‐04 N (<10) was achieved in 50% (five of 10), 50% and 90% (nine of 10) of dogs after D14, D28 and D42, respectively. Likewise, compared to D0, a reduction of the regional CADESI‐04 was seen after 28 (P = 0.013) and 42 (P = 0.003) days (Figure 1). In particular, on D0, a median (range) regional CADESI‐04 score of 6 (0–15) was recorded. This score decreased to 3 (0–6) on D14, 3 (0–4) on D28 and 0.5 (0–3) on D42.
Figure 1.
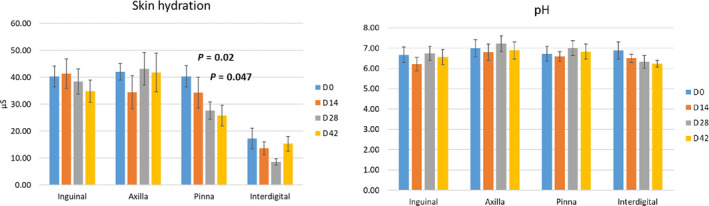
Average of the individual skin hydration and cutaneous pH values.
Hydration values are expressed in microsiemens (µS) as conductance units. Groups were compared using repeated measurements ANOVA with Dunnett's multiple comparison tests. Bars, SE.
Compared to D0, a significant reduction in the pVAS was seen on D42 (P = 0.013) (Figure S2). On D0, the pVAS was 4.9 ± 1.9, whereas on D14 it was 5.2 ± 2.2 and decreased to 4.1 ± 2.1 on D28 and to 2.5 ± 1.1 on D42. A pVAS10‐N (<2) was achieved in 10% (one of 10), 30% (three of 10) and 30% of dogs on D14, D28 and D42, respectively.
On D14, OGATEs of a fair response by seven (70%) owners and a good response by two (20%) owners were recorded (Figure S3). On D28, there was an OGATE equivalent to a fair response by three (30%) owners and a good response by six (60%) owners (Figure S3). On D42, OGATEs of a fair response by one (10%) owner and a good response by six (60%) owners and an excellent response by two (20%) owners were recorded (Figure S3). An OGATE of a poor response was reported by one (10%) owner for the entire duration of the study. An OGATE‐G2E (>2) was achieved in (two) 20%, (six) 60% and 60% of dogs on D14, D28 and D42, respectively.
On D14, two (20%) owners reported the product to be easy to use, whereas eight (80%) owners judged the product very easy to use (Figure S3). Likewise, on D28 and D42, one (10%) owner reported the product to be easy to use, whereas nine (90%) owners judged the product very easy to use (Figure S3).
During the study, major adverse effects were not observed in any dog. Only one dog had a noninfective, papular eruption after 14 days in the study. However, the eruption was not severe enough for the dog to be dropped from the study. The eruption went away after a series of topical shampoos containing oatmeal (not allowed during the study).
Skin barrier function evaluation
A significant change in the skin hydration and pH of the inguinal, axillary and interdigital areas was not seen at any time point (Figure 1). A significant reduction in the skin hydration, not the pH, was seen on the pinnae on D28 (P = 0.02) and D42 (P = 0.047) when compared to D0 (Figure 1).
Skin microbiota evaluation
Microbiome gDNA extractions were processed by using ZymoBIOMICS DNA kits following the user guide. Bacterial and archaeal microbial communities of 40 canine skin samples were characterized by sequencing the V3–V4 regions of PCR‐amplified 16S rRNA genes. An average of 160,137 paired‐end reads for each sample and approximately 159,491 cleaned merged reads (average 440 bp) were obtained for metagenomics analysis. Rarefaction analysis and alpha diversity measures showed that the bacterial communities were sufficiently sampled and that further sequencing would have been unlikely to significantly increase the observed microbial diversity detected.
Altogether, an average of 3,322 unique OTUs, ranging from 963 to 10,850, were obtained from each sample corresponding to about 14 phyla, 28 classes, 57 orders, 95 families and 164 genera of prokaryotes (Figures S4–S12). All microbial communities analysed in both groups (before and after treatment) were dominated by a few microbial genera, including Staphylococcus (Firmicutes), Conchiformibius (Proteobacteria) and Porphyromonas (Bacteroidetes) (Figures S4–S12). The microbial community profiles of the different samples based on the OTU tables and the Bray–Curtis distance model showed some differences between the two groups (Figures S4–S12). However, these differences did not reach a statistically significant level (Figure 2).
Figure 2.
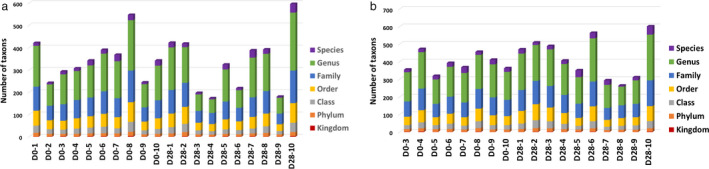
The distribution of operational taxonomic units at different taxonomic levels in the (a) axillae and (b) inguinal samples.
Discussion
To the best of the authors’ knowledge, this is the first study to report a potentially beneficial effect from the topical use of heat‐treated bacteria in the management of cAD. A significant reduction in the clinical score and pruritus was observed for two weeks after the spray was stopped, showing a residual beneficial effect. In addition, most of the owners deemed the product to be easy to very‐easy to use and effectively to manage cAD. An overall improvement of the clinical signs was seen in areas not directly treated by the product (e.g. feet and head). Even so, no changes in the cutaneous microbiota were seen after 28 days of daily application of the compound.
In the present study, the daily application of the spray significantly and relatively rapidly decreased the clinical signs of dogs with mild‐to‐moderate AD without the use of any other therapeutic intervention. The only allowed “intervention” was the use of a T‐shirt when the dog was unsupervised. However, this was not considered sufficient to influence the results because the T‐shirt was not constantly worn and because the dogs were still able to scratch and rub. The amelioration of the disease was not limited only to the treated areas, but also to the rest of the body, suggesting a beneficial effect of the product on the systemic, not only local, immune response. These results are in agreement with previous studies showing an amelioration of the clinical signs of AD in people after oral and topical administration of heat‐killed lactobacilli. 10 , 29 , 30 The present study also is in agreement with a previous human clinical trial that used a topical product containing Vetrosciella filiformis extract in atopic patients. 8 In that study, there was a a significant improvement reported in clinical signs, pruritus and quality of sleep, after only 28 days of daily application of heat‐killed bacteria. 8 Altogether, these results suggest that a short‐term daily application of heat‐killed beneficial bacteria can be used as adjuvant therapy in AD.
Given that no significant changes in the microbiota were detected, it is possible to hypothesize that heat‐treated lactobacilli do not directly interfere with the cutaneous microbiota. Rather, they may have a profound effect on the local and systemic immune response in atopic dogs. Several studies have shown the beneficial direct effect of heat‐killed and treated (by Tyndallization) bacteria on the cutaneous and oral immune response without the potential drawbacks of administering live micro‐organisms (e.g. overgrowth or passage of multidrug‐resistant genes). 12 , 13 , 14
This study had several limitations including the small number of dogs enrolled, wide variety of breeds and ages, as well as the short treatment period. A small number of dogs was chosen because the study was designed to collect preliminary data for future studies. The choice was arbitrary, although based on a hypothesized over‐time success rate of 75% of the product against a placebo effect of 30%. Based on these criteria a minimum number of eight dogs would have been necessary to see a significant clinical difference, and consequently 10 dogs were selected. The wide variation in breed and ages could explain the lack of significant changes in the cutaneous microbiota after treatment; furthermore, it is well‐recognised that the cutaneous microbiota can be highly variable between individuals. Thus, a reduction in interindividual variability would have been more beneficial for the microbiota evaluation. Furthermore, only two microbiological samples were taken (D0 and D28) during the study; it is unknown if microbiota changes would have been evident by D42 when a more significant clinical amelioration was seen. In addition, along with the metagenomic approach, it could have been interesting to simultaneously culture the skin microflora to evaluate for changes in the bacteria population.
Given the open‐label nature of the study and the small number of dogs enrolled, a larger double‐blinded, placebo‐controlled, randomized clinical trial is desirable to confirm the clinical benefits showed in this study and to better evaluate the effects of the LinkSkin spray on the cutaneous microbiota.
Supporting information
Figure S1. Median values for total and regional CADESI‐04 scores and pVAS scores.
Figure S2 . Median values for pVAS scores
Figure S3 . OGATE and ease of administration of lactobacilli‐based spray
Figure S4. The distribution of the top 20 operational taxonomic units at genus level in the axillae samples
Figure S5. PCoA at the genus level in the axillae samples based on Bray–Curtis dissimilarity matrices
Figure S6 . The distribution of the top 20 OTUs at the family level in the axillae samples
Figure S7. PCoA at family level in the axillae samples based on Bray–Curtis dissimilarity matrices
Figure S8 . The distribution of the top 20 OTUs at the genus level in the inguinal samples
Figure S9. PCoA at the genus level in the inguinal samples based on Bray–Curtis dissimilarity matrices
Figure S10. The distribution of the top 20 OTUs at the family level in the inguinal samples
Figure S11 . Figure S11 PCoA at the family level in the inguinal samples based on Bray–Curtis dissimilarity matrices
Source of Funding: This study was funded by DRN srl; this included provision of the product and financial support for the laboratory work; they had no influence on the study design and the preparation of this manuscript.
Conflict of Interest: Domenico Santoro has received reimbursements and consultation fees from DRN srl. None of the other authors has declared any conflicts of interest.
References
- 1. Santoro D, Marsella R, Pucheu‐Haston CM, et al. Review: Pathogenesis of canine atopic dermatitis: skin barrier and host‐micro‐organism interaction. Vet Dermatol 2015; 26: 84‐e25. [DOI] [PubMed] [Google Scholar]
- 2. Pucheu‐Haston CM, Santoro D, Bizikova P, et al. Review: Innate immunity, lipid metabolism and nutrition in canine atopic dermatitis. Vet Dermatol 2015; 26: 104‐e28. [DOI] [PubMed] [Google Scholar]
- 3. Pucheu‐Haston CM, Bizikova P, Eisenschenk MN, et al. Review: The role of antibodies, autoantigens and food allergens in canine atopic dermatitis. Vet Dermatol 2015; 26: 115‐e30. [DOI] [PubMed] [Google Scholar]
- 4. Pucheu‐Haston CM, Bizikova P, Marsella R, et al. Review: Lymphocytes, cytokines, chemokines and the T‐helper 1‐T‐helper 2 balance in canine atopic dermatitis. Vet Dermatol 2015; 26: 124‐e32. [DOI] [PubMed] [Google Scholar]
- 5. Saridomichelakis MN, Olivry T. An update on the treatment of canine atopic dermatitis. Vet J 2016; 207: 29–37. [DOI] [PubMed] [Google Scholar]
- 6. Olivry T, DeBoer DJ, Favrot C, et al. Treatment of canine atopic dermatitis: 2015 updated guidelines from the International Committee on Allergic Diseases of Animals (ICADA). BMC Vet Res 2015; 11: 210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Santoro D. Therapies in canine atopic dermatitis: an update. Vet Clin North Am Small Anim Pract 2019; 49: 9–26. [DOI] [PubMed] [Google Scholar]
- 8. Guéniche A, Hennino A, Goujon C, et al. Improvement of atopic dermatitis skin symptoms by Vitreoscilla filiformis bacterial extract. Eur J Dermatol 2006; 16: 380–384. [PubMed] [Google Scholar]
- 9. Gueniche A, Knaudt B, Schuck E, et al. Effects of nonpathogenic gram‐negative bacterium Vitreoscilla filiformis lysate on atopic dermatitis: a prospective, randomized, double‐blind, placebo‐controlled clinical study. Br J Dermatol 2008; 159: 1,357–1,363. [DOI] [PubMed] [Google Scholar]
- 10. Blanchet‐Réthoré S, Bourdes V, Mercenier A, et al. Effect of a lotion containing the heat‐treated probiotic strain Lactobacillus johnsonii NCC 533 on Staphylococcus aureus colonization in atopic dermatitis. Clin Cosmet Investig Dermatol 2017; 10: 249–257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Lee SH, Yoon JM, Kim YH, et al. Therapeutic effect of tyndallized Lactobacillus rhamnosus IDCC 3201 on atopic dermatitis mediated by down‐regulation of Immunoglobulin E in NC/Nga Mice. Microbiol Immunol 2016; 60: 468–476. [DOI] [PubMed] [Google Scholar]
- 12. Adams CA. The probiotic paradox: live and dead cells are biological response modifiers. Nutr Res Rev 2010; 23: 37–46. [DOI] [PubMed] [Google Scholar]
- 13. Taverniti V, Guglielmetti S. The immunomodulatory properties of probiotic microorganisms beyond their viability (ghost probiotics: proposal of paraprobiotic concept). Genes Nutr 2011; 6: 261–274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Piqué N, Berlanga M, Miñana‐Galbis D. Health benefits of heat‐killed (tyndallized) probiotics: an overview. Int J Mol Sci 2019; 20: 2534. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Olivry T, Saridomichelakis M, Nuttall T, et al. Validation of the Canine Atopic Dermatitis Extent and Severity Index (CADESI)‐4, a simplified severity scale for assessing skin lesions of atopic dermatitis in dogs. Vet Dermatol 2014; 25: 77‐e25. [DOI] [PubMed] [Google Scholar]
- 16. Hensel P, Santoro D, Favrot C, et al. Canine atopic dermatitis: detailed guidelines for diagnosis and allergen identification. BMC Vet Res 2015; 11: 196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Hillier A, Lloyd DH, Weese JS, et al. Guidelines for the diagnosis and antimicrobial therapy of canine superficial bacterial folliculitis (antimicrobial guidelines working group of the International Society for Companion Animal Infectious Diseases). Vet Dermatol 2014; 25: 163‐e43. [DOI] [PubMed] [Google Scholar]
- 18. Hill PB, Lau P, Rybnicek J. Development of an owner‐assessed scale to measure the severity of pruritus in dogs. Vet Dermatol 2007; 18: 301–308. [DOI] [PubMed] [Google Scholar]
- 19. Rybníček J, Lau‐Gillard PJ, Harvey R, et al. Further validation of a pruritus severity scale for use in dogs. Vet Dermatol 2009; 20: 115–122. [DOI] [PubMed] [Google Scholar]
- 20. Olivry T, Bensignor E, Favrot C, et al. Development of a core outcome set for therapeutic clinical trials enrolling dogs with atopic dermatitis (COSCAD'18). BMC Vet Res 2018; 14: 238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Dramard V, Kern L, Hofmans J, et al. Effect of l‐theanine tablets in reducing stress‐related emotional signs in cats: an open‐label field study. Ir Vet J 2018; 71: 21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Cobiella D, Archer L, Bohannon M, et al. Pilot study using five methods to evaluate skin barrier function in healthy dogs and in dogs with atopic dermatitis. Vet Dermatol 2019; 30: 121‐e34. [DOI] [PubMed] [Google Scholar]
- 23. Pierezan F, Olivry T, Paps JS, et al. The skin microbiome in allergen‐induced canine atopic dermatitis. Vet Dermatol 2016; 27: 332‐e82. [DOI] [PubMed] [Google Scholar]
- 24. Martin M. Cutadapt removes adapter sequences from high‐throughput sequencing reads. EMBnet.journal 2011; 17: 10–12. [Google Scholar]
- 25. Caporaso JG, Kuczynski J, Stombaugh J, et al. QIIME allows analysis of high‐throughput community sequencing data. Nat Methods 2010; 7: 335–336. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Quast C, Pruesse E, Yilmaz P, et al. The SILVA ribosomal RNA gene database project: improved data processing and web‐based tools. Nucleic Acids Res 2013; 41: D590–D596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Rognes T, Flouri T, Nichols B, et al. VSEARCH: A versatile open source tool for metagenomics. PeerJ 2016; 4: e2584. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Yao JQ, Yu FH. DEB: A web interface for RNA‐seq digital gene expression analysis. Bioinformation 2011; 7: 44–45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Moroi M, Uchi S, Nakamura K, et al. Beneficial effect of a diet containing heat‐killed Lactobacillus paracasei K71 on adult type atopic dermatitis. J Dermatol 2011; 38: 131–139. [DOI] [PubMed] [Google Scholar]
- 30. Yamamoto K, Yokoyama K, Matsukawa T, et al. Efficacy of prolonged ingestion of Lactobacillus acidophilus L‐92 in adult patients with atopic dermatitis. J Dairy Sci 2016; 99: 5,039–5,046. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Figure S1. Median values for total and regional CADESI‐04 scores and pVAS scores.
Figure S2 . Median values for pVAS scores
Figure S3 . OGATE and ease of administration of lactobacilli‐based spray
Figure S4. The distribution of the top 20 operational taxonomic units at genus level in the axillae samples
Figure S5. PCoA at the genus level in the axillae samples based on Bray–Curtis dissimilarity matrices
Figure S6 . The distribution of the top 20 OTUs at the family level in the axillae samples
Figure S7. PCoA at family level in the axillae samples based on Bray–Curtis dissimilarity matrices
Figure S8 . The distribution of the top 20 OTUs at the genus level in the inguinal samples
Figure S9. PCoA at the genus level in the inguinal samples based on Bray–Curtis dissimilarity matrices
Figure S10. The distribution of the top 20 OTUs at the family level in the inguinal samples
Figure S11 . Figure S11 PCoA at the family level in the inguinal samples based on Bray–Curtis dissimilarity matrices


